Abstract
The purpose of this study was to investigate the molecular characteristics of 83 clinical Cryptococcus neoformans/C. gattii species complex isolated in Beijing, China, between 2007 and 2013. Restriction fragment length polymorphism of the gene URA5 (URA5-RFLP), multilocus sequence typing (MLST), and automated repetitive polymerase chain reaction (rep-PCR; DiversiLab system) were performed to genotype these cryptococcal isolates. There was an excellent correlation amongst the three methods; however, PU157 was assigned as VNII according to URA5-RFLP, while it was classified as VNI by the DiversiLab system analysis. PU157 was finally identified as VNB by seven-locus MLST analysis. Moreover, though AD hybrids could not be processed by MLST, ideal results could be obtained by the DiversiLab system. The genotype VNI accounted for 95.2 % (79/83) of isolates. Besides one strain of VNB, VNIII, and VGI each, a strain of VGII was detected in our study, which was isolated from a patient from the temperate region in North China. In addition, the most common MLST sequence type (ST) was ST5, accounting for 91.6 % (76/83), followed by ST31, ST63, ST182, ST295, ST296, and ST332. ST295, ST296, and ST332 were new STs. Except for isolate PU157 (VNB), identical results were obtained quickly and accurately through the DiversiLab system compared to MLST and URA5-RFLP. The discovery of VNB and VGII in the temperate climate regions of China suggested that the population structure of C. neoformans and C. gattii should be explored more extensively. Our results also showed that the DiversiLab system can be used in the genotyping of C. neoformans and C. gattii.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Cryptococcosis is a life-threatening mycosis. It is mainly caused by Cryptococcus neoformans and C. gattii, which can affect not only immunocompromised patients, but also immunocompetent individuals [1]. Candida spp., Aspergillus spp., and Cryptococcus spp. are the leading causes of fungal disease on the global scale [2, 3]. C. neoformans can be found worldwide, mostly in pigeon droppings, while C. gattii was formerly discovered in Eucalyptus camaldulensis and originally considered to be restricted to tropical or subtropical regions [4]. However, as C. gattii was discovered in an outbreak on Vancouver Island in 1999, which eventually spread to British Columbia, Canada, and the US Pacific Northwest, and as the three independent outbreaks of C. gattii were found in North America, it has gained the attention of many countries [1, 5–8].
Generally, there are eight molecular types involved in the C. neoformans/C. gattii species complex [9]. C. neoformans can be divided into C. neoformans var. grubii (serotype A; VNI and VNII), C. neoformans var. neoformans (serotype D; VNIV), and a hybrid between them (serotype AD; VNIII). C. gattii is composed of VGI, VGII, VGIII, and VGIV, which correspond to serotypes B or C [10]. Based on amplified fragment length polymorphism (AFLP) genotyping, C. neoformans var. grubii can also be divided into VNI, VNII, and VNB [9, 11, 12]. The hybrids between C. neoformans and C. gattii (serotype BD) were also reported [13, 14].
Several molecular techniques have been used to study the epidemiology of the C. neoformans/C. gattii species complex, such as karyotype analysis [15], polymerase chain reaction (PCR) fingerprinting by using M13 [10, 16], restriction fragment length polymorphism of the gene URA5 (URA5-RFLP) [10, 17–19], AFLP [20, 21], random amplification of polymorphic DNA (RAPD) [16], multilocus sequence typing (MLST) [9, 11, 22], and multilocus microsatellite typing (MLMT) [23], etc.
Automated repetitive PCR (DiversiLab system) [24] has been used in the strain identification and typing of some species of yeasts and molds [25–30]. The genotyping of C. neoformans and C. gattii using the DiversiLab system has not yet been reported.
The aims of this study were: (1) to reveal the molecular population structure of C. neoformans and C. gattii in China from 2007 to 2013 and (2) to verify the feasibility of the DiversiLab system used in the genotyping of C. neoformans and C. gattii.
