Abstract
Methane hydroxylation by metal-oxo oxidants is one of the Holy Grails in biomimetic and biotechnological chemistry. The only enzymes known to perform this reaction in Nature are iron-containing soluble methane monooxygenase and copper-containing particulate methane monooxygenase. Furthermore, few biomimetic iron-containing oxidants have been designed that can hydroxylate methane efficiently. Recent studies reported that μ-nitrido-bridged diiron(IV)-oxo porphyrin and phthalocyanine complexes hydroxylate methane to methanol efficiently. To find out whether the reaction rates are enhanced by replacing iron by ruthenium, we performed a detailed computational study. Our work shows that the μ-nitrido-bridged diruthenium(IV)-oxo reacts with methane via hydrogen atom abstraction barriers that are considerably lower in energy (by about 5 kcal mol‒1) as compared to the analogous diiron(IV)-oxo complex. An analysis of the electronic structure implicates similar spin and charge distributions for the diiron(IV)-oxo and diruthenium(IV)-oxo complexes, but the strength of the O‒H bond formed during the reaction is much stronger for the latter. As such a larger hydrogen atom abstraction driving force for the Ru complex than for the Fe complex is found, which should result in higher reactivity in the oxidation of methane.
Graphic abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
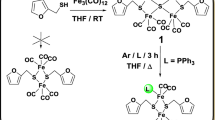
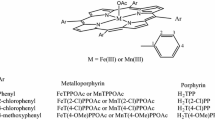
Heme monoxygenases are common enzymes in biology with a variety of functions related to biosynthesis and biodegradation. In general, they react through oxygen atom transfer to substrates on an iron(III)-heme co-factor that binds molecular oxygen, but uses two reduction and two protonation equivalents in the catalytic cycle. The most extensively studied heme monoxygenases are the cytochromes P450, which initiate the biodegradation of drug molecules in the liver as well as the biosynthesis of hormones [1,2,3,4,5,6,7,8,9,10]. During their catalytic cycle, the iron(III)-heme reacts with molecular oxygen and using two external electrons and protons, a high-valent iron(IV)-oxo heme cation radical species called Compound I (Cpd I) is formed [11,12,13]. Although Cpd I is able to hydroxylate a large range of aliphatic and aromatic C–H bonds, it is not known to hydroxylate methane, which has the strongest C–H bond in nature. However, work of Sorokin et al. on biomimetic porphyrin and phthalocyanine complexes (Fig. 1) found evidence of methane hydroxylation by μ-nitrido-bridged diiron(oxo) porphyrin and phthalocyanine [14,15,16] and as such these complexes are unique and highly reactive as well as the supramolecular diiron phthalocyanine–porphyrin conjugates recently published [17, 18]. In previous work, the synthesis of several μ-nitrido-bridged diiron(III) phthalocyanine and porphyrin complexes was reported, and using terminal oxidants such as hydrogen peroxide or m-chloroperbenzoic acid, they were converted to a μ-nitrido-bridged diiron(IV)-oxo species [19]. These short-lived intermediates were efficient in a reaction with aliphatic substrates (cyclohexane, adamantane, and ethylbenzene) leading to substrate hydroxylation [19]. Furthermore, methane hydroxylation to methanol was observed with several complexes, which implicates that these oxidants are more powerful than cytochrome P450 Cpd I [20, 21].
Unprecedented reactivity of µ-nitrido diiron tetrapyrrolic complexes has initiated synthetic development of this platform involving different metals supported by various macrocyclic ligands [17, 18, 22,23,24,25,26,27]. In parallel, several detailed computational studies on μ-nitrido-bridged diiron(IV)-oxo phthalocyanine and porphyrin complexes have been reported by us and others [28,29,30,31,32]. In general, these studies showed that the electron-donating ability of the μ-nitrido group lowers the acidity of the corresponding iron-hydroxo species, and consequently, the strength of the O–H bond of the iron(III)-hydroxo group is large. As the driving force for a hydrogen atom abstraction reaction is larger when a stronger O–H bond is formed [33,34,35], this implies that a significant enhancement of the rate constant for hydrogen atom abstraction will be observed. In this context, it is of great interest to probe how the nature of metal sites might influence on the catalytic properties of µ-nitrido binuclear construction. To gain further insight into the properties and reactivities of μ-nitrido-bridged dimetal-oxo porphyrins and phthalocyanines, we decided to create the analogous diruthenium complexes, and compare the structure, electronic properties, and catalysis with the diiron complexes. We predict that these diruthenium(IV)–oxo phthalocyanine complexes if they can be formed will react with methane even more efficiently than their corresponding iron complexes.
