Abstract
Introduction
Overexpression studies have been commonly used to yield significant advances in cell biology. In vitro osteoclast culturing involves the differentiation of bone marrow-derived monocyte macrophage precursors (BMMs) in medium supplemented with macrophage colony-stimulating factor and receptor activator of nuclear factor-kB ligand (RANKL) into mature osteoclasts. Retroviral vectors are the gold standards for efficient gene delivery into BMMs. While this strategy is effective in BMMs that are in the early stages of differentiation, it is ineffective in RANKL-treated BMMs such as mono- and multinucleated osteoclasts. This study attempted to enhance gene delivery into differentiated BMMs using liposome-mediated RNA transfection.
Material and methods
BMMs were transfected with an EYFP overexpression plasmid or EYFP RNA by lipofection, or transduced with a retroviral vector expressing EYFP. EYFP expression was assessed by flow cytometry.
Results
We performed overexpression analyses using enhanced yellow fluorescent protein (EYFP). Although EYFP expression was observed 24 h after infection of BMMs with a recombinant retrovirus containing EYFP, expression of EYFP was observed within 3 h of transfection with EYFP RNA. Moreover, the efficiency of EYFP RNA for gene delivery into BMMs was comparable to that of retroviral transduction of EYFP. In contrast, while very few BMMs stimulated by RANKL for two days expressed EYFP after retroviral infection, more than half of the cells expressed EYFP after transfection with EYFP RNA.
Conclusion
RNA-mediated gene delivery is quick and easy method for performing gain-of-function analyses in primary osteoclast precursors and mature osteoclasts.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Bone is a type of connective tissue comprising organic proteins, such as collagen, and inorganic mineral hydroxyapatite. It undergoes constant remodeling by osteoclasts and osteoblasts during our adult life [1,2,3]. Osteoclasts are multinucleated cells of the monocyte/macrophage lineage that degrade bone matrix. Osteoclastogenesis is tightly regulated by the binding of RANKL and macrophage colony-stimulating factor (M-CSF) to their respective receptors, RANK and c-Fms, on osteoclast precursors. RANKL is a cytokine that is crucial for cell cycle progression and osteoclast differentiation [4,5,6]; while, M-CSF is important for the survival and motility of osteoclast lineage cells and expression of RANK [7,8,9]. Signaling involving RANK and c-Fms has been extensively studied using gain- and loss-of-function approaches in cultured osteoclasts. Gain-of-function experiments have been used to identify modulators of RANK and c-Fms signaling [10,11,12,13], genes that promote survival [14,15,16,17], and transcriptional programs involved in osteoclast maturation [18,19,20,21].
Retrovirus-mediated gene transduction in osteoclasts cultured in vitro is the gold standard for gain-of-function analyses. However, as retroviruses are incapable of infecting non-dividing cells, this strategy is only effective in the early differentiation stages of proliferating osteoclasts, such as bone marrow-derived monocyte macrophage precursors (BMMs). Therefore, retrovirus-mediated gene transduction is not recommended for cells in the late stage of differentiation such as mono- and multinucleated osteoclasts. Alternatively, adenovirus-mediated gene delivery is a useful approach for introducing genes into RANKL-stimulated BMMs. However, the process of generating recombinant adenoviruses, including construction of an adenovirus plasmid containing the gene of interest, recombination of the plasmid with the adenoviral genome, and packaging and purification of recombinant adenovirus at high titers, is laborious and time consuming. Furthermore, the size of the transgene is substantially limited due to the small packaging capacity of the viral particle.
This study aims to tackle this problem and provides a quick and easy method for gain-of-function analyses in primary osteoclast precursors and mature osteoclasts using RNA-mediated gene introduction. Our findings are predicted to substantially help future research in bone cell biology based on gain-of-function analyses.