Materials and methods
Isolates
A total of 83 clinical Cryptococcus spp. from 76 different patients from Peking Union Medical College Hospital (PUMCH) and Beijing Di-Tan Hospital (BDTH) in Beijing, China between 2007 and 2013 were studied. The clinical information of the 83 isolates can be found in Table 1. The following standard reference isolates of C. neoformans and C. gattii were included: WM148 (= CBS10085 = ATCC MYA-4564, VNI, serotype A), WM626 (= CBS10084 = ATCC MYA-4565, VNII, serotype A), WM628 (= CBS10080 = ATCC MYA4566, VNIII, serotype AD), WM629 (= CBS10079 = ATCC MYA-4567, VNIV, serotype D), WM179 (= CBS10078 = ATCC MYA-4560, VGI, serotype B), WM178 (= CBS10082 = ATCC MYA-4561, VGII, serotype B), WM161 (= CBS10081 = ATCC MYA-4562, VGIII, serotype B), WM779 (= CBS10101 = ATCC MYA-4563, VGIV, serotype C) [16], and H99 (= CBS10515 = CBS8710, VNI, serotype A) [15]. All the isolates were maintained on Sabouraud dextrose agar medium (Becton Dickinson, Sparks, MD, USA) during our investigation.
Phenotype identification
All the isolates were subcultured onto Sabouraud dextrose agar medium at 37 °C for 48–72 h and identified by VITEK 2 Compact (bioMérieux SA, France). Each isolate was also inoculated on canavanine-glycine-bromthymol blue (CGB) agar [31] at 37 °C for at least 1 month.
DNA extraction
Isolates were grown on Sabouraud dextrose agar medium at 37 °C for 48–72 h. Genomic DNA was then extracted using the UltraClean® Microbial DNA Isolation Kit (MO BIO Laboratories, Carlsbad, CA, USA), following the manufacturer’s instructions.
Determination of serotype by PCR
The primer pairs JOHE2596/JOHE3241 and JOHE2596/JOHE3240 [32] were selected to separately amplify the genomic DNA of all the isolates to determine their serotype. JOHE2596/JOHE3241 is specific for serotype A, while JOHE2596/JOHE3240 is specific for serotype D.
URA5-RFLP analysis
The URA5 gene of each isolate was amplified with primers URA5 and SJ01 [10]. PCR reactions and URA5-RFLP analysis were performed according to previously described methods [10].
MLST analysis
MLST analysis consists of seven unlinked loci, including six housekeeping genes, namely, CAP59, GPD1, LAC1, PLB1, SOD1, URA5, and the non-coding region IGS1 [9]. Each locus of every specimen was bidirectionally sequenced. These sequences were uploaded to the MLST Database for the Cryptococcus neoformans/C. gattii species complex (http://mlst.mycologylab.org). An allele number was assigned to each sequence. Seven allele type (AT) numbers and a sequence type (ST) number will be given to each specimen after being compared to the MLST Database website and new AT and ST numbers will be assigned for the new sequences.
The concatenated sequences of the seven loci were aligned with the MEGA v.6.06 software (http://megasoftware.net) [33], along with those of reference cryptococcal isolates reported in previous studies [10, 11, 32, 34, 35]. The phylogenetic tree was constructed using the maximum likelihood method based on the Kimura two-parameter model with 1,000 replications in the bootstrap test.
Each sequence of the seven MLST loci for all tested isolates except isolate PU43 was submitted to GenBank to obtain accession numbers.
rep-PCR DNA fingerprinting
All the samples and reference isolates were performed rep-PCR using the DiversiLab® Fungal Kit (bioMérieux SA, France). The amplified fragments were separated by electrophoresis in a microfluidics DNA LabChip (bioMérieux SA, France) on an Agilent 2100 bioanalyzer (Agilent Technologies, Palo Alto, CA, USA). Further data analysis was performed with the web-based DiversiLab software v.3.4 (http://pumc.diversilab.com). The band-based modified Kullback–Leibler distance method was used for calculating the percent similarities.
