Abstract
Background
Upregulated Kindlin-2 expression in hepatocellular carcinoma (HCC) correlates with metastasis and poor prognosis. In this study, we investigated the molecular mechanism of Kindlin-2 in HCC.
Methods
Kindlin-2 downstream pathways were explored through microRNA sequencing. The Kindlin-2–miR-1258–TCF4 axis was verified using bisulfite sequencing, a luciferase reporter assay, quantitative real-time PCR, and rescue assays. Binding of TCF4 to the Kindlin-2 promoter was confirmed by promoter activity analysis and chromatin immunoprecipitation.
Results
MiRNA sequencing identified miR-1258 as a downstream effector of Kindlin-2. MiR-1258 expression was increased following Kindlin-2 knockdown and decreased after Kindlin-2 overexpression. Next, we identified transcription factor 7 like 2 (TCF7L2 or TCF4) as a target of miR-1258 and found that Kindlin-2 upregulated TCF4 expression by epigenetically suppressing miR-1258 in HCC. Furthermore, our results suggest that TCF4 binds to the Kindlin-2 promotor to enhance its transcription. Therefore, Kindlin-2–miR-1258–TCF4 interaction creates a positive feedback loop. Functional assays and animal experiments demonstrated critical roles of miR-1258 and TCF4 in HCC cell migration in vitro and HCC metastasis in vivo. In HCC tissues, Kindlin-2 expression correlated negatively with miR-1258 expression and positively with TCF4 expression. Meanwhile, miR-1258 expression correlated negatively with TCF4 expression.
Conclusions
This study illustrates a novel integrin-independent signaling pathway, Kindlin-2–miR-1258–TCF4, that regulates HCC invasion and metastasis and identifies Kindlin-2 as a promising therapeutic target in HCC.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Kindlin-2, a member of the Kindlin family also known as fermitin family homolog 2 (FERMT2), is essential for integrin activation and several physiological processes [1]. Kindlin-2 has gained attention recently due to its impact on tumor cell migration, which promotes tumor metastasis [2]. We previously demonstrated that Kindlin-2 expression is upregulated in hepatocellular carcinoma (HCC) and correlates with poor prognosis, and furthermore, we showed that Kindlin-2 enhances HCC cell adhesion, migration, and invasion in vitro [3]. In esophageal squamous cell carcinoma, Kindlin-2 downregulation inhibits the activation of integrin β1 and cancer invasion [4]. In gastric cancer, Kindlin-2 promotes cancer invasion and proliferation by activating integrin β1 and integrin β3. Furthermore, Kindlin-2 downregulation decreases the expression of integrin β3 [5]. Kindlin-2 contributes to epithelial–mesenchymal transition (EMT) through the microRNA (miR)-200 family to further promote tumor progression. Kindlin-2 was reported as a possible target of miR-200b, which leads to the inhibition of Kindlin-2 expression. Meanwhile, Kindlin-2 can epigenetically suppress the miR-200 family through inducing hypermethylation of the miR-200 promoter [4, 6,7,8,9]. In addition, Kindlin-2 participates in tumor-associated signaling pathways, including the transforming growth factor beta (TGF-β), Wnt/β-catenin, epidermal growth factor receptor (EGFR), and Hedgehog signaling pathways [3, 10,11,12,13]. Nevertheless, the precise mechanism by which Kindlin-2 promotes HCC invasion and metastasis remains unclear. Dissecting the Kindlin-2 signaling pathway will be instrumental for understanding HCC pathogenesis.
In the present study, we analyzed the molecular mechanism of Kindlin-2 in HCC. We identified miR-1258 by high-throughput sequencing as novel target of Kindlin-2. Furthermore, we uncovered that transcription factor 7 like 2 (TCF7L2, also referred to as T-cell-specific transcription factor 4 [TCF4]) acts as transcription factor to regulate Kindlin-2 transcription and observed that Kindlin-2–miR-1258–TCF4 signaling regulates HCC invasion and metastasis.
Methods
Cell culture
The human HCC cell lines LM3 and 97H were purchased from the Liver Cancer Institute, Fudan University (Shanghai, China). HepG2, HepG2.2.15, Hep3B, and Huh7 cell lines were acquired from the American Tissue Culture Collection (ATCC, Manassas, VA, USA). The HEK293T cell line was provided by Dr Xu Lin, Key Laboratory of Gastrointestinal Cancer (Fujian Medical University), Ministry of Education. All cell lines were maintained in Dulbecco’s modified Eagle’s medium (DMEM, Gibco, NY, USA) supplemented with 10% fetal bovine serum, 100 U/ml penicillin, and 100 mg/ml streptomycin (Gibco), and cultured in a humidified incubator maintained at 37 °C with 5% CO2.
Tissue specimens
A total of 47 fresh HCC specimens were obtained from HCC patients who underwent hepatic resection at Fujian Provincial Hospital (Fuzhou, China) during the period from June 2020 to December 2020. HCC diagnosis was confirmed by two independent histopathologists. Upon resection, specimens were immediately snap frozen in liquid nitrogen and then were stored at – 80 °C until RNA extraction. None of the patients received preoperative therapy. The study protocol was reviewed and approved by the Ethics Review Committee of Fujian Medical University (Reference No. SQ2015–036–01). Each enrolled patient signed an informed consent form before study commencement.
Western blotting
Total protein was separated on a 10% sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS-PAGE) gel and then transferred onto polyvinylidene difluoride (PVDF) membranes (Millipore, Billerica, MA, USA). After blocking, the membranes were incubated with primary antibodies specific to Kindlin-2 (Ab74030, Abcam, Cambridge, MA, USA; 1:1000), β-catenin (8480, Cell Signaling Technology; 1:1000), TCF4 (2569, Cell Signaling Technology; 1:1000), DNMT1 (ab188453, Abcam; 1:1000), DNMT3a (ab188470, Abcam; 1:2000), DNMT3b (ab79822, Abcam; 1:1000), and β-actin (4970, Cell Signaling Technology; 1:3000) overnight at 4 °C. The next day, the membranes were incubated with the corresponding horseradish peroxidase (HRP)-conjugated immunoglobulin G (IgG) secondary antibody (Cell Signaling Technology). Blots were visualized using an enhanced chemiluminescence kit (Thermo Fisher, Waltham, MA, USA). Experiments for detecting the expression level of each protein were performed in triplicate, and the values of blots for each time were measured using QuantityOne software (Bio-Rad, Hercules, CA, USA). The most representative images of blots are shown, and triplicate values from the quantitative data are plotted.