Materials and methods
Cell culture
In vitro osteoclast differentiation was described previously [20, 22]. Briefly, for in vitro differentiation, bone marrow-derived cells from C57BL6/J mice were cultured with 10 ng/mL M-CSF (Miltenyi Biotec) for two days and used as osteoclast precursor cells and BMMs. Subsequently, cells were further cultured with 50 ng/mL RANKL (Peprotech) in the presence of 10 ng/mL M-CSF for three days. Animal study was approved by the Institutional Animal Care and Use Committee of both Doshisha University and Osaka University.
Gene transfer of EYFP
pcDNA3-EYFP and its derivatives were constructed by inserting DNA fragments encoding EYFP, 4-amino acid sequence (Pro-Glu-Ser-Thr, PEST) [23], a destabilizing domain derived from FKBP12 protein [24], and DNA fragments of adenylate-uridylate (AU)-rich elements (ARE) [25] into pcDNA3 vector. For RNA transfection experiment, synthetic capped RNA was made with the mMESSAGE mMACHINE T7 ULTRA Transcription kit (Thermo) using linearized DNA of the pcDNA3-EYFP derivatives and then purified by RNeasy Mini kit (Qiagen). RNA transfections were performed with the Lipofectamine MessengerMAX (Thermo) according to the manufacturer’s instructions. Three hours after transfection with RNA, BMMs were stimulated with 50 ng/mL RANKL. For the analysis of EYFP derivatives containing AU-rich elements and PEST sequences, BMMs were transfected with the EYFP-derivative RNA by lipofection for 3 h and then cultured with 50 ng/mL RANKL in the presence of 10 ng/mL M-CSF. Three and 48 h after RANKL stimulation, flow cytometry analysis was performed. For the analysis of the EYFP derivative containing the DD domain, BMMs were transfected with EYFP-derivative RNA by lipofection for 3 h and cultured with RANKL and M-CSF in the presence of 0.5 μM Shield-1 for one day. Then, cells were further cultured in the presence and absence of Shield-1 for one day and subjected to flow cytometry analysis.
For retroviral gene transfer experiment, the retroviral vector pMX-EYFP was constructed by inserting DNA fragments encoding EYFP into the pMX vector. Retroviral packaging was performed by transfecting the plasmids into Plat-E cells using FuGENE6 as described previously [26]. Ten hours after transduction with retroviruses, BMMs were stimulated with RANKL.
For DNA transfection experiment, pcDNA3-EYFP derivatives were transfected using FuGENE HD (Promega) according to the manufacturer’s instructions. Twenty-four hours after transfection with DNA vectors, BMMs were stimulated with 50 ng/mL RANKL.
Flow cytometry analysis
The analysis was described previously [21]. Briefly, single-cell suspensions were incubated with anti-CD16/CD32 for ten minutes, and then stained with Phycoerythrin-conjugated anti-CD11b (M1/70; eBioscience) in flow cytometry buffer (1 × phosphate-buffered saline [PBS], 4% heat-inactivated fetal calf serum, and 2 mM EDTA) for fifteen minutes. Stained cells were analyzed on a FACSCanto II Flow Cytometer (BD Biosciences). FACS data were statistically analyzed with FlowJo software (TreeStar Inc.).
Quantitative RT-PCR analysis
Total RNA and cDNA were prepared using the RNeasy Mini Kit (Qiagen) and Superscript III reverse transcriptase (Invitrogen), respectively, according to the manufacturers’ instructions. Real-time PCR was performed with a Thermal Cycler Dice Real Time System (TaKaRa Bio) using SYBR Premix EX Taq (TaKaRa Bio). The primer sequences were described previously [21].
Statistical analysis
All data are expressed as mean ± standard error of the mean (SEM). Statistical analysis was performed using the unpaired two-tailed Student’s t test for comparisons between two groups (*P < 0.05; **P < 0.01; ***P < 0.001; NS, not significant).
