Abstract
Among amino acids, leucine is a potential signaling molecule to regulate cell growth and metabolism by activating mechanistic target of rapamycin complex 1 (mTORC1). To reveal the critical structures of leucine molecule to activate mTORC1, we examined the structure–activity relationships of leucine derivatives in HeLa S3 cells for cellular uptake and for the induction of phosphorylation of p70 ribosomal S6 kinase 1 (p70S6K), a downstream effector of mTORC1. The activation of mTORC1 by leucine and its derivatives was the consequence of two successive events: the cellular uptake by l-type amino acid transporter 1 (LAT1) responsible for leucine uptake in HeLa S3 cells and the activation of mTORC1 following the transport. The structural requirement for the recognition by LAT1 was to have carbonyl oxygen, alkoxy oxygen of carboxyl group, amino group and hydrophobic side chain. In contrast, the requirement for mTORC1 activation was more rigorous. It additionally required fixed distance between carbonyl oxygen and alkoxy oxygen of carboxyl group, and amino group positioned at α-carbon. l-Configuration in chirality and appropriate length of side chain with a terminal isopropyl group were also important. This confirmed that LAT1 itself is not a leucine sensor. Some specialized leucine sensing mechanism with rigorous requirement for agonistic structures should exist inside the cells because leucine derivatives not transported by LAT1 did not activate mTORC1. Because LAT1–mTOR axis is involved in the regulation of cell growth and cancer progression, the results from this study may provide a new insight into therapeutics targeting both LAT1 and leucine sensor.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
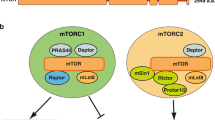
Amino acids are not just the building blocks of proteins and the substrates of metabolic and biosynthetic reactions, but they are also the signaling molecules to regulate cellular metabolism and cell growth. One of the well-recognized cellular signaling pathways mobilized by amino acids is the mechanistic target of rapamycin (mTOR) signaling pathway (Dann and Thomas 2006; Wu 2009; Jewell and Guan 2013). mTOR is an evolutionarily conserved serine/threonine kinase that resides in the center of mTOR signaling pathway and controls cell cycle, growth, metabolism and survival (Brown et al. 1994; Sabatini et al. 1994; Wullschleger et al. 2006). It forms two distinct complexes; mTOR complex 1 (mTORC1) and mTOR complex 2 (mTORC2) (Brown et al. 1994; Sabatini et al. 1994; Wullschleger et al. 2006). While the mechanism of mTORC2 activation is not well understood, the upstream signals and downstream targets of mTORC1 have been extensively studied. mTORC1 regulates cell growth by integrating signals from growth factors, energy status and nutrients such as amino acids (Corradetti and Guan 2006; Jewell et al. 2013). Upon the stimulation by amino acids, the small GTPases RagA/B become active and forms a heterodimeric complex with RagC/D to promote the translocation of mTORC1 to lysosomal membrane surface where Rheb locates and activates mTORC1 (Kim et al. 2008; Sancak et al. 2008, 2010; Bar-Peled et al. 2012). The activated mTORC1 directly phosphorylates two key regulators of translation initiation; p70 ribosomal S6 kinase 1 (p70S6K) and initiation factor 4E binding protein 1 (4E-BP1), whose phosphorylation levels are empirically used as indicators to monitor mTORC1 activity (Brown et al. 1995; Brunn et al. 1997; Xu et al. 1998). Because mTOR is a central regulator of cell growth, over-activation of mTOR leads to cancer progression (Sabatini 2006; Zoncu et al. 2011).
It has been proposed that the level of activation of mTOR by amino acids depends on intracellular amino acid availability (Beugnet et al. 2003). Intracellular amino acids are maintained by autophagic proteolysis during nutrient deprivation, whereas, in nutrient-rich condition, amino acids are provided by the uptake of extracellular amino acids via transporters on the plasma membrane (Mortimore and Poso 1987; Christensen 1990; Kuma et al. 2004; Bröer 2008). Among amino acids occurring in animal body, leucine is the most effective signaling molecule to activate mTORC1 (Xu et al. 1998; Shigemitsu et al. 1999; Stipanuk 2007, Li et al. 2011). Leucine can be transported by Na+-dependent or Na+-independent plasma membrane transporters corresponding to those of amino acid transport systems such as L, y+L, B0, B0,+ and b0,+ (Christensen 1990). Among them, a system L transporter LAT1 (SLC7A5) is proposed to be responsible for the cellular uptake of leucine in many conventional culture cell lines that have been used to study mTOR signaling pathway (Kim et al. 2002; Fuchs and Bode 2005; Nicklin et al. 2009). LAT1 forms a heterodimeric complex with the heavy chain of 4F2 cell surface antigen (4F2hc/CD98hc/SLC3A2) via a disulfide bond and mediates obligatory exchange of large neural amino acids in a Na+-independent manner (Kanai et al. 1998). Leucine-induced mTOR activation is attenuated when LAT1 is inhibited or knocked-down (Nicklin et al. 2009; Yamauchi et al. 2009; Sinclair et al. 2013). Furthermore, LAT1 is highly expressed not only in immortalized culture cell lines but also in various cancer tissues (Yanagida et al. 2001; Fuchs and Bode 2005; Kaira et al. 2009, 2012). The upregulation of LAT1 could enable high amino acid supply and activation of mTOR for cancer cell growth, and control the metabolic reprogramming in an immune response (Sinclair et al. 2013). LAT1 has, thus, been proposed to be a potential molecular target for cancer therapy and immune regulation therapy.
Because of the importance of leucine to activate mTORC1, the structure–activity relationship of leucine derivatives has been studied to reveal critical chemical structures in leucine. It has been shown that the amino group and the carbonyl oxygen of carboxyl group are essential for leucine to activate mTORC1 (Shigemitsu et al. 1999; Lynch et al. 2000). Furthermore, to be the l-isomer and to have the side chain with branched-alkyl group are also critical (Shigemitsu et al. 1999; Lynch et al. 2000). However, exact structural requirements to activate mTORC1 remain unclear, because the contribution of leucine transport through the plasma membrane has not been taken into account even though leucine is supplemented extracellularly and also because a limited number of leucine derivatives have been tested so far. In the present study, we have examined 25 leucine derivatives to determine the critical chemical structures in leucine to be transported by plasma membrane LAT1 as well as to activate mTORC1 in HeLa S3 cells. We have clarified that amino group, carbonyl oxygen and alkoxy oxygen of carboxyl group are required for leucine derivatives to be transported by LAT1, whereas to have fixed position of amino group at α-carbon, l-configuration and a neutral side chain with a branched alkane, particularly terminal isopropyl group, is additionally important for the activation of mTORC1.
