Abstract
Background
Gestational diabetes mellitus is an endocrine and metabolic disorder that appears for the first time during pregnancy and causes varying degrees of short- and/or long-term effects on the mother and child. The etiology of the disease is currently unknown and isobaric tags for relative and absolute quantitation proteomics approach, the present study attempted to identify potential proteins in placental tissues that may be involved in the pathogenesis of GDM and adverse foetal pregnancy outcomes.
Methods
Pregnant women with GDM hospitalised were selected as the experimental group, and pregnant women with normal glucose metabolism as the control group. The iTRAQ protein quantification technology was used to screen the differentially expressed proteins between the GDM group and the normal control group, and the differentially expressed proteins were analysed by GO, KEGG, PPI, etc., and the key proteins were subsequently verified by western blot.
Results
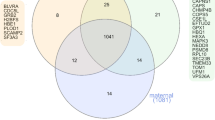
Based on the proteomics of iTRAQ, we experimented with three different samples of placental tissues from GDM and normal pregnant women, and the total number of identified proteins were 5906, 5959, and 6017, respectively, which were similar in the three different samples, indicating that the results were reliable. Through the Wayne diagram, we found that the total number of proteins coexisting in the three groups was 4475, and 91 differential proteins that could meet the quantification criteria were strictly screened, of which 32 proteins were up-regulated and 59 proteins were down-regulated. By GO enrichment analysis, these differential proteins are widely distributed in extracellular membrane-bounded organelle, mainly in extracellular exosome, followed by intracellular vesicle, extracellular organelle. It not only undertakes protein binding, protein complex binding, macromolecular complex binding, but also involves molecular biological functions such as neutrophil degranulation, multicellular organismal process, developmental process, cellular component organization, secretion, regulated exocytosis. Through the analysis of the KEGG signaling pathway, it is found that these differential proteins are mainly involved in HIF-1 signaling pathway, Glycolysis/Gluconeogenesis, Central carbon metabolism in cancer, AMPK signaling pathway, Proteoglycans in cancer, Protein processing in endoplasmic reticulum, Thyroid cancer, Alcoholism, Glucagon signaling pathway.
Discussion
This preliminary study helps us to understand the changes in the placental proteome of GDM patients, and provides new insights into the pathophysiology of GDM.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Gestational diabetes mellitus is an endocrine and metabolic disease that occurs for the first time during pregnancy and is characterized by disorders of glucose metabolism [1]. According to a 2019 study by the International Study Group on Diabetes and Pregnancy, the prevalence of GDM during pregnancy in pregnant women around the world is 17.8 percent [2]. The prevalence varies by diagnostic criteria, region, and ethnicity [3]. The prevalence of GDM in developed countries such as Europe and the United States is 5–8% [4], while in developing countries it is as high as 15–20% [5]. In 2013, a multicenter study in China showed that the prevalence of GDM in pregnant women during pregnancy was as high as 17.5% [6]. However, with the changes in people’s lifestyles, the improvement of living standards, the increase in overweight and obesity, and the increase in the number of older pregnant women due to the full liberalization of the three-child policy, the prevalence of GDM may gradually increase [7].
GDM is one of the most common complications of pregnancy, with varying degrees of short- and/or long-term effects on both the mother and the infant [8]. The short-term effects of GDM on the pregnant woman include complications of hypertensive disorders of pregnancy, multiple pregnancies, premature rupture of membranes, infections, obstructed shoulder dystocia, post-partum hemorrhage and an increase in cesarean section rates [9]. The short-term effects on the fetus are that hyperglycemia in the early stages of pregnancy can lead to abnormal embryonic development or even death, and the incidence of miscarriage can be as high as 15–30% [10]. In the second and third trimesters of pregnancy, it can lead to fetal growth restriction, macrosomia, preterm labor, fetal distress, and even intrauterine fetal death; in the perinatal period, it can also lead to complications such as neonatal hypoglycemia and neonatal respiratory distress syndrome. The maternal hyperglycemic environment is not only harmful to the mother and child during pregnancy, but also has many adverse effects in the postpartum period and even in the future [11]. For example, mothers and their offspring will have an increased risk of developing type 2 diabetes and cardiovascular-related diseases in the long term [12, 13]. A retrospective analysis of the long-term effects of gestational diabetes mellitus in pregnant women from abroad [14] showed that the long-term risk of gestational diabetes mellitus in pregnant women was 20 times higher than that of normal pregnant women, the risk of ischemic heart disease was 2.8 times higher than that of normal pregnant women, and the risk of hypertension was twice as high as that of normal pregnant women. Recently, there have also been studies on the long-term effects of GDM on newborns. High blood glucose in GDM can stimulate an inflammatory response that can lead to abnormal fetal brain development, resulting in memory loss and cognitive deficits [15]. Other studies have shown that pregnant women with gestational diabetes will increase their offspring’s asthma risk. The placenta is an important organ unique to pregnancy that mediates between the fetus and the mother. In addition to its important duties such as the material exchange between mother and child, defense barrier, and immune regulation, it also has powerful secretory and synthetic functions [16], which play an important role in gestation and ensure the smooth progress of the entire pregnancy process [17]. Gestational diabetes mellitus is diagnosed by OGTT at 24–28 weeks when the placental function gradually develops and matures; after delivery (and delivery of the placenta), most patients do not need any treatment, and glucose metabolism disorders can be rapidly returned to normal. These clinical characteristics suggest that the pathogenesis of GDM is closely related to placental tissue.
