Abstract
We have studied the diversity of culturable halophilic Archaea at Rambla Salada, Murcia (south-eastern Spain). We made 8 samplings at different places in this habitat during the years 2006 and 2007 and isolated a total of 49 strains, which were identified by means of phenotypic tests and the hypervariable V1–V3 region of the 16S rRNA gene sequences (around 500 bp). The ribosomal data showed that the isolates belonged to 12 genera within the Halobacteriaceae family, with Haloferax and Natrinema being the most abundant. Five strains showed less than 97% sequence identity with validly described species and may well represent new taxa. All the strains grew best with around 25% w/v salts, required high concentrations of NaCl and magnesium and produced red to pink colonies. They were facultative anaerobes with both respiratory and fermentative metabolisms. The diversity of the archaeal community was analysed with the MOTHUR package. We identified 14 OTUs at the 3% genetic distance level and found quite high diversity. Rarefaction curves of richness estimators and diversity indices demonstrated that our collection of isolates represented the archaeal community at Rambla Salada that can be isolated under the conditions used in this work. This is the first report to be published on the culturable archaea at Rambla Salada, an area of considerable ecological interest.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
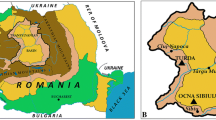
Rambla Salada is a saline “rambla” (a steep-sided river bed, normally dry but subject to flash flooding) located in Murcia (southeastern Spain). Rambla Salada has been declared a protected area by the Murcian regional government (BORM 10/09/1998), a place of community interest (LIC) by the European Union and a protected wildfowl zone (ZEPA). It is an athalassohaline habitat and includes areas of soil, water and sediments with different salt contents, deriving mainly from Miocene gypsiferous marls in the Fortuna basin (Muller and Hsü 1987). Nowadays, the habitat is seriously threatened by human activities that induce changes in the natural hydrology and salinity levels: inputs of freshwater, nutrients, pesticides and other pollutants are dramatically changing its biodiversity.
Velasco et al. (2006) first studied the primary producers and macro-invertebrates at Rambla Salada and demonstrated that their community composition was closely linked to salinity. Nevertheless, to our knowledge there have been no ecological studies so far describing the population of microorganisms inhabiting this rambla, although our group has in the past discovered two new halophilic bacterial species there: Idiomarina ramblicola (Martínez-Cánovas et al. 2004) and strain R53 of Halomonas cerina (González-Domenech et al. 2008). Thus, we undertook an analysis of the community of prokaryotes that live in various environments in the different areas of the Rambla Salada with the aims: firstly, of quantifying the archaeal community; secondly, of isolating a significant number of archaeal strains of those that represent the community of these organisms; finally, of identifying them and ascertaining their diversity.
Materials and methods
Sample collection and physical–chemical determinations
We took samples from four different zones in Rambla Salada (Murcia, southeastern Spain) (Table 1): soil, sediment and water at the Finca de la Salina (site 1), water and sediment from a saline groundwater spring (site 2), soil from the Humedal de Derramadores (site 3) and water and sediment from the Tajo-Segura interconnecting canal (site 4). We collected a total of 32 samples over 2 years (February and June 2006, and February and November 2007). The samples were taken aseptically and stored at 4°C until study in the laboratory (always within 24 h). The soils and sediments were suspended in sterile 25% w/v NaCl solution (1 g in 9 ml), thoroughly homogenized by stirring and then serially diluted (up to 10−6). Water samples were directly diluted in sterile 25% w/v NaCl solution. As much as 100 μl of dilutions 10−3, 10−4, 10−5 and 10−6 were surface-plated on MY medium (Moraine and Rogovin 1966) supplemented with 30% w/v sea-salt solution (Rodríguez-Valera et al. 1985, 1981) and incubated at 41°C for 3 weeks.
pH and conductivity were measured at each sampling point. Conductivity was determined with an ECmeter (TetraConR 325), which automatically calculates salinity.
Counts and selection of the strains
Counts were made in those plates containing between 30 and 300 colonies. A collection of 50 colonies, chosen on the basis of their different appearances, were re-isolated by streaking on a fresh casamino acids medium (medium 2) following the recommendations of Oren (2006) and grown at the same temperature for the same length of time. The sites where each strain was isolated are shown in Table 2.