Materials and methods
Materials
l-[14C]Leucine was purchased from Moravek Biochemicals (Brea, CA, USA). Standard amino acids, 2-amino-2-norbornane-carboxylic acid (BCH), triiodothyronine (T3), 4-methylvaleric acid, transferrin and anti-β-actin antibody were from Sigma-Aldrich (St. Louise, MO, USA). 2-Aminoisobutyric acid (MeAIB) and l-leucinol were from Tokyo Chemical Industry (Tokyo, Japan). α-Methyl-l-leucine, l-leucine hydroxamate and l-leucic acid were from Bachem (Bubendorf, Switzerland). l-Norvaline and l-norleucine were from Wako Pure Chemical Industries (Osaka, Japan). 4-Methyl-l-norleucine and 5-methyl-d-norleucine were from Amatek Chemical (Jiangsu, China). 5-Methyl-l-norleucine was from Toronto Research Chemicals (Ontario, Canada). l-Leucine 2-propoxyethyl ester [H-Leu-O(CH2)2OPr] and l-leucinylmethoxymethane (H-Leu-CH2OMe) were synthesized as described in the Supplementary method. Other leucine derivatives were from Watanabe Chemical Industries (Hiroshima, Japan). Anti-human LAT1 antibody recognizing N-terminal residues of LAT1 was generated in the previous study (Morimoto et al. 2008). Anti-p70S6K and anti-4F2hc antibodies were purchased from Santa Cruz Biotechnology (Santa Cruz, CA, USA). Other primary antibodies were from Cell Signaling Technology (Danvers, MA, USA). Secondary antibodies conjugated with horseradish peroxidases (HRP) and Cy5 were from Jackson ImmunoResearch (West Grove, PA, USA). Unless specially denoted, other chemicals were purchased from Wako Pure Chemical Industries (Osaka, Japan). The leucine derivatives used in this study are classified for convenience into four groups upon the modification sites: group 1, carboxyl group-modified; group 2, amino group-modified; group 3, modified on α-carbon; group 4, side chain-modified (Fig. 1).
Cell culture
Human cell lines HeLa S3 and T24 were maintained in minimum essential medium (MEM) supplemented with 10 % fetal bovine serum (FBS, Invitrogen; NY, USA). The cells were cultured at 37 °C with 5 % CO2 and humidity. The mouse cell line stably expressing human LAT1 (S2-LAT1) has been established as described previously (Morimoto et al. 2008). S2-LAT1 cells were maintained in RITC80-7 medium (Research Institute for the Functional Peptide; Yamagata, Japan) containing 5 % FBS, 14 mM HEPES, 10 mg/l transferrin, 80 U/l insulin, 10 μg/l recombinant human EGF (Funakoshi; Tokyo, Japan) and 0.78 mM G418 sulfate at 33 °C with 5 % CO2 and humidity.
Immunoblotting of membrane proteins
HeLa S3 and T24 cells were washed twice with ice-cold PBS (pH 7.4) and homogenized in PBS containing complete EDTA-free protease inhibitor cocktail (Roche Diagnostics GmBH, Mannheim, Germany) by a Teflon-potter homogenizer. Whole lysate were centrifuged at 1000×g for 5 min to eliminate unbroken cells. The supernatant was collected and further centrifuged at 10,000×g for 10 min. The supernatant was then collected and centrifuged at 391,000×g for 30 min to sediment membranes. The pellet was dissolved in PBS containing 1 % n-dodecyl-β-d-maltoside (DDM). Whole processes were performed on ice or at 4 °C. For immunoblotting, 10 µg of membrane fraction was mixed with sample buffer containing 50 mM Tris pH 6.8, 2 % SDS with 100 mM DTT for reducing condition or without DTT for non-reducing condition. Protein samples in the sample buffer without heat treatment were resolved in 10 % SDS-PAGE. The proteins in SDS–polyacrylamide gel were transferred electrophoretically onto Hybond-P PVDF membrane (GE Healthcare; Buckinghamshire, UK). Membranes were blocked at room temperature for 1 h with Tris-buffered saline Tween-20 (TBS-T) containing 5 % skim milk and incubated overnight at 4 °C with primary antibodies. The membranes were then washed with TBS-T and incubated at room temperature for 1 h with appropriate secondary antibodies conjugated to horseradish peroxidase. Signals were developed by ECL Plus Western Blotting Detection System (GE Healthcare) and visualized using LAS-4000 mini Ver. 2.0 (Fujifilm Corporation; Tokyo, Japan).
Immunostaining
HeLa S3 cells were cultured on glass coverslip (Iwaki; Tokyo, Japan) for 2 days. Cells were fixed with cold methanol for 15 min. The coverslips were rinsed twice with cold PBS and incubated with 5 % skim milk in PBS at 4 °C for 1 h. The coverslips were, then, incubated with primary antibodies in PBS containing 5 % skim milk at 4 °C overnight. After the reaction with primary antibodies, the coverslips were washed three times with cold PBS and incubated with appropriate secondary antibodies conjugated with Alexa Fluor 488 or Cy5 at 4 °C for 1 h. After the washing with cold PBS, the coverslips were mounted on glass slide using Fluoromount/Plus (Diagnostic Biosystems; Pleasanton, CA, USA) and imaged by using Carl Zeiss laser scanning confocal microscope LSM-510 META (Heidelberg, Germany).
Uptake and inhibition experiments
HeLa S3 cells or S2-LAT1 cells were seeded onto 24-well plates (1 x 105 cells/well) and cultured for 2 days to obtain ~90 % confluence. The cells were washed three times with 37 °C pre-warmed Na+-free Hank’s balanced salt solution (HBSS) containing 125 mM choline-Cl, 25 mM HEPES, 4.8 mM KCl, 1.2 mM MgSO4, 1.2 mM KH2PO4, 1.3 mM CaCl2, and 5.6 mM glucose (pH 7.4) and further incubated in the same buffer at 37 °C for 10 min. l-Leucine uptake was measured for 1 min at 37 °C in the same buffer containing 1 µM l-[14C]leucine (2 MBq/mmol). In the inhibition experiment, test compounds was added with l-[14C]leucine. Uptake was terminated by removing the solution followed by washing three times with ice-cold Na+-free HBSS. In the experiments to measure l-[14C]leucine uptake in the presence of Na+, standard HBSS containing 125 mM NaCl was used instead of Na+-free HBSS. The cells were lysed with 0.1 N NaOH and the lysates were mixed with Emulsifier safe cocktail (PerkinElmer; Waltham, MA, USA). The radioactivity was measured with β-scintillation counter (LSC-3100, Aloka; Tokyo, Japan).
Efflux measurements
HeLa S3 cells or S2-LAT1 cells on 24-well plates were preloaded with l-[14C]leucine by incubating in Na+-free HBSS containing 1 µM l-[14C]leucine (2 MBq/mmol) at 37 °C for 10 min. After washing three times with pre-warmed Na+-free HBSS, efflux of radioactivity was induced by incubating the cells with Na+-free HBSS in the presence or absence of indicated concentration of test compounds for 1 min at 37 °C. The medium was, then, collected and its radioactivity was counted. The cells were lysed with 0.1 N NaOH and the radioactivity of cell lysate was counted. The l-[14C]leucine efflux values were expressed as percentage radioactivity (radioactivity of the medium/(radioactivity of the medium + radioactivity of the cells)).
Cell treatment and immunoblotting of signaling pathway proteins
To examine the effect of leucine and leucine derivatives on the phosphorylation of signaling pathway proteins, HeLa S3 cells were seeded on 60-mm plates and cultured for 2 days to obtain ~50 % confluence. The cells were washed twice with Dulbecco’s Phosphate Buffered Saline (DPBS, pH 7.4) and maintained in MEM deprived of FBS for 16 h. Then, the cells were washed twice with DPBS and maintained in DPBS for 2 h for amino acid starvation. In order to stimulate the cells with l-leucine or test compounds, DPBS was replaced by MEM in which amino acids were substituted by indicated concentration of l-leucine or 400 µM of test compounds plus 0.5× amino acids without l-leucine. Because l-glutamine and some other amino acids were necessary for extracellularly applied l-leucine to activate mTOR (Nicklin et al. 2009), 0.5× amino acids without l-leucine were added in the treatment media. The 0.5× amino acids without l-leucine set at the half of those of MEM were composed of 300 µM l-arginine, 50 µM l-cystine, 1,000 µM l-glutamine, 100 µM l-histidine, 200 µM l-isoleucine, 200 µM l-lysine, 50 µM l-methionine, 100 µM l-phenylalanine, 200 µM l-threonine, 25 µM l-tryptophan, 100 µM l-tyrosine and 200 µM l-valine. To avoid the chemical degradation, all leucine derivatives were dissolved freshly in the media just before use for each experiment. As a control, the cells were incubated in MEM containing 0.5× amino acids without l-leucine. To examine the effect of BCH, 20 mM BCH was added to the incubation medium.
