Abstract
Knowledge of protein flexibility is vital for deciphering the corresponding functional mechanisms. This knowledge would help, for instance, in improving computational drug design and refinement in homology-based modeling. We propose a new predictor of the residue flexibility, which is expressed by B-factors, from protein chains that use local (in the chain) predicted (or native) relative solvent accessibility (RSA) and custom-derived amino acid (AA) alphabets. Our predictor is implemented as a two-stage linear regression model that uses RSA-based space in a local sequence window in the first stage and a reduced AA pair-based space in the second stage as the inputs. This method is easy to comprehend explicit linear form in both stages. Particle swarm optimization was used to find an optimal reduced AA alphabet to simplify the input space and improve the prediction performance. The average correlation coefficients between the native and predicted B-factors measured on a large benchmark dataset are improved from 0.65 to 0.67 when using the native RSA values and from 0.55 to 0.57 when using the predicted RSA values. Blind tests that were performed on two independent datasets show consistent improvements in the average correlation coefficients by a modest value of 0.02 for both native and predicted RSA-based predictions.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Protein flexibility is essential for function, allowing structural rearrangements in the allosteric regulation (del Sol et al. 2009). It is also believed to play an important role in protein structure evolution (Tokuriki and Tawfik 2009), enzyme catalysis (Eisenmesser et al. 2005), molecular recognition, allosteric regulation, and antigen–antibody interactions (Dodson and Verma 2006). An accurate knowledge of flexibility would be desirable in the context of building more accurate computational drug design platforms through utilization of flexible protein–ligand docking (B-Rao et al. 2009) and for refining homology modeling-based predictions of protein structures (Han et al. 2008).
The structural data that are deposited in the Protein Data Bank (PDB) (Berman et al. 2000) are mostly solved with X-ray crystallography and provide information on the atomic mobility. The B-factor, also known as Debye–Waller temperature factor or atomic displacement parameter, measures the atomic flexibility of the crystallographic structures (Halle 2002). This parameter can be used to measure structural flexibility of proteins and hence it provides very useful information about the protein dynamics and insights to further stimulate in-depth functional studies (Yuan et al. 2003; Schnell et al. 2004; Yang and Bahar 2005; Fontana et al. 2008; Kwansa and Freeman 2010). B-factor was also broadly investigated from a variety of viewpoints including the relations between mobility and thermal stability (Vihinen 1987) and between domain folding and flexibility (Liu and Rost 2004; Díaz-Espinoza et al. 2007; Mackereth and Sattler 2012), in the context of applications in the prediction of active sites and binding sites (Gutteridge et al. 2003; Neuvirth et al. 2004; Han et al. 2012), identification of protein pockets and cavities (Panjkovich and Daura 2010), and determination of folding rates (Gao et al. 2010). It was also utilized as a benchmark to evaluate computational modeling of flexibility including molecular dynamic simulation (Scheraga et al. 2007), normal mode analysis (Tozzini 2005), Gaussian network model (GNM) (Liu and Karimi 2007), and machine learning-based prediction models (Yuan et al. 2005; Schlessinger and Rost 2005; Pan and Shen 2009; Zhang et al. 2009).
The relative solvent accessibility (RSA), which is defined by the solvent-accessible surface area (ASA) of a residue in the protein structure divided by the ASA observed in an extended conformation (Gly-X-Gly or Ala-X-Ala) (Ahmad et al. 2003), was previously shown to be correlated with the flexibility expressed with B-factors (Zhang et al. 2009). More specifically, a linear regression was applied to model the relation between B-factors and RSA values using a sliding window. This simple model was shown to outperform other existing models which used more inputs and more complex predictors such as support vector machines (Yuan et al. 2005), neural networks (Schlessinger and Rost 2005) and GNMs (Liu and Karimi 2007) to predict B-factors. This suggests that usage of RSA alone provides a strong predictive input for prediction of B-factor values. However, the average correlation coefficients (ACC) between the actual and predicted B-factors, which were around 0.65 for the structure-based methods (where RSA was extracted from structure) and 0.55 for the sequence-based modeling (where RSA was predicted from the sequence) (Zhang et al. 2009), remain distant from the upper bound of 0.80 estimated by Radivojac et al. (2004). In this study, we investigate whether the existing RSA-based linear model that uses RSA (Zhang et al. 2009) can be further improved by introducing other information, such as amino acid (AA) specificity. We hypothesize that besides the previously considered inclusion of the local RSA (with respect to the sequence) that improves prediction of the residue flexibility, consideration of the AA types may also contribute to the improved prediction of the residue flexibility. To this end, we propose a two-stage prediction method by embedding an approach based on the specificity of AA pairs/dipeptides into the original RSA-based linear model. Our method utilizes linear regression in both stages to assure that the underlying model is simple and easy to comprehend, which in turn reduces possibility of overfitting this model into the training dataset. We consider the structure-based prediction that uses the native/actual RSA as the input and the sequence-based prediction using RSA that is predicted from the sequence as the input; the former can be used to estimate an upper bound of the prediction quality of the sequence-based predictor and can be used to compare with other existing structure-based methods.
The number of possible AA pairs is rather large and equal to 400, which results in relatively high dimensionality of inputs of the second stage. Therefore, we reduce this dimensionality by grouping “related” AA together. Similar AA alphabet reduction schemes have been applied in related areas including prediction of intrinsically disordered proteins (Weathers et al. 2004), prediction of subcellular localization (Oğul and Mumcuoğu 2007), fold assignment (Peterson et al. 2009), GPCR classification (Davies et al. 2008) and prediction of secretory proteins (Zuo and Li 2010), to name but a few, and resulted in improved prediction quality. We used particle swarm optimization (PSO) (Kennedy and Eberhart 1995), a widely used population-based algorithm, to perform this reduction with the aim of improving the prediction performance.
Materials and methods
B-factor, solvent accessibility, secondary structure and datasets
B-factor
Experimental B-factor of an atom is defined as \(8\pi\, {<}u^{2}{>}\)using the isotropic mean square displacement, \(u^{2}\), averaged over the lattice (Schlessinger and Rost 2005). In this study, the B-factor value of Cα atom for a residue was used to represent the residue flexibility. Since B-factor values depend on the experimental resolution, crystal contacts, and refinement procedures, they are normalized to allow comparisons between different structures. Following the approach used in (Parthasarathy and Murthy 1997; Schlessinger and Rost 2005; Zhang et al. 2009), B-factors of the Cα atoms in a given chain that were extracted from PDB files (Berman et al. 2000) are normalized using:
where B is the raw B-factor, \(\bar{B}\) is the average B-factor, and \(\sigma\) is the standard deviation of B-factors for all Cα atoms in a given chain.
Solvent accessibility and secondary structure
Several methods were developed for the prediction of RSA (Ahmad et al. 2003; Yuan and Huang 2004; Wang et al. 2007). The proposed two-stage linear models use the RSA values of residues in a local window as inputs. The native ASA values were computed with the DSSP program (Kabsch and Sander 1983). The sequence-based prediction of the ASA values was performed using SPINE-X; this choice was motivated by high quality of predictions generated by this method (Faraggi et al. 2012). The RSA value of a residue is its ASA divided by the ASA observed in an extended conformation (Ahmad et al. 2003). We also used DSSP program to compute the three state secondary structure (SS), i.e., helix (H), strand (E), coil (C), for the considered proteins. These results were used to analyze relation between predictive performance of residue flexibility in the context of the underlying secondary structures.
Datasets
We developed three datasets to design and comparatively evaluate the predictive performance of our proposed models. The first dataset is a subset of protein chains that were solved by X-ray crystallography and were deposited to the Protein Data Bank (PDB) (Berman et al. 2000) between March 2011 and March 2012. The entire corresponding protein set was clustered with the NCBI’s BLASTCLUST (Altschul et al. 1997) at 25 % sequence identity. The first dataset, named PDB632, includes 632 protein chains with local 25 % pairwise sequence identity derived by selecting one chain with length ≥60 in each cluster. PDB632 set was used to design our predictive models and to study the relation between the solvent accessibility and the flexibility. The limits on the protein deposition date are imposed to minimize overlap and similarity with the training set used by the SPINE-X program that is used to predict the RSA values.
