Abstract
Aspirin-exacerbated respiratory disease (AERD) is a nonallergic clinical syndrome characterized by a severe decline in forced expiratory volume in one second (FEV1) following the ingestion of non-steroidal anti-inflammatory drugs (NSAIDs) such as aspirin. The effects of genetic variants have not fully explained all of the observed individual differences to an aspirin challenge despite previous attempts to identify AERD-related genes. In the present study, we performed genome-wide association study (GWAS) and targeted association study in Korean asthmatics to identify new genetic factors associated with AERD. A total of 685 asthmatic patients without AERD and 117 subjects with AERD were used for the GWAS of the first stage, and 996 asthmatics without AERD and 142 subjects with AERD were used for a follow-up study. A total of 702 SNPs were genotyped using the GoldenGate assay with the VeraCode microbead. GWAS revealed the top-ranked variants in 3′ regions of the HLA-DPB1 gene. To investigate the detailed genetic effects of an associated region with the risk of AERD, a follow-up targeted association study with the 702 single nucleotide polymorphisms (SNPs) of 14 genes was performed on 802 Korean subjects. In a case–control analysis, HLA-DPB1 rs1042151 (Met105Val) shows the most significant association with the susceptibility of AERD (p = 5.11 × 10−7; OR = 2.40). Moreover, rs1042151 also shows a gene dose for the percent decline of FEV1 after an aspirin challenge (p = 2.82 × 10−7). Our findings show that the HLA-DPB1 gene polymorphism may be the most susceptible genetic factor for the risk of AERD in Korean asthmatics and confirm the importance of HLA-DPB1 in the genetic etiology of AERD.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Aspirin-exacerbated respiratory disease (AERD) is characterized by asthma, nasal polyps, chronic rhinosinusitis, and exacerbated bronchial obstruction after the ingestion of non-steroidal anti-inflammatory drugs in about 10–20 % of adult asthmatics (Babu and Salvi 2000; Macy et al. 2007; Sturtevant 1999). The main pathophysiology of AERD is the inhibition of the cyclooxygenase (COX) pathway by aspirin that results in the decreased synthesis of prostaglandin E2 (PGE2) and the overproduction of cysteinyl leukotrienes (cysLTs) (Bennett 2000). Several genetic studies have shown that the polymorphisms of genes in the arachidonate pathways, such as cysteinyl leukotriene receptor 1 (Kim et al. 2007), cysteinyl leukotriene receptor 2 (Park et al. 2005), thromboxane A2 receptor (Kim et al. 2005b), and prostaglandin E2 receptor subtype 2 (Jinnai et al. 2004), leukotriene C4 synthase (LTC4S) (Sanak and Szczeklik 2001) and arachidonate 5-lipoxygenase (ALOX5) (Choi et al. 2004b), are associated with AERD. However, the effects of AERD-related gene variants have partly explained the individual differences in the airway obstruction following aspirin challenge.
Studies have identified alterations in immune responses (Delaney and Kay 1976; Lee et al. 2006), and alterations in the immune and inflammatory gene responses as predisposing factors of the disease; genetic variants such as FcεR1β (Kim et al. 2006), HLA (Choi et al. 2004a; Dekker et al. 1997; Park et al. 2004), TBX21 (Akahoshi et al. 2005), IL-4 (Kim et al. 2008), NLRP3 (Hitomi et al. 2009) PPAR-gamma (Oh et al. 2009) genes were reported to be associated with AERD. The data suggests that genetic variants on the extra-arachidonate pathways may be more related to the development of AERD than previously thought.
Since genome-wide association study (GWAS) emerged as a high throughput technology to genotype genomic nucleotide variations, several genes (e.g., ORMDL3, PDE4D, and IL1RL1) have been discovered to be susceptible to asthma and asthma-related phenotypes (Gudbjartsson et al. 2009; Himes et al. 2009; Moffatt et al. 2007). Our previous GWAS using a low-density BeadChip showed that a nonsynonymous single nucleotide polymorphism (SNP) of CEP68, rs7572857 G>A (Gly74Ser) could be an important marker of AERD susceptibility (Kim et al. 2010). However, this 100K BeadChip could cover a small portion of genetic variants over the whole genome. Recent technical improvements have developed a denser BeadChip of 657,366 (660W) that cover a wider range of genetic variants. To identify new genetic variants associated with AERD, the present study performed GWAS using the 660W chip as well as a replication study to evaluate the associations between genetic variants and the risk of AERD in the Korean population.