Due to the lack of early pregnancy diagnostic markers, the diagnosis of GDM usually occurs between 24 and 28 weeks, when the foetus may already be affected to varying degrees. It is therefore imperative to discover diagnostic markers for GDM [18]. By identifying specific biomarkers associated with GDM, we can develop more effective and accurate diagnostic tools for use early in pregnancy. Early detection of GDM will facilitate timely intervention and support, thereby reducing adverse outcomes for both mother and foetus. Proteomic approaches offer a promising avenue to explore the pathogenesis of GDM. By analysing the abundance and activity of proteins in maternal serum, placental tissues and other relevant biological samples, we can gain a deeper understanding of the underlying molecular changes in GDM.
Methods
Patients and tissue samples
We selected pregnant women with gestational diabetes mellitus as the experimental group and pregnant women with normal glucose metabolism as the control group, and matched them by age, gestational week, pregnancy and delivery. This study was approved by the ethics committee of Affiliated Hospital of Guizhou Medical University (Document ID:2019 Ethical approval NO.221) and was implemented in accordance with the Declaration of Helsinki. All research subjects signed a written informed consent. According to the 2014 Guidelines for the Diagnosis and Treatment of Diabetes Mellitus in Combined Pregnancy of the Obstetrics and Gynecology Section of the Chinese Medical Association, all pregnant women underwent a 75-g oral glucose tolerance test (OGTT) during the 24–28 weeks of pregnancy. Inclusion criteria: (i) age between 18 and 35 years old; (ii) a single pregnancy conceived naturally; (iii) 75 g OGTT test at 24–28 weeks of pregnancy; (iv) regular and standardized prenatal checkups during pregnancy without complications or sequelae during pregnancy; and (v) all terminated the pregnancy by cesarean section.
Collection of placental tissue specimens: after removing the meconium and amniotic membrane, 1 cm3 chorionic lobules were cut from the maternal surface close to the umbilical cord (to avoid hemorrhage, necrosis, and calcification), and washed with cold saline to remove contaminated blood. Placentas from GDM patients and controls were collected using the method described above. For proteomics studies, placental tissues were immediately frozen in a refrigerator at 80 °C.
Protein extraction and digestion
Frozen tissues were dissolved in lysis buffer containing 7 M urea, 2 M thiourea, 1% (w/v) dithiothreitol (DTT), and 1% (v/w) protease inhibitor, and lysed with an ultrasonic tissue disrupter from Ningbo Xinzhiqe Instrumentation Research Institute for 10 min, and then lysed sufficiently for 1 h at 4 °C to obtain the placental protein extract. Then, the insoluble molecules were removed by centrifugation at 12,000×g for 1 h at 4 °C. The supernatant was collected and the concentration of human serum albumin was determined by the Bradford method. 150 µg of protein from each sample was reduced and alkylated and digested with trypsin for 14 h at 37 °C (mass spectrometry grade; Promega, Madison, WI, USA). After desalting, peptides were dried by centrifugation in a vacuum concentrator (Savant™ DNA SpeedVac™ Concentrator Kit, DNA 120, 230 V 50 Hz) for 6 h. Peptides were then solubilized in liquid chromatography-tandem mass spectrometry (LC–MS) buffer (solution water: acetonitrile: formic acid, 98:2:0.1, v/v/v), and quantified by BCA quantitation.
Statistical analysis
A high-resolution mass spectrometer (AB Sciex Triple TOFTM 6600) was used for detection. The mass spectrometry data obtained were analyzed by searching for proteins in the UniProt database, protein identification using ProteinPilot™ Software 5.0.1, searching for proteins in the International Swissprot database, and analyzing the peak area integrals of the reporter ions in a relatively quantitative manner, to obtain all the proteins in each dose group in a Relative proportions.
The screening criteria for differentially expressed proteins: (1) The total number of proteins was counted when the number of peptides was ≥ 2; (2) The multiplicity of differences was ≥ 1.5-fold or ≤ 0.67-fold, with a statistical test of p value < 0.05; and (3) the first two replications were performed with proteins of the same criteria. Among the differentially expressed proteins, those with T: C average folding folds ≥ 1.5-fold were up-regulated proteins, and those ≤ 0.67-fold were down-regulated proteins.
Bioinformatics analysis
Bioinformatics analysis was performed using the DAVID analysis system and the String database (http://stringdb.org/, version 9.1). A collection of DEPs was annotated using GO (Gene Ontology), and biological process (BP) and cellular component (CC) were included in GO annotations. KEGG (Kyoto Encyclopedia of Genes and Genomes) pathways associated with these DEPs were analyzed. p value of enrichment analysis was controlled by FDR (false discovery rate) (p < 0.05). Cytohubba software (v3.9.1, http://chianti.ucsd.edu/cytoscape-3.5.1/) was used for ClueGO analysis and screening of hub genes. ClueGO analysis included BP, molecular functions (MFs) analysis, and KEGG, REACTOME and Wiki pathway analysis [19, 20]. VEEN diagram was drawn by using bioinformatics online software (http://www.bioinformatics.com.cn/login/).
HE staining of placental tissue
The HE staining experiment was as follows: placental tissue was dehydrated, hyalinized, waxed, embedded, sectioned, patched, skimmed with water, HE stained, washed, differentiated, rinsed, stained with eosin, dehydrated, hyalinized, and mounted for observation.
Western blot analysis
Total protein (15 μg) was separated by sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS-PAGE) and transferred to a polyvinylidene fluoride (PVDF) membrane. The membranes were closed with 5% milk for 1 h at room temperature, washed and incubated overnight at 4 °C with the following primary antibody: anti-EEF2 antibody (1:8000 dilution; Proteintech Group; No:20107-1-AP) anti-IGF1R antibody (1:1000 dilution; Proteintech Group; No: 20254-1-AP), anti-SLC2A1 antibody (1:1000 dilution; Proteintech Group; No:21829-1-AP). The membranes were washed with tris buffered saline and Tween 20 (TBS-T) buffer solution (3 times for 10 min each) and then incubated with horseradish peroxidase (HRP) coupled diabody at room temperature for 1 h. Immunoreactive bands were visualized with enhanced chemiluminescence (ECL) detection reagent by X-ray film. The intensity of the obtained bands was quantitatively analyzed using ImageJ software.