Results
Phenotypic identification
All 83 isolates were identified as C. neoformans by VITEK 2 Compact. CGB medium inoculated with PU8 turned completely blue after being incubated at 37 °C for 5 days, with the same result for PU99. CGB medium inoculated with the remaining 81 isolates remained yellow a month later. With CGB medium, PU8 and PU99 were identified as C. gattii, while the other 81 isolates were C. neoformans.
Determination of serotype
PU43 could be amplified not only by JOHE2596/JOHE3241 but also by JOHE2596/JOHE3240, and was confirmed to be serotype AD. However, neither of the two primer pairs could amplify the DNA of PU8 and PU99, which were deduced to be serotypes B or C. The remaining 80 isolates (96.4 %), including PU157, could be amplified by JOHE2596/JOHE3241 but not JOHE2596/JOHE3240, and proved to be serotype A.
URA5-RFLP analysis
The URA5-RFLP profiles of nine isolates (PU1, PU28, PU35, PU41, PU48, PU55, PU146, PU161, and PU162) were identical to those of H99 and WM148, which belong to VNI, while PU157, PU43, PU99, and PU8 were assigned as VNII, VNIII, VGI, and VGII, respectively (Fig. 1). The URA5-RFLP patterns of the remaining 70 clinical isolates in this study were the same as those of H99 and WM148. Altogether, 79 out of 83 isolates (95.2 %) were identified as VNI.
MLST analysis
A total of seven different MLST STs were recognized, three (ST295, ST296, and ST332) of which were novel and four (ST5, ST31, ST63, and ST182) of which had been reported previously. The most common ST was ST5 (VNI), which accounted for 91.6 % (76/83). ST31, ST63, and ST296 belonged to VNI, while ST295, ST332, and ST182 represented VNB (PU157), VGI (PU99), and VGII (PU8), respectively (Table 1). All the sequences submitted to GenBank acquired their accession numbers between KF864215 and KF890168.
Some loci of isolate PU43 had double peaks in their peak sequence diagram because of the AD hybrid, and PU43 was not analyzed in MLST. In total, the sequence length of the seven concatenated MLST loci was 4,272 base pairs. The phylogenetic tree constructed by the maximum likelihood method showed that most isolates (PU1, PU35, PU48, and PU161) clustered with K1 [34], which correlated with the M5 genotype identified by 12 MLST loci typing [11]. Only one isolate (PU162) clustered closely with WM148 and H99, which belonged to the M1 genotype [11]. Only one isolate (PU146) clustered with K54 [34], the M4 genotype [11]. PU157 was even closer to the VNB cluster than the VNI cluster, which was, therefore, identified as VNB. PU8 was classified as VGII and PU99 was determined as VGI (Fig. 2). The remaining 73 tested samples not shown in Fig. 2 had identical sequences of the seven MLST loci with PU1 et al., and were assigned as VNI (Table 1).
Phylogenetic tree constructed using the maximum likelihood method with 1,000 replications in the bootstrap test based on the concatenated seven MLST loci (CAP59, GPD1, IGS1, LAC1, PLB1, SOD1, and URA5) of part of the tested isolates identified as PU and other global cryptococcal isolates from previous studies. Only the bootstrap values >50 % are shown
DiversiLab typing
An 85 % similarity threshold was chosen to determine the genotype of tested isolates. Isolates among the VNI group showed polymorphism. The nine reference isolates and all 83 cryptococcal isolates in our study were successfully genotyped with the DiversiLab system. The dendrogram showed that most of the isolates (n = 79; 95.2 %) clustered together with WM148 (VNI), and were classified as VNI. Samples with strain similarity (≥99 %) were condensed in the dendrogram due to space constraints. PU43 and MW628, the AD hybrids, clustered together in the dendrogram. PU8 clustered with WM178 (VGII), while PU99 clustered with WM179 (VGI). PU43, PU99, and PU8 were determined as VNIII, VGI, and VGII, respectively (Fig. 3). PU157 was identified as VNI by the DiversiLab system analysis.
