Abstract
The increasing resistance of bacteria and fungi to currently available antibiotics is a major concern worldwide, leading to enormous efforts to develop new antibiotics with new modes of actions. Antibacterial peptide CM4 (ABP-CM4) is a small cationic peptide with broad-spectrum activities against bacteria, fungi, and tumor cells, which may possibly be used as a promising candidate for a new antibiotic. For pharmaceutical applications, a large quantity of antimicrobial peptides needs to be produced economically. In this communication, the progress in the structural characteristics, heterologous production, and biological evaluation of ABP-CM4 are reviewed.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The increasing resistance of bacteria and fungi to currently available antibiotics is a major concern worldwide, leading to enormous efforts to develop new antibiotics with new modes of actions (Makovitzki et al. 2006). One potential source of novel antibiotics is the antimicrobial peptides (AMPs) which are relatively small molecules that are less than 100 amino acids in length and have a broad spectrum of antimicrobial activity. They serve as an ancient defense mechanism against pathogenic microorganisms that easily come in contact with the host through the environment (Sugiarto and Yu 2004). AMPs are now considered a fundamental component of the innate immune system (Radek and Gallo 2007). In contrast to the adaptive immune response, the innate immune response is immediate, nonspecific, and diverse (Gallo and Nizet 2003; Ganz 2003; Nizet and Gallo 2003).
In the past decades, a large number of AMPs have been isolated and purified from various species of organisms, such as defensins, cathelicidins, histatins, and cecropins (Koczulla and Bals 2003). Defensins are a broadly dispersed group of cationic peptides containing cysteine-rich conserved motifs originally identified in human and rabbit neutrophils (Radek and Gallo 2007). In contrast to the defensins, Cathelicidins were identified solely in mammalian species (Dorschner et al. 2001; Gallo et al. 1997). PR-39 became the first AMP found in mammalian skin, specifically porcine wound fluid (Gallo et al. 1994). Histatins are a group of AMPs, found in the saliva of man and some higher primates, which possess antifungal properties (Helmerhorst et al. 1997). Cecropins are positively charged peptides that were originally isolated from the blood lymph of the giant silk moth (Hultmark et al.1980). Cecropins have the ability to form specific amphipathic alpha-helices which allow them to target nonpolar lipid cell membranes. Upon membrane targeting, they form ion-permeable channels subsequently resulting in cell depolarization, irreversible cytolysis, and finally death (Boman 2003). ABP-CM4, an antibacterial peptide isolated from the hemolymph of the silkworm Bombyx mori, belongs to the cecropins family (Tu et al. 1989). ABP-CM4 kills bacteria, tumors, and fungi by permeabilizing the cell membranes without being toxic to mammalian cells (Chen et al. 2010; Zhang et al. 1997; Xu and Zhang 2001). It would be important for control of resistant pathogen bacterial and fungal infections, but would allow conducting additional studies on their molecular interactions and antimicrobial mechanisms, as well on their eventual use in public health care. Therefore, the purpose of this communication is to revise the state of the art on the study of ABP-CM4 concerning its structural characteristics, expression systems, and biological activities of the recombinant products.
Structural characteristics of ABP-CM4
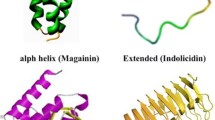
AMPs are small, positively charged, amphipathic molecules (which possess both hydrophobic and hydrophilic regions) of variable amino acid composition and length (6–100 amino acids). Based on their secondary structure, AMPs are grouped into four major classes: β-sheet, α-helical, loop, and extended peptides (Giuliani et al. 2007). The molecular mass of ABP-CM4 was 3876.64 confirmed by MALDI-TOF MS. ABP-CM4 is a cationic peptide containing Arg1,16, Lys3,6,7,10,21, Glu9, and Asp17, which results in a net charge of +5 at neutral pH (Chen et al. 2010).
