Abstract
Immature cells of etiolated apices of sprouts growing from Helianthus tuberosus (H. t.) tubers showed Ca2+-dependent transglutaminase (TG, EC 2.3.2.13) activity on fibronectin (more efficiently) and dimethylcasein as substrates. Three main TG bands of about 85, 75 and 58 kDa were isolated from the 100,000×g apices supernatant through a DEAE-cellulose column at increasing NaCl concentrations and immuno-identified by anti-TG K and anti-rat prostate gland TG antibodies. These three fractions had catalytic activity as catalyzed polyamine conjugation to N-benzyloxycarbonyl-L-γ-glutaminyl-L-leucine (Z-L-Gln-L-Leu) and the corresponding glutamyl-derivatives were identified. The amino acid composition of these TG proteins was compared with those of several sequenced TGs of different origin. The composition of the two larger bands presented great similarities with annotated TGs; in particular, the 75 kDa form was very similar to mammalian inactive EPB42. The 58 kDa form shared a low similarity with other TGs, including a maize sequence of similar molecular mass, which, however, did not present the catalytic triad in the position of all annotated TGs. A 3D model of the H. t. TGs was built adopting TG2 as template. These novel plant TGs are hypothesized to be constitutive and discussed in relation to their possible roles in immature cells. These data suggest that in plants, multiple TG forms are active in the same organ and that plant and animal enzymes probably are very close not only for their catalytic activity but also structurally.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
So far, nine different transglutaminases (TGs) are identified in animals: they have different molecular mass, many of them are tissue-specific with specific functions and developmental expression and are involved in cell differentiation, apoptosis and stabilization of cellular and extracellular structure (Griffin et al. 2002; Lorand and Graham 2003; Beninati et al. 2009). TGs are highly expressed in actively growing tumor cells (Chhabra et al. 2009) with a low transamidating activity; on the contrary, TGs show increased catalytic activity in differentiated cells (Lentini et al. 2004). Moreover, TGs are involved in several other diseases (Tabolacci et al. 2012).
The presence of TG activity in plants has been found in lower and higher plants (reviewed by Serafini-Fracassini et al. 1995; Del Duca and Serafini-Fracassini 2005; Serafini-Fracassini and Del Duca 2008). The plant TG physiological role appears related to photosynthesis, fertilization, response to abiotic and biotic stresses, senescence and programmed cell death. In chloroplasts, plant TG appears to stabilize the photosynthetic complexes and ribulose bisphosphate carboxylase/oxygenase, being regulated by light and other factors, possibly exerting a positive effect on photosynthesis and photo-protection. In the cytosol, plant TG modifies cytoskeletal proteins, as observed in pollen tube, which regulates the growth (Del Duca et al. 2009). Preliminary reports suggest an involvement also in the pollen tube cell wall construction/organization (Di Sandro et al. 2010).
Similar to animal TGs, plant enzymes are Ca2+-dependent and able to produce glutamyl-polyamine (PA) derivatives. Most of them respond to animal TG antibodies and the first one cloned, sequenced from Arabidopsis thaliana (AtPng1p, Q9FGY9), showed in its sequence the TG catalytic triad. The plant TG shows little sequence homology with the best-known animal enzymes; however, they share a possible structural homology (Della Mea et al. 2004a; Serafini-Fracassini et al. 2009).
To date most of the TG activities have been found in mature plant cells, in which possibly only one or more particular TGs related to the differentiation program were expressed. Few data reported the presence of plant TGs in meristematic cells and some were observed in the adventitious secondary meristematic cells, for example, during the synchronous cell-cycle of explanted parenchyma of Helianthus tuberosus (H. t.) tuber grown in invitro colture (Mossetti et al. 1987; Serafini-Fracassini et al. 1988; Del Duca et al. 2000a) or in the Zea mays embryogenetic tissue also cultivated in vitro (Bernet et al. 1999). In these de-differentiated tissues containing mature and meristematic cells, different TG forms could be present (Del Duca et al. 2000b).
Anyway, researches dealing with the possible presence of TG in meristematic cells (Serafini-Fracassini et al. 1989) are still in their infancy; among the few papers published, none reported data about the cloned plant proteins with verified TG activity as the ability to form (γ-glutamyl)-PAs and immuno-recognition by animal TGs antibodies. The presence of a TG-like activity in meristematic primary tissue of etiolated H. t. sprout apices was deduced by the recovery of TCA-insoluble PAs, assuming that these amines were covalently linked to proteins (Serafini-Fracassini et al. 1988). Such apices are composed by staminal cells surrounded by the rapidly dividing cells of the primary apical meristem as well as by their derived cells; all are protected by leaf primordia, whose immature cells are still multiplying and developing etioplasts. The catalytic reaction exerted by the total crude homogenate was found similar to that reported for TGs isolated from animal cells, except for the fact that the widely used TG substrate, dimethylcasein (DMC), was poorly labelled by the plant catalytic activity at the usual 0.3 mM Ca2+ concentration (Serafini-Fracassini et al. 1988).
The aim of the present study was to verify if one or more TG forms constitutively exist in the H. t. immature primary system presented. This has being achieved by isolation of proteins, by animal TG antibodies immuno-recognition, by detecting the transamidating catalytic activity through a highly reactive synthetic substrate (Folk et al. 1980) and identifying the glutamyl-derivatives formed (Del Duca et al. 1995). These preliminary information, compared with those already described in differentiated plant tissues (Del Duca and Serafini-Fracassini 2005; Serafini-Fracassini and Del Duca 2008) will clarify which TG forms are expressed also in the H. t. primary meristems. The amino acid composition (aac) of these forms has been compared to that of some TGs of plant and animal origin and the high composition similarity has been useful for proposing a putative 3D model.
