Abstract
Multiple antibiotic resistance regulator (MarR) family proteins are widely conserved transcription factors that control bacterial resistance to antibiotics, environmental stresses, as well as the regulation of virulence determinants. Escherichia coli MarR, the prototype member of this family, has recently been shown to undergo copper(II)-catalyzed inter-dimer disulfide bond formation via a unique cysteine residue (Cys80) residing in its DNA-binding domain. However, despite extensive structural characterization of the MarR family proteins, the structural mechanism for DNA binding of this copper(II)-sensing MarR factor remains elusive. Here, we report the crystal structures of DNA-bound forms of MarR, which revealed a unique, concerted generation of two new helix–loop–helix motifs that facilitated MarR’s DNA binding. Structural analysis and electrophoretic mobility shift assays (EMSA) show that the flexibility of Gly116 in the center of helix α5 and the extensive hydrogen-bonding interactions at the N-terminus of helix α1 together assist the reorientation of the wHTH domains and stabilize MarR’s DNA-bound conformation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The MarR family of transcription regulators modulates diverse bacterial detoxification responses to antibiotics, oxidative species, organic reagents as well as metal ions [1,2,3,4]. Despite high structural homology of the triangular-shaped MarR dimer proteins with winged helix–turn–helix (wHTH) DNA-binding motifs, members of this family of proteins share low sequence similarities (<25% on average) and the underlying derepression mechanism varies significantly [3, 5, 6]. A protein conformation unsuitable for DNA binding, including displacing or twisting of the DNA-binding motifs, could be enforced by binding of phenolic ligands near the protein–DNA interface (e.g., S. enterica SlyA) [7] or dimer interface (e.g., M. thermoautotrophicum MTH313) [8], or allosterically induced by covalent modification of cysteine residues, such as formation of intermolecular disulfide bonds (e.g., X. campestris OhrR, XcOhrR) [9], intramolecular disulfide bonds (e.g., M. tuberculosis MosR) [10], sulfenic acid (e.g., B. subtilis OhrR, BsOhrR) [11], or phosphorylation (e.g., S. aureus SarZ) [12]. Furthermore, many MarR members are inducible by various small molecule effectors, but how such structurally dissimilar compounds directly bind to the same protein and disrupt the protein–DNA interactions remains elusive [2,3,4].
Escherichia coli MarR is the prototype member of MarR family and regulates the marRAB operon which controls various cellular responses, including DNA repair, outer membrane permeability, and oxidative stress response [2]. Previous study indicates that diverse phenolic compounds, especially salicylic acid (SAL), act as negative regulator of MarR’s DNA binding, which induce the dissociation of MarR from its promoter DNA [13, 14]. However, due to the high concentration of SAL required for derepression of MarR, whether it is the common signaling molecule that induces MarR to regulate bacterial resistance to diverse, structurally unrelated antibiotics remains elusive [5, 13]. Our recent work showed that MarR may undergo a redox-based mechanism for derepression [15]. Disulfide bonds can be formed between MarR dimers by copper(II)-catalyzed oxidation on Cys80, a highly active cysteine residue located in its DNA-binding domain (Fig. 1a). Nevertheless, the structural mechanism for the binding of this MarR protein with its cognate DNA remains to be further illustrated. Herein, we present the structural and mechanistic study of their interactions.