Statistical analysis
ImageJ analysis software (National Institutes of Health) and GraphPad Prism software (version 9.1.0; were used to statistically analyze and quantify the plots of protein bands). A comparison of the two groups was performed using t test. All data were expressed as mean ± standard deviation (SD), and a difference of p < 0.05 was considered statistically significant.
Results
Patient information and hematoxylin–eosin staining of placental tissue
Table 1 summarizes the clinical information of each group. Compared with CTR group, there were no statistically significant differences in age, BMI, fetal birth weight and gestational age among the GDM groups (p > 0.05). The results of HE staining of placental tissue from GDM patients and controls are shown in Fig. 1. Compared with controls, patients with GDM have an increased number of chorioallantoic vessels, increased syncytiotrophoblast nodules and increased fibrous exudates, villous interstitial vascular overfilling.
The morphological changes of placental tissue in GDM patients were observed by hematoxylin–eosin staining. Compared with controls, patients with GDM have an increased number of chorioallantoic vessels, increased syncytiotrophoblast nodules (red arrow) and increased fibrous exudates (black arrow), villous interstitial vascular overfilling (blue arrow) (colour figure online)
Screening of differentially expressed proteins in placental tissues of GDM/CTR group
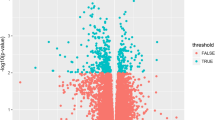
According to the results, the number of proteins with peptide number ≥ 2 in each group was counted into the total number of proteins. The number of proteins with peptide number ≥ 2 in the first group was 5906, the number of proteins with peptide number ≥ 2 in the second group was 5959, and the number of proteins with peptide number ≥ 2 in the third group was 6017. By means of the Wayne diagram, all proteins from the three groups of iTRAQ were merged, and there were a total of 4475 proteins in the three experimental groups. In unsupervised PCA analysis, there was some overlap between GDM and CON groups despite the general trend towards separation of placenta protein profiles (Fig. 2A). Cases and controls could be distinguished using orthogonal partial least squares discriminant analysis (OPLS-DA) (Fig. 2B). In addition, the intercept of the regression line on the Y-axis of 200 permutations test was negative, indicating that the model has not been over-fitted Fig. 2C. When the multiplicity of difference is ≥ 1.5 or ≤ 0.67, and the p value is < 0.05 by statistical test, it can be regarded as differential proteins. After screening, 91 proteins could meet the strict quantification criteria, including 32 up-regulated proteins and 59 down-regulated proteins (Fig. 2D). These DEPs were used for layer clustering analysis, the results showed that the GDM group could be well distinguished from the CON group.
Overview of the expressions of placental tissue proteins in the GDM/CON group. A PCA analysis of GDM and CON groups. B PLS-DA analysis of GDM and CON groups. C 200 random replacement tests. D The distribution of proteins in the GDM/CON group was depicted by volcano plots; there were 91 DEPs, 32 proteins were up-regulated and 59 proteins were down-regulated. The number in parentheses is the number of DEPs
Analysis of differentially expressed proteins in GDM/CTR group based on GO enrichment and KEGG pathway
Using the OmicsBean database, 91 differential proteins from three groups of GDM and normal placental tissues were analyzed by GO enrichment, including biological processes (BP), cellular components (CC) and molecular functions (MF). In biological processes (BP), these proteins were mainly involved in neutrophil degranulation cellular component organization, biological regulation, single-organism cellular process, developmental process, response to chemical, regulated exocytosis, secretion and other biological processes (Fig. 3A). In cellular components (CC), these differential proteins were widely distributed and localized mainly in extracellular membrane-bounded organelle, followed by the extracellular exosome, extracellular vesicle, extracellular organelle, membrane-bounded vesicle, extracellular region part extracellular space (Fig. 3B). In terms of molecular functions (MF), these differential proteins are mainly involved in binding functions, such as protein binding, protein complex binding, macromolecular complex binding, identical protein binding, poly(A) RNA binding, extracellular matrix structural constituent, conferring compression resistance, cell adhesion molecule binding, nucleosome binding, cadherin binding (Fig. 3C).
GO enrichment analysis of the differentially expressed proteins in the GDM/CON group. A The top 20 categories of enriched biological process (BP) associated with DEPs in the GDM/CON group. B The top 20 enriched cell component (CC) categories associated with DEPs in the GDM/CON group. C The top 20 molecular functions (MF) categories associated with DEPs in the GDM/CON group
In order to better understand the biological pathways of differential proteins, we analyzed the KEGG signaling pathway of differential proteins and found that the main enrichment pathways of differential proteins were HIF-1 signaling pathway, Glycolysis / Gluconeogenesis, Central carbon metabolism in cancer, AMPK signaling pathway, Proteoglycans in cancer, Protein processing in endoplasmic reticulum, Thyroid cancer, Alcoholism, Glucagon signaling pathway (Fig. 4A). The metabolic pathways with the highest proportion, IGF1R, SLC2A1, PFKL were highly expressed in the GDM group. The PPI network of DEPs was obtained by searching the OmicsBean database (Fig. 4B).
KEGG pathway analysis of the differentially expressed proteins and protein–protein interaction network analysis of differentially expressed proteins in the GDM/CON group. A The significantly enriched KEGG pathways linked to DEPs in the GDM/CON group. B Protein–protein interaction network analysis of differentially expressed proteins in the GDM/CON group
CluGO-based differential expression protein analysis of key pathways in the GDM/CON group
The hub genes of GDM/CON group were analyzed using Cytohubba software, and 12 algorithms were selected for screening. Finally, the top 10 hub genes, including GAPDH, CANX, DCN, H3F3B, EEF1A1, H1F0, EEF2, IGF1R, SLC2A1 and S100A8 were enriched (Fig. 5A). They are associated with metabolic process.
