Abstract
Mutations in the FBN1 gene are the major cause of Marfan syndrome (MFS), an autosomal dominant connective tissue disorder, which displays variable manifestations in the cardiovascular, ocular, and skeletal systems. Current molecular genetic testing of FBN1 may miss mutations in the promoter region or in other noncoding sequences as well as partial or complete gene deletions and duplications. In this study, we tested for copy number variations by successively applying multiplex ligation-dependent probe amplification (MLPA) and the Affymetrix Human Mapping 500 K Array Set, which contains probes for ∼500,000 single-nucleotide polymorphisms (SNPs) across the genome. By analyzing genomic DNA of 101 unrelated individuals with MFS or related phenotypes in whom standard genetic testing detected no mutation, we identified FBN1 deletions in two patients with MFS. Our high-resolution approach narrowed down the deletion breakpoints. Subsequent sequencing of the junctional fragments revealed the deletion sizes of 26,887 and 302,580 bp, respectively. Surprisingly, both deletions affect the putative regulatory and promoter region of the FBN1 gene, strongly indicating that they abolish transcription of the deleted allele. This expectation of complete loss of function of one allele, i.e. true haploinsufficiency, was confirmed by transcript analyses. Our findings not only emphasize the importance of screening for large genomic rearrangements in comprehensive genetic testing of FBN1 but, importantly, also extend the molecular etiology of MFS by providing hitherto unreported evidence that true haploinsufficiency is sufficient to cause MFS.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Marfan syndrome (MFS; MIM# 154700) is a prevalent connective tissue disorder with variable manifestations in the skeletal, ocular, and cardiovascular systems (De Paepe et al. 1996). MFS is inherited in an autosomal dominant manner and caused by mutations in the gene encoding the 350-kD extracellular matrix protein fibrillin-1 (FBN1; MIM# 134797) in the majority of cases (Dietz et al. 1991). Recently, heterozygous mutations in the genes coding for transforming growth factor beta receptors I (TGFBR1; MIM# 190181) and II (TGFBR2; MIM# 190182) have also been reported in patients with MFS-related disorders, such as MFS type 2 (MFS2; MIM# 154705), Loeys-Dietz aortic aneurysm syndrome (LDS; MIM# 609192), and familial thoracic aortic aneurysms and dissections (TAAD; MIM# 132900), indicating genetic heterogeneity in MFS and its related conditions (Mizuguchi et al. 2004; Loeys et al. 2005; Pannu et al. 2005; Mátyás et al. 2006; Loeys et al. 2006).
In classical MFS patients, mutation analyses have failed to detect FBN1 involvement in at least 10% of cases, suggesting not only that the disease-causing mutation occurs in a different gene but also demonstrate the limitations of commonly used standard PCR-based screening approaches. Indeed, current molecular genetic testing of FBN1, although powerful, may miss mutations in the promoter region or in other noncoding sequences as well as deletions/duplications of the entire gene or part of it. Accordingly, in the locus-specific mutation databases more than 600 unique FBN1 mutations but only 12 (2.0%) large deletions have been registered (UMD, http://www.umd.be:2030; HGMD, http://www.hgmd.cf.ac.uk/ac/gene.php?gene = FBN1). In contrast, the relative frequency of large deletions available in the Human Gene Mutation Database (http://www.hgmd.cf.ac.uk) containing 64,251 mutations in 2,362 genes (version 09/12/2006) is significantly higher (5.6%), suggesting that more large FBN1 deletions await detection.
In this study, we tested this hypothesis by applying the recently introduced multiplex ligation-dependent probe amplification (MLPA) technique, which is suitable for the identification of large deletions and less laborious than methods traditionally used for detection of gross rearrangements, such as Southern blot and cytogenetic techniques. MLPA relies on sequence-specific probe hybridization to genomic DNA, followed by amplification of the hybridized probe with universal primers, and semi-quantitative analysis of the resulting PCR products (Schouten et al. 2002). Here, we report on large deletions affecting the 5′ region of the FBN1 gene in two unrelated MFS patients. The breakpoints of the deletions identified by MLPA were narrowed down by means of high-density single-nucleotide polymorphism (SNP) arrays and characterized by using long-range PCR, subsequent cycle sequencing, and transcript analyses. Our data provide hitherto unreported evidence for true haploinsufficiency in patients with MFS.
Subjects and methods
Patients
A cohort of 101 unrelated patients with suspected MFS (∼40%) or MFS-like phenotypes (∼60%) involving monosymptomatically or predominantly the cardiovascular (∼39%), skeletal (∼18%) or ocular (∼3%) system was selected for this study. In this cohort, previous denaturing high-performance liquid chromatography (DHPLC) analysis and/or sequencing of all exons and flanking intronic regions of FBN1, TGFBR1, and TGFBR2 revealed no disease-causing sequence variant (Mátyás et al. 2002a, 2006). Data on the clinical phenotypes of patients were collected from medical records or during physical examinations by one of the authors (B.S, T.C.).