Materials and methods
Materials
Etiolated apices were excised at the dormancy release from tuber sprouts of Helianthus tuberosus L. cv OB1, selected and grown in the Botanical Garden of Bologna University and stored in moist sand at 4 °C. Where not otherwise specified, chemicals were purchased from Sigma-Aldrich (Milano, Italy).
Extraction
Apices were rapidly ground in an ice-cold mortar with 2 volumes per gm (fresh weight) of 50 mM Tris–HCl buffer pH 8 containing 45 mM DTT and 2 % PVPP. After filtration through four layers of cheesecloth, extracts were centrifuged at 27,000×g for 30 min and the supernatant (SN) used directly for the enzyme assay.
Separation of H. t. TG by DEAE-cellulose column
The separation of H. t. TG by ion-exchange chromatography has been performed according to Kim et al. (1990) with some modifications reported below. The 27,000×g SN was centrifuged at 100,000×g for 30 min and the SN was loaded onto a DEAE-cellulose column (DE-52, Whatman, 16 × 200 mm) equilibrated with 50 mM Tris–HCl buffer pH 8. Proteins were eluted on a stepwise gradient obtained using NaCl (from 0 to 0.6 M) in 50 mM Tris–HCl buffer pH 8. 5-ml fractions were collected, dialyzed at 4 °C overnight against 50 mM Tris–HCl buffer pH 8 and concentrated by dialysis against PEG. Samples were aliquoted and lyophilized. For the TG assay, sample aliquots (1 μg prot μl–1) were resuspended in Tris–HCl buffer pH 8.
TG activity assay
The activity was detected in the crude extract, in its 27,000×g SN, in the 100,000×g SN as well as in the DEAE fractions resuspended in buffer. Two methods were used: the radiolabelled assay and the microplate assay.
To determine TG activity with the radiolabelled assay, aliquots of 50 μl (1 μg prot μl−1) of the above-reported samples were incubated for 30 min at 30 °C in the presence of 14 μl of [1,4(n)-3H] spermidine (Spd) (520 kBq, 555 GBq/mmol) (Amersham Pharmacia, Milan) taken to dryness, 10 μl of 2, 30 or 40 mM Spd in 0.5 M Tris–HCl pH 8.5 and 40 μl of 0.1 mM NaOH with or without 50 mM (final concentration) Z-L-Gln-L-Leu as a synthetic dipeptide substrate.
The amount of radiolabelled Spd, covalently incorporated into the acyl donor substrate by TGs has been performed as reported by Del Duca et al. (1995).
The microplate assay was performed to compare the affinity of the 27,000×g SN enzyme of the sprout apices for two different substrates N, N’-DMC and fibronectin (FN), according to the method reported by Lilley et al. (1998) with minor modifications. The level of biotinylated cadaverine conjugated to DMC (used at 10 mg ml−1, final concentration 400 μM) or FN (used 5 μg ml−1, final concentration 23 μM)-coated 96-well microtitre plates (NUNC Life Technologies, UK) was expressed as Ca2+-dependent increase in A450, after subtraction of the value of the EGTA-treated control. Specific activity was determined as a change in A450 of 0.1 hr−1 mg−1 of proteins.
Identification of the γ-glutamyl-derivatives produced by H. t. TG catalysis
The TCA-soluble and -insoluble fractions obtained from the TG radiolabelled assay were extensively washed with anhydrous ethylether and proteolytically digested according to a method previously reported (Beninati et al. 1988). The ion-exchange chromatographic separation of (γ-glutamyl)-PA derivatives from the proteolytic digest was performed on an LKB Alpha Plus 4151 amino acid analyzer equipped with a 4.5 × 90 mm column (Dionex-DC-6A) packed with Ultropac 8 resin Na+ form, using the five buffer system previously reported by Folk (1980). The identity of conjugated PA was determined by the release of free PA after acid hydrolysis of the ion-exchange chromatographic fractions containing the presumptive derivatives, identified by the chromatographic retention time of γ-glutamyl-PA standards obtained by published procedures reported by Folk (1980) and purified according to Beninati et al. (1988).
Sodium-dodecyl-sulphate–polyacrylamide gel electrophoresis (SDS-PAGE)
DEAE-fractions were dissolved in 60 mM Tris buffer pH 6.8 containing 4 % SDS, 10 mM 2-mercaptoethanol and 10 % glycerol and separated by a discontinuous gradient gel (9–16 % acrylamide) according to the method of Schägger and von Jagow (1987). The gels were stained with Coomassie-blue R. Alternatively proteins were blotted on 0.2 μm pore nitrocellulose sheets (Hoefer, Milan, Italy).
Western-blot analysis
Transfer of proteins was performed with a Trans-Blot Transfer Cell (Bio Rad, Milano, Italy) overnight at 30 V in a transfer buffer of 25 mM Tris pH 8.3, 192 mM glycine, 20 % (v/v) methanol. The nitrocellulose was treated as reported in Di Sandro et al. (2010) and incubated overnight at room temperature with a rabbit polyclonal antibody against rat prostate gland TG (a kind gift of Prof. C. Esposito, University of Salerno) or with rabbit monoclonal antibody against the sequences containing the active site of human epidermal insoluble TG K, prepared according to the suggestions of Dr. S.I. Chung (N.I.H., Bethesda, USA), both diluted with saline Tris-buffer containing 5 % non-fat dry milk. The secondary antibody was incubated for 2 h at room temperature with alkaline phosphatase-conjugated protein A diluted with TBST, and the TG-immunoreactive bands were visualized using the BCIP/NBT substrate system (Bio Rad) following the directions of the manufacturer. The negative control was performed using only the secondary anti-mouse and anti-chicken antibodies. Moreover, the H.t. crude extract has been tested with another additional animal TGs antibody, anti-TG2 (ID-10) and with anti plant TG (AtPng1p) antibody.