After cell treatment, the cells were washed twice with ice-cold DPBS and lysed in ice-cold lysis buffer containing 50 mM Tris–Cl (pH 7.4), 120 mM NaCl, 20 mM NaF, 1 mM EDTA, 6 mM EGTA, 20 mM β-glycerophosphate (Calbiochem; La Jolla, CA, USA), 1 mM DTT, 0.5 mM phenylmethanesulfonyl fluoride, 1 mM Na3VO4, 1 % NP-40 and complete EDTA-free protease inhibitor cocktail. The soluble fraction of cell lysates was prepared by centrifugation at 20,400×g for 30 min. Protein concentration of soluble fraction was determined by BCA protein assay (Pierce Biotechnology; Rockford, IL, USA). To measure phospho-S2448 mTOR, mTOR, phospho-T389 p70S6K, p70S6K, phospho-T37/46 4E-BP1, phospho-S65 4E-BP1, 4E-BP1, phospho-T180/182 p38, p38, phospho-T202/204 p44/42, p44/42, phospho-T183/185 stress-activation protein kinase/Jun amino–amino terminal kinase (SAPK/JNK) and SAPK/JNK, the soluble fraction containing 20 μg of total protein was mixed with sample buffer containing 50 mM Tris pH 6.8, 2 % SDS and 100 mM DTT. Then, the protein samples were heated at 95 °C for 5 min and resolved in 10–15 % SDS-PAGE. Immunoblotting was performed as described above. The signals were quantified with Multi Gauge Ver. 3.1 (Fujifilm Corporation).
The concentration-dependent effect of l-leucine on p70S6K phosphorylation was evaluated by plotting l-leucine-induced p70S6K phosphorylation against l-leucine concentration. The l-leucine-induced p70S6K phosphorylation was obtained by subtracting the level of p70S6K phosphorylation in the absence of l-leucine from that in the presence of indicated concentration of l-leucine. EC50 value was calculated by fitting the concentration-dependence of l-leucine-induced p70S6K phosphorylation to the following equation using SigmaPlot 13 (HULINKS): E = E max × C/(C + EC50), where E, E max and C represent the effect of l-leucine (the value for l-leucine induced p70S6K phosphorylation), maximal effect and l-leucine concentration, respectively.
Results
Transporter for leucine uptake in HeLa S3 cells
The uptake of l-[14C]leucine was measured in the presence or absence of Na+ with or without inhibitors to reveal transporters responsible for l-leucine uptake in HeLa S3 cells. As shown in Fig. 2a, the rate of l-[14C]leucine uptake measured without inhibitors in the absence of Na+ was similar to that measured in the presence of Na+, indicating that l-leucine uptake in HeLa S3 cells is Na+-independent. The uptake was inhibited by BCH but not by l-arginine (Fig. 2a). Among Na+-independent amino acid transport systems, systems L and b0,+ are leucine-transporting ones (Christensen 1990; Bröer 2008). Because BCH inhibits system L but not b0,+ whereas l-arginine, a substrate of system b0,+, inhibits system b0,+ but not L, it is suggested that the l-leucine uptake in HeLa S3 cells is mediated by system L (Christensen 1990; Bröer 2008). Among four system L isoforms, LAT1, LAT2, LAT3 and LAT4, we concluded that LAT1 is responsible for the l-leucine uptake, because the l-leucine uptake was inhibited by α-methyl-l-tyrosine (AMT) and triiodothyronine (T3), both of which inhibit LAT1 specifically (Morimoto et al. 2008; Khunweeraphong et al. 2012; Wiriyasermkul et al. 2012), whereas the l-leucine uptake was not affected by glycine that inhibits LAT2 but not others (Fig. 2a; Bröer 2008).
l-Leucine uptake and the responsible transporter in HeLa S3 cells. a l-[14C]Leucine uptake in HeLa S3 cells. The uptake of l-[14C]leucine (1 µM) was measured for 1 min in the presence (closed column) or absence (open column) of Na+. Inhibition of l-[14C]leucine uptake by 1 mM BCH (2-amino-2-norbornane-carboxylic acid), 1 mM AMT (l-α-methyl-tyrosine), 30 µM T3 (triiodothyronine), 1 mM l-arginine or 1 mM glycine was examined. Minus the uptake of l-[14C]leucine (1 µM) without inhibitors. Data represent mean ± SEM, n = 4. b Expression of endogenous LAT1 and 4F2hc in HeLa S3 and T24 cells. Immunoblotting of crude membrane fractions prepared from HeLa S3 and T24 cells were performed as described in the “Materials and methods” using antibodies against LAT1 (left panel) and 4F2hc (right panel). The crude membrane fractions were treated with (+DTT) or without (−DTT) 100 mM DTT for reducing or non-reducing condition, respectively, prior to SDS-PAGE. The closed arrow in the left panel indicates LAT1 monomer. Open and closed arrows in the right panel indicate 4F2hc monomers in HeLa S3 cells, whereas single star represents 4F2hc monomer in T24 cells. Different sizes of 4F2hc monomers in Hela S3 and T24 cells are presumably due to different glycosylated forms (Fort et al. 2007). Open arrowheads, filled arrowheads and double stars in the left and right panels indicate heterodimeric complexes of LAT1 and 4F2hc, which correspond with 4F2hc monomers indicated by open arrow, filled arrow and single star in the right panel, respectively. c Immunofluorescence of endogenous LAT1 in HeLa S3 cells. The scale bar indicates 10 µm
The functional expression of LAT1 was then confirmed by immunoblotting using crude membrane fraction prepared from HeLa S3 cells. The 37 and 70–100 kDa bands corresponding to LAT1 and 4F2hc, respectively, were detected under reducing condition (Fig. 2b). Compared with T24 cells, previously characterized to express high level of LAT1 (Kim et al. 2002), HeLa S3 cells expressed equivalent amount or more of LAT1 and 4F2hc. On the other hand, LAT2 was not detected in HeLa S3 cells (data not shown). The different sizes of 4F2hc between HeLa S3 and T24 cells on the acrylamide gel are likely due to different glycosylated forms (Yanagida et al. 2001; Fort et al. 2007; Khunweeraphong et al. 2012). Under the non-reducing condition, formation of LAT1 and 4F2hc functional-complex was confirmed by the size-shift of the bands detected using anti-LAT1 and anti-4F2hc antibodies (Fig. 2b). Immunostaining of HeLa S3 cells further confirmed the localization of LAT1 on the plasma membrane, where LAT1 exerts its function as a transporter to take up amino acids into the cells (Fig. 2c).