Results
To compare the efficacy of liposome and retrovirus-mediated gene delivery into osteoclast precursors, we overexpressed EYFP as our test gene of interest. When bone marrow cells are cultured with M-CSF for two days to generate BMMs that are stimulated by RANKL (with M-CSF) for three days, cells differentiate into multinucleated osteoclast-like cells (OCLs) positive for the osteoclast marker tartrate-resistant acid phosphatase capable of bone resorption [21]. Prior to treatment with RANKL, BMMs were transfected with EYFP-overexpressing plasmid or capped EYFP RNA by lipofection, or transduced with a retroviral vector expressing EYFP (Fig. 1A). Both EYFP RNA transfection and retroviral transduction yielded a higher efficiency of EYFP expression as compared to plasmid transfection (Fig. 1b). We detected fluorescence in EYFP RNA-transfected BMMs at 3 h and 8 h following lipofection. However, EYFP expression was barely detectable in retrovirus-transduced or plasmid-transfected BMMs. One day after gene delivery, the EYFP fluorescence was observed in BMMs transfected with EYFP RNA and transduced with the retroviral vector. Gene expression profiling revealed that EYFP RNA reached a maximum level at three hours post-transfection of EYFP RNA and was barely detectable by twenty-four hours post-transfection (Fig. 2a). In contrast, EYFP mRNA levels increased in retrovirus-transduced cells 24 h after infection. Two days after RANKL stimulation, flow cytometry showed that 69.5 ± 1.7% and 62.4 ± 3.8% of CD11b-positive cells transfected with EYFP RNA and retrovirus-transduced cells were positive for EYFP protein expression, respectively (Fig. 2b).
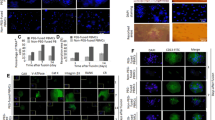
Gene delivery of the exogenous protein, EYFP, into osteoclast precursors. a Schematic of the protocol for gene delivery into bone marrow-derived monocyte macrophage precursors (BMMs). For retroviral transduction, BMMs were transduced with a retroviral vector expressing EYFP for 10 h. For lipofection, BMMs were incubated with lipid–RNA or –DNA complexes for 3 h or 24 h, respectively. b Representative fluorescence images of EYFP in BMMs transfected with the EYFP-overexpressing plasmid or EYFP RNA by lipofection, or transduced with a retroviral vector expressing EYFP. Scale bar 100 μm
Expression of EYFP in osteoclast precursors and mature osteoclasts. a Gene expression profiles of EYFP in BMMs transfected with EYFP. BMMs were transfected with the EYFP RNA by lipofection for 3 h or transduced with a retroviral vector expressing EYFP for 10 h, and then cultured with RANKL in the presence of M-CSF. Gene expression was analyzed at each time point after RANKL stimulation. Percentage of EYFP-expressing cells (gated using CD11b+). BMMs were transfected with an EYFP overexpression plasmid for 24 h or EYFP RNA for 3 h by lipofection, or transduced with a retroviral vector expressing EYFP for 10 h, and then cultured with RANKL in the presence of M-CSF. Two days after RANKL stimulation, flow cytometry analysis was performed. c Representative fluorescence images of EYFP in BMMs subjected to gene introduction. BMMs were transfected with EYFP RNA for 3 h by lipofection, or transduced with a retroviral vector expressing EYFP for 10 h, and then cultured with RANKL in the presence of M-CSF for three days. Scale bar 100 μm
When BMMs transfected with EYFP RNA or transduced with retroviral vector were further treated with M-CSF and RANKL, the fluorescent signal of EYFP was observed in multinucleated cells three days after gene delivery (Fig. 2c). However, the fluorescence intensity of EYFP was reduced in BMMs transfected with EYFP RNA, consistent the negligible detection of exogenous EYFP RNA (Fig. 2a). Together, these results indicate that the efficiency of gene delivery by EYFP RNA lipofection and retroviral transduction of EYFP was comparable in BMMs. Transfection of RNA is a feasible approach to overexpress exogenous proteins through different stages of osteoclast differentiation (very early to late stage of osteoclasts).