Amino acid analysis
Amino acid analysis was performed directly on the samples obtained by electroblotting onto nitrocellulose membranes the SDS-PAGE protein bands corresponding to the three enzymes and then cutting the corresponding region. The three samples were acid hydrolyzed by 6 N HCl for 12 h at 110 °C. The hydrolysate was dried and resuspended in sample diluents and analysed by a LKB model Alfa Plus 4151 amino acid analyzer.
Protein assay
Proteins were determined according to the method of Bradford (1976).
Homology modelling of the H. t. 75 kDa protein
The putative three-dimensional model of the H. t. 75 kDa was modelled by comparison with the three-dimensional structure of the human tissue TGs [Protein Data Bank (PDB) code 3LY6], and known with a 0.314-nm resolution (Han et al. 2010). The target sequence was obtained from the consensus sequence of the four annotated mammalian TGs (TGM2_HUMAN, TGM2_MOUSE, TGM2_BOVIN and TGM2_CAVCU) by the VisCoSe tool, available at the web address http://bio.math-inf.uni-greifswald.de/viscose/ (Spitzer et al. 2004). Alignment of the target with the template was performed with CLUSTAL Ω (http://www.ebi.ac.uk/Tools/msa/clustalo/) (Thompson et al. 1994), Matrix Blosum, default gap penalty and manually checked. The resulted pairwise sequence identity was 93 %. Modelling was performed with the program Modeller version 9.8 (Marti-Renom et al. 2000). For a given alignment, 15 model structures were built and were evaluated with the PROCHECK suite of programs (Laskowski et al. 1993).
Statistics
The values reported are expressed as mean ± SD and represent one of at least 3 or 4 experiments undertaken in triplicate. Differences between sample sets were determined by the Student’s t test with 95 % confidence limits. When indicated statistical analysis was performed using GraphPad Prism (version 5.03 Windows GraphPad Software Inc., La Jolla, CA, USA). All determinations were repeated at least three times.
Results
The enzyme activity and separation of TG-active fractions by a DEAE–cellulose column
To evaluate the TG catalytic activity of the etiolated apices extract, its 27,000×g SN was tested in the microplate assay by the conjugation of the biotinylated cadaverine to the tissue TG substrates DMC and FN at increasing Ca2+ concentrations. As shown in Fig. 1, FN was very efficiently recognized at low Ca2+ concentrations reaching the highest activity level at 0.75 mM CaCl2; DMC, utilized at higher concentration when compared with FN (400 μM DMC in respect to 23 μM FN) reached a plateau only at 10 mM CaCl2 (Fig. 1).
To evaluate the (γ-glutamyl)-derivatives, the TG activity was also measured by the radioactive assay incubating the crude extract with the dipeptide N-benzyloxycarbonyl-L-α-glutaminyl-L-leucine (Z-L-Gln-L-Leu) and with a mixture of spermidine (Spd)/(3H)Spd, as this amine-donor substrate is, among the PAs, the more efficiently conjugated (Serafini-Fracassini et al. 1988). Since Z-L-Gln-L-Leu is TCA-soluble, the incorporation of radiolabelled Spd was determined in the TCA-soluble fraction through the identification of (γ-glutamyl)-derivatives (Table 1). Z-L-Gln-L-Leu is an excellent concentration-dependent amine-acceptor substrate for H. t. TG (not shown) and thus was used for the subsequent determinations of TG activity in the DEAE protein fractions. The dipeptide was assayed also with the 27,000×g SN and its 100,000×g SN; in the last two fractions the specific activity was similar (Table 1).
The anion-exchange DE-52 DEAE-cellulose chromatography of the proteins from the 100,000×g soluble fraction allowed to separate three distinct protein fractions eluted with 0.1, 0.2 and 0.3 M NaCl, which presented the capacity to bind labelled Spd, fractions 0.3 and 0.1 being those with the highest specific activity (Table 1). The isolation rate fold obtained was higher (37 %) for the proteins collected in the 0.3-M fraction of the DEAE elution in respect to those collected at 0.1 M (18 %) and 0.2 M (5 %) salt concentration. In the remaining fractions (0.4–0.6 M) no activity was detected (data not shown).
Figure 2 shows the ion-exchange chromatographic separation of (γ-glutamyl)-PAs, obtained by incubating the DEAE-fractions eluted with 0.1, 0.2 and 0.3 M NaCl with and without Z-L-Gln-L-Leu. The TCA SN of the DEAE chromatographic fractions incubated with the dipeptide showed the most significant difference in respect of the sample incubated without the dipeptide, due to the production of N 1-mono-(γ-glutamyl)-Spd (Fig. 2, peak I) and N 8-mono-(γ-glutamyl)-Spd (Fig. 2, peak II) derivatives; Spd, in fact, being an asymmetric molecule, could bind to the dipeptide glutamyl-residue by both the butyl- or propyl-amino terminals. The two peaks are clearly separated only in the sample eluted by 0.1 M NaCl fraction.
Distribution of radioactive products catalysed by the DEAE-fractions from Helianthus tuberosus sprout apices. Ion-exchange chromatographic separation of the labelled products of the TG assay obtained from the proteolytic digestion of proteins present in 0.1, 0.2 and 0.3 M NaCl DEAE-fractions, deriving from 100,000×g SN. Fractions were incubated with [3H]-Spd and with (white symbols) or without (black symbols) N-benzyloxycarbonyl-L-α-glutaminyl-L-leucine in the presence of 0.2 mM Spd. The peaks indicated in the chromatograms correspond to the retention times of N 1-mono-(γ-glutamyl)-Spd (peak I) and N 8-mono-(γ-glutamyl)-Spd (peak II)
Immuno-staining of H. t. TG active fractions eluted by DEAE-cellulose column
DEAE-fractions having TG activity were dialysed and analysed by SDS-PAGE under reducing conditions by a discontinuous gradient gel from 9 to 16 % acrylamide and immuno-blotted as shown in Fig. 3. Proteins of molecular range of about 75 kDa (DEAE-fraction, 0.1 M NaCl), 85 kDa (0.2 M NaCl) and 58 kDa (0.3 M NaCl) were recognized by both the poly specific antibody to soluble rat prostate gland TG and the monospecific antibody to human epidermal insoluble TG K (directed against the active site to detect all the proteins with this sequence and in particular mammalian TG1); both these antibodies were shown to immuno-recognize plant TGs (Del Duca et al. 1994, 2000a). Other lower molecular range proteins, namely a 32 kDa (0.2 M NaCl) and a 20 kDa (0.3 M NaCl), were also immuno-recognized. Two of these protein bands share MW similar to those of other TGs as shown in Table 3 and described below. Other antibodies, like those raised against AtPng1p (Della Mea et al. 2004a) and against purified guinea pig liver TG (Di Sandro et al. 2010) also recognized these three main bands (not shown).
