Abstract
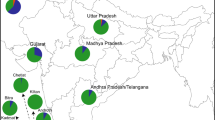
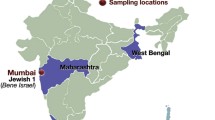
Migration and admixture history of populations have always been curious and an interesting theme. The West Coast of India harbours a rich diversity, bestowing various ethno-linguistic groups, with many of them having well-documented history of migrations. The Roman Catholic is one such distinct group, whose origin was much debated. While some historians and anthropologists relating them to ancient group of Gaud Saraswat Brahmins, others relating them for being members of the Jews Lost Tribes in the first Century migration to India. Historical records suggests that this community was later forcibly converted to Christianity by the Portuguese in Goa during the Sixteenth Century. Till date, no genetic study was done on this group to infer their origin and genetic affinity. Hence, we analysed 110 Roman Catholics from three different locations of West Coast of India including Goa, Kumta and Mangalore using both uniparental and autosomal markers to understand their genetic history. We found that the Roman Catholics have close affinity with the Indo-European linguistic groups, particularly Brahmins. Additionally, we detected genetic signal of Jews in the linkage disequilibrium-based admixture analysis, which was absent in other Indo–European populations, who are inhabited in the same geographical regions. Haplotype-based analysis suggests that the Roman Catholics consist of South Asian-specific ancestry and showed high drift. Ancestry-specific historical population size estimation points to a possible bottleneck around the time of Goan inquisition (fifteenth century). Analysis of the Roman Catholics data along with ancient DNA data of Neolithic and bronze age revealed that the Roman Catholics fits well in a basic model of ancient ancestral composition, typical of most of the Indo-European caste groups of India. Mitochondrial DNA (mtDNA) analysis suggests that most of the Roman Catholics have aboriginal Indian maternal genetic ancestry; while the Y chromosomal DNA analysis indicates high frequency of R1a lineage, which is predominant in groups with higher ancestral North Indian (ANI) component. Therefore, we conclude that the Roman Catholics of Goa, Kumta and Mangalore regions are the remnants of very early lineages of Brahmin community of India, having Indo-Europeans genetic affinity along with cryptic Jewish admixture, which needs to be explored further.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The West Coast India extends from the northern part, the State of Gujarat to the southernmost part of India, the State of Kerala. This narrow stretch of landmass between the Western ghats in east and the Arabian sea in the West is a place of rich cultural and ethno-linguistic diversity. Since ancient times, major evolutionary forces have left their footprints in the genomes of populations that are inhabiting in West Coast. From the arrival of the first modern human to Indian subcontinent from Africa ~ 65 ka (Forster and Matsumura 2005; Kivisild et al. 2003; Thangaraj et al. 2005) to the migration of the inhabitants of Indus Valley civilization after its degradation (Dutt et al. 2018) further southwards and eastwards in the form of various tribes and finally to the integration of Steppe ancestry into genome in the form of ANI-ASI admixture (Moorjani et al. 2013; Narasimhan et al. 2019; Reich et al. 2009), all these events framed the basic genetic layers of ancestry in the human populations of this region. Apart from these major migration and admixture events, populations of this region came in contact with different groups, including the Jews, the Parsees and the Muslims who were traders from Middle East and with Europeans. Among these groups, Parsees and Jews of the West Coast have been previously studied (Behar et al. 2010; Chaubey et al. 2016, 2017; Cohen et al. 1980). But, still the origin and affinity of many ethnologically and culturally distinct groups living in this region since a long time remains poorly resolved and debatable. The Roman Catholics is one of the major communities, for which many Schools of thoughts have emerged from time to time about their origin, however, these are mainly emphasizing either historical or anthropological aspects only.
Although most of the written records of the origin and culture of the groups converted to Roman Catholicism were destroyed during Portuguese rule, especially in the sixteenth century (Conlon et al. 1977), still oral tradition along with cultural and linguistic affinity of this community relates its origin from Gaud Saraswat Brahmin, whose origin in turn, according to some school of thoughts, is near the mythical Saraswathi river region. Many scholarly evidences point to their arrival to Konkan coast from Patliputra and Bengal (Conlon et al. 1977; Farias 1999; Keṇī and Murgaon Mutt Sankul 1998), where they prospered as a major elite Brahminical group until the arrival of Portuguese to West Coast. With the Portuguese establishing themselves in Goa in 1510, thousands of these Hindus fled from Goa because of forced conversions, going southwards or northwards, and they established themselves as one of the major communities of Karnataka, Kerala and Maharashtra (Farias 1999). The Brahmins/Hindus, who remained in Goa were made to convert and formed the present Roman Catholic community but they were subjected later on to severe persecution under the dreadful “Inquisition” (Tribunal do Santo Officio) established in 1560 (Farias 1999). Due to intense torture under the ‘Rigour of Mercy’ of the Inquisition with its dreaded ‘Auto de Fe’, the Brahmin population got severely reduced and many of these newly converted Christians also fled southwards in a second wave. Maratha incursions of the Maratha Raja Shambhaji into the land as also famines, floods and the plagues were other reasons for a series of migrations out of Goa. About 5000 Christians fled to Kanara from Bardez, Salsette and Tiswadi in 1739 (Farias 1999). The only genetic study with group related to Roman Catholics done till date was on comparatively broader groups of Gaud Saraswat Brahmin using classical genetic markers of ABO and Rhesus blood groups (Bhatia HF—Shanbhag et al. 1976). Hence, to get deeper insight into the origin and affinity of Roman Catholics of West Coast India, we performed a detailed population genetic study using mtDNA, Y chromosomal DNA and genome wide autosomal markers.
Materials and methods
Blood samples were collected from 110 individuals from the Roman Catholic community inhabited in Goa, Kumta and Mangalore in the West Coast of India, following all ethical guideline and with informed written consent. For this project we followed approved guidelines, applied protocols and sought permission of Institute Ethical Committee of CSIR-CCMB, Hyderabad, India. All the 110 samples were sequenced for control and coding region of mtDNA and 72 male samples were genotyped for 26 Y chromosomal DNA markers, described elsewhere (Karafet et al. 2008). A total of 48 samples were genotyped for 633,994 markers on Affymetrix Human Origin array following manufacturers prescribed protocols.
Sequencing of hypervariable and control region of mtDNA was carried out using Sanger sequencing method. Sequences were edited and assembled with the revised Cambridge reference sequence (rCRS) (Andrews et al. 1999) using AutoAssembler. Variations observed were used to assign the haplogroup using phylotree build 17 (van Oven and Kayser 2009) and Haplogrep2 (Weissensteiner et al. 2016). For Y chromosomal DNA analysis, genotyping was done for 26 biallelic markers (Supplementary Table 1a) and sequences were compared with reference sequences to mark the variations for assigning the haplogroups of 72 male individuals.
For analysis with autosomal markers, we filtered genotyped markers using Plink 1.9 (Purcell et al. 2007), to include only those markers having genotyping success of > 99% and minor allele frequency > 1%. We filtered the related individuals in our dataset by removing one individual from first-degree and second-degree relative pair utilizing KING-robust (Manichaikul et al. 2010) feature implemented in Plink2 (Chang et al. 2015). To prepare the data for Principal Component Analysis and Admixture (Alexander et al. 2009) analysis, affected by background linkage disequilibrium, we further thinned the marker set by applying LD pruning of markers with strong LD (r2 > 0.4, 200 SNPs window, sliding window of 25 SNPs at a time). LD pruned marker set comprised of 156,810 autosomal SNPs.
