Abstract
Adaptation to exercise training is a complex trait that may be influenced by genetic variants. We identified 36 single nucleotide polymorphisms (SNPs) that had been previously associated with endurance or strength performance, exercise-related phenotypes or exercise intolerant disorders. A MassARRAY multiplex genotyping assay was designed to identify associations with these SNPs against collected endurance fitness phenotype parameters obtained from two exercise cohorts (Gene SMART study; n = 58 and Hawaiian Ironman Triathlon 2008; n = 115). These parameters included peak power output (PP), a time trial (TT), lactate threshold (LT), maximal oxygen uptake (VO2 max) in recreationally active individuals and a triathlon time-to-completion (Hawaiian Ironman Triathlon cohort only). A nominal significance threshold of α < 0.05 was used to identify 17 variants (11 in the Gene SMART population and six in the Hawaiian Ironman Triathlon cohort) which were significantly associated with performance gains in highly trained individuals. The variant rs1474347 located in Interleukin 6 (IL6) was the only variant with a false discovery rate < 0.05 and was found to be associated with gains in VO2 max (additional 4.016 mL/(kg min) for each G allele inherited) after training in the Gene SMART cohort. In summary, this study found further evidence to suggest that genetic variance can influence training response in a moderately trained cohort and provides an example of the potential application of genomic research in the assessment of exercise trait response.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Currently, robust identification of genetic variants associated with exercise phenotypes is limited by a lack of reproducible results. Family and twin studies have estimated high heritability for various exercise performance metrics (e.g. muscle mass: 40%, anaerobic power: 70–80%, aerobic exercise: 50%) (Issurin 2017). However, a wide variety of environmental (e.g. diet, sleep), psychological, and epigenetic factors may also influence exercise responses (den Hoed et al. 2013). In addition, within-subject variability (i.e. the variable response of a given individual to the same exercise training) considerably limits the identification of genetic variants with potentially small effects on exercise response (Voisin et al. 2019).
To date, only two genetic signatures have consistently shown an association with exercise responses; the Alpha-actinin-3 stop gain variant (ACTN3) p.Arg577Ter and the Angiotensin converting enzyme (ACE) insertion/deletion (I/D) in intron 16 (Houweling et al. 2018; Yan et al. 2018). Genome-wide association studies (GWAS) have helped discern genomic loci associated with training response; however, these usually contain a low number of participants, and/or evidence of association with exercise psychology-related phenotypes (Bouchard 2011). Studies in metabolic and cardiovascular disorders such as diabetes or arterial hypertension have further complicated participant ability to perform exercise training at duration and intensity and as such participants can be classified as having exercise intolerant disorders (McCoy et al. 2017). As exercise training yields a host of health benefits, understanding which genetic and molecular processes contribute to these responses might be helpful to the development of personalised exercise therapeutics (e.g. exercise dosing to minimise risk of adverse response within exercise intolerant disorders) (Buford et al. 2013).
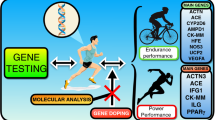
In this study, we investigated candidate genes previously implicated in exercise response in cohorts of varying fitness levels. We used a highly trained cohort (triathlon) and a moderately trained, longitudinal cohort of high-intensity interval endurance training (HIIT) exercise training. We hypothesised that many, if not all, of the candidate SNPs would be found to be associated with triathlon performance and response to 4 weeks of endurance exercise training, regardless of age or baseline fitness level.