DNA extraction, PCR amplification of 16S rRNA genes and sequencing
Genomic DNA was extracted from log-phase cells according to Marmur (1961) with the modification developed by Martín-Platero et al. (2007). The hypervariable V1–V3 regions of the 16S rRNA gene sequences (around 500 bp) were then determined using the method described by Burns et al. (2004) with the specific primers for Archaea: F1 as forward primer (5′-ATTCCGGTTGATCCTGC-3′) (Ihara et al. 1997) and 1492R as reverse primer (5′-ACGGHTACCTTGTTACGACTT′-3′) (Grant et al. 1999). PCR amplifications were made using 50 μl reaction mixtures containing 20–100 ng of template DNA, 10 pmol of each primer (Sigma®), 0.2 mM of dNTP mix (Bioline®), 2 mM of MgCl2, 5× PCR buffer (Bioline®) and 1.25 U of BioTaqTM DNA polymerase (Bioline®). Amplified PCR products were purified with the Illustra GFX DNA and Gel Band Purification kit (GE Healthcare®) and sequenced directly.
Sequence analysis
The sequences obtained were identified by a similarity-based search using the EzTaxon server 2.1. (http://147.47.212.35:8080/index.jsp) (Chun et al. 2007). Thereafter, the sequences were aligned using Clustal X (Thompson et al. 1997). To study the phylogenetic relationship among the isolates and other species of Halobacteriaceae, we applied neighbour-joining (NJ) and maximum parsimony (MP) criteria using the MEGA version 4 software (Tamura et al. 2007). Confidence levels for the phylogenetic trees were assessed by bootstrapping with 1000 replicates. The sequence of the type strain of Methanospirillum hungatei JF-1T was used as outgroup.
Phenotypic characterization
We carried out the phenotypic tests described by Oren et al. (1997), which are the minimal standards for the description of new taxa in the order Halobacteriales.
Diversity measures and rarefaction analysis
Sequence alignments of the 16S rRNA genes allowed us to construct a distance matrix using MOTHUR (http://www.mothur.org/) (Schloss et al. 2009), a software package integrating an improved version of DOTUR (Schloss and Handelsman 2005). Once the matrix was generated, we conducted an OTU-based analysis to study archaeal diversity. The clustering algorithm was furthest neighbour. We carried out rarefaction studies taking the default value of 1000 as the number of randomizations. We also calculated the Shannon (H′) diversity, the reciprocal of Simpson’s indexes (Simpson 1949; Magurran 1996) and Chao 1 and ACE species-richness estimators (Chao 1987).
Nucleotide sequence accession numbers
The sequences reported in this study have been submitted to the GenBank database under accession numbers HQ659121 to HQ659169.
Results
Physical–chemical measurements
Salinity in the different zones (sites 1–4) and samples (water, soil and sediment) taken at Rambla Salada ranged from 1.6 to 8.3% w/v in 2006 and from 1.5 to 3.4% w/v in 2007 with the exception of the water sampled at site 2 (natural groundwater spring), the salt content of which remained between 14 and 15.7% w/v. The pH ranged from 6.3 to 8.7 (Table 1).
Microbial counts and selection of the archaeal strains
Microbial counts (UFC/ml or UFC/g) revealed values of around 104 (1.2–4.3 × 104) in February 2006 and around 106 (1.2–2.6 × 106) in June 2006 and February and November 2007. We chose 50 isolates on the basis of the different appearances of their colonies (26 and 24 strains were chosen in 2006 and 2007, respectively). Phenotypic tests and ribosomal data (see below) proved that 49 of these 50 isolates were strains of archaea. This result suggested that the microbial counts could be attributed almost entirely to archaeal strains. The great majority of the colonies were red to pink, which is the norm among these microorganisms.
Phylogenetic analyses
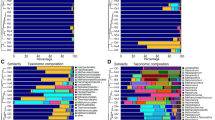
The use of specific primers for the hypervariable region of the 16S rRNA gene of Archaea and subsequent sequencing of the PCR product allowed us to determine a preliminary phylogeny of the isolates. Both NJ and MP methods gave similar clusters, supported by bootstrap values of above 70% (Fig. 1). Phylogenetic analyses indicated that all the strains were related to different genera within the Halobacteriaceae, with Haloferax and Natrinema being predominant with 10 representatives each. In addition, the 16S rRNA genes of strains M2-2d, M2-7b, M3-1c, M4-6a and M4-6b showed a similarity of less than 97% with other archaeal species, which leads us to surmise that they probably constitute new taxa (Table 2).