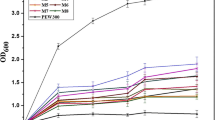
The secondary structure and three-dimensional structure prediction of ABP-CM4 showed the N-terminus to be a typical amphipathic α-helix (Jones 1999; Ward et al. 2004; Zdobnov and Apweiler 2001), which was an important structural parameter for anticancer activity (Yang et al. 2004). Charged residues are grouped on one face of the helix and hydrophobic residues are grouped on the opposite face (Fig. 1a). Then the secondary structure of ABP-CM4 was estimated by CD spectroscopy in a solution containing different concentrations of 2,2,2-trifluoroethanol (TFE) from 0 to 50% (v/v). It was found from the CD spectra that ABP-CM4 displayed a negative band at 200 nm in water, suggesting largely random coil conformation. However, in the presence of 20% TFE in water, the shape of the curves was increasingly characteristic of α-helical secondary structure. This peptide exhibited a positive band at 195 nm and two negative bands at 208 and 222 nm in 30 and 50% TFE in water, suggesting a well-defined α-helical conformation (Fig. 1b). Then the CD spectra of ABP-CM4 in POPG vesicles (at peptide:lipid ratios of 1:25 and 1:50) were estimated (Supplementary Fig. 1S). These spectra suggest that the peptide does not have the same structure in POPG or in TFE. These spectral differences were (1) the dichroic intensities in 200 nm are very different (positive in POPG but negative in TFE); (2) the negative band in 222 nm is displaced to a larger wavelength in POPG than in TFE; and (3) the α-helical signature is weaker in POPG than in TFE. The CD study confirms the molecular feature of ABP-CM4 as a linear α-helix in membrane-mimicking solvents, which is consistent with other α-helix ABPs (Bechinger et al. 1997; Wu et al. 2009). Owing to its special sequence and dimensional structure, ABP-CM4 is insensitive to denaturants, detergents, organic solvents, extreme temperatures, and pH. The results of the thermal stability test showed that temperatures as high as 95°C for 30 min had no influence on the activity of ABP-CM4; however, temperatures above 100°C (30 min) decreased the antimicrobial activity of ABP-CM4 by 15.5%. Varying the pH from 2.0 to 10.0 had no effect on the activity of ABP-CM4; however, pH values above 12.0 decreased the antimicrobial activity of ABP-CM4 by 12.5%. And 25.5% loss of potency was observed at pH 12.0 above 100°C (30 min) (Fig. 2 and Supplementary Fig. 2S). These features strengthen its potential as a microbicide, as well as providing a simple heat precipitation process to separate it from other host proteins when it is expressed in heterologous bacteria or cells.
The secondary structure analysis and three-dimensional structure prediction of ABP-CM4. a Secondary structure analysis of ABP-CM4 by CD spectroscopy. Far-UV CD spectra were conducted at room temperature in TFE/water mixtures at various concentrations of TFE in the range of 0–50% (v/v). Adopted from Chen et al. (2010). b The secondary and three-dimensional structure prediction of ABP-CM4. H helix, E extended-beta, E. coil
Heterologous expression of ABP-CM4
ABP-CM4 is a promising candidate for a new antibiotic. For pharmaceutical applications, a large quantity of AMPs needs to be produced economically. A few years ago, the ways of obtaining ABP-CM4 were (1) isolation from the silkworm, which requires large amounts of material from the source with the problem of having very low yields of the interest peptide; and (2) chemical synthesis of peptides, which generate high costs. Therefore, genetic engineering became a great strategy to produce large amounts of interesting ABP-CM4 with low cost, to produce possible variants using site-directed mutagenesis, and to elucidate the antimicrobial mechanisms and to improve the final yields.
Expression of ABP-CM4 in Escherichia coli
The Escherichia coli expression system is still the most commonly used because of its high level of expression, the relative simplicity of the DNA manipulations, and the short time required to produce a product. However, alleviating the toxicity and proteolytic degradation of an expressed antibacterial peptide in the E. coli system and purifying the peptide easily are common problems. Many fusion partners working as a carrier have been used to express and purify ABP-CM4. The function of the carrier protein is to protect the small cationic peptides against proteolytic degradation (Piers et al. 1993). In addition, the fusion system provides several advantages: (1) they protect the hybrid when proteases are exported to the external environment, (2) the carrier protein might show affinity for a specific ligand, which enables easy purification not only might prevent AMP toxicity against the host cell, and (3) the carrier protein can finally be released by specific proteolytic action, leaving no extra amino acids on the C- or N-terminal regions.
Earlier attempts to produce recombinant ABP-CM4 in our laboratory using the Escherichia coli expression system with the PET-28a vector and synthesized as a fusion protein have been only partially successful for an extra fusion partner left at the N-terminus (Li et al. 1999). Subsequently, we synthesized ABP-CM4 using the plasmid of pET-32a, with T7 promoter and thioredoxin (TrxA) as a fusion partner because of its reported compatibility with expression of foreign proteins in soluble form (LaVallie et al. 2000). However, this expression system yielded only 1.2–1.4 mg target protein per liter of culture and an extra proline or glycine left at the N-terminus (Li et al. 2007; Zhou and Zhang 2009). Substitution of the TrxA fusion partner with intein-mediated expression system resulted in intact ABP-CM4, but this expression system yielded only 2.1 mg target protein per 6 g wet weight (Chen et al. 2008a, b). In addition, the purification process based on HPLC could not meet the requirement of mechanism determination and other large-scale operations.