Immunolabel analysis of proteins of DEAE-fractions from Helianthus tuberosus sprout apices. Fractions 0.1, 0.2 and 0.3 eluted from DEAE column were separated on SDS-PAGE according to Schägger and von Jagow (1987) and blotted on nitrocellulose. The antibody cross-reaction was performed with the polyclonal anti-rat prostate TG (anti-TG 4, Abp) and the monoclonal anti-human keratinocyte TG (anti-TG K, Abk) as primary antibodies
Amino acid composition of H. t. TGs
The aac of the three isolated proteins, recognized by immuno-blotting, was analysed by an amino acid analyzer and it was found to be very consistent between individual isolation runs. The results are shown in Table 2, where the amount of each aa is compared among the three different proteins (each protein is indicated with its correspondent MW).
On the basis of the % aac and of the MW of the different proteins in the eluted bands, all endowed with TG activity as shown above, and using also AACompIdent (which allows to compare the aac of a protein with the individual composition of all the proteins in the UniProtKB/Swiss-Prot databases), we retrieved from the database the proteins annotated and validated with TG activity (with the exception of inactive EPB42), closest in aac and MW to the H. t. proteins as well as the only two annotated sequences of plant origin: the recombinant TG of Arabidopsis thaliana, AtPng1p (Q9FGY9) (PNG1_ARATH) and a sequence detected in Zea mays (Q6KF70), the latter present only in Trembl database, not included in SwissProt. Table 3 shows a detailed description of the reference proteins for each eluted band.
Table 4 reports the list of the percentage of each aa of the H. t. proteins compared with those of selected TGs. The aac of the H. t. 85 kDa enzyme was compared with that of the human factor XIII (FXIIIA) and Peptide:N-glycanase_Arabidopsis thaliana (PNG1_ARATH); the 75 kDa aac was compared with four TG2 (TGM2) of different origin, the human TG3 (TGM3) and the inactive EPB42. The H. t. 58 kDa was compared with the Zea mays sequence (Q6KF70) being the TG with the closest MW. The 85 and 75 kDa H. t. proteins are characterized by an aa frequency that well compares with the reference TGs (Table 4). As an example, only 4 aa differed for a decimal of percentage each, over a total of 18 aa identified, between the 75 kDa protein and the erythrocyte 4.2 band (EPB42) whose structure is unknown at the moment.
The MW of the H. t. 58 kDa protein is lower than that of all the other TGs, and its aac is also different from that of Zea mays (Q6KF70_MAIZE). These data corroborate also at the molecular level the consistency of the experimentally determined function of the H. t. proteins and show that each band contains proteins whose MW and aac are fully consistent with animal and plant TGs fully described before.
We did not get enough material to perform a sequencing experiment. However, considering all the observations detailed above, a “consensus sequence” for H. t. 75 kDa protein has been obtained on the basis of all the mammalian TG sequences reported in Table 3 (EPB42, being an inactive protein, has not been considered for the consensus sequence) (Spitzer et al. 2004). The alignments of the TG core domain of all the reference TGs (Table 3) including H. t. consensus sequence are shown in Fig. 4. Noticeably Q6KF70_MAIZE, which includes in its entire sequence only two cysteins, does not contain the catalytic triad in the core domain, but in the protein C terminus as previously reported (Villalobos et al. 2004) and shown in Table 3.
Multiple sequence alignment of the core transglutaminase domain. The different reference TGs (Table 3) are aligned with the consensus sequence of the 75 kDa protein with Clustal Ω (http://www.ebi.ac.uk/Tools/msa/clustalo/). Only the transglutaminase core is reported; residues involved in the catalytic triad (Cys, His, Asp) are highlighted with a blue box for all the aligned TGs, with the exception of the Zea mays sequences whose catalytic triad is in the C terminus (Villalobos et al. 2004) and of the human EPB42 (not active). Residues are coloured according to their physicochemical properties (small hydrophobic and aromatic with the exception of Y are coloured in red, acidic in blue, basic in magenta, Hydroxyl + sulfhydryl + amine + G depicted in green). For the whole alignment see supplementary materials
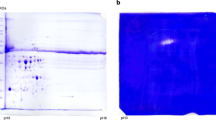
With the consensus sequence at hand a putative model based on the 3D structure of TGM2_HUMAN (PDB code: 3LY6, R factor: 3.14 Å) was computed. The model is shown in Fig. 5.