For comparative analysis with modern Eurasians, we merged our dataset with published DNA dataset from throughout the world (Mallick et al. 2016; Moorjani et al. 2013; Nakatsuka et al. 2017; Reich et al. 2009). The merged dataset comprised of 2984 individuals genotyped at 422,810 markers after filtering for missingness. For inferring ancient ancestral contribution to Roman Catholics, we merged our dataset with ancient DNA dataset of 765 individuals published elsewhere (Allentoft et al. 2015; Broushaki et al. 2016; Damgaard et al. 2018; de Barros Damgaard et al. 2018; Haak et al. 2015; Lazaridis et al. 2014, 2016; Mathieson et al. 2015; Meyer et al. 2012; Narasimhan et al. 2019; Raghavan et al. 2014; Yang et al. 2017).
We first performed PCA on pruned data with Smartpca package of EIGENSOFT 7.2.1 (Patterson et al. 2006) with default settings. For PCA analysis with aDNA data, we projected the ancient samples on PCA space with modern individuals with parameters lsqproject: YES and shrinkmode: YES. We used unsupervised clustering algorithm ADMIXTURE (Alexander et al. 2009) to infer global genetic contribution as ancestral components in Roman Catholics. We first performed Cross Validation for K value ranging from 3 to 14 for 25 times and found K = 4 as best suited value.
F-statistics were calculated using package AdmixTool (Patterson et al. 2012). qp3Pop (Patterson et al. 2012) was used to compute F3 admixture statistics in the form (X, Palliyar; Roman Catholics), where X is any modern South Asian or Eurasian population. We calculated F4 statistics using qpDstat (Patterson et al. 2012) with F4mode: YES, for different combinations of modern and ancient source populations.
To infer ancient ancestral contributions to Roman Catholics ancestry we also performed Distal and Proximal modelling with aDNA data using qpAdm (Haak et al. 2015) and constructed fitted graph topology using qpGraph (Patterson et al. 2012). For this, we first ran qpfstats (Patterson et al. 2012) for precomputation of F-statistics with setting allsnps: YES, and then ran qpAdm (Haak et al. 2015) and qpGraph (Patterson et al. 2012) on the outputs of qpfstats (Patterson et al. 2012). For distal modelling, as Left group we used Andamanese—hunter gatherers, Iranian—Neolithic farmers, Eastern European—hunter gatherers and Neolithic Anatolian, and as rightgroup we included Ethiopia_4500BP_published.SG, ANE, Ust_Ishim, Nautufian, Dai.DG, Belt_Cave_Mesolithic, Western European—hunter gatherers. Whereas in proximal modelling as left groups, we used AHG, Indus periphery, and Western_Steppe_MLBA/Central_Steppe_MLBA and as Right group we used Ethiopia_4500BP_published.SG, Ganj_Dareh_N, EEHG, PPNB, Dai.DG, Anatolia_N and WEHG as reference. We also calculated D statistics with different ancient sources to assess allele sharing between Roman Catholics and different ancient source populations.
We used ALDER (Loh et al. 2013) to infer admixture dates based on weighted LD statistics using modern West Eurasian and South Asian groups as source populations. We also used ChromoPainter (Lawson et al. 2012), FineStructure (Lawson et al. 2012) and GlobeTrotter (Hellenthal et al. 2014) to infer fine level population clustering and admixture timings using haplotype-based approach. We first phased our data with SHAPEIT4 (Delaneau et al. 2019) using default parameters. Chromosome painting was performed using ChromoPainter (Lawson et al. 2012), first by doing 10 EM iteration with five randomly selected chromosomes with subset of surrogate and target individuals to infer global mutation rate (µ) and switch rate parameters (Ne). Then, we ran two separate ChromoPainter (Lawson et al. 2012) run based on downstream analysis: (i) for FineStructure (Lawson et al. 2012) analysis we used ChromoPainter (Lawson et al. 2012) with a switch to paint all recipients with all donor haplotypes (ii) for GlobeTrotter (Hellenthal et al. 2014) run we used ChromoPainter (Lawson et al. 2012) to paint target and surrogate individuals with all donor haplotypes individuals using estimated fixed value of global mutation rate and switch rate parameters. Then, using combined output of ChromoPainter (Lawson et al. 2012), we performed our main FineStructure (Lawson et al. 2012) and GlobeTrotter (Hellenthal et al. 2014) using default parameter described by the authors.
We used IBDne (Browning and Browning 2015) to infer ancestry-specific historical effective population size (Ne). For this, we phased our data with Beagle5.1 (Browning et al. 2018) and ran Refined-IBD (Browning and Browning 2013) to detect IBD segments and gaps were filled by Merge-IBD-segments utility in Beagle. Then individual ancestry was assigned with RFMix v-1.5.4 (Maples et al. 2013) and IBDne (Browning and Browning 2015) was run using combined data across chromosomes by setting 2 cM IBD length as threshold.
Maximum Likelihood Tree was generated using TreeMix v.1.12 (Pickrell and Pritchard 2012) to infer placement of Roman Catholics in the global context. Runs of Homozygosity was performed using PLINK v1.9 (Purcell et al. 2007). We used three different homozygous windows of 1000 kb, 2500 kb and 5000 kb with minimum 50 consecutive SNPs. All plots were done using R statistical package.
Results
Frequency-based clustering and admixture
Principal Component Analysis (PCA) using Smartpca (Patterson et al. 2006) on the LD pruned dataset places all of the Roman Catholics (from Goa, Kumta and Mangalore) on the South Asian cline, the majority among Indo-Europeans with a few of them also clustering among Dravidian groups (Fig. 1a). Whereas, in PCA with aDNA reference, Roman Catholics samples were subdivided into three clusters (Fig S1), one of these clusters was not among but closer to modern North West Indian and North Indian Indo-European groups clustering together among Swat Primitive Grave Type Iron Age individuals. Second cluster was among modern Indo-Europeans having moderate ANI ancestry and along with some Indus Periphery individuals. Third cluster was completely among Dravidian and some Indus Periphery individuals. Thus, suggesting that the Roman Catholics of West Coast represents a genetically heterogenous group having varying proportions of Steppe and Ancestral South Indian-related ancestry.
PCA, ADMIXTURE and Admixture F3 analysis. A Biplot of principal component analysis (PCA) of Roman Catholics with modern Eurasian populations with first two components. Inset is biplot of the population averages of first two principal components. B Stacked barplot of the ADMIXTURE analysis with K = 4 with global populations ordered geographically. C Admixture F3 (X, Palliyar; Roman Catholic) gradient map, showing the affinity of Roman Catholics with West Eurasian and South Asian population groups. Greener colour means more affinity and yellow, orange and red indicate moderate, lesser and least affinity, respectively
To get a clearer picture of ancestral components, we used model-based Admixture (Alexander et al. 2009) algorithm. At the best K value of 4, we found that the Roman Catholics carry South Asian ancestral components in a proportion intermediate to extreme Indo-European groups (having higher ANI components) from Pakistan as well as North India and Dravidians (Fig. 1b).