Functional analysis of differentially expressed proteins and Hub gene analysis in the GDM/CON group and Western blot analysis of differentially expressed proteins in placental tissue of GDM patients and controls. A Based on 12 hybrid algorithms, the horizontal stack bar depicts the top 10 hub genes in the GDM/CON group. B Western blot of EEF2, SLC2A1, IGF1R and β-ACTIN. The relative protein expression (n = 3). Each bar representing the standard deviation (SD; p < 0.05). The image analyzed by Western blot analysis on the left corresponds to the semi-quantitative histogram on the right. *p < 0.05, **p < 0.01, *** p < 0.001
Validation of the proteomics results by Western blot analysis
Three important proteins, i.e., IGF1R, EEF2 and SLC2A1 were selected for Western blot analysis verification. These 3 proteins were involved in different signal transduction pathways. As shown in Fig. 5B, SLC2A1 and IGF1R are down-regulated in the GDM group, EEF2 is up-regulated, and the differences were statistically significant, which was consistent with the results of proteomic analysis.
Discussion
The placenta, a vital organ in pregnancy, plays a crucial role in regulating various metabolic activities between the mother and the fetus [21]. Any damage or impairment to its structure and function can have significant consequences on the outcome of pregnancy and pose potential risks to the long-term health of the offspring [22,23,24]. In this study, we collected placental tissues from both patients with GDM and the control group. By performing HE staining and examining the morphologic and histologic changes in the placenta of GDM patients, we conducted a thorough analysis. Through meticulous microscopic observation, we discovered several notable abnormalities in the placentas of GDM patients, including an increased number of chorioallantoic vessels, elevated levels of syncytiotrophoblast nodules, and excessive fibrous exudates. Additionally, we observed villous interstitial vascular overfilling. The syncytiotrophoblast layer, which plays a crucial role in facilitating nutrient and gas exchange between the mother and the fetus, exhibited an escalated formation of nodules in GDM patients. These nodules serve as indicators of degeneration or local thickening within the placenta. Moreover, fibrous exudates, denoting the accumulation of fibrous tissue within the placental tissue, were found to be more prevalent in GDM patients. This excessive deposition can potentially impact the structure and function of the placenta. Furthermore, we noted that villous interstitial vascular overfilling is likely a consequence of heightened blood flow to the placenta, triggered by elevated maternal glucose levels associated with GDM. Placental dysplasia is related to various pathological conditions during pregnancy, such as GDM and Intrahepatic cholestasis of pregnancy [25].
The iTRAQ proteomics used in this study has the advantages of high protein throughput, good quantitative stability, and accuracy. The method has high sensitivity and reliable detection results. In addition, it does not affect the post-translational modification of proteins and the source of samples is not restricted [26]. Analysis of proteomics results showed that supervised OPLS-DA analysis was able to distinguish the GDM group from the control group. The results showed significant alterations in placental protein expression patterns in GDM and healthy pregnant women. A total of 91 DEPs, including 32 up-regulated and 59 down-regulated proteins, were identified in the GDM/CTR group. According to the bioinformatics results, these DEPs were mainly involved in the HIF-1 signaling pathway, Glycolysis/Gluconeogenesis, Protein processing in the endoplasmic reticulum AMPK signaling pathway, Apoptosis, arachidonate transport, cellular senescence. Among the metabolic pathways with the highest ratio, IGF1R, SLC2A1, and PFKL were down-regulated in the GDM group. cytohubba found the top 10 DEPs with the highest weighting values based on 12 algorithms. The 10 central genes with the highest weight values were found to be GAPDH, SLC2A1, LDHB, CANX, EEF1A1, H3F3B, IGF1R, S100A8, S100A10 and EEF2 whose molecular functions include glycolysis, glucose transport, glucose metabolism, cell proliferation and angiogenesis.
In humans, there is no significant placental glucose xenobiosis, so adequate glucose transfer from the mother to the fetus is necessary for normal fetal growth [27]. Glucose transfer occurs through glucose transporters embedded in placental syncytiotrophoblast microvilli [28] and basement membranes human placental glucose transport in fetoplacental growth [29] and metabolism [30]. In our research, we have discovered a significant decrease in the expression levels of the glucose transporter protein SLC2A1 [31], which is responsible for transporting glucose. The SLC2A1 protein, also known as the type 1 glucose transporter (GLUT1), GLUT1 is responsible for transporting glucose across cell membranes and is critical for glucose uptake in various tissues including the placenta. Decreased levels of GLUT1 may result in impaired glucose transport and utilization in patients with gestational diabetes [32]. This may lead to elevated glucose levels in the blood due to the inability of cells to efficiently take up and utilize glucose. In addition, GLUT1 function is associated with insulin sensitivity [27]. Insulin resistance, common in gestational diabetes, further impedes the function of GLUT1. Reduced activity of GLUT1 in response to insulin can exacerbate glucose intolerance, leading to elevated blood glucose levels.
