Abstract
The typical chloroplast genome consists of two inverted repeat (IR) and two single-copy (SC) regions. Each region has its own pattern of molecular evolution. Many studies have shown that the IR region has a low substitution rate. The plastid rps12 gene encodes ribosomal protein small subunit 12, which is composed of 5′-rps12 (exon 1) and 3′-rps12 (exon 2–3); 3′-rps12 is near the IR region and enters or leaves the IR region as it expands and contracts, enabling study of the variation in its substitution rate. We analyzed the evolutionary rate and adaptive evolution of the rps12 genes of 118 ferns and 2 lycophytes in a phylogenetic context using the maximum likelihood method. We found that the substitution rate of exon 2–3 was significantly lower in the IR region than in the LSC (1–11 times higher in the LSC), and also lower than that of exon 1 in the LSC region (1.9–6.7 times higher for exon 1). In addition, there was more GC content at the third codon position, and the rps12 gene encoded 123 amino acids, with 107 negative selection sites and 4 positive selection sites. These results prove that the substitution rate decreases after the sequence fragments enter the IR region and demonstrate the highly conserved nature of the rps12 gene.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Chloroplast genomes in most plants consist of two inverted repeat (IR) regions. The large single-copy (LSC) region and the small single-copy (SSC) region. In most non-seed plant lineages, the IR region is 10-15 kb in size and generally contains four rRNA genes and five tRNA genes (Zhu et al. 2016). In ferns, some genes enter or leave the IR region with expansion and contraction of the region, including rps7, psbA, 3′-rps12, ycf2, ndhB (Li et al. 2016). The rps12 gene is very special, it encodes ribosomal protein small subunit 12 and consists of 5′-rps12 and 3′-rps12. Interestingly, the two parts are located a distance that is as far as half the size of the chloroplast genome (~ 70 kb) apart: 5′-rps12 contains only exon 1 and is located in the LSC region, while 3′-rps12 usually contains exon 2, introns and exon 3 and is located in the IR region. Some fern lineages have lost introns or 3′-rps12 enters the LSC region. During translation of rps12 gene, trans-splicing occurs between the 5′-rps12 and 3′-rps12 transcripts (Norihiro et al. 1987).
Different lineages and different genomes have different molecular evolutionary rates (Bromham and Penny 2003). Changes in the rate are often related to biological characteristics or population size. IR region may differ from the SC region in evolutionary rates due to its double-copy nature. Some studies have shown that genes in the IR region have lower substitution rates. In angiosperms, the IR region has lower synonymous and non-synonymous substitution rates than the SC region (Maier et al. 1995; Wolfe et al. 1987). Similarly, a low substitution rate has been found in the IR region in Cycas revoluta (Wu and Chaw 2015). In Leguminosae with loss of IR region, genes originally located in the IR region have synonymous substitution rates similar to those in the SC region (Perry and Wolfe 2002). Genes in the SC region were recently found to have substitution rates 3.7 times those of genes in the IR region. After a gene is transferred from the SC region to the IR region or vice versa, the substitution rates are consistent with other genes in the host region (Zhu et al. 2016). Gene translocation into the plastid IR region shows lower substitution rates and higher GC content (Li et al. 2016). However, the Ginkgo biloba ycf2 gene does not have a higher substitution rate after entering the SC region from the IR region (Lin et al. 2012). Note that some previous studies have ignored some of the interfering factors by comparing different genes or the same genes between different plants. Different genes have different biological functions and evolve in fundamentally different ways. Different species also experience different selection pressures. It is necessary to eliminate these factors to reach more accurate conclusions. Therefore, the rps12 gene with its special structure provides a natural condition for studying the change in substitution rate after the gene enters the IR region by comparing differences in evolution rate between the coding sequences in its two parts.
