Abstract
Thermus strain SA-01, previously isolated from a deep (3.2 km) South African gold mine, is closely related to Thermus strains NMX2 A.1 and VI-7 (previously isolated from thermal springs in New Mexico, USA, and Portugal, respectively). Thermus strains SA-01 and NMX2 A.1 have also been shown previously to grow using nitrate, Fe(III), Mn(IV) or SO as terminal electron acceptors and to be capable of reducing Cr(VI), U(VI), Co(III), and the quinone-containing compound anthraquinone-2,6-disulfonate. The objectives of this study were to determine the phylogenetic positions of the three known metal-reducing Thermus strains and to determine the phylogenetic significance of metal reduction within the genus Thermus. Phylogenetic analyses of 16S rDNA sequences, BOX PCR genomic fingerprinting, and DNA–DNA reassociation analyses indicated that these strains belong to the previously described genospecies T. scotoductus. The morphologies and lipid fatty acid profiles of these metal-reducing strains are consistent with their identification as T. scotoductus; however, the T. scotoductus strains tested in this study evinced a wide intraspecies variability in some other phenotypic traits, e.g., carbon substrate utilization and pigmentation. Iron reduction occurred in all strains of T. scotoductus tested except the mixotrophic, sulfur-oxidizing strain IT-7254. Thermus strains belonging to other species did not reduce Fe(III) to Fe(II) or reduced it only poorly.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Increasing interest in the microbiology of extreme environments has resulted in the isolation and characterization of a great diversity of thermophilic bacteria, including several novel species within the genus Thermus, such as Thermus scotoductus (Kristjansson et al. 1994), Thermus brockianus (Williams et al. 1995), Thermus oshimai (Williams et al. 1996), Thermus igniterrae, and Thermus antranikianii (Chung et al. 2000). Members of the genus Thermus were formerly considered to be strictly aerobic, based on the characteristics of the species Thermus aquaticus (Brock 1984). However, many of the more recently described Thermus isolates have been shown to grow anaerobically using nitrate as the terminal electron acceptor (Ramirez-Arcos et al. 1998a, 1998b; Williams and Sharp 1995; da Costa et al. 2001).
Kieft et al. (1999) isolated Thermus strain SA-01 from groundwater in a deep South African gold mine and characterized it as a facultative anaerobe capable of coupling the oxidation of organic substrates to reduction of a wide range of electron acceptors, including nitrate, Fe(III), Mn(IV), and SO. Suspensions of Thermus strain SA-01 could also reduce Co(III)-EDTA, Cr(VI), U(VI), Tc(VII), and the quinone-containing compound anthraquinone-2,6-disulfonate (AQDS) (Kieft et al. 1999). Similar metabolic versatility has been demonstrated in Thermus strain NMX2 A.1, which was isolated from a thermal spring in New Mexico (Hudson et al. 1989). Analysis of 16S rDNA sequences showed that strains SA-01 and NMX2 A.1 are very closely related (Kieft et al. 1999). Thermus strain VI-7, which was isolated from a thermal spring in Portugal (Santos et al. 1989), is also a close phylogenetic relative of strains SA-01 (Kieft et al. 1999) and NMX2 A.1 (Williams and Sharp 1995) and can also reduce Fe(III) (Kieft et al. 1999). Another close relative is Thermus scotoductus strain IT-7254, which was isolated from a thermal spring in Iceland and was shown to be capable of mixotrophic growth using SO or thiosulfate as energy sources (Skirnisdottir et al. 2001). Oxidation of arsenite and reduction of arsenate have also been shown recently in the genus Thermus (Gihring and Banfield 2001; Gihring et al. 2001). Clearly, some species of Thermus are more metabolically versatile than was known heretofore.
The objectives of this study were to determine the phylogenetic positions of three closely related metal-reducing Thermus strains—SA-01 (Kieft et al. 1999), NMX2 A.1 (Hudson 1986), and VI-7 (Santos et al. 1989)—and to determine the phylogenetic significance of metal reduction in the genus Thermus. We compared the morphological, biochemical, and phylogenetic characteristics of SA-01, NMX2 A.1, and VI-7 with those of related Thermus strains, including T. scotoductus, with particular focus on the occurrence of iron-reducing capabilities.
