Abstract
Porcine reproductive and respiratory syndrome virus (PRRSV) is the most important pathogen in the Korean swine industry. Despite efforts including improved biosecurity and vaccination protocols, the virus continues to circulate and evolve. Based on phylogenetic analysis of open reading frame 5 (ORF5), Korean PRRSVs are known to form not only globally circulating lineages but also country-specific lineages (Lin Kor A, B, and C). To understand the recent epidemiological status of PRRSV in Korea, a total of 1349 ORF5 sequences of Korean PRRSV isolates from 2014 to 2019 were analyzed. Phylogenetic analysis was conducted using the maximum-likelihood method, and temporal changes in the relative prevalence of lineages were investigated. The analysis showed that PRRSV1 and PRRSV2 were both highly prevalent throughout the years examined. Among the PRRSV1 isolates, subgroup A (90.1%) and vaccine-like subgroup C (9.0%) composed most of the population. For PRRSV2 isolates, vaccine-like lineage 5 (36.3%) was dominant, followed by Lin Kor B (25.9%), Kor C (16.6%), lineage 1 (11.6%), and Kor A (9.1%). The PRRSV2 lineage 1 population increased from 2014 (1.8%) to 2019 (29.6%) in Korea due to the continual spread of sublineage 1.8 (NADC30-like) and introduction of sublineage 1.6 into the country. Additional genetic analysis, including analysis of non synonymous and synonymous mutations, revealed evidence of diversification and positive selection in immunologically important regions of the genome, suggesting that current vaccination is failing and promoting immune-mediated selection. Overall, these findings provide insights into the epidemiological and evolutionary dynamics of cocirculating viral lineages, and constant surveillance of PRRSV occurrence is needed.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Porcine reproductive and respiratory syndrome (PRRS), first recognized in the United States in 1987 [21], is one of the most important epidemic diseases that affects the global pig industry [16, 47]. The causative agent, porcine reproductive and respiratory syndrome virus (PRRSV), belonging to the genus Betaarterivirus in the family Arteriviridae, is an enveloped, single-stranded positive-sense RNA virus [28]. Its genome is approximately 15 kb in length, containing a untranslated region (UTR), at least 11 open reading frames (ORFs), a 3’-UTR, and a 3’ poly(A) tail [19, 44]. PRRSV is classified into two genotypes, PRRSV1 (European genotype) and PRRSV2 (North American genotype), which share only approximately 60% sequence identity at the nucleotide level.
The PRRSV ORF5 gene encodes the major viral envelope protein GP5, which plays an important role in viral assembly, infectivity, and induction of neutralizing antibody production [7, 50, 66]. Based on genetic relationships among ORF5 sequences and the global PRRSV classification system, PRRSV2 is classified into nine lineages (L1-L9), and PRRSV1 is divided into four subtypes (sub1-sub4) [55, 56].
In South Korea, PRRSV2 was first detected in 1994, while PRRSV1 has spread rapidly since its first detection in 2005 [23, 45]. PRRSV1 in Korea has been recognized to be highly prevalent and is classified into subgroups A (sub1A), B (sub1B), and C (sub1C) of subtype 1 (pan-European PRRSV1), with the majority belonging to subgroup A [20, 24]. PRRSV2 has spread across the country for decades and has formed a unique country-specific clade containing the lineages Kor A (LKA), B (LKB), and C (LKC), which are distinct from the prevalent global PRRSV2 strains and commercial vaccine strains [20, 32]. The majority of the Korean PRRSV2 field isolates have been reported to belong to the Korean lineages and lineage 5. Lineage 5 isolates in Korea are closely related to the Ingelvac PRRS MLV vaccine virus, which has been used commercially in Korea since 1996. It has been reported that LKA and LKC were present in Korea before 2010, while LKB was first reported between 2014 and 2016 [20, 22].
PRRSV2 lineage 1, including viruses with the ORF5 restriction fragment length polymorphism (RFLP) 1-7-4, is known to have a long epidemic history since it initially appeared in Canada in the 1990s [56, 59]. After the initial outbreak in North America in the early 2000s (representative strain: MN184) [14], a new cluster of lineage 1 spread from Canada to the United States and China in 2013 (representative strain: NADC30) [57]. The RFLP 1-7-4 lineage (representative strain: NADC34) started to emerge in the United States in 2014 and caused dramatic abortion ‘storms’ in sow herds and high mortality among piglets [1, 60]. NADC34-like strains were subsequently detected in Peru from 2015 to 2017 as endemic forms [54], and in China since 2017 [39, 68]. In Korea, LKC was reported to be the sole lineage 1 strain in Korea until 2014. It is a country-specific sublineage of lineage 1 and might be related to MN184 [20, 49]. NADC30-like viruses were first identified in Korea in 2014, and knowledge regarding the recent epidemiological status of Korean lineage 1 viruses is lacking.
Here, we collected ORF5 sequence data from clinical samples and analyzed Korean PRRSV ORF5 sequences from 2014 to 2019. We investigated the dynamics of the prevalence of different lineages and the genetic characteristics of Korean PRRSV isolates to provide a more comprehensive understanding of current epidemiology of PRRSV in Korea.
Materials and methods
Sequencing and data processing
From 2015 to 2019, a total of 749 PRRSV ORF5 sequences were obtained from clinical samples (including lung, lymph node, and serum samples), which were collected through Jeonbuk National University Veterinary Diagnostic Center (JBNU-VDC) (GenBank: MW716541-MW717289). The procedures for tissue sample processing, RNA extraction, primer usage, RT-PCR, and ORF5 sequencing were described previously [20]. A total of 604 Korean PRRSV ORF5 sequences that were submitted to the NCBI GenBank database since 2014 were included in the dataset for a better description of PRRSV dynamics in Korea (Table 1). For lineage classification and identification of evolutionary relationships, 118 reference sequences were collected and used as “anchor” sequences.