Materials and methods
Ethics statement
A written informed consent was obtained from all subjects to participate in the study. The protocols used in this study were approved by the Soonchunhyang Bucheon Hospital’s ethics committee. 1,138 DNA were obtained from the biobank at the Genome Research Center for Allergy and Respiratory Diseases (Soonchunhyang Bucheon Hospital), South Korea.
Subjects
Study subjects were recruited from the Asthma Genome Research Center; all subjects were ethnic Koreans and gave their informed consent to participate in the study. A total of 685 asthmatic patients without AERD and 117 subjects with AERD were used for the GWAS of the first stage, and 996 asthmatics without AERD and 142 subjects with AERD (including samples of GWAS) were used for a follow-up study (Table 1). All patients were diagnosed by physicians and met the criteria for asthma according to the Global Initiative for Asthma (GINA) guidelines (1995). All patients had a history of dyspnea and wheezing in the previous 12 months, plus one of the following: (1) >15 % increase in forced expiratory volume in 1 s (FEV1) or >12 % increase plus 200 mL following inhalation of a short-acting bronchodilator; (2) <10 mg/mL PC20 methacholine; and (3) >20 % increase in FEV1 following 2 weeks of treatment with inhaled or systemic corticosteroids. Total IgE was measured by the CAP system (Pharmacia Diagnostics, Uppsala, Sweden). Twenty-four commonly inhaled allergens were used for a skin-prick test. Atopy was defined as a wheal reaction with a diameter of 3 mm or greater than the reaction to histamine.
Questions related to five specific elements of aspirin hypersensitivity (dyspnea, wheezing, nasal blockage, skin eruption, and loss of consciousness after aspirin ingestion) were incorporated in the questionnaires. Oral aspirin challenge (OAC) was performed to all study subjects. The asthmatic patients had experienced no exacerbation of asthma or respiratory tract infection in the 6 weeks preceding oral aspirin challenge (OAC). OAC was performed with increasing doses of aspirin using the methods previously described (Chang et al. 2011; Nizankowska-Mogilnicka et al. 2007). Patients having a history of aspirin hypersensitivity were given a dose of 30 mg and those with no history were started on 100 mg of aspirin orally. Symptoms, external signs (urticaria, angioedema, and rhinorrhea), blood pressure, and FEV1 were documented every 30 min for a period of 2 h. In the absence of any symptoms or signs suggestive of an adverse reaction, 100 mg of aspirin were administered and the same measurements were repeated every hour, with doses of 450 mg given until the patient developed a reaction. The test was deemed negative if no reaction occurred within 4 h after the final dose. Aspirin-induced bronchospasms were calculated as the pre-challenge FEV1 minus the post-challenge FEV1 divided by the pre-challenge FEV1. OAC reactions were categorized into two groups as follows: (1) a 15 % or greater decrease in FEV1 or naso-ocular reactions (AERD); and (2) a decrease of less than 15 % in FEV1 without naso-ocular or cutaneous reactions [aspirin-tolerant asthma (ATA)]. The protocols were approved by the Ethics Committee of the Soonchunhyang University Hospital.
Genome-wide SNP genotyping
Approximately 200 ng of genomic DNA was used to genotype each sample on the Illumina Human660W BeadChip® (Illumina, Inc., San Diego, CA). Samples were processed according to the Illumina Infinium assay manual. Each sample was whole-genome amplified, fragmented, precipitated, and resuspended in an appropriate hybridization buffer. Denatured samples were hybridized on a prepared Human660W BeadChip® for a minimum of 16 h at 48 °C. Following hybridization, the BeadChips were processed for the single-base extension reaction, stained, and imaged on an Illumina Bead Array Reader. Normalized bead intensity data obtained for each sample were loaded into the GenomeStudio® software (Illumina, Inc.), which converted fluorescent intensities into SNP genotypes. SNP clusters for genotype calling were examined for all SNPs using the GenomeStudio® software. For quality control, SNPs that met the following criteria were retained: call rate ≥0.98, minor allele frequency ≥0.05 except for the SNPs of the X chromosome. The overall call rate for all SNPs was 99.89 %.