Materials and methods
Sources of bacterial strains and culture maintenance
The Thermus strains included in this study are listed in Table 1. Thermus strain SA-01 [ATCC 700910; Subsurface Microbial Culture Collection (SMCC; Balkwill 1993) LX-001] was isolated as described previously (Kieft et al. 1999) from groundwater collected at a depth of 3.2 km below land surface in a South African gold mine, where the ambient rock temperature is approximately 60°C. Thermus strain NMX2 A.1 was provided by Hugh Morgan, University of Waikato, Hamilton, New Zealand. Strains CG-2, D1, NH, and VI-7 were provided by Milton S. da Costa, Universidade de Coimbra, Portugal. T. scotoductus strain IT-7254 was provided by Sigurlaug Skirnisdottir, Prokaria Ltd., Reykjavik, Iceland. All other Thermus strains were obtained from the American Type Culture Collection (http://www.atcc.org/).
Thermus strains were routinely cultured under aerobic conditions in TYG medium (5 g tryptone, 3 g yeast extract, and 1 g glucose per liter) (strains SA-01, VI-7, T-351, Thermus filiformis Wai33 A.1T, T. thermophilus HB8T, and T. aquaticus YT-1T), ATCC growth medium 461 (T. scotoductus strain SE-1T), ATCC growth medium 697 (strains NMX2 A.1 and X-1), ATCC growth medium 1598 (IB-57), and medium 166 (Skirnisdottir et al. 2001) (strains NH, D1, and IT-7254). Stock cultures were stored at –80°C in 16% glycerol.
Physiological characteristics
Single carbon growth tests were performed using medium 162 of Degryse et al. (1978), with the various carbon substrates present at a concentration of 20 mM and with yeast extract and tryptone omitted. These cultures were incubated aerobically at 65°C. Anaerobic growth with nitrate as the electron acceptor was evaluated in TYG broth cultures containing 1 g KNO3 l–1 and purged with O2-free N2. A defined basal medium formulated for cultivating Geobacter chapellii and described by Kieft et al. (1999) was used for anaerobic Fe(III) reduction analyses. For these analyses, 10 mM Fe(III) was added as Fe(III)-nitrilotriacetate (NTA) to maintain Fe solubility. Carbon sources (sodium lactate or sodium formate) were added at 10 mM. The headspace gas for these experiments was N2:CO2 (80:20). Cells were first cultivated aerobically in appropriate media and then inoculated, after purging with O2-free N2, into the anaerobic basal medium to achieve a cell density of 5×107 cells ml–1. Cultures were incubated at 65°C with shaking. Reduction of Fe(III)-NTA was determined by measuring Fe(II) in 0.5 N HCl extracts (Kieft et al. 1999) using the ferrozine assay (Stookey 1970).
Lipid analysis
Thermus strains were grown in TYG (strains SA-01 and VI-7), ATCC medium 697 (NMX2 A.1 and T. scotoductus SE-1T), or ATCC medium 461 (T. scotoductus X-1). Each culture was freeze-dried prior to lipid analysis. All solvents were GC grade (Fisher, Pittsburgh, Pa.). All glassware was washed in a 10% (v/v) Micro cleaning solution (VWR Scientific, Pittsburgh, Pa.), rinsed ten times in tap water, then ten times in deionized water. The glassware was then heated at 450°C for 4 h prior to use. Lipids were extracted from duplicate samples (50 mg) using the modified Bligh and Dyer technique (White et al. 1979). Total lipids were fractionated into glyco-, neutral and polar lipids; the polar lipid fraction was then transesterified with mild alkali to recover the phospholipid fatty acids (PLFA) as methyl esters in hexane (Guckert et al. 1985). Trimethyl silyl (TMS) derivatives of glycolipids were generated as described in Wait et al. (1997). The PLFA and TMS-glycolipids were separated, quantified, and identified by GC-MS (Ringelberg et al. 1994). Fatty acids and glycolipids were identified by relative retention times, by comparison with authentic standards (Matreya Inc., Pleasant Gap, Pa.), and by the mass spectra (collected at an electron energy of 70 mV). Fatty acid nomenclature is as described by White and Ringelberg (1998). PLFA and glycolipid mole percent data were arcsine transformed (White and Ringelberg 1998), and then hierarchical cluster analyses were performed with a single linkage method using Minitab version 11 (Minitab, State College, Pa.).
