Abstract
The strain Streptomyces sp. Al-Dhabi-1 was isolated from soil sediments collected from Tharban hot spring in the southern west of Saudi Arabia using actinomycetes isolation agar and starch casein agar at 55 °C. Identification of the isolate was done according to morphological, physiological and biochemical characteristics and 16S rRNA sequence similarity as well. 16S rRNA sequence and blast analyses confirmed that the isolate belonging to the genus Streptomyces. The sequence was submitted to GenBank with accession number (KF815080). Ethyl acetate extract of Streptomyces sp. Al-Dhabi-1 showed good antimicrobial activities against tested pathogenic microbes. Minimum inhibitory concentration results showed that the best values were observed against S. agalactiae (<0.039 mg/ml) and Klebsiella pneumonia (0.125 mg/ml). Minimum inhibitory concentration of Al-Dhabi-1 against fungi; Cryptococcus neoformans (0.078 mg/ml), C. albicans (0.156 mg/ml), A. niger (0.625 mg/ml), and T. mentagrophytes (0.156 mg/ml). GC–MS analysis was used for the chemical profile of ethyl acetate extract. Benzeneacetic acid (16.02 %) and acetic acid 2-phenylethyl ester (10.35 %) were the major compounds among 31 substances found the ethyl acetate extract. According to the results of antimicrobial activity against pathogenic microbes, it is clear that the actinomycetes from hot springs with extreme environments are promising source for antimicrobial compounds.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Discovering of novel metabolites from natural sources which have a potent effect against resistant pathogenic bacteria is considered today as an important field of research. The diversity of natural products makes it one of the most important sources for novel structures with useful biological activities. Several studies are oriented towards isolation of new microorganisms from different habitats in the context of the search for novel antibiotics from new sources (Sujatha et al. 2005).
In Saudi Arabia, there are ten geothermal springs available (Khiyami et al. 2012). Geothermal systems are populated by diverse thermophilic bacteria and archaea (Kaur et al. 2008).
Actinomycetes produced a variety of bioactive compounds (secondary metabolites) such as antibiotics, herbicides, pesticides, anti-parasitic, and enzymes (Oskay et al. 2004; Hozzein et al. 2011). Streptomyces is the greatest producer genus for discovering antibiotics in the microbial world so far; approximately 80 % of the total antibiotics were produced by Streptomyces sp. (Ravel et al. 2000; Procópio et al. 2012). As a result of the emergence of resistant pathogens and the side effects of antimicrobial agents used at the present time, it becomes necessary to search for new antibiotics continuously (Arai et al. 1976). Therefore, in recent years thermophilic actinomycetes have attracted the attention of interested scientists. Owing to their diversity in biological activities and production of novel chemical compounds such as antibiotics like other microorganisms that inhabit extreme environments actinomycetes are promising (Kumar and Kannabiran 2010; Uzel et al. 2011).
The present study was aimed at isolating Streptomyces sp. Al-Dhabi-1 from hot spring and testing it for its antimicrobial activity. Identification of the isolate was done based on morphological properties, physiological and biochemical tests, in addition to analysis of 16S rRNA sequencing.
Materials and methods
Isolation, cultivation and maintenance of thermophilic actinomycetes
The strain Streptomyces sp. Al-Dhabi-1 was isolated from Tharban hot spring (52 °C) in Asir region southern west of Saudi Arabia (19°2′20.37″N, 41°40′45.13″E). Ten grams of dry sediments were added into 100 ml sterile distilled water and shaken for 30 min. Streptomyces sp. Al-Dhabi-1 was isolated using micro dilution technique. The sediment sample (0.5 ml from the dilution) was spread over onto following media actinomycetes isolation agar (AIA) (Himedia) and starch casein agar (SCA) (per liter: starch 10 g, KNO3 2 g, casein 0.3 g, NaCl 2 g, K2HPO4 2 g, MgSO4 7 H2O 0.05 g, CaCO3 0.02 g, FeSO4 7 H2O 0.01 g, agar 18 g, and pH 7.2) supplemented with 0.05 % cycloheximide to prevent the growth of fungi. The plates were incubated at 55 °C for 3–5 days. Streptomyces sp. Al-Dhabi-1 was subcultured on AIA and SCA until pure culture was obtained. The pure culture with 50 % (v/v) glycerol was prepared and stored in the freezer (Uzel et al. 2011).