Ferns are the second largest group of vascular plants, with an estimated 10,578 extant species (PPG I 2016). With the development of next-generation sequencing, it is relatively easy to obtain sequences. This study examined 118 ferns and 2 lycophytes species and calculated the evolutionary rates of the rps12 codons to investigate the change in gene substitution rate after entering the IR region. We also assessed the selection process this particular gene has experienced in the evolution of ferns using selection pressure and adaptive evolution analyses.
Materials and methods
DNA extraction and sequencing sections have been removed.
Sequence data
We downloaded the complete chloroplast genome sequences of 120 plants (118 ferns and 2 lycophytes) from NCBI (Online Resource 1) and imported the sequences into Geneious Prime 2019.0.4 (Kearse et al. 2012) to extract the rbcL, matK and rps12 genes. In addition, the coding sequence (rps12-CDS), exon 1, exon 2 and exon 3 (exon 2–3) of rps12 were extracted. The sequences were aligned and manually corrected using the ClustalW or ClustalW (codons) module in MEGA7.0 (Kumar et al. 2016), and stored as electronic supplementary material (Online Resource 2–6).
Relative GC content
The relative GC content at each position of the rps12 codon in each plant was calculated. That is, the ratio of GC content at each position of rps12 codon to the GC content of plastids genome of corresponding species, so as to reduce the influence caused by the difference in GC content of chloroplast genome between species. Significance was analyzed using two-tailed t-tests.
Construct phylogenetic trees
Based on the complete chloroplast genome and the tandem data set of rbcL and matK (Hollingsworth et al. 2011; Li et al. 2016), two maximum likelihood trees were constructed using RaxmlGUI2 (GTRGAMMAI model and 100 Boostraps) (Stamatakis 2014) and MEGA 7.0 (GTR + G + I model and 1000 bootstraps), respectively. As the phylogenetic position of some fern groups is controversial, when determining the final tree, the Pteridophyte Phylogeny Group (PPG) I system was referred to at the order and family levels (PPG I 2016). The phylogenetic relationships of genera and species were determined from the two constructed trees.
Calculate the rate of evolution and hypothesis test
The rate of evolution was calculated using HyPhy 2.2.4 (Pond et al. 2005) with the partition type set to nucleotide, the HKY85 substitution model and local parameters; the transition (trst) and transversion (trsv) rates and ratio (trsv/trst) were calculated. The synonymous (dS) and non-synonymous (dN) substitution rates and ω (dN/dS) were calculated using the codon partition type, MG94 × HKY85_3 × 4 substitution model and local parameters. For the codon type, the substitution mode (MG94 × HKY85 × 3_4 × 2_rate), parameters (rate Het.) and rate classes (4) were used to analyze selective pressure. The Wilcoxon rank-sum test and Mann–Whitney U-test were performed with IBM SPSS Statistics 22.0.
The hypothesis test first saves the original data as a full model, then constrains the exon 1 value of each species branch to equal exon 2–3 and saves it as a null model after optimization, before finally performing a likelihood ratio test. Hypothesis testing was mainly used to test for significant difference in the evolutionary rate between exon 1 and exon 2–3.
Identification of positive amino acid sites and alignment of amino acid sequences
The positive amino acid selection sites of the rps12 gene were identified using codeml in the PAML 4.9 package (Yang 2007). Three models were used to calculate the number of parameters and likelihood values and estimate the value of the parameters and positively selected sites: the branch, site and branch-site models. The Chi-square test was performed using chi2.exe. The two-ratio model (Model2) in the branch model was used to calculate the ω values of the foreground branch and background branches.
To understand the rps12 sequence characteristics, we added rps12 amino acid sequences from six species (Table 1) and aligned 126 amino acid sequences using MEGA7.0. Finally, the online SWISS-MODEL tool was used to predict the structure of RPS12 protein (https://www.swissmodel.expasy.org/).