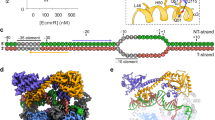
Structure of MarRC80S-DNA complex. a Sequence of E. coli MarR protein. Secondary structure elements of MarR with (MarR-DNA) and without DNA (Free MarR) are illustrated as tubes for α-helices and arrows for β-strands, respectively. Important residues involved in interactions with DNA are colored in cyan; Gly116 is colored in red; and Cys80 is colored in green. b Structure of MarRC80S-DNA complex. The secondary structure elements of one subunit are colored according to the scheme in a; DNA is colored in deep gray
Results
Structure of the MarR-DNA complex
Despite the previous structural characterization of several MarR proteins in the presence and absence of DNA [7,8,9,10,11,12, 16, 17], the structural investigation on DNA binding of E. coli MarR protein, the prototypical member of MarR family, is still limited. An SAL-bound MarR structure has been reported so far, which exhibited a conformation incapable of DNA binding [16]. In our recent work, structure of a reduced MarR mutant without ligand bound was solved [15]. Nevertheless, this structure also adopts a mode incapable of DNA binding. Taken together, MarR takes a conformation unfavored for DNA binding in the DNA-free state and the structural mechanism of interactions between MarR and its cognate DNA remains unknown. To elucidate this mechanism, we set out to solve the structure of DNA-bound MarR. We identified a condition to obtain crystals for WT-MarR-DNA complex at first; however, it was later found to be without DNA after processing the diffraction data, which is likely due to oxidization-triggered MarR-DNA dissociation. Moreover, the attempt to obtain the crystal using a highly reduced condition was failed, because the As(V) ion (in cacodylate) utilized in the crystallization buffer was not compatible with the reducing agent. To overcome this problem, we utilized MarRC80S mutant, which rendered MarR oxidization insensitive but did not impair its DNA binding [15], to generate crystal of the MarR-DNA complex. A 21 base-pair (bp) palindromic DNA sequence (20 bp with 1 bp sticky end) containing the binding site between –35 and –10 element of the previously reported marRAB promoter was chosen which can specifically bind to one equivalent of MarR dimer protein with an apparent affinity at 1 nM [13]. To our delight, the crystal of MarRC80S-DNA complex gave a much higher quality and was solved at 2.7 Å (Fig. 1b, Fig. S1, and Table 1). The asymmetric unit of the P61 crystal form contains one MarR dimer bound to one double-strand DNA molecule. Similar to MarR without DNA, the MarR dimer protein in the DNA-binding complex comprises six α-helices and two β-strands, with helices α1, α5, and α6 forming the dimerization domain, and helices α2, α3, α4, and the two β-strands forming the wHTH DNA-binding domain (Fig. 1a, b) [5, 7,8,9,10,11,12, 15,16,17]. In the wHTH domain, the helix–turn–helix portion consists of helices α3 and α4 with α4 being the DNA recognition helix, and the wing is constituted by the antiparallel β-sheet. The MarR-bound DNA is still a B-form double helix in general, but is globally bent upward towards the bottom of the MarR dimer by 8.9°, resulting in the shortening of the end-to-end distance of the 21-bp DNA by 2.4 Å. To accommodate the α4 and α4′ recognition helices, the major grooves are widened from 10.5 to 13.0 Å and deepened from 5.4 to 6.5 Å, whereas the minor grooves are narrowed near the ends of the binding site. In addition, interactions of the wing motif and the minor grooves cause local overtwisting of bases Thy3/Ade4 and Ade4/Cyt5 to 42.1° and 24.2°, respectively.
There are a total of 43 residues on MarR dimer that gives approximately 88 interactions with the 21 bp DNA (Fig. 2). The two α4 helices in the wHTH domains are inserted into the major grooves, while the two wing regions make close contacts with the minor grooves, including part of −10 regions (TATTAT) and its twofold related sequence on both ends of the DNA (Fig. 2). More specifically, MarR dimer makes a nearly identical set of contacts with the palindrome sequences TTGCC (Fig. 2) through insertion of N termini of the α4 recognition helices into the major grooves at the center of palindromic sequence. The major direct contacts with the DNA bases in this region are provided by Arg73 and Asp67: Arg73 forms bidentate hydrogen bonds between the side-chain NH groups and O6s of Gua14 and Gua8′, respectively, while Asp67 has direct base contact to N4 of Cyt15 (Fig. S2). In addition, Arg77 makes water-mediated base contacts to N7 of Gua13 and