Methods
The work presented here uses computational methods and procedures as reported and discussed previously on biomimetic model complexes that reproduced experimental data well [36, 37]. Overall, density functional theory (DFT) approaches were used as implemented in the Gaussian-09 program package [38]. The full potential energy profile was calculated with two unrestricted DFT methods, namely the hybrid density functional method UB3LYP [39, 40] and the pure density functional UBP86 [41, 42], for all geometry optimizations, geometry scans, and frequencies. Geometry optimizations and potential energy scans were performed with a double-ζ quality LACVP basis set (with core potential) on ruthenium and 6-31G on the rest of the atoms, basis set BS1 [43, 44]. All local minima and transition states were optimized without constraints and characterized with an analytical frequency that confirmed the status of the structures with all transition states having a single imaginary frequency for the correct mode. Calculations include a polarized continuum model (CPCM) as implemented in Gaussian using a dielectric constant of ε = 35.688 mimicking acetonitrile. Energies were improved through a single-point calculation with an LACV3P + (with core potential) basis set on ruthenium and 6-311 + G* on the rest of the atoms: basis set BS2. These methods were used previously and reproduced experimentally determined free energies of activation and kinetic isotope effects well [45, 46]. In the past, we validated our computational methods and showed that these procedures can reproduce experimental free energies of activation to within 3 kcal mol−1. Moreover, changing the basis set for geometry optimizations from BS1 to BS2 gave little changes to the optimized geometries, relative energies, and chemoselectivities of the reaction [47,48,49]. Finally, the effect of dispersion on the optimized geometries of μ-nitrido-bridged diiron(IV)-oxo porphyrins was tested for the defluorination reaction of C6F6 and found to give little changes in geometry and energetics, and hence, dispersion was not used in this work [50].
Results and discussion
In this work, we focus on the chemical properties of the μ-nitrido bound diruthenium(IV)-oxo porphyrazine (Pz) complex 2,4,6[O=RuIV(Pz+·)NRuIV(Pz)]0 (or 2,4,6[O=RuV(Pz)NRuIV(Pz)]0), 1, whereby all side chains of the macrocycle are abbreviated to hydrogen atoms. The complex is charge neutral and was calculated in all low-lying doublet, quartet, and sextet spin states using two density functional theory methods (UB3LYP and UBP86). In addition, the reactivity patterns of the complexes with methane was compared with the analogous diiron(IV)-oxo complex 2,4,62 reported previously [29,30,31].
Before we show the results on the catalytic properties of oxidant 2,4,61, let us investigate the electronic and structural properties of the reactant species in more detail. Figure 2 displays the optimized geometries and relative energies of 2,4,61. In both complexes, the doublet spin state is the ground state and well separated from the quartet and sextet spin states by at least 10 kcal mol−1. This is independent on the density functional method chosen and implicates that the quartet and sextet spin states will play no role in catalysis. As such, the reactivity with substrates is expected to take place on the doublet spin state only and the oxidants will react through single-state reactivity [51, 52] selectively. Mononuclear iron(IV)-oxo oxidants, by contrast often have close-lying spin-state surfaces, where reactivity patterns appear on multiple accessible electronic and spin states. It is not surprising that the ruthenium complexes react through single-state-reactivity patterns as RuIV=O complexes tend to have well separated metal 4d orbitals and hence usually stabilize low-spin states [53,54,55]. Indeed, previous studies on mononuclear RuIV=O complexes showed the high-spin states to be considerably higher in energy than the lower spin states [56] in agreement with what is seen here.