To increase the yield of ABP-CM4, inclusion body (IB) expression in the cytoplasm of E. coli was attempted with Npro mutant (EDDIE) fusion technology (Cheng et al. 2010). Following refolding by a conventional process and purification by Ni2+-chelating chromatography column and cation exchange chromatography column, about 6 mg/L of ABP-CM4 was produced in shaking flask culture. Although it offers an alternative way of producing large quantities of antibacterial peptides, the cleavage rate of His-EDDIE-CM4 fusion protein reached only 50%, which was caused by the first amino acid of ABP-CM4. How to enhance the cleavage rate of His-EDDIECM4 fusion protein need be further studied. The heterogeneity of ABP-CM4 from IB exposes the weakness of this type of expression. Another new technology of biosynthesis of ABP-CM4 using the elastin-like peptide (ELP) tags was also tried, but the low yield and cleavage rate were disappointing (Shen et al. 2010). Finally, we turned our attention to the chaperon-fused expression system. Fortunately, this system with hexahistidine and small ubiquitin-related modifier (SUMO) double-tagged ABP-CM4 leads to a soluble fusion protein in the cytoplasm of E. coli at a level of >18% of the total soluble protein. With this construct, intact and native ABP-CM4 can be rapidly purified in a soluble and biologically active form by two rounds of affinity chromatography with one round of SUMO protease digestion in between according to Gao et al. (2010). In a 5-L bioreactor using conventional fed-batch culture (FBC) in a manner similar to that described previously (Ma et al. 2006), about 24 mg/L native ABP-CM4 was afforded (Li et al. 2009). On this basis, we described a procedure of producing a larger quantity of recombinant ABP-CM4 by multimerization of ABP-CM4 gene with the fusion partner TrxA or SUMO, about 26–48 mg/L of ABP-CM4 was produced in shaking flask culture (Zhou et al. 2009; Li et al. 2011). These were the highest yield of ABP-CM4 reported to date. Overexpression of soluble SUMO–CM4 in the cytoplasm of E. coli to obtain intact, homogeneous, and activated ABP-CM4 by simple downstream processes makes this system promising for practical applications. By comparison of the expression of ABP-CM4 with other cecropin peptides (Campo et al. 2008; Chen et al. 2009; Hong et al. 2008; Jan et al. 2010; Zhang et al. 2010), the SUMO fusion technology potentially could be employed as a new way for the production of recombinant cecropin peptides (Supplementary Table 1S).
In eukaryotic host
Pichia pastoris is an alternative host that, like E. coli, can be grown cheaply and rapidly, possesses certain posttranslational modification pathways, is able to secrete more efficiently, and permits the production of r-proteins without intense process development. Introduction of the coding sequence for ABP-CM4 into P. pastoris demonstrates the potential for expression of ABP-CM4 in a eukaryotic system. After inducing about 72 h with 0.5% methanol at 20°C, supplied with 2% casamino acids to avoid proteolysis, approximately 40 mg ABP-CM4 was secreted into 1 L of medium. Recombinant ABP-CM4 was purified through size-exclusion chromatography and 15 mg pure active ABP-CM4 was obtained from 1 L culture (Zhang et al. 2006). During the production of ABP-CM4, we found that the antibacterial activity of secreted ABP-CM4 obtained by liquid fermentation, 250 rpm, 20°C in YPD media for 144 h, was dropping along the culture time, from 17 mm inhibition zone at 72 h to 9 mm zone at 144 h. The diminished antibacterial activity was correlated with the low stability of ABP-CM4 at 20°C and proteolytic activity inside the fermentation media more than with the inhibition of P. pastoris growth due to ABP-CM4 expression. Furthermore, the yield in P. pastoris was 15 mg/L, which is low compared with cytoplasmic expression in E. coli. Another eukaryotic expression of ABP-CM4 with green fluorescent protein (GFP) fusion protein using the expression vectors pcDNA3 was also tried, but the low yield and an extra GFP fusion partner left at the N-terminus were disappointing (Chen et al. 2008a, b).
In conclusion, ABP-CM4, a promising microbicide candidate with a special primary structure that is sensitive to proteolytic activity, has been expressed in E. coli, yeast, and human myeloid leukemia K562 cells (Table 1). At this moment though, the soluble expression in the cytoplasm of E. coli might be the only feasible option.