Materials and methods
The Gene SMART cohort
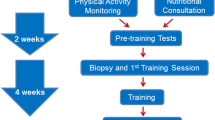
The Gene SMART (Skeletal Muscle Adaptive Response to Training) study design has previously been described (Yan et al. 2017). The study is ongoing with currently > 100 moderately trained participants who were sampled for blood and skeletal muscle (vastus lateralis) at several time points: before, immediately after and 3 h after a single bout of high-intensity endurance exercise (HIIE), and after 4 weeks of high-intensity interval training (HIIT) (Yan et al. 2017). Exercise-related phenotypic measurements were collected before and after the completion of the exercise training intervention (e.g. lactate threshold (LT, in Watts), peak power output (PP, in Watts), maximal oxygen uptake (VO2max, in mL/min/kg body weight, from graded exercise tests), and a time trial measurement (TT, in min). All participants gave informed consent, and the study was approved by the Victoria University Ethics Committee (Approval number: HRE13-233). Subsequently, the study was also approved by the Queensland University of Technology (QUT) Human Research Ethics Committee (Approval number: 1600000342). All procedures performed in studies involving human participants were in accordance with the ethical standards of the respective institutions research committees, and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards. At the time of collection, n = 77 participants had participated in the study, with n = 58 completing the entire 4-week training programme. Genomic DNA was extracted and purified from whole blood using the QIAamp DNA blood midi kit (Qiagen, Hilden, Germany) according to the manufacturer’s instructions for participants who completed the study. Samples that failed genotypic analysis or had a large amount of missing phenotypic data were removed from further analysis, leaving a final sample size of n = 52 (Age = 30.95 ± 8.17 years). In the moderately trained cohort (Gene SMART, n = 58), we focused on the response to an exercise training programme (longitudinal analysis). Specifically, we measured the change in (Δ = post − pre) measurement for each endurance fitness trait as a representation of response to exercise training.
Highly trained (Ironman) cohort
Ironman triathlons consist of a 3.86 km swim, a 180.25 km bike ride, followed by the completion of a full marathon (42.2 km). The 2008 Hawaiian Ironman Triathlon population has been previously described as an elite endurance cohort based on their eligibility and participation in the event (Grealy et al. 2013, 2015). Due to the intensity of this endurance event, only highly trained individuals that completed it were included in this study. To avoid genetic confounding, we analysed only the triathlon participants who self-identified as male and Caucasian (Age = 43.81 ± 11.39 years). This was performed solely on the triathlon group as the Gene SMART population was already homogeneously male. All procedures performed in studies involving human participants were in accordance with the ethical standards of the QUT human Research Ethics Committee (approval number: 1300000499), and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards. Saliva samples (OG-250 Oragene Kit, DNA Genotek Inc.) and questionnaires were collected prior to the event; time-to-completion measurements for each event were collected from the publicly available online event webpage. Genomic DNA was extracted as per manufacturer instructions and described previously (Grealy et al. 2013). In the highly trained cohort (Ironman, n = 115), we focused on endurance performance, the result of months or years of training (cross-sectional analysis). Specifically, we used the time-to-completion of the running event, the biking event, the swimming event, and the total event.
Genotype method and SNP selection
The SNPs investigated in this study (Table 1) were included based on conformity of one of three criteria. The first was that SNPs chosen had to have been previously associated with elite athletic status, exercise responses with reasonable replication, or exercise traits at baseline. This resulted in 11 SNPs chosen, though it should be noted that we were unable to genotype the ACE I/D variant (rs4340) using the MassARRAY and previous work failed to identify an association with baseline fitness levels in the Gene SMART cohort (Yan et al. 2018). The second criteria encompassed SNPs previously investigated but less consistently associated with performance, i.e. studies with equivalent numbers of negative studies or studies related to exercise psychology. The third criteria included SNPs associated with exercise intolerant disorders and non-exercise respiratory, muscular, or energy storage phenotypes such as hypertension (HT), cardiovascular disease (CVD), or type 2 diabetes mellitus (T2DM). The experimental methodology for sample preparation and genotype analysis was performed using the Agena Biosciences MassARRAY, a matrix-assisted laser desorption ionisation time-of-flight (MALDI-TOF) mass spectrometer, which has been described elsewhere (Jurinke et al. 2002; Ellis and Ong 2017). An internal genotyping control SNP (rs17602729, AMPD1), previously validated in our endurance cohort, was used to ensure the MassARRAY system correctly identified genotypes (Grealy et al. 2015).
Data processing
The output files from the MassARRAY platform were converted to PLINK format and analysed for correct genotypic identification (calling). For the Gene SMART and Ironman populations, respectively, SNPs were excluded from further analysis if they exceeded the following thresholds: (1) SNPs that had a calling rate < 80% (> 20% missing data) (n = 5, n = 3); (2) SNPs with a minor allele frequency < 2% (n = 1, n = 2); (3) SNPs determined not in Hardy–Weinberg equilibrium (n = 1, n = 1) (Ellis and Ong 2017). Subsequent analysis was performed on n = 29 SNPs for the Gene SMART population and n = 30 SNPs for the Ironman population.