Phylogenetic analysis
A total of 1471 ORF5 nucleotide sequences were aligned using Clustal Omega [58] and subjected to recombination screening using Recombination Detection Program (RDP) v.4.96 [41]. Seven potential recombinants were removed, and 1464 sequences were used for phylogenetic analysis. A maximum-likelihood (ML) phylogenetic tree was constructed using RAxML-NG [26] with 1,000 bootstrap replicates and the GTRGAMMA nucleotide substitution model. The lineages were classified based on the genetic relationships among the ORF5 sequences and anchor sequences, using the classification system reported by Shi et al. [56, 57]. We used Microreact [3], R [18], ggplot2, tidyverse [64], and ggtree [67] to clean and plot data and trees.
All of the Korean lineage 1 sequences (n = 77) were extracted from the lineage 1 dataset to investigate the epidemiology of Korean lineage 1 strains. An ML tree was built with the addition of reference sequences from other sublineages and recently reported NADC34-like strains. To explore the introduction timeline of lineage 1 strains within Korea, data regarding the number of pigs imported each year from North America (United States and Canada) to Korea were collected from the website of the Korean Animal and Plant Quarantine Agency (https://www.qia.go.kr/).
Amino acid variation and selection pressure analysis
Nucleotide and amino acid (aa) sequence identity between lineages was calculated using MEGA software [29]. To graphically summarize the variability in aa sequences, we constructed alignments of sequence logos derived from the corresponding aa sequence alignments for each lineage using the WebLogo tool (http://weblogo.berkeley.edu/). As the frequency plot option describes the relative frequency of each aa at each position and provides a richer and more precise description than a consensus sequences, an alignment of WebLogo graphical outputs was constructed [6, 40]. Amino acid diversity was calculated as sequence entropy at each codon using the Shannon Entropy-One tool implemented in the HIV database tool (https://www.hiv.lanl.gov/).
To analyze the selection pressure on the ORF5 gene of Korean PRRSV isolates, HyPhy version 2.5.2 [52] was used to infer positive selection for each codon applying single-likelihood ancestor counting (SLAC) [25]. Sites were inferred to be positively selected at a p-value ≤ 0.05.
Results
Lineage classification
After removal of the seven recombinant sequences and anchor sequences from the dataset, the remaining 1349 sequences were classified into nine different lineages (Fig. 1A and Supplementary Table S1). First, 51.0% (688 sequences) were grouped into PRRSV1, and the other 49.0% (661) were grouped into PRRSV2. Out of the 688 sequences in the PRRSV1 group, 90.1% (620) were classified in sub1A, 0.9% (6) in sub1B, and 9.0% (62) in sub1C. Among the PRRSV2 sequences, 36.3% (240) were classified as L5, 25.9% (171) as LKB, 16.6% (110) as LKC, 11.6% (77) as L1, and 9.1% (60) as LKA. Only three sequences (<1%) could be grouped into L8. The within- and between-lineage nucleotide and aa pairwise distances are shown in Table 2. Generally, the within-lineage variation was smaller than the between-lineage/sublineage distances, but the within-lineage variation of LKA, LKC, and sub1A was higher than 10%.
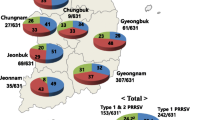
Phylogenetic tree showing clustering and genotyped of Korean PRRSV isolates based on ORF5 sequences and the tendency of PRRSV lineage occurrence. (A) Midpoint-rooted maximum-likelihood phylogeny reconstructed using the GTRGAMMA model based on the Korean PRRSV dataset (n = 1349) and reference anchor sequences (n = 115) of the PRRSV ORF5 gene. Colors represent different lineages, and differences in hues within a color represent anchor versus Korean sequences. The prototype strains were Lelystad for PRRSV1 and VR2332 for PRRSV2. Representatives and vaccines are indicated by arrows and labeled. (B) Pie chart showing the relative frequency of Korean ORF5 sequences according to lineage over the years.
Temporal dynamics of the prevalence of cocirculating lineages
Temporal changes in frequency of occurrence of different lineages were examined to understand PRRSV epidemics in Korea (Fig. 1B and Supplementary Table S1). For PRRSV1, the trend with sub1A as the predominant group (82.5-96.1%) and sub1C as the minor group (2.8-17.5%) was unchanged throughout 2014-2019. Sub1B viruses were detected only in 2014 (two sequences) and 2016 (four sequences), with very small populations. However, for PRRSV2, the relative frequency of each lineage changed over the years, and the increase in L1 occurrence was noteworthy. From 2014 to 2016, the pattern of L5 as the predominant lineage (46.7-34.1%), followed by LKB (29.4-24.3%), LKC (20.0-17.8%), and LKA (14.7-4.4%), was unchanged. Interestingly, the occurrence of L1 started to increase in 2014 (1.8%), and it comprised the second-largest population (29.6%) of PRRSV in 2019, after L5 (31.1%).
Genotyping and epidemiology of PRRSV2 lineage 1 circulating in Korea
To examine the genetic diversity of the increasingly prevalent PRRSV2 lineage 1 in Korea, a phylogenetic tree was reconstructed with Korean L1 isolates and L1 reference strains. Phylogenetic analysis demonstrated that Korean L1 strains can be grouped into two clades based on the ORF5 gene (Fig. 2). One clade (81.8%, 63/77 sequences) could be classified within sublineage 1.8 (NADC30-like) and was designated Kor L1 clade I. Viruses within Kor L1 clade I were first identified in 2014 and have been detected continuously ever since. Notably, the other clade (clade II) (16.9%, 13/77) grouped into sublineage 1.6, which has not been reported in Korea, and started to be detected among isolates in 2018. No NADC34-like strains were detected.