We further proceeded with the Admixture F3 statistics using qp3Pop implementation in AdmixTool (Patterson et al. 2012). We used Palliyar as proxy to ASI as described elsewhere (Narasimhan et al. 2019) and other West Eurasian and South Asian populations as second source population. We found that Roman Catholics show higher admixture F3 with most of West Eurasian population groups, a trend similar to Indo-European groups (Fig. 1c) and also D-statistics with west Eurasian sources using qpDstat was also consistent with PCA and Admixture findings. Maximum Likelihood tree constructed using TreeMix v.1.12 (Pickrell and Pritchard 2012) shows that Roman Catholics from Goa, Kumta and Mangalore are placed between Indian Indo-European caste groups and Dravidian caste groups along with Havik and Karnataka Brahmins (Fig S2).
Linkage disequilibrium-based admixture analysis
We used weighted linkage disequilibrium based inference of admixture timing implemented in ALDER (Loh et al. 2013). We further used Indian Austroasiatic groups as proxy for South Asian ancestry as Austroasiatic speakers were already inhabited in India, much before the arrival of Steppe ancestry or Iranian farmer related ancestry. In other words, they have relatively very low proportions of west Eurasian ancestry (Narasimhan et al. 2019) and our implementation of ALDER was to query on west Eurasian admixture and its timing, therefore we used Austroasiatic speakers for Indian source group of AASI (Ancient Ancestral South Indian) ancestry. This was also implemented in a recent study (Pathak et al. 2018). We tested for timing of admixture of different West Eurasian components into Roman Catholic genome. Best fitted date estimates of West Eurasian admixture was around ~ 85 generations (~ 2500 years) back with French, which corresponds to introduction of Steppe ancestry (ANI component) into the subcontinent (Moorjani et al. 2013) (Table S2a). This suggests that there was no further admixture occurred during Portuguese inquisition. While testing for different West Eurasian admixture events, we also found many good fits (z = 7.20, p = 6.2e−13) with Jews populations (Ashkenazi Jew and Yemenites Jew) but the estimated time was further back to ~ 94 generations (Table S2a). This time frame overlaps with something similar to what was hypothesized by some of the historians, who were working on Indian Jewish diaspora and their historical records (Slapak et al. 1995; Weil et al. 2002). As described by some historians who are working on Indian Jews diaspora (Farias 1999; Slapak et al. 1995; Weil et al. 2002), some Jewish group came to India either after capturing of Jerusalem (700 BC) or as traders during King Solomon’s reign (970 BC). Accordingly, ALDER result suggests an admixture time of 94 generations, i.e., around 2800 YBP, considering 30 years per generation, starting from 1950 AD. But all these tests were failed with Roman Catholics from Kumta and Mangalore using reference genotypes but fits were obtained using PCA loadings instead of reference genotypes for both the groups (Table S2a).
Fine level admixture inference using haplotype-based approach
To gain deeper insight into admixture history of Roman Catholics, we further applied the haplotype-based approach implemented in FineStructure (Lawson et al. 2012) and GlobeTrotter (Hellenthal et al. 2014). FineStructure (Lawson et al. 2012) grouped all individuals into 56 distinct clusters with hierarchical tree having 2 basal branch of West Eurasian groups and South Asian groups (Fig S3). A total of 23 Roman Catholics were placed in one group of 6 clusters and located between Indo-Europeans and Dravidians in South Asian clusters while the remaining individuals were seen among Dravidian clusters. We also examined the co-ancestry matrix (Fig S4), which shows that the Roman Catholics have greater intrapopulation drift than any North West or North Indian Indo-European groups but lesser than Kalash, Palliyar and Ulladan.
GlobeTrotter (Hellenthal et al. 2014) admixture analysis suggested a one date admixture (~ 75 generations before) with two sources (Supplementary Table S2b). The best guess for minority contributing first source was Ukrainian (32%) and majority contributing second source was represented by Kuruba (68%). First admixing source which contributes 32% of ancestry can be represented as mixture of Ukrainian, Georgian, Estonian, Iranian Jew, Georgian Jew, Kalash, Libyan Jew, Druze and Ashkenazi Jew with fractions contributed as 0.50, 0.31, 0.09, 0.02, 0.02, 0.01, 0.01, 0.01 and 0.005, respectively. While second admixing source contributing 68% of ancestry can be best represented by mixture of Kuruba, Sikh_Jatt, Ho_Orissa, Palliyar, Asur and Ulladan with fractions of 0.50, 0.25, 0.10, 0.05, 0.04 and 0.02, respectively. So unlike ALDER (Loh et al. 2013) results, GlobeTrotter (Hellenthal et al. 2014) run suggests a one date admixture event representing typical of ANI-ASI admixture.
Ancestry-specific historical effective population size (Ne) and runs of Homozygosity
We used IBDne (Browning and Browning 2015) to trace the ancestry-specific historical population size dynamics with time in Roman Catholic population. Historical records suggest that during Portuguese colonisation in the West Coast and the Goan inquisition, the Roman Catholics faced severe torture and even extermination (by ‘Auto-de-Fe’: means burning at the stake) (Farias 1999) and hence, possibility of population bottleneck. In our analysis, we found that two of the three-ancestry curves (European and Ancestral South Indian) showed a depression near 15–20 generations before present (Fig. 2). Although bootstrap confidence interval (95% CI) is slightly broader, which can be either because of small sample size or small bottleneck event as mentioned in an earlier study (Browning and Browning 2015). Small bottleneck results from many post-bottleneck coalescences between haplotypes. So, this can be a probable bottleneck with significant change in population size of Roman Catholics during the event of Goan inquisition.
Ancestry-specific effective population size estimation. Y-axis show ancestry specific effective population size (Ne) on log scale. X-axis show generations before present. Blue solid line shows estimated ancestry-specific effective population sizes, and the blue shaded region indicates 95% bootstrap confidence intervals
We also performed Runs of homozygosity (RoH) test using Plink 1.9 (Purcell et al. 2007) for three different window sizes of 1000 kb, 2500 kb and 5000 kb. Comparing the mean RoH length to mean RoH number, we found that Roman Catholics from all three places have higher distribution than most of the Indo Europeans from North India (Fig S5a–c). Among three groups of Roman Catholics, Kumta was higher in mean length and mean number of RoH distribution. This suggests that Roman Catholics from Goa and Mangalore had high level of gene flow from surrounding populations than Kumta, with later being more isolated as a population. Our interpretation of Kumta Catholics being comparatively isolated group than other two is based on RoH analysis which tells that Kumta group is comparatively more inbreeding (thus lesser gene flow) than other two groups but unfortunately frequency-based methods like PCA and Admixture analysis failed to capture this and we could not find similar finding in these methods. Here, we followed the interpretation made in a previous study (Pathak et al 2018), which also implemented RoH method.