RNA extraction
Total RNA, including miRNAs, was isolated using TRIzol (Invitrogen, Carlsbad, CA, USA) according to the manufacturer’s instructions. RNA quality and quantification were analyzed by NanoDrop ND-1000 and Qubit 4.0 (Thermo Fisher). RNA integrity and genomic DNA contamination were evaluated using denaturing agarose gel electrophoresis.
miRNA sequencing
A total of 200 ng RNA were subjected to small RNA library construction using the NEB Multiplex Small RNA Library Prep Set for Illumina (New England BioLabs, Ipswich MA, USA) following the manufacturer’s instructions. Briefly, total RNA was subjected 3′ and 5′ adaptor ligation, followed by cDNA synthesis and PCR amplification. The purified library was examined by Agilent 2100 Bioanalyzer using Agilent DNA 1000 chip kit (Agilent, Santa Clara, CA, USA) for quality control and then sequenced using the Illumina NextSeq 500 platform (IIIumina, San Diego, CA, USA).
Bioinformatics analysis
Sequencing files were processed using an miRNA analysis pipeline. Initially, barcode, adaptor, and low-quality reads were removed, and the trimmed reads were aligned against the miRBase V21 to identify known or novel miRNAs through miRDeep2 (version 0.0.8). Next, mapping results were normalized by counts per million reads (CPM), and the differentially expressed miRNAs among different groups were screened using edgeR package. The cut-off value for differentially expressed miRNAs was determined as log2 (fold change) greater than 1.5 with a P value ≤ 0.05 and an intra-group mean CPM ≥ 1. The TargetScan and miRDB databases were analyzed to predict the target genes of differentially expressed miRNAs.
Quantitative real-time polymerase chain reaction (qRT-PCR)
For miRNA analysis, total RNA was reversely transcribed using the miRNA First Strand cDNA Synthesis Kit (Vazyme, Nanjing, China). qRT-PCR was performed using miRNA Universal SYBR qPCR Master Mix (Vazyme) on a LightCycler 480 (Roche, Basel, Switzerland). To assess differential gene expression at the mRNA level, total RNA was converted into cDNA using the PrimeScript™ RT reagent kit (Takara, Kyoto, Japan). Next, qRT-PCR was carried out using SYBR Master Mix (Takara) on an ABI7500 (Applied Biosystems, Foster City, CA, USA). The primer sequences are listed in Table S1. Relative gene expression was calculated using the comparative threshold (2−ΔΔCt) method.
Establishment of stable cell lines
The Kindlin-2 full-length open reading frame (ORF, Genebank: NM_006832) was cloned into the lentiviral GV358 vector (GENECHEM, Shanghai, China), which harbored a C-terminal Flag-tag. Using the packaging plasmids pHelper 1.0 and pHelper 2.0 (GENECHEM), lentivirus expressing Kindlin-2 with Flag-tag was generated and used to infect Huh7 cells. As control, lentivirus GV358 vector without the Kindlin-2 ORF was packaged and used to infect Huh7 cells as described above. Following 72 h of infection, cells stably expressing Kindlin-2 or not were selected with 2 μg/ml puromycin (Gibco). Cells stably overexpressing Kindlin-2 were designated as Huh7/LV-K2, whereas control cells were named Huh7/LV-ctrl.
Two short-hairpin RNAs (shRNAs) targeting Kindlin-2 and a negative control shRNA (Table S1) were synthesized and cloned into the lentiviral GV248 vector (GENECHEM). Packaging of lentivirus expressing shRNA, infection, and selection of HCC cells was performed as described above. Cells with stable Kindlin-2 silencing were designated as LM3/sh-K2-1 and LM3/sh-K2-2, whereas control shRNA cells were named LM3/sh-ctrl. Kindlin-2 expression in all stable cell lines was verified by western blotting (Fig. S1a).
Lentiviruses (GV654 vector, GENECHEM) containing miR-1258, miR-1258 sponge, or shRNA targeting TCF4 (Table S1) and lentiviruses (CV557 vector, GENECHEM) containing the full-length ORF of TCF4 (Genebank: NM_001146274) were constructed and used to infect LM3 or Huh7 cells as described above. Stable cells were selected with 400 µg/ml hygromycin B (Roche). Cells stably expressing miR-1258 were named as LM3/LV-1258, whereas with stable miR-1258 suppression were labeled as Huh7/LV-1258inhibitor. Cells with stable TCF4 overexpression were termed Huh7/LV-TCF4, whereas those with stable TCF4 knockdown were labeled as LM3/LV-shTCF4. The expression of miR-1258 or TCF4 among all stable clones was verified by qRT-PCR (Fig. S1b) or western blotting (Fig. S1c-S1f). These cells were used for rescue assays in vitro and in vivo.
MiR-1258 mimic, miR-1258 inhibitor, siRNA, and transient transfection
MiR-1258 mimic, miR-1258 inhibitor, and their corresponding controls were purchased from Ribobio (Guangzhou, China). Small interfering RNAs (siRNAs) targeting DNA methyltransferase 3 alpha (DNMT3A), β-catenin, TCF4, and their corresponding scrambled controls were acquired from Ribobio. The TCF4 expression plasmid pcDNA3.1-TCF4 was generated by insertion of the full-length ORF of TCF4 into the Kpn I and Xho I sites of pcDNA3.1 vector (Invitrogen). Huh7, LM3, or HEK293T cells were transfected with miR-1258 mimic, inhibitor, or plasmid using Lipofectamine 3000 (Invitrogen) according to the manufacturer’s instructions. After 48 h, the transfected cells were used for different assays.
Animal model
The Kindlin-2 knockout cell line, 97H/ko-K2, was established via clustered regularly interspaced short palindromic repeats (CRISPR)-associated protein 9 (CRISPR/Cas9) as detailed previously [3]. Lentivirus expressing luciferase (GENECHEM) was used to infect LM3/sh-K2-1, 97H/ko-K2, LM3/LV-1258, LM3/sh-K2 + 1258 inhibitor, LM3/sh-TCF4, and LM3/sh-K2 + TCF4 cells to develop cells stably expressing luciferase. Male BALB/C nude mice (4–5 weeks old) were injected with 200 μl of cell suspension (1 × 107 cell/ml) through the tail vein (n = 5 per group). Tumor metastasis was observed 9 weeks after injection via bioluminescent imaging (IVIS Lumina II, Caliper Life Sciences, Hopkinton, MA, USA), and the luminescent value of metastatic lesions in each mouse lung was measured. Next, mice were sacrificed by CO2 asphyxiation, and tumors as well as the lung tissue were dissected and subjected to histopathological analysis. The luminescent value for each group was calculated for statistical comparison. All animal experiments were conducted in accordance with the Care and Use of Laboratory Animals by the National Institutes of Health. Our experimental protocol was revised and approved by the Animal Care and Use Committee of Fujian Medical University (Approval number: FJMUIACUC2018042). All measures were undertaken to minimize animal discomfort.
