Abstract
The neural stem cells of Drosophila, called neuroblasts, have the ability to self-renew and at the same time produce many different types of neurons and glial cells. In the central brain and ventral ganglia, neuroblasts are specified and delaminate from the neuroectoderm during embryonic development under the control of proneural and neurogenic genes. In contrast, in the optic lobes, neuroepithelial cells are transformed into neuroblasts postembryonically by a spatial wave of proneural gene expression. Central brain and ventral nerve cord neuroblasts manifest a short embryonic proliferation period followed by a stage of quiescence and then undergo a prolonged postembryonic proliferation period during which most of the differentiated neurons of the adult CNS are generated. While most neuroblasts belong to a type I class that produces neuronal lineages through non-self-renewing ganglion mother cells, a small subset of type II neuroblasts generates exceptionally large neuronal lineages through self-renewing intermediate progenitor cells that have a transit amplifying function. All neuroblasts in the CNS generate their neural progeny through an asymmetric cell division mode in which the interplay of apical complex and basal complex molecules in the mitotically active progenitor results in the segregation of cell fate determinants into the smaller more differentiated daughter cell. Defects in this molecular control of asymmetric cell division in neuroblasts can result in brain tumor formation. Proliferating neuroblast lineages in the developing CNS utilize transcription factor cascades as a generic mechanism for temporal patterning and birth order-dependent determination of differential neural cell fate. This contributes to the generation of a remarkable diversity of cell types in the developing CNS from a surprisingly small set of neural stem cell-like precursors.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In humans, as in all other higher animals, the central nervous system manifests the highest level of structural and functional complexity of any organ system. The huge diversity of neural cell types that characterize the complex circuits of the nervous system is produced by neural stem cells. During normal development, neural stem cells produce defined sets of neural progeny composed of specific cell types that interconnect to form functional circuitry. Understanding the molecular mechanisms that underlie this process, and give rise to the astonishing number and diversity of precisely defined cell types in the nervous system, is one of the most challenging problems in biology. In recent years, important contributions to the understanding of the molecular mechanisms involved in neural stem cell biology have been made in several vertebrate and invertebrate neurogenetic model systems, including the fruit fly Drosophila (Homem and Knoblich 2012).
In Drosophila, the neural stem cells, called neuroblasts, are similar to vertebrate neural stem cells in their ability to self-renew and to produce many different types of neurons and glial cells. The Drosophila central nervous system (CNS), which can be divided into the central brain and optic lobe in the head and the ventral nerve cord (VNC) in the trunk region, consists of thousands of diverse neuronal cells, which are arranged in complicated neural circuits. All of these neuronal cells are generated by a remarkably restricted set of neuroblasts through precisely controlled proliferation and differentiation processes during development. In the last decade, significant progress has been made in understanding the generic developmental mechanisms that operate in these neuroblasts during their normal proliferation. Moreover, some insight into the molecular events by which deregulated neuroblast proliferation can lead to the formation of brain tumors has also been obtained.
In this review, we consider some of the recent insights into the mechanisms by which these neuroblasts give rise to diverse neural lineages in CNS development. We first describe the generic series of events that result in the formation, proliferation and termination of neuroblasts in the CNS. We then examine the diversity of neuroblast types with a special focus on the role of transit amplifying neuroblast lineages in brain development. Subsequently, we describe a central feature of all neuroblasts; namely, their ability to self-renew and generate differentiated daughter cells through asymmetric cell divisions, and we also assess how deregulation of this division mode can lead to tumorigenesis. Finally, we review the role of temporal patterning in neuroblasts for the orderly generation of different neural cell types during developmental progression.
The life history of a Drosophila neuroblast
The basic proliferative elements involved in building the Drosophila CNS are the stem cell-like multipotent neural progenitors referred to as neuroblasts. In the VNC and central brain, neuroblasts first arise by delamination from the neuroectoderm during embryonic development (Fig. 1a). In the embryonic neuroectoderm, groups of cells are singled out as proneural clusters through the expression of genes of the achaete–scute complex and are daughterless. In these clusters, neuroblasts become specified by Notch-dependent lateral inhibition from neighboring non-neuroblast cells, in a process in which proneural gene activity is restricted to only the presumptive neuroblast, but not in its neighbors (Artavanis-Tsakonas and Simpson 1991; Campos-Ortega 1993; Hartenstein and Wodarz 2013; Skeath and Thor 2003). Additionally, members of the Sox transcription factor family have also been reported to be involved in the formation of neuroblasts in a Notch-independent manner (Buescher et al. 2002; Overton et al. 2002). Following their specification, the neuroblasts of the VNC and central brain delaminate from the neuroectoderm, enlarge, and begin to proliferate during the short period of late embryogenesis to produce a small set of neurons that make up the simple larval CNS. These embryonically generated neurons are referred to as primary neurons and each neuroblast generates 10–20 primary neurons during embryonic development (Larsen et al. 2009; Lovick et al. 2013).
Neurogenesis in the CNS of Drosophila which occurs in two distinct periods: at embryonic and larval stage. a Neuroblasts of the ventral nerve cord derive from the neuroectoderm (NE) by delamination. Proliferating neuroblasts self-renew and generate one ganglion mother cell (GMC) by asymmetric division. The GMC, in turn, divides once more to produce two postmitotic cells, neurons or glial cells. b Neuroblasts in the postembryonic CNS. Schematic view of the Drosophila CNS in the third instar larva. Different types of neuroblasts are distributed in three anatomically different regions, the central brain (CB), optic lobe (OL), and ventral nerve cord (VNC). The central brain has three different types of neuroblasts, Type I, Type II, and mushroom body (MB) neuroblast
In the central brain and in the thoracic ganglia, most embryonic neuroblasts enter quiescence in the late embryonic stage (Egger et al. 2008; Younossi-Hartenstein et al. 1996). Exceptions are the four neuroblasts that generate the intrinsic neurons of the mushroom body, along with a fifth brain neuroblast, which do not undergo quiescence, and divide continuously throughout all larval stages to generate exceptionally large lineages of neurons in the adult CNS. Neuroblast entry into quiescence is mediated by intrinsically acting Hox genes as well as by temporal identity factors (Tsuji et al. 2008). Following quiescence, most of the remaining neuroblasts enlarge and restart cell division in the late first instar or early second instar of the larva. Re-entrance of the neuroblasts into the cell cycle is triggered by extrinsic signals, including nutritional or hormonal signals such as ecdysone (Colombani et al. 2012; Randhawa and Cohen 2005). Interestingly, the fat body and a glial cell niche mediate this process. In the presence of nutrition, an unknown secreted molecule from the fat body triggers release of the Drosophila insulin-like protein (Dilp) from glial cells. Through the insulin receptor (InR), Dilp activates the PI3K/AKT–Target of Rapamycin (TOR) signaling pathway in neuroblasts and this, in turn, induces the neuroblasts to exit quiescence, increase volume, and re-enter the cell cycle (Chell and Brand 2010; Shim et al. 2013; Sousa-Nunes et al. 2011). In contrast to the neuroblasts that undergo quiescence and reactivation, in the abdominal ganglia many of the embryonic neuroblasts are eliminated at late embryogenesis through programmed cell death.