Homology modelling of the Helianthus tuberosus 75 kDa protein. a Structural comparison of the human TGM2 (Protein Data Bank code 3LY6) depicted in blue and of the putative H. t. 75 kDa protein, in red. N and C-terminus are reported. The position of the catalytic triad is highlighted with the green circle. b Zoom on the catalytic triad C277, H335 and D358
Discussion
The presence of TG-like activity in H. t. sprout apices was first reported in 1988 (Serafini-Fracassini et al. 1988). At that time, however, it was not characterized since the enzyme assayed was scarcely able to catalyze the incorporation of PAs into an exogenous substrate, like DMC, and this rendered difficult to purify and characterize it. In the present paper we have overcome this gap by the use of other exogenous substrates like FN and the synthetic dipeptide Z-L-Gln-L-Leu, specific substrates for tissue TG (Griffin et al. 2002; Folk and Chung 1985). Both are excellent concentration-dependent amine-acceptor substrates for H. t. TGs. Z-L-Gln-L-Leu allowed to isolate the glutamyl-PA derivatives confirming the functional identification of the isolated proteins of H. t. sprout apices as TGs (Table 1). The isolation rate and the specific activity were higher for the H. t. 58 kDa containing fraction when compared with the H. t. 75 and 85 kDa ones. Thus, in this primary meristem at least three main TG forms separated and immunostained with anti-TG 1 (TG K) and -TG 4 (rat prostatic gland TG) antibodies are active.
The ion-exchange chromatographic elution pattern of these enzymatic forms appears very similar to those reported both in chondrosarcoma cells by Chung et al. (1988) and in B16–F10 melanoma cells by Beninati et al. (1993).
The cross-reactivity of H. t. TGs with the mammalian TGs antibodies should suggest that a similarity exists between at least a part of the sequences of H. t. enzymes and the TGs of animal origin. The monoclonal antibody should recognise the catalytic triad of the active site that is shown to be conserved also in the sequenced plant TG AtPgn1p which presents a low-sequence homology to the animal TGs; however, its core domain presents a folding similar to human TGM2 as the catalytic triad has a spatial position perfectly superimposable with TGM2 (Serafini-Fracassini et al. 2009). The cross-reactivity of the animal (nematode but more frequently assayed mammal) TG antibodies with plant TGs has been reported in several organs and type of plants (Del Duca et al. 1994, 2000a; Della Mea et al. 2007). A similarity between plant and animal TGs is also supported by the ability of their catalytic activity which can cross-recognize at least some of their respective substrates, thus suggesting a similarity also in their specificity (Del Duca and Serafini-Fracassini 2005; Serafini-Fracassini et al. 2009).
Amino acid composition
The aac of the three TG forms were compared allowing to suggest that they are molecules different each other, which, however, sometimes shared unexpected similarities with mammalian TGs. Phylogenetic tree analysis of the known animal TGs confirms that two main branches probably arose from a common ancestral gene: one lineage includes TG1 and FXIIIA, the second comprises the genes for erythrocyte band 4.2, TG2, TG3, TG5, TG6 and TG7. The band 4.2 is on a separate branch near to TG2 (Lorand and Graham 2003).
The tool AACompIdent, which allows to compare the aac of a protein against the Swissprot-Trembl databases, has been applied to H. t. composition and retrieved the closest proteins stored in databases. The experimental data showed that the H. t. forms had aac close to those of TGs annotated in Swissprot. The H. t. 75 kDa enzyme aac was practically identical (0.29 %) to that of the erythrocyte EPB42, which is an inactive TG form due to the absence of the TGs catalytic active site, but showing a general sequence homology with the active forms. On the contrary, the H. t. 75 kDa TG is active, as shown by the production of the glutamyl-derivatives reported in Fig. 2. Thus, the small difference could in part be related to the catalytic triad. In animal cells other inactive TGs are known; in the extracellular matrix (ECM) TG2 exerts a role in the communication between the ECM and intracellular signalling molecules. In many cases, after secretion, TG2 becomes a non-enzymically active structural component of the ECM, forming a close association with the glycoprotein FN and heparin sulphate proteoglycans important in cellular adhesion and migration (Verderio et al. 2003; Scarpellini et al. 2009).
The H. t. 75 kDa appears to be distinct in respect to the H. t. 85 kDa, thus suggesting that it is not a fragment of the last one. The 3D model for the H. t. 75 kDa form built adopting the animal TGM2 as template, (four TGM2 sequences were considered for the consensus sequence, see Table 3) showed a considerable homology especially in the catalytic domain. This model appears to be even more suitable than that proposed for PNG1_ARATH (Serafini-Fracassini et al. 2009; Tasco et al. 2003). The H. t. 85 kDa TG and the TGM2 were also very similar for their aac and for a very similar number of residues. TGM2 is a widespread mammalian enzyme which exerts multiple catalytic activities being tissue ubiquitous (Beninati et al. 2009; Lorand and Graham 2003; Griffin et al. 2002). TGM2 and H. t. 85 kDa TG appeared more similar to each other than TGM2 to EPB42_HUMAN and PNG1_ARATH (the 82 kDa purified recombinant AtPng1p).
The AtPng1p was codified on the sequence of an 86 kDa enzyme isolated from the microsomal fraction of Arabidopsis thaliana plant, in which also other double bands of lower mass were detected by western blotting (Della Mea et al. 2004a). The present data allowed us to hypothesise that also the structure of the H. t. 85 kDa TG and TGM2 molecules could be very similar. The aac of the H. t. 85 kDa enzyme shares a greater difference, with respect to TGM2, with the AtPng1p recombinant enzyme (PNG1_ARATH), even though their molecular mass is similar. A similar difference in aac was observed in respect of the Factor XIIIA subunit, a cytosolic and extracellular enzyme whose role is very specialised, related to blood coagulation and bone growth. These comparisons as well as their catalytic capacities suggest that the 75 and 85 kDa H. t. enzymes should be rather conserved during the evolution. The third enzyme (58 kDa), on the contrary, seems to have different characteristics. The 58 kDa one is the most frequently detected MW for the TGs in plants and not reported for animal ones. The H. t. 58 kDa form has a considerable difference in aac also with the known proteolysed 52 kDa fragment of human TG2 and 3 and with the 53 kDa fragment of the Arabidopsis thaliana recombinant enzyme, AtPng1p, whose N-ter fragment has been retrieved by homology (not shown). It cannot be excluded that the H. t. 58 kDa could derive by cleavage of the H. t. 75 or 85 kDa forms. This hypothesis could be supported by the presence in the same DEAE fraction of the 58 kDa of a band of around 20 kDa. Protein cleavage has been reported by Kim et al. (1993) for TG3 which, under denaturating conditions, is dissociated into two forms with 50 and 27 kDa. A protease-induced modulation of cellular TG activity has been proposed by Chung et al. (1988). In this view, Bures and Goldsmith (1978) have suggested an insoluble–soluble translocation mechanism by which TG might be compartmentalized changing its conformational state (Gundemir and Johnson 2009).