The other two dataset are independent of the PDB632 dataset and are used to perform blind tests. The first dataset is based on sequences that were solved by X-ray crystallography and were deposited in PDB between April 2012 and December 2012. Similarly as in the case of the PDB632 dataset, BLASTCLUST (Altschul et al. 1997) was applied to the union of this set and PDB632 with the local identity threshold at 25 %. The new blind set was constructed by selecting one chain of length ≥60 from each cluster that did not contain sequences from the PDB632 dataset. This set, called PDB704, includes 704 chains that, as a result, have local 25 % identity with each other and also with the chains from the PDB632 dataset. The second independent dataset was created utilizing the same procedure, however, it includes subset of protein sequences that were deposited in PDB between January 2013 and June 2013. Therefore, proteins in this set also share low identity (<25 %) with the sequences used in the SPINE-X program and in the PDB632 dataset. More specifically, BLASTCLUST was applied to the union of the set deposited in PDB between January 2013 and June 2013, PDB632 and the training set used in SPINE-X program with the local identity threshold at 25 %. The second blind set, called PDB208, has 208 sequences that were obtained by selecting one chain of length ≥60 from each corresponding cluster that has no chains from the PDB632 dataset and the training set used in SPINE-X.
PDB704 and PDB208 were used for blind tests to validate and comparatively evaluate our proposed two-stage linear models that were trained on the PDB632 dataset. The low identity between sequences from the PDB632 and from PDB704 and PDB208 datasets allows for an unbiased (by the similarity to the training datasets) evaluation.
When preparing the data to test the models, the normalization of B-factors and the computation/prediction of the ASA values using DSSP/SPINE-X programs were performed for each chain in the PDB632, PDB704 and PDB208 datasets. For the SPINE-X program, the authors reported correlation coefficient of 0.74 between the predicted and the actual RSA values on its training set (Faraggi et al. 2009, 2012). To compare, the RSA predictions with SPINE-X on the PDB632, PDB704 and PDB208 sets yielded correlation coefficients of 0.71, 0.69 and 0.69, respectively, and thus we believe that the SPINE-X and the proposed models that utilize its predictions do not overfit these three datasets. The PDB IDs of the PDB632, PDB704 and PDB208 datasets are listed in the Supplementary Material as Table S4, Table S5 and Table S6, respectively.
Two-stage RSA-based linear models for residue flexibility modeling
We used linear regression to investigate the relation between the RSA and the residue flexibility expressed using the normalized B-factor. We created models over a local window in the protein sequence as the first stage of the prediction for B′-factor (the normalized B-factor). Similar design has been used in our previous work (Zhang et al. 2009). The model is defined as:
where b is the intercept and \(\hat{B}_{i}^{'}\) represents the estimated (predicted) B′-factor of the central residue i using RSA values in the window size of h = 0, 1, 2,…, (the window includes 2h + 1 residues), and where weights w k are determined using the least square fit between the predicted B′-factor and the actual B′-factor values.
In the first stage of our predictive model, the window size of 9 (i.e., h = 4) was selected to predict the B′-factors. The optimization procedure for the window size and the corresponding weights w k for the optimized h value are discussed in our previous work (Zhang et al. 2009). In the linear model expressed in Eq. (2), RSA i corresponds to either the actual RSA derived with DSSP (denoted by DsspRSA i ) or the RSA predicted using SPINE-X (denoted by PredRSA i ).
From Eq. (2), the feature vector with the window size of 9 for residue i is (RSA i−4, RSA i−3, RSA i−2, RSA i−1, RSA i , RSA i+1, RSA i+2, RSA i+3, RSA i+4). This vector does not contain information about AA properties, i.e., the model processes all AA types in the same manner. Therefore, we designed the second-stage linear model, which considers AA types, as follows:
where f A(i+k),A(i) corresponds to a given AA pair (A(i + k), A(i)) and measures relation of this specific pair with the flexibility of the central residue i, and A(i) denotes the corresponding AA type of residue i. Given the AA alphabet AA20 = {A, R, N, D, C, Q, E, G, H, I, L, K, M, F, P, S, T, W, Y, V}, then we consider 400 (=20 × 20) factors f p,q (p, q ∈ AA20) that are estimated from the training dataset PDB632. The weights {w k } reflect the positional (in the sequence window) contributions, while the factors {f p,q } measure the contribution of the AA pair specificity, to the flexibility of the central (in the window) residue. If both the weights and the factors are unknown, however, the model shown in Eq. (3) is not linear and thus they cannot be simultaneously estimated by the least square fit. Our approach to solve this problem is that we first estimate the weights shown in (2), which corresponds to the first-stage linear model, and then we learn the factors {f p,q } when weights {w k } are given and inputted into the model (3). However, for two different positions i + k1 and i + k2 in the same local window of the central residue i, it is possible that pairs (A(i + k1), A(i)) and (A(i + k2), A(i)) (k1, k2 ∈{−4, −3, −2, −1, 0, 1, 2, 3, 4} and k1 ≠ k2) are identical, i.e., f A(i+k1), A(i) = f A(i+k2), A(i). Thus, the factors cannot be estimated from the form shown in (3). The model in (3) was then transformed to the following new linear form:
where X p,q is the sum of all w k · RSA i+k values associated with the same item f p,q . The Eq. (4) has linear form that could be modeled using the least square fit. In addition, for unknown factors (f p,q ) p,q∈AA20, the feature vector is (X p,q ) p,q ∈AA20 with the dimensionality of 400. Figure 1 shows how we estimate the B′-factor for the central residue ‘A’ using a sliding window for a given local subsequence ‘AREDATRAN’. This estimation is expressed and simplified as follows:
Thus, for the feature vector (X p,q ) p,q∈AA20, we have X AA = w −4·RSA i−4 + w 0·RSA i + w 3·RSA i+3, X RA = w −3·RSA i−3 + w 2·RSA i+2, X EA = w −2·RSA i−2, X DA = w −1·RSA i−1, X TA = w 1·RSA i+1, X NA = w 4·RSA i+4, and others Xp, q = 0.
If the central residue is close to the C-terminus or N-terminus of the sequence, the sliding window of this residue may extend beyond the sequence. We pad the sequence with the gap symbol ‘-’ that represents a virtual AA and that is used to extend the window beyond the sequence termini. Thus, the considered AA alphabet is extended to AA21 = AA20∪{-}. The corresponding factor vector in (4) is (f p,q ) p,q∈AA21 based on the feature vector (X p,q ) p,q ∈AA21 that has 441 dimensions.
Inspired by the finding that use of the reduced alphabet improves the fold assignment (Peterson et al. 2009) and GPCR classification (Davies et al. 2008), we hypothesize that an optimized AA alphabet AAn with n groups (1 < n < 21) obtained by reducing AA21 may also result in an increase of the prediction performance for the prediction of the residue flexibility. When using AAn as the AA alphabet, the factor vector in (4) is (f p,q ) p,q∈AAn and is associated with the feature vector (X p,q ) p,q ∈AAn with dimensionality of n × n. The computations of the weights in both the first-stage and the second-stage linear models were performed using the scikit-learn method (Pedregosa et al. 2011).