Structural requirements to be transported by LAT1
Because LAT1 is responsible for l-leucine transport in HeLa S3 cells, we examined the effect of leucine derivatives on the l-leucine transport in HeLa S3 cells to characterize their interaction with LAT1. We performed inhibition experiments in which the uptake of 1 µM l-[14C]leucine was measured in the presence of 1 mM test compounds. We, furthermore, conducted efflux experiments to examine whether 1 mM test compounds induce the efflux of preloaded l-[14C]leucine. The efflux experiments were intended to determine whether the compounds that inhibit leucine uptake are transportable substrates or non-transportable blockers of LAT1. Because LAT1 mediates obligatory exchange of amino acids, the uptake of extracellularly applied substrates induces the efflux of preloaded l-[14C]leucine (Kim et al. 2002; Wiriyasermkul et al. 2012). To induce efflux of radioactivity is, therefore, indicating that the test compounds are transported by LAT1.
In the inhibition experiments, we first confirmed that l-leucine itself and system L inhibitor BCH highly inhibited l-[14C]leucine uptake (Fig. 3a). Among leucine derivatives in group 1 with carboxyl group modifications, l-leucinol did not inhibit l-[14C]leucine uptake, whereas leucine esters (H-Leu-OMe, H-Leu-OEt, H-Leu-OAll, H-Leu-O(CH2)2OPr, H-Leu-OtBu and H-Leu-OBzl) inhibited l-[14C]leucine uptake to a high or medium extent (Fig. 3a). l-Leucinylmethoxymethane (H-Leu-CH2OMe) and l-leucine hydroxamate (H-Leu-NHOH) also inhibited l-[14C]leucine uptake but the inhibition was weaker than the leucine esters (Fig. 3a).
The effects of leucine derivatives on leucine uptake and leucine efflux in HeLa S3 cells. a Inhibition of l-[14C]leucine uptake by leucine derivatives. The uptake of 1 μM l-[14C]leucine was measured for 1 min in the presence or absence (minus) of 1 mM leucine derivatives. The ordinate indicates % inhibition in which complete inhibition of l-[14C]leucine uptake was taken as 100 % whereas no inhibition was 0 %. Data represent mean ± SEM, n = 4–6. b Efflux of preloaded l-[14C]leucine induced by leucine derivatives. The efflux of preloaded l-[14C]leucine was measured for 1 min in the presence or absence (minus) of extracellularly applied leucine derivatives (1 mM). The radioactivity released from the cells was expressed as percent of the total preloaded radioactivity (% radioactivity). Data represent mean ± SEM, n = 4–6
In the efflux experiments, we evaluated the assay condition by checking the effect of l-leucine, a typical substrate of LAT1, and BCH, a competitive inhibitor but transported by LAT1. As shown in Fig. 3b, both l-leucine and BCH induced high level of efflux of preloaded l-[14C]leucine. Consistent with the results from inhibition experiments, l-leucinol did not induce the efflux of preloaded l-[14C]leucine compared with the background efflux measured in the absence of test compounds (Fig. 3b). In contrast, the leucine esters induced the efflux at the similar level to that of l-leucine, so that they are transported by LAT1 efficiently. H-Leu-CH2OMe and H-Leu-NHOH induced lower level of efflux. Therefore, it is suggested that the carbonyl oxygen is indispensable, alkoxy oxygen of carboxyl group is important but not essential and hydrogen atom in the carboxyl group is not necessary to be recognized by LAT1. H-Leu-CH2OMe and H-Leu-NHOH are less efficiently transported by LAT1 probably due to the shift of oxygen atom corresponding to alkoxy oxygen to less appropriate position distant from the carbonyl oxygen.
As for group 2 with amino group modifications, none of the compounds exhibited significant inhibition and efflux (Fig. 3a, b). Not only the compounds lacking nitrogen atom (l-leucic acid and 4-methylvaleric acid) but also the ones whose hydrogen atom of the amino group is replaced by functional groups (Me-Leu-OH, For-Leu-OH and Ac-Leu-OH) failed to exert any effect. In addition, compounds with both amino group and carboxyl group modifications showed no inhibition and efflux (Supplementary Fig. 1a–c). Thus, the amino group (NH2 or NH3 +) is critical for the interaction with LAT1. To be noticed, l-β-homoleucine in group 3, which has an amino group at β-carbon of carboxylic acid, was transported by LAT1, supporting the importance of amino group around or near the α-carbon (Fig. 3a, b).
l-β-Homoleucine, α-methyl-l-leucine (H-α-Me-Leu-OH) and d-leucine in group 3 with modifications on α-carbon and all the side chain-modified leucine derivatives (group 4) strongly inhibited l-[14C]leucine uptake and induced high level of efflux, indicating that they are efficiently transported by LAT1 (Fig. 3a, b). Such a substrate selectivity as preferring large neutral amino acids with hydrophobic side chains and accepting α-methyl-amino acids and d-leucine is consistent with that of LAT1 characterized by expressing it in Xenopus oocytes and culture cells (Kanai et al. 1998; Yanagida et al. 2001; Uchino et al. 2002; Morimoto et al. 2008; Khunweeraphong et al. 2012; Wiriyasermkul et al. 2012).
To confirm the results obtained with HeLa S3 cells, we carried out the similar experiments on the exogenously expressed human LAT1 using S2-LAT1 cells stably expressing human LAT1 that we previously established and characterized (Morimoto et al. 2008). Experiments were done with the leucine derivatives at 100 μM corresponding to the Ki value of l-leucine. All the leucine derivatives showed similar effects in S2-LAT1 cells except H-Leu-CH2OMe, H-Leu-NHOH and H-α-Me-Leu-OH, which exerted less inhibition and induced less efflux in S2-LAT1 cells (Supplementary Fig. 2a, b) probably due to the lower concentration of compounds used for S2-LAT1 cells compared with that for HeLa S3 cells (1 mM). H-Leu-CH2OMe, H-Leu-NHOH and H-α-Me-Leu-OH are supposed to be the substrates of LAT1 with lower affinity compared with the other LAT1 substrates in group 4 and group 1.
Activation of mTORC1 by l-leucine
To examine whether l-leucine and l-leucine derivatives taken up by LAT1 into HeLa S3 cells activates mTORC1, we measured the phosphorylation of mTORC1 pathway proteins. HeLa S3 cells were cultured in serum- and amino acid-deprived media, and then incubated in MEM containing 0.5× amino acids with or without l-leucine. As shown in Fig. 4, l-leucine increased the phosphorylation of mTOR and downstream p70S6K and 4E-BP1. Furthermore, their phosphorylation was inhibited by BCH, confirming that it was mediated by LAT1. In contrast, l-leucine did not alter the phosphorylation of p38 mitogen-activated protein kinase (MAPK), p44/42 MAPK and SAPK/JNK (Fig. 4). Thus, MAPK pathways are not involved in leucine-induced p70S6K phosphorylation but LAT1-mediated l-leucine uptake stimulates mTORC1 signaling to activate p70S6K, although p70S6K is known to be activated by MAPK-signaling pathways as well as mTOR pathway (Iijima et al. 2002; Lehman and Gomez-Cambronero 2002).