Optimized geometries of 2,4,61 (left-hand side) and 2,4,62 (right-hand side) as obtained in Gaussian-09 at UB3LYP/BS1 (UPB86/BS1). Bond lengths are in angstroms and relative energies (calculated with BS2 basis set with zero-point energy (ZPE) correction) in kcal mol‒1. Data for 2,4,62 taken from Ref. [29]
In structures 2,42, the Fe1‒O and Fe2‒μ–N distances were found to be about 1.65 Å in length, which indicates that both bonds will be formally a double bond. In the ruthenium complexes, both of these bonds have significantly elongated with respect to those of the diiron complexes as expected for a heavier element. However, the Ru1‒O distances are significantly longer than the Ru2‒μ–N distances, which implicates that they have different bonding character. Furthermore, the ruthenium atom of the Ru1‒O group is located below the plane through the four nitrogen atoms of the equatorial ligand, while in the iron complexes, the Fe1 atom remains above the plane. Finally, particularly in the low-spin state, the bridging nitrogen atom is close to the center of the Ru1‒Ru2 interaction, whereas in the corresponding diiron(IV)-oxo species, it is closer to Fe2 than to Fe1.
To understand the differences in geometry between the diiron and diruthenium complexes, we analyzed the molecular orbitals, which are displayed in Fig. 3. The orbitals are dominated by the π-interactions in the xz and yz molecular planes, where we take the z-axis along the Ru–O bond. Thus, the 4dxz and 4dyz atomic orbitals on both Ru atoms interact with the 2px and 2py atomic orbitals on the oxo and bridging nitrogen atoms to form four sets of orbitals: π1,x/π1,yπ2,x/π2,yπ*3,x/π*3,yπ*4,x/π*4,y. The lowest two sets of orbitals represent the bonding interactions for the Ru‒O and Ru‒N interaction. The π*3,x and π*3,y orbitals have a bonding interaction between the top Ru atom and the axial ligand, but are antibonding for the Ru‒O and Ru‒N interactions. The doublet spin state for both 21 and 22 has orbital occupation π 21,x π 21,y π 22,x π 22,y π* 23,x π* 13,y π* 04,x π* 04,y . These orbital occupations are quite different from typical mononuclear heme complexes, i.e., FeIV=O(heme+·) or P450 Cpd I, that have a heme radical with singly occupied a2u orbital. In the μ-nitrido-bridged complexes, by contrast, the a2u orbitals are lower in energy and are doubly occupied.
Group spin densities of the doublet spin-state reactants give dominant oxo radical character (ρO = 0.90 at UB3LYP and 0.56 for the UBP86 calculation). Nevertheless, in both cases, the radical refers to a singly occupied π*3,y molecular orbital. These two results give S2 values of 0.792 and 0.775 and hence include very little multiconfiguration perturbations.
Subsequently, we investigated methane hydroxylation by 2,4,61 and 2,4,62 and the results are depicted in Fig. 4. Similar to methane hydroxylation by iron(IV)-oxo complexes [57,58,59,60,61,62,63], the reaction is stepwise with an initial hydrogen atom abstraction (via transition state TSHA) to form a radical intermediate (IHA). Thereafter, an OH rebound barrier (via transition state TSreb) gives alcohol product complexes (PHA). The free energies obtained with B3LYP and BP86 are very similar particularly for the transition states and also analogous structures are found. Therefore, the density functional method appears to have little effect on the structure and energies of the reaction mechanism. This contrast the spin-state ordering and relative energies of mononuclear iron and manganese-oxo complexes that often give strong variations depending on the density functional method chosen and particularly the amount of Hartree–Fock Exchange that is included in the method [45, 64, 65]. In all cases, the doublet spin state is well below the quartet and sextet spin state, and hence, the reaction takes place via single-state reactivity on the doublet spin-state surface and no spin crossing to another spin state is expected. Thus, the doublet spin hydrogen atom abstraction barrier is 7.8 (10.2) kcal mol−1 above isolated reactants as calculated with UB3LYP (UBP86), while the quartet spin barriers are at 30.0 (32.0) kcal mol−1 and the sextet spin ones at 57.7 (61.8) kcal mol−1. At room temperature, the quartet and sextet barriers will be inaccessible and the reaction will take place on a dominant doublet spin state only. Therefore, we focus on the doublet spin results from Fig. 4 in the following only.