Midpoint-rooted maximum-likelihood phylogenetic tree constructed based on the ORF5 gene sequences of 77 Korean PRRSV2 L1 isolates and 119 reference PRRSV strains. Korean L1 isolates are indicated by a blue triangle (▲). Chinese, Peruvian, and American PRRSV2 L1 isolates are indicated by open red circles (○), green squares (□), and purple triangles (△), respectively.
To explore the introduction of lineage 1 virus from the North American region to Korea, the introduction of each sublineage was examined based on virus detection date and the yearly pig trade numbers from the United States and Canada to Korea (Fig. 3). Korea has continuously imported pigs from those regions, with the first peak in the number of importations occurring in 2011, and the second in 2017 (Fig. 3B). After the first import peak, the first lineage 1 virus was detected in Korea and identified as sublineage 1.8 (NADC30-like) in 2014. Subsequently, sublineage 1.6 was first detected in 2018, immediately after the second import peak (Fig. 3A). These sublineage 1 viruses were identified to originate from an external virus introduction rather than from L5 or Korean lineages that were mainly circulating in Korea, as seen in the evolutionary tree (Fig. 1A).
A timeline of PRRSV2 L1 and pig introductions from North America. (A) The timeline shows the introduction of the sublineage 1 of L1 viruses first detected in Korea. Red stars (★) indicate peaks in the number of swine imported throughout the years. (B) The number of swine imported from the United States and Canada into Korea each year
Amino acid analysis of Korean PRRSV1
Figure 4 shows the overall entropy plot for PRRSV1 GP5 (aa 1-201) from 688 Korean PRRSV1 isolates together with the alignment of sequence logos obtained for each subgroup. The neutralizing epitope at aa 29-35 was found to be conserved, with a low level of variability. Korean PRRSV1 sub1C showed polymorphism at V/A32 and slight variation at aa positions 33 and 35 in the neutralizing epitope (WSFADGN or WSFVDGN). In contrast, the neutralizing epitope of Korean PRRSV1 sub1A was relatively conserved (WSFADGN), with slight variation at aa positions 30-33 and 35. Hypervariable sites were observed with higher aa entropy (diversity) in the signal peptide region (aa positions 4, 8, and 9), putative T cell epitope 1 (56, 60, and 63), hypervariable domain (103, 104, and 105), putative T cell epitope 2 (126), and putative B cell epitope 3 (173 and 174). By SLAC analysis, a total of 22 codon sites were found to be positively selected in Korean PRRSV1 sub1A, while only one site was found in sub1C, which was within the signal peptide region (Table 3). More than half of the positively selected sites among sub1A isolates (13 sites) were found in the signal peptide region. Interestingly, three sites (positions 31, 35 and 36) within or near the neutralizing epitope, two sites (56, 60) within T cell epitope 1, and four sites (104, 105, 106 and 116) within or near the hypervariable domain were found to be positively selected in sub1A.
Amino acid diversity (entropy plot) and alignment of sequence logos produced with the 688 Korean PRRSV1 GP5 sequences including all subgroups. Amino acids are numbered from the start of the GP5 domain (aa 1-201). In the sequence logos, the height of each aa letter at each position is proportional to the frequency in all of the sequences from the corresponding subgroup. Amino acids are color-coded: blank, nonpolar; green, polar uncharged; red, polar with positive charge; blue, polar with negative charge. The subgroup is indicated on the left-hand side. Boxes with dashed black lines show biologically significant regions, and boxes with red lines show antigenic regions [12, 31].
Amino acid analysis of Korean PRRSV2
An entropy plot with the sequence logo alignment of each lineage/sublineage of PRRSV2 GP5 (aa 1-200) from 661 Korean PRRSV2 strains showed that the principal neutralizing epitopes (PNEs) of each lineage were conserved for L5 (SSHLQLIYNLTLCELNG) and L1/LKC (SSHLQLIYNLTICELNG) (Fig. 5). However, variations were found in the PNE within LKA at aa positions 38 (K/H/Y/N), 39 (L/F/S), and 47 (L/I), and within LKB at position 38 (N/K/T/H). Hypervariable sites among PRRSV2 GP5 sequences were mostly located in the decoy epitope region (aa positions 27-30) and known hypervariable regions (32-35 and 57-59) near the PNE. Minor variations were observed in the signal peptide (aa positions 3, 10, 13, 15, 16, and 23-26), TM2 (102 and 104), putative T cell epitope 1 (121, 124, 127, and 128), T cell epitope 2 (151), and B cell epitope (185). Subsequent positive selection pressure analysis of PRRSV2 GP5 revealed that all lineages showed positive selection near the PNE region, especially in the decoy epitope region and hypervariable region 1. Most of the positively selected sites were located in the first third of PRRSV GP5.
Amino acid diversity (entropy plot) and alignment of sequence logos with the 661 Korean PRRSV2 GP5 sequences (aa 1-200), including all lineages and sublineages. The meaning of the height and color of the sequence logos is explained in the legend to Fig. 4. Biologically significant regions and antigenic regions are indicated [44].
Discussion
In this study, the circulation, emergence, prevalence of different lineages, and genetic characteristics of PRRSV were investigated using a total of 1349 PRRSV ORF5 sequences obtained from Korean field isolates circulating on swine farms nationwide over a span of 6 years (2014-2019). By classifying the ORF5 sequences into phylogenetic lineages based on a global classification system [55, 56] and previously known unique Korean lineages [20], the diversification and temporal dynamics of the PRRSV population were investigated. Through further analysis of lineage 1 viruses, the first detection and emergence of sublineage 1.6 in Korea was identified. The alignment of sequence logos, entropy plots, and positive selection pressure analysis revealed sites with aa differences among different lineages, and nonsynonymous mutations that led to aa changes in the protein at those sites were favoured.