Morphological, physiological and biochemical Characteristics
The pure isolate of Streptomyces sp. Al-Dhabi-1 grown in starch casein broth at 50 °C for 3 days. After 3 days Al-Dhabi-1 smear of culture was taken in a sterile glass slide and heated gently over a flame. The smear was covered with crystal violet for 1 min and washed gently in slow running tap water. Gram’s iodine solution was applied over the smear for 1 min and washed with running tap water. Alcohol solution was used to decolorize the smear. The slide was washed with water and counter stain safranine was flooded over the smear for 2 min, then the slide was washed, drained, air dried, and viewed under microscope. The culture retaining the violet color indicated that it was Gram-positive organism.
Morphological, physiological and biochemical properties were determined based on standard methods. The International Streptomyces Project (ISP) media were used as following ISP medium 2, ISP medium 3, ISP medium 4 and ISP medium 5 to determine the Arial mass color and soluble pigments; ISP medium 6 and ISP medium 7 were used for melanoid pigments, whereas ISP medium 9 was used to determine utilization of carbon source as recommended by Shirling and Gottlieb (1966). Utilization of nitrogen source, degradation of starch, gelatin and DNA were done according to Williams et al. (1983) and sensitivity to antibiotics was carried out as described by Petrova and Vlahov (2007). Ability to grow at different NaCl concentrations, temperatures and pH were done using ISP medium 2.
Molecular identification of active isolate using 16S rRNA
Streptomyces sp. Al-Dhabi-1 was grown in yeast peptone glucose broth (5 g peptone, 3 g yeast extract, 3 g NaCl, and 10 g glucose in 1 liter distilled water) until sufficient mycelium was formed for DNA extraction. High molecular weight genomic DNA was extracted from the isolate using NucleoSpin® Tissue kit (Macherey–Nagel, Duren, Germany) according to the manufacturer’s instructions. The 16S rRNA was amplified and sequenced by (Macrogen Inc., Korea) using primers F 27 (5′-AGAGTTTGATCCTGGCTCAG-3) and R 1492 (5′-TACGGCTACCTTGTTACGACTT-3′) (Dalisay et al. 2013).
Sequence of 16S rRNA of strain Al-Dhabi-1 was aligned with others retrieved from GenBank using Clustal W (Thompson et al. 1994). Phylogenetic analysis was performed using the software package MEGA 5 (Tamura et al. 2011). The resulting data were processed using the maximum-parsimony method (Fitch 1971) and the neighbor-joining method (Saitou and Nei 1987) for analysis of resultant data as well. The topology of the resultant unrooted tree was evaluated by bootstrap assay of the neighbor-joining tree, performed in 1000 replications.
Antimicrobial activity
Preliminary antimicrobial activity
Preliminary Antimicrobial screening assay was carried out for isolate Al-Dhabi-1 on Modified Nutrient Glucose Agar (MNGA) medium. The spore suspension of thermophilic Streptomyces sp. Al-Dhabi-1 was inoculated in a straight line on prepared agar plates and incubated at 50 °C for 7 days. After 7 days the following pathogenic microbes (ATTC) were used for preliminary screening; Bacillus cereus, Escherichia coli, Enterococcus faecalis, Klebsiella pneumoniae, Proteus vulgaris, Staphylococcus epidermidis, Salmonella typhimurium, Staphylococcus aureus, Pseudomonas aeruginosa and Streptococcus agalactiae, and yeast: Cryptococcus neoformans, Candida albicans. The above microbes were grown in nutrient broth at 37 °C for 24 h. The grown microbes were streaked perpendicular (‘T’-streak) to the strain Streptomyces sp. Al-Dhabi-1. The plates were incubated at 37 °C for 24 h. After the incubation period the antimicrobial activity was determined based on the distance of inhibition between the margin of thermophilic actinomycetes and pathogenic microbes. (Duraipandiyan and Ignacimuthu 2009).