Multiplex ligation-dependent probe amplification (MLPA)
A total of 100 ng of genomic DNA of each patient was screened by MLPA using the SALSA kits P065 (probes for FBN1 and TGFBR2), P066 (probes for FBN1), and P148 (probes for both TGFBR1 and TGFBR2), commercially available from MRC-Holland (Amsterdam, The Netherlands), on an ABI PRISM 310 or 3100 Genetic Analyzer (Applied Biosystems, Rotkreuz, Switzerland) according to the manufacturers’ instructions. Each MLPA signal was normalized and compared to the corresponding peak area obtained in a control DNA sample. Deviations more than 30% were suspected as alterations and verified by repeated MLPA analysis.
High-density microarray analyses
In order to narrow down the breakpoints of the deletions identified by MLPA, we used the high-density GeneChip Human Mapping 500 K Array Set (Affymetrix, Santa Clara, CA, USA), which contains ∼500,000 genomewide SNPs, according to the manufacturer’s manual. After hybridization and scanning, raw data were processed using the GCOS 1.4 software (Affymetrix, Santa Clara, CA, USA) and probe cell intensities were calculated and summarized for the respective probe sets by means of the MAS5 algorithm (Hubbell et al. 2002). SNP allele calls, marker intensity values, and quality control measures were computed by the GTYPE software (Affymetrix, Santa Clara, CA, USA) using the integrated RLMM algorithm (Rabbee and Speed 2006). Quality control measures were considered before performing the statistical analysis, including appropriate numbers of present calls (>97%) and control SNP performance (Modified Partitioning Around Medoids [MPAM] algorithm Call Rate [MCR] close to or higher than 90%). For the determination of regions with loss of heterozygosity and decreased signal intensity, advanced data analyses were performed using the academic software tools dChip (Harvard University; http://www.biosun1.harvard.edu/complab/dchip) and CNAG2.0 (University of Tokyo; http://www.genome.umin.jp).
Amplification and sequencing of the junctional fragments
Primers flanking the predicted deletions were designed based on decreased MLPA and array signal intensities and were used in long-range PCR (44F 5′-ACGAACCTTTCAAATATTCCCCC-3′, 44R 5′-TGGGAGTGATGGGACAACTGAG-3′, 70F 5′-ACTGCTGCAGAGTGCCTGATA-3′, 70R 5′-GGGCTATGTGCTCTGGCTATT-3′). Briefly, 100 ng of DNA and Buffer 3 were used with the Expand Long Template PCR System (Roche Diagnostics, Rotkreuz, Switzerland) according to the manufacturer’s instructions with the following cycling profile: 5 min initial denaturation at 92°C; 10 cycles of 10 s at 92°C, 30 s at 58.5°C (for 44F/44R) or 59.3°C (for 70F/70R), and 8 min at 68°C; followed by 25 cycles of 15 s at 92°C, 30 s at 58.5°C (for 44F/44R) or 59.3°C (for 70F/70R), and 8 min (+20 s/cycle) at 68°C; finished by a 7 min final extension step at 68°C. Amplicons were treated with ExoSAP-IT (USB Corporation, Cleveland, OH, USA) and sequenced using internal primers by means of a BigDye Terminator Cycle Sequencing Ready Reaction kit v1.1 on an ABI PRISM 3100 Genetic Analyzer (Applied Biosystems, Rotkreuz, Switzerland). Sequences were analyzed using SeqScape 2.5 (Applied Biosystems, Rotkreuz, Switzerland) and the genome browser of the University of California, Santa Cruz (UCSC; http://www.genome.ucsc.edu). Sequence variants were referenced with respect to the human genome reference sequence (NCBI build 36.1, March 2006) according to the guidelines of the HGVS (http://www.hgvs.org/mutnomen).