direct hydrogen bond to phosphate group of Gua12 (Fig. S2). Another important direct contact with the DNA bases was from β-wing. Arg94′ forms hydrogen bond between the guanidinium NH groups and O2 atom of Thy3. Arg94, dyadic mate of Arg94′, donates hydrogen bond to N3 atom of Ade3′, since the DNA is not fully symmetrical (Fig. S3). The sequence reads CATACT in the sense direction and GAATAT in the opposite strand. Additional interactions between the wing and the sugar-phosphate DNA backbone are hydrogen bonds between N atom of Val96 and phosphate of Ade5′ and between Arg86 and phosphate of Thy6′ (Fig. S3). The third DNA-binding element, a helix–helix motif (HH), is composed by helices α1 and α2 and their connecting residues that make contacts with the phosphate backbones of DNA. Residues that are involved in DNA binding within this region include Gln23, Thr39, and Gln42, which form hydrogen bonds with Gua14′, Gua12, and Gua12, respectively (Fig. S4). Analysis of van der Waals interactions is represented in Supporting Information (Supplementary notes and Fig. S5). Consistent with our structural result, the previous study indicated that mutations of Arg73, Arg77, and Arg94 affected the MarR–DNA interactions [18, 19]. In addition, since Cys80 is located at helix α4 that makes contacts with the major groove of DNA, we closely inspected the potential interactions of Ser80 with DNA, which may modulate the susceptibility of MarR to copper oxidation. Closed view of Ser80 showed that it is located at the C-terminus of helix α4, which stretched out towards the solution. This unique location, in conjunction with the short length of the Ser80 side chain (almost the same for cysteine), allowed negligible interaction of Ser80 with DNA (Fig. S6). The distance between the Oγ atom on Ser80 and the nearest phosphate group is 6.8 Å, which excluded direct interactions (Fig. S6). The lack of electron density of water molecules near Ser80 indicates that there are also no stable water-mediated interactions (Fig. S6). Thus, the mutation of Cys80 to Ser did not affect MarR’s DNA binding and the sensitivity of MarR to copper oxidation may not be directly modulated by an interaction between Cys80 and DNA.
Schematic representation of the interactions of MarR with its operator DNA in MarRC80S-DNA’s complex structure. Orange lines indicate water-mediated interactions between MarR protein and DNA; cyan lines indicate van der Waals contacts, and magenta lines indicate direct electrostatic interactions between MarR protein and DNA. Residues from helices α4s and the β-wings of MarR protein are marked in red and green, respectively. DNA bases of the palindrome sequence TTGCC and its complementary sequence are colored in cyan and that of the partial –10 region sequence TATT and its twofold-related sequence are colored in yellow
Overall, MarRC80S has a similar DNA-binding pattern as other MarR family members. With the overall triangular shape, each subunit of the MarR dimer locks onto the marRAB operator by an HTH motif, an extended wing, and the HH interaction region. R.m.s.d. of superpositions between DNA-bound MarRC80S dimer and the dimeric structures of other DNA-bound MarR family proteins BsOhrR [11], S. aureus MepR (SaMepR) [17], SlyA [7], MosR [10], and S. coelicolor SCO3205 [20] are 1.4, 1.2, 2.1, 2.9, and 1.3 Å, respectively. Major differences among these proteins are the length and direction of α1 and α6 helices, leading to variation in the dimerization domains. In contrast, structural alignments of the wHTH domain between MarR and BsOhrR, MepR, SlyA, MosR, or SCO3205 generate r.m.s.d. 1.1, 0.6, 0.7, 0.5, and 0.6 Å, respectively, indicating high similarity of this domain among these proteins (Fig. 3a). Noteworthy, the superposition between MarR and S. tokodaii ST1710 gave a much higher r.m.s.d. (5.5 Å for dimer protein), because it interacts with DNA through the wing motif that is different from other MarR family proteins [21]. In addition, sequence alignment among these aforementioned proteins showed that residues in α4 helix that make direct contacts with DNA bases have low conservation, since these residues are responsible for the specific protein–DNA recognition (Fig. S7A). For example, Asp67 and Arg73 from MarR were both utilized for specific binding to the palindromic sequence of TTGCC, but are not both conserved in other MarR family members. Although there are similar amino acids at the corresponding residues in MosR (Glu69 and Arg75) and SlyA (Glu59 and Arg65), neither of the acidic residues (Glu69 in MosR and Glu59 in SlyA) showed