Media optimization and antimicrobial metabolites production
For the production of bioactive compounds from the isolate Streptomyces sp. Al-Dhabi-1, the following media were used; Antibiotic Production Medium (APM), Micromonospora Medium (M3), Modified nutrient glucose medium (MNG), M6 media and yeast extract malt medium (YM). The pH of the medium was adjusted to pH-7.0 using 1 M HCl and 1 M NaOH. The culture was grown with continuous shaking on a rotary shaker (150 rpm) at 45 °C for 13 days. After growth the fermented broth was centrifuged and the supernatant was used as crude antibiotic extract. To study the influence of incubation periods the culture was maintained in the optimized production media for up to 16 days (Valan Arasu et al. 2012).
Extraction of antimicrobial metabolites
Modified nutrient glucose medium (MNG) was used for the extraction of antimicrobial compounds based on media optimization result. The strain Streptomyces sp. Al-Dhabi-1 was cultivated in MNG broth medium in a rotary shaker (150 rpm) at 45 °C for 10 days. To obtain the supernatant, the filtration of broth culture was done using Whatman No. 1 filter paper. The filtrate pH was adjusted to two using 0.1 N HCl to extract the majority of compounds in the filtrate. The ethyl acetate as an organic solvent was used for extracting the bioactive compounds by adding an equal volume (v/v) of the solvent to culture filtrate. The mixture of filtrate with organic solvent was appearing in two layers, the organic layer which contained the secondary metabolites and the aqueous layer. The crude extract was obtained after concentrating the solvent using rotary evaporator (IKA Rotary Evaporator model) at 60 °C and stored at 4 °C for antimicrobial assay.
Cup plate method
Antimicrobial activities for crude extracts were determined using the agar well diffusion method (Chaudhary et al. 2013). Crude extract of the isolate Al-Dhabi-1 was prepared for antibacterial activity as follows: 100 mg of crude extract was dissolved in 500 µl of dimethyl sulfoxide (DMSO). Sterile Mueller–Hinton Agar (MHA) (Hi-media, Mumbai) (25 ml) was poured into petri plates. 100 µl of suspension containing 108 CFU/ml of test bacteria was swabbed onto solidified media. Wells were made using sterile cork borer (5 mm in diameter) and filled with 25 µl of prepared extract (the concentration is 5 mg), 25 µl of streptomycin (the concentration is 10 µg) as positive control and 25 µl of DMSO as negative control and left at room temperature for 30 min for allowing the compounds to diffuse through the agar. The plates were incubated at 37 °C for 24 h. The zone of inhibition was recorded in millimeters.
Minimum inhibitory concentration
The crude extract (10 mg) was dissolved in 1 ml of dimethyl sulfoxide (DMSO): water (1:9) and used for minimum inhibitory concentration (MIC), using standard broth microdilution method (Duraipandiyan and Ignacimuthu 2009) and the MIC was calculated. Mueller–Hinton broth (Himedia, Mumbai) was prepared and sterilized by autoclaving at 121 °C, 15 lbs for 15 min. The required concentration of the extract (5, 2.5, 1.25, 0.625, 0.312, 0.156, and 0.0781 mg/ml) was added to the 96 well micro titer plate containing 0.2 ml broth. The 5 μl of inoculums culture suspension was introduced into the respective wells and the final inoculum size was 1 × 105 cfu/ml. The plates were incubated at 37 °C for 24 h. Positive control and solvent control (DMSO) were also included. MIC was determined as the lowest concentration of the extract which inhibited complete growth.