Reduction of AA alphabet
Particle swarm optimization (PSO) has been successfully applied in several areas such as image processing (Niu and Shen 2006), parameter optimization (Meissner et al. 2006), and Quantitative Structure–Activity Relationship (QSAR) modeling (Lin et al. 2005). Each particle in PSO is initialized at a random position in a given search space. The position of particle i is given by a vector x i = (x i1, x i2, …, x iD ), where D is the dimensionality of the problem. Velocity of a given particle is represented by the vector v i = (v i1, v i2, …, v iD ). PSO is an iterative algorithm in which the best position of the ith particle in previous iteration t is denoted by p i = (p i1, p i2, …, p iD ), and the best particle among all particles in the population is represented as p g = (p g1, p g2, …, p gD ). The “goodness” of a given particle is evaluated by the fitness function defined in the Sect. 2.4. The particle updates its velocity and position according to Eqs. (5) and (6), respectively:
where d is the dth dimension of a particle, w is the inertia weight, c 1 and c 2 are two positive constants called learning factors, and r 1 and r 2 are random numbers in the (0,1) range (Kennedy and Eberhart 1995).
The PSO is used to optimize the AA grouping, i.e., to reduce the AA alphabet, to improve the prediction of residue flexibility. Here, the dimensionality of a particle equals to 21 (i.e., D = 21), where x ij in the position vector x i = (x i1, x i2, …, x iD ) that represents the group index of the jth amino acid in AA21. The value of x ij ranges from 1 to the maximum group number m (2 ≤ m ≤ 20). Since the PSO algorithm is implemented in a continuous search space, the value of x ij (1 ≤ j ≤ 21) is a real number. We use a rule that if n − 0.5 ≤ x i j < n + 0.5 for a positive integer n ≤ m, then the jth amino acid in AA21 should be grouped into the nth group of AA21. For example, if m = 5 and the position vector of particle i is (3.66, 3.4, 5.0, 2.28, 4.6, 3.8, 1.2, 3.3, 4.0, 3.7, 1.1, 2.88, 4.52, 2.76, 4.34, 5.2, 2.12, 3.32, 4.21, 2.23, 3.56), then the corresponding vector of category indices for AA21 is (4, 3, 5, 2, 5, 4, 1, 3, 4, 4, 1, 3, 5, 3, 4, 5, 2, 3, 4, 2, 4), which implies a reduced alphabet AA5 = {EL, DTV, RGKFW, AQHIPY-, NCMS} by grouping AA21. The reduced alphabet is applied to the Eq. (4) based on AA5 instead of AA20, resulting in the factor vector (f p,q ) p,q∈AA5 and the feature vector (X p,q ) p,q ∈AA5, both having dimensionality of 5 × 5.
The parameters of the PSO-based optimizer were set as follows: the inertia weight w = 0.8, the learning factors c 1 = 2 and c 2 = 2, the population size of particles NP = 20, the maximum group number m = {2, 3,…, 20}, the range of each element in the position vector x ij = (1, m), and the maximum number of iterations Iter = 20.
Fitness function and performance evaluation
A number of studies (Kurgan et al. 2008) were focused on the real-value predictions of various protein descriptors including solvent exposure expressed as RSA (Ahmad et al. 2003; Wang et al. 2007) and residue depth (Zhang et al. 2008), and residue flexibility expressed as B-factor (Yuan et al. 2005). For such real-value prediction, Pearson correlation coefficient (CC) is usually used to evaluate the predictive performance. The other commonly used criterion is the mean absolute error, but due to the normalization of the raw B-factor values, the measure cannot be used to evaluate the quality of the flexibility predictors. The CC is defined as
where x i is the observed B′-factor and y i is the predicted B′-factor for the ith residue in the sequence. If CC is close to 1, then {x i } and {y i } are fully correlated. If CC is close to 0 then the two variables are not correlated, and in the case when CC is close to −1 then the variables are anticorrelated. The absolute CC values quantify the degree of the correlation.
Similarly as in our previous work (Zhang et al. 2009), the correlation is measured at the protein chain level. The CC value is computed for each chain separately and next these values are averaged to compute the correlation over a given dataset. We use the term average correlation coefficient (ACC) to refer to the CC at the chain level. To evaluate the ability of the proposed models to generalize to blind datasets, we performed fivefold cross validation on the training dataset PDB632.
To reduce the AA21 using the PSO methods, we used a fitness function to assess the performance of each particle. The position vector of a particle was converted to a reduced AA alphabet and the ACC derived from the fivefold cross validation on the PDB632 dataset under the reduced alphabet was used to define the fitness function of the particle.
Results
We used the PDB632 dataset to design and compute the two-stage linear models with the PSO-optimized AA grouping. All computations, except when testing the model on the blind/independent datasets, were performed based on the fivefold cross validation on the training dataset PDB632. The ACC values were used to evaluate the prediction quality and to determine the fitness of a particle in the PSO algorithm. This procedure was applied to both DsspRSA-based and PredRSA-based linear models, which represent the structure-based and sequence-based prediction of the residue flexibility, respectively. When testing on the independent PDB704 and PDB208 dataset, the prediction model was computed using the entire PDB632 dataset.
Results for the optimized reduced amino acid alphabets
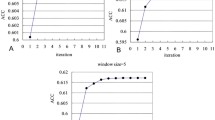
The results of the search for optimal (resulting in the highest ACC based on the cross validation on the PDB632 dataset) AA grouping using PSO with varying maximum group numbers (m) are shown in Supplementary Material as Table S1 for the prediction using native RSA and in Table S2 for the prediction using predicted RSA. Of all possible m values, the highest ACC for the prediction with the native RSA on the PDB632 dataset is 0.665 and is associated with several values of m, i.e., m = 12, 14, 16, 17, 19 and 20, as shown in Table S1. The actual sizes of the reduced alphabets could be based on any number in {1, 2, …, m} that should be smaller than or equal to m. Smaller size of the reduced alphabet would lead to a simpler prediction model. Therefore, when m = 20 with the highest ACC equal to 0.665 when utilizing for the native RSA, the PSO optimizer generated a reduced alphabet with the smallest size of 9. This alphabet is {NQEHKPS, L, WY, R, -, I, G, V, ADCMFT}. Similarly, when considering prediction using the RSA values (i.e., when predicting from the sequence), the smallest size of the reduced alphabet is 7 with the corresponding highest ACC of 0.572 when m = 19. This alphabet is {CILKMSV, Y, -, FW, G, R, ANDQEHPT}. These two reduced alphabets were used to build the models in the second stage and were evaluated on the blind PDB704 and PDB208 datasets.
PSO is an optimization algorithm that introduces randomness when initializing and updating the particles. As a result, the “optimal” reduced alphabets are not unique, depending on these initial conditions. However, multiple experiments with the varying maximum group number (m) show certain trends concerning the resulting AA groupings. As seen in Table S1 and Table S2, several AAs (including ‘-’) demonstrate a more consistent tendency to group as they are inclined to belong to a certain group that includes only one or two AAs. Moreover, Fig. 2 shows the count distribution of each AA that was clustered belonging to an individual or two-symbol group. The result that stands out is the terminal gap symbol ‘-’, which is expected since the terminal residues tend to be more flexible. When compared with the other AAs, four AAs including Glycine (G), Phenylalanine (F), Tyrosine (W) and Tryptophan (Y) usually belong to a group that contains only one or two AAs for both the DsspRSA- and PredRSA-based models. This is possibly due to the fact that Glycine has the smallest side chain with the lowest molecular weight, while Phenylalanine, Tyrosine and Tryptophan have aromatic rings with the largest molecular weights.
When using AA21 without the alphabet reduction, the ACC values between the native and predicted B′-factors evaluated based on fivefold cross validation for the two-stage linear models are 0.661 and 0.568 for the structure-based and sequence-based predictions, respectively. The proposed method that uses optimized reduced AA alphabet improves ACC values when compared with the method without the reduction for AA alphabet. Although the improvement is relatively small, the reduction of the AA alphabet also decreases the dimensionality of the linear model in the second stage, resulting in a simplified predictive model.