Neighbour-joining phylogenetic tree based on 16S rRNA gene sequences, showing the position of the archaeal isolates with respect to other members of the family Halobacteriaceae. The sequence of the type strain of Methanospirillum hungatei JF-1T was used as outgroup. Bar 1% sequence divergence. Common clusters resulting from both neighbour-joining and maximum-parsimony methods show bootstrap values at the corresponding nodes (in that order). The 5 strains with less than 97% similarity to validly described species are shaded in grey. GenBank/EMBL/DDBJ accession numbers are given in parenthesis
Phenotypic characterization
We used a total of 78 phenotypic tests to characterize the strains, in accordance with the minimal standards for the description of new taxa in the order Halobacteriales (Oren et al. 1997) (Table S1). All the strains were either Gram-negative rods or pleiomorphic, and extremely halophilic, growing best with 25% w/v sea salt. They required magnesium. They grew best between 37 and 41°C. Colonies ranged from pink to red in colour. They were facultative anaerobes. All fermented glucose and arginine and some of them respired with nitrate. They were resistant to ampicillin, chloramphenicol, erythromycin, nalidixic acid, penicillin and tetracycline.
Diversity measures and rarefaction analyses
Using the clustering algorithm implemented in the MOTHUR package, we identified 14 OTUs at the 3% distance level. We used rarefaction curves to compare the relative richness between the archaeal population from each sampling season, 2006 and 2007. The rarefaction analyses at 97% grouping stringency revealed that diversity was higher in 2006 than in 2007 (Fig. 2). Chao 1 and ACE rarefaction curves tend to be parallel to the x-axis, indicating a representative sampling under the conditions used (Fig. 3). The Chao 1 and ACE estimators predicted between 15 and 17 species at 97% grouping stringency: at a 95% confidence interval, the values for Chao 1 ranged between 14.18 and 26.47 and for ACE between 14.51 and 29.16, taking the whole sampling area into account (Fig. 3). Thus, the values for the predicted number of OTUs are quite close to the number of OTUs observed (see above). Furthermore, both the Chao 1 and ACE estimators were higher in 2006 (18.5 and 17.82) than in 2007 (8.2 and 11.3) (Table 3).
We also assessed diversity by means of Simpson’s and Shannon’s diversity indexes, using the MOTHUR programme and obtained values of 0.09 (reciprocal value of Simpson’s index 10.05) and 2.35 for the total sampling area, respectively (Table 3). These values reflect quite high archaeal diversity. Both indices were also higher in 2006 (10 and 2.16) than in 2007 (5 and 1.74), as seen previously. In comparison to other studies of archaeal populations in hypersaline habitats (Clementino et al. 2008; Baati et al. 2008, 2010; Pasić et al. 2005), we found lower values for the richness estimators and similar values for diversity indices (Table 3).
Discussion
All extremely halophilic archaea, or haloarchaea, cultured to date belong to the Halobacteriaceae family within the order Halobacteriales in the phylum Euryarchaeota. They are found extensively in such saline environments as salt lakes and saltern crystallizer ponds and also in saline soils (Oren 1994; Burns et al. 2004; Grant et al. 2001; Maturrano et al. 2006; Pasić et al. 2005; Dave and Desai 2006). In recent years, some publications have also described their presence in medium-to-low-salinity environments or even non-saline habitats (Aller and Kemp 2008; Cambon-Bonavita et al. 2009). Furthermore, new molecular ecology techniques have found archaea belonging to the phylum Crenarchaeota in saline habitats, but they remain to be uncultured.
Our study has demonstrated that Rambla Salada is host to a substantial density and diversity of culturable halophilic archaea belonging to the Halobacteriaceae, even in zones of low and medium salinity (see Table 2) and that they represent a diverse group of taxa belonging to different genera and species.
As far as the total counts are concerned, they were high and quite similar to those obtained in other hypersaline habitats, such as solar salterns in Alicante (Spain) (Rodríguez-Valera et al. 1985, 1981) and in San Francisco (CA, USA) (Litchfield et al. 1999), where the values were 104 and 105–106 UFC/ml, respectively.