Potential energy landscape of methane hydroxylation by 2,4,61 as obtained with DFT. Data obtained through full geometry optimization with UB3LYP [UBP86] level of theory. Free energies (at BS2 level of theory) are in kcal mol‒1 with solvent, thermal, entropic, and ZPE corrections included. Optimized geometries give bond lengths in angstroms and the imaginary frequency in cm‒1
Optimized geometries of the rate-determining doublet spin transition states (2TSHA) are given in Fig. 4. The transition states are late with long C‒H distances (1.375 and 1.471 Å at B3LYP and BP86 level of theory) and short O‒H distances (1.157 and 1.117 Å at B3LYP and BP86 level of theory). Late transition states often related to high energy barriers. Thus, for a series of hydrogen atom abstraction barriers by the same metal(IV)-oxo oxidant, it was shown that the barrier height correlated with the strength of the C‒H bond that was broken [58, 59, 66, 67]. It was found that reactions with substrates with strong C‒H bonds gave more product-like transition states, whereas with substrates with weak C‒H bonds, more reactant-like transition states were found. As methane has a strong C‒H-bond strength with bond dissociation energy (BDECH,methane = 101.6 kcal mol‒1 at UB3LYP level of theory), it is not surprising that the hydrogen atom abstraction barriers are high.
The rate-determining step in the reaction mechanism is hydrogen atom abstraction with a free energy of activation of 7.8 kcal mol−1, which is well lower in free energy here than that found for the analogous μ-nitrido-bridged diiron(IV)-oxo phthalocyanine complexes reported before [29], where a value of 15.7 kcal mol−1 was found. Therefore, the diruthenium complex is expected to react with hydrogen atom abstraction barriers that are almost 8 kcal mol−1 lower in free energy, which would correspond to a rate enhancement of over 106. Clearly, the diruthenium(IV)-oxo species is a considerably better oxidant that the corresponding diiron(IV)-oxo species. We will analyze the differences in structure and reactivity in detail in the following. Note that the rebound barrier is 7.2 (3.7) kcal mol−1 in energy above the radical intermediate 2IHA as calculated at UB3LYP (UPB86) level of theory. These barriers are considerable and may implicate a finite lifetime of the radical intermediates, which in the case of ethene activation by iron(IV)-oxo complexes was shown to lead to by-products [68, 69]. Furthermore, the radical could be released from the intermediate complex as dissipate into solution as suggested for nonheme iron reactivities [70].
Figure 5 gives the orbital energy changes during the methane hydroxylation reaction on the doublet spin state in a valence bond description. Thus, we describe electrons as a dot and a line bordered by two dots is a bonding orbital occupied by two electrons. These schemes were used previously to rationalize regioselectivities through analysing the electronic configurations of oxidants [34, 71,72,73]. As mentioned above in Fig. 3, the μ-nitrido-bridged diruthenium(IV)-oxo complex has electronic configuration of π* 23,x π* 13,y . Upon abstraction of a hydrogen atom from substrate, a σCH bond of methane is broken and splits into atomic orbitals: 2pC and 1sH. The hydrogen atom pairs up with one electron from the π*3,y molecular orbital to form the σO–H orbital with two electrons, while the πy set of orbitals splits into a new set of three orbitals (π′1,yπ′2,yπ*′3,y) that only spread over the Ru, N, and Ru atoms and contain four electrons. During the OH rebound process, also the π orbitals along the x-axis lose the oxygen contribution and split into a new set of orbitals π′1,xπ′2,xπ*′3,x with four electrons. One electron from the Ru‒O interaction pairs up with the radical on the CH3 group to form the new σO–C orbital, whereas the second one is promoted to a virtual π orbital on the porphyrazine group. As a consequence, the product has spin density on the ligand but not on the metals.