Comparison of the two-stage linear models with sliding window-based methods
The literature includes a wide variety of sequence-based predictors of various structural aspects of residues in proteins including secondary structures (Jones 1999; Zhang et al. 2011), contact maps (Cheng and Baldi 2007; Tegge et al. 2009), domain boundaries (Liu and Rost 2004; Li et al. 2012), solvent accessibility (Nguyen and Rajapakse 2006; Chen et al. 2008; Faraggi et al. 2012), B-factors (Yuan et al. 2005), and disordered regions (Jin and Dunbrack 2005; Mizianty et al. 2010; Zhang et al. 2012b). Most of these methods encode the input features using a sliding window where the predictions are preformed for the central residue. Each position in the sliding window is usually characterized by a number of features, such as AA type and evolutionary information extracted from the multiple alignment profiles. Thus, we compare the proposed models with the sliding window-based approaches. These approaches are implemented using linear regression and they include DsspRSA9, DsspRSA9 + AA9, DsspRSA9 + AA9 + PSSM9 when considering the structure-based (based on native RSA values) methods; and PredRSA9, PredRSA9 + AA9, PredRSA9 + AA9 + PSSM9 for the sequence-based methods (that use RSA values predicted from sequence); note that RSA denotes relative solvent accessibility, AA represents amino acid type, PSSM is the position specific scoring matrix generated by PSI-BLAST (Altschul et al. 1997), and 9 is the window size. For the AA type, each position in the sliding window is coded by a 20-dimension vector with value of 1 for one element where the residue type equals to a given AA type and otherwise 0. The PSSM for each residue in the sliding window is a 20-dimension vector where the score values are normalized with commonly utilized logistic function 1/(1 + exp(−x)). We use “RSA9” to investigate both structure-based and sequence-based predictions. The results are summarized in Fig. 3 and Table 1.
Comparison of the ACC values between the one-stage and the two-stage linear models based on fivefold cross validation on the PDB632 dataset. The x axis shows the two types of predictions where inputs are either structure-based using the actual RSA values or sequence-based using the predicted RSA values. The CC values are averaged over the folds of the cross validation and the corresponding standard deviations are shown using error bars
Figure 3 plots the ACC values of the two-stage linear models based on the fivefold cross validation on the PDB632 dataset together with standard deviations calculated over the fivefold shown as the error bars. We observe that the first-stage models (i.e., DsspRSA9 and PredRSA9) achieve ACCs of 0.65 and 0.55 for the structure-based and the sequence-based methods, respectively. The results of the proposed models that include the second stage provide modest but consistent (with low error bars) improvements in ACC values by 0.02 for both the structure-based and sequence-based methods.
Table 1 compares the prediction quality of the two-stage linear models and the sliding window-based methods based on the fivefold cross validations on the PDB632 dataset. The CC values between the native and the predicted B-factors are given for each fold and are averaged over the fivefold. By adding the features coded by AA type, as shown in the table, the sliding window-based method RSA9 + AA9 achieved an improvement in ACC by 0.01 when compared with the RSA9 model. However, no significant improvement is observed by adding other features including PSSM, i.e., the RSA9 + AA9 + PSSM9 model does not show further improvement when compared with the RSA9 + AA9 model. This is probably since RSA values provide strong input for the prediction of the B-factors and since prediction of RSA by SPINE-X that we utilize already includes the PSSM values.
As shown in Table 1, the predictions of the proposed models provided consistent and modest improvement of the CC values for each fold when compared with the sliding window-based methods including RSA9, RSA9 + AA9 and RSA9 + AA9 + PSSM9. Consequently, the paired t tests performed at the 95 % significance level, which compare pairs of ACC values from the two-stage linear models and the sliding window-based methods over the fivefold of the cross-validation procedures, reveal that the differences are significant. The corresponding p values for both the structure-based and sequence-based cases are below 0.0001. To sum up, the proposed two-stage models provide statistically significantly better B-factor predictions with the modest magnitude of the improvements when compared with the sliding window-based methods.
Comparison of the two-stage linear models with other existing methods for prediction of B′-factor
Table 2 summarizes the prediction quality, measured based on the ACC between the native and the predicted B′-factor values, of several existing methods for prediction of B′-factor values. We include four methods that predict B′-factors from protein structures: GNM (Kundu et al. 2002); a parameter-free Gaussian network model (pfGNM) (Yang et al. 2009); weighted contact number (WCN) method (Lin et al. 2008) and our previous method DsspRSA9 (Zhang et al. 2009). These methods are compared against our corresponding two-stage linear model. The prediction with the GNM, pfGNM and WCN is described in (Zhang et al. 2012a). We report the result of our previous PredRSA9 method as a representative method that predicts B′-factors from the protein sequences. We note that the web servers for other existing sequence-based B′-factor predictors, such as the neural network approach from (Schlessinger and Rost 2005), support vector regression method from (Yuan et al. 2005) and the two-stage SVR from (Pan and Shen 2009), are either unavailable or do not work and thus we were unable to include their results. However, we include results generated by several predictors of disordered regions. Disordered regions can be perceived as regions with high flexibility and they are defined as regions that have no coordinates according to the ‘REMARK465’ in the corresponding PDB files. Predictions generated by these methods were shown to be correlated with the B-factor values (Radivojac et al. 2004; Jin and Dunbrack 2005; Worch and Stolarski 2008). We used the probability scores generated by four disorder predictors: IUPred (Dosztányi et al. 2005a, b) (for prediction of short disordered regions), SPINE-D (Zhang et al. 2012b), DisEMBL (Linding et al. 2003) and ESpritz (Walsh et al. 2012) to assess their correlation with the native B′-factor values. DisEMBL also provides probability scores for the predictions of two types of relatively flexible residues, i.e., hot loops and coils, for which the coordinates are provided. Coils are composed of residues with states T, S, B and I assigned by the DSSP program (Kabsch and Sander 1983). Hot loops are the coils with high B-factors. The three corresponding DisEMBL models are denoted as DisEMBL-Remark465, DisEMBL-coil and DisEMBL-hotloop, respectively. Similarly, ESpritz is an ensemble of protein disorder predictors trained on three different types of data, which were derived from X-ray structures in PDB, experimental data deposited in DisProt database (Sickmeier et al. 2007) and nuclear magnetic resonance (NMR) structures in PDB. These three ESpritz models are called ESpritz-X-ray, ESpritz-DisProt and ESpritz-NMR, respectively. We also include the DynaMine method that predicts backbone N–H S2 order parameter (\(S_{\text{RCI}}^{2}\)) values, which are estimated from chemical shifts. A value of 1.0 for the S 2 order parameter means complete order (stable conformation), whereas a value of 0.0 represents fully random bond vector movement (highly dynamic). We note that computation of the ACC values between the native B-factors and the predicted disorder probability values was done before (Jin and Dunbrack 2005; Zhang et al. 2009), while we are the first to examine the relation between the native B-factors and the predictions of hot loops, coils, DisProt disorder, NMR mobility and the S 2 order parameter.
The structure-based methods, including DsspRSA9, GNM, pfGNM and WCN yield ACC values of 0.65, 0.57, 0.63 and 0.62 on the PDB632 dataset, respectively, while the proposed method achieves ACC value of 0.67. Among the existing structure-based methods, DsspRSA9 provides the highest ACC value of 0.65. Moreover, blind tests on the PDB704 and PDB208 datasets when using our model trained on PDB632 dataset confirm the modest improvements offered by our structure-based predictor. Table 2 shows that the ACC value derived by the proposed two-stage linear model is 0.65 and 0.62 on the PDB704 and PDB208 datasets, respectively, when compared with the best considered existing method DsspRSA9 that obtains ACC of 0.63 and 0.60. The proposed method also outperforms GNM, pfGNM and WCN that have ACC values of 0.53, 0.60 and 0.59 for the PDB704 dataset and 0.51, 0.57 and 0.56 for the PDB208 dataset, respectively.