interactions with DNA bases [7, 10] (Fig. S7a). On the other hand, residues in the β-wing region, such as Arg86, Asp92, and Arg94, are highly conserved, facilitating MarR family proteins to bind A/T rich regions besides the palindromic signature sequence [17] (Fig. S7b). Residues Thr39 and Gln42 in HH motif are also conserved, consistent with their roles in facilitating DNA binding (Fig. S7c). Furthermore, as described above, alignment of promoter sequences for MarR family proteins showed a highly conserved A/T base pairs next to the palindromic regions, which corresponds to the highly conserved residues in the β-wing of MarR. The palindromic regions itself are less conserved due to the demand of specific interactions with the recognition α4 helices of MarR members (Fig. S7d). There seems no simple relationship between the sequences of recognition α4 helices and the corresponding promoters, since in addition to the primary sequence of α4 helices, the flexibility of amino acid side chain also affect the hydrogen bonding between α4 helices and their promoter DNA. For example, while MarR, SlyA, and MosR all utilize an Arg residue for hydrogen bonding with their promoter DNA, the proton acceptors are not the same: Arg73 forms bidentate hydrogen bonds with O6s of Gua14 and Gua8′ in MarR, Arg65 forms bidentate hydrogen bonds with O6 and N7 atoms of Gua14 in SlyA, and Arg75 forms bidentate hydrogen bonds with O6, N7 atoms of Gua12, and O6 atom of Thy13 in MosR. The multiple possibility of hydrogen-bonding formation between amino acids and DNA bases makes it difficult to analyze the mode of DNA sequence recognition by MarR family regulators. Taken together, with the exception of ST1710, the molecular mechanisms of MarR–DNA interactions are highly similar to other structurally characterized protein–DNA complexes within MarR family, and the structural differences among these proteins are mainly at the dimerization interface.
Structural comparison of MarRC80S-DNA complex with MarRC80S and other MarR family members. a Superimposition of the subunits of MarR family members (MarR, BsOhrR, SaMepR, SlyA, MosR, and SCO3205) from their respective DNA complex structures centered at the wHTH domain. MarR is colored in blue, while all other proteins are colored in yellow. The kink regions within MarR’s helices α5 and α1 are marked in red box. b Superposition of MarRC80S (orange) and MarRC80S-DNA(blue). Cα atoms within the dimerization domain (residues 126–144) are used for superposition, and the arrow indicates the translocation of the wing as measured between the Cα atoms of residue Asn91. c 90°-rotated view of b along the x-axis. The intersubunit distances between α4 and α4′ (top), and tip-to-tip (bottom) are marked above the arrows. The curved arrow indicates the rotation of α4 for MarR protein with and without DNA. d Helix-to-loop transition of MarR protein on helices α5 and α1 in the DNA-bound state. Comparison of MarR-DNA complex (blue) with MarRC80S (orange) shows two notable conformational changes on helices α5 and α1. Residues that generate the new helix in N-terminus are colored in red. The red arrow indicates the new helix formation in the N-terminus, the black arrow indicates the movement of wHTH domain, and the purple arrows indicate α5 breakage. e Close-up view of “kinked” helices α5 (top) and α1 (bottom) formation upon MarR-DNA binding. f Potential hydrogen bonding and electrostatic interactions (black dashed lines) among the newly formed helix and its nearby residues
Structural comparisons between MarR with and without DNA binding
We next inspected the difference between the structures of MarRC80S and MarRC80S-DNA complexes to investigate the mechanism of MarR’s DNA binding. Structural comparisons between MarRC80S and the MarRC80S-DNA complex revealed a large conformational change (r.m.s.d. = 4.896 Å for 263 Cα atoms) (Fig. 3b). Superposition of these two structures at the dimerization domain showed that the wHTH motif undergoes ~27° rotation and the wing’s tip has a ~14 Å translocation upon DNA binding. Such movements reoriented the originally neighboring α4 and α4′ helices and expanded their distance to ~29 Å, an optimal distance for binding to two consecutive major grooves on a B-form DNA (Fig. 3c). In addition, the tip-to-tip distance in MarR dimer was increased from ~55 to ~68 Å, which is also favored for interacting with minor grooves of B-form DNA (Fig. 3c). These results suggest that MarR undergoes a switching from the DNA-free, “closed” state to DNA-bound, and “open” state by reorientation of its wHTH motifs.
