Abstract
Amino acid sequence alignments of the transcriptional regulator AfeR, which is involved in type 1 quorum sensing (QS) in Acidithiobacillus ferrooxidans bacteria, with other acyl homoserine lactone (AHL)-dependent QS regulators, revealed the presence of strictly or highly conserved residues located in the active site of these proteins. As a consequence, a model of AfeR was constructed to study the binding mode of long-chain AHLs using molecular dynamics and subsequent rigid ligand docking. This study, performed on the tetradecanoyl homoserine lactone C14-AHL, showed that the binding mode involved a curved conformation. Based on these results, the binding mode of tetradec-7-Z enoyl homoserine lactone, an AHL that is conformationally constrained due to the presence of the cis double bond, was investigated. This mono-unsaturated AHL with its preferential curved shape conformation was found to be particularly well adapted to the active site of AfeR. These results should be helpful in the rational design of QS modulators with potential biotechnological applications and especially in the improvement of industrial bioleaching from ores.

Left Overall folding of the AfeR model with secondary structure; right best docking results obtained for tetradecanoyl homoserine lactone (C14-AHL)
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Quorum sensing (QS) is a communication system that allows bacteria to adapt their behaviour to their cell density [1–5]. This system regulates the expression of genes encoding important phenotypes such as biofilm formation, virulence and bioluminescence. QS is based on signalling molecules called auto-inducers, including cyclic peptides in Gram-positive bacteria [6], acyl homoserine lactone (AHLs) in Gram-negative bacteria, and 4,5-dihydroxy-2,3-pentanedione (DPD)-derived compounds in both Gram-positive and -negative bacteria [7]. QS has recently aroused great interest as a novel target for interference with biological functions of bacteria, with potential medical or biotechnological applications [8–12]. The design and synthesis of QS modulators has been demonstrated to be an effective target to interact with the development of bacteria [13–15].
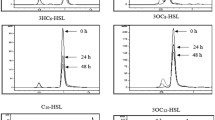
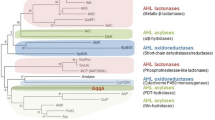
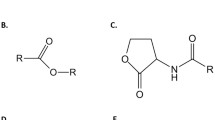
Acidithiobacillus ferrooxidans is an acidophilic Gram-negative bacterium able to oxidise ferrous ion as an energy source. This biological characteristic is used directly in biomining operations for the extraction of copper and other metals from ores (bioleaching) [16, 17]. Since the first observation of QS in Vibrio fischeri, a bioluminiscent bacterium [18], a QS system type AI-1 complex involving an AHL synthase (LuxI protein family) and an AHL-dependent transcriptional regulator (LuxR proteins family) has been characterised in several Gram-negative bacteria. With the discovery of a functional QS type AI-1 system in A. ferrooxidans [19, 20], QS has become a potential target to improve bioleaching [21]. This functional QS type AI-1 system is based on AfeR, a transcriptional regulator protein that binds the corresponding AHLs, and on AfeI, which synthesises long-chain AHLs that act as auto-inducers. These include 3-hydroxy-octanoyl homoserine lactone (3-hydroxy-C8-AHL), 3-hydroxy-decanoyl homoserine lactone (3-hydroxy-C10-AHL), dodecanoyl homoserine lactone (C12-AHL), 3-oxo-dodecanoyl homoserine lactone (3-oxo-C12-AHL), 3-hydroxy- dodecanoyl homoserine lactone (3-hydroxy-C12-AHL), tetradecanoyl homoserine lactone (C14-AHL), 3-oxo- tetradecanoyl homoserine lactone (3-oxo-C14-AHL), 3-hydroxy- tetradecanoyl homoserine lactone (3-hydroxy-C14-AHL), and 3-hydroxy-hexadecanoyl (3-hydroxy-C16-AHL) (Fig. 1) [19].
We recently reported a molecular modelling study of four proteins belonging to the LuxR protein family (TraR, SdiA, LuxR and LasR), and the binding mode of QS modulators [22, 23]. Here, in order to define new methods for bioleaching improvement, we extended this work to include a 3D-structural study of the AfeR protein from A. ferrooxidans. A study of the binding mode of long-chain AHLs via conformational analysis using molecular dynamics (MD) and subsequent rigid ligand docking (RLD) is presented.