Overall, PRRSV1 and PRRSV2 were found to be circulating simultaneously in Korea. In previous PRRSV prevalence studies in Korea, the positive rates of PRRSV1 and PRRSV2 were reported to be 29.4% and 54.4% from the samples collected from 2007 to 2008 [35], and 38.4% and 37.4%, respectively, from the samples collected from 2013 to 2016 [20]. Although the positive rate could not be determined in this study, 51% and 49% of the ORF5-positive samples collected in the fields were found to be PRRSV1 and PRRSV2, respectively, which is consistent with the trend of PRRSV1 becoming prevalent in Korea following its first isolation in 2005, as reported previously.
All of the Korean PRRSV1 isolates were grouped into subgroups A, B, and C of subtype 1, and the majority of Korean PRRSV1 isolates belonged to subgroup A (90.1%), while subgroup B (0.9%) and subgroup C (9.0%) viruses formed minor populations, consistent with previous reports [20, 24]. It was noted that the subgroup C viruses, which were grouped with PRRSV1 live vaccine strains (Porcilis® PRRS and UNISTRAIN® PRRS MLV), became more prevalent from 2014 (2.8%) to 2019 (17.5%). Considering that live PRRSV1 vaccines have been used commercially in Korea since 2014 [31], it has been suggested that the emergence of these vaccine-like viruses might be related to previously known safety issues of current modified live vaccines (MLVs), such as reversion to virulence and persistence and circulation in the field of vaccine-derived strains [46]. Indeed, outbreaks of PRRSV1 strains that shared a common evolutionary ancestor with a European PRRSV vaccine strain have been reported recently in Hungary [42], China [5], and Taiwan [36], raising concerns about the safety of the current MLVs. Recently, there have been several reports that mosaic recombinants derived from recombination between the PRRSV1 vaccine and the predominant circulating field strains have caused outbreaks and have shown increased viremia and transmissibility [10, 30, 70]. Although recombination between PRRSV field strains has been reported less frequently for PRRSV1 than for PRRSV2 [30], the increasing circulation of vaccine-related strains in Korea indicates that such spread could pose a potential outbreak threat by increasing the chance of recombination with field strains. Further research applying whole-genome sequencing is needed to identify characteristics of Korean PRRSV1 genomes, including recombination signals.
For Korean PRRSV2, viruses were classified into lineages 1, 5, and 8, as well as Korean lineages LKA, LKB, and LKC. The majority of Korean PRRSV2 field strains belonged to lineage 5, which is closely related to the Ingelvac® PRRS MLV, and Korean lineages as reported previously [20]. Considering that the PRRSV2 MLV was licensed in 1996 and had been the only option for controlling PRRSV2 in Korea before 2014 [4, 31], the consistent prevalence of vaccine-like PRRSV2 is explainable as the circulation of vaccine-related variants. Among the specifically Korean lineages, LKB has comprised the largest population (17.8-33.3%) since its first identification in 2014 [20], followed by LKC (10.8-20.0%) and LKA (2.2-14.7%). The continuous circulation of these viruses indicates that these lineages might be sufficiently immunologically distinct to overcome herd immunity [48] and escape from the immunity induced by nationwide PRRSV2 MLV usage. PRRSV vaccines are expected to offer better protection against wild viral variants that have a higher degree of similarity to the original parental vaccine isolate [9, 11]. Although our understanding of heterologous cross-protection is limited for PRRSV, the emergence and sequential dominance of different variants leading to lineages and sublineages is consistent with the theory of multistrain dynamics [13, 27]. Continuous surveillance of these Korean lineages and further research to evaluate the cross-protection capability of commercial PRRSV2 MLVs against these lineages are needed in the future.
PRRSV2 lineage 1 viruses have mainly been epidemic in the United States, while unique endemic clusters have developed in some countries, including Korea in the case of LKC [57]. While the connection of lineage 1 viruses between the United States and other countries remains unclear, worldwide hog trading and artificial insemination are considered to be the most likely causes [59]. Lineage 1 viruses have caused severe epidemic situations outside the United States, including the spread of NADC30-like viruses in China and Taiwan since 2013 and 2018, respectively [37, 69], and the introduction of RFLP 1-7-4 virus (sublineage 1.5, NADC34-like) in Peru [54] and China [68]. In Korea, the first nonendemic lineage 1 virus was detected and identified as NADC30-like virus in 2014, but the current epidemiologic status has been unknown since 2016 [20]. In this study, temporal surveillance detected an increase in the prevalence of lineage 1 virus since 2017, and phylogenetic analysis, together with swine-trade statistics, revealed that sublineage 1.6, which has been reported to be a Canadian-origin (Quebec and Ontario) PRRSV2 sublineage that is prevalent mainly in North America [34, 57], was newly introduced into Korea. Despite the fact that NADC34-like viruses have not yet been detected in Korea, the population of newly introduced sublineage 1.6 and NADC30-like viruses in the field is growing. A recent study demonstrated an NADC30-like PRRSV outbreak in a pig herd that had been vaccinated with an MLV [37]. Although the vaccination status at the individual farm level was not investigated in this study, the observation of active circulation of NADC30-like viruses indicates that vaccination with the current MLVs is failing to provide sufficient cross-protection against those viruses despite extensive MLV vaccination on most PRRSV-positive or endangered farms in Korea. Vaccine protection experiments and continuous surveillance should be performed to track the epidemiological impact of the emerging novel sublineage 1 viruses in Korea.