Results
Characteristics of rps12 gene
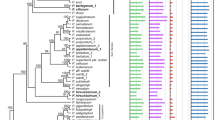
The size of the rps12 gene ranges from 372 (no introns) to 985 bp (Pteris vittata). The rps12 coding sequence in all species is the same size, 372 bp, and is divided into three exons: exon 1 is 114 bp, exon 2 is 232 bp, and exon 3 is 26 bp. In all, 106 plants also had introns which ranged in size from 447 to 613 bp. Figure 1 shows the phylogenetic tree of the 120 species; according to the location of exon 2–3, its divided into two groups: in the IR region (IR-109) and in the LSC region (LSC-11). Figure 2 shows the double-tailed t-test results of the relative GC content. The mean relative GC content in the first and second codon positions was lower in IR-109 than in LSC-11 (Pfirst = 0.658 Psecond = 0.032), while the relative content in the third codon position of the rps12 gene was higher (Pthird = 0).
Phylogenetic relationships of ferns
The phylogenetic relationships of the complete chloroplast genome construction are chaotic (Online Resource 7). The base group of ferns (Equisetales, Ophioglossales and Schizaeales) did not form a monophyletic branch. Lindsaeaceae and Saccoloma inaequale and Neocheiropteris palmatopedata in Polypodiales are located at the base of ferns. In the tree constructed with the concatenated data set (rbcL and matK) shows that the phylogenetic relationships among the various groups are relatively clear (Online Resource 8): the basal group forms a monophyletic branch, Equisetales and Marattiales have low bootstrap support, and Salviniales and Cyathales as well as Diplopterygium glaucum and Hymenophyllales are sister groups. In two trees, Polypodiales, Rhachidosoraceae, Cystopteridaceae, Dennstaedtiaceae, Lindsaeaceae and Saccolomataceae have uncertain positions. We referred to the PPG I system at the order and family levels, and manually adjusted the tree to obtain the final phylogenetic tree for analyses of evolutionary rates (Fig. 1).
Analysis of the rate of evolution
For IR-109, it can be intuitively observed numerically that 80% (for trst), 94% (for trsv), 94% (for ratio), 84% (for dN), 76% (for dS) and 86% (for ω) of the species branches had exon 1 values greater than or equal to exon 2–3 values (Online Resource 9). The parameter mean value of exon 1 was 1.9–6.7 times that of exon 2–3, and there were significant differences, except in ratio (P = 0.05) (Fig. 3a). For LSC-11, only trst was significantly different (P = 0.028) and exon 2–3 had twice the value of exon 1 (Fig. 3b).
For exon 1, the mean value in IR-109 was lower than in LSC-11, and there were significant differences in trsv and ratio (Fig. 4a). For exon 2–3, the mean value of LSC-11 was 1–11 times that of IR-109, and the difference was significant (Fig. 4b). For rps12-CDS, the mean value of LSC-11 (except ω, P = 0.893) was 1.6–6 times that of IR-109, with a significant difference (Fig. 4c).
The results of hypothesis testing are shown in Fig. 5. For all four parameters, more than 80% of the branches of species that could be effectively constrained were acceptable for hypothesis testing.
Hypothesis testing. Set the original data as an alternative hypothesis, and the null hypothesis is to constrain the value of exon2-3 to be equal to exon1 and perform the likelihood ratio test of the two hypotheses. accept: P > 0.05. refuse: P < 0.05. unable: the number of the species branches that cannot be effectively constrained by assumptions. The number represents the number of branches of a species
Selection pressure and Identification of positively selected amino acid sites
With HyPhy, setting the Bayes factor value as greater than 20, we detected 2 positively selected sites (17G, 74L) and 107 negatively selected sites (Fig. 6).