rep-PCR-based dendrogram and virtual gel image of the DiversiLab system for the 83 clinical and nine standard reference isolates of C. neoformans and C. gattii. a Number of isolates for which the similarity was no less than 99 % compared with the isolates adjacent to them. b Varieties. c Genotype concluded by the DiversiLab system. d Genotype determined by seven-loci MLST (CAP59, GPD1, IGS1, LAC1, PLB1, SOD1, and URA5) analysis. e Genotype assigned by URA5-RFLP. f ST generated by the seven MLST loci. g –, no varieties. h NA, not applicable
Discussion
Until now, the reports concerning the molecular epidemiology of the Cryptococcus species complex in China were insufficient and less comprehensive. Chen et al. [17] analyzed 129 clinical Cryptococcus spp. isolates from China using URA5-RFLP and PCR fingerprinting M13 analyses, and showed that 93.0 % (120/129) belonged to C. neoformans VNI, while 7.0 % (9/129) were C. gattii VGI. With almost the same methods, Feng et al. [18] studied 115 clinical cryptococcal isolates in China and revealed that 89.6 % (103/115) represented VNI, along with two isolates of VNIII, one VNIV, eight VGI, and one VGII.
Our data also showed that VNI was the most common genotype. Meanwhile, we revealed the MLST STs of the 82 isolates of C. neoformans and C. gattii in our study. A total of 76 VNI (91.6 %) were identified as MLST ST5, followed by one each of ST31, ST63, and ST296 for VNI, one ST295 for VNB, one ST332 for VGI, and one ST182 for VGII. Some studies have shown that ST5 was the most frequent ST in China and other Asian countries [19, 36]. A total of 476 isolates of C. neoformans var. grubii from eight Asian countries, including 86 Chinese clinical Cryptococcus isolates from the Second Military Medical University, were analyzed by MLST analysis. Seventy-four of 86 Chinese cryptococcal isolates were identified as ST5, followed by five ST53, two ST194, and one each of ST31, ST93, ST186, ST191, and ST195 [36]. However, the three dominating MLST STs in Thailand were ST44, ST45, and ST46 [37].
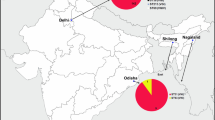
C. gattii has not yet been reported in temperate regions in China. A strain of C. gattii VGII (PU8) from the temperate climate region located on longitude 115° ~ 116° and latitude 37° ~ 38° in China was detected in our study. Few VGII isolates had been reported in China previously, all of which were distributed in the subtropical climate zones of South China [18, 38]. Our VGII (ST = 182) strain was different from XH91 reported as VGII in China in 2008 [18]. It was also different from the prevalent outbreak strain VGIIb (ST = 7) on Vancouver Island from 1999 to 2003 [5], though they clustered closely in Fig 2.
C. neoformans VNB has not yet been reported in Asian countries till now. It had been found in Botswana [11], Italy [12], Portugal, Rwanda, Brazil, and Venezuela [32] etc. A strain of C. neoformans VNB (PU157) was discovered in our study. PU157 was finally determined as C. neoformans VNB by seven-loci MLST analysis. It belonged to MLST ST295, a novel ST. According to the URA5-RFLP results, the VNII-1 isolates of C. neoformans were identified as VNII. The phylogenetic tree based on the combined sequences of GPD1, IGS1, PLB1, and URA5 revealed that those VNII-1 isolates were VNB [39]. Similar results occurred in our study; PU157 was first assigned as VNII according to its URA5-RFLP pattern (Fig. 1), and it seemed to belong to VNI by the DiversiLab system typing (Fig. 3). In the phylogenic tree constructed on the base of the concatenated seven MLST loci, PU157 clustered relatively closer to VNB (Fig. 2).