The effect of l-leucine on mTORC1 and MAPK pathways. After serum- and amino acid deprivation as described in the “Materials and methods”, HeLa S3 cells were incubated in MEM containing ×0.5 amino acids with or without 400 μM l-leucine for 120 min. The effect of BCH was studied by adding 20 mM BCH during the incubation period. mTORC1 signaling pathway proteins, mTOR, p70S6K and 4E-BP1, are shown in a. MAPK signaling pathway proteins, p38 MAPK (p38), p44/42 MAPK (p44/42) and stress-activation protein kinase/Jun amino–amino terminal kinase (SAPK/JNK), are in b. l-leucine increased the phosphorylation of mTORC1 pathway proteins, whereas MAPK pathway proteins were not affected by l-leucine. BCH completely suppressed the phosphorylation induced by l-leucine. Antibodies against 4E-BP1 and phosphorylated (T37/46 and S65) 4E-BP1 recognized three bands, “α”, “β” and “γ”, which represent hypophosphorylated, phosphorylated and hyperphosphorylated 4E-BP1, respectively (a)
The dependence of p70S6K phosphorylation on incubation time and l-leucine concentration was examined. As shown in Fig. 5a, the phosphorylation of p70S6K increased over the incubation period and peaked at 90–120 min. The level of phosphorylation was dependent on the concentration of l-leucine (Fig. 5b, c). The EC50 value to induce p70S6K phosphorylation was 33.9 µM for l-leucine (Fig. 5c).
Phosphorylation of p70S6K induced by l-leucine. a Time-dependent phosphorylation of p70S6K induced by l-leucine. After serum- and amino acid deprivation as described in the “Materials and methods”, HeLa S3 cells were incubated in MEM containing ×0.5 amino acids with (plus) or without (minus) 400 μM l-leucine for the indicated time periods. Phosphorylated p70S6K (upper panel) and total p70S6K (lower panel) were measured by immunoblotting using anti-phospho-p70S6K and anti-p70S6K antibodies, respectively. p85 indicates p85S6K, another kinase isoform derived from alternative splicing of the same mRNA as that for p70S6K. Non-specific bands under the band for p70S6K (“star” in the lower panel) sometimes appeared dependent on the lot of antibodies used. b Concentration-dependent phosphorylation of p70S6K. HeLa S3 cells were incubated for 120 min in MEM containing ×0.5 amino acids with indicated concentration of l-leucine. Phosphorylated p70S6K and total p70S6K were detected as described above. “Star” in the lower panel indicates non-specific bands as described in a. c Dependence of p70S6K phosphorylation on l-leucine concentration. Signal intensities of phosphorylated p70S6K in b were quantified and normalized by the signals of total p70S6K. The values were plotted against l-leucine concentration. EC50 was calculated to be 33.9 μM by fitting to E = E max × C/(C + EC50), where E, E max and C represent the effect of l-leucine (the value of l-leucine-induced p70S6K phosphorylation), maximal effect and l-leucine concentration, respectively. au arbitrary unit in which E max value is set to 1.0
Structural requirements for mTORC1 activation
To reveal the critical structures in a leucine molecule to activate mTORC1, we examined the effects of leucine derivatives on the levels of p70S6K phosphorylation. After the pretreatment of HeLa S3 cells with serum- and amino acid-deprived media, the cells were incubated with test compounds at 400 µM for 120 min, in which l-leucine could induce maximal response (Fig. 5c). In each set of experiment shown in Fig. 6a, the effects of test compounds on p70S6K phosphorylation were normalized with respect to that of l-leucine and shown in Fig. 6b.
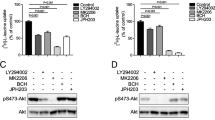
The effects of leucine derivatives to induce p70S6K phosphorylation. a p70S6K phosphorylation induced by leucine derivatives. After serum- and amino acid deprivation as described in the “Materials and methods”, HeLa S3 cells were incubated in MEM containing ×0.5 amino acids with 400 μM l-leucine or indicated leucine derivatives for 120 min. As a control, the cells were incubated as described above except that the incubation media did not contain l-leucine or leucine derivatives (minus). To examine the effect of BCH on p70S6K phosphorylation induced by l-leucine or leucine derivatives, 20 mM BCH was added in the incubation media with l-leucine or leucine derivatives. Phosphorylation of p70S6K was measured as described in the legend to Fig. 5a. b Quantified level of p70S6K phosphorylation. The band intensities obtained by densitometry were shown in the presence (open column) or absence (closed column) of BCH. In each set of experiment shown in a, the effects of test compounds on p70S6K phosphorylation were normalized with respect to that of l-leucine and shown in b. 100 % corresponds to the amount of phospho-p70S6K in the presence of 400 μM l-leucine. Data represent mean ± SEM from three independent experiments. Gray horizontal line indicates the level of background phosphorylation of p70S6K measured in the absence of l-leucine or leucine derivatives and in the absence of BCH. “Star” indicates statistical significance (p < 0.05) compared with control (in the absence of l-leucine or leucine derivatives and in the absence of BCH)
Among leucine derivatives in group 1, all leucine esters (H-Leu-OMe, H-Leu-OEt, H-Leu-OAll, H-Leu-O(CH2)2OPr, H-Leu-OtBu and H-Leu-OBzl) induced high level of phosphorylation on p70S6K, whereas l-leucinol did not (Fig. 6a, b), indicating that the hydrogen atom of carboxyl group of the compounds is replaceable, whereas the carbonyl oxygen is crucial to increase p70S6K phosphorylation when leucine derivatives are applied extracellularly. Substitution of the hydrogen atom in carboxyl group with methyl, ethyl, allyl, propoxyethyl or benzyl group did not remarkably attenuate the effect compared with that of l-leucine, whereas the substitution with tert-alkyl group (H-Leu-OtBu) decreased p70S6K phosphorylation to the half. H-Leu-CH2OMe and H-Leu-NHOH did not induce significant level of p70S6K phosphorylation. Therefore, similar to the requirements to be transported by LAT1, the carbonyl oxygen is indispensable, whereas the hydrogen atom in the carboxyl group can be substituted for the activation of mTORC1. However, it may also be possible that leucine esters taken up into cells were hydrolyzed to leucine by intracellular esterases and subsequently activated mTORC1.
In group 2 with amino group-modifications, none of the compounds induced p70S6K phosphorylation (Fig. 6a, b). Compounds with both amino group and carboxyl group modifications did not induce the phosphorylation (Supplementary Fig. 1d). Thus, the amino group (NH2 or NH3 +) is indispensable for extracellularly applied leucine derivatives to activate mTORC1.
In group 3 with modifications on α-carbon, l-β-homoleucine with an amino group at β-carbon of carboxylic acid did not induce the phosphorylation of p70S6K, suggesting the importance of amino group positioned at α-carbon. The leucine derivatives that have methyl group at α-carbon (H-α-Me-Leu-OH) weakly but significantly increase p70S6K phosphorylation. It suggests that the hydrogen atom on α-carbon can be replaced at least with a methyl group (Fig. 6b). As shown in Fig. 6a, b, p70S6K phosphorylation is also induced by d-isomers such as d-leucine (group 3) and 5-methyl-d-norleucine [H–d-Nle(5-Me)-OH] (group 4). The levels of induced phosphorylation were lower than those of the corresponding l-isomers (Fig. 6b).