Fine-mapping study
A total of 14 genes (Supplementary Table 1), which included the SNPs showing significant association signals, were selected. SNPs on these genes were selected using the database of Asian populations from the International HapMap Project database (http://hapmap.ncbi.nlm.nih.gov/) and the NCBI database (http://www.ncbi.nlm.nih.gov). SNP selection was based on the following scheme. At first, candidate SNPs were extracted in the intragenic region including 2 kb of the 5′ region of each gene using Asian data of International HapMap database, and then LD structures of each gene were analyzed using SNPs with over 5 % of minor allele frequencies. Among SNPs in almost absolute LD (|D′| = 1 and r 2 > 0.95), one representative SNP was selected. A total of 702 SNPs were selected from the sequential processing and genotyped using the GoldenGate assay with the VeraCode microbead (Illumina, Inc.) (Lin et al. 2009). This was followed by a scan using the BeadXpress® system (Illumina, Inc.).
Statistics
For association analysis, genotype distributions between asthmatics with and without AERD were compared using logistic regression analyses adjusted for age, sex, atopic status, smoking status, and BMI as covariates using the HelixTree® software (Golden Helix Inc., Bozeman, MT, USA). Regression models were used for an association analysis of the percent decrease of FEV1 after an aspirin challenge. Haploview v4.2 software downloaded from the Broad Institute (http://www.broadinstitute.org/mpg/haploview) was used to determine the LD of the candidate genes (Barrett et al. 2005). Haplotypes were first estimated using the PHASE software (Stephens et al. 2001) and then computed by logistic analyses using the HelixTree® software.
Results
Genome-wide associations
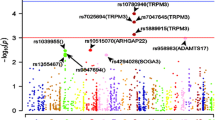
A total of 430,486 SNPs passed our quality control and were used for association analysis with the risk of AERD in 117 subjects with AERD and 685 ATA subjects. The results of the genome-wide association analyses of all SNPs with the risk of AERD are represented in a Manhattan plot (Fig. 1). rs2281389 (p = 5.6 × 10−6; OR = 2.41; Table 2) on the intergenic area adjacent to the 3′ region of HLA-DPB1 was the most strongly associated SNP, although it failed to meet the statistical significance (1.17 × 10−7) when adjusted by a Bonferroni correction (Table 2). The 49 top-ranked SNPs associated with the risk of AERD are listed in Table 2. In the association analysis with the percent decline of FEV1 after an aspirin challenge, all 49 SNPs showed significant associations with the percent decline of FEV1 after an aspirin challenge (Table 2); subsequently, a follow-up study selected 14 genes based on two kinds of association analyses (Supplementary Table 1, Table 2).
p values of GWAS. The Manhattan plot shows p values for AERD versus ATA using the logistic regression analysis. X-axis represents the SNP markers in each chromosome. A total of 430,487 SNPs are presented. The highest p value (p = 5.6 × 10−6; OR = 2.41) was observed in rs2281389 on the intergenic area adjacent to the 3′ region of HLA-DPB1 on 6p21.3
Follow-up fine mapping study
Total 702 SNPs in the 14 genes were genotyped in 142 AERD and 996 ATA subjects. Their clinical characteristics are summarized in Table 1. Three SNPs were failed in genotyping. Among 699 SNPs, HLA-DPB1 gene polymorphisms were most significantly associated with the risk of AERD. Among HLA-DPB1 polymorphisms, rs1042151 (Met105Val) in exon 2 showed the highest susceptibility to AERD (p = 5.11 × 10−7; OR = 2.40; Fig. 2; Supplementary Table 2). Two SNPs among the top ranked in the GWAS (rs2281389 and rs3117230 in 3′ region of HLA-DPB1) were in almost absolute LDs (|D′| = 1 and r 2 = 0.97) with the rs1042151 (Met105Val). The association of these two SNPs observed in GWAS analysis was well replicated in the follow-up analysis (p = 8.85 × 10−7 and 1.37 × 10−6; OR = 2.35 and 2.31, respectively; Fig. 2; Table 3). In addition, a nonsynonymous SNP (Met322Thr) of TSC1 in GWAS and follow-up study showed significant association with the risk of AERD (p = 1.57 × 10−5 and 2.06 × 10−5; OR = 2.49 and 2.26, respectively; Supplementary Table 3).