Transwell migration and invasion assays
Migration and invasion assays were performed using 6.5 mm Transwell culture chambers (Millipore) with an 8 μm pore size polyester membrane insertion. For the invasion assay, the lower side of the membrane was pre-coated with Matrigel (Corning, Corning, NY, USA). Then, 200 µL cell suspension (2.5 × 105 cells/mL in serum-free DMEM) was added to each Transwell chamber. The chambers were placed in wells of 24-well plates that were filled with 750 µL DMEM with 10% serum for incubation under routine conditions. After 24 h, the media were removed, and the chambers were rinsed with phosphate-buffered saline (PBS). After removal of the cells on the upper surface of the membrane, the migrating or invading cells were fixed with 4% paraformaldehyde. Then, the membrane was stained using crystal violet, and the numbers of migrating or invading cells were counted in five randomly selected visual fields at 200 × magnification. Each experiment was performed in triplicate.
Bisulfite sequencing
Genomic DNA was extracted from LM3/sh-K2-1 cells, Huh7/LV-K2 cells, and their respective controls using the DNeasy Blood and Tissue Kit (Qiagen, Valenica, CA, USA). Genomic DNA was modified with sodium bisulfite using EpiTect Bisulfite Kit (Qiagen) per the manufacturer’s instructions. The miR-1258 promoter contains 21 CpG islands on chromosome 2. The CpG islands were amplified with Taq DNA Polymerase (Takara) and primers designed by MethPrimer 2.0 (Table S1). PCR products were purified using a DNA Purification Kit (TINGEN, Beijing, China) and cloned into pMD18-T vector (Takara). Then, the recombinant was transformed into E. coli DH5α competent cells, and IPTG (TINGEN) and X-gal (TINGEN) were used for blue/white screening. A total of 8 clones were chosen randomly for sequencing, and the percentage of methylated CpG dinucleotides was calculated.
Luciferase reporter assay
Segments of wild-type or mutant 3′ untranslated region (3′-UTR) of TCF4 were cloned into a pSI-Check2 luciferase reporter plasmid (Promega, WI, USA). Next, wild-type or mutant TCF4 3′-UTR and miR-1258 mimic or its control were co-transfected into HEK293T cells. After 48 h, Renilla and firefly luciferase signals were detected using the Dual-Luciferase Reporter Assay system (Promega) following the instructions of the manufacturer.
Analysis of promoter activity
Possible binding sites between the Kindlin-2 promoter and TCF4 were predicted by Jaspar 2020. Segments of the Kindlin-2 wild-type promoter (from the transcription start site to 2000 bp upstream) or the mutant promoter were cloned into a pGL3-basic luciferase reporter vector. Next, the Kindlin-2 wild-type promoter or mutant promoter plasmids, pcDNA3.1-TCF4 plasmids, and the pRL-TK Renilla luciferase-expressing vector (Promega) were co-transfected into HEK293T cells. The pRL-TK Renilla luciferase-expressing vector was used as an internal control for normalizing the transfection efficiency. After 48 h, the Renilla and firefly luciferase activities were detected using the Dual-Luciferase Reporter Assay system (Promega) following the manufacturer’s protocol.
Chromatin immunoprecipitation (ChIP) assay
Nuclear fractions were extracted using the Nuclear Extraction Kit (Abcam) following the manufacturer’s instructions. Immunoprecipitation of the nuclear extract was performed using the SimpleChIP Plus Enzymatic Chromatin IP Kit (Cell Signaling Technology). Immunoprecipitation was performed using antibodies specific to TCF4 (2569, Cell Signaling Technology; 1:50), and nonspecific IgG (Cell Signaling Technology). Next, the immunoprecipitated promoter region of Kindlin-2 was amplified by PCR and visualized by agarose gel electrophoresis. The primer sequences used are listed in Table S1.
Statistical analysis
Data for continuous variables with a normal distribution are presented as mean ± standard deviation (SD), and categorical data were presented as proportions. Differences in the means between groups were analyzed by Student’s t test, while the associations of miRNA levels and relative gene expression were performed using Spearman’s rank correlation. All statistical analyses were conducted using SPSS version 19.0 (SPSS, Inc., Armonk, NY, USA), and a P value < 0.05 was indicative of statistical significance.
Results
Downregulation of Kindlin-2 inhibited HCC metastasis in vivo
To evaluate the role of Kindlin-2 in HCC metastasis in vivo, we injected nude mice with either 97H/ko-K2-luc or LM3/sh-K2-1-luc cells in which luciferase was stably expressed and Kindlin-2 was downregulated using CRISPR/Cas9 and shRNA technology, respectively, or their respective controls (97H/wt-luc and LM3/sh-ctrl-luc). Tumor metastasis was monitored using bioluminescent imaging. Among the 5 mice of 97H control group, 4 mice showed tumor metastases (80%; 4/5) 9 weeks after injection, whereas no mice in the 97H/ko-K2-luc group had tumor metastases (0%; 0/5). In the LM3/sh-K2 group, metastases occurred in 2 mice (40%; 2/5), compared with 4 mice (80%; 4/5) in the control group. The luminescent value of metastatic lesions was much lower in the Kindlin-2 knockout or knockdown group than in the control group (Fig. 1a, b; P < 0.01 and P < 0.05). Metastatic foci were observed in the lungs of mice of the 97H/wt-luc group (Fig. 1c) and the LM3/sh-K2-luc as well as LM3/sh-ctrl-luc groups (Fig. 1d). Hematoxylin and eosin (H&E) staining provided histopathological evidence of tumor metastasis in the lungs (Fig. 1e, f). These results suggest that the downregulation of Kindlin-2 inhibited HCC metastasis in vivo.