Leucine derivatives in group 4 with side-chain modifications induced p70S6K phosphorylation at various levels (Fig. 6a, b). They have side-chains of n-alkyl or branched-alkyl groups with different length and varied branching (Fig. 1). Among the leucine derivatives with a side chain of three-carbon length [l-leucine, l-isoleucine, γ-methyl-l-leucine [H-Leu(Me)-OH] and l-norvaline (H-Nva-OH)], l-leucine and l-isoleucine induced p70S6K phosphorylation although the level of phosphorylation by l-isoleucine was much less than that by l-leucine (Fig. 6a, b). l-Leucine and l-isoleucine have a branching in the side chain, whereas H-Nva-OH dose not, suggesting that to hold a branched side chain but not with a tertiary butyl structure is important to activate mTORC1. It also suggests that a structure with branching at the end of the alkyl group (to have a terminal isopropyl group) most effectively activates mTORC1. This was supported by the observation on the leucine derivatives with a side chain of four- or five-carbon length [4-methyl-l-norleucine (H-Nle(4-Me)-OH), l-norleucine (H-Nle-OH], l-2-amino-heptanoic acid [H-Ahep(2)-OH] and 5-methyl-l-norleucine [H-Nle(5-Me)-OH)]. Among them, H-Nle(5-Me)-OH which has a branching (an isopropyl group) at the end of the alkyl group induced relatively high level of p70S6K phosphorylation (Fig. 6a, b). Concerning the length of side chains, it was concluded that the side chain with three-carbon length is optimal because l-leucine with three-carbon-length side chain is more effective than H-Nle(5-Me)-OH with four-carbon-length side chain although they have the same branching at the end of side-chains. Similarly, l-isoleucine with three-carbon-length side chain was slightly effective but H-Nle(4-Me)-OH with four-carbon-length side chain was not. Therefore, to have a side chain with three-carbon length and to have a terminal isopropyl group are required to activate mTORC1 effectively. It is also noted that BCH, which is recognized by LAT1 as a substrate, did not activate mTORC1, probably because of its bulky bicycloheptane structure in the side chain (Fig. 4).
Finally, the p70S6K phosphorylation induced by l-leucine and leucine derivatives were almost completely suppressed by BCH that competitively inhibits system L, confirming that the phosphorylation is due to LAT1-mediated uptake of l-leucine and leucine derivatives (Figs. 4, 6).
Discussion
Leucine is a potential signaling molecule to activate mTORC1. We found that the leucine derivatives which activated mTORC1 (Fig. 6b) were transported by LAT1 (Fig. 3b) and the mTORC1 activation was suppressed by BCH inhibiting LAT1, which indicates that the leucine derivatives are taken up into cells by LAT1 and subsequently activate mTORC1 in HeLa S3 cells. This is in agreement with the previous study showing that the uptake of leucine by LAT1 is the rate-limiting step for the activation of mTORC1 (Nicklin et al. 2009). Because leucine derivatives were not able to activate mTORC1 unless they were transported by LAT1, the recognition of the compounds by LAT1 as substrates should be considered as the primary step of leucine-induced mTORC1 activation. We, thus, attempted to the specify structural requirements for two steps to activate mTORC1, one for the recognition by LAT1 and the other for mTORC1 activation after the compounds are taken up into the cells. The results from the present study indicate that carbonyl oxygen, alkoxy oxygen of carboxyl group, amino group and hydrophobic side chain are necessary for the compounds to be recognized by LAT1 (Fig. 7a). On the other hand, the position of amino group at α-carbon, the l-configuration in chirality and the side chain of appropriate length with a terminal isopropyl group are proposed to be additionally important to activate mTORC1 after the compounds enter the cells (Fig. 7b).
Proposed important moieties for the recognition by LAT1 and for the activation of mTORC1. The important moieties for the recognition by LAT1 (a) and for the activation of mTORC1 (b) are proposed on the structure of leucine. Dark gray highlight indicates the structures of leucine that are crucial and indispensable. Light gray highlight indicates the structures of leucine that are important but can be modified. a LAT1 recognizes carbonyl oxygen, alkoxy oxygen and amino group of leucine (dark gray highlight). The side chain of leucine can be replaced by the other hydrophobic side chains. The distance between carbonyl oxygen and the oxygen corresponding to alkoxy oxygen (light gray highlight) can be increased, while that of leucine is the optimal. b For leucine to activate mTORC1, carbonyl oxygen, alkoxy oxygen and amino group positioned at α-carbon (dark gray highlight) are indispensable. The terminal isopropyl group is optimal to activate mTORC1, but the branching can be positioned at β-carbon although the agonistic activity is highly retarded (light gray highlight). The length of the side chain is also critical: three-carbon-length is optimal for the side chain (light gray highlight)
The structure–activity relationship regarding the substrate recognition by LAT1 was studied previously (Uchino et al. 2002; Geier et al. 2013). It was proposed that carboxyl group, amino group and hydrophobic side chain are important (Uchino et al. 2002; Geier et al. 2013), which is consistent with our results obtained in the present study. In our study, by analyzing variety of leucine derivatives more comprehensively, we furthermore specified important moieties in the carboxyl group. We found that carbonyl oxygen and alkoxy oxygen, but not hydrogen of the carboxyl group, are the key moieties for leucine to be recognized by LAT1 (Fig. 7a). Carbonyl oxygen and alkoxy oxygen do not necessarily have to be close to each other, although the strongest interaction with LAT1 occurs when these two oxygen atoms are adjacent (Figs. 3a, 7a). Notably, amino group is not necessary to be fixed at α-carbon as evidenced by high level of l-β-homoleucine uptake (Fig. 3b). To the best of our knowledge, this is the first report to reveal the recognition of β-amino acids by LAT1. Concerning γ-amino acids, previous studies showed that LAT1 recognizes gabapentin, pregabalin and baclofen (van Bree et al. 1988; Su et al. 2005; Geier et al. 2013). Therefore, LAT1 recognizes not only α- but also β- and γ-amino acids.
Although the crystal structure of LAT1 is not available, the already solved crystal structure of its bacterial ortholog arginine:agmatine antiporter AdiC and that of bacterial leucine transporter LeuT (Yamashita et al. 2005; Gao et al. 2010) are useful to speculate the mechanism of substrate recognition by LAT1. The protein folding and pseudosymmetry of AdiC are similar to those of LeuT (so called LeuT-fold) and their proposed substrate recognition mechanisms are comparable. Substrates of both transporters bind to their substrate binding sites via hydrogen bonds at their carbonyl oxygen, alkoxy oxygen and amino group. To bind with the side chains of substrates, AdiC interacts with the guanidinum group of substrate arginine probably through cation-π interaction, whereas the binding pocket of LeuT binds the side chain of leucine via van der Waals forces. Therefore, we propose that the substrate binding site of LAT1, similar to those of AdiC and LeuT, recognizes carbonyl oxygen, alkoxy oxygen and amino group of the substrates. LAT1 likely interacts with carbonyl oxygen and alkoxy oxygen via hydrogen bonds rather than ionic interactions, because all the leucine esters, whose ester groups are not charged, were recognized by LAT1. The binding pocket of LAT1 might be similar to that of LeuT that accepts the hydrophobic side chain. Moreover, the binding site, which recognizes the carboxyl group of substrates, is presumably flexible to accept carbonyl oxygen and an additional oxygen atom corresponding to alkoxy oxygen even though the distance between two oxygen atoms is varied.