Regional association plots showing signals in follow-up association study of HLA-DPB1 and 3′ expanded region. SNPs are plotted by position on chromosome against association with risk of AERD (−log10 p value). Circles mean the SNPs of GWAS and squares mean the added SNPs for follow-up study. The figures highlight the most significantly associated SNP after the follow-up study. Estimated recombination rates, averaged across populations from HapMap DB, are plotted in blue to reflect the local LD structure. SNPs surrounding the most significant SNP are color-coded to reflect their LD with this SNP as in the insert. Genes and the plotted SNPs, the direction of transcription, and the LDs among SNPs are noted below the plots. Haplotype blocks are parsed by |D′| values (|D′| > 0.9) (color figure online)
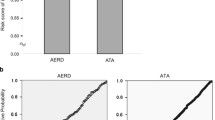
The associations between genotype distribution of each SNP and the percent decline of FEV1 after aspirin challenge were also analyzed using the linear regression analyses. HLA-DPB1 rs1042151 (Met105Val) showed the most significant association in gene dose-dependent manner, e.g. the mean values of percent decline of FEV1 after aspirin challenge were 6.28 % in Met/Met, 10.15 % in Met/Val, and 17.27 % in Val/Val (p = 2.82 × 10−7; Supplementary Table 4). SNPs in other candidate genes showed similar pattern in their associations of the percent decline of FEV1 after aspirin challenge to the analyses of the risk of AERD (Supplementary Table 5).
Discussion
In the present study to identify a new genetic predisposition for the risk of AERD, GWAS with 430,486 SNPs was tried in 802 asthmatics. In fine mapping, 702 SNPs were genotyped in 1,138 subjects. As a result, a nonsynonymous SNP in exon 2 of HLA-DPB1, rs1042151 (Met105Val), showed the highest association with the risk of AERD. The significance remained after a Bonferroni correction.
The HLA-DPB1 gene has been implicated in AERD. Dekker and colleagues (Dekker et al. 1997) first reported that the DPB1*0301 frequency increased in 118 AERD patients when compared with 114 ATA subjects (19.5 vs. 4.4 %, respectively). The association of DPB1*0301 with AERD was replicated in a Korean population study; 13.8 % in 76 AERD and 4.1 % in 73 ATA (p = 0.004) (Choi et al. 2004a). The reason for the high frequency in the Caucasian population may be due to racial differences and the genotyping methods between the studies. Dekker and colleagues performed HLA-DPB l typing of 19 recognized alleles (Bodmer et al. 1990) by a restriction enzyme analysis of the exon 2 amplified by PCR (Dekker and Easteal 1990). The Korean study (Dekker et al. 1997) performed a high-resolution genotyping of HLA-DPB l alleles on amplified gene products using 21 recognized locus specific primers (Van der Zwan and Tilanus 2000) and automatic direct sequencing. In the present study, the minor allele frequency of rs1042151 A>G is 17.7 % in 142 AERD and 8.3 % in 996 ATA. The data indicates that the minor allele frequency of rs1042151 A>G in the present study is higher than the frequency of DPB1*0301 in AERD (13.8 %) and in ATA (4.1 %) of the previous Korean population study (Choi et al. 2004a). The differences may originate from the HLA-DPB1 alleles other than HLA-DPB1*0301 having the minor allele of rs1042151 A>G. Because more than hundred alleles have been described for DPB1 locus (http://hla.alleles.org/data/txt/dpb_prot.txt), frequency of other HLA-DPB1 alleles may be important information in the study of immune responses. There are 40 DPB1 alleles having valine at 76 AA in exon 2. Among them, the DPB1*0301 may be the most frequent allele having rs1042151 (Met105Val) in exon 2 in Koreans (41) and Caucasians (22). Dekker and collaborators reported a higher frequency of DPB1*0101 allele and lower frequency of 0401 in AERD compared to ATA (22). HLA-DPB1*0101 have valine while HLA-DPB1*0401 have methionine in exon 2; subsequently, the summation of DPB1*0101 and DPB1*0301 may increase the frequency of the minor allele of rs1042151 A>G. In addition, high-density sequencing analysis of HLA-DPB1 alleles would provide more detailed information on the immunogenetic mechanism of AERD. A Korean study of 152 subjects with AERD reported the frequency of 10 alleles in HLA-DPB1 (Kim et al. 2005a). The minor allele frequencies of HLA-DPB1* rs1042151 A>G in AERD are 13.8 % (DPB1*0301), 2 % (DPB1*0901), and 2 % (DPB1*1401), respectively. The summation of the alleles frequency is 17.8 %, which is almost similar to the allele frequency of rs1042151 A>G (17.7 %) in the present study. In addition, they genotyped DPB1*0301 in 188 subjects with aspirin-induced urticaria and found no difference in the frequency of DPB1*0301 compared to that of the normal control (Kim et al. 2005a). The data suggest that aspirin-induced urticaria may have different immunologic backgrounds from AERD.
In our previous low-density BeadChip GWAS (Kim et al. 2010), we could not find any association of HLA-DPB1 with AERD. This result is caused by the low SNP coverage of the region that includes the HLA-DPB1 gene; rs1042151 was excluded in the contents of the low-density BeadChip. In addition, CEP68 gene polymorphisms were not replicated in the present study. In our previous study, the nonsynonymous rs7572857 G>A variant of the CEP68 gene showed the most significant association with AERD compared to ATA controls in co-dominant model (p = 6.0 × 10−5). In the present study, the Illumina Human660W contains rs2302647 and rs10496123 SNPs on the CEP68 gene instead of rs7572857 SNP. The two SNPs rs2302647 and rs10496123 showed a statistically insignificant association with AERD both in the present and in the previous study. Because the 660 Chip includes only one-thirties of the SNP contents of the 100K chip (3,178 SNPs), combination of the two results would provide more additive information on the genetic variants related to human diseases.
Additional in silico analyses were performed to investigate the potential functions of the nonsynonymous rs1042151 of HLA-DPB1. Using the ELM program (http://elm.eu.org/index.html), the amino acid change from the methionine to valine of the rs1042151 was not predicted as a functional linear motif. We applied the eQTL browser (http://eqtl.uchicago.edu/cgi-bin/gbrowse/eqtl/) to expect whether rs1042151 could regulate the expression of protein(s) and/or mRNA(s). Intriguingly, rs1042151 was found to act as a potential cis-regulator for HLA-DPB1 expression with a high eQTL score of 36.83 (Supplementary Fig. 1). In addition, the SNP function prediction (http://snpinfo.niehs.nih.gov/snpfunc.htm) indicates that the rs1042151 could be located in an exonic splicing enhancer (ESE) region (Supplementary Table 6). The findings suggest that HLA-DPB1 rs1042151 could be a functional nonsynonymous variant for AERD development.
In conclusion, we demonstrated that SNPs on 3′ regions of the HLA-DPB1 gene were significantly associated with the risk of AERD on a GWAS in a Korean asthmatic cohort. The genetic effects of HLA-DPB1 polymorphisms were scrutinized for AERD via fine mapping and rs1042151 (Met105Val) was revealed to be the most important genetic variant for the risk of AERD. The findings confirm previous studies and provide new insight into the genetic factors associated with AERD.