The majority of the neurons that make up the adult central brain and VNC, termed secondary or adult-specific neurons, are generated by neuroblasts postembryonically during a prolonged period of intense proliferative activity which typically lasts from the end of the first larval instar until late larval/early pupal stages (Ito and Hotta 1992; Prokop and Technau 1991; Truman and Bate 1988). Thus, the development of the VNC and central brain is accomplished in two distinct periods of neurogenesis: a brief first period in embryonic stages and an extensive second period in larval stages. In the central brain, approximately 90 % of the neurons present in the adult brain are produced postembryonically by a stereotyped array of 100 embryonically derived neuroblast pairs (Technau et al. 2006; Urbach and Technau 2004). Neurogenesis in the adult central brain and ventral nerve cord has not been reported; however, a recent study indicates that adult neurogenesis can occur in the optic lobes after acute damage (Fernandez-Hernandez et al. 2013). This unexpected proliferative capability of progenitors in the Drosophila adult optic lobes may provide a useful model for studying the mechanisms that control neural stem cell proliferation in adult brain homeostasis and repair.
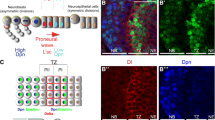
While the neuroblasts of the central brain and VNC, which can be further divided into type I and type II neuroblasts (see below), arise from the neuroectoderm of the early embryo, the neuroblasts of the optic lobe (OL) are generated from the neuroepithelial cells of the optic anlagen in larval stages (Fig. 1b). During early larval development, the embryonic optic placode generated by invagination of the OL primordium in the early embryonic stage expands dramatically in size through symmetric cell divisions, and becomes segregated into two separate epithelia termed inner proliferation center (IPC) and outer proliferation center (OPC). At the medial edge of the OPC, the neuroepithelial cells of the neuroectoderm are sequentially converted into neuroblasts of the medulla, which represents the largest neuropile of the OL (Egger et al. 2007). The dynamic transition of neuroectodermal cells to neuroblasts is triggered by a synchronized medial to lateral wave of expression of the proneural gene lethal of Scute (l’sc), which is more refined by integration of Notch signaling. (Egger et al. 2010, 2011). This neuroepithelium-to-neuroblast transition by the proneural wave is negatively regulated by JAK/STAT signaling and positively regulated by Fat-Hippo signaling (Reddy et al. 2010; Yasugi and Mizuno 2008; Yasugi et al. 2008).
Tight regulation of the precise time at which neuroblasts stop their proliferative divisions is critical for achieving the correct balance of early versus late-born neuronal fate and for determining the final number of neurons in the mature CNS. In the VNC and central brain, termination of neuroblast proliferation occurs either through apoptosis or by terminal differentiation (Reichert 2011). Since neuroblasts end their proliferative periods at different times in different regions of the developing CNS, the molecular mechanisms for terminating proliferation are varied for distinct neuroblasts. For example, a pulse of Hox protein expression leads to elimination of specific embryonic and postembryonic neuroblasts in the abdominal ganglia of the VNC, and the activation of pro-apoptotic genes, such as reaper, grim, and hid is involved in this process (Bello et al. 2003; Peterson et al. 2002). Hox gene expression in these neuroblasts is suppressed until the appropriate time by the Polycomb group (PcG) genes (Bello et al. 2007). In contrast, the mushroom body neuroblasts of the central brain, which do not undergo quiescence and continue proliferating until the end of the pupal stage, are prevented from premature cell cycle exit by mechanisms that involve the Tailless (Tll) transcription factor and the leucine-zipper protein Bunched (Kurusu et al. 2009; Siegrist et al. 2010). In the central brain and thoracic ganglia, most neuroblasts disappear due to terminal differentiation, which involves step-wise changes of the neuroblast’s cellular properties, including shrinkage of cell size, attenuation of the cell cycle, and expression of the homeodomain transcription factor Prospero (Pros), to terminate their proliferation. Pros promotes terminal differentiation of neuroblasts by inducing genes required for cell cycle exit and terminal differentiation (Maurange et al. 2008). In many cases, the timing of cell cycle exit of neuroblasts is controlled by the expression of a series of transcription factors (temporal transcription factor series; see below), which is also important for generating different cell types in a given neuroblast lineage (Almeida and Bray 2005; Cenci and Gould 2005; Maurange et al. 2008).
Diversity of neuroblast types in the CNS
With few exceptions, almost all neuroblasts in the CNS generate their postmitotic neural progeny through secondary progenitors, that can be either non-self-renewing or self-renewing (Reichert 2011). The so-called type I neuroblasts generate non-self-renewing secondary progenitors, referred to as ganglion mother cells (GMCs). Each stem cell-like division of the parent type I neuroblast (which self-renews) gives rise to one GMC which in turn divides only once to produce two postmitotic daughter cells, either neurons or glial cells (Fig. 2a). Due to the asymmetric segregation of the Notch signaling inhibitor Numb during this terminal GMC division, one of its daughter cells has active Notch signaling (“Notch-On”) while the other daughter has inhibited Notch signaling (“Notch-Off”). This difference translates into lineage-specific differences in the cellular and molecular properties of the two daughters such as axonal targeting, dendritic innervation, or survival. Since each type I neuroblast gives rise to numerous GMCs during its period of proliferative activity, its lineage of neural progeny comprises two “hemilineages”, one of which is Notch-On while the other is Notch-Off (Karcavich and Doe 2005; Karcavich 2005). This generic binary mechanism of asymmetric Notch signaling operating in all neuroblast lineages is an important factor in generating the remarkable neural diversity in the CNS and notably in the central brain and OL of Drosophila (Kumar et al. 2009; Li et al., 2010, 2013; Truman et al. 2010).
Different types of neuroblasts and their proliferation modes. a Type I neuroblasts (NB) divide asymmetrically to generate one neuroblast and one ganglion mother cell (GMC). The neuroblast self-renews and the GMC divides terminally into two neurons or glia. b Type II neuroblasts, eight of which are present in each hemisphere of the larval brain, divide asymmetrically to generate one self-renewing neuroblast and one immature intermediate neural precursor (INP) with transit amplifying function. The INP matures through expression of genes that inhibit dedifferentiation and promote lineage progression. Mature INPs produce one immature INP and one GMC through another asymmetric division. c Optic Lobe neuroblasts are generated by transition from neuroepithelial cells (NE) to neuroblasts induced at the medial edge of the outer proliferation center by a proneural wave. They proliferate in the type I mode
All the neuroblasts in the VNC and most of the neuroblasts in the central brain belong to the type I class. Although their characterization is still incomplete, the neuroblasts that generate the medulla neurons of the optic lobe also appear to belong to the type I class (Fig. 2c). In contrast, 8 neuroblasts located in the central brain hemispheres belong to a different class referred to as type II (Bello et al. 2008; Boone and Doe 2008; Bowman et al, 2008). These type II neuroblasts can be distinguished from type I neuroblasts by the absence of expression of the proneural transcription factor Asense and the cell fate determinant Pros (Bello et al. 2008; Boone and Doe 2008). Type II neuroblasts generate their lineages of neural progeny through transit amplifying self-renewing secondary progenitors called intermediate neural progenitors (INPs). Each INP undergoes a limited series of proliferative divisions, in each of which it self-renews and generates a GMC that divides once more to produce two postmitotic neural cells (Fig. 2b). Since each type II neuroblast generates numerous INPs and each INP generates several GMCs, a marked amplification of proliferation ensues, and lineages that are 4- to 5-fold larger than any type I lineages are produced. These remarkably large type II neuroblast lineages comprise up to 500 neural cells and, hence, make a substantial contribution to the complex circuitry of the central brain (Bello et al. 2008; Reichert 2011). For example, type II neuroblasts generate numerous neural cells, neurons, and glia, which contribute to an extensive midline neuropile structure, the central complex of the Drosophila central brain (Izergina et al. 2009; Viktorin et al. 2011). Moreover, and more strikingly, they also contribute to the optic lobe by generating glial cells, which migrate out of the central brain and differentiate into lobula giant glial cells (Viktorin et al. 2013). Interestingly, the pronounced amplification of proliferation achieved in type II neuroblast lineages is balanced by extensive programmed cell death in these lineages, and this likely helps to generate the precise number of differentiated neurons needed in corresponding brain circuitry (Jiang and Reichert 2012).