Transcript analyses
Primary fibroblasts were available only from Patient 70. Total RNA was isolated by the RNeasy mini kit (Qiagen, Hilden, Germany) from cultured fibroblasts. Prior to RNA extraction, fibroblasts were incubated with 10 μg/ml cycloheximide (Actidione; Sigma, St. Louis, MO, USA) for 15 h in order to inhibit potential nonsense-mediated mRNA decay. Reverse transcription (RT)-PCR was used to amplify a complementary DNA (cDNA) fragment (1,484 bp) containing the polymorphism c.8,930C > T (rs1042078, average heterozygosity 0.490 ± 0.071, dbSNP build 126) that is located in the 3′ untranslated region (UTR) of the FBN1 gene and was detected during the mutation screening at genomic DNA level. RT-PCR was performed by means of the OneStep RT-PCR kit (Qiagen, Hilden, Germany) using 125 ng RNA and primers specific to the FBN1 exons 63 (5′-AGGGCGGTTACCTGTGTG-3′) and 65 (5′-CCCCTTGTTGACAGGAATGAC-3′). RT-PCR product was sequenced by using an internal primer in exon 65 (5′-AGCACCATTACAAACCCTCACA-3′) as described above. The presence of allele-specific transcripts was considered according to the procedure previously outlined (Mátyás et al. 2002b; Qiu et al. 2003).
Results
Multiplex ligation-dependent probe amplification (MLPA)
In 2 out of 101 previously “mutation-negative” patients with MFS or related phenotypes, MLPA analyses resulted in abnormal patterns. In Patient 44, MLPA revealed reduced relative peak areas for both probes of FBN1 exon 1 (Fig. 1), suggesting the deletion of this exon. Only the father’s DNA was available for MLPA analysis, which revealed a normal profile (data not shown). In Patient 70, MLPA showed reduced relative peak areas of fragments corresponding to exons 1–9 and 12–16 (Fig. 1), suggesting the deletion of exons 1–16. The parents of Patient 70 were not available for testing. All other patients showed relative peak areas within the range defined as normal. MLPA for the TGFBR1 and TGFBR2 genes (kit P148) showed normal relative peak areas in all patients (data not shown). Taken together, the relative frequency of gross FBN1 deletions in patients with suspected MFS in whom exon-by-exon screening revealed no mutation in the genes FBN1, TGFBR1, and TGFBR2 can be estimated as 2/101 (0.3–7.7%, P = 0.05; according to VassarStats, http://www.faculty.vassar.edu/lowry/prop1.html). Patients 44 and 70 were sporadic cases and fulfilled the diagnostic criteria of Ghent nosology (Table 1).
Results of semiquantitative MLPA analyses in Patients 44 and 70. The normal range of normalized relative peak areas (white bars) is given by dotted lines (±0.30). Values out of this range are marked by black bars and numbers of the corresponding FBN1 exons (Ex). a Normalized relative peak areas measured with the P065 kit, which consists of 25 MLPA probes for FBN1 and eleven control MLPA probes located on chromosomes 1, 2, 4, 5, 8, 11, and 16 (fragments 1, 2, 6, 13, 19, 25, 31, 32, 36, 40, and 43) as well as probes for all seven TGFBR2 exons (fragments 8, 12, 16, 20, 24, 28, and 34). b Normalized relative peak areas measured with the P066 kit, which consists of MLPA probes for 30 FBN1 exons, in two of which (exons 1 and 6) a probe of P065 is located as well, and eleven control MLPA probes located on chromosomes 1, 3, 5, 6, 13, 16, and 22 (fragments 1, 2, 7, 12, 17, 22, 27, 31, 36, 40, and 41). Taken together the results of the P065 and P066 kits, MLPA revealed reduced relative peak areas of fragments corresponding to the FBN1 exon 1 in Patient 44 and to the FBN1 exons 1–9 and 12–16 in Patient 70 (note that there are no MLPA probes for the FBN1 exons 10 and 11 as well as 20, 22, 27 32, 37, 39, 48, 51, 59, and 61)
Identification and characterization of breakpoints
Loss of heterozygosity and decreased signal intensities upon high-density SNP array analyses confirmed the MLPA results and narrowed down the deletion breakpoints. In Patient 44, loss of heterozygosity occurred in a region spanning 94.5 kb between the SNPs rs1678982 and rs784408, while reduction of signal intensities indicated a deletion of less than 44.4 kb between rs2018854 and rs931781 (Fig. 2a). Similarly, in Patient 70 the deletion was localized to a region of ∼317 kb between rs11070644 and rs2899422 by considering decreased array signal intensities (Fig. 3a).