Antimicrobial and anticancer activity assays of ABP-CM4
The most intriguing feature of ABP-CM4 is its destructive effect on bacteria, fungi, and tumors without damaging normal cells.
Antibacterial activity assays
The antibacterial activity of the ABP-CM4 was evaluated by determining its minimal inhibitory concentration (MIC) against selected bacteria (Lee et al. 2002). The results indicated that recombinant ABP-CM4 and native ABP-CM4 have almost the same antimicrobial activity against E. coli K12D31, Salmonella spp. and P. aeruginosa (Table 2). The antibiotic mechanism of amphipathic antibacterial peptides with α-helical structures is not clearly understood. Cell structure disrupted by pore formation (Ludtke et al. 1996; Shai and Oren 2001) or ion channel generation (Huang 2000) maybe is the most likely mechanism. In our study, ABP-CM4 can kill the bacteria by forming transmembrane pores on the biofilm (Fig. 3).
Scanning electron micrographs (SEM, X5, 000) of ABP-CM4’s effect on bacteria surface morphology and live/dead assay. a SEM of E. coli K12D31 incubated with PBS. b SEM of E. coli K12D31 incubated with 10 μM ABP-CM4. c Live/dead staining assay of normal E. coli K12D31. d Live/dead staining assay of E. coli K12D31 incubated with 10 μM ABP-CM4
Antifungal activity assays
The ABP-CM4 showed a twofold and fourfold greater antifungal activity against A. niger than did cecropin A (Delucca et al. 1997) and cecropin B (Delucca et al. 1998), and was also more effective against A. niger than AMPs from mammals, such as defensins (DeLucca and Walsh 1999). In addition to A. niger, ABP-CM4 is also effective against T. viride, G. saubinetii, P. chrysogenum (Table 2).
Lee et al. (2002) reported that HP (2-20) may exert its antifungal activity by disrupting the structure of the cell membrane via pore formation or directly by interacting with the lipid bilayer in a salt-dependent manner. In our study, the cell wall regeneration capacity of protoplasts treated with ABP-CM4 was much lower than that of the control and suggested that the peptide act on the fungal plasma membrane. These results demonstrated that the prime target of ABP-CM4 action was the plasma membrane, not the cell wall. ABP-CM4 antifungal activity is characterized by the disruption of cell membranes and the cellular cytoskeleton, internal structural changes within the cell including decreased mitochondrial integrity, and binding to fungal DNA (Zhang et al. 2008). The sum of these interactions can lead to cell death (Fig. 4).
Scanning electron micrographs (SEM, X3, 000) and Transmission electron micrographs (TEM, X12, 000) of ABP-CM4’s effect on cell surface morphology and cellular organelles. A. niger cells were incubated with PBS (a) or 8 μM ABP-CM4 at 30°C for 16 h (b), normal ultrastructure of A. niger (c) or A. niger treated with ABP-CM4 for 8 h (d). Adopted from Zhang et al. (2008)
Anticancer activity assays
Recently, a growing number of studies have shown that some of the cationic ABPs, which are toxic to bacteria but not to normalmammalian cells, exhibit a broad spectrum of cytotoxicity against cancer cells (Hoskin and Ramamoorthy 2008). Until now, several amphiphilic helical peptides, such as BMAP-27, BMAP-28, cecropin B, maganins, LL-37, and aurein 1.2, exhibited anticancer activity. Our studies have shown ABP-CM4 has a selective anticancer activity in several leukemia cell lines but does not affect normal cells (Table 3). Difference of binding activity between leukemia cells and normal cells results from membrane differences implicated in contributing to the selective cytotoxicity (Chen et al. 2010).
So far, at least three anticancer mechanisms have been proposed, including (1) cell membrane lytic effect; (2) activation of intrinsic pathways of apoptosis via mitochondrial membrane disruption; (3) certain peptides are potent inhibitors of blood vessel development, which is associated with tumor progression (Zaiou 2007). In our study, ABP-CM4 can disturb the tumor cellular membrane leading to leakage (Chen et al.2010; Wang et al. 1998; Zhang et al. 1997). By comparison of the anticancer activities of ABP-CM4 with other known antitumor peptides (Chen et al. 2005; Eliassen et al. 2006; Lehmann et al. 2006; Müller et al. 2002; Papo et al. 2006; Sharma 1992; Shin et al. 1999), most antitumor peptides are membrane-targeted leading to cell lysis (supplementary Table 2S), which shows potential for synergy with current cancer treatments. In addition to leukemia cells, ABP-CM4 is also effective against the growth of SHG-44, HeLa, and HepG2 cells (Cheng et al. 2010; Li et al. 2010). Our results indicated that ABP-CM4 has the potential for development as a novel anticancer agent.