We also did a thermochemical analysis on the hydrogen atom and electron abstraction ability of the μ-nitrido-bridged diiron and diruthenium-oxo complexes, see Fig. 6. First, we calculated the bond dissociation energy of the O–H bond (BDEOH) in the MIV(OH) complex (M = Fe, Ru) as defined in Eq. (1), where we compare the energy of the MIV(OH) complex relative to that of the MV=O complex and a separate hydrogen atom. For the iron complex, a value of 86.7 kcal mol−1 was reported for the structure without axial ligand and 82.3 kcal mol−1 when an axial acetate was present [29]. Interestingly, using the same methods and techniques, a value of 134.9 kcal mol−1 is found for the RuIV(OH) system. Therefore, based on the relative BDE values, the μ-nitrido-bridged diruthenium-oxo complex is expected to be a considerably better oxidant than the corresponding iron complex and should react with methane even faster. The relative energies of the hydrogen atom abstraction transition states discussed above indeed confirm this:
Technically, a hydrogen atom abstraction is the sum of a proton transfer and an electron transfer; therefore, we split the BDEOH further into the sum of the acidity of the reduced oxidant (ΔGacid), the electron affinity (EA) of the starting complex, and the ionization energy of a hydrogen atom (IEH). The latter was taken from the literature [74]. Interestingly, the acidity of the iron and ruthenium-hydroxo complexes are alike and the differences in electron affinity compensates for stronger O–H-bond formation. As a particularly strong O–H bond is formed after hydrogen atom abstraction, this results in a large driving force for hydrogen atom abstraction and consequently low-energy hydrogen atom abstraction barriers.
Conclusions
Computational studies on a μ-nitrido-bridged diruthenium(IV)–oxo porphyrazine complex were performed and its reactivity with methane investigated. Our studies show that the complex is in a doublet spin ground state that is well separated from other spin states and with significant radical character on the oxo group. The electronic configuration of the μ-nitrido-bridged diruthenium(IV)-oxo complex is analogous to the corresponding diiron complex; however, it reacts with substrate with considerably lower barriers due to a more favorable hydrogen atom abstraction reaction. These differences are rationalized with thermochemical cycles and valence bond schemes.
Abbreviations
- DFT:
-
Density functional theory
- Cpd I:
-
Compound I
- BDE:
-
Bond dissociation energy
- EA:
-
Electron affinity
- IE:
-
Ionization energy
References
Sono M, Roach MP, Coulter ED, Dawson JH (1996) Chem Rev 96:2841–2888. https://doi.org/10.1021/cr9500500
Groves JT (2003) Proc Natl Acad Sci USA 100:3569–3574. https://doi.org/10.1073/pnas.0830019100
Ortiz de Montellano PR (2004) Cytochrome P450: structure, mechanism and biochemistry, 3rd edn. Kluwer Academic, New York