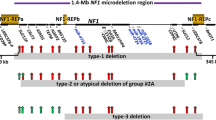
Breakpoint analyses in Patient 44. a Schematic representation of the genome in the region of deletion as well as an overview of the results of the MLPA and SNP array analyses. The open arrow below the gene name indicates the direction of transcription. Exons are specified by bars and marked with the corresponding number or character (cf. C, A, and B are alternatively spliced exons of FBN1, while exon 1, also known as exon M, harbors the presumptive translation initiation codon). The region with decreased normalized MLPA signals (Fig. 1) is indicated by a yellow bar and the positions of the two MLPA probes located in exon 1 of FBN1 are denoted by arrows (filled triangle). The regions with loss of heterozygosity and decreased SNP array signal intensity both gained by the Affymetrix GeneChip Human Mapping 500 K Array Set are indicated by white and blue bars, respectively (note that the flanking SNPs are not included in the regions, displaying an open interval). The positions of SNPs tested in the array set are denoted by vertical lines. Note the large region (∼67 kb) with loss of heterozygosity not associated with the deletion. Primers designed for long-range PCR (LR primers 44F and 44R) and the deleted region of the genomic DNA (brown bar) are indicated. b Sequences of the long-range PCR product spanning the breakpoint junction of the deletion. Uppercase letters indicate sequences of FBN1 intron 1, lowercase letters denote intergenic sequences. Due to identical sequences at the site of breakpoints, the break and rejoining could have occurred at three positions as indicated by arrows (open triangle). The dotted line marks the most telomeric position of the possible breakpoints. All nucleotide positions are described in relation to the human genome reference sequence (NCBI build 36.1)
Breakpoint analyses in Patient 70. Symbols and labels are as used in Fig. 2. a Schematic representation of the genome in the region of deletion as well as overview of the results of the MLPA and SNP array analyses. Note the extended region (∼180 kb) with loss of heterozygosity not deleted. Forward and reverse primers designed for long-range PCR are indicated (LR primers 70F and 70R). For clarity, exons C, A, and B of FBN1 are not shown (cf. Fig. 2). b Sequences of the long-range PCR product spanning the breakpoint junction of the deletion. Uppercase letters indicate sequences of FBN1 intron 16, and lowercase letters denote sequences of CEP152 intron 2
Based on these data, forward and reverse primers were designed for long-range PCR, which resulted in fragments of ∼6.5 and ∼8.5 kb in Patient 44 and Patient 70, respectively. Due to the size of normal alleles (33.2 kb using primers 44F/44R and 311.3 kb with primers 70F/70R), amplifications were observed from the deleted alleles only, but not from the normal alleles of either Patient 44, his father, Patient 70, or several controls (data not shown). Subsequent sequencing of the long-range PCR products identified a deletion of 26,887 bp in Patient 44 and 302,580 bp in Patient 70 (Figs. 2b, 3b).
Transcript analyses
Both deletions affect exon 1 (345 bp), which contains the translation initiation ATG codon at nucleotide positions 182–184, and a ∼4-kb upstream region of the FBN1 gene, which may harbor the promoter, thus strongly indicating that they abolish the transcription/translation of the respective deleted allele (Figs. 2, 3). This expectation of complete loss of function of one allele (true haploinsufficiency) was confirmed by RT-PCR sequence analysis in Patient 70 being heterozygous for the c.8,930C > T polymorphism in the FBN1 3′UTR, which showed the transcription of only one allele (Fig. 4). Appropriate clinical samples were not available from Patient 44 to verify this finding in affected tissue.
Partial reverse sequence of the FBN1 3′ untranslated region flanking the polymorphism c.8,930C > T (g.46,490,165 NCBI build 36.1; arrowed). Note that the cDNA sequence of Patient 70 heterozygous for c.8,930C > T lacks allele C (black) and is indistinguishable from the cDNA sequence of Control 2 homozygous for allele T (green), while in the controls the results at gDNA and cDNA levels are consistent with respect to each other. Complementary DNA sequences were derived from cultured fibroblast mRNA stabilized by the translation inhibitor cycloheximide
Discussion
In this study, we applied for the first time MLPA and high-density SNP arrays to analyze genomic DNA samples of patients with MFS or related phenotypes in whom standard molecular testing detected no mutation. Our results contribute to the molecular etiology of MFS by providing previously unprecedented evidence for true haploinsufficiency in patients. Our data significantly extend the number of large FBN1 deletions, which have previously been detected by cDNA or Southern blot analyses (Table 2), demonstrating that comprehensive genetic testing of FBN1 should include screening for large genomic rearrangements by using an appropriate method.
The P065-P066 MLPA kit used in this study contains probes for 53 of the 65 FBN1 exons. Since not all FBN1 exons were analyzed, a deletion or duplication of non-tested exons cannot be excluded. Furthermore, copy-number neutral rearrangements as well as those affecting no MLPA-specific sequences could not be detected because such mutations do not change the relative peak area of MLPA probes. In contrast to traditionally used techniques, MLPA is a simple method to detect gross deletions. However, sequence variations in the probe-binding regions as well as probe and DNA quality may significantly affect MLPA results. Accordingly, it is highly recommended to confirm MLPA results by an independent method. Both deletions described here were also detectable by quantitative real-time PCR (data not shown). Like MLPA, the Mapping 500 K Array Set cannot detect balanced rearrangements and is sensitive to handling and DNA quality in our experience. Deletions that affect only a small proportion of templates (mosaicism) would also be expected to present detection problems for both MLPA and SNP array technologies.