Downregulation of Kindlin-2 inhibited HCC metastasis in vivo. Effect of Kindlin-2 knock out in 97H cells or knock down in LM3 cells on tumor metastasis following injection into the tail vein of nude mice. Bioluminescence imaging showing that Kindlin-2 knock out (97H/ko-K2) (a) or knock down (LM3/sh-K2) (b) significantly inhibited the metastatic capacity of HCC cells. c, d Representative images of the gross lung morphology demonstrating the macroscopic metastatic foci. Yellow circles indicate metastatic nodules. e, f Representative H and E staining images of metastatic nodules in the lung. Data are mean ± standard deviation (SD); **P < 0.01, *P < 0.05
Kindlin-2 epigenetically suppresses miR-1258 expression
We used high-throughput sequencing to compare the miRNA expression profiles between Kindlin-2 knockdown and control cells. MiR-1258 was significantly upregulated in LM3/sh-K2 cells (Fig. 2a; P < 0.001). Furthermore, the expression of miR-1258 increased following Kindlin-2 knockdown and decreased after Kindlin-2 overexpression (Fig. 2b; P < 0.01). Next, we detected miR-1258 expression levels in different HCC cell lines. Compared to that in the highly metastatic cell lines (97H and LM3 cells), miR-1258 expression was elevated in the less metastatic cell lines including Huh7, HepG2, HepG2.2.15, and Hep3B cells, which showed the opposite tendency compared with Kindlin-2 expression (Fig. 2c).
Kindlin-2 epigenetically suppresses miR-1258 expression. a Heatmap of differential miRNA expression profiles between LM3 cells with Kindlin-2 knockdown and control cells obtained by high-throughput sequencing. Green color indicates lower than mean intensity, while red represents higher than mean intensity. MiR-1258 was the top upregulated contender. b Relative miR-1258 expression by qRT-PCR in LM3 and Huh7 cells infected with sh-K2-1 and shK2-2 as well as LV-K2, respectively. U6 was used as an internal control. c Relative Kindlin-2 and miR-1258 expression by qRT-PCR among different cell lines. d Relative miR-1258 expression by qRT-PCR in LM3 and Huh7 cells infected with sh-K2-1 and sh-K2-2 as well as LV-K2, respectively, and treated with 5-aza-CdR (inhibitor of DNMT). e Schematic diagram of miR-1258 promoter. The miR-1258 promoter contains 21 CpG islands on chromosome 2. f Effect of Kindlin-2 on the methylation of the miR-1258 promoter. Bisulfite sequencing analysis revealed that Kindlin-2 improved the methylation rate of CpG islands in the miR-1258 promoter. Data are mean ± SD; ***P < 0.001, **P < 0.01, and *P < 0.05
DNA methyltransferase (DNMT) plays important roles in DNA methylation, and Kindlin-2 could interact with DNMT3A [6] or DNMT1 [14]. Therefore, we detected the effect of Kindlin-2 on DNMT expression. The levels of DNMT1, DNMT3A, and DNMT3B protein were not altered by ectopic expression or knockdown of Kindlin-2 (Fig. S2). Then, we investigated the regulation of miR-1258 expression via DNMT. In this experiment, Huh7 and LM3 cells were treated with 5-aza-2’-deoxycytidine (5-aza-CdR, DNMT inhibitor). Compared with the miR-1258 expression level in the LV-K2 group (Huh7 cells overexpressing Kindlin-2), 5-aza-CdR treatment increased miR-1258 expression (Fig. 2d; P < 0.05). Elevated miR-1258 expression was also observed in both Kindlin-2 shRNA-treated and 5-aza-CdR-treated LM3 cells (Fig. 2d; P < 0.01).
To verify that Kindlin-2 can epigenetically regulate miR-1258 expression, miR-1258 methylation was analyzed by bisulfite sequencing in Kindlin-2–overexpressing or knockdown cells (Fig. 2e). Kindlin-2 ectopic expression improved the methylation rate of CpG islands in the miR-1258 promoter (Fig. 2f). On the contrary, Kindlin-2 knockdown resulted in partial demethylation of miR-1258 promoter (Fig. 2f). Collectively, these findings indicate that Kindlin-2 has a role in the epigenetic suppression of miR-1258 in HCC.
Kindlin-2 regulated TCF4 expression by directly targeting miR-1258
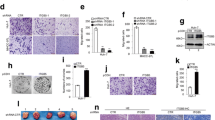
Next, we aimed to explore the function of miR-1258 in HCC. Target genes of miR-1258 were predicted by analysis of the miRDB and TargetScan databases. Among all the potential target genes, TCF4 (a key member of the Wnt/β-catenin signaling pathway) drew our attention, because our previous research showed that Kindlin-2 promotes HCC invasion and metastasis through Wnt/β-catenin signaling. We speculated that miR-1258 might function by regulating TCF4 and determined possible binding sites between miR-1258 and the 3′-UTR of TCF4 (Fig. 3a). Next, western blotting confirmed the downregulation of TCF4 expression in Huh7 cells with increased miR-1258 expression. Interestingly, the addition of a miR-1258 inhibitor resulted in the upregulation of TCF4 expression (Fig. 3b). A similar trend was observed in the LM3 cells (Fig. 3b). A luciferase reporter assay plasmid containing the wild-type TCF4 3’-UTR was constructed (Fig. 3c). We introduced four point mutations to create the mutant TCF4 3′-UTR (Fig. 3c). In comparison to cells co-transfected with the mutant TCF4 3′-UTR and miR-1258 mimic, cells co-transfected with miR-1258 mimic and wild-type TCF4 3′-UTR had significantly reduced luciferase activity (Fig. 3d; P < 0.001). In agreement, these results reveal that miR-1258 directly targets TCF4 in HCC.