Salinity is one of the most important driving forces of diversity for both macro- and microorganisms (Auguet et al. 2010; Lozupone and Knight 2007; Tamames et al. 2010). According to Velasco et al. (2006), this is reflected in the composition of the communities of primary producers and macro-invertebrates at Rambla Salada. Our work has also demonstrated that the diversity of archaea is affected to some extent by this factor.
In our study, salinity was automatically calculated from the conductivity measurements made in situ during each sampling season. Generally, the salinity gradient reached its highest values in 2006. At site 2, a natural spring, the salinity values were practically constant throughout the sampling period, which might be expected from a permanent flow of saline groundwater. We found the highest biodiversity in 2006, which, according to the physical–chemical parameters measured, may well be related to higher salinity. Thus, in 2006 we isolated 12 genera (Haladaptatus, Haloarcula, Halococcus, Haloferax, Halogeometricum, Halomicrobium, Halorhabdus, Halorubrum, Halostagnicola, Haloterrigena, Natrialba and Natrinema), whilst in 2007 only 8 genera were identified (Haladaptatus, Halococcus, Haloferax, Halomicrobium, Halostagnicola, Haloterrigena, Natrialba and Natrinema). We found several taxa that were isolated only during one of the sampling periods. Thus, Halorubrum aidingense, Haloarcula argentinensis, Haloarcula quadrata, Halogeometricum borinquense and Halorhabdus tiamatea were isolated in 2006, whilst Halomicrobium mukohataei, Haloferax prahovense, Halococcus hamelinensis and Haladaptatus paucihalophilus were isolated in 2007.
On the other hand, it should be pointed out that in none of our samples did we find the extreme halophilic bacterium Salinibacter, the most significant features of which are similar to archaea (Antón et al. 2002).
As shown in Table 3, higher diversity was related to higher salinity, which, according to their diversity indices, is also the trend observed in hypersaline archaeal populations from other habitats (Clementino et al. 2008; Baati et al. 2008, 2010; Pasić et al. 2005).
In this study we did not detect all the archaea living in the habitat and the introduction of more isolation media, such as those mentioned by Burns et al. (2004), would probably allow the cultivation of a greater number of taxa, but rarefaction analyses using various richness estimators suggest that the total number of sequences studied within the area of Rambla Salada covers most of the culturable archaea under our chosen conditions. Moreover, although the Chao 1 and ACE estimators normally underestimate true richness when sample sizes are small (Hughes et al. 2001), we found in our study that the estimated value was quite similar to that observed.
In general terms, the diversity of haloarchaea at Rambla Salada was similar to that in other saline environments (Oren 2002; Burns et al. 2004; Baati et al. 2008, 2010; Clementino et al. 2008; Ozcan et al. 2007). Nevertheless, the predominant taxa were different. In solar salterns, one of the most thoroughly studied types of hypersaline habitat, the predominant population tends to be made up of strains belonging to the genera Haloferax, Halorubrum, Halococcus, Haloterrigena, Haloarcula, Natrialba and Halobacterium (Oren 2002). In saltern crystallizers, the predominant archaea are Halorubrum and Haloquadratum (Oren 2002). In Rambla Salada, however, we isolated more strains belonging to the genera Natrinema and Haloferax, this latter often being found in habitats with low salinity, although it grows in media containing 1.0–5.1 M NaCl and grows best at 2.5 M NaCl (Oren 2011). Nevertheless, these taxa were not dominant when the archaeal population was analysed by molecular methods. In fact, Oueragli (communication personal) found that Haloarcula is the most abundant taxon in Rambla Salada.
This study is the first to describe the culturable halophilic-archaeal community at Rambla Salada. Our results confirm the presence of a substantial biodiversity and density of archaea in this environment. We have in addition discovered a number of strains that may well constitute new taxa and are being subject to further scrutiny in our laboratory.