A recent study demonstrated that antibodies can impose strong positive selective pressure on PRRSV by targeting specific viral subpopulations while allowing the establishment of other subpopulations [63]. It has also been noted that PRRSV vaccination resulted in an increase in the genetic heterogeneity of PRRSV and the emergence of new glycosylation sites in the viral population [31]. In our study, pairwise distance analysis showed that ORF5 of Korean PRRSV1 subgroup A viruses was highly divergent at the nucleotide (10.7%) and aa levels (12.4%). For subgroup C, or the vaccine-like subgroup, the pairwise distance within the group was 3.7% and 6.0% at the nucleotide and aa level, respectively, indicating more diversity than the previously reported pairwise distance (< 1%) of subgroup C viruses in Korea at the nucleotide level [20]. Alignment of sequence logos showed that diversification of Korean PRRSV1 subgroups A and C could be detected in previously identified putative T cell and B cell epitopes [8, 43, 61, 65]. Subgroup A viruses did not show much variation in the neutralizing epitope region, while subgroup C viruses showed variation at aa position 32 within the neutralizing epitope region. Although the positive selection signal was not detected in the region from subgroup C (which might be due to the small sample size), the selection signals of subgroup A supported immunological selection among Korean PRRSV1 strains. Aggregated data suggested that the nationwide PRRSV1 MLV usage might have caused immunological selection to become a driver of PRRSV1 diversification [44].
Likewise, Korean PRRSV2 shows large pairwise distances at the nucleotide and aa levels, not only between lineages but also within lineages. Subsequently, alignment of sequence logos, entropy plots, and positive selection analysis showed variations and sites under positive selective pressure within or near two hypervariable regions located near the PNE. The PNE is known to be located between aa positions 36 and 52 in GP5 and forms an ectodomain that can induce antibody production during PRRSV infection [15, 51]. The flanking hypervariable regions can modulate the accessibility of neutralizing antibodies specific for the PNE [53], including N-linked glycosylation sites such as N34, N44, and N51 [2]. Generally, glycosylation may alter protein-protein interactions and whether these proteins induce a humoral or cellular immune response in the host [38], and a previous study found evidence that these glycosylation sites in PRRSV play a role in glycan shielding, which enables the virus to evade a neutralizing immune response [62]. Although glycosylation site prediction was not performed in this study, a previous report revealed that the N-glycosylation pattern of Korean PRRSV2 is highly diverse and changeable, while it has been conserved in Korean PRRSV1 [20]. Therefore, it is suggested that the strong preference for mutation of flanking hypervariable regions and epitopes might be associated with limited protective efficacy of PRRSV2 MLV vaccination against heterologous viruses in the field, as shown in previous studies [17, 33].
In the present study, we examined the epidemic status and characteristics of Korean PRRSV1 and PRRSV2. Since the ORF5 gene represents approximately 5% of the whole genome of PRRSV, further studies utilizing whole-genome analysis to better understand evolutionary dynamics elsewhere in the genome are needed. However, ORF5 sequencing could still provide a fast and extensive view of concurrent PRRSV outbreaks and introductions. Expected limited cross-protection efficacy and foreign introduction of new sublineages have contributed to the constant circulation of MLV vaccine variants, establishment of country-specific Korean lineages, and emergence of newly introduced sublineage 1 viruses in Korea despite nationwide MLV vaccination. Therefore, further study will be required to understand the genomic characteristics of the prevailing PRRSV lineages and the cross-protection efficacy of current commercial vaccines against the prevalent PRRSV strains in Korea. To minimize the circulation of MLV strains in the field and possible recombination between vaccines and field strains, which can trigger the emergence of novel genotypes and outbreaks, enhanced biosecurity and proper use of live vaccines will be necessary. Furthermore, strict surveillance should be performed routinely to prevent the potential introduction of new viruses.
References
Alkhamis MA, Perez AM, Murtaugh MP, Wang X, Morrison RB (2016) Applications of Bayesian phylodynamic methods in a recent US porcine reproductive and respiratory syndrome virus outbreak. Front Microbiol 7:67
Ansari IH, Kwon B, Osorio FA, Pattnaik AK (2006) Influence of N-linked glycosylation of porcine reproductive and respiratory syndrome virus GP5 on virus infectivity, antigenicity, and ability to induce neutralizing antibodies. J Virol 80:3994–4004
Argimón S, Abudahab K, Goater RJ, Fedosejev A, Bhai J, Glasner C, Feil EJ, Holden MT, Yeats CA, Grundmann H (2016) Microreact: visualizing and sharing data for genomic epidemiology and phylogeography. Microb Genom. https://doi.org/10.1099/mgen.0.000093
Cha S-H, Choi E-J, Park J-H, Yoon S-R, Song J-Y, Kwon J-H, Song H-J, Yoon K-J (2006) Molecular characterization of recent Korean porcine reproductive and respiratory syndrome (PRRS) viruses and comparison to other Asian PRRS viruses. Vet Microbiol 117:248–257