Meunier B, de Visser SP, Shaik S (2004) Chem Rev 104:3947–3980. https://doi.org/10.1021/cr020443g
Denisov IG, Makris TM, Sligar SG, Schlichting I (2005) Chem Rev 105:2253–2277. https://doi.org/10.1021/cr0307143
Kadish KM, Smith KM, Guilard R (2010) Handbook of porphyrin science. World Scientific Publishing Co., New Jersey
Ortiz de Montellano PR (2010) Chem Rev 110:932–948. https://doi.org/10.1021/cr9002193
Grogan G (2011) Curr Opin Chem Biol 15:241–248. https://doi.org/10.1016/j.cbpa.2010.11.014
Poulos TL (2014) Chem Rev 114:3919–3962. https://doi.org/10.1021/cr400415k
Huang X, Groves JT (2018) Chem Rev 118:2491–2553. https://doi.org/10.1021/acs.chemrev.7b00373
Rittle J, Green MT (2010) Science 330:933–937. https://doi.org/10.1126/science.1193478
Ogliaro F, Cohen S, de Visser SP, Shaik S (2000) J Am Chem Soc 122:12892–12893. https://doi.org/10.1021/ja005619f
Shaik S, Kumar D, de Visser SP, Altun A, Thiel W (2005) Chem Rev 105:2279–2328. https://doi.org/10.1021/cr030722j
Sorokin AB, Kudrik EV, Bouchu D (2008) Chem Commun. https://doi.org/10.1039/b804405h
Sorokin AB, Kudrik EV, Alvarez LX, Afanasiev P, Millet JMM, Bouchu D (2010) Catal Today 157:149–154. https://doi.org/10.1016/j.cattod.2010.02.007
Sorokin AB (2013) Chem Rev 113:8152–8191. https://doi.org/10.1021/cr4000072
Yamada Y, Morita K, Mihara N, Igawa K, Tomooka K, Tanaka K (2019) New J Chem 43:11477–11482. https://doi.org/10.1039/c9nj02210d
Mihara N, Yamada Y, Takaya H, Kitagawa Y, Igawa K, Tomooka K, Fujii H, Tanaka K (2019) Chem Eur J 25:3369–3375. https://doi.org/10.1002/chem.201805580
Afanasiev P, Sorokin AB (2016) Acc Chem Res 49:583–593. https://doi.org/10.1021/acs.accounts.5b00458
Kudrik EV, Afanasiev P, Alvarez LX, Dubourdeaux P, Clémancey M, Latour J-M, Blondin G, Bouchu D, Albrieux F, Nefedov SE, Sorokin AB (2012) Nat Chem 4:1024–1029. https://doi.org/10.1038/nchem.1471
Colomban C, Kudrik EV, Afanasiev P, Sorokin AB (2014) J Am Chem Soc 136:11321–11330. https://doi.org/10.1021/ja505437h
So S-C, Cheung W-M, Chiu W-H, de Vere-Tucker M, Sung HH-Y, Williams ID, Leung W-H (2019) Dalton Trans 48:8340–8349. https://doi.org/10.1039/c9dt00244h
Cheung W-M, Ng W-M, Wong W-H, Lee HK, Sung HH-Y, Williams ID, Leung W-H (2018) Inorg Chem 57:9215–9222. https://doi.org/10.1021/acs.inorgchem.8b01229
Cheung W-M, Chiu W-H, de Vere-Tucker M, Sung HH-Y, Williams ID, Leung W-H (2017) Inorg Chem 56:5680–5687. https://doi.org/10.1021/acs.inorgchem.7b00281
Stuzhin PA, Ivanova SS, Dereven’kov I, Makarov SV, Silaghi-Dumitrescu R, Homborg H (2012) Macroheterocycles 5:175–177. https://doi.org/10.6060/mhc2012.120360s
Colomban C, Kudrik EV, Sorokin AB (2017) J Porphyrins Phthalocyanines 21:345–353. https://doi.org/10.1142/S1088424617500274
İşci Ü, Dumoulin F, Sorokin AB, Ahsen V (2014) Turk J Chem 38:923–949. https://doi.org/10.3906/kim-1407-47
İşci Ü, Faponle AS, Afanasiev P, Albrieux F, Briois V, Ahsen V, Dumoulin F, Sorokin AB, de Visser SP (2015) Chem Sci 6:5063–5075. https://doi.org/10.1039/c5sc01811k