In the case of the sequence-based methods, the two-stage linear model provides the best result on the PDB632 dataset with ACC = 0.57, while the other methods sorted in the descending order by their absolute ACC values are PredRSA9 (ACC = 0.55), SPINE-D (0.38), DisEMBL-Remark465 (0.37), DynaMine (0.36), ESpritz-X-ray (0.32), DisEMBL-hotloop (0.32), ESpritz-NMR (0.31), IUPred (0.28), DisEMBL-coil (0.24), and ESpritz-DisProt (0.13). While performing the blind tests on the PDB704 and PDB208 datasets using our models trained on the PDB632 dataset, we again observe consistent improvements when compared with the second-best PredRSA9 method. The ACC values generated by our sequence-based approach are 0.56 and 0.54 on the PDB704 and PDB208 datasets, respectively, compared to 0.54 and 0.52 obtained by the PredRSA9 method. The blind tests on the PDB208 dataset for the sequence-based predictions result in smaller ACC values than those on the PDB704 and PDB632 datasets. This is also true for the structure-based methods. Therefore, the lower quality results on the PDB208 set when compared with the other two datasets are not influenced by the quality of the RSA predictions generated using the SPINE-X program, and may be due to the higher degree of difficulty of this dataset. To sum up, the proposed two-stage linear models provide consistent (over three datasets) improvements in ACC values by a modest margin of 0.02 when compared with the corresponding structure-and sequence-based one-stage linear models (i.e., DsspRSA9 and PredRSA9). In addition, as expected, the predictions of IUPred, SPINE-D, DisEMBL-Remark465, DisEMBL-coil, and DisEMBL-hotloop provide lower ACC values than those of the sequence-based two-stage linear model on the three datasets: PDB632, PDB704 and PDB208. This is due to the fact that IUPred, SPINE-D, ESpritz-X-ray, DisEMBL-Remark465, DisEMBL-coil, DisEMBL-hotloop, ESpritz-DisProt, ESpritz-NMR and DynaMine predict the disordered regions, coils, hot loops, DisProt disorder, NMR mobility and S2 order parameter, respectively, rather than the B′-factors. However, the ACC values that range between 0.3 and 0.4 that are achieved by IUPred, SPINE-D, ESpritz-X-ray, ESpritz-NMR, DisEMBL-Remark465 and DynaMine show a modest correlation between the predicted propensity of intrinsic disorder and the native B-factor values.
We performed paired t tests at the 95 % significance level, which compare pairs of ACC values for the same sequences predicted by the proposed two-stage linear model and each of the existing methods. The calculations were done separately for the structure-based and the sequence-based predictors. The resulting p values for both blind tests on the PDB704 and PDB208 datasets are below 0.0001, which implies that the proposed methods provide statistically significant improvements for the prediction of the B-factors, i.e., the improvements are consistent across different proteins although the corresponding magnitude could be modest.
Figure 4 directly compares results for individual proteins between the best-performing existing one-stage and the proposed two-stage models based on the ACC values obtained on the PDB704 and PDB208 datasets. The (proposed) two-stage models provide higher ACC values for majority of the predicted sequences when compared with the one-stage models, i.e., most of the points are located above the diagonal red line. More specifically, in the case of the PDB704 dataset, 511 out of 704 proteins for the structure-based models (Fig. 4a) and 473 out of 704 proteins for the sequence-based models (Fig. 4b) have higher ACC values for the proposed predictors. Similar findings are true for the PDB208 dataset where 164 out of 208 proteins for the structure-based models (Fig. 4c) and 143 out of 208 proteins for the sequence-based models (Fig. 4d) are above the diagonal. Furthermore, we performed paired t tests at the 95 % significance level to compare pairs of ACC values for the same sequences predicted by the one-stage and the proposed two-stage linear models. The p values for both blind tests on the PDB704 and PDB208 datasets are below 0.0001, which suggests that the differences between the one-stage and the proposed two-stage linear models are statistically significant.
Comparison of the ACC values at the sequence level between the one-stage and the two-stage linear models based on the blind tests on the PDB704 and PDB208 datasets using the models trained on the PDB632 dataset. a and b show results for the PDB704 dataset, and c and d correspond to the results on the PDB208 dataset. a and c concern the structure-based predictions, while b and d plot correspond to the sequence-based predictions
Table 3 analyzes relation between predictive quality and secondary structures of the input proteins. It lists the ACC values for proteins that are enriched in helices, strands and coils for the two independent datasets: PDB704 and PDB208. For the proposed sequence-based methods, the ACC values of the proteins with high helix contents of at least 0.4 are higher than the results on the entire dataset. This suggests that results for the helix-rich proteins are characterized by stronger predictive performance. On the other hand, we note a lower predictive performance for the strand- and coil-rich proteins for the sequence-based predictions. This is also true in the case of the structure-based predictions for the proteins enriched in coils. The lower predictive quality for the proteins with high coil content is likely because coils have a relative wide B′-factor profile; see Fig. 2a in (Zhang et al. 2009). In the case of the sequence-based predictions for the strand-rich proteins, a possible explanation comes from the fact that these regions are difficult to identify in the sequence since they are based on long-range (w.r.t. the distance in the sequence) interactions (Faraggi et al. 2012). Table 3 also compares results of the proposed predictor with the PredRSA9 (for sequence-based predictions) and DsspRSA9 (for the structure-based predictions) models. The results demonstrate that the improvements associated with use of our model are consistent across different protein subsets that are enriched in each of the three types of secondary structures. Moreover, we compared the mean SS content (SSC) values for the proteins for which predictive quality is higher vs. lower when compared with the one-stage models (PredRSA9 and DsspRSA9), see Table S3 in the Supplementary Materials. The corresponding differences in the content values were assessed with the two-sided t test at the 95 % significance level showing that they are not significant (p values are above 0.2). This confirms results from Table 3 that the improvements are consistent regardless of the composition of the secondary structures in the input protein.
Additionally, we assessed two-class prediction of rigid vs. flexible residues based on thresholding the native and putative B′-factors. According to the normalization Eq. (1), the mean B′-factor for a given chain is zero. If the threshold is set at the mean B’-factor (0.0), the residues with the B’-factor above zero are regarded as flexible and the residues with B′-factor below zero are considered as rigid. Similarly as in the work by Yuan et al. (2005), we used the mean value as the threshold. We evaluated two-class predictions on the PDB704 and PDB208 datasets using the two-stage and one-stage models that were trained on the PDB632 dataset. The overall accuracies (the ratio of the correctly predicted residues to all residues considered in the dataset) on the PDB704 dataset are 72.3 and 71.6 % for the structure-based two-stage and one-stage models, respectively, and 68.1 and 67.5 % for the sequence-based two-stage and one-stage models, respectively. Similarly, the accuracies on the PDB208 dataset are 72.4 and 71.7 % for the structure-based two-stage and one-stage models, respectively, and 68.4, 67.5 % for the sequence-based two-stage and one-stage models, respectively. We conclude that the proposed two-stage model offers better predictive quality when compared with the one-stage model in the context of predicting rigid vs. flexible residues. The same result observation is true for other choices of threshold values (data not shown).
Prediction models
Given the optimized AA groupings for the DsspRSA- and PredRSA-based models, respectively, we analyze the corresponding prediction models to investigate the relation with flexibility between AA types. As explained above, we utilize two linear regression models to predict the residue flexibility. Similar to our previous work (Zhang et al. 2009), we estimated the weights for the first-stage linear model using the least square fit trained on the PDB632 dataset. Specifically, the linear model for the DsspRSA9 predictor follows
where i represents the ith residue in the protein sequence and \(\hat{B}_{i} '\) denotes B′-factor prediction for the ith residue. The model that uses predicted RSA values, PredRSA9, in the first-stage linear regression, is shown below
The weights in the above two models differ from the values estimated in our previous work (Zhang et al. 2009) since we used a new training dataset PDB632.