Previous studies suggested that to have carbonyl oxygen, amino group, l-configuration in chirality and side chain of leucine is important to increase the phosphorylation of 4E-BP1, a downstream of mTORC1 (Shigemitsu et al. 1999; Lynch et al. 2000). Our findings in the present study were basically in agreement with this proposal, but, by using wider variety of leucine derivatives, we described more detailed structures in leucine critical to activate mTORC1. We found that the carbonyl oxygen, alkoxy oxygen of carboxyl group, amino group, position of amino group at α-carbon, l-configuration in chirality and appropriate length of side chain with a terminal isopropyl group are important features to activate mTORC1 when the compound is applied extracellularly. As already indicated, the activation of mTORC1 by extracellularly applied leucine is the consequence of two successive events: step 1, transport by LAT1; step 2, activation of mTORC1 following the transport. Because carbonyl oxygen, alkoxy oxygen of carboxyl group and amino group are required for step1, it is not concluded whether they are also required for step 2 or just only for step 1. However, step 2 exhibits more rigorous spatial requirements concerning carboxyl and amino groups than that of step 1. Strict requirement for the distance between carbonyl oxygen and oxygen atom corresponding to alkoxy oxygen of carboxyl group, and strict requirement for the position of amino group at α-carbon are the characteristic features for step 2. As for the chirality, although LAT1 transports both l- and d-leucine (Fig. 3b and Supplementary Fig. 2b), its preference for l-configuration becomes evident at lower concentration of the compounds (Yanagida et al. 2001). We propose that step 2 would be further more rigorous in the preference for l-configuration, because the apparent difference in mTOR activation between stereoisomers looks more evident even at higher concentration (400 µM in Fig. 6b) compared with the difference between stereoisomers in the interaction with LAT1 at 100 µM (Supplementary Fig. 2), although we cannot make a direct comparison between the uptake experiments and the mTOR activation experiments due to the possible competition of extracellularly applied test compounds with LAT1 substrates contained in 0.5× amino acids used for the mTOR activation experiments. The difference between step 1 and step 2 is the most evident in the requirement for the side chain. To have a hydrophobic and bulky side chain is enough for step 1, whereas the appropriate length of side chain with a terminal isopropyl group is preferred for step 2 (Fig. 7a, b).
In this study, we showed that leucine esters strongly activated mTORC1 (Fig. 6a, b). However, because of the compound stability inside the cells, it is not concluded whether the hydrogen of carboxyl group is dispensable in step 2 or not. It might be possible that, once they entered the cells via LAT1, they were hydrolyzed into leucine by intracellular esterases and subsequently activated mTORC1, although we cannot rule out the idea that the esters themselves activate mTORC1. In fact, H-Leu-CH2OMe and H-Leu-NHOH, which are not the substrates of esterases, seemed to increase p70S6K phosphorylation slightly although it was not statistically significant (Fig. 6b), which let us speculate that a negative charge at the carboxyl group may not necessarily be essential in step 2. Further studies using the stable derivatives that would activate mTOR but not dissociate at carboxyl group should be considered in the future.
The proteins or other molecules called “leucine sensor” that recognizes leucine and links to mTORC1 signaling pathway have been proposed to be located at extracellular surface and/or intracellular compartment (Shigemitsu et al. 1999; Lynch et al. 2000; Beugnet et al. 2003; Kanazawa et al. 2004; Gulati and Thomas 2007). In our study, we propose that leucine sensor is located inside the cells at least in HeLa S3 cells, because leucine derivatives which are not transported by LAT1 did not activate mTORC1. LAT1 itself is not likely a leucine sensor, because the structural requirement for mTORC1 activation is much more rigorous than that for the recognition by LAT1. We propose that the requirement for above-described step 2 is that for the agonists of leucine sensor.
Recently, leucyl-tRNA synthetase (LRS) has been identified as a candidate of the intracellular leucine sensor (Han et al. 2012). The crystal structures of the complex of LRS and leucyl-adenylate analog demonstrated that the substrate binding site interacts with carbonyl oxygen and amino group of the substrate via hydrogen bonds, whereas the information on the alkoxy oxygen was missing due to the substitution of alkoxy oxygen to nitrogen atom in leucyl-adenylate analog (Cusack et al. 2000; Lincecum et al. 2003). Moreover, the mechanisms of recognition of leucine side chain by LRS have not been well understood (Han et al. 2012). Further study on the ligand recognition by LRS is required to determine whether the structure–activity relationship for the activation of mTORC1 obtained in the present study is due to the interaction of ligands with LRS.
In conclusion, we have shown that l-leucine is transported into the cells by LAT1 to activate mTORC1. We have revealed the important moieties of leucine for the interaction with LAT1 and for the activation of mTORC1. Because leucine-sensing mTOR signaling pathway controls cell growth and promotes cancer progression, LAT1-mTOR axis can be an appealing target for cancer therapy. Our study may provide a new insight into the novel anti-tumor agents that target both LAT1 and leucine sensor.
References
Bar-Peled L, Schweitzer LD, Zoncu R, Sabatini DM (2012) Ragulator is a GEF for the rag GTPases that signal amino acid levels to mTORC1. Cell 150(6):1196–1208
Beugnet A, Tee AR, Taylor PM, Proud CG (2003) Regulation of targets of mTOR (mammalian target of rapamycin) signalling by intracellular amino acid availability. Biochem J 372(Pt 2):555–566
Bröer S (2008) Amino acid transport across mammalian intestinal and renal epithelia. Physiol Rev 88(1):249–286
Brown EJ, Albers MW, Shin TB et al (1994) A mammalian protein targeted by G1-arresting rapamycin-receptor complex. Nature 369(6483):756–758
Brown EJ, Beal PA, Keith CT et al (1995) Control of p70 s6 kinase by kinase activity of FRAP in vivo. Nature 377(6548):441–446
Brunn GJ, Hudson CC, Sekulic A et al (1997) Phosphorylation of the translational repressor PHAS-I by the mammalian target of rapamycin. Science 277(5322):99–101
Christensen HN (1990) Role of amino acid transport and countertransport in nutrition and metabolism. Physiol Rev 70(1):43–77
Corradetti MN, Guan KL (2006) Upstream of the mammalian target of rapamycin: do all roads pass through mTOR? Oncogene 25(48):6347–6360
Cusack S, Yaremchuk A, Tukalo M (2000) The 2 Angstrom crystal structure of leucyl-tRNA synthetase and its complex with a leucyl-adenylate analogue. EMBO J 19(10):2351–2361
Dann SG, Thomas G (2006) The amino acid sensitive TOR pathway from yeast to mammals. FEBS Lett 580(12):2821–2829
Fort J, de la Ballina LR, Burghardt HE et al (2007) The structure of human 4F2hc ectodomain provides a model for homodimerization and electrostatic interaction with plasma membrane. J Biol Chem 282(43):31444–31452
Fuchs BC, Bode BP (2005) Amino acid transporters ASCT2 and LAT1 in cancer: partners in crime? Semin Cancer Biol 15(4):254–266
Gao X, Zhou L, Jiao X et al (2010) Mechanism of substrate recognition and transport by an amino acid antiporter. Nature 463(7282):828–832
Geier EG, Schlessinger A, Fan H et al (2013) Structure-based ligand discovery for the large-neutral amino acid transporter 1, LAT-1. Proc Natl Acad Sci USA 110(14):5480–5485