Conclusions and perspectives
ABP-CM4 is a promising microbicide candidate that was discovered 22 years ago. It has displayed an impressive potential as a microbicide and could even be a novel anticancer agent. Using the present technology, it may be possible to produce ABP-CM4 on a large scale and at low cost. The soluble cytoplasm expression of native ABP-CM4, plus the established know-how of the bioprocesses of E. coli produced biologics in the industrial sector, might provide a ready solution for the manufacture of this microbicide candidate. Other approaches provide a backup for future manufacturing of the drug.
Recent studies have demonstrated that ABP-CM4 has the potential to inhibit cellular cytokine and NO released by binding directly to LPS or by blocking the binding of LPS to LPS-binding protein (Lin et al. 2008). This makes it an attractive drug candidate for treatment of endotoxin shock and sepsis caused by bacterial infection. Further studies are needed to elucidate the exact mechanism involved in its cytotoxicity and its selectivity tested in different assay and pharmacological models.
References
Bechinger B (1997) Structure and functions of channel-forming peptides: magainins, cecropins, melittin and alamethicin. J Membr Biol 156:197–211. doi:10.1007/s002329900201
Boman HG (2003) Antibacterial peptides: basic facts and emerging concepts. J Intern Med 254:197–215. doi:10.1046/j.1365-2796.2003.01228.x
Campo S, Manrique S, García-Martínez J, San Segundo B (2008) Production of cecropin A in transgenic rice plants has an impact on host gene expression. Plant Biotechnol J 6:585–608. doi:10.1111/j.1467-7652.2008.00339.x
Chen J, Xu XM, Underhill CB, Yang S, Wang L, Chen Y, Hong S, Creswell K, Zhang L (2005) Tachyplesin activates the classic complement pathway to kill tumor cells. Cancer Res 65:4614–4622. doi:10.1158/0008-5472.CAN-04-2253
Chen YQ, Zhang SQ, Jiao B, Qiu W, Zhang J (2008a) Construction and expression of eukaryotic expression vector of Antibacterial Peptide CM4 and Green Fluorescent Protein fusion gene. Lett Biotechnol 19:8–10. doi:cnki:SUN:SWTX.0.2008-01-005
Chen YQ, Zhang SQ, Li BC, Qiu W, Jiao B, Zhang J, Diao ZY (2008b) Expression of a cytotoxic cationic antibacterial peptide in Escherichia coli using two fusion partners. Protein Expr Purif 57:303–311. doi:10.1016/j.pep.2007.09.012
Chen X, Zhu F, Cao Y, Qiao S (2009) Novel expression vector for secretion of cecropin AD in Bacillus subtilis with enhanced antimicrobial activity. Antimicrob Agents Chemother 53:3683–3689. doi:10.1128/AAC.00251-09
Chen YQ, Min C, Sang M, Han YY, Ma XA, Xue XQ, Zhang SQ (2010) A cationic amphiphilic peptide ABP-CM4 exhibits selective cytotoxicity against leukemia cells. Peptides 31:1504–1510. doi:10.1016/j.peptides.2010.05.010
Cheng XW, Lu WG, Zhang SQ, Cao P (2010) Expression and purification of antimicrobial peptide CM4 by N-pro fusion technology in E. coli. Amino Acids 39:1545–1552. doi:10.1007/s00726-010-0625-0
Delucca AJ, Walsh TJ (1999) Antifungal peptides: novel therapeutic compounds against emerging pathogens. Antimicrob Agents Chemother 43:1–11
Delucca AJ, Bland JM, Jacks TJ, Grimm C, Cleveland TE, Walsh TJ (1997) Fungicidal activity of cecropin A. Antimicrob Agents Chemother 41:481–483
Delucca AJ, Bland JM, Jacks TJ, Grimm C, Walsh TJ (1998) Fungicidal and binding properties of the natural peptides cecropin B and dermaseptin. Med Mycol 36:291–298. doi:10.1080/02681219880000461
Dorschner RA, Pestonjamasp VK, Tamakuwala S, Ohtake T, Rudisill J, Nizet V, Agerberth B, Gudmundsson GH, Gallo RL (2001) Cutaneous injury induces the release of cathelicidin anti-microbial peptides active against group A Streptococcus. J Invest Dermatol 117:91–97. doi:10.1046/j.1523-1747.2001.01340.x