Recently, considerable insight into the mechanisms that control proliferation and lineage progression in type II neuroblast lineages, and notably in their INP sublineages, has been obtained. Immediately following their generation, INPs are in an immature state characterized cellularly by mitotic inactivity and arrest in the G2 phase and molecularly by the absence of expression of Asense and the bHLH-O transcription factor Deadpan (Bowman et al. 2008). During the following 4–5 h of cell cycle arrest, INPs mature and acquire the restricted developmental potential necessary for several ensuing asymmetric cell divisions. During each of these cell divisions the mature Asense- and Deadpan-positive INPs self-renew and generate a GMC which gives rise to two neuronal or glial cells (Bayraktar et al. 2010). During the initial asymmetric division of the type II neuroblast, the cell fate determinants Brain tumor (Brat) and Numb are segregated into the INP daughter where they play an essential role in establishing INP potential (Bowman et al. 2008). Numb specifies INP identity by antagonizing the Notch pathway. Brat, on the other hand, contributes to the identity of INPs by blocking their potential dedifferentiation into neuroblast-like progenitors, and this process is likely to be mediated by suppressing the action of the self-renewal factor Klumpfuss through attenuation of β-catenin/Armadillo activity (Berger et al. 2012; Komori et al. 2014; Xiao et al. 2012). Additional restriction of INP dedifferentiation potential is mediated by dFezf/Earmuff (Erm), which is expressed in mature INPs and prevents their dedifferentiation by activating Prospero (Pros) to limit proliferation as well as by antagonizing Notch signaling (Weng et al. 2010; Weng and Lee 2011). Mutation in any one of the genes that encode these INP specifying molecules including Brat, Numb or Erm results in the failure of neural differentiation and overgrowth of Type II neuroblasts or INPs (see below) (Bowman et al. 2008; Weng et al. 2010). Recently, several new genes involved in proliferation and differentiation of type I and type II neuroblasts have been identified by genome-wide transgenic RNAi screening (Neumuller et al. 2011). Further investigation of these new candidate genes is likely to result in additional information concerning the mechanisms that control neurogenesis in different neuroblast types.
Neuroblasts proliferate in a stem cell mode
A defining feature of stem cells is their ability to self-renew and at the same time generate daughter cells, that are committed to further differentiation, in one and the same cell cycle. This feature is usually linked to the ability of stem cells to undergo asymmetric cell divisions. All of the neural stem cell-like neuroblasts in the developing CNS of Drosophila, be they type I, type II, or OL neuroblasts, divide in an asymmetric stem cell mode (Benito-Sipos et al. 2011; Brody and Odenwald 2000; Egger et al. 2008; Isshiki et al. 2001; Kambadur et al. 1998; Karlsson et al. 2010; Reichert 2011; Touma et al. 2012; Tran and Doe 2008). Indeed, many of the basic cellular processes and molecular mechanisms that operate in asymmetric stem cell division have been elucidated in the Drosophila neuroblast models (Januschke and Gonzalez 2008; Knoblich 2008; Schaefer and Knoblich 2001; Wu et al. 2008; Zhong and Chia 2008). While type I and type II neuroblasts differ in some aspects of their asymmetric cell division modes, a fundamental property of the asymmetric divisions manifested by these neuroblasts is the unequal segregation of proteins that assign cell polarity and cell fate to the two asymmetric daughter cells, the self-renewing neuroblast and the more differentiated daughter cell (GMC or INP) (Doe 2008; Homem and Knoblich 2012; Knoblich 2008; Neumuller and Knoblich 2009). This unequal segregation of molecular determinants involves two major molecular complexes that act in the neuroblast during the cell cycle (Fig. 3).
Asymmetric cell division of neuroblasts. a Through asymmetric cell division neuroblasts self-renew and simultaneously generate a more differentiated GMC. In the mitotically active neuroblast, a Par3/Par6/aPKC protein complex localized asymmetrically at the apical cortex is linked to the Pins/ Gαi/Mud protein complex via the scaffolding protein Inscuteable. Cell fate determinants including Pros, Brat, and Numb are asymmetrically localized at the basal cortex together with their adaptor proteins, Mira and Pon. During asymmetric cell division, these cell fate determinants are exclusively segregated into the GMC where they induce various differentiation events. b The apical protein complexes mediate the basal localization of cell fate determinants through protein phosphorylation cascades. Aur-A phosphorylates Par6 to activate aPKC in the complex. aPKC phosphorylates Lgl, Numb, and Mira. Phosphorylated Mira carries Pros and Brat to the basal cortex. Polo is also involved in asymmetric protein distribution by phosphorylating Numb and Pon. A apical, B basal
A so-called apical complex is essential for determining the axis of polarity and the orientation of the mitotic spindle in the neuroblast. This apical complex consists of the Par3/Par6/aPKC subcomplex and the Pins/Gαi/Mud subcomplex, both of which are localized in the apical region of the neuroblast and are linked via Inscuteable (Insc) protein. The Pins/Gαi/Mud protein complex is required for proper spindle orientation. Mud binds directly to astral microtubules so that Pins/Gαi/Mud–Insc–par3/Par6/aPKC can exert a pulling force on the spindle of the dividing neuroblast (Izumi et al. 2006; Kraut and Campos-Ortega 1996; Kraut et al. 1996; Siller et al. 2006; Speicher et al. 2008). The Par3/Par6/aPKC complex is involved in setting up and maintaining the apical–basal axis of polarity in the neuroblast. This complex is also responsible for the basal localization of cell fate determinants through sequential phosphorylation events that occur in the apical region of the neuroblast (Betschinger et al. 2003; Knoblich 2008; Wirtz-Peitz et al. 2008; Yamanaka et al. 2006). For example, the mitotically active kinase Aurora A (Aur-A) phosphorylates Par6 resulting in activation of aPKC which then phosphorylates specific cell fate determinants located in the apical region of the neuroblast’s cell cortex resulting in their release from the cortex apically and, hence, in their basal accumulation (Fig. 3b).