Unfortunately, the only other plant sequences found in database with the signature of TG (Q6KF70, as well as of the associated Q6KF61) and published to have a binding activity (Villalobos et al. 2004) did not show the characteristic active site nor an aac comparable to that of the TG family. Having a considerable difference with the H. t. 58 kDa, it is clear that the two proteins only share a similarity in the MW and location as both have been found, but not exclusively, in differentiated chloroplasts (Del Duca et al. 1994, 2000a; Villalobos et al. 2001; Sobieszczuk-Nowicka et al. 2007). Other protein sequences from various plants can be found in database classified as TGs only on the base of their homology with TG domain, but not experimentally assayed for the enzymatic activity. Thus, we have no bona fide sufficient elements to find similarities with confirmed TG forms. It can be hypothesised that this TG, which is active as confirmed by many experimental data, eventually was not codified by chromosomal DNA. If these proteins have been codified by the plastid genome, they should share a similarity with those of cyanobacteria, but this cannot be verified, as the sequences for TG unfortunately are not available in cyanobacteria genome.
Possible roles and cell location
It could be presumed that the multiple TG forms in H. t. sprouts exert different functions, possibly related to the type of cells where they are compartmentalized.
Meristems of H. t. etiolated sprout apices consist of a primary meristem protected by leaf primordia whose cells have etioplasts which, in the light, rapidly organize differentiated chloroplasts. Light, in fact, regulates the expression of the genes for several plastidial proteins concomitantly to the maturation of the thylakoids, a process in which TG seems to participate (Del Duca et al. 1993; Dondini et al. 2003; Della Mea et al. 2004b). During the light-induced greening of etioplasts of Cucumis sativus endosperm in addition to a 58 kDa form, also two 30 and 77 kDa bands were immuno detected in the membranes; the latter form dramatically decreased at the end of the differentiation to mature chloroplasts and appears involved in the conjugation of PAs to the organizing photosynthetic membrane system (Sobieszczuk-Nowicka et al. 2007). Thus, the similar 32 and 75 kDa TG forms detected could derive from the etioplasts of the H. t. apex leaflets.
A 58 kDa protein was the first and most frequently immuno-recognized protein in chloroplasts and leaf extracts by antibodies raised against different animal TGs (Del Duca et al. 1994, 1995; Della Mea et al. 2004b; Serafini-Fracassini and Del Duca 2008). The location of this enzyme in thylakoids, the identification of the photosynthetic antenna complexes as substrates and the formation of glutamyl-PAs suggest that a TG could be involved in the photosynthesis.
Also in tuber parenchyma grown in in vitro and exposed to light, two 58 and 90 kDa immuno-recognized SDS-PAGE bands were rapidly synthesized under light exposure (Del Duca et al. 2000a). In Nicotiana tabacum corolla, undergoing programmed cell death, the 58 kDa form, immuno-recognised by three antibodies raised against nematode, plant (Arabidopsis thaliana) and mammalian TGs has been detected in the soluble, microsomal, plastidial and cell wall fractions (Della Mea et al. 2007). Thus, the 58 kDa form is widespread either in meristematic or differentiated cells and is not exclusive of plastids.
The most apical part of etiolated sprouts, in addition to the staminal cells of the fundamental meristem, consists of meristematic cells characterized by a high rate of proliferation to give rise to the new stem, when tuber dormancy is released, as in our experiments. In these processes cytoskeletal proteins play a key role. Actin and tubulin are substrates of apple pollen purified 70 kDa TG (as well as of guinea pig liver TG) immuno-recognised by an anti-TG2 antibody, which controls the ability of actin and tubulin to assemble and interact with motor proteins thus regulating the development of pollen tubes (Del Duca et al. 2009). In addition, apple TGs of 70 and 75 kDa and their substrates co-localized on the pollen tube surface. TG-specific inhibitors and an anti-TG monoclonal antibody blocked pollen tube growth, consistent with a role of extracellular TG as modulator of cell wall building and strengthening (Di Sandro et al. 2010). A similar role could also be exerted by H. t. TGs in the new cell walls of the dividing cells of the H. t. apical meristem and in elongating derived cells.
An 86 kDa TG was isolated from the microsomal fraction of Arabidopsis (Della Mea et al. 2004a). A similar subcellular location in the H. t. sprout apices could be hypothesized as these cells are growing and differentiating; thus their Golgi apparatus is actively involved in the cell wall building; TG could be transported towards the sites where it will exert its function.
In conclusion, these data confirm that in H. t. immature cells there are at least three immuno-recognized TGs expressed and active. These TG forms are probably the same found also in the differentiated cells of other plant systems and thus are constitutive enzymes. Two of these TGs share a high degree of molecular mass and aac similarity with two mammalian TGs. This could suggest that these TGs are enzymes phylogenetically very conserved.