Kindlin-2 regulated TCF4 expression by directly targeting miR-1258. a Schematic diagram predicting possible binding sites for miR-1258 in the 3′-UTR of TCF4. b Western blotting analyzing the effect of miR-1258 on TCF4 expression in HCC cells. c Sequence alignment of wild-type and mutant TCF4 3′-UTR with miR-1258. d Relative luciferase activity of HEK293T cells co-transfected with the wild-type or mutant TCF4 3′-UTR reporter genes and miR-1258 mimic or control. e–h Western blotting analysis of TCF4 expression in HCC cells. e Western blotting detecting the impact of Kindlin-2 knockdown (LM3 cells) or overexpression (Huh7 cells) on TCF4 expression. f Western blotting demonstrating the impact of miR-1258 inhibitor or miR-1258 mimic on TCF4 expression in cells with Kindlin-2 knockdown (LM3 cells) or overexpression (Huh7 cells). g Western blotting demonstrating the impact of 5-aza-cdR treatment and transient transfection of shRNA targeting DNMT3A on TCF4 expression in Huh7 cells overexpressing Kindlin-2. h Western blotting demonstrating the impact of transient transfection of shRNA targeting β-catenin on TCF4 expression in Huh7 cells overexpressing Kindlin-2. wt, wild type, mut, mutant. Data are mean ± SD; ***P < 0.001, ns: P > 0.05
To explore the role of Kindlin-2 in regulating TCF4, we examined the difference in TCF4 expression in HCC cells with Kindlin-2 knockdown or ectopic expression. Compared with the expression levels in the control groups, Kindlin-2 shRNA downregulated TCF4 expression in LM3 cells and Kindlin-2 overexpression upregulated TCF4 levels in Huh7 cells (Fig. 3e). Transfection with a miR-1258 inhibitor partially restored the TCF4 expression in LM3 cells, but transfection with the miR-1258 mimic downregulated TCF4 expression in Huh7 cells (Fig. 3f). Similarly, both 5-aza-cdR treatment and transient transfection with DNMT3A siRNA in Huh7 cells overexpressing Kindlin-2 downregulated TCF4 (Fig. 3g). Kindlin-2 can activate Wnt/β-catenin signaling and upregulate β-catenin expression. Thus, Kindlin-2 can form a tripartite complex with β-catenin and TCF4. To investigate whether the effect of upregulation of Kindlin-2 on TCF4 relied on β-catenin, we analyzed the impact of β-catenin siRNA on TCF4 expression. Our results clearly show that β-catenin knockdown did not affect TCF4 expression (Fig. 3h). Taken together, these experiments show that Kindlin-2 can upregulate TCF4 expression by inhibiting miR-1258 in HCC independent of β-catenin signaling.
Kindlin-2–miR-1258–TCF4 form a feedback loop
Using the Jaspar algorithm, we predicted four potential binding sites in the Kindlin-2 promoter for TCF4, two sites in the sense chain and two in the anti-sense chain (Fig. 4a). Next, western blotting and qRT-PCR confirmed significant upregulation of Kindin-2 in cells with ectopic TCF4 expression (Fig. 4b, c; P < 0.001). Conversely, Kindin-2 expression was markedly downregulated with TCF4 siRNA (Fig. 4d, e; P < 0.001). We constructed two plasmids containing a mutant promoter in the sense and anti-sense chains, respectively. The wild-type or mutant promoter plasmids, TCF4 plasmids, and the Renilla luciferase-expressing vector were co-transfected into HEK293T cells (Fig. 4f). Luciferase activity increased upon the co-transfection of TCF4 with the wild-type Kindlin-2 promoter compared to the mutant1 + 4 (Fig. 4g; P < 0.001). However, luciferase activity was similar between cells transfected with the wild-type Kindlin-2 promoter and the mutant2 + 3 (Fig. 4g). ChIP assay further verified the direct binding of TCF4 with the promoter in the sense chain of Kindlin-2 (Fig. 4h and Fig. S3). These results indicate that TCF4 functions as a transcription factor that enhances Kindlin-2 expression and the TCF4–Kindlin-2 binding sites are those located in its sense chain.
Kindlin-2–miR-1258–TCF4 interaction forms a feedback loop. a Schematic diagram of the predicted potential binding sites of Kindlin-2 promoter with TCF4. Sites 1 and 4 are in the sense chain, and sites 2 and 3 are in the anti-sense chain. The effect of TCF4 on Kindin-2 expression was analyzed by western blotting (b) and qRT-PCR (c) in Huh7, LM3, and HEK293T cells. The effect of siRNA targeting of TCF4 on Kindin-2 expression was analyzed by western blotting (d) and qRT-PCR (e) in Huh7 and HEK293T cells. f Sequence alignment of wild-type and mutant Kindlin-2 promoter with TCF4. Mut1 + 4 contained mutant sites 1 and 4 in the sense chain. Mut2 + 3 contained mutant sites 2 and 3 in the anti-sense chain. g Relative luciferase activity of HEK293T cells co-transfected with the wild-type or mutant Kindlin-2 promoter reporter genes and TCF4 expression plasmids or control. h ChIP assay confirming the direct binding of TCF4 with the promoter of Kindlin-2 in the sense chain. i Western blotting of LM3 cells transfected with miR-1258 mimic or inhibitor along with TCF4 siRNA or pcDNA-TCF4, indicating that miR-1258 negatively regulated Kindlin-2 expression through TCF4. wt, wild type, mut mutant. Data are mean ± SD; ***P < 0.001
Furthermore, we transfected HCC cells with miR-1258 mimic or inhibitor along with TCF4 siRNA or pcDNA3-TCF4 to further examine the relationship between TCF4 and Kindlin-2. Transfection with miR-1258 mimic downregulated Kindlin-2 and TCF4 expression, but TCF4 ectopic expression restored Kindlin-2 expression (Fig. 4i). On the contrary, transfection with miR-1258 inhibitor induced Kindlin-2 expression (Fig. 4i). These results suggest that miR-1258 can negatively regulate Kindlin-2 expression through TCF4 in HCC.
MiR-1258 and TCF4 are involved in Kindlin-2-regulated HCC invasion and metastasis
To further prove that Kindlin-2 elicits its role in HCC via miR-1258, we performed Transwell migration and invasion assays in vitro. Upregulation of miR-1258 inhibited cell migration and invasion through Matrigel (Fig. 5a; both P < 0.001), whereas miR-1258 downregulation significantly improved the migration and invasion capacities of the cells (Fig. 5a; both P < 0.001). Conversely, the migration and invasion capacities were apparently enhanced after transfection of LM3 cells with Kindlin-2 knockdown with miR-1258 inhibitor (Fig. 5b; both P < 0.001). For the in vivo metastasis assay, the stable LM3/miR-1258-luc cells were injected into the tail vein of nude mice (n = 5 for each group). In comparison with that in the control group, a lower luminescent value for tumor metastasis was detected upon miR-1258 overexpression (Fig. 5c; P < 0.05). Next, mice were injected with LM3/sh-K2 + 1258inhibitor cells. The metastatic capacity of LM3/sh-K2 cells was strengthened by miR-1258 inhibitor (Fig. 5d; P < 0.05). Therefore, miR-1258 attenuated the metastasis-promoting effect of Kindlin-2.