Three major cell fate determinants, Numb, Brat, and Pros, and two adaptor proteins, Miranda (Mira) and Partner-of-Numb (Pon) make up the so-called basal complex in the proliferating neuroblast. During asymmetric cell division of the neuroblast, these basally localized proteins are segregated into the smaller daughter cell, where they act in promoting differentiation and suppressing proliferation. Numb is a membrane bound Notch inhibitor containing a phosphoserine-binding domain. Numb participates in specifying GMC fate by promoting endocytosis of Notch, thus maintaining Notch at a lower level in the GMC than in the neuroblast (Bowman et al. 2008; Rhyu et al. 1994; Spana and Doe 1996; Spana et al. 1995; Uemura et al. 1989; Wang et al. 2007; Zhong et al. 1996). Pros is involved in specifying neuronal and glial cell types in the developing nervous system, and during asymmetric cell division of the neuroblast, Pros is segregated together with Mira into the GMC. Upon completion of cell division, Mira is degraded and Pros is released from the cortex and enters into the nucleus, where it specifies GMC identity by promoting the expression of GMC-specific genes and repressing the expression of neuroblast-specific genes (Atwood and Prehoda 2009; Choksi et al. 2006; Ikeshima-Kataoka et al. 1997; Li and Vaessin 2000; Shen et al. 1997). Thus, Pros negatively regulates the expression of cell cycle genes such as cyclin A, cyclin E, and string, a Drosophila homolog of Cdc25, and positively regulates the expression of dacapo, a cyclin-dependent kinase inhibitor. Pros also activates many genes involved in terminal differentiation of neurons such as fasciclin II and netrin B (Choksi et al. 2006). Brat, an NHL containing translation regulator, is thought to regulate ribosomal protein biosynthesis and to inhibit the transcription factor Myc at the posttranscriptional level. Like Pros, Brat is exclusively segregated with Mira into the GMC during mitosis and contributes to GMC specification by decreasing protein synthesis (Bello et al. 2006; Betschinger et al. 2006; Lee et al. 2006b).
As in other stem cell lineages, maintaining the precise balance between self-renewal and differentiation in asymmetrically dividing neuroblast lineages is essential to ensure normal development of the CNS as well as to prevent accumulation of aberrant neural stem cell-like progenitors. Indeed, recent studies using Drosophila neuroblasts have shown that defects in the key molecular mechanisms involved in asymmetric cell division control can result in loss of differentiated cells and uncontrolled overgrowth of neuroblast-like cells leading to brain tumor formation (Bello et al. 2007; Caussinus and Gonzalez 2005; Chang et al. 2012; Knoblich 2008) (Fig. 4). Notably, mutations in genes that result in defects in function or asymmetric localization of cell fate determinants such as mutations in Pros, Numb, Brat or in their adaptors Mira and Pon result in massive tumorous overproliferation in the brain due to the production of supernumerary self-renewing daughters at the expense of differentiated cells (Bello et al. 2006; Betschinger et al. 2006; Choksi et al. 2006; Wang et al. 2006a). Neural tumors also result from mutation of other genes involved in asymmetric cell division such as discs large (dlg), lethal giant larva (lgl), and scribble (scrib) or the genes encoding the Aur-A and Polo kinases (Beaucher et al. 2007; Lee et al. 2006a; Ohshiro et al. 2000; Peng et al. 2000; Reichert, 2011; Wang et al. 2006b, 2007;). All the resulting neural tumor cells undergo massive overgrowth upon transplantation into wild-type hosts, kill the host within weeks, and become immortalized and can be serially transplanted into successive hosts over years (Beaucher et al. 2007; Caussinus and Gonzalez 2005). These transplanted cells can also exhibit metastatic behavior, migrating away from the site of the primary tumor, passing through several cell layers, and establishing secondary colonies. As might be expected, type II neuroblasts are more susceptible to tumorigenesis, since their lineages comprise two cell types with self-renewing capability, namely neuroblasts and INPs.
Defects in asymmetric cell division of neuroblasts cause tumorigenesis. Defects in the molecular machinery involved in asymmetric cell division, including mutations of cell fate determinant genes, pros and brat, cause tumor cell-like overgrowth. While the mutant neuroblasts often still divide asymmetrically, their secondary progenitor progeny (GMC in type I neuroblasts and INP/GMC in type II neuroblasts) do not generate differentiated neural cells but rather revert to neuroblast-like cells that continue to divide in an uncontrolled manner. a Normal neuroblast proliferation leading to differentiated neural cells. b Mutant neuroblast overproliferation leading to tumorigenesis
Temporal patterning of neuroblast proliferation
The ensemble of neuroblasts in the Drosophila CNS can give rise to an astounding diversity of neural cell types. While the molecular mechanisms that make this possible are not completely understood, the requirement of both positional and temporal information in proliferating neuroblasts for the generation of different neural cell types in its lineal progeny has been firmly established. Positional information is provided to each neuroblast of the central brain and VNC by the early embryonic expression of anteroposterior and dorsoventral patterning genes (Bossing et al. 1996; Broadus and Doe 1995; Doe 1992; Doe and Technau 1993; Schmidt et al. 1997; Urbach and Technau 2003). These two sets of developmental control genes, which include the Hox genes, the gap genes, the segment polarity genes, and the columnar genes, establish a Cartesian grid-like molecular coordinate system in the neuroectoderm, from which the neuroblasts derive. As a result, each neuroblast acquires a specific combination of developmental control genes, which contribute to the specific identity of the neuroblast. As shown by an enormous body of genetic evidence, this “combinatorial code” of transcription factors can directly influence the neural cell types that a given neuroblast generates (Skeath 1999; Skeath and Thor 2003; Technau et al. 2006; Urbach and Technau 2004).
In addition to positional information, temporal information is also required in neuroblasts, notably for the generation of different cell types in their lineage of progeny at different times during the proliferation process. The time at which a given progeny is produced and exits the cell cycle is referred to as its birth date, and different progeny are generated by the parent neuroblast in a fixed birth order. The basic molecular mechanism that links birth order to neuronal fate involves a stereotyped temporal series of transcription factors expressed in the parent neuroblast. This temporal transcription factor series was first identified in the proliferating embryonic neuroblasts of the VNC (Fig. 5a), where a serial cascade of transient expression of the five transcription factors Hunchback (Hb), Krüppel (Kr), Pdm, Castor (Cas), and Grainyhead (Grh) takes place (Baumgardt et al. 2009; Benito-Sipos et al. 2010; Brody and Odenwald 2000; Grosskortenhaus et al. 2005, 2006; Isshiki et al. 2001; Kambadur et al. 1998; Novotny et al. 2002; Pearson and Doe 2003). The temporal transition of transcription factors is facilitated by cross-regulation among these transcription factors, which usually involves both positive feedforward regulation and negative feedback regulation (Baumgardt et al. 2009; Nakajima et al. 2010). However, this cross-regulation is not necessarily required for temporal series progression since loss of one of the transcription factors Hb, Kr, or Pdm does not result in a blockage of the temporal series but only in the skipping of one temporal identity (Brody and Odenwald 2000; Grosskortenhaus et al. 2006; Isshiki et al. 2001; Maurange et al. 2008; Tran and Doe 2008). The specific molecular signals that control the switch in expression from one transcription factor to the next are still unclear.