Identification of active principle using gas chromatography-mass spectrometry (GC–MS) analysis
The active ethyl acetate extract was quantified using gas chromatograph (GCMS-Shimadzu) equipped with a CPB-capillary column (mm inner diameter × 50 m length) mass spectrometer (ion source 200 °C, RI 70 eV) programmed at 40–280 °C with a rate of 4 °C/min. Injector temperature was 280 °C; carrier gas was He (20 psi). Sample volumes of 1 μl were injected with a split ratio of 25:1 using a hot needle. GC–MS analysis was carried out at Sargam Laboratory Service, Private Ltd, Chennai-600089, India.
Results
The genus Streptomyces has been studied and many reports have been published by earlier researchers relating to their morphology, physiology, biochemical, and biological properties in the medical field. Very limited reports have been published on thermophilic actinomycetes dealing with antimicrobial properties. Especially, in Saudi Arabia, there is not much more report on actinomycetes, particularly on thermophilic Streptomyces sp. Thermophilic Streptomyces sp. Al-Dhabi-1 was isolated from sediment soil collected from Tharban hot spring, Asir region, Saudi Arabia, using actinomycetes isolation agar (AIA) (Himedia, Mumbai) and starch casein agar.
Morphological, physiological and biochemical characteristics
The strain Streptomyces sp. Al-Dhabi-1 was filamentous gram-positive actinobacteria. Cultural characteristic of Al-Dhabi-1 showed that the isolate grew on different media (ISP2, ISP3, ISP4, ISP5, and MNGA) (Fig. 1); it formed green and whitish color mycelium and on the reverse side it showed yellowish-brown and white colors. The strain Al-Dhabi-1 never showed diffusible and melanoid pigments on any medium (Table 1).
The results of physiological and biochemical properties revealed that the strain Streptomyces sp. Al-Dhabi-1 grew at temperature between 37 and 55 °C; the optimal temperature was 50 °C, at pH between 6 and 11 with the normal pH of 7.0 and growth at 4 % NaCl concentration. The obtained results (Table 2) were compared to the relevant characteristics in Bergey’s Manual of Systematic Bacteriology (Whitman et al. 2012) and numerical classification of thermophilic streptomycetes (Goodfellow et al. 1987) for identifying the strain Al-Dhabi-1. This showed that the thermophilic strain Al-Dhabi-1 belonged to the genus Streptomyces.
Sequencing and phylogenetic analysis
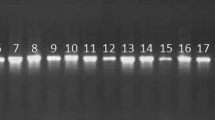
The sequencing result of 16S rRNA gene of strain Al-Dhabi-1 revealed that a total 1314 bp were obtained (Fig. 2). The sequence was submitted to GenBank with accession number (KF815080). The phylogenetic tree of strain Al-Dhabi-1 (Fig. 3) based on the neighbor-joining method formed a single clade with two strains of Streptomyces megaspores (16S rRNA gene sequence similarity of 98 %). The molecular characterization result of Al-Dhabi-1 revealed that the strain belonged to Streptomyces.
Relationship between Streptomyces sp. Al-Dhabi-1 and the closest phylogenetic relative (21) species in the single best tree inferred from neighbor-joining analysis based on partial 16S rRNA sequences (1314 nt). Numbers on the nodes indicate bootstrap values. The number between parentheses is the GenBank accession number
In vitro antimicrobial assay
In primary screening, the antimicrobial activity of Streptomyces sp. Al-Dhabi-1 was determined by the cross-streak method on Modified Nutrient Glucose Agar (MNGA) medium. The strain exhibited the significant activity against ATTC cultures (Table 3). Results of media optimization revealed that MNG medium was found to be the most suitable medium for antimicrobial compounds production. About 1 g was obtained as crude extract using ethyl acetate.
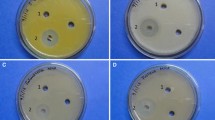
The ethyl acetate extract of Streptomyces sp. Al-Dhabi-1 was tested against pathogenic microbes using the well diffusion method. Results revealed that good antimicrobial activity was observed in the ethyl acetate extract of strain Al-Dhabi-1 (Fig. 4). The extract inhibited the growth of ATTC bacteria and fungi at a concentration of 5 mg/well for B.cereus (15 mm), E. coli (11 mm), E. Faecalis (16 mm), K. Pneumoniae (9 mm), P. vulgaris (9 mm), S. epidermidis (15 mm), S. typhimurium (14 mm), S. aureus (14 mm), P. aeruginosa (14 mm), S. agalactiae (18 mm) C. neoformans (15 mm), C. albicans (14 mm), A. niger (24 mm), and T. mentagrophytes (17 mm) (Table 4).