Two helix–loop–helix motifs are newly generated in concert with MarR’s DNA binding
DNA binding resulted in a significant conformational change on helix α5, the long linkage helix that connects MarR’s wHTH domain with its dimerization domain. Two shorter helices [α5a (residues 102–115) and α5b (residues 117–125)] appeared with the central residues (115–117) looped-out towards the solvent (Figs. 3a, d and S8a, b). From DNA-free MarRC80S, the i + 4 → i hydrogen bonds are formed in residue pairs Leu119/Val115 and His120/Gly116 on helix α5, indicating a typical α-helical conformation. By contrast, upon DNA binding, the α-helical structure in this region was disrupted due to the non-canonical i + 3 → i hydrogen bonding between residues Asp118 and Val115, as well as the missing i + 4 → i hydrogen bond between the backbone of Gly116 and His120 (Fig. 3e). Interestingly, these looping-out residues are centered at Gly116, which has been previously shown as a key residue in controlling MarR’s derepression activity [18, 22, 23]. For example, mutation of Gly116 on MarR significantly reduced the susceptibility of E. coli bacteria to tigecycline antibiotics or organic solvents [22, 23]. We speculated that the backbone flexibility of Gly116 might be the structural basis of conformational change upon MarR’s DNA binding. For further verification, we performed EMSA analysis on the MarRG116V mutant and the results indicated that Gly116-to-Val mutation effectively attenuated affinity of MarR with the 42-bp double-strand cognate DNA that has the same sequence of −41 to +1 region of marRAB promoter and contains MarR-binding site in the middle part [2], which proved the critical role of Gly116 in facilitating α5 breakage (Fig. 4a). Noteworthy, the helix-to-loop transition in helix α5 was also observed in both additional ROS-responsive members, such as XcOhrR, as well as non-ROS-responsive members, such as SaMepR and B. cereus MepR (BcMepR), within MarR family. However, the helix breakage occurred within the induced, non-DNA-binding conformation for all these proteins, instead of the “repressed”, DNA-binding conformation for MarR, indicating a different mode-of-action among these MarR members [9, 17, 24, 25] (Figs. 3a, d, e, S8a, b). Given the important role of helix α5 in connecting the dimerization domain and the DNA-binding domain of MarR family proteins, the splitting and reformation of helix α5 may be a general regulatory mechanism for a subgroup of MarR proteins.
Besides the conformational change in helix α5, a new single-turn α-helix was generated between residues Leu7 and Ile11 at MarR’s N-terminus upon DNA binding. These residues were within a flexible loop region in front of helix α1 in DNA-free MarRC80S, but were converted into a short helix in the presence of DNA (Figs. 1a, 3d–f, S8a, b). This new helix is perpendicular to helix α1, with the connecting loop bent towards the breakage region on helix α5. Interestingly, this single-turn α-helix was formed in concert with helix α5 breakage, which is not the case for XcOhrR, SaMepR, or BsOhrR proteins (Figs. 1a, 3a, S9) [9, 11, 17]. In XcOhrR, whereas two 310 helices are present in the reduced form with intact α5 helix, both of them disappeared in the oxidized form when helix α5 was broken [9]. In BsOhrR, the N-terminal residues form a 310 helix in both the DNA-binding and non-DNA-binding conformations, with little difference between these two states [11] (r.m.s.d. = 0.349 Å for Cα atoms of residues 8–16); while in SaMepR, fewer N-terminal residues were found in front of helix α1 without any secondary structures in the presence and absence of DNA [17]. Extensive hydrogen bonding and electrostatic interactions between side chains of residues, such as Arg16-Ser48, Glu10-His112′, and Lys44-His19′, were found in this looping-out region (Fig. 3f), which may stabilize the DNA-bound MarR conformation. This observation is also consistent with previous work showing that mutation of these hydrogen-bond-forming residues (e.g., Arg16, His19, Lys44, and Ser48) led to decreased DNA-binding ability or upregulation of MarR’s downstream genes [18, 19]. To further verify the functional role of this newly formed N-terminal helix in stabilizing the MarR-DNA complex, we created two mutations at the potential hydrogen-bond-forming residues on the newly formed N-terminal helix, respectively, and assessed their DNA-binding ability by EMSA. MarRN9A, which lacks the amide group for hydrogen-bonding formation on Asn9, and MarRE10R, which have opposite charge on Glu10, were selected. Whereas MarRN9A only exhibited slightly changed DNA affinity as that of WT-MarR, MarRE10R displayed considerable attenuation of DNA-binding ability (Fig. 4b). Therefore, it is conceivable that certain residues, such as Glu10 at the N-terminal helix, are important for stabilization of MarR’s DNA-binding conformation. Taken together, our results showed that the two newly generated helix–loop–helix motifs in the linkage region may play essential roles in facilitating the switching of MarR from the “closed” to “open” states.