Admixture modelling with ancient sources
Further, we performed admixture modelling with distant and proximal ancient sources using qpAdm (Haak et al. 2015) and tried to fit our population group in the basic framework of South Asian-specific graph topology using qpGraph (Patterson et al. 2012). For distal modelling, we used Neolithic and pre-Copper Age West Eurasian and South Asian populations (Andamanese hunter gatherers, Ganj_Dareh_N, Anatolia_N, EEHG, WEHG and WSHG) from which South Asia derives most of their ancestry. We found that best fit was obtained for Goan (p = 0.25), Kumta (p = 0.39), and Mangalorean Catholics (p = 0.53) (Supplementary Table 2c) with a combination of AHG, Ganj_Dareh_N, Anatolia_N and EEHG (Fig. 3a). In proximal modelling we found that best fit was with AHG, Indus_Periphery and Central Steppe MLBA/Western Steppe MLBA as sources of ancestry for Goan (p = 0.59), Kumta (p = 0.77) and Mangalorean Catholics (p = 0.9) (Fig. 3b–c) (Supplementary Table 2d, e).
Admixture modelling to infer contribution of ancient ancestral west Eurasian source populations. A Distal modelling using AHG, Iran_N, Anatolia_N and EEHG as distant source groups of Goan, Kumta and Mangalorian catholics and their comparison with four modern Indo–European populations. Each colour represents individual distant ancestral source and numbe indicates individual ancestral contribution in terms of fraction of total ancestry. B Proximal modelling with AHG, Indus_Periphery and western Steppe MLBA groups. C Proximal modelling with AHG, Indus_Periphery and Central Steppe MLBA groups
To model our population into admixture graph topology using qpGraph (Patterson et al. 2012), we used the basic graph framework for South Asia shown by Narasimhan et al., 2019 (Narasimhan et al. 2019), incorporating most of the South Asian linguistics groups. We obtained the fitted graph toplogy for Goan (worst F_stat = − 2.896) (Fig. S6) and Mangalorean Catholics (worst F_stat = 2.831) (Fig. S7) but for Kumta Catholic the fit was not obtained with the above framework (worst F_stat = 3.134) (Fig. S8).
Distribution of mtDNA and Y chromosomal haplogroups
mtDNA analysis using hypervariable and coding region sequences reveals four major mtDNA haplogroups; M37, M30, M2 and M52, in decreasing order of frequency (Supplementary Table1c). The M37 was with the highest frequency of 12% in the population, which is the most prevalent haplogroup in Gujarat. The M2 is the oldest Indian-specific mtDNA haplogroup. Less frequency of mtDNA haplogroup observed were; M33, U2b, M5a, M35, M57 and M64. Of these, M64 is exclusively present in the Nihali population of Madhya Pradesh, which is a linguistic isolate group in India.
Among Y chromosomal haplogroups, the most prevalent was R1a with a frequency of 46%, while the second most frequent haplogroups were L1a (15%) and J2 (15%), followed by H2 (13%) (Supplementary Table 1b). High prevalence of R1a haplogroup and significantly higher z-score (2-ref z-score > 7) for west Eurasian groups corroborates with their origin from ancestral Brahmin population of North and North West India (Cordaux et al. 2004; Sharma et al. 2009; Zerjal et al. 2007). Studies of fine mapping of R1a phylogeny with downstream markers will better reveal their origin and affinities.
Discussion
According to the historical records and oral traditions, Roman Catholics were part of Konkani speaking larger Gaud Saraswat community before arrival of Portuguese to the West Coast. They were inhabiting the Konkan coast along with some other Hindu, Jewish and Muslim communities. With Portuguese ascendancy with the goal of forming the Estado d’India in the sixteenth century, most of these people were compelled to convert to Christianity and many of them either got penalised if they did not convert or escaped by fleeing southwards and admixed with local populations (Farias 1999; Keṇī and Murgaon Mutt Sankul 1998). One School of thought relates the GSB to some Jews of the early lost tribes of Israel, who came to India as refugees after the capture and destruction of Jerusalem by Assyrians in BC 722 (Weil et al. 2002). Others relate this migration of those who came as traders in King Solomon’s times (BC 977-937) when the Jews came by the sea route to the West Coast of India and had arrived in Kerala (Farias 1999). The later Bene Israel Jews of Maharashtra came in BC 175—while others say they came in 70 AD (Schreiber et al. 2003; Slapak et al. 1995). Later some of these migrant groups also got converted to Christianity with the arrival of Portuguese to West Coast in the sixteenth century (Farias 1999). So, these groups may be part of broader Roman Catholic community of the Konkan coast.
We are able to show in PCA (Patterson et al. 2006) (Fig. 1a) and Admixture (Alexander et al. 2009) (Fig. 1b) analysis that Roman Catholics shares South Asian ancestry and that they show affinity with Indian Indo-European populations. In Admixture (Alexander et al. 2009) analysis, Roman Catholics shows two major components (ANI and ASI) which is present in most of the Indo-Europeans and Dravidian castes and tribes. Only proportions of these two major components vary among Indo Europeans and Dravidians. Among three Roman Catholic groups from Goa, Kumta and Mangalore also these two components vary in proportions. Some of the Goan Catholics individuals are similar to Indo–Europeans, showing more yellow (representing ANI ancestry) component than red (representing ASI signal) in Admixture analysis, but these proportions vary among other individuals from the three places, indicating some Dravidian admixture with local groups after their arrival to the West Coast. Formal test of admixture with F3-statistics (Patterson et al. 2012) (Fig. 1c) suggests that Roman Catholics have higher West Eurasian-specific admixture which is reflected in higher affinity with European, Middle Eastern, North West Indian and North Indo–European caste groups.
In addition to Indo–European-specific ancestry, reflected in Admixture (Alexander et al. 2009) and F3-statistics (Patterson et al. 2012), we further tested the hypothesis of dubious origin of Roman Catholics from Jewish community of the Konkan Coast using linkage disequilibrium-based method implemented in ALDER (Loh et al. 2013). This test of admixture showed indications of Roman Catholics having some Jewish component as reflected by a significant good fit for weighted LD decay curve with Ashkenazi Jews and Yemenite Jews, which we could not find in other Indo Europeans (UP_Brahmins, Bhumihar, Sikh_Jatts Haryana Brahmins, Karnataka Brahmins and Havik) (Supplementary Table 2a). Admixture dates correspond to the arrival of the first Jews to the West Coast as traders during King Solomon’s reign (BC 977-937) (Weil et al. 2002).
We further tested the above finding with haplotype-based methods implemented in FineStructure (Lawson et al. 2012), which captured mostly South Asia-specific ancestry among Roman Catholics, with significant intrapopulation genetic drift higher than most of the South Asian caste and tribal groups. This drift was lesser than populations showing very high genetic drift due to long-term isolation, such as Kalash, Ulladan and Palliyar (Supplementary Fig. S4). Analysis of time of admixture with GlobeTrotter (Hellenthal et al. 2014) suggests only one date of admixture for ANI ancestry, ~ 75 generation ago (Supplementary Table 2b). Keeping the differences observed in the above two approaches in mind, we could not come up with a clear and conclusive evidence for Jewish admixture, because of the complete absence of signal in haplotype-based analysis. Further evidence can be gathered only with more robust admixture modelling approach using proper aDNA reference for a homogenous ancient Jews diaspora, as most of the contemporary Jews population are having majority of either Middle Eastern or European admixture.