Minimum inhibitory concentration (MIC) was carried out to determine the active concentration of Streptomyces sp. Al-Dhabi-1 ethyl acetate extract that inhibited the growth of tested microorganism. MIC of the strain Streptomyces sp. Al-Dhabi-1 extract against pathogenic microbes was as follows; B. cereus (0.312 mg/ml), E. coli (0.625 mg/ml), E. faecalis (0.625 mg/ml), K. pneumoniae (1.56 mg/ml), P. vulgaris (0.312 mg/ml), S. epidermidis (0.312 mg/ml), S. typhimurium (0.312 mg/ml), S. aureus (0.312 mg/ml), P. aeruginosa (0.312 mg/ml) and S. agalactiae (0.078), C. neoformans (0.078 mg/ml), C. albicans (0.156 mg/ml), A. niger (0.625 mg/ml), and T. mentagrophytes (0.156 mg/ml) (Table 5). Significant MIC values of the strain Streptomyces sp. Al-Dhabi-1 extract were observed against S. agalactiae (<0.039 mg/ml), K. pneumonia (1.25 mg/ml) and C. neoformans (0.078 mg/ml).
GC–MS analysis of ethyl acetate extract
The active ethyl acetate extract of Streptomyces sp. Al-Dhabi-1 was studied using GC–MS chromatograph (Fig. 5). The GC–MS spectrum revealed the presence of many compounds (Table 6). The major compounds were: butanoic acid, 3-methyl ester (1.72 %), butanoic acid, 2-methyl ester (1.71 %), isopropyl alcohol (4.56 %), phenylethyl alcohol (1.11 %), acetic acid, 2-phenylethyl ester (10.35 %), benzeneacetic acid (16.02 %), benzoic acid 2-amino-methyl ester (5.19 %), 6-amino-1,3,5-triazine-2,4(1H,3H)-dione (1.08 %), pyrrolo[1,2-a]pyrazine-1,4-dione (2.34), l-leucine, N-cyclopropylcarbonyl (3.92 %), pyrrolo[1,2-a]pyrazine-1,4-dione (5.74 %), sebacic acid 2 6-dimethoxyphenyl (2.40 %), 2,5-piperazinedione, 3-methyl-6-[phenylmethyl (1.00 %), 2,5-piperazinedione, 3,6-bis(2-methylpropyl) (3.73 %)], pyrrolo[1,2-a]pyrazine-1,4-dione, hexahydro-3-[phenylmethyl (2.07 %), pyrimidine-2(1H)-thione, 4,4,6-trimethyl-1-(1-phenylethyl) (1.98 %], and cyclohexanecarboxylic acid, 4-heptyl-, 4-fluorophenyl ester (1.48 %).
Discussion
Actinomycetes are widespread in natural environments and mesophilic species have been and continue to be extensively screened for their potential for producing useful natural products. The isolated thermophilic Streptomyces sp. Al-Dhabi-1 showed good antimicrobial activity against tested microbes. Previously many researchers have reported that Streptomyces produced antimicrobial metabolites (Boudemagh et al. 2005; Fguira et al. 2005; Saravana Kumar et al. 2014; Govindarajan et al. 2014).