Temporal patterning of neuroblast proliferation. a Embryonic neuroblasts in the VNC express a temporal series of the transcription factors, Hb, Kr, Pdm, Cas, and Grh as they age. The temporal transcription factor expressed in the neuroblast is inherited by its GMC and specifies the identity of its two neural cell progenies. During embryogenesis, a transient burst of Svp expression is required for the switch from Hb to Kr expression. Cas expression is maintained through quiescence and defines the temporal identity of the larval neuroblast until Svp is re-expressed. b Serial expression of Hth, Klu, Ey, Slp, D, and Tll transcription factors in the medulla neuroblasts of the OL during postembryonic development. c Combinatorial temporal patterning in type II neuroblast lineages. In addition to a temporal series expressed in the type II neuroblasts, a second different temporal series comprising D, Grh, and Ey is expressed in each INP. Thus, two axes of temporal transcription factor cascades interact to generate a large diversity of neural cell types in these lineages
Each of the transcription factors in this temporal series is expressed in the proliferating neuroblast during a specific time window, and the GMC that is generated by the neuroblast during that time window inherits the expression of that transcription factor. In consequence, the neurons that derive from the GMC inherit and maintain the expression of the same transcription factor, which is both required and sufficient for their birth order-dependent neuronal specification (Homem and Knoblich 2012; Li et al. 2014). While the positional information acquired by each neuroblast in a neurogenic array is distinct, the temporal information manifested in proliferating neuroblasts has a more generic character. Many of the neuroblasts in the embryonic VNC manifest the same temporal series of Hb, Kr, Pdm, Cas, and Grh expression. However, since different neuroblasts generate different lineal cell types, this temporal series does not control neural cell type per se. Rather, it specifies birth order-dependent neural identity, which together with positional identity provided by spatial combinations of transcription factor expression (and with hemilineage-specific Notch signaling) is translated into the specific neural cell types produced in a neuroblast lineage.
Temporal specification is not limited to embryogenesis but also occurs during postembryonic neurogenesis. In VNC neuroblasts, two transcription factors, Cas and Sevenup (Svp), act in a postembryonic temporal series; Cas expression in late embryonic neuroblasts is maintained in postembryonic neuroblasts after exit from quiescence and is followed by a wave of Svp expression (Maurange et al. 2008; Zhu et al. 2006). Other members of the postembryonic temporal series must also exist; however, they have not yet been identified. A more complete characterization of a postembryonic temporal series has been carried out in OL development where a different temporal series of transcriptional factors has been identified (Fig 5b). In the OL neuroblasts of the developing medulla, a temporal transcription factor series composed of Homothorax (Hth), Klumpfuss (Klu), Eyeless (Ey), Sloppy-paired (Slp), Dichaete (D), and Tailless (Tll) is expressed (Li et al. 2013; Suzuki et al. 2013). Moreover, crossregulatory interactions are required between some, but not all, of these transcription factors. Mutational inactivation or overexpression of individual members of this temporal series in OL neuroblasts affects birth order-dependent expression of different neuronal markers in the neural cells that are generated by these progenitors, implying that the temporal transcription factors control OL neuronal fate. An interesting concatenation of two different temporal transcription factor series is seen during postembryonic development in type II neuroblast lineages (Bayraktar and Doe 2013). The type II neuroblasts themselves serially express the transcription factors D/Cas and Svp, and more temporal transcription factors are likely to exist as well in these neuroblasts. In addition, each INP daughter cell generated by a type II neuroblast also expresses its own series of temporal transcription factors, namely D, Grh, and Ey, in the sublineage of cells that it generates. Mutation or overexpression of the temporal transcription factors in INPs demonstrate the requirement of these factors in fate determination of the lineal neural progeny in INP sublineages, and also show that the sequential expression of these transcription factors is tightly controlled by cross-regulation mechanisms. This type of combinatorial temporal patterning composed by two different axes of temporal transcription factor cascades leads to a larger diversity of neurons and glial cells in complex neural lineages of type II neuroblasts.
Taken together, these findings indicate that virtually all neuroblast lineages in the developing CNS utilize transcription factor cascades as a generic mechanism for temporal patterning and determination of neural cell fate. The specific transcription factor combinations utilized in type I, type II, and OL neuroblasts differs. However, the functional role of the resulting temporal information, integrated together with positional information and binary Notch signaling, is a common one, namely the generation of the remarkable diversity of cell types in the developing CNS from a surprisingly small set of neural stem cell-like precursors. These features exemplify two emerging principles in the molecular control of specification: proliferation and differentiation in neural stem cell lineages. First, the same fundamental developmental process can involve different sets of controlling factors such as the different combinations of transcription factors that make up the temporal series in type I, type II, and OL neuroblasts. Second, the same controlling factor can be involved in widely different developmental processes in neuroblast lineages including, but not limited to, specification of positional information, control of lineage progression, temporal patterning and cell fate. If similar considerations hold for neural stem cell lineages in vertebrate brain development, we would predict that while the basic developmental processes are likely to be mechanistically similar and evolutionarily conserved, the specific identity of the molecular players involved in these fundamental processes will be more divergent.
Conclusion
Drosophila neuroblasts have emerged as an excellent model for understanding the cellular molecular mechanisms involved in neural stem cell self-renewal and differentiation. The genetic basis for the generation of these neural stem cells from the neuroectoderm as well as many of the mechanisms that operate in these primary progenitors during their asymmetric proliferative cell divisions have been elucidated. Moreover, the processes that integrate amplification of proliferation with restricted lineage progression in transit amplifying intermediate progenitors are beginning to be understood. Finally, insight into the combinatorial molecular code that imparts positional and temporal information to neural stem cells as well as the role of these two types of information in specifying the diversity of differentiated neural cell types generated by individual neural stem cells is being obtained. Given the remarkable conservation of molecular mechanisms involved in nervous system development in Drosophila and vertebrates including mammals, the investigations of all of these features of neural stem cell biology in the fly model is likely to help in understanding the roles of neural stem cells in generating the highly complex human brain. From this perspective, the use of the Drosophila model for unraveling the mechanisms underlying neural stem cell derived brain tumors may also lead to important insight into the aberrant molecular mechanisms that cause brain tumors in human patients.