MiR-1258 is involved in Kindlin-2-regulated HCC invasion and metastasis. a MiR-1258 abrogated Kindlin-2-induced Huh7 cell migration and invasion. b MiR-1258 inhibitor restored Kindlin-2 knockdown-mediated inhibition of LM3 migration and invasion. c Bioluminescence imaging of tumor metastasis in nude mice. MiR-1258 inhibited the metastasis capacity of HCC cells. d Bioluminescence imaging of tumor metastasis in nude mice. The metastasis capacity of LM3/sh-K2 cells was strengthened by miR-1258 inhibitor. Data are mean ± SD; ***P < 0.001, *P < 0.05
Furthermore, we confirmed the involvement of TCF4 in Kindlin-2-induced HCC progression in vitro and in vivo. Ectopic expression of TCF4 enhanced the migration and invasion capacities of the cells (Fig. 6a; P < 0.01 and P < 0.001), whereas TCF4 knockdown significantly weakened the migration, invasion, and metastasis capacities of HCC cells (Fig. 6b and c; P < 0.001 and P < 0.01). Also, the inhibition of migration, invasion, and metastasis of LM3 cells with Kindlin-2 knockdown could be restored by TCF4 overexpression (Fig. 6b and d; P < 0.001 and P < 0.05).
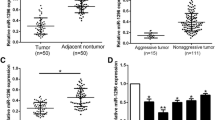
TCF4 is involved in Kindlin-2-regulated HCC invasion and metastasis. a TCF4 downregulation abrogated Kindlin-2-induced Huh7 cell migration and invasion. b TCF4 ectopic expression restored Kindlin-2 knockdown-mediated inhibition of LM3 migration and invasion. c Bioluminescence imaging of tumor metastasis in nude mice. TCF4 enhanced the metastasis capacity of HCC cells. d Bioluminescence imaging of tumor metastasis in nude mice. The metastasis capacity of LM3/sh-K2 cells was strengthened by TCF4. e Pearson’s correlation analysis demonstrating the relationship between Kindlin-2 and miR-1258. Kindlin-2 expression was negatively correlated with miR-1258 expression. f Pearson’s correlation analysis demonstrating the relationship between Kindlin-2 and TCF4. Kindlin-2 expression was positively correlated with TCF4 expression. g Pearson’s correlation analysis demonstrating the relationship between miR-1258 and TCF4. MiR-1258 expression was negatively correlated with TCF4 expression. Data are mean ± SD; ***P < 0.001, **P < 0.01, and *P < 0.05
We also analyzed miR-1258, TCF4, and Kindlin-2 expression in HCC clinical samples by qRT-PCR. Pearson’s correlation analysis revealed a negative correlation between Kindlin-2 and miR-1258 expression (Fig. 6e; P < 0.01) as well as a positive correlation between Kindlin-2 and TCF4 expression (Fig. 6f; P < 0.001). Meanwhile, miR-1258 expression was negatively correlated with TCF4 expression (Fig. 6g; P < 0.01). Collectively, these results illustrate the importance of miR-1258 for Kindlin-2’s promotion of HCC invasion and metastasis.
Discussion
Advanced-stage HCC is associated with high risks of recurrence and metastasis and is usually associated with poor patient prognosis [15]. Kindlin-2 is a ubiquitously expressed cytoskeletal protein that can promote HCC progression and metastasis [3]. In this study, we focused on elucidating the molecular mechanisms of Kindlin-2 signaling in HCC to provide insight for the development of new therapeutic strategies. In this study, we further verified the impact of Kindlin-2 on HCC metastasis in vivo. Moreover, we demonstrated that Kindlin-2 formed a positive feedback loop with miR-1258 and TCF4 to regulate HCC invasion and metastasis.
The Kindlin family consists of three members, Kindlin-1, -2, and -3 [1]. Kindlin-2 plays important roles in embryonic and organ development as well as in tumor progression [2]. In HCC patients, high Kindlin-2 expression is correlated with aggressive clinicopathological features, including tumor encapsulation, microvascular invasion, extrahepatic metastasis, and poor prognosis. Functional experiments showed that Kindlin-2 enhances the invasion and motility of HCC cells in vitro [3, 16]. In the present study, we explored the effect of Kindlin-2 on HCC metastasis in vivo. In agreement with previous reports, our results demonstrate that Kindlin-2 downregulation significantly inhibited HCC tumor metastasis in nude mice.
Together with talin, the Kindlin family of proteins can activate integrins [17]. And Kindlin-2 can elicit its function through an integrin-dependent signaling pathway. In addition to integrin-dependent signaling, Kindlin-2 functions via integrin-independent pathways [18]. Kindlin-2 was reported to promote breast cancer invasion by forming a complex with DNMT3A, which leads to the epigenetic suppression of miRNA200 [6]. In the same context, our results demonstrated that miR-1258 was significantly upregulated in Kindlin-2 knockdown HCC cells. MiR-1258 was previously reported to inhibit the invasion and migration of HCC [19, 20]. Indeed, we observed lower miR-1258 expression in highly metastatic HCC cell lines. In addition, miR-1258 reduced the migration and invasion capacities of HCC cells in vitro as well as tumor metastasis in vivo. Meanwhile, miR-1258 expression was negatively correlated with Kindlin-2 expression in HCC cell lines as well as tissue specimens from HCC patients. Previous studies showed that miR-1258 is regulated by circular RNAs or long non-coding RNAs [21,22,23,24]. Interestingly, methylation of miR-1258 was recently reported in myeloma, breast, and ovarian cancers [25, 26]. Nevertheless, there is no solid evidence suggesting the epigenetic repression of miR-1258. We postulated that Kindlin-2 might inhibit miR-1258 expression by inducing promoter hypermethylation. Upon treatment with a pharmacological inhibitor of DNMT (—5-aza-CdR), we observed an increase in miR-1258 expression in LM3 and Huh7 cells (with Kindlin-2 overexpression). However, ectopic expression or knockdown of Kindlin-2 did not affect the DNMT expression levels in HCC cells, unlike the finding by Yu et al. that ectopic expression of Kindlin-2 could increase DNMT levels in breast cancer cells. The reason for this inconsistency may be differences in the biological properties of the different type tumor cell lines. The possibility of Kindlin-2 to form a complex with DNMT3A might enrich DNMT3A to the promoter of genes to influence gene expression [6, 27]. Furthermore, bisulfite sequencing confirmed that Kindlin-2 improved the methylation rate of CpG islands in the miR-1258 promoter. Taken together, these results suggest that Kindlin-2 epigenetically suppresses miR-1258 expression in HCC.