The deletions presented here generated junction fragments with short stretches of identical sequences at the sites of breakpoints (Figs. 2b, 3b), a phenomenon that has been described not only in FBN1 but also in other genes (Giacalone and Francke 1992; Otto et al. 2000; Liu et al. 2001). The precise mechanism of this phenomenon, however, remains to be elucidated. Both deletions comprise exon 1 and a ∼4-kb upstream region of FBN1, a surprising finding not reported before, which is most likely due to the limitations of previously used screening methods. A priori, one might expect that these deletions affect the regulatory or promoter regions of FBN1. Indeed, both deleted regions include the three alternatively spliced exons of the FBN1 gene (C, A, and B; Fig. 2a), which are thought to have regulatory function (Corson et al. 1993; Biery et al. 1999). Furthermore, although little is known about the FBN1 promoter region, very recent studies indicates that the FBN1 gene is controlled by a single promoter with a start site located up to 500 bases upstream (5′) of the ATG start codon (11th International Congress of Human Genetics, http://www.ichg2006.com/abstract/488.htm), which is affected by both deletions as well.
In general, no transcript will be produced when the promoter is deleted, unless alternative down- or up-stream promoter is activated. The latter occurs when a deletion fuses two genes that are in the same transcriptional orientation. Since the deletion in Patient 70 involves also the neighboring CEP152 gene, which encodes the centrosomal protein of 152 kDa with unknown function (Fig. 3a), we made use of a polymorphism in the FBN1 3′UTR to determine whether or not both alleles are transcribed. In Patient 70, sequence analysis of RT-PCR products revealed transcripts from only one allele (Fig. 4). This finding is consistent with the expectation that the deletion involves the regulatory and promoter regions and thus prevents transcription. The impact of the partial deletion of CEP152, if any, is unknown. In Patient 44, the activation of an alternative cryptic promoter, e.g. downstream of the deletion in intron 1 of FBN1, could not be excluded. The quantification of the total amount of FBN1 transcripts was not within the scope of this study, but, similar to other modifying factors, differences in normal FBN1 expression may explain some of the clinical variability seen in Patients 44 and 70 (Table 1), as previously demonstrated in a family with MFS (Hutchinson et al. 2003). Although, except of the father of Patient 44, neither the parents nor other family members (with no hints for MFS by history) were available for genetic testing, negative family histories suggest de novo occurrence of the deletions.
Interestingly, neither Patient 44 nor Patient 70 presents any sign of ectopia lentis. This is in accordance with the observation that MFS patients with a premature termination codon (PTC) mutation, which can lead to functional haploinsufficiency, i.e., preferential degradation of mutant transcripts due to nonsense-mediated mRNA decay (NMD), have a distinctly lesser chance of developing ectopia lentis than individuals with FBN1 cysteine substitutions (Schrijver et al. 2002; Rommel et al. 2005). However, the clinical findings in Patients 44 and 70 could also be explained by the lack of ocular involvement seen in about one third of MFS patients. In this context, mutations resulting in functional haploinsufficiency can also be associated with mild phenotypes which may fail to meet the diagnostic criteria for MFS (Dietz et al. 1993). However, such association between mutant transcript abundance and disease severity could not be found in Patients 44 and 70, similar to previously reported patients with classic severe MFS who harbored very low levels of PTC-containing mutant transcripts (Halliday et al. 1999).