The isolate was confirmed as Streptomyces sp. by morphology, physiology and molecular characteristics. The Streptomyces sp. Al-Dhabi-1 was identified by the use of 16S rRNA gene sequence analysis for the molecular identification. The results showed high similarity of 98 % (E value 0.0) with 16S rRNA gene of S. megaspores. Contrarily to the traditional methods of actinomycetes identification, which are slow, difficult and very expensive, the molecular approach for identification is often used for their speed and efficiency (Kim et al. 1999; Labeda and Kroppenstedt 2000). The ethyl acetate extract of Streptomyces sp. Al-Dhabi-1 inhibited the growth of bacteria and fungi. Maximum zone of inhibition was observed against S. epidermidis (15 mm), S. typhimurium (14 mm), S. aureus (14 mm), P. aeruginosa (14 mm), S. agalactiae (18 mm) C. neoformans (15 mm), C. albicans (14 mm), A. niger (24 mm), and T. mentagrophytes (17 mm). Abussaud et al. (2013) investigated the antimicrobial activity in 8 thermophilic Streptomyces strains isolated from hot springs; the strains inhibited the growth of E. coli, S. aureus and C. albicans. Sahin (2003) reported that a total of 74 different Streptomyces isolates were recovered from 46 soil samples. From these isolates, 45.9 % isolates exhibited antibacterial activity and 29.4 % of the Streptomyces isolates inhibited the growth of S. epidermidis.
The production of antimicrobial metabolites was optimized using different media. The isolate Al-Dhabi-1 grew well in MNGA medium and produced metabolites. The media optimization study has been reported in our earlier work (Al-Dhabi et al. 2014). Most Streptomyces and other Actinomycetes are used in the production of a diverse array of antibiotics including aminoglycosides, macrolides, β-lactams, peptides, polyenes, polyether, tetracyclines, etc. (Subramani and Aalbersberg 2012). Al-Bari et al. (2005) reported that thermophilic Streptomyces bangladeshensis produced bis-(2-ethylhexyl) phthalate which exhibited antibacterial and antifungal activities against pathogenic microbes. Streptomyces sp. Al-Dhabi-1 produced an array of secondary metabolites; the major secondary metabolites were Acetic acid, 2-phenylethyl ester (10.35 %), benzeneacetic acid (16.02 %), and benzoic acid 2-amino-methyl ester (5.19 %). Previous researcher has shown that benzeneacetic acid showed antimicrobial properties (Tayade and Jadhao 2012).
The present study revealed that Streptomyces sp Al-Dhabi-1 isolated from hot spring of Saudi Arabia showed good antimicrobial activity against tested microbes in preliminary screening. Fandi et al. (2014) reported that thermophilic isolates isolated from hot spring of Jordan. The isolates were showed antimicrobial activity against pathogenic bacteria. Maataoui et al. (2014) have studied that antimicrobial Streptomyces strain isolated from deteriorated wood. The isolate was identified by 16S rRNA technique. The ethyl acetate extract of Streptomyces sp-H2 inhibited the growth of bacteria S. aureus. Anusuya and Geetha (2012) reported that isolation of thermophilic actinomycetes from banana waste. The isolates isolated at 50–55 °C temperature. The isolate Streptomyces sp. Al-Dhabi-1 also isolated same condition of temperature of 52 °C.
The ethyl acetate extract was taken from fermented broth of Al-Dhabi-1; it showed significant antimicrobial activity against bacteria and fungi at lowest concentration. The chemical profile was analysed using ethyl acetate extract; it revealed the diversity of secondary metabolites. In future we may isolate the active molecules from Streptomyces sp. Al-Dhabi-1 ethyl acetate extract to be used as drugs for the control of microbes causing infectious diseases.