References
Aller JY, Kemp PF (2008) Are Archaea inherently less diverse than Bacteria in the same environments? FEMS Microbiol Ecol 65:74–87
Antón J, Oren A, Benlloch S, Rodríguez-Valera F, Amann R, Rossselló-Mora R (2002) Salinibacter ruber gen. nov., sp. nov., a novel, extremely halophilic member of the Bacteria from saltern crystallizer ponds. Int J Syst Evol Microbiol 52:485–491
Auguet JC, Barberan A, Casamayor EO (2010) Global ecological patterns in uncultured Archaea. ISME J 4:182–190
Baati H, Guermazi S, Amdouni R, Gharsallah N, Sghir A, Ammar E (2008) Prokaryotic diversity of a Tunisian multipond solar saltern. Extremophiles 12:505–518
Baati H, Guermazi S, Gharsallah N, Sghir A, Ammar E (2010) Novel prokaryotic diversity in sediments of Tunisian multipond solar saltern. Res Microbiol 161:573–582
Burns DG, Camakaris HM, Janssen PH, Dyall-Smith ML (2004) Combined use of cultivation-dependent and cultivation-independent methods indicates that members of most haloarchaeal groups in an Australian crystallizer pond are culturable. Appl Environ Microbiol 70:5258–5265
Cambon-Bonavita MA, Nadalig T, Roussel E, Delage E, Duperron S, Caprais JC, Boetius A, Sibuet M (2009) Diversity and distribution of methane-oxidizing microbial communities associated with different faunal assemblages in a giant pockmark of the Gabon continental margin. Deep sea res part II: topical studies in oceanography 56:2248–2258
Chao A (1987) Estimating the population size for capture–recapture data with unequal catchability. Biometrics 43:783–791
Chun J, Lee JH, Jung Y, Kim M, Kim S, Kim BK, Lim YW (2007) EzTaxon: a web-based tool for the identification of prokaryotes based on 16S ribosomal RNA gene sequences. Int J Syst Evol Microbiol 57:2259–2261
Clementino MM, Vieira RP, Cardoso AM, Nascimento AP, Silveira CB, Riva TC, González AS, Paranhos R, Albano RM, Ventosa A, Martins OB (2008) Prokaryotic diversity in one of the largest hypersaline coastal lagoons in the world. Extremophiles 12:595–604
Dave SR, Desai HB (2006) Microbial diversity at marine saltern near Bhavnagar, Gujarat, India. Curr Sci 90:497–500
González-Domenech CM, Martínez-Checa F, Quesada E, Béjar V (2008) Halomonas cerina sp. nov., a moderately halophilic, denitrifying, exopolysaccharide-producing bacterium. Int J Syst Evol Microbiol 58:803–809
Grant S, Grant WD, Jones BE, Kato C, Li L (1999) Novel archaeal phylotypes from an East African alkaline saltern. Extremophiles 3:139–145
Grant WD, Kamekura M, McGenity TJ, Ventosa A (2001) Order I. Halobacteriales. In: Garrity GM (ed) Bergey’s manual of systematic bacteriology V.I. The Archaea and deeply branching and phototrophic bacteria. Springer, New York, pp 213–235
Hughes JB, Hellmann JJ, Ricketts TH, Bohannan BJ (2001) Counting the uncountable: statistical approaches to estimating microbial diversity. Appl Environ Microbiol 67:4399–4406
Ihara K, Watanabe S, Tamura T (1997) Haloarcula argentinensis sp. nov. and Haloarcula mukohataei sp. nov., two new extremely halophilic archaea collected in Argentina. Int J Syst Bacteriol 47:73–77
Litchfield CD, Irby A, Vreeland RH (1999) The microbial ecology of solar salt plants. In: Oren A (ed) Microbiology and biogeochemistry of hypersaline environments. CRC Press, Boca Ratón, pp 39–52
Lozupone CA, Knight R (2007) Global patterns in bacterial diversity. Proc Natl Acad Sci USA 104:11436–11440
Magurran AE (1996) Ecological diversity and its measurement. Chapman and Hall, London
Marmur J (1961) A procedure for the isolation of deoxyribonucleic acid from microorganisms. J Mol Biol 3:207–208
Martínez-Cánovas MJ, Béjar V, Martínez-Checa F, Páez R, Quesada E (2004) Idiomarina fontislapidosi sp. nov. and Idiomarina ramblicola sp. nov., isolated from inland hypersaline habitats in Spain. Int J Syst Evol Microbiol 54:1793–1797
Martín-Platero AM, Valdivia E, Maqueda M, Martínez-Bueno M (2007) Fast, convenient, and economical method for isolating genomic DNA from lactic acid bacteria using a modification of the protein “salting-out” procedure. Anal Biochem 366:102–104