Molecular genetic analyses
Genomic DNA was isolated from cultures incubated overnight in TYG at 65°C. Cells were harvested by centrifugation and washed once with 5 M NaCl. The DNA isolation and purification DNA was carried out with the QIAGEN Genomic Tip 100G kit (Qiagen Inc., Valencia, Calif.). The isolated genomic DNA was mixed with 0.1-mm glass beads and sheared by bead beating (Mini BeadBeader, BioSpec Products, Bartlesville, Okla.) to a size range of 3 to 6 kb. DNA base compositions were determined (by Marin Biologic Contract/Research Laboratories, Tiburon, Calif.) by measuring the thermal melt temperatures (Marmur and Doty 1962).
Repetitive DNA fingerprinting was performed following the method of Louws et al. (1994). The PCR primer derived from the repetitive sequence BOX A1R (5′-CTACGGCAAGGCGACGCTGACG-3′) was used in PCR amplification of Thermus DNA (purified with the QIAGEN Genomic Tip column as described above). The PCR mixtures contained 2.5 units of Taq polymerase, 1.5 mM MgCl2, 50 mM KCl, and 10% dimethyl sulfoxide in a 50-μl reaction volume. The PCR protocol was 80°C for 5 min (hot start); 35 cycles of 94°C for 1 min, 44°C for 30 s, and 65°C for 6 min; followed by a final extension step at 65°C for 20 min. PCR products (25 μl of each) were electrophoresed in 1.5% agarose to resolve the amplified DNA fragments.
Thermus cultures were sent to the Deutsche Sammlung von Mikroorganismen und Zellkulturen (DSMZ), GmbH, Braunschweig, Germany (http://www.dsmz.de/) for DNA-DNA reassociation analyses. DSMZ uses the hydroxyapatite chromatography method for DNA isolation (Cashion et al. 1977) and the method of De Ley et al. (1970) for DNA-DNA reassociation as modified by Huss et al. (1983) and Escara and Hutton (1980).
16S rRNA gene sequences for Thermus strains SA-01, NMX2 A.1, and VI-7 were obtained from the GenBank/EMBL database. These sequences were initially aligned to all sequences for Thermus strains available from both the Ribosomal Database Project (RDP; Maidak et al. 2001) and GenBank/EMBL databases. The sequences were aligned by hand, based on the secondary structure of the 16S rRNA molecule (Gutell 1994; Gutell et al. 1994). The resulting alignment was analyzed using the distance matrix technique to generate a preliminary phylogenetic tree (not shown). Duplicate sequences and those not needed for comparison to strains SA-01, NMX2 A.1, and VI-7 (based on evaluation of the tree) were then removed, leaving 26 sequences in the alignment for a more detailed analysis. The final alignment of 1,368 bases included sequences for Thermus strains SA-01, NMX2 A.1, and VI-7; representative sequences for all recognized species of Thermus; and sequences for several other eubacteria that were used for comparison or as an outgroup during analyses. The phylogenetic positions of Thermus strains SA-1, NMX2 A.1, and VI-7 were analyzed using the distance matrix, maximum-likelihood, and parsimony methods. Distance matrix analysis was performed with the PHYLIP group of computer programs (Felsenstein 1993). Maximum-likelihood analysis was performed with the fastDNAml program (Olsen et al. 1994). The PAUP* software program (PAUP* 4.0, Beta Version 4.0b4a; Swofford 2000) was used for parsimony analysis.