References
Abussaud MJ, Alanagreh L, Abu-Elteen K (2013) Isolation, characterization and antimicrobial activity of Streptomyces strains from hot spring areas in the northern part of Jordan. Afr J Biotechnol 12:7124–7132
Al-Bari MA, Bhuiyan MS, Flores ME, Petrosyan P, Garcia-Varela M, Islam MA (2005) Streptomyces bangladeshensis sp. nov., isolated from soil, which produces bis-(2-ethylhexyl)phthalate. Int J Syst Evol Microbiol 55:1973–1977
Al-Dhabi NA, Duraipandiyan V, Valan Arasu M, Ponmurugan K, Ignacimuthu S (2014) Antifungal metabolites from sponge associated marine Streptomyces sp. Strain (ERIMA-01). J Pure Appl Microbiol 8:115–128
Anusuya D, Geetha M (2012) Isolation of thermophilic Actinomycetes from Banana waste compost and their biochemical characteristics. Int J Sci Res 3:315–317
Arai T, Yazawa K, Mikami Y (1976) Isolation and characterization of satellite antibiotics, mimosamycin and chlorocarcins from Streptomyces lavendulae, streptothricin source. J Antibiot 29:398–407
Boudemagh A, Kitouni M, Boughachiche F, Hamdiken H, Oulmi L, Reghioua S, Zerizer H, Couble A, Mouniee D, Boulahrouf A, Boiron P (2005) Isolation and molecular identification of actinomycete microflora, of some Saharian soils of south east Algeria (Biskra, EL-Oued and Ourgla)study of antifungal activity of isolated strains. J Mycol Med 15:39–44
Chaudhary HS, Yadav J, Shrivastava AR, Singh S, Singh AK, Gopalan N (2013) Antibacterial activity of actinomycetes isolated from different soil samples of Sheopur (a city of central India). J Adv Pharm Technol Res 4:118–123
Dalisay DS, Williams DE, Wang XL, Centko R, Chen J, Andersen RJ (2013) Marine sediment-derived Streptomyces bacteria from British Columbia, Canada are a promising microbiota resource for the discovery of antimicrobial natural products. PLoS One 8:e77078
Duraipandiyan V, Ignacimuthu S (2009) Antibacterial and antifungal activity of Flindersine isolated from the traditional medicinal plant, Toddalia asiatica (L.) Lam. J Ethnopharmacol 123:494–498
Fandi K, Muaikel NA, Momani FA (2014) Antimicrobial activities of some thermophiles isolated from Jordan hot springs. Int J Chem Environ Biol Sci 2:57–60
Fguira LFB, Fotso Serge, Ameur-Mehdi R, Mellouli L, Laatsch H (2005) Purification and structure elucidation of antifungal and antibacterial activities of newly isolated Streptomyces sp. strain US80. Res Microbiol 156:341–347
Fitch WM (1971) Towards defining the course of evolution: minimum change for a specific tree topology. Syst Zool 20:406–416
Goodfellow M, Lacey J, Todd C (1987) Numerical classification of thermophilic streptomycetes. J Gen Microbiol 133:3135–3149
Govindarajan G, Satheeja S, Jebakumar VSRD (2014) Antimicrobial potential of phylogenetically unique actinomycete, Streptomyces sp. JRG-04 from marine origin. Biologicals 42:305–311
Hozzein WN, Rabie W, Ali MIA (2011) Screening the Egyptian desert actinomycetes as candidates for new antimicrobial compound and identification of a new desert Streptomyces strain. Afr J Biotechnol 10:2295–2301
Kaur G, Mountain B, Pancost R (2008) Organic geochemistry, microbial membrane lipids in active and inactive sinters from Champagne Pool, New Zealand: elucidating past geothermal chemistry and microbiology. Org Geochem 39:1024–1028
Khiyami MA, Ehab AS, Shehata MM, Bahklia AH (2012) Thermo-aerobic bacteria from geothermal springs in Saudi Arabia. Afr J Biotechnol 11:4053–4062
Kim BN, Sahin DE, Minnikin J, Zakrewska-Czerwinska M, Mordarski M, Goodfellow M (1999) Classification of thermophilic Streptomycetes including the description of Streptomyces thermoalcalitolerans sp. nov. Int J Syst Bacteriol 49:7–17
Kumar S, Kannabiran K (2010) Antifungal activity of Streptomyces VITSVK5 spp. against drug resistant Aspergillus clinical isolates from pulmonary tuberculosis patients. J Mycol Med 20:101–107