Materials and methods
All calculations were performed on a Dell OPTIPLEX GW 620 PC equipped with a double processor, and using Sybyl 7.2 package for Linux [24], ArgusLab [25] and YASARA [26] software. Multiple amino acid sequence alignments were generated with T-COFFEE [27] with the protein sequences of AfeR (NCBI protein code AAV53702), TraR Agrobacterium tumefaciens (NCBI protein code 1L3LA), SdiA from Escherichia coli (NCBI protein code 2AVXA), LuxR from Vibrio fischeri (NCBI protein code CAA68561) and LasR from Pseudomonas aeruginosa (NCBI protein code AAA25874). The protein model of AfeR was created using SWISS-MODEL [28] with Clustal W [29]. The natural ligand of TraR, 3-oxo-octanoyl homoserine lactone, was manually docked in the active site of AfeR (see Fig. 4) and residues within a distance of 5 Å from the ligand were selected to generate the docking box. Conformation analyses were carried out using MD at 400 K and 600 K with the TRIPOS force field and the following parameters: step: 1 fs, length 10,000 steps. AHLs were drawn with Sybyl and minimised with the TRIPOS force field. The resulting conformers were used as initial conformers for MD simulations. Preferential conformers of AHLs, generated as a result of MD, were minimised, classified and selected according to their relative conformation-shape to obtain the representative preferential conformers shown in Fig. 5. These conformers were then docked as rigid ligands in the docking box. Docking results evaluated with the empirical binding free energy function [30] were analysed for the conformers in which the ligand was best positioned to describe the interactions between the ligand and the active site of AfeR; hydrogen bonds were assigned within a distance of 3 Å. Docking experiments were performed with the docking module of ArgusLab with the following parameters: Docking box: X = Y = Z = 15 Å, ligand option: rigid; calculation type: Dock; Docking engine: GADock (Lamarckian Genetic Algorithm) [30]; default advanced parameters.
Results and discussion
Molecular insights into AfeR
Multiple amino acid sequence alignment of the transcriptional regulator AfeR with sequences of the other AHL-dependent transcriptional regulators TraR, SdiA, LasR and LuxR showed that strictly conserved residues located in the active sites of the proteins TraR, LuxR and LasR are also conserved in AfeR (Tyr58, Trp63, Asp75, Trp90, Ala105, Gly113). This observation strongly suggests that the shape of the active site of AfeR should be similar to that of TraR [31] (Fig. 2). Indeed, models of LuxR and LasR proteins based on TraR have already been reported for the docking of QS AHL-derived compounds [32, 33] and unrelated-AHL QS modulators [34]. In these cases, a LuxR or a LasR model derived from the TraR structure was used based on amino acid sequence alignment of the LuxR-protein family. Such molecular modelling studies have been subsequently corroborated by the NMR-resolved structure of SdiA [35] and the X-ray structure of LasR [36], both of which show a similar folding to TraR and SdiA.
Multiple amino acid sequence alignment of AfeR homologues (TraR from Agrobacterium tumefaciens, SdiA from Escherichia coli, LasR from Pseudomonas aeruginosa and LuxR from Vibrio fischeri). Strictly conserved residues located in the active site are boxed in red and conserved residues are boxed in blue. The multiple sequence alignment was generated using T-COFFEE [27]
Based on the amino acid sequence alignment, we then constructed a model of AfeR with Clustal W, using SWISS-MODEL (Fig. 3). The monomer model embodies a receiver domain with the active site (upper part) and a regulatory domain (lower part) that interacts with DNA to regulate gene transcription. By analogy with TraR, the active site with the entrance pocket for natural ligands is located between three α-helices and several β-sheets, as illustrated in Fig. 3. The active site is located on the opposite side of this region, and an α-helix is involved in the dimerisation process. In the LuxR protein family, this process is due to binding of the auto-inducer and leads to the formation of an AHL-transcriptional regulator-DNA complex.
In order to validate the AfeR model derived from TraR, the natural ligand of TraR, 3-oxo-octanoyl homoserine lactone (3-oxo-C8-AHL), was docked within the putative active site of AfeR deduced from the entire protein with strictly conserved residues located in the active site of other proteins in the LuxR family (Fig. 2). As expected, the docking result showed a very similar binding mode of 3-oxo-C8-AHL within the active site of AfeR compared with that in the active site of TraR [31] (Fig. 4b). Following docking of 3-oxo-C8-AHL, amino acids in the active site of AfeR were determined by selecting amino acids within a distance of 5 Å from the ligand (Fig. 4). The active site of the AfeR model comprises two hydrophobic pockets, one interacting with the alkyl chain of AI-1, and a smaller one interacting with the lactone ring. As with other proteins in the LuxR family, two conserved tyrosine residues (Tyr58 and Tyr66 represented in red in Fig. 4) are located at the entrance of the large hydrophobic pocket constituted by hydrophobic residues such as Leu54, Pro170, Ala38. Residue Asp75, located at the junction between the two pockets, is equivalent to the aspartic acid residues in the LuxR family that play an important role by forming a hydrogen bond with the NH group of the natural ligand. Tryptophan residues Trp90 and Trp62 are also important conserved residues in the LuxR family, the first being involved in a bond with the carbonyl group of the amide function, and the second being located in the hydrophobic pocket interacting with the lactone ring.