Abbreviations
- aa:
-
Amino acid
- aac:
-
Amino acid composition
- AtPng1p:
-
Arabidopsis thaliana peptide N-glycanase
- BCIP/NBT:
-
5-bromo-4-chloro-3-indolylphosphate/nitro blue tetrazolium
- DEAE:
-
Diethylaminoethyl
- DMC:
-
N,N’-dimethylcasein
- DTT:
-
Dithiotreitol
- FXIIIA:
-
Factor XIIIA subunit
- FN:
-
Fibronectin
- H. t :
-
Helianthus tuberosus
- PAs:
-
Polyamines
- PNG1_ARATH:
-
Peptide: N-glycanase_Arabidopsis thaliana
- Put:
-
Putrescine
- PVPP:
-
Polyvinylpolypyrrolidone
- Q6KF61 and Q6KF70:
-
Cloned maize sequences
- SDS-PAGE:
-
Sodium dodecyl-sulphate-gel electrophoresis
- SN:
-
Supernatant
- Spd:
-
Spermidine
- TBST:
-
Saline Tris-buffer containing 0.05 % Tween 80
- TCA:
-
Trichloroacetic acid
- TGAS_STRMB:
-
TG from Streptomyces mobaraensis
- TGs:
-
Transglutaminases
- TGM2:
-
Mammalian TG2
- Z-L-Gln-L-Leu:
-
N-benzyloxycarbonyl-L-γ-glutaminyl-L-leucine
- EPB42:
-
Erythrocyte membrane protein band 4.2
References
Beninati S, Piacentini M, Cocuzzi ET, Autuori F, Folk JE (1988) Covalent incorporation of polyamines as gamma-glutamyl derivatives into CHO cell protein. Biochim Biophys Acta 952:325–333
Beninati S, Abbruzzese A, Cardinali M (1993) Differences in the post-translational modification of proteins by polyamines between weakly and highly metastatic B16 melanoma cells. Int J Cancer 53:792–797
Beninati S, Bergamini CM, Piacentini M (2009) An overview of the first 50 years of transglutaminase research. Amino Acids 36:591–598
Bernet E, Claparols I, Dondini L, Santos MA, Serafini-Fracassini D, Torné JM (1999) Changes in polyamine content, arginine and ornithine decarboxilases and transglutaminase activities during ligh/dark phases of initial differentiation in maize calluses and their chloroplasts. Plant Physiol Biochem 37:899–909
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principles of protein-dye binding. Anal Biochem 78:248–254
Bures DM, Goldsmith LA (1978) Localization of transglutaminase in adult chicken epidermis. Arch Dermatol Res 262:329–332
Chhabra A, Verma A, Mehta K (2009) Tissue transglutaminase promotes or suppresses tumors depending on cell context. Anticancer Res 29:1909–1919
Chung SI, Chang SK, Cocuzzi ET, Folk JE, Kim HC, Lee SY, Martinet N, Nigra T, Sun HS (1988) Modulation of cellular transglutaminase: protease-induced activation. Adv Exp Med Biol 231:1–13
Del Duca S, Serafini-Fracassini D (2005) Transglutaminases of higher, lower plants and fungi. In: The Karger Group ‘Progress in Experimental and Tumor Research’ Basel (ed) Transglutaminases: the family of enzymes with diverse functions, vol 38, pp 223–247
Del Duca S, Favali A, Serafini Fracassini D, Pedrazzini R (1993) Transglutaminase-like activity during greening and growth of Helianthus tuberosus explants. Protoplasma 174:1–9
Del Duca S, Tidu V, Bassi R, Serafini-Fracassini D, Esposito C (1994) Identification of transglutaminase activity and its substrates in isolated chloroplast of Helianthus tuberosus. Planta 193:283–289
Del Duca S, Beninati S, Serafini-Fracassini D (1995) Polyamines in chloroplasts: identification of their glutamyl- and acetyl- derivatives. Biochem J 305:233–237
Del Duca S, Allué Creus J, D’Orazi D, Dondini L, Bregoli AM, Serafini-Fracassini D (2000a) Tuber vegetative stages and cell cycle in Helianthus tuberosus: protein pattern and their modification by spermidine. J Plant Physiol 156:17–25
Del Duca S, Dondini L, Della Mea M, Munoz de Rueda P, Serafini-Fracassini D (2000b) Factors affecting transglutaminase activity catalysing polyamine conjugation to endogenous substrates in the entire chloroplast. Plant Physiol Biochem 38:429–439
Del Duca S, Serafini-Fracassini D, Bonner PLR, Cresti M, Cai G (2009) Effects of post-translational modifications catalyzed by pollen transglutaminase on the functional properties of microtubules and actin filaments. Biochem J 418:651–664
Della Mea M, Caparrós-Ruiz D, Claparols I, Serafini-Fracassini D, Rigau J (2004a) AtPng1p. The first plant transglutaminase. Plant Physiol 135:2046–2054
Della Mea M, Di Sandro A, Dondini L, Del Duca S, Vantini F, Bergamini C, Bassi R, Serafini-Fracassini D (2004b) A Zea mays 39 kDa thylakoidal transglutaminase catalyses the modification by polyamines of light harvesting complex II in a light-dependent way. Planta 219:754–764
Della Mea M, De Filippis F, Genovesi V, Serafini Fracassini D, Del Duca S (2007) The acropetal wave of developmental cell death (DCD) of Nicotiana tabacum corolla is preceded by activation of transglutaminase in different cell compartments. Plant Physiol 144:1211–1222
Di Sandro A, Del Duca S, Verderio E, Hargreaves AJ, Scarpellini A, Cai G, Cresti M, Faleri C, Iorio R, Hirose S, Furutani Y, Coutts IGC, Griffin M, Bonner PLR, Serafini-Fracassini D (2010) An extracellular transglutaminase is required in apple pollen tube growth. Biochem J 429:261–271
Dondini L, Del Duca S, Dall’Agata L, Bassi R, Gastaldelli M, Della Mea M, Di Sandro A, Claparols I, Serafini-Fracassini D (2003) Suborganellar localisation and effect of light on Helianthus tuberosus chloroplast transglutaminases and their substrates. Planta 217:84–95
Folk JE (1980) Transglutaminases. Annu Rev Biochem 49:517–531
Folk JE, Chung SI (1985) Transglutaminases. Methods Enzymol 113:358–375