We further tested the presence of possible population bottleneck in Roman Catholic population based on historical records of their persecution and further migration during the Goan inquisition (Farias 1999; Keṇī and Murgaon Mutt Sankul 1998). Ancestry-specific historical population size estimates with IBDne (Browning and Browning 2015) showed a depression about 15 generations ago (Fig. 2), which corroborates with the significant reduction in population size during the Goan inquisition under Portuguese invasion to West Coast. During this time, there were many waves of migrations of Roman Catholics population further southwards (Keṇī and Murgaon Mutt Sankul 1998) and admixed with local groups. Runs of homozygosity suggested that among three groups of Roman Catholics, Kumta group showed comparatively higher distribution for mean length and mean number of RoH segments, indicating that they remained more isolated compared to other Catholic groups during all migration waves and admixture events.
While estimating the ancient contribution to the Roman Catholics genome using qpAdm (Haak et al. 2015), we found that they exhibit an ancient ancestral composition similar to most of the Indo-European caste and tribal groups, formed after arrival of Steppe ancestry (Supplementary Table 2c–e) (Fig. 3a–c). But the proportion of Steppe component was lesser than most of the North Indian groups with high ANI component (Sikh_Jatt, Brahmins), which may be a result of later admixture events. We used similar graph topology as used for Indian populations in previous study of Narasimhan et al. (2019), but model for Kumta group was not fitted (or marginally fit with worst F_stat of 3.134) in topology, probably because of more level of population-specific drift (Supplementary Fig. S6–8).
Analysis of uniparental markers, revealed that maternal ancestry of Roman Catholics is completely of South Asian-specific with most of the mtDNA haplogroups of Indian origin (Supplementary Table 1c). But their Y chromosomal haplogroup distribution was highly skewed with significantly higher frequency of R1a (Supplementary Table 1b), which is typical of North West Indian or North Indian Indo-European groups, like the caste with priestly status such as Brahmins (Cordaux et al. 2004; Sharma et al. 2009; Zerjal et al. 2007) having high proportion of ANI ancestry (Narasimhan et al. 2019). We also obtained similar pattern with Gaud Saraswat Brahmin from West Coast, from which Roman Catholics share their ancestry (unpublished data). Further, downstream genotyping of R1a markers in future study will shed light on their origin from specific group and a probable Jewish component in their ancestry. The present level of analysis with uniparental markers and genome wide autosomal SNP markers, suggest that they have male biased admixture with the arrival of R1a carrying population and also genetic composition typical of Indo-European populations on the Indian cline, which holds true with all of the populations with affinity with priestly groups on ANI-ASI cline in India. They also exhibit cryptic genetic affinity with some Jews groups like Ashkenazi Jews in our test of admixture using west Eurasian references, which needs to be further validated.
Availability of data and material
Data have been uploaded in the CCMB server http://tdb.ccmb.res.in/bic/database_pagelink.php?page=wcdata and would be made available for the researchers upon request to the corresponding authors.
References
Alexander DH, Novembre J, Lange K (2009) Fast model-based estimation of ancestry in unrelated individuals. Genome Res 19:1655–1664. https://doi.org/10.1101/gr.094052.109
Allentoft ME, Sikora M, Sjogren KG, Rasmussen S, Rasmussen M, Stenderup J, Damgaard PB, Schroeder H, Ahlstrom T, Vinner L, Malaspinas AS, Margaryan A, Higham T, Chivall D, Lynnerup N, Harvig L, Baron J, Della Casa P, Dabrowski P, Duffy PR, Ebel AV, Epimakhov A, Frei K, Furmanek M, Gralak T, Gromov A, Gronkiewicz S, Grupe G, Hajdu T, Jarysz R, Khartanovich V, Khokhlov A, Kiss V, Kolar J, Kriiska A, Lasak I, Longhi C, McGlynn G, Merkevicius A, Merkyte I, Metspalu M, Mkrtchyan R, Moiseyev V, Paja L, Palfi G, Pokutta D, Pospieszny L, Price TD, Saag L, Sablin M, Shishlina N, Smrcka V, Soenov VI, Szeverenyi V, Toth G, Trifanova SV, Varul L, Vicze M, Yepiskoposyan L, Zhitenev V, Orlando L, Sicheritz-Ponten T, Brunak S, Nielsen R, Kristiansen K, Willerslev E (2015) Population genomics of Bronze Age Eurasia. Nature 522:167–172. https://doi.org/10.1038/nature14507
Andrews RM, Kubacka I, Chinnery PF, Lightowlers RN, Turnbull DM, Howell N (1999) Reanalysis and revision of the Cambridge reference sequence for human mitochondrial DNA. Nat Genet 23:147–147. https://doi.org/10.1038/13779
Behar DM, Yunusbayev B, Metspalu M, Metspalu E, Rosset S, Parik J, Rootsi S, Chaubey G, Kutuev I, Yudkovsky G, Khusnutdinova EK, Balanovsky O, Semino O, Pereira L, Comas D, Gurwitz D, Bonne-Tamir B, Parfitt T, Hammer MF, Skorecki K, Villems R (2010) The genome-wide structure of the Jewish people. Nature 466:238–242. https://doi.org/10.1038/nature09103
Bhatia HF, Shanbhag SR, Baxi AJ, Baxi AF, Bapat J, Sathe MS, Sharma RS, Kabeer H, Bharucha ZS, Surlacar L (1976) Genetic studies among endogamous groups of Saraswats in Western India. Hum Hered 26:458
Broushaki F, Thomas MG, Link V, Lopez S, van Dorp L, Kirsanow K, Hofmanova Z, Diekmann Y, Cassidy LM, Diez-Del-Molino D, Kousathanas A, Sell C, Robson HK, Martiniano R, Blocher J, Scheu A, Kreutzer S, Bollongino R, Bobo D, Davudi H, Munoz O, Currat M, Abdi K, Biglari F, Craig OE, Bradley DG, Shennan S, Veeramah K, Mashkour M, Wegmann D, Hellenthal G, Burger J (2016) Early neolithic genomes from the eastern fertile crescent. Science 353:499–503. https://doi.org/10.1126/science.aaf7943
Browning BL, Browning SR (2013) Improving the accuracy and efficiency of identity-by-descent detection in population data. Genetics 194:459–471. https://doi.org/10.1534/genetics.113.150029