Binding mode of long-chain AHLs
To examine the binding mode of a long-chain AHL, C14-AHL, one of the more abundant AHLs produced by A. ferrooxidans [19] and one of the most structurally simple long-chain AHLs, i.e. without 3-oxo or 3-hydroxy functions, was chosen to perform a sequential MD / RLD analysis. This will allow us first to describe the conformation analysis of C14-AHL leading to stable preferential conformers and second, to study the fitting of these conformers into the active site of AfeR. This method was expected to yield information on the structure/conformation-activity relationship of this molecule, and should help in the design of new modulators to interfere with the biological activities of A. ferrooxidans that will eventually lead to enhanced bioleaching.
Molecular dynamics gave access to stable conformers of C14-AHL. All conformers obtained as a result of the calculation could be classified according to the shape adopted by the alkyl chain into representative conformer groups. One group, represented by conformers C14-AHL 2-4 and 6 (Fig. 5a) increasing from conformers C 14 -AHL 2 to 4, were characterised by alkyl chain withdrawal; a second group, e.g. C 14 -AHL 8, 16 (Fig. 5b), had either a more linear alkyl chain. A third type, e.g. the conformer C 14 -AHL 11, had several alkyl chain withdrawals. The conformer C 14 -AHL 15 was found to be intermediate between the linear and the withdrawal shape.
Rigid ligand docking
The ensemble of representative conformers of C14-AHL was then submitted to RLD experiments within the active site of AfeR. Conformers from group A with an alkyl chain withdrawal were found to fit particularly well into the active site, giving a ΔGbind < − 9 kcal mol−1 whereas other representative conformers from group B (not shown) led to ΔGbind > − 9 kcal mol−1. This was especially the case for the linear conformer C14-AHL 16 and the conformer C14-AHL 11, which gave no results. On the other hand, C14-AHL 8 showed a moderate fit (Fig. 6). For C14-AHL 15, a conformer between the linear and the withdrawal or curved shape conformation, the docking experiment led to a ΔGbind of about −8 kcal mol−1.
a Best docking results obtained for C14-AHL. b Superimposition of the docked conformers with the conformation adopted by 3-oxo-C12-AHL (represented in blue) co-crystallised in the active site of LasR [36]. c Schematic overview of the hydrogen bond network between C14-AHL and residues of the active site
This docking study revealed that preferential conformers of C14-AHL with alkyl chain withdrawal, which induces a curved shape conformation, are particularly adapted to the active site of AfeR with a ΔGbind of about – 10 kcal mol−1 in the best case (Fig. 6a). These results are in good agreement with the conformation of 3-oxo-C12-AHL co-crystallised in the active site of the receiver domain of the protein LasR [36]. The superimposition of the docked conformers with the conformation adopted by the 3-oxo-C12-AHL indeed shows a similar orientation of the alkyl chains (Fig. 6b). Strictly conserved residues in the LuxR protein family are involved in the hydrogen network (Fig. 6c).
Implications for the rational design of potential QS modulators for AfeR
According to the conformational analysis of C14-AHL performed using MD and the subsequent docking experiments, conformers with an alkyl chain withdrawal, i.e. with a curved shape conformation are particularly adapted to the active site of the AfeR model. Based on these results, the rational design of QS modulators aimed at enhancing QS activities should be focussed on conformationally restrained AHL analogues with a curved shape-type conformation. A. ferrooxidans is not the only bacterium able to produce long-chain AHLs. In the literature, long-chain AHLs are indeed used as AIs by other bacteria such as Yersinia enterocolitica [37], Staphylococcus aureus [6], Sinorhizobium meliloti [38], and P. aeruginosa [36].