Furthermore, we computed the weights in the second stage following Eq. (4). Tables 4 and 5 list the weights that were calculated using the least square fit for the DsspRSA and PredRSA-based models, respectively. For the structure-based model, the reduced optimal AA alphabet derived by the PSO algorithm is {NQEHKPS, L, WY, R, -, I, G, V, ADCMFT}, where the original AA21 alphabet is clustered into 9 AA groups, G1 = NQEHKPS, G2 = L, G3 = WY, G4 = R, G5 = -, G6 = I, G7 = G, G8 = V, and G9 = ADCMFT. Similarly, for the sequence-based model, the reduced optimal alphabet is {CILKMSV, Y, -, FW, G, R, ANDQEHPT} and its size is 7. The values in the two tables represent the strength of relation with flexibility of the RSA values of for the two types of AA groups. In other words, the value in the ith row and the jth column in Tables 4 and 5 is a factor that represents the relation between RSA and flexibility for a residue with AA type G j on the central residue with AA type G i .
When considering the linear model in the first stage, the weights that represent the relation with flexibility for AAs are set to 1. In the second-stage model, these weights may vary around 1. Since the positions represented by the gap symbol ‘-’, which denotes positions used to pad the window at the sequence terminus, are never predicted (cannot be set as a central residue in the window), as shown in Table 4 for the DsspRSA model, their weights in the G5 row are set to 0. The G5 group promotes/strengthens prediction of increased flexibility for the central residue in a given sliding window, which is shown by the fact that the weights in G5 column of Table 4 (except for the G5 row) are larger than 1. Majority of weights in the G7 column are also larger than 1, revealing that the single AA group G7, which includes glycine (G), also promotes prediction of higher flexibility for the central residue. This could be due to the fact that glycine (G) is the smallest and hydrophobic residue. We observe that five columns corresponding to the G2, G3, G4, G6 and G8 AA groups have the weights that are below 1. The weights are close to zero for the G3 that includes tyrosine (W) and tryptophan (Y). This implies that the AA group G3 promotes prediction of lower values of flexibility for the central residue. Moreover, several other AA groups including G2, G4, G6 and G8 are also biased toward prediction of lower values of flexibility for the central residues, since their weights are below 1 and range between 0.5 and 0.7. These four single AA groups G2, G4, G6 and G8 include leucine (L), arginine (R), isoleucine (I) and valine (V), respectively. With the exception of arginine (R), the other three AAs are hydrophobic (Nguyen and Rajapakse 2006), which may inhibit flexibility of the nearby central residues.
Similar observations can be made for the PredRSA-based model based on the values shown in Table 5. The group G3 that includes the gap symbol used to pad the sequence termini promotes prediction of higher flexibility of the nearby central residue. The G2 group that has tryptophan (Y) promotes prediction of lower values of flexibility for the central residue since majority of the weights in the G2 column are low and range between 0.1 and 0.3. The G4 and G6 groups that are composed of large amino acids F, W and R are also biased toward prediction of lower values of flexibility; the corresponding weights in G4 and G6 columns are below 1 and range between 0.5 and 0.8.
Conclusions
We proposed a novel method that utilizes the two-stage RSA-based linear regressions to predict the residue flexibility expressed by B-factor. The linear model in the first stage uses a generic combination of the RSA values in a sequence window, while the linear equation in the second stage utilizes the new concept of AA pair-based space. Furthermore, particle swarm algorithm was used to reduce the AA alphabet used in the second stage to reduce the complexity of the model and to improve the predictive performance. The improvement was verified empirically by measuring and comparing (with a single-stage and other existing methods) the ACC values between native and predicted B′-factors.
Our empirical results suggest that the original full alphabet composed of the 20 standard AAs does not provide optimal predictive performance. The alphabet size that is reduced to about 5–10 AA groups is sufficient to represent the diversity of the relations between AA types and residue flexibility in the context of predicting the residue flexibility when using our two-stage linear model. This is consistent with an observation that the 20 AAs alphabet is redundant in a structural sense (Riddle et al. 1997; Luthra et al. 2007). Additionally, our empirical results confirm that the AA groupings can be efficiently generated with PSO and that these groupings provide a consistent (over multiple datasets) improvement in the predictive quality. In general, this (PSO-based) optimization approach can be used to generate “optimal” amino acid grouping for other types of related problems, which can possibly result in two benefits: simpler predictive model and improved predictive performance. The groupings developed in this work may not be effective in other problems, say in prediction of secondary structure or intrinsic disorder, and this PSO-based optimization should be repeated. Moreover, we stress that likely several similarly effective groupings can be generated for a given problem and the end-user has to make the ultimate selection.
Apart from B-factors that primarily come from the crystal structures, the residue flexibility can be also measured using Nuclear Magnetic Resonance (NMR) (Yang et al. 2007; Carbonell and Sol 2009; Zhang et al. 2010) and is also related to the intrinsically disordered regions that, for instance, can be annotated as the segments of residues with missing electron density in crystal structures (Ferron et al. 2006; Dosztányi et al. 2010). In the past decade, the intrinsic disorder has received considerable amount of attention due to its important functional roles (Uversky and Dunker 2010; Peng et al. 2013a, b). Recently, a number of predictors of disordered region have been developed and their outputs show a relatively strong correlations with the B-factor values (Radivojac et al. 2004; Jin and Dunbrack 2005; Worch and Stolarski 2008). Our previous work (Zhang et al. 2009) has also shown that the B-factor predictors can be applied to the determination of disordered region although the corresponding predictive quality is usually inferior compared to the results obtained with the existing disorder predictors (Peng and Kurgan 2012). This implies that B-factors and disordered regions are to some extent related, which is also shown in Table 2. The annotation of predicted disordered regions was successfully used as input to predict other aspects of protein structure and function including prediction of domain boundaries (Magnusson et al. 2002; Li et al. 2012; Zhang et al. 2013), certain protein–peptide binding events (Disfani et al. 2012) and propensity of protein chains to be amenable to crystallization (Mizianty and Kurgan 2011). Overall, we believe that the B-factor predictors should also provide useful information for similar predictive efforts, besides the more “natural” applications related to the investigations of proteins dynamics, including characterization of the RMSD (root-mean-square distance) of NMR ensembles (Yang et al. 2007), torsion angle fluctuations (Zhang et al. 2010), and order parameters (Carbonell and Sol 2009).