Amino acid sequence alignment (with Drynaria roosii as the reference sequence). The number 1–123 indicates the amino acid site number. Blue area indicates the sites located in the α-helix region. Green area indicates the sites located in the β-sheet region. "*" represents the consistent sites of 126 amino acid sequence alignment. With Hyphy, under the condition that the Bayes factor is greater than 20, 2 positive selection sites (pink area-17 and red area-74) and 107 negative selection sites (gray area) are screened out. Three positive selective amino acid sites (red area-74, aurantium area 16 and 14) were identified using PAML with a posterior probability of P > 0.95
In the branch model, the F model detected 50 branches with ω > 1: these were used as foreground branches in the branch-site model. In Model2, for three different sequence sets (rps12-CDS, exon 1 and exon 2–3), the likelihood ratio test results of the foreground (LSC-11) and background branches (IR-109) were calculated. The results for M0 and Model2 showed that ω1 of exon 2–3 was 3.3 times that of ω2 (P = 0). In the site model, the likelihood ratio tests between the four pairs of models differed significantly. The results of M1a-M2a were taken as the standard and a posterior probability greater than 0.95 is taken as the condition to identify three positively selected amino acid sites (14Q, 16G, 74L). The likelihood ratio test of the branch-site model, null and alternative was significantly different (P = 0.031), and no positive selective amino acid sites were identified when the posterior probability was greater than 0.95 (Tables 2 and 3).
Amino acid sequence alignment
The amino acid sequences of rps12 from 126 plants were aligned, and the protein structure was predicted using the rps12 amino acid sequence of Drynaria roosii as a reference. As shown in Fig. 6, there were 57 completely consistent sites, all of which experienced negative selection. The predicted protein structure detected negative selection in 38 of the 44 sites in the α-helix and β-sheet regions, and 4 positive selection sites (14Q, 16G, 17G and 74L) in the loop or irregular region.
Discussions
Decreased substitution rates
Consistent with previous studies (Li et al. 2016), the gene substitution rates decreased after entering the IR region. When exon 2–3 enters the IR region, its substitution rate significantly decreased. The substitution rate of exon 1 was 1.9–6.7 times that of exon 2–3. Similarly, that of exon 2–3 in LSC was 1–9 times that in the IR region.
Because exon 1 and exon 2–3 encode RPS12 together, we tested whether the substitution rate of exon 1 may be affected by the decrease in the exon 2–3 substitution rate in the IR region. The results showed that trsv and ratio of exon 1 located in IR-109 were significantly decreased, but no differences consistent with exon 2–3 were seen. In a previous study of 70 species (IR-61 and LSC-9), we found that exon 1 of IR-61 had significantly lower trsv and dN compared to LSC-9. Perhaps conversions and non-synonymous substitutions readily lead to amino acid changes, so exon 1 and exon 2–3 have consistent conversion and non-synonymous substitution rates. Of course, the results may arise from the large difference in the amount of data in the IR-109 and LSC-11 groups. Nevertheless, the substitution rate of rps12-CDS of IR-109 was also significantly decreased, indicating that the substitution rate of the exon 2–3 sequence affected the substitution rate of the gene-coding sequence.
The low rate of evolution in IR may be due to its double-copy-related repair mechanism (Perry and Wolfe 2002). Biased gene transformation can occur throughout the genome; due to its double-copy nature, the frequency of gene transformation is relatively higher in the IR region, and gene transformation can also occur between the two copies in the IR region (Khakhlova and Bock 2006). As a result, only a small number of substitutions can be detected, so they show a lower rate of evolution.
Selection pressure affects the substitution rate and GC content increase
Previous research has suggested that protein-level selection has little or no effect on the rate decrease (Li et al. 2016). We reached different conclusions: selection pressure may influence the decreased substitution rate. Our calculations using HyPhy and the two-ratio model in the PAML branch model show that rps12-CDS and exon 1 both fit the M0 model, showing that the selection pressure experienced by IR-109 and LSC-11 is consistent. However, exon 2–3 fits Model2 (PM0-Model2 = 0), which indicates that exon 2–3 experiences different selection pressures on the LSC-11 (ω = 0.076) and IR-109 (ω = 0.023) branches. Therefore, in LSC-11, relaxed selection would tend to increase the number of nucleotide substitutions per unit time and cause higher nucleotide substitution rates. Compared to LSC-11, IR-109 experienced constrained selection pressure, coupled with the repair effects of gene conversion, and a lower substitution rate was detected.