Results and discussion
Morphological and biochemical characteristics
Thermus strains SA-01, NMX2, and VI-7 and T. scotoductus strains X-1 and SE-1T varied in their pigment production; they also showed considerable variability in their ability to utilize various simple sugars, amino acids, and organic acids as sole C sources (Table 2). None of the substrates tested could be utilized by all five of these strains.
Metal reduction
A previous study demonstrated that Thermus strains SA-01 and NMX2-A.1 were able to couple oxidation of lactate and acetate to Fe(III) reduction for energy and growth (Kieft et al. 1999). These strains could also couple oxidation of formate to reduction of Fe(III) to Fe(II) (Fig. 1). As noted previously, both strains reduced some Fe(III), even in the absence of an exogenous electron donor (Figs. 1, 2a). This was most likely due to oxidation of storage carbohydrates, oxidation of the NTA used to complex and maintain solubility of Fe(III), or both (Kieft et al. 1999). Neither H2 nor acetate stimulated Fe(III) reduction beyond that observed for the cultures that did not receive an electron donor, whereas pyruvate was equally or more effective than lactate as an electron donor (data not shown).
Reduction of 10 mM Fe(III) to Fe(II) during 7 days’ incubation at 65°C. a Reduction by various Thermus strains with 10 mM lactate as the electron donor. Error bars represent one standard deviation (n =3). b Reduction with 10 mM formate as the electron donor (n=1). Values have been corrected for the amount of Fe(II) produced in the absence of lactate or formate as electron donor
Various strains of Thermus showed differences in their abilities to reduce Fe(III) (Fig. 2). Of the T. scotoductus strains tested, only IT-7254 was unable to reduce Fe(III). Strains of T. aquaticus (YT-1T), T. thermophilus (HB8T), and T. filiformis (strains T351 and Wai33 A1T) reduced Fe(III) poorly. Nearly all of these T. scotoductus strains are capable of dissimilatory Fe(III) reduction, the only exception in this study being strain IT-7254. In addition to the ability to grow with Fe(III)-NTA as the sole terminal electron acceptor, SA-01 and NMX2 A.1 can reduce Fe(III)-citrate, hydrous ferric oxide (HFO), Mn(IV), Cr(VI), U(VI), Co(III)-EDTA, and the quinone-containing compound anthraquinone-2,6-disulfonate (AQDS) (Kieft et al. 1999). It remains to be seen whether the other T. scotoductus strains shown here to be iron reducers share this metabolic diversity with SA-01 and NMX2 A.1. Curiously, IT-7254 shows metabolic versatility of a different sort in that it can grow mixotrophically while oxidizing SO or thiosulfate.
Fatty acid composition
The membrane PLFAs of the five strains tested in this study (Table 3) were dominated by terminally branched saturated fatty acids, as has been shown previously for other members of this genus (Hensel et al. 1986; Tenreiro et al. 1995; Wait et al. 1997; Chung et al. 2000; da Costa et al. 2001). Terminally branched saturated fatty acids comprised over 95 mol% of the PLFAs in the five strains tested, with iso-branched 15:0 and 17:0 PLFAs being the dominant forms. Cluster analysis of PLFAs placed T. scotoductus strain SE-1T somewhat distant from the other four strains (data not shown). The glycolipid fatty acid profiles also showed slight differences among Thermus strains SA-01, NMX2 A.1, and VI-7 and the two T. scotoductus strains (Table 4). The finding of long-chain 1,2-diols in T. scotoductus strain X-1 confirms the work of Wait et al. (1997) and extends it to four closely related strains. The function of these diol-linked glycolipids remains unknown. Cluster analysis of the glycolipid fatty acid profiles separated Thermus strains SA-01, NMX2 A.1, and VI-7 from the T. scotoductus strains (data not shown).