Statistical analysis
We measured normality metrics (skewness and kurtosis) for each phenotype in both populations using the ggplot2, tidyverse and moments packages in R, to determine if data transformation was necessary from the raw phenotypic values. We used PLINK V1.90p to perform quantitative linear association tests (95% CI) with both dominant and recessive models for each cohort, adjusting for age. An additive model was considered but did not differ from the results obtained from the dominant model. As this was a candidate gene study, SNPs that had a raw p value < 0.05 were considered nominally significant, while variants that had an adjusted p value (Benjamini–Hochberg false discovery rate (FDR)) < 0.05 were considered significant. This adjustment method represents a good balance between type I and type II errors and as such minimises false positive results. To avoid multiple testing burdens with phenotypic traits, we used a separate hypothesis for each quantitative trait. Effect sizes were determined using raw beta regression coefficient values interpreted as “how much a specific phenotype increased for each additional X allele at the SNP of interest”.
Results
The array genotyping control (AMPD1) was identified to be 100% concordant with the genotyping results from another method (RFLP) in our previous study with the same population, confirming the validity of the MassARRAY data (Table 2).
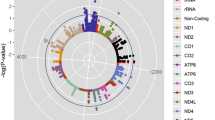
Six variants in five distinct genes were nominally associated with time-to-completion of Ironman events: Nuclear Respiratory Factor 1 (NRF1: rs6949152), Myostatin (MSTN: rs1805086), Major Histocompatibility complex class 1A (HLA-A: rs1061235), Peroxisome proliferator-activated receptor gamma coactivator 1-alpha (PPARGC1α: rs6821591, rs6821591), and SH3 Domain GRB2 Like Endophilin Interacting Protein (SGIP1: rs9633417). The results of our association testing did not change significantly when age was used as a covariate.
In the Gene SMART cohort, eleven variants in nine distinct genes were shown to be nominally associated with gains in endurance fitness following exercise training (Table 3): Adenosine Monophosphate Deaminase 1 (AMPD1: rs17602729), Iodothyronine Deiodinase 1 (DIO1: rs2294512), Bradykinin receptor B2 (BDKRB2: rs1799722), Nuclear Respiratory Factor 2 (NRF2: rs7181866, rs8031031), (COL6A1, rs39796750), Apolipoprotein E (APOE: rs7412), Interleukin 6 (IL6: rs1474347), Mitochondrial uncoupling protein 2 (UCP2: rs660339), and Homeostatic Iron Regulator (HFE: rs1799945).
Interestingly, no variants were identified to be significantly associated with both time-to-completion in the Ironman cohort and the response to endurance exercise training in the Gene SMART cohort. Only rs1474347 in IL6 passed correction for multiple testing using the BH-FDR method (FDR: 0.018). The C allele at rs1474347 was associated with VO2max response within the Gene SMART study with an effect size of − 4.016 mL/(kg min).
Discussion
In the present study, we have successfully replicated previously associated exercise-related SNPs using the combined data from highly trained and moderately trained cohorts. Our main findings identified the rs1474347 in the IL6 gene to be significantly associated with gains in VO2 max in the Gene SMART cohort after multiple testing statistical corrections. In addition, 17 genetic variants were found to be associated with either elite performance or responses to exercise; however, none of these variants were common between these cohorts.
Different genetic signatures likely confer different responses to exercise training via specific molecular pathways. Therefore, variants that influence pathways responsible for adaptation to moderate training may in part differ to those that confer response to high-intensity endurance training. Additionally, moderately trained cohorts typically contain individuals with large variability in environmental factors such as diet, sleep, and habitual physical activity patterns, while the inter-individual variability in these measures is smaller in highly trained cohorts and therefore less likely to confound results (Bonetti and Hopkins 2010).