References
(1995) Global Initiative for Asthma. Global strategy for asthma management and prevention. NHLBI/WHO workshop report. Bethesda. National Institutes of Health
Akahoshi M, Obara K, Hirota T, Matsuda A, Hasegawa K, Takahashi N, Shimizu M, Nakashima K, Cheng L, Doi S, Fujiwara H, Miyatake A, Fujita K, Higashi N, Taniguchi M, Enomoto T, Mao XQ, Nakashima H, Adra CN, Nakamura Y, Tamari M, Shirakawa T (2005) Functional promoter polymorphism in the TBX21 gene associated with aspirin-induced asthma. Hum Genet 117:16–26
Babu KS, Salvi SS (2000) Aspirin and asthma. Chest 118:1470–1476
Barrett JC, Fry B, Maller J, Daly MJ (2005) Haploview: analysis and visualization of LD and haplotype maps. Bioinformatics 21:263–265
Bennett A (2000) The importance of COX-2 inhibition for aspirin induced asthma. Thorax 55(Suppl 2):S54–S56
Bodmer JG, Marsh SG, Parham P, Erlich HA, Albert E, Bodmer WF, Dupont B, Mach B, Mayr WR, Sasazuki T et al (1990) Nomenclature for factors of the HLA system, 1989. Tissue Antigens 35:1–8
Chang HS, Park JS, Jang AS, Park SW, Uh ST, Kim YH, Park CS (2011) Diagnostic value of clinical parameters in the prediction of aspirin-exacerbated respiratory disease in asthma. Allergy Asthma Immunol Res 3:256–264
Choi JH, Lee KW, Oh HB, Lee KJ, Suh YJ, Park CS, Park HS (2004a) HLA association in aspirin-intolerant asthma: DPB1*0301 as a strong marker in a Korean population. J Allergy Clin Immunol 113:562–564
Choi JH, Park HS, Oh HB, Lee JH, Suh YJ, Park CS, Shin HD (2004b) Leukotriene-related gene polymorphisms in ASA-intolerant asthma: an association with a haplotype of 5-lipoxygenase. Hum Genet 114:337–344
Dekker JW, Easteal S (1990) HLA-DP typing by amplified fragment length polymorphisms (AFLPs). Immunogenetics 32:56–59
Dekker JW, Nizankowska E, Schmitz-Schumann M, Pile K, Bochenek G, Dyczek A, Cookson WO, Szczeklik A (1997) Aspirin-induced asthma and HLA-DRB1 and HLA-DPB1 genotypes. Clin Exp Allergy 27:574–577
Delaney JC, Kay AB (1976) Complement components and IgE in patients with asthma and aspirin idiosyncrasy. Thorax 31:425–427
Gudbjartsson DF, Bjornsdottir US, Halapi E, Helgadottir A, Sulem P, Jonsdottir GM, Thorleifsson G, Helgadottir H, Steinthorsdottir V, Stefansson H, Williams C, Hui J, Beilby J, Warrington NM, James A, Palmer LJ, Koppelman GH, Heinzmann A, Krueger M, Boezen HM, Wheatley A, Altmuller J, Shin HD, Uh ST, Cheong HS, Jonsdottir B, Gislason D, Park CS, Rasmussen LM, Porsbjerg C, Hansen JW, Backer V, Werge T, Janson C, Jonsson UB, Ng MC, Chan J, So WY, Ma R, Shah SH, Granger CB, Quyyumi AA, Levey AI, Vaccarino V, Reilly MP, Rader DJ, Williams MJ, van Rij AM, Jones GT, Trabetti E, Malerba G, Pignatti PF, Boner A, Pescollderungg L, Girelli D, Olivieri O, Martinelli N, Ludviksson BR, Ludviksdottir D, Eyjolfsson GI, Arnar D, Thorgeirsson G, Deichmann K, Thompson PJ, Wjst M, Hall IP, Postma DS, Gislason T, Gulcher J, Kong A, Jonsdottir I, Thorsteinsdottir U, Stefansson K (2009) Sequence variants affecting eosinophil numbers associate with asthma and myocardial infarction. Nat Genet 41:342–347