Molecular genetic analysis
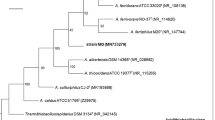
The mole percent G+C contents, as determined by thermal denaturation, were 65.0 (±0.4) for SA-01, 65.8 (±0.6) for NMX2 A.1, and 65.9 (±0.6) for VI-7. These values are very close to the 64–65% reported for T. scotoductus strains X-1 and SE-1 (Kristjansson et al. 1994). Phylogenetic trees of 16S rRNA gene sequences generated by parsimony, maximum-likelihood, and distance methods showed that strains SA-01, NMX2 A.1, and VI-7 clustered with previously described strains of T. scotoductus. Thermus SA-01, NMX2 A.1, and VI-7 sequences were 98.4–99.5% similar to T. scotoductus sequences and <96.8% similar to sequences from strains other than T. scotoductus (1,368 bases compared). The 16S rRNA gene sequence-based phylogenetic analysis thus supports the identification of the metal-reducing strains SA-01, NMX2 A.1, and VI-7 as T. scotoductus.
Because bacterial phylogeny cannot be resolved solely by analysis of 16S rRNA gene sequences if those sequences are more than 97% similar, DNA-DNA reassociation studies and/or other analyses are needed to make species-level comparisons (Fox et al. 1992; Stackebrandt and Goebel 1994). Whole genome DNA-DNA hybridizations revealed significant similarities (76.7–89.9% reassociation) between strains X-1, SE-1, NH, D1, IT-7254, and CG-2 and genomic DNA from Thermus strains SA-01 and NMX2 A.1. Thus, all eight strains appear to belong to the species T. scotoductus, according to the 70% DNA-DNA reassociation criterion (Wayne et al. 1987). Genomic fingerprinting of the BOX A repeat sequence showed close similarities between the three metal-reducing strains (SA-01, NMX2 A.1, and VI-7) and Thermus scotoductus strains X-1 and SE-1T (Fig. 3). In general, the BOX fingerprints data were consistent with the 16S rDNA-based phylogenetic analysis.
Phylogenetic significance and geographic distribution
The morphological, biochemical, and physiological characteristics of metal-reducing strains SA-01, NMX2 A.1, and VI-7 confirm their placement in the genus Thermus. Lipid profiles of SA-01, NMX2 A.1, and VI-7 confirm their tight phylogenetic relationship to each other and further demonstrate a similarity to T. scotoductus strains X-1 and SE-1T. Phylogenetic analyses based on 16S rDNA sequences and genomic DNA-DNA hybridization clearly show that the closely related metal-reducing strains SA-01, NMX2 A.1, and VI-7, plus strain CG-2, fall within the previously described species T. scotoductus. Thermus strains NMX2 A.1 and VI-7 were suggested (though never formally proposed) by Williams and Sharp (1995) to comprise a separate species, “T. imahorii,” based on 16S rRNA gene sequences.
While T. scotoductus comprises a phylogenetically distinct group at the species level, it is phenotypically variable, probably more so than most Thermus species described to date. For example, the organic substrates that can be used for growth vary greatly among T. scotoductus strains (Table 2; Kristjansson et al. 1994). Pigmentation is also variable and thus appears not to be a useful diagnostic character for discriminating T. scotoductus strains, or for all metal-reducing strains within the species. The yellow pigmentation observed in NMX2 A.1 is common in most other Thermus spp., including T. aquaticus, T. thermophilus, T. filiformis, T. brockianus, T. oshimai, T. igniterrae, and T. antranikianii (Chung et al. 2000), but is not a characteristic of previously described T. scotoductus strains (Tenreiro et al. 1995). The soluble melanin-like pigment reported for T. scotoductus strains X-1 and SE-1T (Kristjansson et al. 1994) was not observed in the metal-reducing strains SA-01, NMX2 A.1, and VI-7. Nearly all T. scotoductus strains can reduce iron, while other species appear to reduce iron poorly; thus, dissimilatory metal reduction may be a useful phylogenetic trait for identifying this species and may also suggest a useful strategy for obtaining strains of this species in enrichment culture. However, unequivocal identification of T. scotoductus must rely on phylogenetic analysis.