Chen N, Liu Q, Qiao M, Deng X, Chen X, Sun M (2017) Whole genome characterization of a novel porcine reproductive and respiratory syndrome virus 1 isolate: genetic evidence for recombination between Amervac vaccine and circulating strains in mainland China. Infect Genet Evol 54:308–313
Crooks GE, Hon G, Chandonia J-M, Brenner SE (2004) WebLogo: a sequence logo generator. Genome Res 14:1188–1190
Delisle B, Gagnon CA, Lambert M-È, D’Allaire S (2012) Porcine reproductive and respiratory syndrome virus diversity of Eastern Canada swine herds in a large sequence dataset reveals two hypervariable regions under positive selection. Infect Genet Evol 12:1111–1119
Díaz I, Pujols J, Ganges L, Gimeno M, Darwich L, Domingo M, Mateu E (2009) In silico prediction and ex vivo evaluation of potential T-cell epitopes in glycoproteins 4 and 5 and nucleocapsid protein of genotype-I (European) of porcine reproductive and respiratory syndrome virus. Vaccine 27:5603–5611
Díaz I, Gimeno M, Darwich L, Navarro N, Kuzemtseva L, López S, Galindo I, Segalés J, Martín M, Pujols J (2012) Characterization of homologous and heterologous adaptive immune responses in porcine reproductive and respiratory syndrome virus infection. Vet Res 43:1–15
Eclercy J, Renson P, Lebret A, Hirchaud E, Normand V, Andraud M, Paboeuf F, Blanchard Y, Rose N, Bourry O (2019) A field recombinant strain derived from two type 1 porcine reproductive and respiratory syndrome virus (PRRSV-1) modified live vaccines shows increased viremia and transmission in SPF pigs. Viruses 11:296
Geldhof MF, Vanhee M, Van Breedam W, Van Doorsselaere J, Karniychuk UU, Nauwynck HJ (2012) Comparison of the efficacy of autogenous inactivated Porcine Reproductive and Respiratory Syndrome Virus (PRRSV) vaccines with that of commercial vaccines against homologous and heterologous challenges. BMC Vet Res 8:1–16
Guo Z, Chen X-x, Li R, Qiao S, Zhang G (2018) The prevalent status and genetic diversity of porcine reproductive and respiratory syndrome virus in China: a molecular epidemiological perspective. Virol J 15:1–14
Gupta S, Ferguson N, Anderson R (1998) Chaos, persistence, and evolution of strain structure in antigenically diverse infectious agents. Science 280:912–915
Han J, Wang Y, Faaberg KS (2006) Complete genome analysis of RFLP 184 isolates of porcine reproductive and respiratory syndrome virus. Virus Res 122:175–182
Hanada K, Suzuki Y, Nakane T, Hirose O, Gojobori T (2005) The origin and evolution of porcine reproductive and respiratory syndrome viruses. Mol Biol Evol 22:1024–1031
Holtkamp DJ, Kliebenstein JB, Neumann E, Zimmerman JJ, Rotto H, Yoder TK, Wang C, Yeske P, Mowrer CL, Haley CA (2013) Assessment of the economic impact of porcine reproductive and respiratory syndrome virus on United States pork producers. J Swine Health Prod 21:72
Hu J, Zhang C (2014) Porcine reproductive and respiratory syndrome virus vaccines: current status and strategies to a universal vaccine. Transbound Emerg Dis 61:109–120
Ihaka R, Gentleman R (1996) R: a language for data analysis and graphics. J Comput Graph Stat 5:299–314
Johnson CR, Griggs TF, Gnanandarajah J, Murtaugh MP (2011) Novel structural protein in porcine reproductive and respiratory syndrome virus encoded by an alternative ORF5 present in all arteriviruses. J Gen Virol 92:1107
Kang H, Yu JE, Shin J-E, Kang A, Kim W-I, Lee C, Lee J, Cho I-S, Choe S-E, Cha S-H (2018) Geographic distribution and molecular analysis of porcine reproductive and respiratory syndrome viruses circulating in swine farms in the Republic of Korea between 2013 and 2016. BMC Vet Res 14:160
Keffaber K (1989) Reproduction failure of unknown etiology. Am Assoc Swine Pract Newsl 1:1–9
Kim H, Nguyen V, Kim I, Park J, Park S, Rho S, Han J, Park B (2012) Epidemiologic and phylogenetic characteristics of porcine reproductive and respiratory syndrome viruses in conventional swine farms of Jeju Island as a candidate region for PRRSV eradication. Transbound Emerg Dis 59:62–71
Kim J-Y, Lee S-Y, Sur J-H, Lyoo YS (2006) Serological and genetic characterization of the European strain of the porcine reproductive and respiratory syndrome virus isolated in Korea. Korean J Vet Res 46:363–370
Kim S-H, Roh I-S, Choi E-J, Lee C, Lee C-H, Lee K-H, Lee K-K, Song Y-K, Lee O-S, Park C-K (2010) A molecular analysis of European porcine reproductive and respiratory syndrome virus isolated in South Korea. Vet Microbiol 143:394–400
Kosakovsky Pond SL, Frost SD (2005) Not so different after all: a comparison of methods for detecting amino acid sites under selection. Mol Biol Evol 22:1208–1222
Kozlov AM, Darriba D, Flouri T, Morel B, Stamatakis A (2019) RAxML-NG: a fast, scalable and user-friendly tool for maximum likelihood phylogenetic inference. Bioinformatics 35:4453–4455
Kucharski AJ, Andreasen V, Gog JR (2016) Capturing the dynamics of pathogens with many strains. J Math Biol 72:1–24
Kuhn JH, Lauck M, Bailey AL, Shchetinin AM, Vishnevskaya TV, Bao Y, Ng TF, LeBreton M, Schneider BS, Gillis A, Tamoufe U, Diffo Jle D, Takuo JM, Kondov NO, Coffey LL, Wolfe ND, Delwart E, Clawson AN, Postnikova E, Bollinger L, Lackemeyer MG, Radoshitzky SR, Palacios G, Wada J, Shevtsova ZV, Jahrling PB, Lapin BA, Deriabin PG, Dunowska M, Alkhovsky SV, Rogers J, Friedrich TC, O’Connor DH, Goldberg TL (2016) Reorganization and expansion of the nidoviral family Arteriviridae. Arch Virol 161:755–768