Abbreviations
- aPKC:
-
Atypical protein kinase C
- CNS:
-
Central nerve system
- Gαi:
-
G protein α i subunit 65A
- INP:
-
Intermediate neural progenitor
- Mud:
-
Mushroom body defect
- Par3:
-
Partitioning defect 3
- Par6:
-
Partitioning defect 6
- Pins:
-
Partner of Inscuteable
- Pon:
-
Partner of Numb
References
Almeida MS, Bray SJ (2005) Regulation of post-embryonic neuroblasts by Drosophila Grainyhead. Mech Dev 122:1282–1293
Artavanis-Tsakonas S, Simpson P (1991) Choosing a cell fate: a view from the Notch locus. Trends Genetics 7:403–408
Atwood SX, Prehoda KE (2009) aPKC Phosphorylates Miranda to polarize fate determinants during neuroblast asymmetric cell division. Curr Biol 19:723–729
Baumgardt M, Karlsson D, Terriente J, Diaz-Benjumea FJ, Thor S (2009) Neuronal subtype specification within a lineage by opposing temporal feed-forward loops. Cell 139:969–982
Bayraktar OA, Boone JQ, Drummond ML, Doe CQ (2010) Drosophila type II neuroblast lineages keep Prospero levels low to generate large clones that contribute to the adult brain central complex. Neural Dev 5:26
Bayraktar OA, Doe CQ (2013) Combinatorial temporal patterning in progenitors expands neural diversity. Nature 498:449–455
Beaucher M, Goodliffe J, Hersperger E, Trunova S, Frydman H, Shearn A (2007) Drosophila brain tumor metastases express both neuronal and glial cell type markers. Dev Biol 301:287–297
Bello B, Holbro N, Reichert H (2007) Polycomb group genes are required for neural stem cell survival in postembryonic neurogenesis of Drosophila. Development 134:1091–1099
Bello B, Reichert H, Hirth F (2006) The brain tumor gene negatively regulates neural progenitor cell proliferation in the larval central brain of Drosophila. Development 133:2639–2648
Bello BC, Hirth F, Gould AP (2003) A pulse of the Drosophila Hox protein Abdominal-A schedules the end of neural proliferation via neuroblast apoptosis. Neuron 37:209–219
Bello BC, Izergina N, Caussinus E, Reichert H (2008) Amplification of neural stem cell proliferation by intermediate progenitor cells in Drosophila brain development. Neural Dev 3:5
Benito-Sipos J, Estacio-Gomez A, Moris-Sanz M, Baumgardt M, Thor S, Diaz-Benjumea FJ (2010) A genetic cascade involving klumpfuss, nab and castor specifies the abdominal leucokinergic neurons in the Drosophila CNS. Development 137:3327–3336
Benito-Sipos J, Ulvklo C, Gabilondo H, Baumgardt M, Angel A, Torroja L, Thor S (2011) Seven up acts as a temporal factor during two different stages of neuroblast 5-6 development. Development 138:5311–5320
Berger C, Harzer H, Burkard TR, Steinmann J, van der Horst S, Laurenson AS, Novatchkova M, Reichert H, Knoblich JA (2012) FACS purification and transcriptome analysis of drosophila neural stem cells reveals a role for Klumpfuss in self-renewal. Cell Rep 2:407–418
Betschinger J, Mechtler K, Knoblich JA (2003) The Par complex directs asymmetric cell division by phosphorylating the cytoskeletal protein Lgl. Nature 422:326–330
Betschinger J, Mechtler K, Knoblich JA (2006) Asymmetric segregation of the tumor suppressor brat regulates self-renewal in Drosophila neural stem cells. Cell 124:1241–1253
Boone JQ, Doe CQ (2008) Identification of Drosophila type II neuroblast lineages containing transit amplifying ganglion mother cells. Dev Neurol 68:1185–1195
Bossing T, Udolph G, Doe CQ, Technau GM (1996) The embryonic central nervous system lineages of Drosophila melanogaster. I. Neuroblast lineages derived from the ventral half of the neuroectoderm. Dev Biol 179:41–64
Bowman SK, Rolland V, Betschinger J, Kinsey KA, Emery G, Knoblich JA (2008) The tumor suppressors Brat and Numb regulate transit-amplifying neuroblast lineages in Drosophila. Dev Cell 14:535–546
Broadus J, Doe CQ (1995) Evolution of neuroblast identity: seven-up and prospero expression reveal homologous and divergent neuroblast fates in Drosophila and Schistocerca. Development 121:3989–3996
Brody T, Odenwald WF (2000) Programmed transformations in neuroblast gene expression during Drosophila CNS lineage development. Dev Biol 226:34–44
Buescher M, Hing FS, Chia W (2002) Formation of neuroblasts in the embryonic central nervous system of Drosophila melanogaster is controlled by SoxNeuro. Development 129:4193–4203
Campos-Ortega JA (1993) Mechanisms of early neurogenesis in Drosophila melanogaster. J Neurobiol 24:1305–1327
Caussinus E, Gonzalez C (2005) Induction of tumor growth by altered stem-cell asymmetric division in Drosophila melanogaster. Nat Genet 37:1125–1129
Cenci C, Gould AP (2005) Drosophila Grainyhead specifies late programmes of neural proliferation by regulating the mitotic activity and Hox-dependent apoptosis of neuroblasts. Development 132:3835–3845
Chang KC, Wang C, Wang H (2012) Balancing self-renewal and differentiation by asymmetric division: insights from brain tumor suppressors in Drosophila neural stem cells. BioEssays 34:301–310
Chell JM, Brand AH (2010) Nutrition-responsive glia control exit of neural stem cells from quiescence. Cell 143:1161–1173
Choksi SP, Southall TD, Bossing T, Edoff K, de Wit E, Fischer BE, van Steensel B, Micklem G, Brand AH (2006) Prospero acts as a binary switch between self-renewal and differentiation in Drosophila neural stem cells. Dev Cell 11:775–789
Colombani J, Andersen DS, Leopold P (2012) Secreted peptide Dilp8 coordinates Drosophila tissue growth with developmental timing. Science 336:582–585
Doe CQ (1992) Molecular markers for identified neuroblasts and ganglion mother cells in the Drosophila central nervous system. Development 116:855–863
Doe CQ (2008) Neural stem cells: balancing self-renewal with differentiation. Development 135:1575–1587
Doe CQ, Technau GM (1993) Identification and cell lineage of individual neural precursors in the Drosophila CNS. Trends Neurosci 16:510–514
Egger B, Boone JQ, Stevens NR, Brand AH, Doe CQ (2007) Regulation of spindle orientation and neural stem cell fate in the Drosophila optic lobe. Neural Dev 2:1
Egger B, Chell JM, Brand AH (2008) Insights into neural stem cell biology from flies. Philos Trans R Soc Lond B 363:39–56
Egger B, Gold KS, Brand AH (2010) Notch regulates the switch from symmetric to asymmetric neural stem cell division in the Drosophila optic lobe. Development 137:2981–2987
Egger B, Gold KS, Brand AH (2011) Regulating the balance between symmetric and asymmetric stem cell division in the developing brain. Fly 5:237–241
Fernandez-Hernandez I, Rhiner C, Moreno E (2013) Adult neurogenesis in Drosophila. Cell Reports 3:1857–1865
Grosskortenhaus R, Pearson BJ, Marusich A, Doe CQ (2005) Regulation of temporal identity transitions in Drosophila neuroblasts. Dev Cell 8:193–202
Grosskortenhaus R, Robinson KJ, Doe CQ (2006) Pdm and Castor specify late-born motor neuron identity in the NB7-1 lineage. Genes Dev 20:2618–2627
Hartenstein V, Wodarz A (2013) Initial neurogenesis in Drosophila. Dev Biol 2:701–721