There was a good correlation between the DiversiLab system and MLST as well as URA5-RFLP in genotyping the Cryptococcus species complex. Additionally, the total time consumed in the performance of the DiversiLab system for 12 samples lasted less than 4 h. Of particular note is that the AD hybrid can be recognized by the DiversiLab system, which cannot be handled by MLST. Had a reference strain of VNB been included in our study, PU157 may have had an accurate result from the DiversiLab system.
However, the downfall of the DiversiLab system in the genotyping of Cryptococcus spp. is that, as there is no database for Cryptococcus spp. at present, the reference standard isolates of Cryptococcus spp. need to be included simultaneously for the first performance. And the limitation of our study was that there were not enough cryptococcal isolates with diversity in our collection, which suggested us to further investigate the population structure of C. neoformans and C. gattii in China and validate the ability of the DiversiLab system in genotyping cryptococcal isolates.
In summary, our results showed that the discovery of C. neoformans VNB and C. gattii VGII in the temperate climate regions in China warrant further investigation concerning the molecular epidemiology of C. neoformans and C. gattii. Moreover, our results demonstrated that the DiversiLab system could be used in the genotyping of C. neoformans and C. gattii due to its convenience and effectiveness. However, the resolution and reproducibility of the system needs to be further verified with more cryptococcal isolates of different genotypes.
References
La Hoz RM, Pappas PG (2013) Cryptococcal infections: changing epidemiology and implications for therapy. Drugs 73:495–504. doi:10.1007/s40265-013-0037-z
Liao Y, Chen M, Hartmann T, Yang RY, Liao WQ (2013) Epidemiology of opportunistic invasive fungal infections in China: review of literature. Chin Med J (Engl) 126:361–368
Pfaller MA, Diekema DJ (2010) Epidemiology of invasive mycoses in North America. Crit Rev Microbiol 36:1–53. doi:10.3109/10408410903241444
Ellis DH, Pfeiffer TJ (1990) Ecology, life cycle, and infectious propagule of Cryptococcus neoformans. Lancet 336:923–925
Kidd SE, Hagen F, Tscharke RL, Huynh M, Bartlett KH, Fyfe M, Macdougall L, Boekhout T, Kwon-Chung KJ, Meyer W (2004) A rare genotype of Cryptococcus gattii caused the cryptococcosis outbreak on Vancouver Island (British Columbia, Canada). Proc Natl Acad Sci U S A 101:17258–17263. doi:10.1073/pnas.0402981101
Chaturvedi V, Chaturvedi S (2011) Cryptococcus gattii: a resurgent fungal pathogen. Trends Microbiol 19:564–571. doi:10.1016/j.tim.2011.07.010
Byrnes EJ 3rd, Bildfell RJ, Frank SA, Mitchell TG, Marr KA, Heitman J (2009) Molecular evidence that the range of the Vancouver Island outbreak of Cryptococcus gattii infection has expanded into the Pacific Northwest in the United States. J Infect Dis 199:1081–1086. doi:10.1086/597306
Hagen F, Ceresini PC, Polacheck I, Ma H, van Nieuwerburgh F, Gabaldón T, Kagan S, Pursall ER, Hoogveld HL, van Iersel LJ, Klau GW, Kelk SM, Stougie L, Bartlett KH, Voelz K, Pryszcz LP, Castañeda E, Lazera M, Meyer W, Deforce D, Meis JF, May RC, Klaassen CH, Boekhout T (2013) Ancient dispersal of the human fungal pathogen Cryptococcus gattii from the Amazon rainforest. PLoS One 8:e71148. doi:10.1371/journal.pone.0071148