Kumar S, Stecher G, Li M, Knyaz C, Tamura K (2018) MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol 35:1547–1549
Kvisgaard LK, Kristensen CS, Ryt-Hansen P, Pedersen K, Stadejek T, Trebbien R, Andresen LO, Larsen LE (2020) A recombination between two Type 1 Porcine Reproductive and Respiratory Syndrome Virus (PRRSV-1) vaccine strains has caused severe outbreaks in Danish pigs. Transbound Emerg Dis 67:1786–1796
Kwon T, Yoo SJ, Lee D-U, Sunwoo SY, Sang HJ, Park JW, Kim M-H, Park C-K, Lyoo YS (2019) Differential evolution of antigenic regions of porcine reproductive and respiratory syndrome virus 1 before and after vaccine introduction. Virus Res 260:12–19
Kwon T, Yoo SJ, Sunwoo SY, Lee D-U, Sang HJ, Park JW, Park C-K, Lyoo YS (2019) Independent evolution of porcine reproductive and respiratory syndrome virus 2 with genetic heterogeneity in antigenic regions of structural proteins in Korea. Adv Virol 164:213–224
Labarque G, Van Reeth K, Nauwynck H, Drexler C, Van Gucht S, Pensaert M (2004) Impact of genetic diversity of European-type porcine reproductive and respiratory syndrome virus strains on vaccine efficacy. Vaccine 22:4183–4190
Lambert M-È, Delisle B, Arsenault J, Poljak Z, D’Allaire S (2019) Positioning Quebec ORF5 sequences of porcine reproductive and respiratory syndrome virus (PRRSV) within Canada and worldwide diversity. Infect Gen Evolut 74:103999
Lee C, Kim H, Kang B, Yeom M, Han S, Moon H, Park S, Kim H, Song D, Park B (2010) Prevalence and phylogenetic analysis of the isolated type I porcine reproductive and respiratory syndrome virus from 2007 to 2008 in Korea. Virus Genes 40:225–230
Lin W-H, Kaewprom K, Wang S-Y, Lin C-F, Yang C-Y, Chiou M-T, Lin C-N (2020) Outbreak of porcine reproductive and respiratory syndrome Virus 1 in Taiwan. Viruses 12:316
Lin WH, Shih HC, Wang SY, Lin CF, Yang CY, Chiou MT, Lin CN (2019) Emergence of a virulent porcine reproductive and respiratory syndrome virus in Taiwan in 2018. Transbound Emerg Dis 66:1138–1141
Lisowska E (2002) The role of glycosylation in protein antigenic properties. Cell Mol Life Sci CMLS 59:445–455
Liu J, Wei C, Lin Z, Xia W, Ma Y, Dai A, Yang X (2019) Full genome sequence analysis of a 1-7-4-like PRRSV strain in Fujian Province, China. PeerJ 7:e7859
López-Labrador FX, Moya A, Gonzàlez-Candelas F (2008) Mapping natural polymorphisms of hepatitis C virus NS3/4A protease and antiviral resistance to inhibitors in worldwide isolates. Antivir Ther 13:481
Martin DP, Murrell B, Golden M, Khoosal A, Muhire B (2015) RDP4: Detection and analysis of recombination patterns in virus genomes. Virus Evolut. https://doi.org/10.1093/ve/vev003
Marton S, Szalay D, Kecskeméti S, Forró B, Olasz F, Zádori Z, Szabó I, Molnár T, Bányai K, Bálint Á (2019) Coding-complete sequence of a vaccine-derived recombinant porcine reproductive and respiratory syndrome virus strain isolated in Hungary. Adv Virol 164:2605–2608
Mokhtar H, Eck M, Morgan SB, Essler SE, Frossard J-P, Ruggli N, Graham SP (2014) Proteome-wide screening of the European porcine reproductive and respiratory syndrome virus reveals a broad range of T cell antigen reactivity. Vaccine 32:6828–6837
Murtaugh MP, Stadejek T, Abrahante JE, Lam TT, Leung FC-C (2010) The ever-expanding diversity of porcine reproductive and respiratory syndrome virus. Virus Res 154:18–30
Nam E, Park C-K, Kim S-H, Joo Y-S, Yeo S-G, Lee C (2009) Complete genomic characterization of a European type 1 porcine reproductive and respiratory syndrome virus isolate in Korea. Adv Virol 154:629–638
Nan Y, Wu C, Gu G, Sun W, Zhang Y-J, Zhou E-M (2017) Improved vaccine against PRRSV: current progress and future perspective. Front Microbiol 8:1635
Neumann EJ, Kliebenstein JB, Johnson CD, Mabry JW, Bush EJ, Seitzinger AH, Green AL, Zimmerman JJ (2005) Assessment of the economic impact of porcine reproductive and respiratory syndrome on swine production in the United States. J Am Vet Med Assoc 227:385–392
Paploski IAD, Corzo C, Rovira A, Murtaugh MP, Sanhueza JM, Vilalta C, Schroeder DC, VanderWaal K (2019) Temporal dynamics of co-circulating lineages of porcine reproductive and respiratory syndrome virus. Front Microbiol 10:2486
Park J, Choi S, Jeon JH, Lee K-W, Lee C (2020) Novel lineage 1 recombinants of porcine reproductive and respiratory syndrome virus isolated from vaccinated herds: genome sequences and cytokine production profiles. Adv Virol 165:2259–2277
Pirzadeh B, Gagnon CA, Dea S (1998) Genomic and antigenic variations of porcine reproductive and respiratory syndrome virus major envelope GP5 glycoprotein. Can J Vet Res 62:170
Plagemann P, Rowland R, Faaberg K (2002) The primary neutralization epitope of porcine respiratory and reproductive syndrome virus strain VR-2332 is located in the middle of the GP5 ectodomain. Adv Virol 147:2327–2347
Pond SLK, Muse SV (2005) HyPhy: hypothesis testing using phylogenies Statistical methods in molecular evolution. Springer, pp 125–181
Popescu LN, Trible BR, Chen N, Rowland RR (2017) GP5 of porcine reproductive and respiratory syndrome virus (PRRSV) as a target for homologous and broadly neutralizing antibodies. Vet Microbiol 209:90–96
Ramírez M, Bauermann FV, Navarro D, Rojas M, Manchego A, Nelson EA, Diel DG, Rivera H (2019) Detection of porcine reproductive and respiratory syndrome virus (PRRSV) 1-7-4-type strains in Peru. Transbound Emerg Dis 66:1107–1113