Browning SR, Browning BL (2015) Accurate non-parametric estimation of recent effective population size from segments of identity by descent. Am J Hum Genet 97:404–418. https://doi.org/10.1016/j.ajhg.2015.07.012
Browning BL, Zhou Y, Browning SR (2018) A one-penny imputed genome from next-generation reference panels. Am J Hum Genet 103:338–348. https://doi.org/10.1016/j.ajhg.2018.07.015
Chang CC, Chow CC, Tellier LC, Vattikuti S, Purcell SM, Lee JJ (2015) Second-generation PLINK: rising to the challenge of larger and richer datasets. Gigascience 4:7. https://doi.org/10.1186/s13742-015-0047-8
Chaubey G, Singh M, Rai N, Kariappa M, Singh K, Singh A, Pratap Singh D, Tamang R, Selvi Rani D, Reddy AG, Kumar Singh V, Singh L, Thangaraj K (2016) Genetic affinities of the Jewish populations of India. Sci Rep 6:19166. https://doi.org/10.1038/srep19166
Chaubey G, Ayub Q, Rai N, Prakash S, Mushrif-Tripathy V, Mezzavilla M, Pathak AK, Tamang R, Firasat S, Reidla M, Karmin M, Rani DS, Reddy AG, Parik J, Metspalu E, Rootsi S, Dalal K, Khaliq S, Mehdi SQ, Singh L, Metspalu M, Kivisild T, Tyler-Smith C, Villems R, Thangaraj K (2017) “Like sugar in milk”: reconstructing the genetic history of the Parsi population. Genome Biol 18:ARTN110. https://doi.org/10.1186/s13059-017-1244-9
Cohen T, Levene C, Yodfat Y, Fidel J, Friedlander Y, Steinberg AG, Brautbar C (1980) Genetic studies on Cochin Jews in Israel: 1. Population data, blood groups, isoenzymes, and HLA determinants. Am J Med Genet 6:61–73. https://doi.org/10.1002/ajmg.1320060106
Conlon FF, Conlon FF, South Cf, studies SA, University of California BCfS, Studies SA (1977) A caste in a changing world: the Chitrapur Saraswat Brahmans, 1700–1935. University of California Press
Cordaux R, Aunger R, Bentley G, Nasidze I, Sirajuddin SM, Stoneking M (2004) Independent origins of Indian caste and tribal paternal lineages. Curr Biol 14:231–235. https://doi.org/10.1016/j.cub.2004.01.024
Damgaard PB, Marchi N, Rasmussen S, Peyrot M, Renaud G, Korneliussen T, Moreno-Mayar JV, Pedersen MW, Goldberg A, Usmanova E, Baimukhanov N, Loman V, Hedeager L, Pedersen AG, Nielsen K, Afanasiev G, Akmatov K, Aldashev A, Alpaslan A, Baimbetov G, Bazaliiskii VI, Beisenov A, Boldbaatar B, Boldgiv B, Dorzhu C, Ellingvag S, Erdenebaatar D, Dajani R, Dmitriev E, Evdokimov V, Frei KM, Gromov A, Goryachev A, Hakonarson H, Hegay T, Khachatryan Z, Khaskhanov R, Kitov E, Kolbina A, Kubatbek T, Kukushkin A, Kukushkin I, Lau N, Margaryan A, Merkyte I, Mertz IV, Mertz VK, Mijiddorj E, Moiyesev V, Mukhtarova G, Nurmukhanbetov B, Orozbekova Z, Panyushkina I, Pieta K, Smrcka V, Shevnina I, Logvin A, Sjogren KG, Stolcova T, Taravella AM, Tashbaeva K, Tkachev A, Tulegenov T, Voyakin D, Yepiskoposyan L, Undrakhbold S, Varfolomeev V, Weber A, Wilson Sayres MA, Kradin N, Allentoft ME, Orlando L, Nielsen R, Sikora M, Heyer E, Kristiansen K, Willerslev E (2018) 137 ancient human genomes from across the Eurasian steppes. Nature 557:369–374. https://doi.org/10.1038/s41586-018-0094-2
de Barros DP, Martiniano R, Kamm J, Moreno-Mayar JV, Kroonen G, Peyrot M, Barjamovic G, Rasmussen S, Zacho C, Baimukhanov N, Zaibert V, Merz V, Biddanda A, Merz I, Loman V, Evdokimov V, Usmanova E, Hemphill B, Seguin-Orlando A, Yediay FE, Ullah I, Sjogren KG, Iversen KH, Choin J, de la Fuente C, Ilardo M, Schroeder H, Moiseyev V, Gromov A, Polyakov A, Omura S, Senyurt SY, Ahmad H, McKenzie C, Margaryan A, Hameed A, Samad A, Gul N, Khokhar MH, Goriunova OI, Bazaliiskii VI, Novembre J, Weber AW, Orlando L, Allentoft ME, Nielsen R, Kristiansen K, Sikora M, Outram AK, Durbin R, Willerslev E (2018) The first horse herders and the impact of early Bronze Age steppe expansions into Asia. Science 360:eaar7711. https://doi.org/10.1126/science.aar7711
Delaneau O, Zagury JF, Robinson MR, Marchini JL, Dermitzakis ET (2019) Accurate, scalable and integrative haplotype estimation. Nat Commun 10:5436. https://doi.org/10.1038/s41467-019-13225-y
Dutt S, Gupta AK, Wünnemann B, Yan D (2018) A long arid interlude in the Indian summer monsoon during ∼4350 to 3450 cal. yr BP contemporaneous to displacement of the Indus valley civilization. Quat Int 482:83–92. https://doi.org/10.1016/j.quaint.2018.04.005
Farias KK (1999) The Christian impact in south Kanara. Church History Association of India. ISBN: 9788175251267
Forster P, Matsumura S (2005) Evolution. Did early humans go north or south? Science 308:965–966. https://doi.org/10.1126/science.1113261
Haak W, Lazaridis I, Patterson N, Rohland N, Mallick S, Llamas B, Brandt G, Nordenfelt S, Harney E, Stewardson K, Fu QM, Mittnik A, Banffy E, Economou C, Francken M, Friederich S, Pena RG, Hallgren F, Khartanovich V, Khokhlov A, Kunst M, Kuznetsov P, Meller H, Mochalov O, Moiseyev V, Nicklisch N, Pichler SL, Risch R, Guerra MAR, Roth C, Szecsenyi-Nagy A, Wahl J, Meyer M, Krause J, Brown D, Anthony D, Cooper A, Alt KW, Reich D (2015) Massive migration from the steppe was a source for Indo–European languages in Europe. Nature 522:207. https://doi.org/10.1038/nature14317
Hellenthal G, Busby GBJ, Band G, Wilson JF, Capelli C, Falush D, Myers S (2014) A genetic atlas of human admixture history. Science 343:747–751. https://doi.org/10.1126/science.1243518
Karafet TM, Mendez FL, Meilerman MB, Underhill PA, Zegura SL, Hammer MF (2008) New binary polymorphisms reshape and increase resolution of the human Y chromosomal haplogroup tree. Genome Res 18:830–838. https://doi.org/10.1101/gr.7172008
Keṇī C, Murgaon Mutt Sankul S (1998) Saraswats in Goa and beyond. Murgaon Mutt Sankul Samiti, Vasco-da-Gama, Goa
Kivisild T, Rootsi S, Metspalu M, Mastana S, Kaldma K, Parik J, Metspalu E, Adojaan M, Tolk HV, Stepanov V, Gölge M, Usanga E, Papiha SS, Cinnioğlu C, King R, Cavalli-Sforza L, Underhill PA, Villems R (2003) The genetic heritage of the earliest settlers persists both in Indian tribal and caste populations. Am J Hum Genet 72:313–332. https://doi.org/10.1086/346068