Eliassen LT, Berge G, Leknessund A, Wikman M, Lindin I, Løkke C, Ponthan F, Johnsen JI, Sveinbjørnsson B, Kogner P, Flaegstad T, Rekdal Ø (2006) The antimicrobial peptide, Lactoferricin B, is cytotoxic to neuroblastoma cells in vitro and inhibits xenograft growth in vivo. Int J Cancer 119:493–500. doi:10.1002/ijc.21886
Gallo RL, Nizet V (2003) Endogenous production of antimicrobial peptides in innate immunity and human disease. Curr Allergy Asthma Rep 3:402–409
Gallo RL, Ono M, Povsic T, Page C, Eriksson E, Klagsbrun M, Bernfield M (1994) Syndecans, cell surface heparan sulfate proteoglycans, are induced by a proline-rich antimicrobial peptide from wounds. Proc Natl Acad Sci USA 91:11035–11039
Gallo RL, Kim KJ, Bernfield M, Kozak CA, Zanetti M, Merluzzi L, Gennaro R (1997) Identification of CRAMP, a cathelin-related antimicrobial peptide expressed in the embryonic and adult mouse. J Biol Chem 272:13088–13093. doi:10.1074/jbc.272.20.13088
Ganz T (2003) Defensins: antimicrobial peptides of innate immunity. Nat Rev Immunol 3:710–720. doi:10.1038/nri1180
Gao XL, Chen W, Guo CW, Qian CW, Liu G, Ge F, Huang YD, Kitazato K, Wang YF, Xiong S (2010) Soluble cytoplasmic expression, rapid purification, and characterization of cyanovirin-N as a His-SUMO fusion. Appl Microbiol Biotechnol 85:1051–1060. doi:10.1007/s00253-009-2078-5
Giuliani A, Pirri G, Nicoletto S (2007) Antimicrobial peptides: an overview of a promising class of therapeutics. Cent Eur J Biol 2:1–33. doi:10.2478/s11535-007-0010-5
Helmerhorst EJ, Van t’Hof W, Veerman EC, Simons-Smit I, Nieuw Amerongen AV (1997) Synthetic histatin analogues with broad spectrum antimicrobial activity. Biochem J 326:39–45
Hong SM, Kusakabe T, Lee JM, Tatsuke T, Kawaguchi Y, Kang MW, Kang SW, Kim KA, Nho SK (2008) Structure and expression analysis of the cecropin-E gene from the silkworm, Bombyx mori. Biosci Biotechnol Biochem 72:1992–1998. doi:10.1271/bbb.80082
Hoskin DW, Ramamoorthy A (2008) Studies on anticancer activities of antimicrobial peptides. Biochim Biophys Acta 1778:357–375. doi:10.1016/j.bbamem.2007.11.008
Huang HW (2000) Action of antimicrobial peptides: two-state model. Biochemistry 39:8347–8352. doi:10.1021/bi000946l
Hultmark D, Steiner H, Rasmuson T, Boman HG (1980) Insect immunity purification and properties of three inducible bactericidal proteins from hemolymph of immunized pupae of Hyalophora cecropia. Eur J Biochem 106:7–16. doi:10.1111/j.1432-1033.1980.tb05991.x
Jan PS, Huang HY, Chen HM (2010) Expression of a synthesized gene encoding cationic peptide cecropin B in transgenic tomato plants protects against bacterial diseases. Appl Environ Microbiol 76:769–775. doi:10.1128/AEM.00698-09
Jones DT (1999) Protein secondary structure prediction based on position-specific scoring matrices. J Mol Biol 292:195–202. doi:10.1006/jmbi.1999.3091
Koczulla AR, Bals R (2003) Antimicrobial peptides: current status and therapeutic potential. Drugs 63:389–406
LaVallie ER, Lu ZJ, Diblasio-Smith EA, Collins-Racie LA, McCoy JM (2000) Thioredoxin as a fusion partner for production of soluble recombinant proteins in Escherichia coli. Methods Enzymol 326:322–340
Lee DG, Park Y, Kim HN, Kim HK, Kim PI, Choi BH, Hahm KS (2002) Antifungal mechanism of an antimicrobial peptide, HP (2–20), derived from N-terminus of Helicobacter pylori ribosomal protein L1 against Candida albicans. Biochem Biophys Res Commun 291:1006–1013. doi:10.1006/bbrc.2002.6548