Shi M, Lam TT-Y, Hon C-C, Hui RK-H, Faaberg KS, Wennblom T, Murtaugh MP, Stadejek T, Leung FC-C (2010) Molecular epidemiology of PRRSV: a phylogenetic perspective. Virus Res 154:7–17
Shi M, Lam TT-Y, Hon C-C, Murtaugh MP, Davies PR, Hui RK-H, Li J, Wong LT-W, Yip C-W, Jiang J-W (2010) Phylogeny-based evolutionary, demographical, and geographical dissection of North American type 2 porcine reproductive and respiratory syndrome viruses. J Virol 84:8700–8711
Shi M, Lemey P, Brar MS, Suchard MA, Murtaugh MP, Carman S, D’Allaire S, Delisle B, Lambert M-È, Gagnon CA (2013) The spread of type 2 Porcine Reproductive and Respiratory Syndrome Virus (PRRSV) in North America: a phylogeographic approach. Virology 447:146–154
Sievers F, Wilm A, Dineen D, Gibson TJ, Karplus K, Li W, Lopez R, McWilliam H, Remmert M, Söding J (2011) Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol Syst Biol 7:539
Sun YK, Chen YJ, Cai Y, Li Q, Xie JX, Liang G, Gao Q, Yu ZQ, Lu G, Huang LZ (2020) Insights into the evolutionary history and epidemiological characteristics of the emerging lineage 1 porcine reproductive and respiratory syndrome viruses in China. Transbound Emerg Dis 67:2630–2641
van Geelen AGM, Anderson TK, Lager KM, Das PB, Otis NJ, Montiel NA, Miller LC, Kulshreshtha V, Buckley AC, Brockmeier SL, Zhang J, Gauger PC, Harmon KM, Faaberg KS (2018) Porcine reproductive and respiratory disease virus: Evolution and recombination yields distinct ORF5 RFLP 1-7-4 viruses with individual pathogenicity. Virology 513:168–179
Vanhee M, Van Breedam W, Costers S, Geldhof M, Noppe Y, Nauwynck H (2011) Characterization of antigenic regions in the porcine reproductive and respiratory syndrome virus by the use of peptide-specific serum antibodies. Vaccine 29:4794–4804
Vu HL, Kwon B, Yoon K-J, Laegreid WW, Pattnaik AK, Osorio FA (2011) Immune evasion of porcine reproductive and respiratory syndrome virus through glycan shielding involves both glycoprotein 5 as well as glycoprotein 3. J Virol 85:5555–5564
Wang X (2016) Immunological selection as a driver of porcine reproductive and respiratory syndrome virus (PRRSV) evolution (Doctoral dissertation). University of Minnesota, Minneapolis
Wickham H, Averick M, Bryan J, Chang W, McGowan LDA, François R, Grolemund G, Hayes A, Henry L, Hester J (2019) Welcome to the Tidyverse. J Open Sour Softw 4:1686
Wissink E, van Wijk H, Kroese M, Weiland E, Meulenberg J, Rottier P, van Rijn P (2003) The major envelope protein, GP5, of a European porcine reproductive and respiratory syndrome virus contains a neutralization epitope in its N-terminal ectodomain. J Gen Virol 84:1535–1543
Wissink E, Kroese M, Van Wijk H, Rijsewijk F, Meulenberg J, Rottier P (2005) Envelope protein requirements for the assembly of infectious virions of porcine reproductive and respiratory syndrome virus. J Virol 79:12495–12506
Yu G, Smith DK, Zhu H, Guan Y, Lam TTY (2017) ggtree: an R package for visualization and annotation of phylogenetic trees with their covariates and other associated data. Methods Ecol Evol 8:28–36
Zhang H-L, Zhang W-L, Xiang L-R, Leng C-L, Tian Z-J, Tang Y-D, Cai X-H (2018) Emergence of novel porcine reproductive and respiratory syndrome viruses (ORF5 RFLP 1-7-4 viruses) in China. Vet Microbiol 222:105–108
Zhang H, Leng C, Ding Y, Zhai H, Li Z, Xiang L, Zhang W, Liu C, Li M, Chen J (2019) Characterization of newly emerged NADC30-like strains of porcine reproductive and respiratory syndrome virus in China. Adv Virol 164:401–411
Zhao H, Han Q, Zhang L, Zhang Z, Wu Y, Shen H, Jiang P (2017) Emergence of mosaic recombinant strains potentially associated with vaccine JXA1-R and predominant circulating strains of porcine reproductive and respiratory syndrome virus in different provinces of China. Virol J 14:1–10
Acknowledgments
This study was supported by a Grant (Z-1543069-2017-20-1) from the Animal and Plant Quarantine Agency (QIA) and Animal Disease Management Technology Development Program (320060-02) of the Ministry of Agriculture, Food and Rural Affairs, Republic of Korea.
Funding
This study was supported by a Grant (Z-1543069-2017-20-1) from the Animal and Plant Quarantine Agency (QIA) and Animal Disease Management Technology Development Program (320060-02) of the Ministry of Agriculture, Food and Rural Affairs, Republic of Korea.
Author information
Authors and Affiliations
Contributions
Conceptualization, W.-I.K. and K.-K.L.; methodology, S.-C.K.; software, S.-C.K.; validation, J.-Y.P. and H.-Y.J.; formal analysis, S.-C.K.; investigation, S.-C.K.; resources, G.-S.P., S.-H.K., G.-E.S. and M.-K.K.; data curation, S.-C.K.; writing—original draft preparation, S.-C.K.; writing—review and editing, W.-I.K.; visualization, S.-C.K.; supervision, W.-I.K.; project administration, C.-G.J.; funding acquisition, W.-I.K. All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Additional information
Handling Editor: Sheela Ramamoorthy.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Kim, SC., Jeong, CG., Park, GS. et al. Temporal lineage dynamics of the ORF5 gene of porcine reproductive and respiratory syndrome virus in Korea in 2014–2019. Arch Virol 166, 2803–2815 (2021). https://doi.org/10.1007/s00705-021-05169-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-021-05169-w