Meyer W, Aanensen DM, Boekhout T, Cogliati M, Diaz MR, Esposto MC, Fisher M, Gilgado F, Hagen F, Kaocharoen S, Litvintseva AP, Mitchell TG, Simwami SP, Trilles L, Viviani MA, Kwon-Chung J (2009) Consensus multi-locus sequence typing scheme for Cryptococcus neoformans and Cryptococcus gattii. Med Mycol 47:561–570. doi:10.1080/13693780902953886
Meyer W, Castañeda A, Jackson S, Huynh M, Castañeda E; IberoAmerican Cryptococcal Study Group (2003) Molecular typing of IberoAmerican Cryptococcus neoformans isolates. Emerg Infect Dis 9:189–195. doi:10.3201/eid0902.020246
Litvintseva AP, Thakur R, Vilgalys R, Mitchell TG (2006) Multilocus sequence typing reveals three genetic subpopulations of Cryptococcus neoformans var. grubii (serotype A), including a unique population in Botswana. Genetics 172:2223–2238. doi:10.1534/genetics.105.046672
Cogliati M, Zamfirova RR, Tortorano AM, Viviani MA; Fimua Cryptococcosis Network (2013) Molecular epidemiology of Italian clinical Cryptococcus neoformans var. grubii isolates. Med Mycol 51:499–506. doi:10.3109/13693786.2012.751642
Bovers M, Hagen F, Kuramae EE, Diaz MR, Spanjaard L, Dromer F, Hoogveld HL, Boekhout T (2006) Unique hybrids between the fungal pathogens Cryptococcus neoformans and Cryptococcus gattii. FEMS Yeast Res 6:599–607. doi:10.1111/j.1567-1364.2006.00082.x
Bovers M, Hagen F, Kuramae EE, Hoogveld HL, Dromer F, St-Germain G, Boekhout T (2008) AIDS patient death caused by novel Cryptococcus neoformans × C. gattii hybrid. Emerg Infect Dis 14:1105–1108. doi:10.3201/eid1407.080122
Perfect JR, Ketabchi N, Cox GM, Ingram CW, Beiser CL (1993) Karyotyping of Cryptococcus neoformans as an epidemiological tool. J Clin Microbiol 31:3305–3309
Meyer W, Marszewska K, Amirmostofian M, Igreja RP, Hardtke C, Methling K, Viviani MA, Chindamporn A, Sukroongreung S, John MA, Ellis DH, Sorrell TC (1999) Molecular typing of global isolates of Cryptococcus neoformans var. neoformans by polymerase chain reaction fingerprinting and randomly amplified polymorphic DNA—a pilot study to standardize techniques on which to base a detailed epidemiological survey. Electrophoresis 20:1790–1799. doi:10.1002/(SICI)1522-2683(19990101)20:8<1790::AID-ELPS1790>3.0.CO;2-2
Chen J, Varma A, Diaz MR, Litvintseva AP, Wollenberg KK, Kwon-Chung KJ (2008) Cryptococcus neoformans strains and infection in apparently immunocompetent patients, China. Emerg Infect Dis 14:755–762. doi:10.3201/eid1405.071312
Feng X, Yao Z, Ren D, Liao W, Wu J (2008) Genotype and mating type analysis of Cryptococcus neoformans and Cryptococcus gattii isolates from China that mainly originated from non-HIV-infected patients. FEMS Yeast Res 8:930–938. doi:10.1111/j.1567-1364.2008.00422.x
Mihara T, Izumikawa K, Kakeya H, Ngamskulrungroj P, Umeyama T, Takazono T, Tashiro M, Nakamura S, Imamura Y, Miyazaki T, Ohno H, Yamamoto Y, Yanagihara K, Miyzaki Y, Kohno S (2013) Multilocus sequence typing of Cryptococcus neoformans in non-HIV associated cryptococcosis in Nagasaki, Japan. Med Mycol 51:252–260. doi:10.3109/13693786.2012.708883
Boekhout T, Theelen B, Diaz M, Fell JW, Hop WC, Abeln EC, Dromer F, Meyer W (2001) Hybrid genotypes in the pathogenic yeast Cryptococcus neoformans. Microbiology 147:891–907