It is interesting to note the broad global distribution of the metal-reducing T. scotoductus strains: SA-01 was isolated from ground water collected from a deep subsurface gold mine in South Africa (Kieft et al. 1999); NMX2 A.1 was isolated from a hot spring in Jemez Springs in New Mexico (Hudson 1986); and VI-7 was isolated from a hot spring, the Caldas de Vizela, in Portugal (Santos et al. 1989). This distribution suggests that metal-reducing T. scotoductus exists in hot, subterranean waters worldwide, possibly because of its versatility with regard to the utilization of a wide range of electron acceptors. The observation of broad geographic distribution of Thermus genospecies is consistent with the conclusion of Williams et al. (1996) that the “concept of circumscribed geographical distribution of Thermus species may be more apparent than real.”
References
Balkwill D (1993) DOE makes subsurface cultures available. ASM News 59:504–506
Brock TD (1984) The genus Thermus. In: Krieg NR, Holt JG (eds) Bergey’s manual of systematic bacteriology. Williams & Wilkins, Baltimore, pp 333–337
Brock TD, Freeze H (1969) Thermus aquaticus gen. n. sp. n., a non-sporulating extreme thermophile. J Bacteriol 98:289–297
Cashion P, Hodler-Franklin MA, McCully J, Franklin M (1977) A rapid method for base ratio determination of bacterial DNA. Anal Biochem 81:461–466
Chung AP, Rainey FA, Valente M, Nobre MF, da Costa MS (2000). Thermus igniterrae sp. nov. and Thermus antranikianii sp. nov., two new species from Iceland. Int J Syst Evol Microbiol 50:209–217
da Costa MS, Nobre MF, Rainey F (2001) Genus I. Thermus. In: Garrity G, Boone DR, Castenholz RW (eds) Bergey’s manual of systematic bacteriology, vol 1. The Archaea, cyanobacteria, phototrophs & deeply branching bacteria. Springer, Berlin Heidelberg New York, pp 404–414
Degryse E, Glansdorff N, Pierard A (1978) A comparative analysis of extreme thermophilic bacteria belonging to the genus Thermus. Arch Microbiol 117:189–196
De Ley, J, Cattoir, H, Reynaerts A (1970) The quantitative measurement of DNA hybridization from denaturation curves. Eur J Biochem 12:133–142
Escara JF, Hutton JR (1980) The thermal stability of DNA in dimethylsulphoxide solutions: acceleration of renaturation rate. Biopolymers 19:1315–1327
Felsenstein J (1993) PHYLIP (Phylogeny Inference Package) Version 3.5c., 3.5c edn. University of Washington, Seattle, Washington
Fox GE, Wisotzkey JD, Jurtshuk P J (1992) How close is close: 16S rRNA sequence identity may not be sufficient to guarantee species identity. Int J Syst Bacteriol 42:166–170
Gihring TM, Banfield JF (2001) Arsenite oxidation and arsenate respiration by a new Thermus isolate. FEMS Microbiol Lett 204:335–340
Gihring TM, Druschel G K, McCleskey RB, Hamers RJ, Banfield JF (2001) Rapid arsenite oxidation by Thermus aquaticus and Thermus thermophilus: field and laboratory investigations. Environ Sci Technol 35:3857–3862
Guckert JB, Antworth CP, Nichols PD, White DC (1985) Phospholipid ester-linked fatty acid profiles as reproducible assays for changes in prokaryotic community structure of estuarine sediments. FEMS Microbiol Ecol 31:147–158
Gutell RR (1994) Collection of small subunit (16S- and 16S-like) ribosomal RNA structures. Nucleic Acids Res 22:3502–3507
Gutell RR, Larsen N, Woese CR (1994) Lessons from an evolving rRNA: 16S and 23S rRNA structures from a comparative perspective. Microbiol Rev 58:10–26