Lehmann J, Retz M, Sidhu SS, Suttmann H, Sell M, Paulsen F, Harder J, Unteregger G, Stöckle M (2006) Antitumor activity of the antimicrobial peptide magainin II against bladder cancer cell lines. Eur Urol 50:141–147. doi:10.1016/j.eururo.2005.12.043
Li XL, Dai ZY, Zhang SQ, Zhang LY (1999) Studies on the changes of CM4 gene structure and the expression products in E. coli. Acta Biochim Biophys Sinica 15:387–391. doi:cnki:ISSN:1007-7626.0.1999-03-008
Li BC, Zhang SQ, Dan WB, Chen YQ, Cao P (2007) Expression in Escherichia coli and purification of bioactive antibacterial peptide ABP-CM4 from the Chinese silk worm, Bombyx mori. Biotechnol Lett 29:1031–1036. doi:10.1007/s10529-007-9351-4
Li JF, Zhang J, Song R, Zhang JX, Shen Y, Zhang SQ (2009) Production of a cytotoxic cationic antibacterial peptide in Escherichia coli using SUMO fusion partner. Appl Microbiol Biotechnol 84:383–388. doi:10.1007/s00253-009-2109-2
Li NN, Liu P, Chen SJ, Lin QP, Zhou LF, Zhang SQ (2010) Construction and expression of a novel bioactive IFN-α2b/CM4 fusion protein in Escherichia coli. Microbiol Res 165:116–121. doi:10.1016/j.micres.2009.01.002
Li JF, Zhang J, Zhang Z, Kang CT, Zhang SQ (2011) SUMO mediating fusion expression of antimicrobial peptide CM4 from two joined genes in Escherichia coli. Curr Microbiol 62:296–300. doi:10.1007/s00284-010-9705-3
Lin QP, Zhou LF, Li NN, Chen YQ, Li BC, Cai YF, Zhang SQ (2008) Lipopolysaccharide neutralization by the antibacterial peptide CM4. Eur J Pharmacol 596:160–165. doi:10.1016/j.ejphar.2008.08.017
Ludtke SJ, He K, Heller WT, Harroun TA, Yang L, Huang HW (1996) Membrane pores induced by magainin. Biochemistry 35:13723–13728. doi:10.1021/bi9620621
Ma XY, Zheng WY, Wei DZ, Ma YS, Wang TW, Wang JZ, Liu QH, Yang SL (2006) High-level expression, purification and pro-apoptosis activity of HIV-TAT-survivin (T34A) mutant to cancer cells in vitro. J Biotechnol 123:367–378. doi:10.1016/j.jbiotec.2005.11.018
Makovitzki A, Avrahami D, Shai Y (2006) Ultrashort antibacterial and antifungal lipopeptides. Proc Natl Acad Sci USA 103:15997–16002. doi:10.1073/pnas.0606129103
Müller CA, Markovic-Lipkovski J, Klatt T, Gamper J, Schwarz G, Beck H, Deeg M, Kalbacher H, Widmann S, Wessels JT, Becker V, Müller GA, Flad T (2002) Human α-defensins HNPs-1, -2, and-3 in renal cell carcinoma. Influences on tumor cell proliferation. Am J Pathol 160:1311–1324
Nizet V, Gallo RL (2003) Cathelicidins and innate defense against invasive bacterial infection. Scand J Infect Dis 35:670–676. doi:10.1080/00365540310015629
Papo N, Seger D, Makovitzki A, Kalchenko V, Eshhar Z, Degani H, Shai Y (2006) Inhibition of tumor growth and elimination of multiple metastases in human prostate and breast xenografts by systemic inoculation of a host defense-like lytic peptide. Cancer Res 66:5371–5378. doi:10.1158/0008-5472
Piers KL, Brown MH, Hancock REW (1993) Recombinant DNA procedures for producing small antimicrobial cationic peptides in bacteria. Gene 134:7–13. doi:10.1016/0378-1119(93)90168-3
Radek K, Gallo R (2007) Antimicrobial peptides: natural effectors of the innate immune system. Semin Immunopathol 29:27–43. doi:10.1007/s00281-007-0064-5
Shai Y, Oren Z (2001) From ‘‘carpet’’ mechanism to de-novo designed diastereomeric cell-selective antimicrobial peptides. Peptides 22:1629–1641. doi:10.1016/S0196-9781(01)00498-3
Sharma SV (1992) Melittin resistance: a counterselection for ras transformation. Oncogene 7:193–201
Shen Y, Ai HX, Song R, Liang ZN, Li JF, Zhang SQ (2010) Expression and purification of moricin CM4 and human beta-defensins 4 in Escherichia coli using a new technology. Microbiol Res 165:713–718. doi:10.1016/j.micres.2010.01.002