Homem CC, Knoblich JA (2012) Drosophila neuroblasts: a model for stem cell biology. Development 139:4297–4310
Ikeshima-Kataoka H, Skeath JB, Nabeshima Y, Doe CQ, Matsuzaki F (1997) Miranda directs Prospero to a daughter cell during Drosophila asymmetric divisions. Nature 390:625–629
Isshiki T, Pearson B, Holbrook S, Doe CQ (2001) Drosophila neuroblasts sequentially express transcription factors which specify the temporal identity of their neuronal progeny. Cell 106:511–521
Ito K, Hotta Y (1992) Proliferation pattern of postembryonic neuroblasts in the brain of Drosophila melanogaster. Dev Biol 149:134–148
Izergina N, Balmer J, Bello B, Reichert H (2009) Postembryonic development of transit amplifying neuroblast lineages in the Drosophila brain. Neural Dev 4:44
Izumi Y, Ohta N, Hisata K, Raabe T, Matsuzaki F (2006) Drosophila Pins-binding protein Mud regulates spindle-polarity coupling and centrosome organization. Nat Cell Biol 8:586–593
Januschke J, Gonzalez C (2008) Drosophila asymmetric division, polarity and cancer. Oncogene 27:6994–7002
Jiang Y, Reichert H (2012) Programmed cell death in type II neuroblast lineages is required for central complex development in the Drosophila brain. Neural Dev 7:3
Kambadur R, Koizumi K, Stivers C, Nagle J, Poole SJ, Odenwald WF (1998) Regulation of POU genes by castor and hunchback establishes layered compartments in the Drosophila CNS. Genes Dev 12:246–260
Karcavich R, Doe CQ (2005) Drosophila neuroblast 7-3 cell lineage: a model system for studying programmed cell death, Notch/Numb signaling, and sequential specification of ganglion mother cell identity. J Comp Neurol 481:240–251
Karcavich RE (2005) Generating neuronal diversity in the Drosophila central nervous system: a view from the ganglion mother cells. Dev Dyn 232:609–616
Karlsson D, Baumgardt M, Thor S (2010) Segment-specific neuronal subtype specification by the integration of anteroposterior and temporal cues. PLoS Biol 8:e1000368
Knoblich JA (2008) Mechanisms of asymmetric stem cell division. Cell 132:583–597
Komori H, Xiao Q, McCartney BM, Lee CY (2014) Brain tumor specifies intermediate progenitor cell identity by attenuating beta-catenin/Armadillo activity. Development 141:51–62
Kraut R, Campos-Ortega JA (1996) inscuteable, a neural precursor gene of Drosophila, encodes a candidate for a cytoskeleton adaptor protein. Dev Biol 174:65–81
Kraut R, Chia W, Jan LY, Jan YN, Knoblich JA (1996) Role of inscuteable in orienting asymmetric cell divisions in Drosophila. Nature 383:50–55
Kumar A, Bello B, Reichert H (2009) Lineage-specific cell death in postembryonic brain development of Drosophila. Development 136:3433–3442
Kurusu M, Maruyama Y, Adachi Y, Okabe M, Suzuki E, Furukubo-Tokunaga K (2009) A conserved nuclear receptor, Tailless, is required for efficient proliferation and prolonged maintenance of mushroom body progenitors in the Drosophila brain. Dev Biol 326:224–236
Larsen C, Shy D, Spindler SR, Fung S, Pereanu W, Younossi-Hartenstein A, Hartenstein V (2009) Patterns of growth, axonal extension and axonal arborization of neuronal lineages in the developing Drosophila brain. Dev Biol 335:289–304
Lee CY, Andersen RO, Cabernard C, Manning L, Tran KD, Lanskey MJ, Bashirullah A, Doe CQ (2006a) Drosophila Aurora-A kinase inhibits neuroblast self-renewal by regulating aPKC/Numb cortical polarity and spindle orientation. Genes Dev 20:3464–3474
Lee CY, Wilkinson BD, Siegrist SE, Wharton RP, Doe CQ (2006b) Brat is a Miranda cargo protein that promotes neuronal differentiation and inhibits neuroblast self-renewal. Dev Cell 10:441–449
Li L, Vaessin H (2000) Pan-neural Prospero terminates cell proliferation during Drosophila neurogenesis. Genes Dev 14:147–151
Li S, Wang C, Sandanaraj E, Aw SS, Koe CT, Wong JJ, Yu F, Ang BT, Tang C, Wang H (2014) The SCFSlimb E3 ligase complex regulates asymmetric division to inhibit neuroblast overgrowth. EMBO Rep 15:165–174
Li X, Erclik T, Bertet C, Chen Z, Voutev R, Venkatesh S, Morante J, Celik A, Desplan C (2013) Temporal patterning of Drosophila medulla neuroblasts controls neural fates. Nature 498:456–462
Lin S, Lai SL, Yu HH, Chihara T, Luo L, Lee T (2010) Lineage-specific effects of Notch/Numb signaling in post-embryonic development of the Drosophila brain. Development 137:43–51
Lovick JK, Ngo KT, Omoto JJ, Wong DC, Nguyen JD, Hartenstein V (2013) Postembryonic lineages of the Drosophila brain: I. Development of the lineage associated fiber tracts Dev Biol 384:228–257
Maurange C, Cheng L, Gould AP (2008) Temporal transcription factors and their targets schedule the end of neural proliferation in Drosophila. Cell 133:891–902
Nakajima A, Isshiki T, Kaneko K, Ishihara S (2010) Robustness under functional constraint: the genetic network for temporal expression in Drosophila neurogenesis. PLoS Comput Biol 6:e1000760
Neumuller RA, Knoblich JA (2009) Wicked views on stem cell news. Nat Cell Biol 11:678–679
Neumuller RA, Richter C, Fischer A, Novatchkova M, Neumuller KG, Knoblich JA (2011) Genome-wide analysis of self-renewal in Drosophila neural stem cells by transgenic RNAi. Cell Stem Cell 8:580–593
Novotny T, Eiselt R, Urban J (2002) Hunchback is required for the specification of the early sublineage of neuroblast 7-3 in the Drosophila central nervous system. Development 129:1027–1036
Ohshiro T, Yagami T, Zhang C, Matsuzaki F (2000) Role of cortical tumour-suppressor proteins in asymmetric division of Drosophila neuroblast. Nature 408:593–596
Overton PM, Meadows LA, Urban J, Russell S (2002) Evidence for differential and redundant function of the Sox genes Dichaete and SoxN during CNS development in Drosophila. Development 129:4219–4228
Pearson BJ, Doe CQ (2003) Regulation of neuroblast competence in Drosophila. Nature 425:624–628
Peng CY, Manning L, Albertson R, Doe CQ (2000) The tumour-suppressor genes lgl and dlg regulate basal protein targeting in Drosophila neuroblasts. Nature 408:596–600
Peterson C, Carney GE, Taylor BJ, White K (2002) reaper is required for neuroblast apoptosis during Drosophila development. Development 129:1467–1476
Prokop A, Technau GM (1991) The origin of postembryonic neuroblasts in the ventral nerve cord of Drosophila melanogaster. Development 111:79–88
Randhawa R, Cohen P (2005) The role of the insulin-like growth factor system in prenatal growth. Mol Genet Metab 86:84–90
Reddy BV, Rauskolb C, Irvine KD (2010) Influence of fat-hippo and notch signaling on the proliferation and differentiation of Drosophila optic neuroepithelia. Development 137:2397–2408
Reichert H (2011) Drosophila neural stem cells: cell cycle control of self-renewal, differentiation, and termination in brain development. Results Probl Cell Differ 53:529–546
Rhyu MS, Jan LY, Jan YN (1994) Asymmetric distribution of numb protein during division of the sensory organ precursor cell confers distinct fates to daughter cells. Cell 76:477–491
Schaefer M, Knoblich JA (2001) Protein localization during asymmetric cell division. Exp Cell Res 271:66–74