Maturrano L, Santos F, Rosselló-Mora R, Antón J (2006) Microbial diversity in Maras salters, a hypersaline environment in the Peruvian Andes. Appl Environ Microbiol 72:13887–13895
Moraine RA, Rogovin P (1966) Kinetics of polysaccharide B-1459 fermentation. Biotechnol Bioeng 8:511–524
Muller DW, Hsü KJ (1987) Event stratigraphy and paleoceanography in the Fortuna basin (Southeast Spain): a scenario for the Messinian salinity crisis. Paleoceanography 2:679–696
Oren A (1994) The ecology of the extremely halophilic archaea. FEMS Microbiol Rev 13:415–440
Oren A (2002) Halophilic microorganisms and their environments. Kluwer Academic Publishers, Netherlands
Oren AT (2006) The order Halobacteriales. In: Dworkin M, Falkow S, Rosenberg E, Schleifer K-H, Stackebrandt E (eds) The prokaryotes a handbook on the biology of bacteria. Springer, New York, pp 113–164
Oren A (2011) Diversity of halophiles. In: Horikoshi K (ed) Extremophiles handbook. Springer, Tokyo, pp 309–325
Oren A, Ventosa A, Grant WD (1997) Proposed minimal standards for description of new taxa in the order Halobacteriales. Int J Syst Evol Microbiol 47:233–238
Oren A, Arahal DR, Ventosa A (2009) Emended descriptions of genera of the family Halobacteriaceae. Int J Syst Evol Microbiol 59:637–642
Ozcan B, Ozcengiz G, Coleri A, Cokmus C (2007) Diversity of halophilic archaea from six hypersaline environments in Turkey. J Microbiol Biotechnol 17:985–992
Pasić L, Bartual SG, Ulrih NP, Grabnar M, Velikonja BH (2005) Diversity of halophilic archaea in the crystallizers of an Adriatic solar saltern. FEMS Microbiol Ecol 54:491–498
Rodríguez-Valera F, Ventosa A, Juez G, Imhoff J (1985) Variation of the environmental features and microbial population with salt concentrations in a multipond saltern. Microbiol Ecol 11:107–115
Rodríguez-Varela F, Ruiz-Berraquero F, Ramos-Cormenzana A (1981) Characteristics of the heterotrophic bacterial populations in hypersaline environments of different salt concentration. Microbiol Ecol 7:235–243
Schloss PD, Handelsman J (2005) Introducing DOTUR, a computer program for defining operational taxonomic units and estimating species richness. Appl Environ Microbiol 71:1501–1506
Schloss PD, Westcott SL, Ryabin T, Hall JR, Hartmann M, Hollister EB, Lesniewski RA, Oakley BB, Parks DH, Robinson CJ, Sahl JW, Stres B, Thallinger GG, Van Horn DJ, Weber CF (2009) Introducing MOTHUR: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl Environ Microbiol 75:7537–7541
Simpson EH (1949) Measurement of diversity. Nature 163:688
Tamames J, Abellán JJ, Pignatelli M, Camacho A, Moya A (2010) Environmental distribution of prokaryotic taxa. BMC Microbiol 10:85
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4: molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol Biol Evol 24:1596–1599
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25:4876–4882
Velasco J, Millán A, Hernández J, Gutiérrez C, Abellán P, Sánchez D, Ruíz M (2006) Response of biotic communities to salinity changes in a Mediterranean hypersaline stream. Saline Syst 2:12
Acknowledgments
The authors thank Josefa Velasco and Andrés Millán for their assistance with field work and zone selection for sampling. Our thanks also go to Miguel Angel Núñez of the Explanation Centre at Rambla Salada for his willing assistance in the field. This research was supported by grants from the Dirección General de Investigación Científica y Técnica (CGL2005-05947; CGL2008-02399), from the Plan Andaluz de Investigación and Plan Propio de la Universidad de Granada, Spain. We also thank our colleague Dr J. Trout for revising our English text.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by A. Oren.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Luque, R., González-Domenech, C.M., Llamas, I. et al. Diversity of culturable halophilic archaea isolated from Rambla Salada, Murcia (Spain). Extremophiles 16, 205–213 (2012). https://doi.org/10.1007/s00792-011-0420-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00792-011-0420-1