During pregnancy, the mother’s body undergoes a series of physiologic changes to support the growth needs of the fetus [33]. These changes include adaptations to the cardiovascular, renal, hematologic, respiratory, and metabolic systems. An important metabolic adaptation is insulin sensitivity. During early pregnancy, insulin sensitivity increases, promoting the conversion of glucose to fat storage in preparation for the energy demands of late pregnancy [34]. Insulin resistance in gestational diabetes mellitus (GDM) is defined as a diminished response of the body’s cells to the effects of the hormone insulin. Insulin resistance is a normal physiological adaptation during pregnancy [35], but in gestational diabetes, it becomes more severe and may lead to elevated blood glucose levels. Insulin resistance in GDM is caused by a variety of factors, including hormonal changes, increased placental secretion, adipose tissue dysfunction, and genetic factors. Insulin receptor substrates (IRS) are proteins that play a key role in the insulin signaling pathway [36]. When insulin binds to receptors on the cell surface, it activates a series of intracellular signaling events that regulate various metabolic processes. Insulin receptor substrates, which are key mediators of this signaling cascade, are phosphorylated by the insulin-like growth factor 1 receptor (IGF1R) when stimulated by IGF1 or IGF2 [20, 21], and tyrosine-phosphorylated IRSs interact with the cytoplasmic protein PI3K through its SH2 structural domain.
Activation of PI3K leads to the transmission of functional effects of IGFs [36], such as enhanced glucose transport and enhanced cardiomyocyte contractility. Therefore, when IGF1R is deficient, it affects IRS phosphorylation, which in turn may affect glucose metabolism. Some studies have also suggested that IGF1R may play an important role in the development and progression of gestational diabetes [37]. In addition, the downregulation of IGF1R may also affect fetal growth and development. IGF1R is essential for fetal growth because it regulates cell proliferation and differentiation. Studies have shown that deficient IGF1R signaling leads to intrauterine growth restriction [38] (IUGR) and increases the risk of neonatal malformations [39] (macrosomia of pregnancy), and its role in the development of GDM needs to be further investigated.
Based on the bioinformatics results, we found and glucose metabolism pathway also changed, LDHB is an enzyme involved in the conversion of pyruvate to lactate during anaerobic glycolysis [40], which promotes the anaerobic oxidation of glucose is essential for the maintenance of cellular energy homeostasis. In our study, LDHB levels were significantly increased in the GDM group compared to the control group, and in GDM, there is an increased demand for energy production due to impaired glucose metabolism. As a result, LDHB expression and activity were upregulated to compensate for the higher energy demand. Decreased aerobic oxidation of glucose, which may also be due to mitochondrial hypoxia or dysfunction, is thought to be impaired mitochondrial function that affects the insulin signaling pathway, leading to decreased glucose uptake and utilization in the cell, and elevated LDHB may be a reflection of mitochondrial dysfunction.
Inflammation plays an important role in the onset and development of GDM [41, 42]. During pregnancy, hormonal changes lead to insulin resistance, which means that the body’s cells are less sensitive to insulin. As a result, more insulin is needed to keep blood glucose levels in the normal range. In women with GDM, the pancreas does not produce enough insulin to overcome this insulin resistance [43, 44]. Inflammation is thought to contribute to insulin resistance [45, 46]. Chronic low-grade inflammation can disrupt the normal balance of insulin signaling pathways, impairing cellular uptake of glucose and leading to elevated blood glucose levels. This inflammation can be caused by a variety of factors such as obesity, oxidative stress, and immune system dysfunction [47]. CASP14 is an enzyme that plays a role in skin barrier function and keratinocyte differentiation [48]. CASP14 is expressed primarily in the outermost layer of the skin (stratum corneum) and is involved in the process of skin barrier formation. GDM has been associated with an increased risk of skin-related complications such as dry skin, itchiness GDM is associated with an increased risk of skin-related complications, such as dry skin, pruritus, and infections, and alterations in CASP14 activity or expression in GDM may lead to impaired skin barrier function, thereby increasing susceptibility to these complications. GDM is often characterized by a chronic low-grade inflammatory state. CASP14 has been found to have anti-inflammatory properties and modulate the immune response. Dysregulation of CASP14 in GDM may lead to increased inflammation and altered immune response, which may further affect insulin sensitivity and glucose metabolism. The immunoglobulin kappa constant (IGKC) is the constant region of the light chain of immunoglobulin kappa, a protein produced by B cells as part of the immune response [49]. GDM is associated with a chronic low-grade inflammatory state. Dysregulation of the immune system, including alterations in immunoglobulin secretion, may contribute to this inflammatory state. Changes in IGKC expression or function may affect the immune response and may contribute to inflammation in GDM [50].
We have found changes in the S100 protein family, which includes S100A8, S100A9, S100A10, and S100A11. These proteins are calcium-binding proteins that play a variety of roles in inflammation, immune responses, and cellular processes. Studies have shown that women with GDM have elevated levels of S100A8/A9 [51]. These proteins form a heterodimer called calreticulin that is involved in anti-microbial defense and inflammatory regulation. High levels of S100A8/A9 have been associated with insulin resistance [52, 53] and inflammation, which are factors associated with the development of GDM.
Disruptions in intrauterine glucose homeostasis, can compromise the immune function of pregnant women [54]. This compromised immune function is associated with an increased susceptibility to infections. One proposed mechanism involves the suppressive effects of hyperglycemia on immune cell activity [55]. Additionally, high blood glucose levels can create a more favorable environment for bacterial growth, further enhancing the risk of infection [56]. Pregnant women with underlying glucose metabolism disorders face a double jeopardy. Not only are they more prone to infections, but these infections can also lead to serious complications for both mother and fetus, including fetal growth restriction [57], preterm delivery [58], and increased labor complications. In our study, we identified changes in the glucose-related proteins GLUT1, IGF1R on the placenta, as well as changes in the CASP14, S100 protein family proteins related to immune function, which may be associated with maternal and foetal outcomes in gestational diabetes.