Quesne MG, Senthilnathan D, Singh D, Kumar D, Maldivi P, Sorokin AB, de Visser SP (2016) ACS Catal 6:2230–2243. https://doi.org/10.1021/acscatal.5b02720
Silaghi-Dumitrescu R, Makarov SV, Uta M-M, Dereven’kov IA, Stuzhin PA (2011) New J Chem 35:1140–1145. https://doi.org/10.1039/c0nj00827c
Ansari M, Vyas N, Ansari A, Rajaraman G (2015) Dalton Trans 44:15232–15243. https://doi.org/10.1039/C5DT01060H
Phung QM, Pierloot K (2019) Chem Eur J. https://doi.org/10.1002/chem.201902766 (in press)
Mayer JM (2004) Annu Rev Phys Chem 55:363–390
de Visser SP (2010) J Am Chem Soc 132:1087–1097. https://doi.org/10.1021/ja908340j
Kumar D, Sastry GN, de Visser SP (2011) Chem Eur J 17:6196–6205. https://doi.org/10.1002/chem.201003187
de Visser SP, Quesne MG, Martin B, Comba P, Ryde U (2014) Chem Commun 50:262–282. https://doi.org/10.1039/c3cc47148a
Kumar S, Faponle AS, Barman P, Vardhaman AK, Sastri CV, Kumar D, de Visser SP (2014) J Am Chem Soc 136:17102–17115. https://doi.org/10.1021/ja508403w
Frisch MJ (2013) Gaussian-09, revision C.02. Gaussian Inc, Wallingford CT
Becke AD (1993) J Chem Phys 98:5648–5652. https://doi.org/10.1063/1.464913
Lee C, Yang W, Parr RG (1988) Phys Rev B 37:785–789. https://doi.org/10.1103/PhysRevB.37.785
Becke AD (1988) Phys Rev A 38:3098–3100. https://doi.org/10.1103/PhysRevA.38.3098
Perdew JP (1986) Phys Rev B 33:8822–8824. https://doi.org/10.1103/PhysRevB.33.8822
Hay PJ, Wadt WR (1985) J Chem Phys 82:270–283. https://doi.org/10.1063/1.448799
Ditchfield R, Hehre WJ, Pople JA (1971) J Chem Phys 54:724–728. https://doi.org/10.1063/1.1674902
Cantú Reinhard FG, Faponle AS, de Visser SP (2016) J Phys Chem A 120:9805–9814. https://doi.org/10.1021/acs.jpca.6b09765
Cantú Reinhard FG, Barman P, Mukherjee G, Kumar J, Kumar D, Kumar D, Sastri CV, de Visser SP (2017) J Am Chem Soc 139:18328–18338. https://doi.org/10.1021/jacs.7b10033
Vardhaman AK, Sastri CV, Kumar D, de Visser SP (2011) Chem Commun 47:11044–11046. https://doi.org/10.1039/c1cc13775a
Cheaib K, Mubarak MQE, Sénéchal-David K, Herrero C, Guillot R, Clémancey M, Latour J-M, de Visser SP, Mahy J-P, Banse F, Avenier F (2019) Angew Chem Int Ed 58:854–858. https://doi.org/10.1002/anie.201812724
Barman P, Cantú Reinhard FG, Bagha UK, Kumar D, Sastri CV, de Visser SP (2019) Angew Chem Int Ed 58:10639–10643. https://doi.org/10.1002/anie.201905416
Colomban C, Tobing AH, Mukherjee G, Sastri CV, Sorokin AB, de Visser SP (2019) Chem Eur J. 25:1. https://doi.org/10.1002/chem.201902934
Hirao H, Kumar D, Que L Jr, Shaik S (2006) J Am Chem Soc 128:8590–8606. https://doi.org/10.1021/ja061609o
de Visser SP (2006) J Am Chem Soc 128:15809–15818. https://doi.org/10.1021/ja065365j
Sharma PK, de Visser SP, Ogliaro F, Shaik S (2003) J Am Chem Soc 125:2291–2300. https://doi.org/10.1021/ja0282487
Man W-L, Xie J, Pan Y, Lam WWY, Kwong H-K, Ip K-W, Yiu S-M, Lau K-C, Lau T-C (2013) J Am Chem Soc 135:5533–5536. https://doi.org/10.1021/ja401553d