Hagen F, Chowdhary A, Prakash A, Yntema JB, Meis JF (2014) Molecular characterization of Cryptococcus gattii genotype AFLP6/VGII isolated from woody debris of divi-divi (Caesalpinia coriaria), Bonaire, Dutch Caribbean. Rev Iberoam Micol 31:193–196. doi:10.1016/j.riam.2013.10.007
Fraser JA, Giles SS, Wenink EC, Geunes-Boyer SG, Wright JR, Diezmann S, Allen A, Stajich JE, Dietrich FS, Perfect JR, Heitman J (2005) Same-sex mating and the origin of the Vancouver Island Cryptococcus gattii outbreak. Nature 437:1360–1364. doi:10.1038/nature04220
Pan W, Khayhan K, Hagen F, Wahyuningsih R, Chakrabarti A, Chowdhary A, Ikeda R, Taj-Aldeen SJ, Khan Z, Imran D, Sjam R, Sriburee P, Liao W, Chaicumpar K, Ingviya N, Mouton JW, Curfs-Breuker I, Boekhout T, Meis JF, Klaassen CH (2012) Resistance of Asian Cryptococcus neoformans serotype A is confined to few microsatellite genotypes. PLoS One 7:e32868. doi:10.1371/journal.pone.0032868
Healy M, Huong J, Bittner T, Lising M, Frye S, Raza S, Schrock R, Manry J, Renwick A, Nieto R, Woods C, Versalovic J, Lupski JR (2005) Microbial DNA typing by automated repetitive-sequence-based PCR. J Clin Microbiol 43:199–207. doi:10.1128/jcm.43.1.199-207.2005
Wise MG, Healy M, Reece K, Smith R, Walton D, Dutch W, Renwick A, Huong J, Young S, Tarrand J, Kontoyiannis DP (2007) Species identification and strain differentiation of clinical Candida isolates using the DiversiLab system of automated repetitive sequence-based PCR. J Med Microbiol 56:778–787. doi:10.1099/jmm.0.47106-0
Treviño M, García-Riestra C, Areses P, García X, Navarro D, Suárez FJ, López-Dequidt IA, Zaragoza O, Cuenca-Estrella M (2014) Emerging Trichosporon asahii in elderly patients: epidemiological and molecular analysis by the DiversiLab system. Eur J Clin Microbiol Infect Dis 33:1497–1503. doi:10.1007/s10096-014-2099-6
Healy M, Reece K, Walton D, Huong J, Shah K, Kontoyiannis DP (2004) Identification to the species level and differentiation between strains of Aspergillus clinical isolates by automated repetitive-sequence-based PCR. J Clin Microbiol 42:4016–4024. doi:10.1128/jcm.42.9.4016-4024.2004
Healy M, Reece K, Walton D, Huong J, Frye S, Raad II, Kontoyiannis DP (2005) Use of the Diversi Lab System for species and strain differentiation of Fusarium species isolates. J Clin Microbiol 43:5278–5280. doi:10.1128/jcm.43.10.5278-5280.2005
Pounder JI, Williams S, Hansen D, Healy M, Reece K, Woods GL (2005) Repetitive-sequence-PCR-based DNA fingerprinting using the DiversiLab system for identification of commonly encountered dermatophytes. J Clin Microbiol 43:2141–2147. doi:10.1128/jcm.43.5.2141-2147.2005
Pounder JI, Hansen D, Woods GL (2006) Identification of Histoplasma capsulatum, Blastomyces dermatitidis, and Coccidioides species by repetitive-sequence-based PCR. J Clin Microbiol 44:2977–2982. doi:10.1128/jcm.00687-06
Kwon-Chung KJ, Polacheck I, Bennett JE (1982) Improved diagnostic medium for separation of Cryptococcus neoformans var. neoformans (serotypes A and D) and Cryptococcus neoformans var. gattii (serotypes B and C). J Clin Microbiol 15:535–537