Labeda DP, Kroppenstedt RM (2000) Phylogenetic analis of Saccharothrix and related taxa: proposal for Actinosynnemataceae fam. nov. Int J Syst Evol Microbiol 50:331–336
Maataoui H, Iraqui M, Jihani S, Ibnsouda S, Haggoud A (2014) Isolation, characterization and antimicrobial activity of a Streptomyces strain isolated from deteriorated wood. Afr J Microbiol Res 8:1178–1186
Oskay M, Tamer AÜ, Azeri C (2004) Antibacterial activity of some actinomycetes isolated from farming soils of Turkey. Afr J Biotechnol 3:441–446
Petrova D, Vlahov S (2007) Taxonomic characterization of the thermophilic actinomycete strain 21e-producer of thermostable collagenase. J Cult Collect 5:3–9
Procópio RE, Silva IR, Martins MK, Azevedo JL, Araújo JM (2012) Antibiotics produced by Streptomyces. Braz J Infect Dis 16:466–471
Ravel J, Wellington EMH, Hill RT (2000) Interspecific transfer of Streptomyces giant linear plasmids in sterile amended soil microcosms. Appl Environ Microbiol 66:529–534
Sahin N (2003) Investigation of the antimicrobial activity of some Streptomyces isolates. Turk J Biol 27:79–84
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Saravana Kumar P, Duraipandiyan V, Ignacimuthu S (2014) Isolation, screening and partial purification of antimicrobial antibiotics from soil Streptomyces sp. SCA 7. Kaohsiung J Med Sci 30:435–446
Shirling EB, Gottlieb D (1966) Methods for characterization of Streptomyces species. Int J Syst Bacteriol 16:313–340
Subramani R, Aalbersberg W (2012) Marine actinomycetes: an ongoing source of novel bioactive metabolites. Microbiol Res 167:571–580
Sujatha KVSN, Raju B, Ramana T (2005) Studies on a new marine Streptomyces BT-408 producing polyketide antibiotic SBR-22 effective against methicillin resistant Staphylococcus aureus. Microbiol Res 160:119–126
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Bio; Evol 28:2731–2739
Tayade DT, Jadhao NG (2012) Attempt in the synthesis of 2-[(2,6 di substituted thiocarbamidophenyl)amino] benzeneacetic acid and their antimicrobial study. J Pure Appl Microbiol 6:2025–2028
Thompson JD, Higgins DG, Gibson TJ (1994) CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22:4673–4680
Uzel A, Kocabas EH, Bedir E (2011) Prevalence of Thermoactinomyces thalpophilus and T. sacchari strains with biotechnological potential at hot springs and soils from West Anatolia in Turkey. Turk J Biol 35:195–202
Valan Arasu M, Asha KRT, Duraipandiyan V, Ignacimuthu S, Agastian P (2012) Characterization and phylogenetic analysis of novel polyene type antimicrobial metabolite producing actinomycetes from marine sediments: Bay of Bengal India. Asian Pac J Trop Biomed 2:803–810
Whitman W, Goodfellow M, Kämpfer P, Busse HJ, T ujillo M, Ludwig W, Suzuki K, Parte A (eds) (2012) Bergey’s manual of systematic bacteriology, the actinobacteria, vol 5. Springer, Berlin, pp 1638–1763
Williams ST, Goodfellow M, Alderson EMH, Wellington PHA, Sneath Sachin MJ (1983) Numerical classification of Streptomyces and related genera. J Gen Microbiol 129:1743–1813
Acknowledgments
This work was supported by the National Plan for Science, Technology and Innovation, (MAARIFAH), King Abdulaziz City for Science and Technology, Kingdom of Saudi Arabia, Award Number (11-BIO1873-02).
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by M. da Costa.
Rights and permissions
About this article
Cite this article
Al-Dhabi, N.A., Esmail, G.A., Duraipandiyan, V. et al. Isolation, identification and screening of antimicrobial thermophilic Streptomyces sp. Al-Dhabi-1 isolated from Tharban hot spring, Saudi Arabia. Extremophiles 20, 79–90 (2016). https://doi.org/10.1007/s00792-015-0799-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00792-015-0799-1