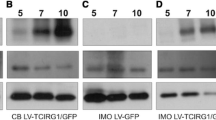
Next, we compared the efficacy of gene delivery by liposome transfection and virus transduction in fully differentiated osteoclasts. Osteoclast lineage-committed cells were induced by treating BMMs with M-CSF and RANKL. In vitro osteoclast differentiation results in the formation of multinucleated OCLs that exhibit characteristics of mature osteoclasts three days after RANKL stimulation [20]. We found that transfection with the EYFP overexpression plasmid or capped EYFP RNA by lipofection, or transduction with a retroviral vector expressing EYFP, of BMMs prior to and one day following RANKL-stimulated produced similar results (data not shown). Fluorescence from EYFP was observed within 3 h of transfection with EYFP RNA but not with the EYFP overexpression plasmid. In contrast, fluorescence from EYFP was observed 24 h after incubation with the EYFP-containing retrovirus. There was a significant difference in the level of fluorescence in liposome-transfected or retrovirus-transduced BMMs stimulated with RANKL for two days (Fig. 3a). Very few cells expressed EYFP 24 h after infection with the recombinant retrovirus (Fig. 3b), while approximately 50% of cells expressed EYFP after transfection with EYFP RNA (Fig. 4). Interestingly, fluorescence from EYFP was detected in multinucleated OCLs as well as mononucleated cells (Fig. 3b). These results suggest that RNA-based gene delivery is effective in osteoclast precursors and mature osteoclasts.
Gene delivery of EYFP in fully differentiated osteoclasts. a Schematic of the protocol for gene delivery into BMMs two days after treatment with RANKL. For retroviral transduction, RANKL-treated BMMs were transduced with the retroviral vector expressing EYFP for 10 h. For lipofection, RANKL-treated BMMs were incubated with lipid–RNA or –DNA complexes for 3 h or 24 h, respectively. b Representative images for the fluorescence in fully differentiated osteoclasts transfected with the EYFP overexpression plasmid or EYFP RNA by lipofection, or transduced with the retroviral vector expressing EYFP. Scale bar 100 μm
Expression of EYFP in mature osteoclasts. RANKL-treated BMMs for two days were transfected with an EYFP overexpression plasmid for 24 h or EYFP RNA for 3 h by lipofection, or transduced with a retroviral vector expressing EYFP for 10 h, and further cultured with RANKL in the presence of M-CSF. One day after gene introduction, the percentage of EYFP-expressing cells (gated using CD11+) was analyzed by flow cytometry
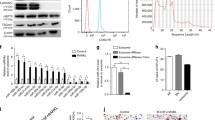
Based on these findings, we aimed to develop a method to transiently overexpress exogenous proteins selectively during the early stage of osteoclast differentiation. We used the following approach: destabilization of exogenous protein at the RNA and protein level during late stage osteoclast differentiation. AREs are a major cis-acting binding element in the untranslated region of labile RNAs [25]. The PEST tag comprises a peptide sequence that is rich in proline, glutamic acid, serine, and threonine, and reduces the half-life of PEST-tagged proteins [23]. BMMs transfected with EYFP RNA harboring either AREs or PEST tags were treated with M-CSF and RANKL showed no reduction in EYFP fluorescence throughout osteoclast differentiation (data not shown). Fluorescence from EYFP containing both the AREs and PEST tags decreased to a slight yet significant extent during the late stages of osteoclast differentiation (Fig. 5a). Next, we used the Shield-1 ligand to rapidly increase the stability of EYFP [24]. This technique involves the fusion of the protein of interest to a destabilizing domain (DD) derived from FKBP12 protein. Addition of the membrane permeable small molecule, Shield-1, that binds to the DD shields the protein from degradation. Removal of Shield-1 leads to rapid degradation of the DD domain along with any fused protein. BMMs were transfected with EYFP RNA carrying the DD domain and treated with M-CSF and RANKL in the presence of Shield-1. One day post-transfection, although RANKL-treated BMMs exhibited fluorescence from EYFP, removal of Shield-1 completely and rapidly reduced the fluorescence from EYFP (Fig. 5b). Taken together, our method of RNA-mediated gene delivery overexpresses exogenous proteins based on the various stages differentiation in primary osteoclasts.