Hensel R, Demharter W, Kandler O, Kroppenstedt RM, Stackebrandt (1986) Chemotaxonomic and molecular-genetic studies of the genus Thermus: evidence for a phylogenetic relationship of Thermus aquaticus and Thermus ruber to the genus Deinococcus. Int J Syst Bacteriol 36:444–453
Hudson JA (1986) The taxonomy and ecology of the genus Thermus. PhD dissertation, University of Waikato, Hamilton, New Zealand
Hudson JA, Morgan HW, Daniel RM (1987) Thermus filiformis sp. nov., a filamentous caldoactive bacterium. Int J Syst Bacteriol 37:431–436
Hudson JA, Morgan HW, Daniel RM (1989) Numerical classification of Thermus isolates from globally distributed hot springs. Syst Appl Microbiol 11:250–256
Huss VAR, Festl H, Schleifer KH (1983) Studies on the spectrometric determination of DNA hybridization from renaturation rates. J Syst Appl Microbiol 4:184–192
Kieft TL, Fredrickson JK, Onstott TC, Gorby YA, Kostandarithes HM, Bailey TJ, Kennedy DW, Li SW, Plymale AE, Spadoni CM, Gray MS (1999) Dissimilatory reduction of Fe(III) and other electron acceptors by a Thermus isolate. Appl Environ Microbiol 65:1214–1221
Kristjansson JK, Hreggvidsson GO, Alfredsson GA (1986) Isolation of halotolerant Thermus spp. from submarine hot springs in Iceland. Appl Environ Microbiol 52:1313–1316
Kristjansson JK, Hjorleifsdottir S, Marteinsson VT, Alfredsson GA (1994) Thermus scotoductus, sp. nov., a pigment-producing thermophilic bacterium from hot tap water in Iceland and Including Thermus sp. X-1. Syst Appl Microbiol 17:44–50
Louws FJ, Fulbright DW, Stephens T, de Bruijn FJ (1994) Specific genomic fingerprints of phytopathogenic Xanthomonas and Pseudomonas pathovars and strains generated with repetitive sequences and PCR. Appl Environ Microbiol 60:2286–2295
Maidak BL, Cole JR, Lilburn TG, Parker CT Jr, Saxman PR, Farris RJ, Garrity GM, Olsen GJ, Schmidt TM, Tiedje JM (2001) The RDP-II (Ribosomal Database Project). Nucleic Acids Res 29:173–174
Marmur J, Doty P (1962) Determination of the base composition of deoxyribonucleic acid from its thermal denaturation temperature. J Mol Biol 5:109–118
Olsen GJ, Matsuda H, Hagstrom R, Overbeek R (1994) fastDNAmL: a tool for construction of phylogenetic trees of DNA sequences using maximum likelihood. Comput Appl Biosci 10:41–8
Oshima T, Imahori K (1974) Description of Thermus thermophilus (Yoshida and Oshima) comb. nov., a non-sporulating thermophilic bacterium form a Japanese thermal spa. Int J Syst Bacteriol 24:102–112
Pask-Hughes RA, Williams RAD (1975) Extremely thermophilic Gram-negative bacteria from hot tap water. J Gen Microbiol 88:321–328
Ramaley RF, Hixson J (1970) Isolation of non-pigmented, thermophilic bacterium similar to Thermus aquaticus. J Bacteriol 103:527–528
Ramirez-Arcos S, Fernandez-Herrero L A, Berenguer J (1998a) A thermophilic nitrate reductase is responsible for the strain specific anaerobic growth of Thermus thermophilus HB8. Biochim Biophys Acta 1396:215–27
Ramirez-Arcos S, Fernandez-Herrero LA, Marin I, Berenguer J (1998b) Anaerobic growth, a property horizontally transferred by an Hfr-like mechanism among extreme thermophiles. J Bacteriol 180:3137–43
Ringelberg DB, Townsend GT, DeWeerd KA, Suflita JM, and White DC (1994) Detection of the anaerobic dechlorinating microorganism Desulfononiletiedjei in environmental matrices by its signature lipopolysaccharide branch-long-chain hydroxy fatty acids. FEMS Microbiol Ecol 14:9–18