Folk JE, Park MH, Chung SI, Schrode J, Lester EP, Cooper HL (1980) Polyamines as physiological substrates for transglutaminases. J Biol Chem 255:3695–3700
Griffin M, Casadio R, Bergamini CM (2002) Transglutaminases: nature’s biological glues. Biochem J 368:377–396
Gundemir S, Johnson GV (2009) Intracellular localization and conformational state of transglutaminase 2: implications for cell death. PLoS ONE 4:e6123
Han BG, Cho JW, Cho YD, Jeong KC, Kim SY, Lee BI (2010) Crystal structure of human transglutaminase 2 in complex with adenosine triphosphate. Int J Biol Macromol 47:190–195
Kim HC, Lewis MS, Gorman JJ, Park SC, Girard JE, Folk JE, Chung SI (1990) Protransglutaminase E from guinea pig skin. Isolation and partial characterization. J Biol Chem 265:21971–21978
Kim IG, Gorman JJ, Park SC, Chung SI, Steinert MP (1993) The deduced sequence of the novel protransglutaminase E (TGase 3) of human and mouse. J Biol Chem 268:12682–12690
Laskowski RA, MacArthur MW, Moss DS, Thornton JM (1993) PROCHECK: a program to check the stereochemical quality of protein structures. J Appl Crystallogr 26:283–291
Lentini A, Abbruzzese A, Caraglia M, Marra M, Beninati S (2004) Protein-polyamine conjugation by transglutaminase in cancer cell differentiation: review article. Amino Acids 26:331–337
Lilley GR, Skill J, Griffin M, Bonner PL (1998) Detection of Ca2+-dependent transglutaminase activity in root and leaf tissue of monocotyledonous and dicotyledonous plants. Plant Physiol 117:1115–1123
Lorand L, Graham RM (2003) Transglutaminases: crosslinking enzymes with pleiotropic functions. Nat Rev Mol Cell Biol 4:140–156
Marti-Renom MA, Stuart A, Fiser A, Sánchez R, Melo F, Sali A (2000) Comparative protein structure modeling of genes and genomes. Annu Rev Biophys Biomol Struct 29:291–325
Mossetti U, Serafini-Fracassini D, Del Duca S (1987) Conjugated polyamines during dormancy and activation of tuber of Jerusalem artichoke. In: Schreiber K, Schuette HR, Sembdner G (eds) Conjugated plant hormones structure metabolism and function. Deutscher Verlag der Wissenschaften, Berlin, pp 369–375
Scarpellini A, Germack R, Lortat-Jacob H, Muramatsu T, Billett E, Johnson T, Verderio E (2009) Heparan sulfate proteoglycans are receptors for the cell-surface trafficking and biological activity of transglutaminase-2. J Biol Chem 284:18411–18423
Schägger H, von Jagow G (1987) Tricine-sodium dodecyl sulfate-polyacrylamide gel electrophoresis for the separation of proteins in the range from 1 to 100 kDa. Anal Biochem 166:368–379
Serafini-Fracassini D, Del Duca S (2008) Tranglutaminases: widespread cross-linking enzymes in plants. Ann Bot 102:145–152
Serafini-Fracassini D, Del Duca S, D’Orazi D (1988) First evidence for polyamine conjugation mediated by an enzymic activity in plants. Plant Physiol 87:757–761
Serafini-Fracassini D, Del Duca S, Torrigiani P (1989) Polyamine conjugation during the cell cycle of Helianthus tuberosus: non enzymatic and transglutaminase-like binding activity. Plant Physiol Biochem 27:659–668
Serafini-Fracassini D, Del Duca S, Beninati S (1995) Plant transglutaminases. Phytochemistry 40:355–365
Serafini-Fracassini D, Della Mea M, Tasco G, Casadio R, Del Duca S (2009) Plant and animal transglutaminases: do similar functions imply similar structures? Amino Acids 36:643–657
Sobieszczuk-Nowicka E, Di Sandro A, Del Duca S, Serafini-Fracassini D, Legocka J (2007) Plastid-membrane-associated polyamines and thylakoid transglutaminases during etioplast-to-chloroplast transformation stimulated by kinetin. Physiol Plant 130:590–600
Spitzer M, Fuellen G, Cullen P, Lorkowski S (2004) VisCoSe: visualization and comparison of consensus sequences. Bioinformatics 20:433–435
Tabolacci C, Lentini A, Provenzano B, Beninati S (2012) Evidences for a role of protein cross-links in transglutaminase-related disease. Amino Acids 42:975–986
Tasco G, Della Mea M, Serafini-Fracassini D, Casadio R (2003) Building a low resolution model of a transglutaminase domain of an hypothetical N-Glycanase from Arabidopsis thaliana. Amino Acids 25:197
Thompson JD, Higgins DG, Gibson TJ (1994) CLUSTALW: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22:4673–4680
Verderio E, Telci D, Okoye A, Melino G, Griffin M (2003) A novel RGD independent cell adhesion pathway mediated by fibronectin-bound tissue transglutaminase rescues cells from anoikis. J Biol Chem 278:42604–42614
Villalobos E, Torné JM, Rigau J, Ollés I, Claparols I, Santos M (2001) Immunogold localization of a transglutaminase related to grana development in different maize cell types. Protoplasma 216:155–163
Villalobos E, Santos M, Talavera D, Rodriguez-Falcon M, Torne JM (2004) Molecular cloning and characterization of a maize transglutaminase complementary DNA. Gene 336:93–104
Acknowledgments
We acknowledge the financial support provided by RFO 2009, 2010. Rat prostate gland TG was a kind gift of prof. C. Esposito, Salerno University.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Beninati, S., Iorio, R.A., Tasco, G. et al. Expression of different forms of transglutaminases by immature cells of Helianthus tuberosus sprout apices. Amino Acids 44, 271–283 (2013). https://doi.org/10.1007/s00726-012-1411-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00726-012-1411-y