We suggest that the induced fit mechanism might be preferred for conformational change of MarR upon DNA binding, because (1) different from the MarR homologue NadR [26], we did not obtain the crystal structure of MarR in a conformation suitable for DNA binding in the absence of DNA. (2) Crystal structure of MarRC80S has highly similar overall topology with that of MarR-SAL complex (r.m.s.d = 1.035 Å for 252 Cα atoms in dimer), both of which are unsuitable for DNA binding (distance between neighboring α4 and α4′ helices ~18 Å), indicating that binding of SAL has little effect on conformational change of MarR (Fig. S10). Together with the fact that only one asymmetric unit was contained in the unit cell of MarRC80S, the flexibility of MarR that allowed conformational change before DNA binding remains unknown. Nevertheless, these speculations may not be sufficient for confirming the induced fit mechanism. Interestingly, recent research showed that both induced fit and conformational selection could take part in protein–DNA interactions [27], which might also be similar for MarR. Indeed, further kinetic and thermodynamic investigations on MarR’s DNA binding are needed to fully clarify whether the induced fit or conformational selection is the main mechanism to drive MarR’s conformational change.
Conclusion
In summary, our study presented here reveals the structural mechanism for interactions of the E. coli MarR with its cognate DNA. Upon DNA binding, MarR coherently forms two new helix–loop–helix motifs, which may be essential for facilitating the reorientation of the wHTH domains and stabilizing MarR’s DNA-bound conformation. Furthermore, the coherent appearance of two helix–loop–helix motifs located in the center of helix α5 and at the N-terminus of helix α1 in DNA-bound state is a unique feature for E. coli MarR that is not observed from other proteins (Fig. 3a), revealing that MarR binds DNA via an unprecedented structural mechanism. Taken together, these DNA-free and DNA-bound MarR structures, in conjunction with the previously characterized copper(II)-oxidized MarR structure, allow us to propose the following induction model for MarR: (i) DNA-free MarR adopts the “closed”, non-DNA-binding conformation, but remains competitive for DNA binding; (ii) the presence of cognate DNA forces MarR to adopt the DNA-bound, “open” conformation, with helical breakage and bending in MarR’s linkage domain; and (iii) MarR can reversibly switch between these two states until copper(II)-mediated oxidation of its Cys80, which converted MarR into the “locked closed” state with two MarR dimers conformationally fixed by a pair of disulfide bonds (Fig. 5).
Proposed “two-state switching” model for MarR’s DNA binding and dissociation. In the absence of DNA, MarR adopts the ‘closed’, non-DNA-binding conformation but remains competitive for DNA binding. In the presence of DNA, MarR switches to the ‘open’ state in which two helical kinks are formed on α1 and α5 to stabilize the MarR–DNA interaction. MarR can reversibly switch between these two states. Copper(II)-induced oxidation drives MarR to a ‘locked closed’ state in which the two intermolecular disulfide bonds between MarR dimers fixed MarR dimer in the closed state. Gray protein scaffold; yellow spheres disulfide bonds between Cys80 residues
Materials and methods
Materials
All solutions were prepared with Milli-Q water. 4-(2-hydroxyethyl)-1-piperazineethanesulfonic acid (HEPES), 2-amino-2-hydroxymethyl-propane-1,3-diol (Tris), sodium chloride, potassium chloride, magnesium chloride, sodium cacodylate, and polyethylene glycol (PEG) with different average molecular weights were purchased from Sigma-Aldrich.
Protein expression and purification
MarR variants were expressed and purified as described previously [15], and were kept in a buffer with the following conditions: 20 mM Tris, pH 7.5, 150 mM NaCl.
Crystallization
For crystallization of MarRC80S-DNA, protein and DNA were premixed at a 1:2 ratio for 1 h at 20 °C. Screening was performed with conditions containing sodium cacodylate, NaCl, MgCl2, and different PEG. Crystals were grown by sitting-drop vapor diffusion at room temperature (10–20 °C). For every drop, 2 μl protein sample (4 mg/ml) was mixed with 2 μl well solution. MarRC80S-DNA crystals appeared in 0.1 M sodium cacodylate pH 5.6–6.2, 0.1 M NaCl, 50 mM MgCl2, and 16–18% PEG 5000 MME.