Taken together, Patient 70, and most likely Patient 44 as well, have only a single functional FBN1 allele, representing true haploinsufficiency. Since the first description of an FBN1 mutation (Dietz et al. 1991), the role of haploinsufficiency in the pathogenesis of MFS has been the subject of intensive investigations and discussions. Mouse models suggested a critical threshold of functional microfibrils in the disease presentation of MFS (Pereira et al. 1999) and provided evidence for a critical contribution of haploinsufficiency by showing that half-normal amounts of fibrillin-1 can be insufficient to initiate productive microfibrillar assembly (Dietz and Mecham 2000; Judge et al. 2004). Furthermore, mouse MFS models showed that mice haploinsufficient in fibrillin-1 have marked dysregulation of transforming growth factor beta (TGF-beta) signaling (Neptune et al. 2003), most likely due to the fact that fibrillin-1 regulates the bioavailability of TGF-beta 1 (Chaudhry et al. 2007). In humans, although several observations supported a traditional dominant-negative mechanism, it has been suggested that functional haploinsufficiency through NMD of most of the mutant mRNAs plays a role in the pathogenic mechanism of MFS (Dietz et al. 1993; Hewett et al. 1994; Nijbroek et al. 1995; Pepe et al. 2001; Caputi et al. 2002; Judge and Dietz 2005; Robinson et al. 2006). However, since mutant transcripts being subject to NMD are often not completely degraded, it has remained unclear, whether the potent dominant-negative effect of the remaining mutant transcripts leads to the disease. Even cytogenetically detectable heterozygous deletions of the entire FBN1 gene, although suggested, failed to demonstrate the role of true haploinsufficiency in MFS, because these chromosomal abnormalities affect not only FBN1 but also several other genes, resulting in different phenotypes do not meet the Ghent criteria (e.g., Hutchinson et al. 2003; Adès et al. 2006). Hence, the findings in this study demonstrate for the first time that true haploinsufficiency, i.e., the complete loss of function of one FBN1 allele, is sufficient to cause MFS in patients. In such cases, increased expression from the normal FBN1 allele may represent a novel therapeutic avenue.
References
Adès LC, Sullivan K, Biggin A, Haan EA, Brett M, Holman KJ, Dixon J, Robertson S, Holmes AD, Rogers J, Bennetts B (2006) FBN1, TGFBR1, and the Marfan-craniosynostosis/mental retardation disorders revisited. Am J Med Genet A 140:1047–1058
Biery NJ, Eldadah ZA, Moore CS, Stetten G, Spencer F, Dietz HC (1999) Revised genomic organization of FBN1 and significance for regulated gene expression. Genomics 56:70–77
Caputi M, Kendzior RJ Jr, Beemon KL (2002) A nonsense mutation in the fibrillin-1 gene of a Marfan syndrome patient induces NMD and disrupts an exonic splicing enhancer. Genes Dev 16:1754–1759
Chaudhry SS, Cain SA, Morgan A, Dallas SL, Shuttleworth CA, Kielty CM (2007) Fibrillin-1 regulates the bioavailability of TGFbeta1. J Cell Biol 176:355–67
Corson GM, Chalberg SC, Dietz HC, Charbonneau NL, Sakai LY (1993) Fibrillin binds calcium and is coded by cDNAs that reveal a multidomain structure and alternatively spliced exons at the 5′ end. Genomics 17:476–484
De Paepe A, Devereux RB, Dietz HC, Hennekam RCM, Pyeritz RE (1996) Revised diagnostic criteria for the Marfan syndrome. Am J Med Genet 62:417–426
Dierlamm J, Schilling G, Michaux L, Hinz K, Penas EM, Seeger D, Hagemeijer A, Hossfeld DK (2003) Deletion of chromosome 15 represents a rare but recurrent chromosomal abnormality in myelocytic malignancies. Cancer Genet Cytogenet 144:1–5
Dietz HC, Mecham RP (2000) Mouse models of genetic diseases resulting from mutations in elastic fiber proteins. Matrix Biol 19:481–488
Dietz HC, Cutting GR, Pyeritz RE, Maslen CL, Sakai LY, Corson GM, Puffenberger EG, Hamosh A, Nanthakumar EJ, Curristin SM, Stetten G, Meyers DA, Francomano CA (1991) Marfan syndrome caused by a recurrent de novo missense mutation in the fibrillin gene. Nature 352:337–339
Dietz HC, McIntosh I, Sakai LY, Corson GM, Chalberg SC, Pyeritz RE, Francomano CA (1993) Four novel FBN1 mutations: significance for mutant transcript level and EGF-like domain calcium binding in the pathogenesis of Marfan syndrome. Genomics 17:468–475