Association between genetic variants and exercise responses in the Gene SMART cohort
Located in an intron of the IL6 gene, the rs1474347 variant has been previously associated with T2D traits in a large-scale study (n = 10,775). The IL6 protein is a pro-inflammatory cytokine with myokinetic (i.e. excreted from skeletal muscle) functions and is responsible for triggering and maintaining immune processes following post-exercise muscle damage (Munoz-Canoves et al. 2013). We found that the C allele at this locus negatively affected the exercise response to the VO2max phenotype (β = − 4.016 mL/kg min) and therefore a homozygous C/C genotype would result in a VO2max loss of − 8.032 mL/kg min. The rs1800795 coding variant within the IL6 gene has shown mixed evidence of exercise associations, i.e. variant C = athleticism, G = power (Ahmetov and Fedotovskaya 2015; Karanikolou et al. 2017). Interestingly, further analysis identified the rs1474347 C allele to be in strong linkage disequilibrium (LD) with the C allele of rs1800795 (R2 = 0.96). As such, it is feasible that the LD identified between these variants has contributed to the mixed evidence reported for association studies implicating the latter variant (rs1800795) in IL6 for exercise traits. We propose that the rs1474347 variant may reduce the expression of IL6 during acute muscle damage and therefore cause a reduced local immune response leading to loss of skeletal muscle remodelling and repair. In addition this variant is also located 2 kb upstream of an uncharacterised long non-coding RNA (lncRNA; LOC541472) and therefore, variants in this region may affect the IL-6 pro-inflammatory pathway or post-translational epigenetic and regulatory processes.
Association between genetic variants and Ironman performance
The run time (42.2 km marathon) and bike time (180.25 km ride) events in the triathlon are largely leg-based exercise activities, and therefore we expected a significant overlap of variants associated with these traits. In contrast, the triathlon swimming event utilises whole body muscle groups; therefore, SNPs seen in this test were anticipated to only be associated with this particular trait. Our findings supported this as the variants associated with the swim time event were not seen in either of the other isolated finishing times, or indeed the total time-to-completion trait. This is also supported by the current literature where elite runners and swimmers are not analysed collectively (Woods et al. 2001). We also note that the two variants nominally associated with the swim time trait are involved in hypoxic events characteristic of swimming (Galy et al. 2008). It is possible that the rs6949152 G allele within the NRF1 gene results in lower activity of NRF transcription factor and therefore increased levels of hypoxia-inducible factor 1 alpha (HIF1α). This would cause reduced oxidative metabolic processing and therefore lead to the increase in swim time that is associated with the variant (β = 0.2459 h). Additionally, the CKMM protein has been shown to exhibit protective effects during mild hypoxia and therefore hypothesised that the rs8111989 variant would increase the functionality of the CKMM protein, resulting in protection against re-oxygenation-induced muscle damage and decreased swim time (Zervou et al. 2017).
Although multiple SNPs examined in this study passed our nominal threshold for significance, which was unexpected given our relatively small sample sizes, all variants nominally significant in each cohort have previously been investigated as causative variants in multiple exercise studies.
Using a MassARRAY design of 36 SNPs, we found a significant association for the rs1474347 SNP in IL6 with the change in VO2max trait in a cohort of moderately trained individuals. Furthermore, 16 other SNPs were shown to have nominal association with exercise response in the Gene SMART cohort, or Ironman performance in highly trained athletes. As such, these markers may be useful in the development of tailored genetic panel screening and therapeutics in sports science, and exercise intolerant disorders. However, to more fully exploit their applicability in this context, confirmation of the genotypic phenotype on gene function is required. While this is outside the purview of this study, we have successfully replicated the significance of several exercise genes in two relatively small exercise study cohorts through nominally significant associations identified in the study cohorts. We were also able to implicate and ascertain directionality of SNPs between the different phenotypic traits. Additionally, the different variants associated with each cohort highlight the need to examine multiple cohorts of differing fitness levels and training capabilities. However, more replication studies are required in conjunction with functional transcriptomic/proteomic studies to confirm the genes and pathways associated with exercise adaptations. The use of multi-centre studies and consortia, such as the Athlome study consortium, would be helpful to better facilitate these efforts to further develop the field of exercise genomics research (Pitsiladis et al. 2016).
Data availability
The datasets generated and analysed in this study will be provided by the corresponding author upon reasonable request.