MiRNAs can induce degradation of target mRNAs through binding with the 3′-UTRs or the coding sequence (CDS), thereby downregulating the expression of target genes [28]. Via analysis of the miRDB and TargetScan databases, TCF4 was predicted as a target gene of miR-1258. To the best of our knowledge, no evidence has been reported that suggests an interaction between Kindlin-2 and TCF4. We performed a dual-luciferase reporter assay to verify the binding of miR-1258 to the 3′-UTR of TCF4. Our results showed that TCF4 expression was decreased in miR-1258-overexpressing cells and increased in miR-1258-silenced cells. On the other hand, Kindlin-2 downregulation decreased TCF4 expression, and its overexpression increased TCF4 expression. Using a miR-1258 inhibitor, we were able to restore the TCF4 expression that was initially downregulated following Kindlin-2 knockdown. Conversely, miR-1258 mimic, a pharmacological inhibitor of DNMT or silencing of DNMT3A were all able to downregulate TCF4 expression that was initially upregulated following Kindlin-2 overexpression. Based on these collective results, we hypothesize that Kindlin-2 regulates TCF4 via miR-1258. Previous research showed that Kindlin-2 can increase β-catenin and activate canonical Wnt signaling [3]. The activation of canonical Wnt signaling depends on β-catenin and TCF through the formation of a transcriptional complex [29]. Kindlin-2 can participate in that transcriptional complex, thereby triggering downstream gene transcription with β-catenin and TCF4 collectively [30]. Furthermore, our functional assays and animal experiments suggested that miR-1258 and TCF4 are critical for Kindlin-2 function in enhancing HCC cell migration in vitro and HCC metastasis in vivo. Moreover, Kindlin-2 expression was negatively correlated with miR-1258 expression, but positively correlated with TCF4 expression in HCC patient tissues. Therefore, it is plausible to speculate that Kindlin-2 promotes HCC progression by regulating the miR-1258–TCF4 signaling axis.
TCF4 is an essential transcription factor that can promote gene transcription [31]. Using the Jaspar algorithm, we predicted possible binding sites in the Kindlin-2 promotor for TCF4. We performed a dual-luciferase reporter assay and ChIP to confirm that TCF4 can act as a transcription factor to enhance Kindlin-2 expression. At the same time, miR-1258 inhibited Kindlin-2 expression in HCC cells, and this inhibition was abolished by TCF4. Therefore, Kindlin-2–miR-1258–TCF4 signaling can actually form a feedback loop in HCC. Specifically, Kindlin-2 can upregulate TCF4 via epigenetically suppressing miR-1258. In turn, TCF4 can upregulate Kindlin-2 via promoting transcription (Fig. 7). This newly uncovered mechanism further enhances our knowledge of HCC pathogeneses.
Schematic illustration demonstrating the Kindlin-2–miR-1258–TCF4 positive feedback loop in HCC metastasis. Kindlin-2 can upregulate TCF4 via epigenetically suppressing miR-1258. Our previous research has demonstrated that Kindlin-2 can increase β-catenin expression and activate canonical Wnt signaling [3]. The formation of a transcriptional complex composed of TCF4 and β-catenin is critical for canonical Wnt signaling activation. Therefore, Kindlin-2 is closely connected with Wnt signaling through both TCF4 and β-catenin. In turn, TCF4 can upregulate Kindlin-2 via promoting its transcription
Conclusions
Kindlin-2 promotes HCC progression through a miR-1258–TCF4 axis. Kindlin-2–miR-1258–TCF-4 signaling forms a positive feedback loop to amplify the function of Kindlin-2 in HCC. Accordingly, Kindlin-2 represents a promising therapeutic target for blocking HCC invasion and metastasis.
Abbreviations
- HCC:
-
Hepatocellular carcinoma
- EMT:
-
Epithelial–mesenchymal transition
- miR:
-
MicroRNA
- SDS-PAGE:
-
Sodium dodecyl sulfate–polyacrylamide gel electrophoresis
- shRNA:
-
Short hairpin RNAs
- CPM:
-
Counts per million reads
- qRT-PCR:
-
Quantitative real-time PCR
- CRISPR:
-
Clustered regularly interspaced short palindromic repeats
- Cas 9:
-
CRISPR-associated protein 9
- ChIP:
-
Chromatin immunoprecipitation
- DNMT:
-
DNA methyltransferase
- 5-aza-CdR:
-
5-Aza-2’-deoxycytidine
- TCF4:
-
Transcription factor 7 like 2
- FERM:
-
4.1-Ezrin-ridixin-moesin
- 3′-UTR:
-
3′-Untranslated region
- CDS:
-
Coding sequence
References
Li H, Deng Y, Sun K, et al. Structural basis of kindlin-mediated integrin recognition and activation. Proc Natl Acad Sci USA. 2017;114:9349–54.
Wang W, Kansakar U, Markovic V, et al. Role of Kindlin-2 in cancer progression and metastasis. Ann Transl Med. 2020;8:901.
Lin J, Lin W, Ye Y, et al. Kindlin-2 promotes hepatocellular carcinoma invasion and metastasis by increasing Wnt/β-catenin signaling. J Exp Clin Cancer Res. 2017;36:134.
Zhang HF, Alshareef A, Wu C, et al. Loss of miR-200b promotes invasion via activating the Kindlin-2/integrin β1/AKT pathway in esophageal squamous cell carcinoma: an E-cadherin-independent mechanism. Oncotarget. 2015;6:28949–60.
Shen Z, Ye Y, Kauttu T, et al. Novel focal adhesion protein kindlin-2 promotes the invasion of gastric cancer cells through phosphorylation of integrin β1 and β3. J Surg Oncol. 2013;108:106–12.
Yu Y, Wu J, Guan L, et al. Kindlin 2 promotes breast cancer invasion via epigenetic silencing of the microRNA200 gene family. Int J Cancer. 2013;133:1368–79.
Ren W, Gao L, Qiang C, et al. Kindlin-2-mediated upregulation of ZEB2 facilitates migration and invasion of oral squamous cell carcinoma in a miR-200b-dependent manner. Am J Transl Res. 2018;10:2529–41.