Guo Z, Guan X, Huang J-S, Tsui W-M, Lin Z, Che C-M (2013) Chem Eur J 19:11320–11331. https://doi.org/10.1002/chem.201300021
Dhuri SN, Seo MS, Lee Y-M, Hirao H, Wang Y, Nam W, Shaik S (2008) Angew Chem Int Ed 47:3356–3359
Ogliaro F, Harris N, Cohen S, Filatov M, de Visser SP, Shaik S (2000) J Am Chem Soc 122:8977–8989. https://doi.org/10.1021/ja991878x
de Visser SP, Kumar D, Cohen S, Shacham R, Shaik S (2004) J Am Chem Soc 126:8362–8363. https://doi.org/10.1021/ja048528h
Shaik S, Kumar D, de Visser SP (2008) J Am Chem Soc 130:10128–10140. https://doi.org/10.1021/ja8019615
Singh D, Kumar D, de Visser SP (2015) Int J Sc Technol 1:26–40. https://doi.org/10.18091/ijsts.v1i2.9516
Yoshizawa K (2006) Acc Chem Res 39:375–382. https://doi.org/10.1021/ar050194t
Rosa A, Ricciardi G (2012) Inorg Chem 51:9833–9845. https://doi.org/10.1021/ic301232r
Tang H, Guan J, Zhang L, Liu H, Huang X (2012) Phys Chem Chem Phys 14:12863–12874. https://doi.org/10.1039/c2cp42423a
Yang T, Quesne MG, Neu HM, Cantú Reinhard FG, Goldberg DP, de Visser SP (2016) J Am Chem Soc 138:12375–12386. https://doi.org/10.1021/jacs.6b05027
Sainna MA, Sil D, Sahoo D, Martin B, Rath SP, Comba P, de Visser SP (2015) Inorg Chem 54:1919–1930. https://doi.org/10.1021/ic502803b
Latifi R, Bagherzadeh M, de Visser SP (2009) Chem Eur J 15:6651–6662. https://doi.org/10.1002/chem.200900211
Latifi R, Valentine JS, Nam W, de Visser SP (2012) Chem Commun 48:3491–3493. https://doi.org/10.1039/c2cc30365e
de Visser SP, Ogliaro F, Shaik S (2001) Angew Chem Int Ed 40:2871–2874
Timmins A, Quesne MG, Borowski T, de Visser SP (2018) ACS Catal 8:8685–8698. https://doi.org/10.1021/acscatal.8b01673
Cho K-B, Hirao H, Shaik S, Nam W (2016) Chem Soc Rev 45:1197–1210
Sainna MA, Kumar S, Kumar D, Fornarini S, Crestoni ME, de Visser SP (2015) Chem Sci 6:1516–1529. https://doi.org/10.1039/C4SC02717
Cantú Reinhard FG, Sainna MA, Upadhyay P, Balan GA, Kumar D, Fornarini S, Crestoni ME, de Visser SP (2016) Chem Eur J 22:18608–18619. https://doi.org/10.1002/chem.201604361
Kaczmarek MA, Malhotra A, Balan GA, Timmins A, de Visser SP (2018) Chem Eur J 24:5293–5302. https://doi.org/10.1002/chem.201704688
NIST Chemistry WebBook, NIST Standard Reference Database, Number 69; Linstrom PJ, Mallard WG (2016) National Institute of Standards and Technology: Gaithersburg, MD. http://webbook.nist.gov. Accessed 1 July 2019
Acknowledgements
MQEM thanks the Government of Malaysia for a studentship. The EU-COST Network for Bioinorganic Reaction Mechanisms (CM1003) is acknowledged for support. ABS is grateful to ANR, France for support (Grant ANR-16-CE29-0018-01).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Mubarak, M.Q.E., Sorokin, A.B. & de Visser, S.P. Properties and reactivity of μ-nitrido-bridged dimetal porphyrinoid complexes: how does ruthenium compare to iron?. J Biol Inorg Chem 24, 1127–1134 (2019). https://doi.org/10.1007/s00775-019-01725-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00775-019-01725-7