In conclusion, the development of GDM may result from abnormalities in maternal glucose transport, glycolysis and insulin signalling. These abnormalities can lead to structural and functional changes in the pancreas, placenta and other tissues and organs, ultimately leading to the development of GDM. Our study, as well as previous studies using high-throughput methods, provides further evidence that the expression of specific proteins or genes is altered in placental tissues of patients with GDM. These molecular changes are likely to play a role in the pathological mechanisms of GDM, leading to the development of adverse pregnancy outcomes and complications associated with GDM. By elucidating the molecular changes that occur in the placenta and other affected tissues, we have gained valuable insight into the mechanisms of GDM. This knowledge is critical to understanding the impact of the disease on maternal health and pregnancy outcomes. Ultimately, these findings provide a foundation for identifying potential targets for therapeutic intervention and prevention strategies to mitigate the adverse effects of GDM. By addressing the underlying molecular dysregulation, we aim to improve maternal health and pregnancy outcomes for those affected by GDM.
Conclusion
In conclusion, we employed a proteomic approach to investigate protein expression profile of the placenta in gestational diabetes mellitus GDM compared to a control group. The placenta plays a crucial role in facilitating maternal and fetal nutrient exchange, and change in these processes may be implicated in the pathogenesis of GDM. Some differential proteins found were closely related to the pathogenesis of GDM, providing new perspectives for GDM study. The combination of DEPs analysis can provide a deeper insight into the underlying mechanisms of GDM. The changes observed in DEP expression can serve as indicators of altered glycolysis, glucose transport, glucose metabolism, cell proliferation and angiogenesis in placental tissues of GDM pregnant women. Despite our study being preliminary and restricted to a small sample size, these findings present a promising avenue for future research. Moving forward, we aim to broaden our cohort study to include a larger number of participants. Such an approach will allow us to identify specific medications that can be effectively used to treat gestational diabetes mellitus. Additionally, we plan to collect maternal peripheral blood in order to validate our observations and explore potential early biomarkers of GDM. Our goal is to gain a more comprehensive understanding of the disease and ultimately improve its diagnosis, management, and prevention.
Data availability
All raw data have been deposited as online resource to the Figshare database: iTRAQ Proteomics analysis of placental tissue with gestational diabetes mellitus (https://doi.org/10.6084/m9.figshare.24521815).
References
McIntyre HD, Catalano P, Zhang C et al (2019) Gestational diabetes mellitus. Nat Rev Dis Primers 5(1):47. https://doi.org/10.1038/s41572-019-0098-8
Lee KW, Ching SM, Ramachandran V et al (2018) Prevalence and risk factors of gestational diabetes mellitus in Asia: a systematic review and meta-analysis. BMC Pregnancy Childbirth 18(1):494. https://doi.org/10.1186/s12884-018-2131-4
Zhu WW, Yang HX, Wei YM et al (2013) Evaluation of the value of fasting plasma glucose in the first prenatal visit to diagnose gestational diabetes mellitus in china. Diabetes Care 36(3):586–590. https://doi.org/10.2337/dc12-1157
Chandna AR, Kuhlmann N, Bryce CA et al (2015) Chronic maternal hyperglycemia induced during mid-pregnancy in rats increases RAGE expression, augments hippocampal excitability, and alters behavior of the offspring. Neuroscience 303:241–260. https://doi.org/10.1016/j.neuroscience.2015.06.063
Sellers EAC, Dean HJ, Shafer LA et al (2016) Exposure to gestational diabetes mellitus: impact on the development of early-onset type 2 diabetes in Canadian first nations and non-first nations offspring. Diabetes Care 39(12):2240–2246. https://doi.org/10.2337/dc16-1148
Tam WH, Ma RCW, Ozaki R et al (2017) In utero exposure to maternal hyperglycemia increases childhood cardiometabolic risk in offspring. Diabetes Care 40(5):679–686. https://doi.org/10.2337/dc16-2397
Ostetrics Subgroup (2022) Chinese Society of Obstetrics and Gynecology Chinese Medical Association Chinese Society of Perinatal Medicine Chinese Medical Association Committee of Pregnancy with Diabetes Mellitus China Maternal and Child Health Association [Guideline of diagnosis and treatment of hyperglycemia in pregnancy [Part two]]. Zhonghua Fu Chan Ke Za Zhi. 57(2):81–90. https://doi.org/10.3760/cma.j.cn112141-20210917-00529
Bell R, Bailey K, Cresswell T et al (2008) Trends in prevalence and outcomes of pregnancy in women with pre-existing type I and type II diabetes. BJOG 115(4):445–452. https://doi.org/10.1111/j.1471-0528.2007.01644.x
Weir TL, Majumder M, Glastras SJ (2024) A systematic review of the effects of maternal obesity on neonatal outcomes in women with gestational diabetes. Obes Rev Publ 25(7):e13747. https://doi.org/10.1111/obr.13747
Mathiesen ER, Ringholm L, Damm P (2011) Stillbirth in diabetic pregnancies. Best Pract Res Clin Obstet Gynaecol 25(1):105–111. https://doi.org/10.1016/j.bpobgyn.2010.11.001