Bovers M, Hagen F, Kuramae EE, Boekhout T (2008) Six monophyletic lineages identified within Cryptococcus neoformans and Cryptococcus gattii by multi-locus sequence typing. Fungal Genet Biol 45:400–421. doi:10.1016/j.fgb.2007.12.004
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: Molecular Evolutionary Genetics Analysis version 6.0. Mol Biol Evol 30:2725–2729. doi:10.1093/molbev/mst197
Choi YH, Ngamskulrungroj P, Varma A, Sionov E, Hwang SM, Carriconde F, Meyer W, Litvintseva AP, Lee WG, Shin JH, Kim EC, Lee KW, Choi TY, Lee YS, Kwon-Chung KJ (2010) Prevalence of the VNIc genotype of Cryptococcus neoformans in non-HIV-associated cryptococcosis in the Republic of Korea. FEMS Yeast Res 10:769–778. doi:10.1111/j.1567-1364.2010.00648.x
Carriconde F, Gilgado F, Arthur I, Ellis D, Malik R, van de Wiele N, Robert V, Currie BJ, Meyer W (2011) Clonality and alpha-a recombination in the Australian Cryptococcus gattii VGII population—an emerging outbreak in Australia. PLoS One 6:e16936. doi:10.1371/journal.pone.0016936
Khayhan K, Hagen F, Pan W, Simwami S, Fisher MC, Wahyuningsih R, Chakrabarti A, Chowdhary A, Ikeda R, Taj-Aldeen SJ, Khan Z, Ip M, Imran D, Sjam R, Sriburee P, Liao W, Chaicumpar K, Vuddhakul V, Meyer W, Trilles L, van Iersel LJ, Meis JF, Klaassen CH, Boekhout T (2013) Geographically structured populations of Cryptococcus neoformans variety grubii in Asia correlate with HIV status and show a clonal population structure. PLoS One 8:e72222. doi:10.1371/journal.pone.0072222
Simwami SP, Khayhan K, Henk DA, Aanensen DM, Boekhout T, Hagen F, Brouwer AE, Harrison TS, Donnelly CA, Fisher MC (2011) Low diversity Cryptococcus neoformans variety grubii multilocus sequence types from Thailand are consistent with an ancestral African origin. PLoS Pathog 7:e1001343. doi:10.1371/journal.ppat.1001343
Tseng HK, Liu CP, Ho MW, Lu PL, Lo HJ, Lin YH, Cho WL, Chen YC; Taiwan Infectious Diseases Study Network for Cryptococcosis (2013) Microbiological, epidemiological, and clinical characteristics and outcomes of patients with cryptococcosis in Taiwan, 1997–2010. PLoS One 8:e61921. doi:10.1371/journal.pone.0061921
Ngamskulrungroj P, Gilgado F, Faganello J, Litvintseva AP, Leal AL, Tsui KM, Mitchell TG, Vainstein MH, Meyer W (2009) Genetic diversity of the Cryptococcus species complex suggests that Cryptococcus gattii deserves to have varieties. PLoS One 4:e5862. doi:10.1371/journal.pone.0005862
Acknowledgments
We would like to thank Prof. Dr. Wieland Meyer (Molecular Mycology Research Laboratory, Westmead Hospital, Australia) for the reference isolates in our study originating from their laboratory. We also thank Dr. Luciana Trilles (Molecular Mycology Laboratory, Westmead Hospital, Australia) for providing the MLST STs of the VNB isolates. This work was supported by the National Key Technologies R&D Program for the 12th Five-Year Plan (grant number 2012ZX10001003).
Conflict of interest
The authors declare that they have no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Dou, HT., Xu, YC., Wang, HZ. et al. Molecular epidemiology of Cryptococcus neoformans and Cryptococcus gattii in China between 2007 and 2013 using multilocus sequence typing and the DiversiLab system. Eur J Clin Microbiol Infect Dis 34, 753–762 (2015). https://doi.org/10.1007/s10096-014-2289-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10096-014-2289-2