References
Ahmad S, Gromiha MM, Sarai A (2003) Real value prediction of solvent accessibility from amino acid sequence. Proteins 50:629–635. doi:10.1002/prot.10328
Altschul SF, Madden TL, Schäffer AA et al (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25:3389–3402
Berman HM, Westbrook J, Feng Z et al (2000) The protein data bank. Nucleic Acids Res 28:235–242
B-Rao C, Subramanian J, Sharma SD (2009) Managing protein flexibility in docking and its applications. Drug Discov Today 14:394–400. doi:10.1016/j.drudis.2009.01.003
Carbonell P, del Sol A (2009) Methyl side-chain dynamics prediction based on protein structure. Bioinformatics 25:2552–2558. doi:10.1093/bioinformatics/btp463
Chen K, Kurgan M, Kurgan L (2008) Sequence based prediction of relative solvent accessibility using two-stage support vector regression with confidence values. J Biomed Sci Eng 01:1–9. doi:10.4236/jbise.2008.11001
Cheng J, Baldi P (2007) Improved residue contact prediction using support vector machines and a large feature set. BMC Bioinform 8:113. doi:10.1186/1471-2105-8-113
Cilia E, Pancsa R, Tompa P et al (2013) From protein sequence to dynamics and disorder with DynaMine. Nat Commun 4:2741. doi:10.1038/ncomms3741
Cilia E, Pancsa R, Tompa P et al (2014) The DynaMine webserver: predicting protein dynamics from sequence. Nucleic Acids Res 42:W264–W270. doi:10.1093/nar/gku270
Davies MN, Secker A, Freitas AA et al (2008) Optimizing amino acid groupings for GPCR classification. Bioinformatics 24:1980–1986. doi:10.1093/bioinformatics/btn382
Del Sol A, Tsai C-J, Ma B, Nussinov R (2009) The origin of allosteric functional modulation: multiple pre-existing pathways. Structure 17:1042–1050. doi:10.1016/j.str.2009.06.008
Díaz-Espinoza R, Garcés AP, Arbildua JJ et al (2007) Domain folding and flexibility of Escherichia coli FtsZ determined by tryptophan site-directed mutagenesis. Protein Sci 16:1543–1556. doi:10.1110/ps.072807607
Disfani FM, Hsu W-L, Mizianty MJ et al (2012) MoRFpred, a computational tool for sequence-based prediction and characterization of short disorder-to-order transitioning binding regions in proteins. Bioinformatics 28:i75–i83. doi:10.1093/bioinformatics/bts209
Dodson G, Verma CS (2006) Protein flexibility: its role in structure and mechanism revealed by molecular simulations. Cell Mol Life Sci 63:207–219. doi:10.1007/s00018-005-5236-7
Dosztányi Z, Csizmok V, Tompa P, Simon I (2005a) IUPred: web server for the prediction of intrinsically unstructured regions of proteins based on estimated energy content. Bioinformatics 21:3433–3434. doi:10.1093/bioinformatics/bti541
Dosztányi Z, Csizmók V, Tompa P, Simon I (2005b) The pairwise energy content estimated from amino acid composition discriminates between folded and intrinsically unstructured proteins. J Mol Biol 347:827–839. doi:10.1016/j.jmb.2005.01.071
Dosztányi Z, Mészáros B, Simon I (2010) Bioinformatical approaches to characterize intrinsically disordered/unstructured proteins. Brief Bioinformatics 11:225–243. doi:10.1093/bib/bbp061
Eisenmesser EZ, Millet O, Labeikovsky W et al (2005) Intrinsic dynamics of an enzyme underlies catalysis. Nature 438:117–121. doi:10.1038/nature04105
Faraggi E, Xue B, Zhou Y (2009) Improving the prediction accuracy of residue solvent accessibility and real-value backbone torsion angles of proteins by guided-learning through a two-layer neural network. Proteins 74:847–856. doi:10.1002/prot.22193
Faraggi E, Zhang T, Yang Y et al (2012) SPINE X: improving protein secondary structure prediction by multistep learning coupled with prediction of solvent accessible surface area and backbone torsion angles. J Comput Chem 33:259–267. doi:10.1002/jcc.21968
Ferron F, Longhi S, Canard B, Karlin D (2006) A practical overview of protein disorder prediction methods. Proteins 65:1–14. doi:10.1002/prot.21075
Fontana A, Spolaore B, Mero A, Veronese FM (2008) Site-specific modification and PEGylation of pharmaceutical proteins mediated by transglutaminase. Adv Drug Deliv Rev 60:13–28. doi:10.1016/j.addr.2007.06.015
Gao J, Zhang T, Zhang H et al (2010) Accurate prediction of protein folding rates from sequence and sequence-derived residue flexibility and solvent accessibility. Proteins 78:2114–2130. doi:10.1002/prot.22727
Gutteridge A, Bartlett GJ, Thornton JM (2003) Using a neural network and spatial clustering to predict the location of active sites in enzymes. J Mol Biol 330:719–734
Halle B (2002) Flexibility and packing in proteins. Proc Natl Acad Sci USA 99:1274–1279. doi:10.1073/pnas.032522499
Han R, Leo-Macias A, Zerbino D et al (2008) An efficient conformational sampling method for homology modeling. Proteins 71:175–188. doi:10.1002/prot.21672
Han L, Zhang Y-J, Song J et al (2012) Identification of catalytic residues using a novel feature that integrates the microenvironment and geometrical location properties of residues. PLoS One 7:e41370. doi:10.1371/journal.pone.0041370
Jin Y, Dunbrack RL Jr (2005) Assessment of disorder predictions in CASP6. Proteins 61(Suppl 7):167–175. doi:10.1002/prot.20734
Jones DT (1999) Protein secondary structure prediction based on position-specific scoring matrices. J Mol Biol 292:195–202. doi:10.1006/jmbi.1999.3091
Kabsch W, Sander C (1983) Dictionary of protein secondary structure: pattern recognition of hydrogen-bonded and geometrical features. Biopolymers 22:2577–2637. doi:10.1002/bip.360221211
Kennedy J, Eberhart R (1995) Particle swarm optimization. In: Proceedings IEEE International Conference on Neural Networks, vol 4, 1995 pp 1942–1948
Kundu S, Melton JS, Sorensen DC, Phillips GN Jr (2002) Dynamics of proteins in crystals: comparison of experiment with simple models. Biophys J 83:723–732. doi:10.1016/S0006-3495(02)75203-X
Kurgan L, Cios K, Zhang H et al (2008) Sequence-based methods for real value predictions of protein structure. Curr Bioinform 3:183–196. doi:10.2174/157489308785909197
Kwansa AL, Freeman JW (2010) Elastic energy storage in an unmineralized collagen type I molecular model with explicit solvation and water infiltration. J Theor Biol 262:691–697. doi:10.1016/j.jtbi.2009.10.024
Li B-Q, Hu L–L, Chen L et al (2012) Prediction of protein domain with mRMR feature selection and analysis. PLoS One. doi:10.1371/journal.pone.0039308
Lin W-Q, Jiang J-H, Shen Q et al (2005) Optimized block-wise variable combination by particle swarm optimization for partial least squares modeling in quantitative structure-activity relationship studies. J Chem Inf Model 45:486–493. doi:10.1021/ci049890i
Lin C-P, Huang S-W, Lai Y-L et al (2008) Deriving protein dynamical properties from weighted protein contact number. Proteins 72:929–935. doi:10.1002/prot.21983
Linding R, Jensen LJ, Diella F et al (2003) Protein disorder prediction: implications for structural proteomics. Structure 11:1453–1459
Liu X, Karimi HA (2007) High-throughput modeling and analysis of protein structural dynamics. Brief Bioinform 8:432–445. doi:10.1093/bib/bbm014
Liu J, Rost B (2004) Sequence-based prediction of protein domains. Nucleic Acids Res 32:3522–3530. doi:10.1093/nar/gkh684
Luthra A, Jha AN, Ananthasuresh GK, Vishveswara S (2007) A method for computing the inter-residue interaction potentials for reduced amino acid alphabet. J Biosci 32:883–889
Mackereth CD, Sattler M (2012) Dynamics in multi-domain protein recognition of RNA. Curr Opin Struct Biol 22:287–296. doi:10.1016/j.sbi.2012.03.013
Magnusson U, Chaudhuri BN, Ko J et al (2002) Hinge-bending motion of d-allose-binding protein from Escherichia coli three open conformations. J Biol Chem 277:14077–14084. doi:10.1074/jbc.M200514200
Meissner M, Schmuker M, Schneider G (2006) Optimized particle swarm optimization (OPSO) and its application to artificial neural network training. BMC Bioinform 7:125. doi:10.1186/1471-2105-7-125
Mizianty MJ, Kurgan L (2011) Sequence-based prediction of protein crystallization, purification and production propensity. Bioinformatics 27:i24–i33. doi:10.1093/bioinformatics/btr229