Expression of EYFP derivatives in osteoclast precursors. a Percentage of EYFP-expressing cells (gated using CD11 +) after transfection with EYFP derivative containing AU-rich elements and PEST sequences. BMMs were transfected with the EYFP derivatives RNA by lipofection for 3 h and then cultured with RANKL in the presence of M-CSF. Three and 48 h after RANKL stimulation, flow cytometry analysis was performed. b Percentage of EYFP-expressing cells (gated using CD11s+) after transfection with EYFP derivative containing DD domain. BMMs were transfected with the EYFP derivatives RNA by lipofection for 3 h and cultured with RANKL and M-CSF in the presence of Shield-1 (0.5 μM) for one day. Then, cells were further cultured in the presence and absence of Shield-1 for one day and were subjected to flow cytometry analysis
Discussion
To the best of our knowledge, this is the first report on a quick and easy method for overexpressing exogenous proteins in primary osteoclast precursors and mature osteoclasts. Overexpression experiments constitute a qualitative method to identify novel proteins and investigate their function. Overexpression of NFATc1 induces osteoclastogenesis even in the absence of RANKL, suggesting that NFATc1 is a master regulator of osteoclast differentiation [18]. Thus, overexpression experiments have become an indispensable technique in molecular biology research.
In primary cell cultures, including the in vitro osteoclast differentiation assay, virus-mediated gene transduction is the gold standard for overexpression experiments; however, this technique is associated with drawbacks, including limited size of the gene of interest and physiological stage of cells undergoing gene transduction. RNA-mediated gene delivery helps to overcome these drawbacks and provides a powerful approach to study osteoclastogenesis. Despite the efficacy of gene delivery by RNA-mediated method, however, adenoviral transduction may still be advantageous in some studies as it has been used extensively for gain-of-function analyses and allows for efficient gene delivery into both osteoclast precursors and mature osteoclasts [27,28,29].
Bacterial CRISPR–Cas9 has been commonly used for genome editing and has emerged as a multifunctional platform for sequence-specific regulation of gene activation and repression, modification of CpG methylation, and visualization of genomic loci [30, 31]. However, since Cas9 is large protein, the virus has a low efficiency of encapsulating Cas9. Moreover, purifying recombinant viruses in high titer is laborious and time consuming. The major advantages our method of RNA-mediated gene delivery include speed and simplicity (regardless of size limitation). Therefore, combining CRISPR with our technique can be applied to cultures of primary osteoclasts.
We used techniques in destabilizing RNA and protein of exogenous genes to successfully overexpress exogenous proteins specifically in differentiating primary osteoclasts. Tools in optogenetics have been developed to enable the optical enhancement of protein stability [32] or degradation [33]. Therefore, protein levels can be rapidly fine-tuned by utilizing optogenetic tools.
References
Takayanagi H (2007) Osteoimmunology: shared mechanisms and crosstalk between the immune and bone systems. Nat Rev Immunol 7:292–304
Lorenzo J, Horowitz M, Choi Y (2008) Osteoimmunology: interactions of the bone and immune system. Endocr Rev 29:403–440
Okamoto K, Takayanagi H (2019) Osteoimmunology. Cold Spring Harbor Perspect Med 9:a03145
Theill LE, Boyle WJ, Penninger JM (2002) RANK-L and RANK: T cells, bone loss, and mammalian evolution. Annu Rev Immunol 20:795–823