Santos MA, Williams RAD, da Costa MS (1989) Numerical taxonomy of Thermus isolates from hot springs in Portugal. Syst App Microbiol 12:310–315
Skirnisdottir S, Hreggvidsson GO, Holst O, Kristjannson JK (2001) Isolation and characterization of a mixotrophic sulfur-oxidizing Thermus scotoductus. Extremophiles 5:45–51
Stackebrandt E, Goebel BM (1994) Taxanomic note: a place for DNA-DNA reassociation and 16S rRNA sequence analysis in the present species definition in bacteriology. Int J Syst Bacteriol 44:846–849
Stookey LL (1970) Ferrozine—a new spectrophotometric reagent for iron. Anal Chem 42:779–781
Swofford DL (2000) PAUP* 4.0, beta version 4.0b4a, Sinauer Associates, Inc., Sunderland, Maryland
Tenreiro S, Nobre MF, da Costa MS (1995a) Thermus silvanus sp. nov. and Thermus chliarophilus sp. nov., two new species related to Thermus ruber but with lower growth temperatures. Int J Syst Bacteriol 45:633–639
Tenreiro S, Nobre MF, Hoste B, Gillis M, Kristjansson JK, da Costa MS (1995b) DNA:DNA hybridization and chemotaxonomic studies of Thermus scotoductus. Res Microbiol 146:315–324
Wait R, Carreto L, Fernandez Nobre M, Ferreira AM, da Costa MS (1997) Characterization of novel long chain 1,2-diols in Thermus species and demonstration that Thermus strains contain both glycerol-linked and diol-linked glycolipids. J Bacteriol 179:6154–6162
Wayne LG, Brenner, DJ, Colwell RR, Grimont PAD, Kandler O, Krichevsky MI, Moore LH, Moore WEC, Murray RGE, Stackebrandt E, Starr MP, Trüper HG (1987) Report of the ad hoc committee on reconciliation of approaches to bacterial systematics. Int J Syst Bacteriol 37:463–464
White DC, Ringelberg DB (1998) Signature lipid biomarker analysis. In: Burlage RS, Atlas R, Stahl D, Geesey G, Sayler G (eds) Techniques in microbial ecology. Oxford University Press, New York, pp 255–272
White DC, Davis WM, Nickels JS, King JD, Bobbie RJ (1979) Determination of the sedimentary microbial biomass by extractable lipid phosphate. Oecologia 40:51–62
Williams RAD, Sharp RJ (1995) The taxonomy and identification of Thermus. In: Sharp RJ, Williams RAD (eds) Thermus species: a biotechnology handbook. Plenum Press, New York, pp 1–42
Williams RAD, Smith KE, Welch SG, Micallef J, Sharp RJ (1995) DNA relatedness of Thermus strains, description of Thermus brockianus sp. nov., and proposal to reestablish Thermus thermophilus (Oshima and Imahori). Int J Syst Bacteriol 46:495–499
Williams RAD, Smith KE, Welch SG, Micallef J (1996) Thermus oshimai sp. nov., isolated from hot springs in Portugal, Iceland, and the azores, and comment on the concept of a limited geographical distribution of Thermus species. Int J Syst Bacteriol 46:403–408
Acknowledgments
We thank Mary McHale and Gwendolyn R. Drake for technical assistance. This research was supported by grant no. EAR9978267 from the National Science Foundation, Life in Extreme Environments (LExEn) Program, and by grants from the National Aeronautics and Space Administration’s Astrobiology Institute.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by J. Wiegel
Rights and permissions
About this article
Cite this article
Balkwill, D.L., Kieft, T.L., Tsukuda, T. et al. Identification of iron-reducing Thermus strains as Thermus scotoductus . Extremophiles 8, 37–44 (2004). https://doi.org/10.1007/s00792-003-0357-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00792-003-0357-0