Recently, novel long-chain AHLs have been identified, or have been found to be efficient, in marine alphaproteobacteria [39]. Interestingly, the structure of some AHLs contains one double bond with a (Z) geometry, as in the case of the 3-oxo-dodec-5-Z-enoyl homoserine lactone (5Z-3-oxo-C12-AHL) and dodec-5-Z-enoyl homoserine lactone (5-Z-C12-AHL) in Mesorhizobium sp. [40] and tetradec-7-Z-enoyl homoserine lactone (7-Z-C14-AHL) in Rhodobacter sphaeroides [39] (Fig. 7a).
a Structure of mono-unsaturated AHLs (7-Z-C14-AHL and 5-Z-C12-AHL). b Docking results obtained for 7-Z-C14-AHL (residues at the active site of AfeR are represented with Van der Waals surface). c Best docking result obtained for 7-Z-C14-AHL. d Schematic overview of the hydrogen bond network between 7-Z-C14-AHL and residues of the active site
Because long-chain AHLs with double bonds have been identified in the literature as AIs, and since double bonds are indeed known to induce a curved shape conformation of alkyl chains [41, 42], we investigated the conformation of 7-Z-C14-AHL in order to subsequently perform docking experiments. As expected, the conformation analysis showed that the presence of the double bond induced a curved shape conformation. Three of the conformers generated led to ligand positions with the best ΔGbind values of less than −12 kcal mol−1 (Fig. 7b). Figure 7c shows the best ligand pose for 7-Z-C14-AHL, which fits particularly well into the active site of the AfeR model. The curved shape conformation is adapted to the shape of the active site, as was demonstrated for the natural ligand C14-AHL (see above). Moreover, analysis of the best ligand pose for 7-Z-C14-AHL shows that, as for C14-AHL, the hydrogen bond network involved strictly conserved residues and is equivalent to that obtained in X-ray structure analysis of AHL-LuxR protein family complexes (Figs. 4b, 7d) [31].
Conformational analysis of 7-Z-C14-AHL showed that the Z double-bond-induced curved shape conformation improved the fitting of this compound in the active site. Thus, the Z geometry of the double bond seems to induce a conformational constraint that optimizes binding of the ligand.
Conclusions
The transcriptional regulator AfeR, which is involved in the type AI-I QS system in the bacterium A. ferrooxidans, presents strictly or highly conserved residues corresponding to residues located in the active site of other AHL-dependent QS regulators. Based on this observation, a model of AfeR was constructed to study its interaction with long-chain AHLs. This work, performed on C14-AHL, revealed that the binding mode involved a curved shape conformation. The study of the binding mode of a long-chain AHL with a conformation constraint due to the presence of a cis double bond strongly suggests improved fitting of this compound into the active site of AfeR. These results might explain the existence of such mono-unsaturated AHLs acting as AIs. In order to evaluate the potential of AHL analogues to modulate QS in the bacterium A. ferrooxidans, in particular for 7-Z-C14-AHL and for other compounds with preferential curved shape conformations, the construction of a reporter strain as a specific biosensor for the QS system of this bacterium with the AfeR gene associated with the reporter gene Lux [22] is currently under investigation.
The present study should be helpful in the rational design of QS modulators for AfeR or in general for long-chain AHL-dependent transcriptional regulators involved in biotechnological applications such as the bioleaching of minerals. Moreover, the method presented here for the study of the binding mode using conformation analysis and subsequent RLD should be useful in the rational design of conformationally constrained bioactive compounds.
References
Camilli A, Bassler BL (2006) Science 311:1113–1116
Gonzalez JE, Keshavan ND (2006) Microbiol Mol Biol Rev 70:859–875
Miller MB, Bassler BL (2001) Annu Rev Microbiol 55:165–199
Reading NC, Sperandio V (2006) FEMS Microbiol Lett 254:1–11
Waters CM, Bassler BL (2005) Annu Rev Cell Dev Biol 21:319–346