Shin SY, Kang JH, Hahm KS (1999) Structure-antibacterial, antitumor and hemolytic activity relationships of cecropin A-magainin 2 and cecropin A-melittin hybrid peptides. J Pept Res 53:82–90. doi:10.1111/j.1399-3011.1999.tb01620.x
Sugiarto H, Yu PL (2004) Avian antimicrobial peptides: the defense role of beta-defensins. Biochem Biophys Res Commun 323:721–727. doi:10.1016/j.bbrc.2004.08.162
Tu YZ, Zhang SQ, Xu YS (1989) Separation, purification of antibacterial CM4 and the research of the structure and character. Sci China B 32:473–480. doi:cnki:ISSN:1006-9291.0.1989-09-006
Wang F, Zhang SQ, Dai ZY (1998) Studies on the action of the ABP-CM4 anti K562 cancer cells by SCGE. Prog Biochem Biophys 1:64–67. doi:cnki:ISSN:1000-3282.0.1998-01-015
Ward JJ, Sodhi JS, McGuffin LJ, Buxton BF, Jones DT (2004) Prediction and functional analysis of native disorder in proteins from the three kingdoms of life. J Mol Biol 337:635–645. doi:10.1016/j.jmb.2004.02.002
Wu JM, Jan PS, Yu HC, Haung HY, Fang HJ, Chang YI, Cheng JW, Chen HM (2009) Structure and function of a custom anticancer peptide, CB1a. Peptides 30:839–848. doi:10.1016/j.peptides.2009.02.004
Xu J, Zhang SQ (2001) The research of the mechanism of antibacterial peptide CM4 component against A. parasiticus. Prog Nat Sci 11:1263–1267. doi:cnki:ISSN:1002-0071.0.2001-12-004
Yang N, Strom MB, Mekonnen SM, Svendsen JS, Rekdal O (2004) The effects of shortening lactoferrin derived peptides against tumour cells, bacteria and normal human cells. J Pept Sci 10:37–46. doi:10.1002/psc.470
Zaiou M (2007) Multifunctional antimicrobial peptides: therapeutic targets in several human diseases. J Mol Med 85:317–329. doi:10.1007/s00109-006-0143-4
Zdobnov EM, Apweiler R (2001) InterProScan-an integration platform for the signature-recognition methods in InterPro. Bioinformatics 17:847–848. doi:10.1093/bioinformatics/17.9.847
Zhang SQ, Jia HW, Dai ZY (1997) Ultrastructure observation of K562 leukemia cells treated with antibacterial peptide CM4 component. Prog Biochem Biophys 24:159–163. doi:cnki:ISSN:1000-3282.0.1997-02-015
Zhang J, Zhang SQ, Wu X, Chen YQ, Diao ZY (2006) Expression and characterization of antimicrobial peptide ABP-CM4 in methylotrophic yeast Pichia pastoris. Proc Biochem 41:251–256. doi:10.1016/j.procbio.2005.06.030
Zhang J, Wu X, Zhang SQ (2008) Antifungal mechanism of antibacterial peptide, ABPCM4, from Bombyx mori against Aspergillus niger. Biotechnol Lett 30:2157–2163. doi:10.1007/s10529-008-9819-x
Zhang YP, Fan T, Ma ZY, Dai JJ, Wang CY, Wu CH, Tian SF, Xiao LZ (2010) Cloning and construction of the expression vector of antimicrobial peptide Cecropin D gene from Bombyx mori. North Seric 31:18–21. doi:cnki:SUN:CYKE.0.2011-02-012
Zhou L, Zhang SQ (2009) TrxA mediating fusion expression of antimicrobial peptide CM4 from multiple joined genes in Escherichia coli. Protein Expr Purif 64:225–230. doi:10.1016/j.pep.2008.11.006
Zhou LF, Lin QP, Li BC, Li NN, Zhang SQ (2009) Expression and purification the antimicrobial peptide CM4 in Escherichia coli. Biotechnol Lett 31:437–441. doi:10.1007/s10529-008-9893-0
Acknowledgments
Published data referred to in this manuscript were supported by a grant of the National Natural Science Foundation of China (Grant No. 30900743) and A Project Funded by the Priority Academic Program Development of Jiangsu Higher Education Institutions (PAPD).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Li, J.F., Zhang, J., Xu, X.Z. et al. The antibacterial peptide ABP-CM4: the current state of its production and applications. Amino Acids 42, 2393–2402 (2012). https://doi.org/10.1007/s00726-011-0982-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00726-011-0982-3