Himes BE, Hunninghake GM, Baurley JW, Rafaels NM, Sleiman P, Strachan DP, Wilk JB, Willis-Owen SA, Klanderman B, Lasky-Su J, Lazarus R, Murphy AJ, Soto-Quiros ME, Avila L, Beaty T, Mathias RA, Ruczinski I, Barnes KC, Celedon JC, Cookson WO, Gauderman WJ, Gilliland FD, Hakonarson H, Lange C, Moffatt MF, O’Connor GT, Raby BA, Silverman EK, Weiss ST (2009) Genome-wide association analysis identifies PDE4D as an asthma-susceptibility gene. Am J Hum Genet 84:581–593
Hitomi Y, Ebisawa M, Tomikawa M, Imai T, Komata T, Hirota T, Harada M, Sakashita M, Suzuki Y, Shimojo N, Kohno Y, Fujita K, Miyatake A, Doi S, Enomoto T, Taniguchi M, Higashi N, Nakamura Y, Tamari M (2009) Associations of functional NLRP3 polymorphisms with susceptibility to food-induced anaphylaxis and aspirin-induced asthma. J Allergy Clin Immunol 124(779-85):e6
Jinnai N, Sakagami T, Sekigawa T, Kakihara M, Nakajima T, Yoshida K, Goto S, Hasegawa T, Koshino T, Hasegawa Y, Inoue H, Suzuki N, Sano Y, Inoue I (2004) Polymorphisms in the prostaglandin E2 receptor subtype 2 gene confer susceptibility to aspirin-intolerant asthma: a candidate gene approach. Hum Mol Genet 13:3203–3217
Kim SH, Choi JH, Lee KW, Shin ES, Oh HB, Suh CH, Nahm DH, Park HS (2005a) The human leucocyte antigen-DRB1*1302-DQB1*0609-DPB1*0201 haplotype may be a strong genetic marker for aspirin-induced urticaria. Clin Exp Allergy 35:339–344
Kim SH, Choi JH, Park HS, Holloway JW, Lee SK, Park CS, Shin HD (2005b) Association of thromboxane A2 receptor gene polymorphism with the phenotype of acetyl salicylic acid-intolerant asthma. Clin Exp Allergy 35:585–590
Kim SH, Ye YM, Lee SK, Choi JH, Holloway JW, Park CS, Park HS (2006) Association of TNF-alpha genetic polymorphism with HLA DPB1*0301. Clin Exp Allergy 36:1247–1253
Kim SH, Ye YM, Hur GY, Lee SK, Sampson AP, Lee HY, Park HS (2007) CysLTR1 promoter polymorphism and requirement for leukotriene receptor antagonist in aspirin-intolerant asthma patients. Pharmacogenomics 8:1143–1150
Kim TH, Chang HS, Park SM, Nam BY, Park JS, Rhim T, Park HS, Kim MK, Choi IS, Cho SH, Chung IY, Park BL, Park CS, Shin HD (2008) Association of angiotensin I-converting enzyme gene polymorphisms with aspirin intolerance in asthmatics. Clin Exp Allergy 38:1727–1737
Kim JH, Park BL, Cheong HS, Bae JS, Park JS, Jang AS, Uh ST, Choi JS, Kim YH, Kim MK, Choi IS, Cho SH, Choi BW, Park CS, Shin HD (2010) Genome-wide and follow-up studies identify CEP68 gene variants associated with risk of aspirin-intolerant asthma. PLoS ONE 5:e13818
Lee SH, Rhim T, Choi YS, Min JW, Kim SH, Cho SY, Paik YK, Park CS (2006) Complement C3a and C4a increased in plasma of patients with aspirin-induced asthma. Am J Respir Crit Care Med 173:370–378
Lin CH, Yeakley JM, McDaniel TK, Shen R (2009) Medium- to high-throughput SNP genotyping using VeraCode microbeads. Methods Mol Biol 496:129–142
Macy E, Bernstein JA, Castells MC, Gawchik SM, Lee TH, Settipane RA, Simon RA, Wald J, Woessner KM (2007) Aspirin challenge and desensitization for aspirin-exacerbated respiratory disease: a practice paper. Ann Allergy Asthma Immunol 98:172–174
Moffatt MF, Kabesch M, Liang L, Dixon AL, Strachan D, Heath S, Depner M, von Berg A, Bufe A, Rietschel E, Heinzmann A, Simma B, Frischer T, Willis-Owen SA, Wong KC, Illig T, Vogelberg C, Weiland SK, von Mutius E, Abecasis GR, Farrall M, Gut IG, Lathrop GM, Cookson WO (2007) Genetic variants regulating ORMDL3 expression contribute to the risk of childhood asthma. Nature 448:470–473