Maternal diabetes during pregnancy and early onset of cardiovascular disease in offspring: population based cohort study with 40 years of follow-up—PubMed. Accessed 25 Oct 2022. https://pubmed.ncbi.nlm.nih.gov/31801789/
Xie W, Wang Y, Xiao S et al (2022) Association of gestational diabetes mellitus with overall and type specific cardiovascular and cerebrovascular diseases: systematic review and meta-analysis. BMJ 378:e070244. https://doi.org/10.1136/bmj-2022-070244
Kautzky-Willer A, Harreiter J, Winhofer-Stöckl Y et al (2019) Gestational diabetes mellitus (update 2019). Wien Klin Wochenschr 131(1):91–102. https://doi.org/10.1007/s00508-018-1419-8
Babaei K, Shams S, Keymoradzadeh A et al (2020) An insight of microRNAs performance in carcinogenesis and tumorigenesis: an overview of cancer therapy. Life Sci 240:117077. https://doi.org/10.1016/j.lfs.2019.117077
Włodarski A, Strycharz J, Wróblewski A et al (2020) The role of micrornas in metabolic syndrome-related oxidative stress. Int J Mol Sci 21(18):6902. https://doi.org/10.3390/ijms21186902
Amiri A, Mahjoubin-Tehran M, Asemi Z et al (2021) Role of resveratrol in modulating micrornas in human diseases: from cancer to inflammatory disorder. Curr Med Chem 28(2):360–376. https://doi.org/10.2174/0929867326666191212102407
Ghafouri-Fard S, Shoorei H, Taheri M (2020) Role of microRNAs in the development, prognosis and therapeutic response of patients with prostate cancer. Gene 759:144995. https://doi.org/10.1016/j.gene.2020.144995
Shen L, Zhao D, Chen Y et al (2019) Comparative proteomics analysis of serum proteins in gestational diabetes during early and middle stages of pregnancy. Proteom Clin Appl 13(5):e1800060. https://doi.org/10.1002/prca.201800060
Shen L, Feng C, Zhang K et al (2019) Proteomics study of peripheral blood mononuclear cells (PBMCS) in autistic children. Front Cell Neurosci 13:105. https://doi.org/10.3389/fncel.2019.00105
ClueGO: a Cytoscape plug-in to decipher functionally grouped gene ontology and pathway annotation networks—PubMed. Accessed 22 Aug 2023. https://pubmed.ncbi.nlm.nih.gov/19237447/
Maltepe E, Fisher SJ (2015) Placenta: the forgotten organ. Annu Rev Cell Dev Biol 31:523–552. https://doi.org/10.1146/annurev-cellbio-100814-125620
Thornton CA, Vance GHS (2002) The placenta: a portal of fetal allergen exposure. Clin Exp Allergy 32(11):1537–1539. https://doi.org/10.1046/j.1365-2222.2002.01543.x
Salas-Lucia F, Stan MN, James H et al (2023) Effect of the fetal THRB genotype on the placenta. J Clin Endocrinol Metab 108(10):e944–e948
Sanchez-Martinez S, Randanne PC, Hawkins-Villarreal A et al (2023) Acute fetal cardiovascular adaptation to artificial placenta in sheep model. Ultrasound Obstet Gynecol 62(2):255–265. https://doi.org/10.1002/uog.26215
Jiang Y, Yin X, Xu Q et al (2023) SWATH proteomics analysis of placental tissue with intrahepatic cholestasis of pregnancy. Placenta 137:1–13. https://doi.org/10.1016/j.placenta.2023.04.009
Martyniuk CJ, Alvarez S, Denslow ND (2012) DIGE and iTRAQ as biomarker discovery tools in aquatic toxicology. Ecotoxicol Environ Saf 76(2):3–10. https://doi.org/10.1016/j.ecoenv.2011.09.020
Green DJ, Park K, Bhatt-Mehta V et al (2021) Regulatory considerations for the mother, fetus and neonate in fetal pharmacology modeling. Front Pediatr 9:698611. https://doi.org/10.3389/fped.2021.698611
What factors determine placental glucose transfer kinetics? PubMed. Accessed 7 Oct 2023. https://pubmed.ncbi.nlm.nih.gov/23886770/
Holme AM, Roland MCP, Lorentzen B et al (2015) Placental glucose transfer: a human in vivo study. PLoS ONE 10(2):e0117084. https://doi.org/10.1371/journal.pone.0117084
Levkovitz R, Zaretsky U, Jaffa AJ et al (2013) In vitro simulation of placental transport: part II. Glucose transfer across the placental barrier model. Placenta 34(8):708–715. https://doi.org/10.1016/j.placenta.2013.05.006
Heilig CW, Deb DK, Abdul A et al (2013) GLUT1 regulation of the pro-sclerotic mediators of diabetic nephropathy. Am J Nephrol 38(1):39–49. https://doi.org/10.1159/000351989
GDM-induced macrosomia is reversed by Cav-1 via AMPK-mediated fatty acid transport and GLUT1-mediated glucose transport in Placenta—PubMed. Accessed 7 Oct 2023. https://pubmed.ncbi.nlm.nih.gov/28125642/
Choudhury AA, Devi RV (2021) Gestational diabetes mellitus—a metabolic and reproductive disorder. Biomed Pharmacother 143:112183. https://doi.org/10.1016/j.biopha.2021.112183
Di Cianni G, Miccoli R, Volpe L, Lencioni C, Del Prato S (2003) Intermediate metabolism in normal pregnancy and in gestational diabetes. Diabetes Metab Res Rev 19(4):259–270. https://doi.org/10.1002/dmrr.390
Liu Y, Kuang A, Talbot O et al (2020) Metabolomic and genetic associations with insulin resistance in pregnancy. Diabetologia 63(9):1783–1795. https://doi.org/10.1007/s00125-020-05198-1
Hanke S, Mann M (2009) The phosphotyrosine interactome of the insulin receptor family and its substrates IRS-1 and IRS-2. Mol Cell Proteom 8(3):519–534. https://doi.org/10.1074/mcp.M800407-MCP200
Zhu W, Shen Y, Liu J et al (2020) Epigenetic alternations of microRNAs and DNA methylation contribute to gestational diabetes mellitus. J Cell Mol Med 24(23):13899–13912. https://doi.org/10.1111/jcmm.15984
Darp RA, de Boo HA, Phua HH et al (2010) Differential regulation of igf1 and igf1r mRNA levels in the two hepatic lobes following intrauterine growth restriction and its treatment with intra-amniotic insulin-like growth factor-1 in ovine fetuses. Reprod Fertil Dev 22(8):1188–1197. https://doi.org/10.1071/RD09292