X-ray data collection, structure determination, and analysis
Cryoprotectant was made by mixing well solution with 90% glycerol at a ratio of 7:2. Stepwise soaking was applied to avoid crystal cracking. The crystals were then flash frozen in liquid nitrogen before data collection. Data sets of MarRC80S-DNA were collected on the BL17U1 beamline at the Shanghai Synchrotron Radiation Facility (Shanghai, China) with wavelength of 1.0055 Å [28]. Data reduction was processed by the CCP4 package [29, 30] and HKL2000 [31], respectively. The structures were solved via molecular replacement using the SlyA protein in complex with DNA [7] (Protein Data Bank code 3Q5F) as the search model with the program Phaser [32]. Refinement was performed with the programs REFMAC5 [33], PHENIX [34], and Coot [35]. PyMOL (Schrödinger, LLC, www.pymol.org) was used to generate all the figures. A summary of the data statistics is shown in Table 1. The analysis of DNA conformation was done by Curves+ [36]. Identification of hydrogen bonds was done by COCOMAPS Server [37], and all atoms within 4 Å of each other are considered to have van der Waals contacts. The atomic coordinates and structure factors have been deposited in the Protein Data Bank as entry 5H3R for MarRC80S-DNA complex.
EMSA
EMSA was performed as described previously [16]. Briefly, different concentrations of MarR proteins (e.g., WT-MarR, MarRG116V) were added to a final volume of 20 μl containing 50 nM annealed double-stranded DNA in binding buffer (20 mM HEPES pH 7.0, 50 mM KCl, 5 mM MgCl2 and 10% glycerol). Sequence of oligonucleotides used for the annealing process was 5′-ATTCATTTGACTTATACTTGCCTGGGCAATATTATCCCCTGC-3′ and its complementary sequence.
References
Cohen SP, Levy SB, Foulds J, Rosner JL (1993) J Bacteriol 175:7856–7862
Alekshun MN, Levy SB (1997) Antimicrob Agents Chemother 41:2067–2075
Wilkinson SP, Grove A (2006) Curr Issues Mol Biol 8:51
Reyes-Caballero H, Guerra AJ, Jacobsen FE, Kazmierczak KM, Cowart D, Koppolu UMK, Scott RA, Winkler ME, Giedroc DP (2010) J Mol Biol 403:197–216
Perera IC, Grove A (2010) J Mol Cell Biol 2:243–254
Kumarevel T (2012) In: Pana M (ed) Antibiotic resistant bacteria—a continuous challenge in the new millennium. INTECH Open Access Publisher, Rijeka, pp 403–418
Dolan KT, Duguid EM, He C (2011) J Biol Chem 286:22178–22185
Saridakis V, Shahinas D, Xu X, Christendat D (2008) J Mol Biol 377:655–667
Newberry KJ, Fuangthong M, Panmanee W, Mongkolsuk S, Brennan RG (2007) Mol Cell 28:652–664
Brugarolas P, Movahedzadeh F, Wang Y, Zhang N, Bartek IL, Gao YN, Voskuil MI, Franzblau SG, He C (2012) J Biol Chem 287:37703–37712
Hong M, Fuangthong M, Helmann JD, Brennan RG (2005) Mol Cell 20:131–141
Sun F, Ding Y, Ji Q, Liang Z, Deng X, Wong CCL, Yi C, Zhang L, Xie S, Alvarez S, Hicks LM, Luo C, Jiang H, Lan L, He C (2012) Proc Natl Acad Sci USA 109:15461–15466
Martin RG, Rosner JL (1995) Proc Natl Acad Sci USA 92:5456–5460
Alekshun MN, Levy SB (1999) J Bacteriol 181:4669–4672
Hao Z, Lou H, Zhu R, Zhu J, Zhang D, Zhao BS, Zeng S, Chen X, Chan J, He C, Chen PR (2014) Nat Chem Biol 10:21–28