Studies have shown that GC content is higher in the IR region than in the SC region. There are many A/T to G/C substitutions in the IR region of cycads (Wu and Chaw 2015). Li et al. (2016) found that the gene GC content increased after entering the IR region. We also found that after exon 2–3 entered the IR region, the GC content of the third codon position significantly increased, indicating that there is a GC bias for gene conversion.
Highly conservative characteristics
Lower substitution rates often indicate that a gene is relatively stable and highly conserved. The fern rps12 coding sequence is 372 bp and the size of the three exon sequences is also constant. At the amino acid level, 87% of the amino acid sites experienced strong negative selection, including 39 of the 44 sites in the α-helix and β-sheet domains, and most of the sites interacting with the S4, S8, 16 and S17 proteins. The four positive selection sites in the loop or irregular region have limited effects on the structure and function of the rps12 gene. From prokaryotes to angiosperms, the amino acid sequence has remained high consistent during evolution. All of these results demonstrate the highly conserved nature of the rps12 sequence. Corresponding to its conservative characteristics, exon 2–3 of the ferns and the ancestral lycophytes is located in the IR region, indicating that exon 2–3 is more inclined to be located in the IR region to maintain a lower substitution rate.
Conclusion
The results of this study of the rps12 gene provide strong evidence for decreased substitution rates and increased GC content when the gene is located in the IR region. The gene is highly conserved, providing a reference for understanding the structural changes and functional variation in the rps12 gene. The decreased substitution rates of gene sequences and relatively high conservation in the IR region seem to characterize the IR region. Study of the IR region provides insights into the basic characteristics of chloroplast genomes and the heterogeneity of substitution rates.
Availability of data and material
Data are available as Electronic Supplementary Material.
References
Bromham L, Penny D (2003) The modern molecular clock. Nat Rev Genet 4:216–224. https://doi.org/10.1038/nrg1020
Hollingsworth PM, Graham SW, Little DP (2011) Choosing and using a plant DNA barcode. PLoS One 6:e19254. https://doi.org/10.1371/journal.pone.0019254
Kearse M, Moir R, Wilson A et al (2012) Geneious Basic: an integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 28:1647–1649. https://doi.org/10.1093/bioinformatics/bts199
Khakhlova O, Bock R (2006) Elimination of deleterious mutations in plastid genomes by gene conversion. Pl J 46:85–94. https://doi.org/10.1111/j.1365-313X.2006.02673.x
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Molec Biol Evol 33:1870–1874. https://doi.org/10.1093/molbev/msw054
Li FW, Kuo LY, Pryer KM, Rothfels CJ (2016) Genes translocated into the plastid inverted repeat show decelerated substitution rates and elevated GC content. Genome Biol Evol 8:2452–2458. https://doi.org/10.1093/gbe/evw167
Lin CP, Wu CS, Huang YY, Chaw SM (2012) The complete chloroplast genome of Ginkgo biloba reveals the mechanism of inverted repeat contraction. Genome Biol Evol 4:374–381. https://doi.org/10.1093/gbe/evs021
Lowe TM, Eddy SR (1997) tRNAscan-SE: a program for improved detection of transfer RNA genes in genomic sequence. Nucl Acids Res 25:955–964. https://doi.org/10.1093/nar/25.5.955
Maier RM, Neckermann K, Igloi GL et al (1995) Complete sequence of the maize chloroplast genome: gene content, hotspots of divergence and fine tuning of genetic information by transcript editing. J Molec Biol 251:614–628. https://doi.org/10.1006/jmbi.1995.0460