Stróżewska W, Durda-Masny M, Szwed A (2022) Mutations in GHR and IGF1R genes as a potential reason for the lack of catch-up growth in SGA children. Genes 13(5):856. https://doi.org/10.3390/genes13050856
Jiang X, Feng N, Zhou Y et al (2022) Slc2a6 regulates myoblast differentiation by targeting LDHB. Cell Commun Signal 20(1):107. https://doi.org/10.1186/s12964-022-00915-2
Pantham P, Aye ILMH, Powell TL (2015) Inflammation in maternal obesity and gestational diabetes mellitus. Placenta 36(7):709–715. https://doi.org/10.1016/j.placenta.2015.04.006
Abell SK, De Courten B, Boyle JA, Teede HJ (2015) Inflammatory and other biomarkers: role in pathophysiology and prediction of gestational diabetes mellitus. Int J Mol Sci 16(6):13442–13473. https://doi.org/10.3390/ijms160613442
Ponzo V, Fedele D, Goitre I et al (2019) Diet-gut microbiota interactions and gestational diabetes mellitus (GDM). Nutrients 11(2):330. https://doi.org/10.3390/nu11020330
Bianco ME, Josefson JL (2019) Hyperglycemia during pregnancy and long-term offspring outcomes. Curr Diab Rep 19(12):143. https://doi.org/10.1007/s11892-019-1267-6
Matulewicz N, Karczewska-Kupczewska M (2016) Insulin resistance and chronic inflammation. Postepy Hig Med Dosw 70:1245–1258
Olefsky JM, Glass CK (2010) Macrophages, inflammation, and insulin resistance. Annu Rev Physiol 72:219–246. https://doi.org/10.1146/annurev-physiol-021909-135846
Berbudi A, Rahmadika N, Tjahjadi AI, Ruslami R (2020) Type 2 diabetes and its impact on the immune system. Curr Diabetes Rev 16(5):442–449. https://doi.org/10.2174/1573399815666191024085838
Hoste E, Denecker G, Gilbert B et al (2013) Caspase-14-deficient mice are more prone to the development of parakeratosis. J Invest Dermatol 133(3):742–750. https://doi.org/10.1038/jid.2012.350
Zou YR, Gu H, Rajewsky K (1993) Generation of a mouse strain that produces immunoglobulin kappa chains with human constant regions. Science 262(5137):1271–1274. https://doi.org/10.1126/science.8235658
Hu Z, Zhang M, Tian Y (2020) Screening and analysis of small molecular peptides in urine of gestational diabetes mellitus. Clin Chim Acta 502:174–182. https://doi.org/10.1016/j.cca.2019.12.024
Oliva K, Barker G, Rice GE, Bailey MJ, Lappas M (2013) 2D-DIGE to identify proteins associated with gestational diabetes in omental adipose tissue. J Endocrinol 218(2):165–178. https://doi.org/10.1530/JOE-13-0010
Blanco-Rojo R, Delgado-Lista J, Lee YC et al (2016) Interaction of an S100A9 gene variant with saturated fat and carbohydrates to modulate insulin resistance in 3 populations of different ancestries. Am J Clin Nutr 104(2):508–517. https://doi.org/10.3945/ajcn.116.130898
Ortega FJ, Mercader JM, Moreno-Navarrete JM et al (2013) Targeting the association of calgranulin B (S100A9) with insulin resistance and type 2 diabetes. J Mol Med 91(4):523–534. https://doi.org/10.1007/s00109-012-0979-8
Chen JL, Dai HF, Kan XC et al (2024) The integrated bioinformatic analysis identifies immune microenvironment-related potential biomarkers for patients with gestational diabetes mellitus. Front Immunol 15:1296855. https://doi.org/10.3389/fimmu.2024.1296855
Wang N, Zhang C (2024) Oxidative stress: a culprit in the progression of diabetic kidney disease. Antioxidants 13(4):455. https://doi.org/10.3390/antiox13040455
Zhang Y, Ni P, Miao Y et al (2024) Vitamin D3 improves glucose metabolism and attenuates inflammation in prediabetic human and mice. J Nutr Biochem 27:109659. https://doi.org/10.1016/j.jnutbio.2024.109659
Vambergue A, Fajardy I (2011) Consequences of gestational and pregestational diabetes on placental function and birth weight. World J Diabetes 2(11):196–203. https://doi.org/10.4239/wjd.v2.i11.196
Cundy T, Gamble G, Townend K et al (2000) Perinatal mortality in type 2 diabetes mellitus. Diabet Med 17(1):33–39. https://doi.org/10.1046/j.1464-5491.2000.00215.x
Funding
This work was supported by National Natural Science Foundation of China (No. 81960284), Science and Technology Support Program of Science and Technology Department of Guizhou Province (No: Qian Ke He Zhi Cheng [2022] Yi Ban 183) and the Shenzhen-Hong Kong Institute of Brain Science-Shenzhen Fundamental Research Institutions (No. 2023SHIBS0003).
Author information
Authors and Affiliations
Contributions
Conception and design: XY, DZ, and LS; experimental operation: XY, JL, QH and XT; provision of materials or patient’s information: LY, XY and GM; collection and assembly of data: FY and HZ; manuscript writing: XY and FY; manuscript revision: DZ and LS. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Ethical approval
This study was approved by the ethics committee of the Affiliated Hospital of Guizhou Medical University (Document ID: 2019 Ethical Review No. 221) and was implemented in accordance with the Declaration of Helsinki. Written informed consent was obtained from all Patients to participate in the study.
Additional information
Managed by Annunziata Lapolla.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Yin, X., Yang, F., Lin, J. et al. iTRAQ proteomics analysis of placental tissue with gestational diabetes mellitus. Acta Diabetol (2024). https://doi.org/10.1007/s00592-024-02321-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00592-024-02321-1