Zhang HF, Zhang K, Liao LD, et al. miR-200b suppresses invasiveness and modulates the cytoskeletal and adhesive machinery in esophageal squamous cell carcinoma cells via targeting Kindlin-2. Carcinogenesis. 2014;35:292–301.
Sossey-Alaoui K, Pluskota E, Szpak D, et al. The Kindlin-2 regulation of epithelial-to-mesenchymal transition in breast cancer metastasis is mediated through miR-200b. Sci Rep. 2018;8:7360.
Sossey-Alaoui K, Pluskota E, Bialkowska K, et al. Kindlin-2 regulates the growth of breast cancer tumors by activating CSF-1-mediated macrophage infiltration. Cancer Res. 2017;77:5129–41.
Wang Z, Yang Y, Cui Y, et al. Tumor-associated macrophages regulate gastric cancer cell invasion and metastasis through TGFβ2/NF-κB/Kindlin-2 axis. Chin J Cancer Res. 2020;32:72–88.
Guo B, Gao J, Zhan J, et al. Kindlin-2 interacts with and stabilizes EGFR and is required for EGF-induced breast cancer cell migration. Cancer Lett. 2015;361:271–81.
Gao J, Khan AA, Shimokawa T, et al. A feedback regulation between Kindlin-2 and GLI1 in prostate cancer cells. FEBS Lett. 2013;587:631–8.
Peng W, Wen HC, Xi Z, et al. Kindlin-2 interacts with and stabilizes DNMT1 to promote breast cancer development. Int J Biochem Cell Biol. 2018;105:41–51.
Llovet JM, Kelley RK, Villanueva A, et al. Hepatocellular carcinoma. Nat Rev Dis Primers. 2021;7:6.
Ge YS, Liu D, Jia WD, et al. Kindlin-2: a novel prognostic biomarker for patients with hepatocellular carcinoma. Pathol Res Pract. 2015;211:198–202.
Sun Z, Costell M, Fässler R. Integrin activation by talin, kindlin and mechanical forces. Nat Cell Biol. 2019;21:25–31.
Zhan J, Zhang H. Kindlins: roles in development and cancer progression. Int J Biochem Cell Biol. 2018;98:93–103.
Hu M, Wang M, Lu H, et al. Loss of miR-1258 contributes to carcinogenesis and progression of liver cancer through targeting CDC28 protein kinase regulatory subunit 1B. Oncotarget. 2016;7:43419–31.
Huang WJ, Tian XP, Bi SX, et al. The β-catenin/TCF-4-LINC01278-miR-1258-Smad2/3 axis promotes hepatocellular carcinoma metastasis. Oncogene. 2020;39:4538–50.
Zou H, Xu X, Luo L, et al. Hsa_circ_0101432 promotes the development of hepatocellular carcinoma (HCC) by adsorbing miR-1258 and miR-622. Cell Cycle. 2019;18:2398–413.
Fang Q, Liu H, Zhou A, et al. Circ_0046599 promotes the development of hepatocellular carcinoma by regulating the miR-1258/RPN2 network. Cancer Manag Res. 2020;12:6849–60.
Li W, Yang X, Shi C, et al. Hsa_circ_002178 promotes the growth and migration of breast cancer cells and maintains cancer stem-like cell properties through regulating miR-1258/KDM7A axis. Cell Transplant. 2020;29:963689720960174.
Zhang W, Wu G, Sun P, et al. circ_SMAD2 regulate colorectal cancer cells proliferation through targeting miR-1258/RPN2 signaling pathway. J Cancer. 2021;12:1678–86.
Wang LQ, Kumar S, Calin GA, et al. Frequent methylation of the tumour suppressor miR-1258 targeting PDL1: implication in multiple myeloma-specific cytotoxicity and prognostification. Br J Haematol. 2020;190:249–61.
Loginov VI, Burdennyy AM, Filippova EA, et al. Hypermethylation of miR-107, miR-130b, miR-203a, miR-1258 genes associated with ovarian cancer development and metastasis. Mol Biol. 2018;52:801–9.
Zheng DL, Zhang L, Cheng N, et al. Epigenetic modification induced by hepatitis B virus X protein via interaction with de novo DNA methyltransferase DNMT3A. J Hepatol. 2009;50:377–87.
Treiber T, Treiber N, Meister G. Regulation of microRNA biogenesis and its crosstalk with other cellular pathways. Nat Rev Mol Cell Biol. 2019;20:5–20.
Caspi M, Wittenstein A, Kazelnik M, et al. Therapeutic targeting of the oncogenic Wnt signaling pathway for treating colorectal cancer and other colonic disorders. Adv Drug Deliv Rev. 2021;169:118–36.
Yu Y, Wu J, Wang Y, et al. Kindlin 2 forms a transcriptional complex with β-catenin and TCF4 to enhance Wnt signalling. EMBO Rep. 2012;13:750–8.
Wedel M, Fröb F, Elsesser O, et al. Transcription factor Tcf4 is the preferred heterodimerization partner for Olig2 in oligodendrocytes and required for differentiation. Nucleic Acids Res. 2020;48:4839–57.
Acknowledgements
This work was supported by grants from the National Natural Science Foundation of China (Grant number: 81802882), Joint Funds for the innovation of Science and Technology, Fujian province (Grant number: 2018Y9104, 2018Y9108), and the Natural Science Foundation of Fujian Province (Grant number: 2018J01251).
Funding
National Natural Science Foundation of China, 81802882, Jie Lin, Joint Funds for the innovation of Science and Technology, Fujian province, 2018Y9104, Wansong Lin, 2018Y9108, Yunbin Ye, Natural Science Foundation of Fujian Province, 2018J01251, Jie Lin.
Author information
Authors and Affiliations
Contributions
LWS and LJ conceived and designed the study as well as performed the laboratory analysis. LY and WYJ performed the animal experiments and the ChIP analysis. LJY and CSP performed the western blotting analysis. CLF performed the qRT-PCR analysis in clinical samples. CH and LL collected the samples. LJ, LWS, CXY, and YYB contributed reagents, materials, and analysis tools. LWS and LJ wrote the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.



Rights and permissions
About this article
Cite this article
Lin, W., Lin, J., Li, J. et al. Kindlin-2–miR-1258–TCF4 feedback loop promotes hepatocellular carcinoma invasion and metastasis. J Gastroenterol 57, 372–386 (2022). https://doi.org/10.1007/s00535-022-01866-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00535-022-01866-8