Norihiro Z, Keita T, Kazuo S et al (1987) Trans splicing in vivo: joining of transcripts from the ‘divided’ gene for ribosomal protein S12 in the chloroplasts. Febs J 210:153–156. https://doi.org/10.1016/0014-5793(87)81326-1
Perry AS, Wolfe KH (2002) Nucleotide substitution rates in legume chloroplast DNA depend on the presence of the inverted repeat. J Molec Evol 55:501–508. https://doi.org/10.1007/s00239-002-2333-y
Pond SL, Frost SD, Muse SV (2005) HyPhy: Hypothesis testing using phylogenies. Bioinformatics 21:676–679. https://doi.org/10.1093/bioinformatics/bti079
PPG I The Pteridophyte Phylogeny group, (2016) A community-derived classification for extant lycophytes and ferns. J Syst Evol 54:563–603. https://doi.org/10.1111/jse.12229
Stamatakis A (2014) RaxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogeies. Bioinformatics 30:1312–1313. https://doi.org/10.1093/bioinformatics/btu033
Wolfe KH, Li WH, Sharp PM (1987) Rates of nucleotide substitution vary greatly among plant mitochondrial, chloroplast, and nuclear DNAs. Proc Natl Acad Sci USA 84:9054–9058. https://doi.org/10.1073/pnas.84.24.9054
Wu CS, Chaw SM (2015) Evolutionary stasis in Cycad plastomes and the first case of plastome GC-biased gene conversion. Genome Biol Evol 7:2000–2009. https://doi.org/10.1093/gbe/evv125
Wyman SK, Jansen RK, Boore JL (2004) Automatic annotation of organellar genomes with DOGMA. Bioinformatics 20:3252–3255. https://doi.org/10.1093/bioinformatics/bth352
Yang Z (2007) PAML 4: phylogenetic analysis by maximum likelihood. Molec Biol Evol 24:1586–1591. https://doi.org/10.1093/molbev/msm088
Zerbino DR, Birney E (2008) Velvet: algorithms for de novo short read assembly using de Bruijn graphs. Genome Res 18:821–829. https://doi.org/10.1101/gr.074492.107
Zhu AD, Guo WH, Gupta SS et al (2016) Evolutionary dynamics of the plastid inverted repeat: the effects of expansion, contraction, and loss on substitution rates. New Phytol 209:1747–1756. https://doi.org/10.1111/nph.13743
Acknowledgements
The authors wish to thank Ziying Liang, and Xiuqin Ma for technical assistance and sharing opinions, ShanShan Liu for the sample. This work was funded by the National Natural Science Foundation of China (31670200, 31770587, 31872670 and 32071781).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Handling Editor: Yunpeng Zhao.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Information on Electronic Supplementary Material
Below is the link to the electronic supplementary material.
Information on electronic supplementary material
Information on electronic supplementary material
Online Resource 1. Plastomes sampled in this study.
Online Resource 2. Alignment of the complete chloroplast genome sequences used in this study.
Online Resource 3. Alignment of the tandem data set of rbcL and matK sequences used in this study.
Online Resource 4. Alignment of the rps12-CDS sequences used in this study.
Online Resource 5. Alignment of the exon1 sequences used in this study.
Online Resource 6. Alignment of the exon2-3 sequences used in this study.
Online Resource 7. Maximum likelihood (ML) tree constructed based on the complete chloroplast genome.
Online Resource 8. Maximum likelihood (ML) tree constructed based on concatenated data sets of rbcL and matK.
Online Resource 9. Distribution of exon 1 and exon 2-3 values for each species in IR-109.
Rights and permissions
About this article
Cite this article
Ping, J., Li, A., Feng, P. et al. The highly conserved rps12 gene in ferns provides strong evidence for decreased substitution rates in the inverted repeat region. Plant Syst Evol 307, 26 (2021). https://doi.org/10.1007/s00606-021-01750-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00606-021-01750-7