Lawson DJ, Hellenthal G, Myers S, Falush D (2012) Inference of population structure using dense haplotype data. PLoS Genet 8:ARTNe1002453. https://doi.org/10.1371/journal.pgen.1002453
Lazaridis I, Patterson N, Mittnik A, Renaud G, Mallick S, Kirsanow K, Sudmant PH, Schraiber JG, Castellano S, Lipson M, Berger B, Economou C, Bollongino R, Fu Q, Bos KI, Nordenfelt S, Li H, de Filippo C, Prufer K, Sawyer S, Posth C, Haak W, Hallgren F, Fornander E, Rohland N, Delsate D, Francken M, Guinet JM, Wahl J, Ayodo G, Babiker HA, Bailliet G, Balanovska E, Balanovsky O, Barrantes R, Bedoya G, Ben-Ami H, Bene J, Berrada F, Bravi CM, Brisighelli F, Busby GB, Cali F, Churnosov M, Cole DE, Corach D, Damba L, van Driem G, Dryomov S, Dugoujon JM, Fedorova SA, Gallego Romero I, Gubina M, Hammer M, Henn BM, Hervig T, Hodoglugil U, Jha AR, Karachanak-Yankova S, Khusainova R, Khusnutdinova E, Kittles R, Kivisild T, Klitz W, Kucinskas V, Kushniarevich A, Laredj L, Litvinov S, Loukidis T, Mahley RW, Melegh B, Metspalu E, Molina J, Mountain J, Nakkalajarvi K, Nesheva D, Nyambo T, Osipova L, Parik J, Platonov F, Posukh O, Romano V, Rothhammer F, Rudan I, Ruizbakiev R, Sahakyan H, Sajantila A, Salas A, Starikovskaya EB, Tarekegn A, Toncheva D, Turdikulova S, Uktveryte I, Utevska O, Vasquez R, Villena M, Voevoda M, Winkler CA, Yepiskoposyan L, Zalloua P et al (2014) Ancient human genomes suggest three ancestral populations for present-day Europeans. Nature 513:409–413. https://doi.org/10.1038/nature13673
Lazaridis I, Nadel D, Rollefson G, Merrett DC, Rohland N, Mallick S, Fernandes D, Novak M, Gamarra B, Sirak K, Connell S, Stewardson K, Harney E, Fu Q, Gonzalez-Fortes G, Jones ER, Roodenberg SA, Lengyel G, Bocquentin F, Gasparian B, Monge JM, Gregg M, Eshed V, Mizrahi A-S, Meiklejohn C, Gerritsen F, Bejenaru L, Blüher M, Campbell A, Cavalleri G, Comas D, Froguel P, Gilbert E, Kerr SM, Kovacs P, Krause J, McGettigan D, Merrigan M, Merriwether DA, O’Reilly S, Richards MB, Semino O, Shamoon-Pour M, Stefanescu G, Stumvoll M, Tönjes A, Torroni A, Wilson JF, Yengo L, Hovhannisyan NA, Patterson N, Pinhasi R, Reich D (2016) Genomic insights into the origin of farming in the ancient Near East. Nature 536:419–424. https://doi.org/10.1038/nature19310
Loh PR, Lipson M, Patterson N, Moorjani P, Pickrell JK, Reich D, Berger B (2013) Inferring admixture histories of human populations using linkage disequilibrium. Genetics 193:1233. https://doi.org/10.1534/genetics.112.147330
Mallick S, Li H, Lipson M, Mathieson I, Gymrek M, Racimo F, Zhao M, Chennagiri N, Nordenfelt S, Tandon A, Skoglund P, Lazaridis I, Sankararaman S, Fu Q, Rohland N, Renaud G, Erlich Y, Willems T, Gallo C, Spence JP, Song YS, Poletti G, Balloux F, van Driem G, de Knijff P, Romero IG, Jha AR, Behar DM, Bravi CM, Capelli C, Hervig T, Moreno-Estrada A, Posukh OL, Balanovska E, Balanovsky O, Karachanak-Yankova S, Sahakyan H, Toncheva D, Yepiskoposyan L, Tyler-Smith C, Xue Y, Abdullah MS, Ruiz-Linares A, Beall CM, Di Rienzo A, Jeong C, Starikovskaya EB, Metspalu E, Parik J, Villems R, Henn BM, Hodoglugil U, Mahley R, Sajantila A, Stamatoyannopoulos G, Wee JT, Khusainova R, Khusnutdinova E, Litvinov S, Ayodo G, Comas D, Hammer MF, Kivisild T, Klitz W, Winkler CA, Labuda D, Bamshad M, Jorde LB, Tishkoff SA, Watkins WS, Metspalu M, Dryomov S, Sukernik R, Singh L, Thangaraj K, Paabo S, Kelso J, Patterson N, Reich D (2016) The Simons Genome Diversity Project: 300 genomes from 142 diverse populations. Nature 538:201–206. https://doi.org/10.1038/nature18964
Manichaikul A, Mychaleckyj JC, Rich SS, Daly K, Sale M, Chen WM (2010) Robust relationship inference in genome-wide association studies. Bioinformatics 26:2867–2873. https://doi.org/10.1093/bioinformatics/btq559
Maples BK, Gravel S, Kenny EE, Bustamante CD (2013) RFMix: a discriminative modeling approach for rapid and robust local-ancestry inference. Am J Hum Genet 93:278–288. https://doi.org/10.1016/j.ajhg.2013.06.020
Mathieson I, Lazaridis I, Rohland N, Mallick S, Patterson N, Roodenberg SA, Harney E, Stewardson K, Fernandes D, Novak M, Sirak K, Gamba C, Jones ER, Llamas B, Dryomov S, Pickrell J, Arsuaga JL, de Castro JM, Carbonell E, Gerritsen F, Khokhlov A, Kuznetsov P, Lozano M, Meller H, Mochalov O, Moiseyev V, Guerra MA, Roodenberg J, Verges JM, Krause J, Cooper A, Alt KW, Brown D, Anthony D, Lalueza-Fox C, Haak W, Pinhasi R, Reich D (2015) Genome-wide patterns of selection in 230 ancient Eurasians. Nature 528:499–503. https://doi.org/10.1038/nature16152
Meyer M, Kircher M, Gansauge MT, Li H, Racimo F, Mallick S, Schraiber JG, Jay F, Prufer K, de Filippo C, Sudmant PH, Alkan C, Fu Q, Do R, Rohland N, Tandon A, Siebauer M, Green RE, Bryc K, Briggs AW, Stenzel U, Dabney J, Shendure J, Kitzman J, Hammer MF, Shunkov MV, Derevianko AP, Patterson N, Andres AM, Eichler EE, Slatkin M, Reich D, Kelso J, Paabo S (2012) A high-coverage genome sequence from an archaic Denisovan individual. Science 338:222–226. https://doi.org/10.1126/science.1224344
Moorjani P, Thangaraj K, Patterson N, Lipson M, Loh PR, Govindaraj P, Berger B, Reich D, Singh L (2013) Genetic evidence for recent population mixture in India. Am J Hum Genet 93:422–438. https://doi.org/10.1016/j.ajhg.2013.07.006
Nakatsuka N, Moorjani P, Rai N, Sarkar B, Tandon A, Patterson N, Bhavani GS, Girisha KM, Mustak MS, Srinivasan S, Kaushik A, Vahab SA, Jagadeesh SM, Satyamoorthy K, Singh L, Reich D, Thangaraj K (2017) The promise of discovering population-specific disease-associated genes in South Asia. Nat Genet 49:1403–1407. https://doi.org/10.1038/ng.3917