References
Ahmetov II, Fedotovskaya ON (2015) Current progress in sports genomics. Adv Clin Chem 70:247–314
Alves AJ, Goldhammer E, Ribeiro F, Eynon N, Cohen SBZ, Duarte JA, Viana JL, Sagiv M, Oliveira J (2013) GNAS A-1121G variant is associated with improved diastolic dysfunction in response to exercise training in heart failure patients. Int J Sports Med 34:274–280
Andrulionyte L, Kuulasmaa T, Chiasson JL, Laakso M, Group S-NS (2007) Single nucleotide polymorphisms of the peroxisome proliferator-activated receptor-alpha gene (PPARA) influence the conversion from impaired glucose tolerance to type 2 diabetes: the STOP-NIDDM trial. Diabetes 56:1181–1186
Arora P, Garcia-Bailo B, Dastani Z, Brenner D, Villegas A, Malik S, Spector TD, Richards B, El-Sohemy A, Karmali M, Badawi A (2011) Genetic polymorphisms of innate immunity-related inflammatory pathways and their association with factors related to type 2 diabetes. BMC Med Genet 12:95
Bonetti DL, Hopkins WG (2010) Variation in performance times of elite flat-water canoeists from race to race. Int J Sports Physiol Perform 5:210–217
Bouchard C (2011) Overcoming barriers to progress in exercise genomics. Exerc Sport Sci Rev 39:212–217
Brondani LA, Assmann TS, Duarte GC, Gross JL, Canani LH, Crispim D (2012) The role of theuncoupling protein 1 (UCP1) on the development of obesity and type 2 diabetes mellitus. Arq Bras Endocrinol Metabol 56:215–225
Brown BM, Rainey-Smith SR, Castalanelli N, Gordon N, Markovic S, Sohrabi HR, Weinborn M, Laws SM, Doecke J, Shen K, Martins RN, Peiffer JJ (2017) Study protocol of the Intense Physical Activity and Cognition study: The effect of high-intensity exercise training on cognitive function in older adults. Alzheimers Dement (N Y) 3:562–570
Burguete-Garcia AI, Martinez-Nava GA, Valladares-Salgado A, Bermudez Morales VH, Estrada-Velasco B, Wacher N, Peralta-Romero J, Garcia-Mena J, Parra E, Cruz M (2014) Association of beta1 and beta3 adrenergic receptors gene polymorphisms with insulin resistance and high lipid profiles related to type 2 diabetes and metabolic syndrome. Nutr Hosp 29:1327–1334
Buford TW, Roberts MD, Church TS (2013) Toward exercise as personalized medicine. Sports Med 43:157–165
Chathoth S, Ismail MH, Vatte C, Cyrus C, Al Ali Z, Ahmed KA, Acharya S, Al Barqi AM, Al Ali A (2018) Association of uncoupling protein 1 (UCP1) gene polymorphism with obesity: a case-control study. BMC Med Genet 19:203
De Moor MH, Liu YJ, Boomsma DI, Li J, Hamilton JJ, Hottenga JJ, Levy S, Liu XG, Pei YF, Posthuma D, Recker RR, Sullivan PF, Wang L, Willemsen G, Yan H, De Geus EJ, Deng HW (2009) Genomewide association study of exercise behavior in Dutch and American adults. Med Sci Sports Exerc 41:1887–1895
Deeny SP, Poeppel D, Zimmerman JB, Roth SM, Brandauer J, Witkowski S, Hearn JW, Ludlow AT, Contreras-Vidal JL, Brandt J, Hatfield BD (2008) Exercise, APOE, and working memory: MEG and behavioral evidence for benefit of exercise in epsilon4 carriers. Biol Psychol 78:179–187
Doney AS, Fischer B, Lee SP, Morris AD, Leese G, Palmer CN (2005) Association of common variation in the PPARA gene with incident myocardial infarction in individuals with type 2 diabetes: a Go-DARTS study. Nucl Recept 3:4
Dong C, Zhou H, Shen C, Yu LG, Ding Y, Zhang YH, Guo ZR (2015) Role of peroxisome proliferatoractivated receptors gene polymorphisms in type 2 diabetes and metabolic syndrome. World J Diabetes 6:654–661