Xing L, Schwarz EM, Boyce BF (2005) Osteoclast precursors, RANKL/RANK, and immunology. Immunol Rev 208:19–29
Ross FP, Teitelbaum SL (2005) alphavbeta3 and macrophage colony-stimulating factor: partners in osteoclast biology. Immunol Rev 208:88–105
Jimi E, Shuto T, Koga T (1995) Macrophage colony-stimulating factor and interleukin-1 alpha maintain the survival of osteoclast-like cells. Endocrinology 136:808–811
Otero K, Turnbull IR, Poliani PL, Vermi W, Cerutti E, Aoshi T, Tassi I, Takai T, Stanley SL, Miller M, Shaw AS, Colonna M (2009) Macrophage colony-stimulating factor induces the proliferation and survival of macrophages via a pathway involving DAP12 and beta-catenin. Nat Immunol 10:734–743
Novack DV, Faccio R (2011) Osteoclast motility: putting the brakes on bone resorption. Ageing Res Rev 10:54–61
Miyazaki T, Sanjay A, Neff L, Tanaka S, Horne WC, Baron R (2004) Src kinase activity is essential for osteoclast function. J Biol Chem 279:17660–17666
Ikeda F, Nishimura R, Matsubara T, Tanaka S, Inoue J, Reddy SV, Hata K, Yamashita K, Hiraga T, Watanabe T, Kukita T, Yoshioka K, Rao A, Yoneda T (2004) Critical roles of c-Jun signaling in regulation of NFAT family and RANKL-regulated osteoclast differentiation. J Clin Investig 114:475–484
Gohda J, Akiyama T, Koga T, Takayanagi H, Tanaka S, Inoue J (2005) RANK-mediated amplification of TRAF6 signaling leads to NFATc1 induction during osteoclastogenesis. EMBO J 24:790–799
Kim JH, Kim K, Jin HM, Song I, Youn BU, Lee SH, Choi Y, Kim N (2010) Negative feedback control of osteoclast formation through ubiquitin-mediated down-regulation of NFATc1. J Biol Chem 285:5224–5231
Akiyama T, Bouillet P, Miyazaki T, Kadono Y, Chikuda H, Chung UI, Fukuda A, Hikita A, Seto H, Okada T, Inaba T, Sanjay A, Baron R, Kawaguchi H, Oda H, Nakamura K, Strasser A, Tanaka S (2003) Regulation of osteoclast apoptosis by ubiquitylation of proapoptotic BH3-only Bcl-2 family member Bim. EMBO J 22:6653–6664
Tanaka S, Miyazaki T, Fukuda A, Akiyama T, Kadono Y, Wakeyama H, Kono S, Hoshikawa S, Nakamura M, Ohshima Y, Hikita A, Nakamura I, Nakamura K (2006) Molecular mechanism of the life and death of the osteoclast. Ann N Y Acad Sci 1068:180–186
Ikeda F, Matsubara T, Tsurukai T, Hata K, Nishimura R, Yoneda T (2008) JNK/c-Jun signaling mediates an anti-apoptotic effect of RANKL in osteoclasts. J Bone Mineral Res 23:907–914
Iwasawa M, Miyazaki T, Nagase Y, Akiyama T, Kadono Y, Nakamura M, Oshima Y, Yasui T, Matsumoto T, Nakamura T, Kato S, Hennighausen L, Nakamura K, Tanaka S (2009) The antiapoptotic protein Bcl-xL negatively regulates the bone-resorbing activity of osteoclasts in mice. J Clin Investig 119:3149–3159
Takayanagi H, Kim S, Koga T, Nishina H, Isshiki M, Yoshida H, Saiura A, Isobe M, Yokochi T, Inoue J, Wagner EF, Mak TW, Kodama T, Taniguchi T (2002) Induction and activation of the transcription factor NFATc1 (NFAT2) integrate RANKL signaling in terminal differentiation of osteoclasts. Dev Cell 3:889–901
Zhao B, Takami M, Yamada A, Wang X, Koga T, Hu X, Tamura T, Ozato K, Choi Y, Ivashkiv LB, Takayanagi H, Kamijo R (2009) Interferon regulatory factor-8 regulates bone metabolism by suppressing osteoclastogenesis. Nat Med 15:1066–1071
Nishikawa K, Nakashima T, Hayashi M, Fukunaga T, Kato S, Kodama T, Takahashi S, Calame K, Takayanagi H (2010) Blimp1-mediated repression of negative regulators is required for osteoclast differentiation. Proc Natl Acad Sci USA 107:3117–3122