Narasimhan VA-O, Patterson NA-OX, Moorjani P, Rohland N, Bernardos RA-O, Mallick SA-O, Lazaridis I, Nakatsuka NA-O, Olalde I, Lipson MA-O, Kim AA-O, Olivieri LA-O, Coppa A, Vidale MA-O, Mallory J, Moiseyev V, Kitov EA-O, Monge JA-O, Adamski NA-O, Alex N, Broomandkhoshbacht NA-O, Candilio FA-O, Callan KA-O, Cheronet OA-O, Culleton BJ, Ferry MA-O, Fernandes DA-O, Freilich SA-O, Gamarra BA-OX, Gaudio DA-O, Hajdinjak M, Harney É, Harper TK, Keating D, Lawson AA-O, Mah MA-O, Mandl KA-O, Michel M, Novak MA-O, Oppenheimer JA-O, Rai N, Sirak K, Slon VA-O, Stewardson K, Zalzala FA-O, Zhang ZA-O, Akhatov G, Bagashev AA-O, Bagnera AA-O, Baitanayev BA-O, Bendezu-Sarmiento J, Bissembaev AA, Bonora GA-OX, Chargynov TT, Chikisheva TA-O, Dashkovskiy PK, Derevianko AA-O, Dobeš M, Douka K, Dubova NA-O, Duisengali MN, Enshin D, Epimakhov AA-O, Fribus AA-O, Fuller DA-OX, Goryachev A, Gromov AA-O, Grushin SA-O, Hanks BA-O, Judd MA-O, Kazizov EA-O, Khokhlov AA-O, Krygin AP, Kupriyanova E, Kuznetsov PA-O, Luiselli DA-O, Maksudov FA-O, Mamedov AM, Mamirov TB, Meiklejohn C, Merrett DA-O, Micheli R, Mochalov O, Mustafokulov SA-O, Nayak A, Pettener DA-O, Potts RA-O, Razhev D, Rykun M, Sarno S, Savenkova TM, Sikhymbaeva KA-O, Slepchenko SM, Soltobaev OA-O, Stepanova NA-O, Svyatko S, Tabaldiev KA-O, Teschler-Nicola M, Tishkin AA-OX, Tkachev VA-O et al (2019) The formation of human populations in South and Central Asia. LID. https://doi.org/10.1126/science.aat7487
Pathak AK, Kadian A, Kushniarevich A, Montinaro F, Mondal M, Ongaro L, Singh M, Kumar P, Rai N, Parik J, Metspalu E, Rootsi S, Pagani L, Kivisild T, Metspalu M, Chaubey G, Villems R (2018) The genetic ancestry of modern Indus valley populations from northwest India. Am J Hum Genet 103:918–929. https://doi.org/10.1016/j.ajhg.2018.10.022
Patterson N, Price AL, Reich D (2006) Population structure and eigenanalysis. PLoS Genet 2:e190. https://doi.org/10.1371/journal.pgen.0020190
Patterson N, Moorjani P, Luo YT, Mallick S, Rohland N, Zhan YP, Genschoreck T, Webster T, Reich D (2012) Ancient admixture in human history. Genetics 192:1065. https://doi.org/10.1534/genetics.112.145037
Pickrell JK, Pritchard JK (2012) Inference of population splits and mixtures from genome-wide allele frequency data. PLoS Genet 8:e1002967. https://doi.org/10.1371/journal.pgen.1002967
Purcell S, Neale B, Todd-Brown K, Thomas L, Ferreira MA, Bender D, Maller J, Sklar P, de Bakker PI, Daly MJ, Sham PC (2007) PLINK: a tool set for whole-genome association and population-based linkage analyses. Am J Hum Genet 81:559–575. https://doi.org/10.1086/519795
Raghavan M, Skoglund P, Graf KE, Metspalu M, Albrechtsen A, Moltke I, Rasmussen S, Stafford TW Jr, Orlando L, Metspalu E, Karmin M, Tambets K, Rootsi S, Magi R, Campos PF, Balanovska E, Balanovsky O, Khusnutdinova E, Litvinov S, Osipova LP, Fedorova SA, Voevoda MI, DeGiorgio M, Sicheritz-Ponten T, Brunak S, Demeshchenko S, Kivisild T, Villems R, Nielsen R, Jakobsson M, Willerslev E (2014) Upper Palaeolithic Siberian genome reveals dual ancestry of Native Americans. Nature 505:87–91. https://doi.org/10.1038/nature12736
Reich D, Thangaraj K, Patterson N, Price AL, Singh L (2009) Reconstructing Indian population history. Nature 461:489–494. https://doi.org/10.1038/nature08365
Schreiber M, Schiff AI, Klenicki L (2003) The Shengold Jewish encyclopedia. Schreiber Pub
Sharma S, Rai E, Sharma P, Jena M, Singh S, Darvishi K, Bhat AK, Bhanwer AJ, Tiwari PK, Bamezai RN (2009) The Indian origin of paternal haplogroup R1a1* substantiates the autochthonous origin of Brahmins and the caste system. J Hum Genet 54:47–55. https://doi.org/10.1038/jhg.2008.2
Slapak O, סלפק ע, Yisrael M, Yiśraʼel M, ישראל מ (1995) The Jews of India: a story of three communities. Israel Museum
Thangaraj K, Chaubey G, Kivisild T, Reddy AG, Singh VK, Rasalkar AA, Singh L (2005) Reconstructing the origin of Andaman Islanders. Science 308:996. https://doi.org/10.1126/science.1109987
van Oven M, Kayser M (2009) Updated comprehensive phylogenetic tree of global human mitochondrial DNA variation. Hum Mutat 30:E386–E394. https://doi.org/10.1002/humu.20921
Weil S, Wayl Š, Ṿail S, Publications M, Arts NCftP (2002) India’s Jewish heritage: ritual, art and life-cycle. Marg Publications
Weissensteiner H, Pacher D, Kloss-Brandstatter A, Forer L, Specht G, Bandelt HJ, Kronenberg F, Salas A, Schonherr S (2016) HaploGrep 2: mitochondrial haplogroup classification in the era of high-throughput sequencing. Nucleic Acids Res 44:W58-63. https://doi.org/10.1093/nar/gkw233
Yang MA, Gao X, Theunert C, Tong H, Aximu-Petri A, Nickel B, Slatkin M, Meyer M, Paabo S, Kelso J, Fu Q (2017) 40,000-year-old individual from asia provides insight into early population structure in Eurasia. Curr Biol 27(3202–3208):e9. https://doi.org/10.1016/j.cub.2017.09.030
Zerjal T, Pandya A, Thangaraj K, Ling EY, Kearley J, Bertoneri S, Paracchini S, Singh L, Tyler-Smith C (2007) Y-chromosomal insights into the genetic impact of the caste system in India. Hum Genet 121:137–144. https://doi.org/10.1007/s00439-006-0282-2
Acknowledgements
We thank all the study participants, who volunteered in this study. LK acknowledge the Council of Scientific and Industrial Research (CSIR) for the Fellowship. NR thank the Director, Birbal Sahni Institute of Palaeosciences, and was supported by In-house project-9.3 of BSIP, Lucknow. K.T was supported by the JC Bose Fellowship, SERB, DST and CSIR, Ministry of Science and Technology, Government of India.
Funding
This study is funded by Council of Scientific and Industrial Research (CSIR), Ministry of Science and Technology, Government of India.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
None.
Ethics approval
The study was approved by Institute Ethical Committee of CSIR-CCMB, Hyderabad, India.
Informed consent
Informed written consent was obtained from all the participants involved in the study.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Kumar, L., Farias, K., Prakash, S. et al. Dissecting the genetic history of the Roman Catholic populations of West Coast India. Hum Genet 140, 1487–1498 (2021). https://doi.org/10.1007/s00439-021-02346-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00439-021-02346-4