Schmidt H, Rickert C, Bossing T, Vef O, Urban J, Technau GM (1997) The embryonic central nervous system lineages of Drosophila melanogaster. II. Neuroblast lineages derived from the dorsal part of the neuroectoderm. Dev Biol 189:186–204
Shen CP, Jan LY, Jan YN (1997) Miranda is required for the asymmetric localization of Prospero during mitosis in Drosophila. Cell 90:449–458
Shim J, Gururaja-Rao S, Banerjee U (2013) Nutritional regulation of stem and progenitor cells in Drosophila. Development 140:4647–4656
Siegrist SE, Haque NS, Chen CH, Hay BA, Hariharan IK (2010) Inactivation of both Foxo and reaper promotes long-term adult neurogenesis in Drosophila. Curr Biol 20:643–648
Siller KH, Cabernard C, Doe CQ (2006) The NuMA-related Mud protein binds Pins and regulates spindle orientation in Drosophila neuroblasts. Nat Cell Biol 8:594–600
Skeath JB (1999) At the nexus between pattern formation and cell-type specification: the generation of individual neuroblast fates in the Drosophila embryonic central nervous system. BioEssays 21:922–931
Skeath JB, Thor S (2003) Genetic control of Drosophila nerve cord development. Curr Opin Neurobiol 13:8–15
Sousa-Nunes R, Yee LL, Gould AP (2011) Fat cells reactivate quiescent neuroblasts via TOR and glial insulin relays in Drosophila. Nature 471:508–512
Spana EP, Doe CQ (1996) Numb antagonizes Notch signaling to specify sibling neuron cell fates. Neuron 17:21–26
Spana EP, Kopczynski C, Goodman CS, Doe CQ (1995) Asymmetric localization of numb autonomously determines sibling neuron identity in the Drosophila CNS. Development 121:3489–3494
Speicher S, Fischer A, Knoblich J, Carmena A (2008) The PDZ protein Canoe regulates the asymmetric division of Drosophila neuroblasts and muscle progenitors. Curr Biol 18:831–837
Suzuki T, Kaido M, Takayama R, Sato M (2013) A temporal mechanism that produces neuronal diversity in the Drosophila visual center. Dev Biol 380:12–24
Technau GM, Berger C, Urbach R (2006) Generation of cell diversity and segmental pattern in the embryonic central nervous system of Drosophila. Dev Dyn 235:861–869
Touma JJ, Weckerle FF, Cleary MD (2012) Drosophila Polycomb complexes restrict neuroblast competence to generate motoneurons. Development 139:657–666
Tran KD, Doe CQ (2008) Pdm and Castor close successive temporal identity windows in the NB3-1 lineage. Development 135:3491–3499
Truman JW, Bate M (1988) Spatial and temporal patterns of neurogenesis in the central nervous system of Drosophila melanogaster. Dev Biol 125:145–157
Truman JW, Moats W, Altman J, Marin EC, Williams DW (2010) Role of Notch signaling in establishing the hemilineages of secondary neurons in Drosophila melanogaster. Development 137:53–61
Tsuji T, Hasegawa E, Isshiki T (2008) Neuroblast entry into quiescence is regulated intrinsically by the combined action of spatial Hox proteins and temporal identity factors. Development 135:3859–3869
Uemura T, Shepherd S, Ackerman L, Jan LY, Jan YN (1989) numb, a gene required in determination of cell fate during sensory organ formation in Drosophila embryos. Cell 58:349–360
Urbach R, Technau GM (2003) Molecular markers for identified neuroblasts in the developing brain of Drosophila. Development 130:3621–3637
Urbach R, Technau GM (2004) Neuroblast formation and patterning during early brain development in Drosophila. BioEssays 26:739–751
Viktorin G, Riebli N, Popkova A, Giangrande A, Reichert H (2011) Multipotent neural stem cells generate glial cells of the central complex through transit amplifying intermediate progenitors in Drosophila brain development. Dev Biol 356:553–565
Viktorin G, Riebli N, Reichert H (2013) A multipotent transit-amplifying neuroblast lineage in the central brain gives rise to optic lobe glial cells in Drosophila. Dev Biol 379:182–194
Wang H, Cai Y, Chia W, Yang X (2006a) Drosophila homologs of mammalian TNF/TNFR-related molecules regulate segregation of Miranda/Prospero in neuroblasts. EMBO J 25:5783–5793
Wang H, Ouyang Y, Somers WG, Chia W, Lu B (2007) Polo inhibits progenitor self-renewal and regulates Numb asymmetry by phosphorylating Pon. Nature 449:96–100
Wang H, Somers GW, Bashirullah A, Heberlein U, Yu F, Chia W (2006b) Aurora-A acts as a tumor suppressor and regulates self-renewal of Drosophila neuroblasts. Genes Dev 20:3453–3463
Weng M, Golden KL, Lee CY (2010) dFezf/Earmuff maintains the restricted developmental potential of intermediate neural progenitors in Drosophila. Dev Cell 18:126–135
Weng M, Lee CY (2011) Keeping neural progenitor cells on a short leash during Drosophila neurogenesis. Curr Opin Neurobiol 21:36–42
Wirtz-Peitz F, Nishimura T, Knoblich JA (2008) Linking cell cycle to asymmetric division: Aurora-A phosphorylates the Par complex to regulate Numb localization. Cell 135:161–173
Wu PS, Egger B, Brand AH (2008) Asymmetric stem cell division: lessons from Drosophila. Semin Cell Dev Biol 19:283–293
Xiao Q, Komori H, Lee CY (2012) klumpfuss distinguishes stem cells from progenitor cells during asymmetric neuroblast division. Development 139:2670–2680
Yamanaka T, Horikoshi Y, Izumi N, Suzuki A, Mizuno K, Ohno S (2006) Lgl mediates apical domain disassembly by suppressing the PAR-3-aPKC-PAR-6 complex to orient apical membrane polarity. J Cell Sci 119:2107–2118
Yasugi S, Mizuno T (2008) Molecular analysis of endoderm regionalization. Dev Growth Differ 50(Suppl 1):S79–96
Yasugi T, Umetsu D, Murakami S, Sato M, Tabata T (2008) Drosophila optic lobe neuroblasts triggered by a wave of proneural gene expression that is negatively regulated by JAK/STAT. Development 135:1471–1480
Younossi-Hartenstein A, Nassif C, Green P, Hartenstein V (1996) Early neurogenesis of the Drosophila brain. J Comp Neurol 370:313–329
Zhong W, Chia W (2008) Neurogenesis and asymmetric cell division. Curr Opin Neurobiol 18:4–11
Zhong W, Feder JN, Jiang MM, Jan LY, Jan YN (1996) Asymmetric localization of a mammalian numb homolog during mouse cortical neurogenesis. Neuron 17:43–53
Zhu S, Lin S, Kao CF, Awasaki T, Chiang AS, Lee T (2006) Gradients of the Drosophila Chinmo BTB-zinc finger protein govern neuronal temporal identity. Cell 127:409–422
Acknowledgments
We thank Yanrui Jiang for critical reading of the manuscript. This work was supported by grants from the Swiss National Research Program 63 (4063 L 128006) and the Swiss National Science Foundation (31003A 140607) as well as by grants from the Global Research Laboratory Program (NRF-2009-00424), Brain Research Program (NRF-2009-0081465), and Stem Cell Research Program (NRF-2006-2004289) of the Korean Ministry of Science, ICT, and Future Planning (MSIP).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Kang, K.H., Reichert, H. Control of neural stem cell self-renewal and differentiation in Drosophila . Cell Tissue Res 359, 33–45 (2015). https://doi.org/10.1007/s00441-014-1914-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00441-014-1914-9