Fukushima Y, Wakui K, Nishida T, Nishimoto H (1990) Craniosynostosis in an infant with an interstitial deletion of 15q [46,XY,del(15)(q15q22.1)]. Am J Med Genet 36:209–213
Giacalone JP, Francke U (1992) Common sequence motifs at the rearrangement sites of a constitutional X/autosome translocation and associated deletion. Am J Hum Genet 50:725–741
Halliday D, Hutchinson S, Kettle S, Firth H, Wordsworth P, Handford PA (1999) Molecular analysis of eight mutations in FBN1. Hum Genet 105:587–597
Hewett D, Lynch J, Child A, Firth H, Sykes B (1994) Differential allelic expression of a fibrillin gene (FBN1) in patients with Marfan syndrome. Am J Hum Genet 55:447–452
Hubbell E, Liu WM, Mei R (2002) Robust estimators for expression analysis. Bioinformatics 18:1585–1592
Hutchinson S, Furger A, Halliday D, Judge DP, Jefferson A, Dietz HC, Firth H, Handford PA (2003) Allelic variation in normal human FBN1 expression in a family with Marfan syndrome: a potential modifier of phenotype? Hum Mol Genet 12:2269–2276
Judge DP, Dietz HC (2005) Marfan’s syndrome. Lancet 366:1965–1976
Judge DP, Biery NJ, Keene DR, Geubtner J, Myers L, Huso DL, Sakai LY, Dietz HC (2004) Evidence for a critical contribution of haploinsufficiency in the complex pathogenesis of Marfan syndrome. J Clin Invest 114:172–181
Kainulainen K, Sakai LY, Child A, Pope FM, Puhakka L, Ryhanen L, Palotie A, Kaitila I, Peltonen L (1992) Two mutations in Marfan syndrome resulting in truncated fibrillin polypeptides. Proc Natl Acad Sci USA 89:5917–5921
Liu W, Schrijver I, Brenn T, Furthmayr H, Francke U (2001) Multi-exon deletions of the FBN1 gene in Marfan syndrome. BMC Med Genet 2:11
Loeys B, De Backer J, Van Acker P, Wettinck K, Pals G, Nuytinck L, Coucke P, De Paepe A (2004) Comprehensive molecular screening of the FBN1 gene favors locus homogeneity of classical Marfan syndrome. Hum Mutat 24:140–146
Loeys BL, Chen J, Neptune ER, Judge DP, Podowski M, Holm T, Meyers J, Leitch CC, Katsanis N, Sharifi N, Xu FL, Myers LA, Spevak PJ, Cameron DE, De Backer J, Hellemans J, Chen Y, Davis EC, Webb CL, Kress W, Coucke P, Rifkin DB, De Paepe AM, Dietz HC (2005) A syndrome of altered cardiovascular, craniofacial, neurocognitive and skeletal development caused by mutations in TGFBR1 or TGFBR2. Nat Genet 37:275–281
Loeys BL, Schwarze U, Holm T, Callewaert BL, Thomas GH, Pannu H, De Backer JF, Oswald GL, Symoens S, Manouvrier S, Roberts AE, Faravelli F, Greco MA, Pyeritz RE, Milewicz DM, Coucke PJ, Cameron DE, Braverman AC, Byers PH, De Paepe AM, Dietz HC (2006) Aneurysm syndromes caused by mutations in the TGF-beta receptor. N Engl J Med 355:788–798
Mátyás G, De Paepe A, Halliday D, Boileau C, Pals G, Steinmann B (2002a) Evaluation and application of denaturing HPLC for mutation detection in Marfan syndrome: identification of 20 novel mutations and two novel polymorphisms in the FBN1 gene. Hum Mutat 19:443–456
Mátyás G, Giunta C, Steinmann B, Hossle JP, Hellwig R (2002b) Quantification of single nucleotide polymorphisms: a novel method that combines primer extension assay and capillary electrophoresis. Hum Mutat 19:58–68
Mátyás G, Arnold E, Carrel T, Baumgartner D, Boileau C, Berger W, Steinmann B (2006) Identification and in silico analyses of novel TGFBR1 and TGFBR2 mutations in Marfan syndrome-related disorders. Hum Mutat 27:760–769
Mizuguchi T, Collod-Beroud G, Akiyama T, Abifadel M, Harada N, Morisaki T, Allard D, Varret M, Claustres M, Morisaki H, Ihara M, Kinoshita A, Yoshiura K, Junien C, Kajii T, Jondeau G, Ohta T, Kishino T, Furukawa Y, Nakamura Y, Niikawa N, Boileau C, Matsumoto N (2004) Heterozygous TGFBR2 mutations in Marfan syndrome. Nat Genet 36:855–860
Neptune ER, Frischmeyer PA, Arking DE, Myers L, Bunton TE, Gayraud B, Ramirez F, Sakai LY, Dietz HC (2003) Dysregulation of TGF-beta activation contributes to pathogenesis in Marfan syndrome. Nat Genet 33:407–11
Nijbroek G, Sood S, McIntosh I, Francomano CA, Bull E, Pereira L, Ramirez F, Pyeritz RE, Dietz HC (1995) Fifteen novel FBN1 mutations causing Marfan syndrome detected by heteroduplex analysis of genomic amplicons. Am J Hum Genet 57:8–21