Nishikawa K, Iwamoto Y, Kobayashi Y, Katsuoka F, Kawaguchi SI, Tsujita T, Nakamura T, Kato S, Yamamoto M, Takayanagi H, Ishii M (2015) DNA methyltransferase 3a regulates osteoclast differentiation by coupling to an S-adenosylmethionine-producing metabolic pathway. Nat Med 21:281
Nishikawa K, Iwamoto Y, Ishii M (2013) Development of an in vitro culture method for stepwise differentiation of mouse embryonic stem cells and induced pluripotent stem cells into mature osteoclasts. J Bone Miner Metab 32:331
Li X, Zhao X, Fang Y, Jiang X, Duong T, Fan C, Huang CC, Kain SR (1998) Generation of destabilized green fluorescent protein as a transcription reporter. J Biol Chem 273:34970–34975
Banaszynski LA, Chen LC, Maynard-Smith LA, Ooi AG, Wandless TJ (2006) A rapid, reversible, and tunable method to regulate protein function in living cells using synthetic small molecules. Cell 126:995–1004
Lagnado CA, Brown CY, Goodall GJ (1994) AUUUA is not sufficient to promote poly(A) shortening and degradation of an mRNA: the functional sequence within AU-rich elements may be UUAUUUA(U/A)(U/A). Mol Cell Biol 14:7984–7995
Morita S, Kojima T, Kitamura T (2000) Plat-E: an efficient and stable system for transient packaging of retroviruses. Gene Ther 7:1063–1066
Tanaka S, Takahashi T, Takayanagi H, Miyazaki T, Oda H, Nakamura K, Hirai H, Kurokawa T (1998) Modulation of osteoclast function by adenovirus vector-induced epidermal growth factor receptor. J Bone Mineral Res 13:1714–1720
Kobayashi Y, Take I, Yamashita T, Mizoguchi T, Ninomiya T, Hattori T, Kurihara S, Ozawa H, Udagawa N, Takahashi N (2005) Prostaglandin E2 receptors EP2 and EP4 are down-regulated during differentiation of mouse osteoclasts from their precursors. J Biol Chem 280:24035–24042
Matsubara T, Ikeda F, Hata K, Nakanishi M, Okada M, Yasuda H, Nishimura R, Yoneda T (2010) Cbp recruitment of Csk into lipid rafts is critical to c-Src kinase activity and bone resorption in osteoclasts. J Bone Mineral Res 25:1068–1076
Hsu PD, Lander ES, Zhang F (2014) Development and applications of CRISPR-Cas9 for genome engineering. Cell 157:1262–1278
Komor AC, Badran AH, Liu DR (2017) CRISPR-based technologies for the manipulation of eukaryotic genomes. Cell 168:20–36
Mondal P, Krishnamurthy VV, Sharum SR, Haack N, Zhou H, Cheng J, Yang J, Zhang K (2019) Repurposing protein degradation for optogenetic modulation of protein activities. ACS Synth Biol 8:2585–2592
Bonger KM, Rakhit R, Payumo AY, Chen JK, Wandless TJ (2014) General method for regulating protein stability with light. ACS Chem Biol 9:111–115
Acknowledgments
This work was supported by Grants-in-Aid for Scientific Research (B) from the Japan Society for the Promotion of Science (JSPS) (18H02614 to K. N.); and grants from the Takeda Science Foundation (to K.N.); and the Food Science Institute Foundation (Ryoushoku-kenkyukai to K. N.); and Suzuken Memorial Foundation (to K. N.); and Terumo Life Science Foundation (to K. N.); and Fuji Foundation For Protein Research (to K. N.); and The Tojuro Iijima Foundation for Food Science and Technology (to K. N.); and Nakatani Foundation (to K. N.).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
All authors state that they have no conflicts of interest.
Ethical approval
Animal study was approved by the Institutional Animal Care and Use Committee of both Doshisha University and Osaka University.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
About this article
Cite this article
Nishikawa, K., Ishii, M. Novel method for gain-of-function analyses in primary osteoclasts using a non-viral gene delivery system. J Bone Miner Metab 39, 353–359 (2021). https://doi.org/10.1007/s00774-020-01161-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00774-020-01161-7