Nizankowska-Mogilnicka E, Bochenek G, Mastalerz L, Swierczynska M, Picado C, Scadding G, Kowalski ML, Setkowicz M, Ring J, Brockow K, Bachert C, Wohrl S, Dahlen B, Szczeklik A (2007) EAACI/GA2LEN guideline: aspirin provocation tests for diagnosis of aspirin hypersensitivity. Allergy 62:1111–1118
Oh SH, Park SM, Park JS, Jang AS, Lee YM, Uh ST, Kim YH, Choi IS, Kim MK, Park BL, Shin HD, Park CS (2009) Association analysis of peroxisome proliferator-activated receptors gamma gene polymorphisms with aspirin hypersensitivity in asthmatics. Allergy Asthma Immunol Res 1:30–35
Park HS, Kim SH, Sampson AP, Lee KW, Park CS (2004) The HLA-DPB1*0301 marker might predict the requirement for leukotriene receptor antagonist in patients with aspirin-intolerant asthma. J Allergy Clin Immunol 114:688–689
Park JS, Chang HS, Park CS, Lee JH, Lee YM, Choi JH, Park HS, Kim LH, Park BL, Choi YH, Shin HD (2005) Association analysis of cysteinyl-leukotriene receptor 2 (CYSLTR2) polymorphisms with aspirin intolerance in asthmatics. Pharmacogenet Genomics 15:483–492
Sanak M, Szczeklik A (2001) Leukotriene C4 synthase polymorphism and aspirin-induced asthma. J Allergy Clin Immunol 107:561–562
Stephens M, Smith NJ, Donnelly P (2001) A new statistical method for haplotype reconstruction from population data. Am J Hum Genet 68:978–989
Sturtevant J (1999) NSAID-induced bronchospasm—a common and serious problem. A report from MEDSAFE, the New Zealand Medicines and Medical Devices Safety Authority. NZ Dent J 95:84
Van der Zwan RE, Tilanus MG (2000) Sequence-based typing for HLA-DPB1 strategy for ABI sequencing equipment. In: Tilanus MGJ, Hansen JA, Heuley CK (eds) IHWG technical manual. International Histocompatibility Working Group, Seattle, TM14A, pp 1–4
Acknowledgments
This study was supported by a grant of the Korea Healthcare Technology R&D Project, Ministry for Health, Welfare & Family Affairs, Republic of Korea (A090548 & 010249), and by the National Research Foundation of Korea (NRF) grant funded by the Korea Government (MEST) (No. 2009-0080157). The DNA samples were generously provided by Soonchunhyang University, Bucheon Hospital Biobank, a member of the National Biobank of Korea, supported by the Ministry of Health, Welfare and Family Affairs, Republic of Korea.
Conflict of interest
The authors declare that they have no conflicts of interest.
Author information
Authors and Affiliations
Corresponding authors
Electronic supplementary material
Below is the link to the electronic supplementary material.
439_2012_1247_MOESM2_ESM.tif
Supplementary Figure 1 Supplementary Figure 1. The eQTL result of rs1042151 of HLA-DPB1. Red box indicates eQTL rs1042151 of HLA-DPB1 derived from the eQTL browser (http://eqtl.uchicago.edu/cgi-bin/gbrowse/eqtl/) (TIFF 2722 kb)
Rights and permissions
About this article
Cite this article
Park, B.L., Kim, TH., Kim, JH. et al. Genome-wide association study of aspirin-exacerbated respiratory disease in a Korean population. Hum Genet 132, 313–321 (2013). https://doi.org/10.1007/s00439-012-1247-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00439-012-1247-2