Mizianty MJ, Stach W, Chen K et al (2010) Improved sequence-based prediction of disordered regions with multilayer fusion of multiple information sources. Bioinformatics 26:i489–i496. doi:10.1093/bioinformatics/btq373
Neuvirth H, Raz R, Schreiber G (2004) ProMate: a structure based prediction program to identify the location of protein–protein binding sites. J Mol Biol 338:181–199. doi:10.1016/j.jmb.2004.02.040
Nguyen MN, Rajapakse JC (2006) Two-stage support vector regression approach for predicting accessible surface areas of amino acids. Proteins 63:542–550. doi:10.1002/prot.20883
Niu Y, Shen L (2006) An adaptive multi-objective particle swarm optimization for color image fusion. In: Wang T-D, Li X, Chen S-H et al (eds) Simulated evolution and learning. Springer, Berlin Heidelberg, pp 473–480
Oğul H, Mumcuoğu EU (2007) Subcellular localization prediction with new protein encoding schemes. IEEE/ACM Trans Comput Biol Bioinform 4:227–232. doi:10.1109/TCBB.2007.070209
Pan X-Y, Shen H-B (2009) Robust prediction of B-factor profile from sequence using two-stage SVR based on random forest feature selection. Protein Pept Lett 16:1447–1454
Panjkovich A, Daura X (2010) Assessing the structural conservation of protein pockets to study functional and allosteric sites: implications for drug discovery. BMC Struct Biol 10:9. doi:10.1186/1472-6807-10-9
Parthasarathy S, Murthy MR (1997) Analysis of temperature factor distribution in high-resolution protein structures. Protein Sci 6:2561–2567. doi:10.1002/pro.5560061208
Pedregosa F, Varoquaux G, Gramfort A et al (2011) Scikit-learn: machine learning in Python. J Mach Learn Res 12:2825–2830
Peng Z-L, Kurgan L (2012) Comprehensive comparative assessment of in silico predictors of disordered regions. Curr Protein Pept Sci 13:6–18
Peng Z, Oldfield CJ, Xue B et al (2013a) A creature with a hundred waggly tails: intrinsically disordered proteins in the ribosome. Cell Mol Life Sci. doi:10.1007/s00018-013-1446-6
Peng Z, Xue B, Kurgan L, Uversky VN (2013b) Resilience of death: intrinsic disorder in proteins involved in the programmed cell death. Cell Death Differ 20:1257–1267. doi:10.1038/cdd.2013.65
Peterson EL, Kondev J, Theriot JA, Phillips R (2009) Reduced amino acid alphabets exhibit an improved sensitivity and selectivity in fold assignment. Bioinformatics 25:1356–1362. doi:10.1093/bioinformatics/btp164
Radivojac P, Obradovic Z, Smith DK et al (2004) Protein flexibility and intrinsic disorder. Protein Sci 13:71–80. doi:10.1110/ps.03128904
Riddle DS, Santiago JV, Bray-Hall ST et al (1997) Functional rapidly folding proteins from simplified amino acid sequences. Nat Struct Biol 4:805–809
Scheraga HA, Khalili M, Liwo A (2007) Protein-folding dynamics: overview of molecular simulation techniques. Annu Rev Phys Chem 58:57–83. doi:10.1146/annurev.physchem.58.032806.104614
Schlessinger A, Rost B (2005) Protein flexibility and rigidity predicted from sequence. Proteins 61:115–126. doi:10.1002/prot.20587
Schnell JR, Dyson HJ, Wright PE (2004) Structure, dynamics, and catalytic function of dihydrofolate reductase. Annu Rev Biophys Biomol Struct 33:119–140. doi:10.1146/annurev.biophys.33.110502.133613
Sickmeier M, Hamilton JA, LeGall T et al (2007) DisProt: the database of disordered proteins. Nucleic Acids Res 35:D786–D793. doi:10.1093/nar/gkl893
Tegge AN, Wang Z, Eickholt J, Cheng J (2009) NNcon: improved protein contact map prediction using 2D-recursive neural networks. Nucleic Acids Res 37:W515–W518. doi:10.1093/nar/gkp305
Tokuriki N, Tawfik DS (2009) Protein dynamism and evolvability. Science 324:203–207. doi:10.1126/science.1169375
Tozzini V (2005) Coarse-grained models for proteins. Curr Opin Struct Biol 15:144–150. doi:10.1016/j.sbi.2005.02.005
Uversky VN, Dunker AK (2010) Understanding protein non-folding. Biochim Biophys Acta 1804:1231–1264. doi:10.1016/j.bbapap.2010.01.017
Vihinen M (1987) Relationship of protein flexibility to thermostability. Protein Eng 1:477–480
Walsh I, Martin AJM, Di Domenico T, Tosatto SCE (2012) ESpritz: accurate and fast prediction of protein disorder. Bioinformatics 28:503–509. doi:10.1093/bioinformatics/btr682
Wang J-Y, Lee H-M, Ahmad S (2007) SVM-Cabins: prediction of solvent accessibility using accumulation cutoff set and support vector machine. Proteins 68:82–91. doi:10.1002/prot.21422
Weathers EA, Paulaitis ME, Woolf TB, Hoh JH (2004) Reduced amino acid alphabet is sufficient to accurately recognize intrinsically disordered protein. FEBS Lett 576:348–352. doi:10.1016/j.febslet.2004.09.036
Worch R, Stolarski R (2008) Stacking efficiency and flexibility analysis of aromatic amino acids in cap-binding proteins. Proteins 71:2026–2037. doi:10.1002/prot.21882
Yang L-W, Bahar I (2005) Coupling between catalytic site and collective dynamics: a requirement for mechanochemical activity of enzymes. Structure 13:893–904. doi:10.1016/j.str.2005.03.015
Yang L-W, Eyal E, Chennubhotla C et al (2007) Insights into equilibrium dynamics of proteins from comparison of NMR and X-ray data with computational predictions. Structure 15:741–749. doi:10.1016/j.str.2007.04.014
Yang L, Song G, Jernigan RL (2009) Protein elastic network models and the ranges of cooperativity. Proc Natl Acad Sci USA 106:12347–12352. doi:10.1073/pnas.0902159106
Yuan Z, Huang B (2004) Prediction of protein accessible surface areas by support vector regression. Proteins 57:558–564. doi:10.1002/prot.20234
Yuan Z, Zhao J, Wang Z-X (2003) Flexibility analysis of enzyme active sites by crystallographic temperature factors. Protein Eng 16:109–114
Yuan Z, Bailey TL, Teasdale RD (2005) Prediction of protein B-factor profiles. Proteins 58:905–912. doi:10.1002/prot.20375
Zhang H, Zhang T, Chen K et al (2008) Sequence based residue depth prediction using evolutionary information and predicted secondary structure. BMC Bioinform 9:388. doi:10.1186/1471-2105-9-388
Zhang H, Zhang T, Chen K et al (2009) On the relation between residue flexibility and local solvent accessibility in proteins. Proteins 76:617–636. doi:10.1002/prot.22375
Zhang T, Faraggi E, Zhou Y (2010) Fluctuations of backbone torsion angles obtained from NMR-determined structures and their prediction. Proteins 78:3353–3362. doi:10.1002/prot.22842
Zhang H, Zhang T, Chen K et al (2011) Critical assessment of high-throughput standalone methods for secondary structure prediction. Brief Bioinform 12:672–688. doi:10.1093/bib/bbq088
Zhang H, Shi H, Hanlon M (2012a) A large-scale comparison of computational models on the residue flexibility for NMR-derived proteins. Protein Pept Lett 19:244–251
Zhang T, Faraggi E, Xue B et al (2012b) SPINE-D: accurate prediction of short and long disordered regions by a single neural-network based method. J Biomol Struct Dyn 29:799–813
Zhang X, Lu L, Song Q et al (2013) DomHR: accurately identifying domain boundaries in proteins using a hinge region strategy. PLoS One 8:e60559. doi:10.1371/journal.pone.0060559
Zuo Y-C, Li Q-Z (2010) Using K-minimum increment of diversity to predict secretory proteins of malaria parasite based on groupings of amino acids. Amino Acids 38:859–867. doi:10.1007/s00726-009-0292-1
Acknowledgments
This work was supported by the National Natural Science Foundation of China (grant no. 61170099) and the Zhejiang Provincial Natural Science Foundation of China (grant no. Y1110840) to H.Z., and by Discovery grant by Natural Sciences and Engineering Research Council (NSERC) of Canada to L.K.
Conflict of interest
The authors declare that they have no competing financial interests.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Zhang, H., Kurgan, L. Improved prediction of residue flexibility by embedding optimized amino acid grouping into RSA-based linear models. Amino Acids 46, 2665–2680 (2014). https://doi.org/10.1007/s00726-014-1817-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00726-014-1817-9