Alekshun MN, Levy SB, Mealy TR, Seaton BA, Head JF (2001) Nat Struct Biol 8:710–714
Birukou I, Seo SM, Schindler BD, Kaatz GW, Brennan RG (2014) Nucleic Acids Res 42:2774–2788
Alekshun MN, Kim YS, Levy SB (2000) Mol Microbiol 35:1394–1404
Duval V, McMurry LM, Foster K, Head JF, Levy SB (2013) J Bacteriol 195:3341–3351
Stevenson CE, Assaad A, Chandra G, Le TB, Greive SJ, Bibb MJ, Lawson DM (2013) Nucleic Acids Res 41:7009–7022
Kumarevel T, Tanaka T, Umehara T, Yokoyama S (2009) Nucleic Acids Res 37:4723–4735
Watanabe R, Doukyu N (2012) AMB Express 2:1–11
Linkevicius M, Sandegren L, Andersson DI (2013) J Antimicrob Chemother 68:2809–2819
Birukou I, Tonthat NK, Seo SM, Schindler BD, Kaatz GW, Brennan RG (2013) mBio 4:e00528–e00613
Kim MI, Cho MU, Hong M (2015) Biochem Biophys Res Commun 458:644–649
Liguori A, Malito E, Lo Surdo P, Fagnocchi L, Cantini F, Haag AF, Brier S, Pizza M, Delany I, Bottomley MJ (2016) PLoS Pathog 12:e1005557
Chu X, Liu F, Maxwell BA, Wang Y, Suo Z, Wang H, Han W, Wang J (2014) PLoS Comput Biol 10:e1003804
Wang QS, Yu F, Huang S, Sun B, Zhang KH, Liu K, Wang ZJ, Xu CY, Wang SS, Yang LF, Pan QY, Li L, Zhou H, Cui Y, Xu Q, Earnest T, He JH (2015) Nucl Sci Tech 26:12–17
Potterton E, Briggs P, Turkenburg M, Dodson E (2003) Acta Crystallogr Sect D Biol Crystallogr 59:1131–1137
Winn MD, Ballard CC, Cowtan KD, Dodson EJ, Emsley P, Evans PR, Keegan RM, Krissinel EB, Leslie AGW, McCoy A, McNicholas SJ, Murshudov GN, Pannu NS, Potterton EA, Powell HR, Read RJ, Vagin A, Wilson KS (2011) Acta Crystallogr D 67:235–242
Otwinowski Z, Minor W (1997) In: Carter CW Jr, Sweet RM (eds) Methods in enzymology, Volume 276: Macromolecular Crystallography, part A. Academic Press, New York, pp 307–326
McCoy AJ, Grosse-Kunstleve RW, Adams PD, Winn MD, Storoni LC, Read RJ (2007) J Appl Crystallogr 40:658–674
Murshudov GN, Vagin AA, Dodson EJ (1997) Acta Crystallogr D Biol Crystallogr 53:240–255
Adams PD, Afonine PV, Bunkoczi G, Chen VB, Davis IW, Echols N, Headd JJ, Hung L-W, Kapral GJ, Grosse-Kunstleve RW, McCoy AJ, Moriarty NW, Oeffner R, Read RJ, Richardson DC, Richardson JS, Terwilliger TC, Zwart PH (2010) Acta Crystallogr Sect D Biol Crystallogr 66:213–221
Emsley P, Lohkamp B, Scott WG, Cowtan K (2010) Acta Crystallogr Sect D Biol Crystallogr 66:486–501
Lavery R, Moakher M, Maddocks JH, Petkeviciute D, Zakrzewska K (2009) Nucleic Acids Res 37:5917–5929
Vangone A, Spinelli R, Scarano V, Cavallo L, Oliva R (2011) Bioinformatics 27:2915–2916
Acknowledgments
We thank all staff members of Shanghai Synchrotron Radiation Facility (SSRF). This work was supported by research grants from the National Natural Science Foundation of China (21225206, 21432002, and 21521003) and National Key Research and Development Program (2016YFA0501500).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Zhu, R., Hao, Z., Lou, H. et al. Structural characterization of the DNA-binding mechanism underlying the copper(II)-sensing MarR transcriptional regulator. J Biol Inorg Chem 22, 685–693 (2017). https://doi.org/10.1007/s00775-017-1442-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00775-017-1442-7