van Leeuwen W, van Nieuwenhuizen W, Gijzen C, Verbrugh H, van Belkum A (2000) J Bacteriol 182:5721–5729
Chen X, Schauder S, Potier N, Van Dorsselaer A, Pelczer I, Bassler BL, Hughson FM (2002) Nature 415:545–549
Persson T, Givskov M, Nielsen J (2005) Curr Med Chem 12:3103–3115
Raffa RB, Iannuzzo JR, Levine DR, Saeid KK, Schwartz RC, Sucic NT, Terleckyj OD, Young JM (2005) J Pharmacol Exp Ther 312:417–423
Rasmussen TB, Givskov M (2006) Int J Med Microbiol 296:149–161
Smith JL, Fratamico PM, Novak JS (2004) J Food Prot 67:1053–1070
Geske GD, Wezeman RJ, Siegel AP, Blackwell HE (2005) J Am Chem Soc 127:12762–12763
Pomianek ME, Semmelhack MF (2007) ACS Chem Biol 2:293–295
Reverchon S, Chantegrel B, Deshayes C, Doutheau A, Cotte-Pattat N (2002) Bioorg Med Chem Lett 12:1153–1157
Geske GD, O’Neill JC, Blackwell HE (2007) ACS Chem Biol 2:315–319
Rohwerder T, Gehrke T, Kinzler K, Sand W (2003) Appl Microbiol Biotechnol 63:239–248
Olson GJ, Brierley JA, Brierley CL (2003) Appl Microbiol Biotechnol 63:249–257
Eberhard A, Burlingame AL, Eberhard C, Kenyon GL, Nealson KH, Oppenheimer NJ (1981) Biochemistry 20:2444–2449
Farah C, Vera M, Morin D, Haras D, Jerez CA, Guiliani N (2005) Appl Environ Microbiol 71:7033–7040
Rivas M, Seeger M, Holmes DS, Jedlicki E (2005) Biol Res 38:283–297
Valenzuela L, Chi A, Beard S, Orell A, Guiliani N, Shabanowitz J, Hunt DF, Jerez CA (2006) Biotechnol Adv 24:197–211
Frezza M, Castang S, Estephane J, Soulere L, Deshayes C, Chantegrel B, Nasser W, Queneau Y, Reverchon S, Doutheau A (2006) Bioorg Med Chem 14:4781–4791
Soulere L, Frezza M, Queneau Y, Doutheau A (2007) J Mol Graph Model 26:581–590
SYBYL Molecular Modeling System, Tripos Associates, St Louis, MO
Thompson MA (2004) ArgusLaB 4.0.1, Seattle, WA planetaria Software LLC
Krieger E, Koraimann G, Vriend G (2002) Proteins 47:393–402
Notredame C, Higgins DG, Heringa J (2000) J Mol Biol 302:205–217
Schwede T, Kopp J, Guex N, Peitsch MC (2003) Nucleic Acids Res 31:3381–3385
Thompson JD, Higgins DG, Gibson TJ (1994) Nucleic Acids Res 22:4673–4680
Morris GM, Goodsell DS, Halliday RS, Huey R, Hart WE, Belew RK, Olson AJ (1998) J Comp Chem 19:1639–1662
Zhang RG, Pappas T, Brace JL, Miller PC, Oulmassov T, Molyneaux JM, Anderson JC, Bashkin JK, Winans SC, Joachimiak A (2002) Nature 417:971–974
Jog GJ, Igarashi J, Suga H (2006) Chem Biol 13:123–128
Castang S, Chantegrel B, Deshayes C, Dolmazon R, Gouet P, Haser R, Reverchon S, Nasser W, Hugouvieux-Cotte-Pattat N, Doutheau A (2004) Bioorg Med Chem Lett 14:5145–5149
Muh U, Hare BJ, Duerkop BA, Schuster M, Hanzelka BL, Heim R, Olson ER, Greenberg EP (2006) Proc Natl Acad Sci USA 103:16948–16952
Yao Y, Martinez-Yamout MA, Dickerson TJ, Brogan AP, Wright PE, Dyson HJ (2006) J Mol Biol 355:262–273
Bottomley MJ, Muraglia E, Bazzo R, Carfi A (2007) J Biol Chem 282:13592–13600
Atkinson S, Chang CY, Sockett RE, Camara M, Williams P (2006) J Bacteriol 188:1451–1461
Llamas I, Keshavan N, Gonzalez JE (2004) Appl Environ Microbiol 70:3715–3723
Wagner-Dobler I, Thiel V, Eberl L, Allgaier M, Bodor A, Meyer S, Ebner S, Hennig A, Pukall R, Schulz S (2005) Chembiochem 6:2195–2206
Krick A, Kehraus S, Eberl L, Riedel K, Anke H, Kaesler I, Graeber I, Szewzyk U, Konig GM (2007) Appl Environ Microbiol 73:3587–3594
Saiz L, Klein ML (2001) Biophys J 81:204–216
Arab K, Rossary A, Soulere L, Steghens JP (2006) Br J Nutr 96:811–819
Acknowledgements
Financial support from MENESR and CNRS are gratefully acknowledged. This work was also supported by ECOS/CONICYT C05 B04, FONDECYT 1040676, ICM P05-001-F grants and ICM project P-05-001-F.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Soulère, L., Guiliani, N., Queneau, Y. et al. Molecular insights into quorum sensing in Acidithiobacillus ferrooxidans bacteria via molecular modelling of the transcriptional regulator AfeR and of the binding mode of long-chain acyl homoserine lactones. J Mol Model 14, 599–606 (2008). https://doi.org/10.1007/s00894-008-0315-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00894-008-0315-y