Otto E, Betz R, Rensing C, Schatzle S, Kuntzen T, Vetsi T, Imm A, Hildebrandt F (2000) A deletion distinct from the classical homologous recombination of juvenile nephronophthisis type 1 (NPH1) allows exact molecular definition of deletion breakpoints. Hum Mutat 16:211–223
Pannu H, Fadulu VT, Chang J, Lafont A, Hasham SN, Sparks E, Giampietro PF, Zaleski C, Estrera AL, Safi HJ, Shete S, Willing MC, Raman CS, Milewicz DM (2005) Mutations in transforming growth factor-beta receptor type II cause familial thoracic aortic aneurysms and dissections. Circulation 112:513–520
Pepe G, Giusti B, Evangelisti L, Porciani MC, Brunelli T, Giurlani L, Attanasio M, Fattori R, Bagni C, Comeglio P, Abbate R, Gensini GF (2001) Fibrillin-1 (FBN1) gene frameshift mutations in Marfan patients: genotype-phenotype correlation. Clin Genet 59:444–450
Pereira L, Lee SY, Gayraud B, Andrikopoulos K, Shapiro SD, Bunton T, Biery NJ, Dietz HC, Sakai LY, Ramirez F (1999) Pathogenetic sequence for aneurysm revealed in mice underexpressing fibrillin-1. Proc Natl Acad Sci USA 96:3819–3823
Pramparo T, Mattina T, Gimelli S, Liehr T, Zuffardi O (2005) Narrowing the deleted region associated with the 15q21 syndrome. Eur J Med Genet 48:346–352
Qiu P, Soder GJ, Sanfiorenzo VJ, Wang L, Greene JR, Fritz MA, Cai XY (2003) Quantification of single nucleotide polymorphisms by automated DNA sequencing. Biochem Biophys Res Commun 309:331–338
Rabbee N, Speed TP (2006) A genotype calling algorithm for affymetrix SNP arrays. Bioinformatics 22:7–12
Robinson PN, Arteaga-Solis E, Baldock C, Collod-Beroud G, Booms P, De Paepe A, Dietz HC, Guo G, Handford PA, Judge DP, Kielty CM, Loeys B, Milewicz DM, Ney A, Ramirez F, Reinhardt DP, Tiedemann K, Whiteman P, Godfrey M (2006) The molecular genetics of Marfan syndrome and related disorders. J Med Genet 43:769–787
Rommel K, Karck M, Haverich A, von Kodolitsch Y, Rybczynski M, Muller G, Singh KK, Schmidtke J, Arslan-Kirchner M (2005) Identification of 29 novel and nine recurrent fibrillin-1 (FBN1) mutations and genotype-phenotype correlations in 76 patients with Marfan syndrome. Hum Mutat 26:529–539
Schouten JP, McElgunn CJ, Waaijer R, Zwijnenburg D, Diepvens F, Pals G (2002) Relative quantification of 40 nucleic acid sequences by multiplex ligation-dependent probe amplification. Nucleic Acids Res 30:e57
Schrijver I, Liu W, Odom R, Brenn T, Oefner P, Furthmayr H, Francke U (2002) Premature termination mutations in FBN1: Distinct effects on differential allelic expression and on protein and clinical phenotypes. Am J Hum Genet 71:223–237
Shur N, Cowan J, Wheeler PG (2003) Craniosynostosis and congenital heart anomalies associated with a maternal deletion of 15q15-22.1. Am J Med Genet A 120:542–546
Singh KK, Elligsen D, Liersch R, Schubert S, Pabst B, Arslan-Kirchner M, Schmidtke J (2007) Multi-exon out of frame deletion of the FBN1 gene leading to a severe juvenile onset cardiovascular phenotype in Marfan syndrome. J Mol Cell Cardiol 42:352–356
Acknowledgments
We are grateful to the patients, the father of Patient 44, and the referring physicians for participating in this study. We thank Marzanna Künzli, Ulrich Wagner, Philippe Reuge for technical support and assistance; Angelika Schwarze for cell cultures; Melanie Maudrich for initial mutation analysis of the FBN1 gene; members of the Institute of Medical Genetics, University of Zurich, for constructive discussions. This work was supported by the Functional Genomics Center Zurich and grants from the Foundation for Research at the Medical Faculty and Research Funding of the University of Zurich (to G.M.), Swiss Heart Foundation (to G.M.), Swiss National Science Foundation (NF 3200B0-109370/1 to B.S.), and the Wolfermann-Nägeli-Stiftung Zurich (to B.S.).
Author information
Authors and Affiliations
Corresponding author
Additional information
Gábor Mátyás and Sira Alonso have contributed equally to this work.
Rights and permissions
About this article
Cite this article
Mátyás, G., Alonso, S., Patrignani, A. et al. Large genomic fibrillin-1 (FBN1) gene deletions provide evidence for true haploinsufficiency in Marfan syndrome. Hum Genet 122, 23–32 (2007). https://doi.org/10.1007/s00439-007-0371-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00439-007-0371-x