Gulati P, Thomas G (2007) Nutrient sensing in the mTOR/S6K1 signalling pathway. Biochem Soc Trans 35(Pt 2):236–238
Han JM, Jeong SJ, Park MC et al (2012) Leucyl-tRNA synthetase is an intracellular leucine sensor for the mTORC1-signaling pathway. Cell 149(2):410–424
Iijima Y, Laser M, Shiraishi H et al (2002) c-Raf/MEK/ERK pathway controls protein kinase C-mediated p70S6K activation in adult cardiac muscle cells. J Biol Chem 277(25):23065–23075
Jewell JL, Guan KL (2013) Nutrient signaling to mTOR and cell growth. Trends Biochem Sci 38(5):233–242
Jewell JL, Russell RC, Guan KL (2013) Amino acid signalling upstream of mTOR. Nat Rev Mol Cell Biol 14(3):133–139
Kaira K, Oriuchi N, Imai H et al (2009) L-type amino acid transporter 1 (LAT1) is frequently expressed in thymic carcinomas but is absent in thymomas. J Surg Oncol 99(7):433–438
Kaira K, Sunose Y, Arakawa K et al (2012) Prognostic significance of L-type amino-acid transporter 1 expression in surgically resected pancreatic cancer. Br J Cancer 107(4):632–638
Kanai Y, Segawa H, Miyamoto K et al (1998) Expression cloning and characterization of a transporter for large neutral amino acids activated by the heavy chain of 4F2 antigen (CD98). J Biol Chem 273(37):23629–23632
Kanazawa T, Taneike I, Akaishi R et al (2004) Amino acids and insulin control autophagic proteolysis through different signaling pathways in relation to mTOR in isolated rat hepatocytes. J Biol Chem 279(9):8452–8459
Khunweeraphong N, Nagamori S, Wiriyasermkul P et al (2012) Establishment of stable cell lines with high expression of heterodimers of human 4F2hc and human amino acid transporter LAT1 or LAT2 and delineation of their differential interaction with alpha-alkyl moieties. J Pharmacol Sci 119(4):368–380
Kim DK, Kanai Y, Choi HW et al (2002) Characterization of the system L amino acid transporter in T24 human bladder carcinoma cells. Biochim Biophys Acta 1565(1):112–121
Kim E, Goraksha-Hicks P, Li L et al (2008) Regulation of TORC1 by Rag GTPases in nutrient response. Nat Cell Biol 10(8):935–945
Kuma A, Hatano M, Matsui M et al (2004) The role of autophagy during the early neonatal starvation period. Nature 432(7020):1032–1036
Lehman JA, Gomez-Cambronero J (2002) Molecular crosstalk between p70S6K and MAPK cell signaling pathways. Biochem Biophys Res Commun 293(1):463–469
Li F, Yin Y, Tan B et al (2011) Leucine nutrition in animals and humans: mTOR signaling and beyond. Amino Acids 41(5):1185–1193
Lincecum TL Jr, Tukalo M, Yaremchuk A et al (2003) Structural and mechanistic basis of pre- and posttransfer editing by leucyl-tRNA synthetase. Mol Cell 11(4):951–963
Lynch CJ, Fox HL, Vary TC et al (2000) Regulation of amino acid-sensitive TOR signaling by leucine analogues in adipocytes. J Cell Biochem 77(2):234–251
Morimoto E, Kanai Y, Kim DK et al (2008) Establishment and characterization of mammalian cell lines stably expressing human L-type amino acid transporters. J Pharmacol Sci 108(4):505–516
Mortimore GE, Poso AR (1987) Intracellular protein catabolism and its control during nutrient deprivation and supply. Annu Rev Nutr 7:539–564
Nicklin P, Bergman P, Zhang B et al (2009) Bidirectional transport of amino acids regulates mTOR and autophagy. Cell 136(3):521–534
Sabatini DM (2006) mTOR and cancer: insights into a complex relationship. Nat Rev Cancer 6(9):729–734
Sabatini DM, Erdjument-Bromage H, Lui M et al (1994) RAFT1: a mammalian protein that binds to FKBP12 in a rapamycin-dependent fashion and is homologous to yeast TORs. Cell 78(1):35–43
Sancak Y, Peterson TR, Shaul YD, Lindquist RA et al (2008) The Rag GTPases bind raptor and mediate amino acid signaling to mTORC1. Science 320(5882):1496–1501
Sancak Y, Bar-Peled L, Zoncu R et al (2010) Ragulator-Rag complex targets mTORC1 to the lysosomal surface and is necessary for its activation by amino acids. Cell 141(2):290–303
Shigemitsu K, Tsujishita Y, Miyake H et al (1999) Structural requirement of leucine for activation of p70 S6 kinase. FEBS Lett 447(2–3):303–306
Sinclair LV, Rolf J, Emslie E et al (2013) Control of amino acid-transport by antigen receptors coordinates the metabolic reprogramming essential for T cell differentiation. Nat Immunol 14(5):500–508
Stipanuk MH (2007) Leucine and protein synthesis: mTOR and beyond. Nutr Rev 65(3):122–129
Su TZ, Feng MR, Weber ML (2005) Mediation of highly concentrative uptake of pregabalin by L-type amino acid transport in Chinese hamster ovary and Caco-2 cells. J Pharmacol Exp Ther 313(3):1406–1415
Uchino H, Kanai Y, Kim DK et al (2002) Transport of amino acid-related compounds mediated by L-type amino acid transporter 1 (LAT1): insights into the mechanisms of substrate recognition. Mol Pharmacol 61(4):729–737
van Bree JB, Audus KL, Borchardt RT (1988) Carrier-mediated transport of baclofen across monolayers of bovine brain endothelial cells in primary culture. Pharm Res 5(6):369–371
Wiriyasermkul P, Nagamori S, Tominaga H et al (2012) Transport of 3-fluoro-l-alpha-methyl-tyrosine by tumor-upregulated L-type amino acid transporter 1: a cause of the tumor uptake in PET. J Nucl Med 53(8):1253–1261
Wu G (2009) Amino acids: metabolism, functions, and nutrition. Amino Acids 37(1):1–17
Wullschleger S, Loewith R, Hall MN (2006) TOR signaling in growth and metabolism. Cell 124(3):471–484
Xu G, Kwon G, Marshall CA et al (1998) Branched-chain amino acids are essential in the regulation of PHAS-I and p70 S6 kinase by pancreatic beta-cells. A possible role in protein translation and mitogenic signaling. J Biol Chem 273(43):28178–28184
Yamashita A, Singh SK, Kawate T et al (2005) Crystal structure of a bacterial homologue of Na+/Cl− dependent neurotransmitter transporters. Nature 437(7056):215–223
Yamauchi K, Sakurai H, Kimura T et al (2009) System L amino acid transporter inhibitor enhances anti-tumor activity of cisplatin in a head and neck squamous cell carcinoma cell line. Cancer Lett 276(1):95–101
Yanagida O, Kanai Y, Chairoungdua A et al (2001) Human L-type amino acid transporter 1 (LAT1): characterization of function and expression in tumor cell lines. Biochim Biophys Acta 1514(2):291–302
Zoncu R, Efeyan A, Sabatini DM (2011) mTOR: from growth signal integration to cancer, diabetes and ageing. Nat Rev Mol Cell Biol 12(1):21–35
Acknowledgments
The authors would like to thank Yasuhiro M. Umemura for his valuable advice and Yoko Tanaka for her technical assistance. This work was supported in part by Grant-in-Aid for Scientific Research and Grant-in-Aid for Young Scientists from Japan Society for the Promotion of Science (JSPS), by Regional Innovation Strategy Support Program, Grant-in-Aid for the Scientific Research on Innovative Areas “HD Physiology”, and Grant-in-Aid for Scientific Research on Priority Areas “Transportsome” from Ministry of Education, Culture, Sports, Science and Technology of Japan (MEXT), and by grants from the Osaka Medical Research Foundation for Intractable Diseases, the Advanced Research for Medical Products Mining Program of the National Institute of Biomedical Innovation, and the Ajinomoto Amino Acid Research Program.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Additional information
Handling Editor: E. I. Closs.
S. Nagamori and P. Wiriyasermkul contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Nagamori, S., Wiriyasermkul, P., Okuda, S. et al. Structure–activity relations of leucine derivatives reveal critical moieties for cellular uptake and activation of mTORC1-mediated signaling. Amino Acids 48, 1045–1058 (2016). https://doi.org/10.1007/s00726-015-2158-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00726-015-2158-z