den Hoed M, Brage S, Zhao JH, Westgate K, Nessa A, Ekelund U, Spector TD, Wareham NJ, Loos RJ (2013) Heritability of objectively assessed daily physical activity and sedentary behavior. Am J Clin Nutr 98:1317–1325
Ellis JA, Ong B (2017) The MassARRAY((R)) system for targeted SNP genotyping. Methods Mol Biol 1492:77–94
Evans D, Aberle J, Wendt D, Wolf A, Beisiegel U, Mann WA (2001) A polymorphism, L162V, in the peroxisome proliferator-activated receptor alpha (PPARalpha) gene is associated with lower body mass index in patients with non-insulin-dependent diabetes mellitus. J Mol Med (Berl) 79:198–204
Flavell DM, Ireland H, Stephens JW, Hawe E, Acharya J, Mather H, Hurel SJ, Humphries SE (2005) Peroxisome proliferator-activated receptor alpha gene variation influences age of onset and progression of type 2 diabetes. Diabetes 54:582–586
Fredriksson J, Anevski D, Almgren P, Sjogren M, Lyssenko V, Carlson J, Isomaa B, Taskinen MR, Groop L, Orho-Melander M, Botnia Study G (2007) Variation in GYS1 interacts with exercise andgender to predict cardiovascular mortality. PLoS One 2:e285
Fuku N, Alis R, Yvert T, Zempo H, Naito H, Abe Y, Arai Y, Murakami H, Miyachi M, Pareja-Galeano H, Emanuele E, Hirose N, Lucia A (2016) Muscle-Related Polymorphisms (MSTN rs1805086 and ACTN3 rs1815739) Are Not Associated with Exceptional Longevity in Japanese Centenarians. PLoS One 11:e0166605
Galy O, Hue O, Chamari K, Boussana A, Chaouachi A, Prefaut C (2008) Influence of performance level on exercise-induced arterial hypoxemia during prolonged and successive exercise in triathletes. Int J Sports Physiol Perform 3:482–500
Garatachea N, Santiago C, Yvert T, Verde-Rello Z, Fiuza-Luces C, Santos-Lozano A, Gomez-Gallego F, Lucia A (2015) Genetic variants in the PPARD-PPARGC1A-NRF-TFAM mitochondriogenesis pathway are neither associated with muscle characteristics nor physical performance in elderly. Ricyde-Revista Internacional De Ciencias Del Deporte 11:196–208
Grealy R, Smith CL, Chen T, Hiller D, Haseler LJ, Griffiths LR (2013) The genetics of endurance: frequency of the ACTN3 R577X variant in Ironman World Championship athletes. J Sci Med Sport 16:365–371
Grealy R, Herruer J, Smith CL, Hiller D, Haseler LJ, Griffiths LR (2015) Evaluation of a 7-gene genetic profile for athletic endurance phenotype in ironman championship triathletes. PLoS ONE 10:e0145171
He Z, Hu Y, Feng L, Li Y, Liu G, Xi Y, Wen L, Lucia A (2008) NRF-1 genotypes and endurance exercise capacity in young Chinese men. British J of Sports Med 42(5):361–366
He ZH, Hu Y, Li YC, Gong LJ, Cieszczyk P, Maciejewska-Karlowska A, Leonska-Duniec A, Muniesa CA, Marin-Peiro M, Santiago C, Garatachea N, Eynon N, Lucia A (2015) PGC-related gene variants and elite endurance athletic status in a Chinese cohort: a functional study. Scand J Med Sci Sports 25:184–195
Hennis PJ, O'Doherty AF, Levett DZ, Grocott MP, Montgomery HM (2015) Genetic factors associated with exercise performance in atmospheric hypoxia. Sports Med 45:745–761
Houweling PJ, Papadimitriou ID, Seto JT, Perez LM, Coso JD, North KN, Lucia A, Eynon N (2018) Is evolutionary loss our gain? The role of ACTN3 p.Arg577Ter (R577X) genotype in athletic performance, ageing, and disease. Hum Mutat 39:1774–1787
Issurin VB (2017) Evidence-based prerequisites and precursors of athletic talent: a review. Sports Med 47:1993–2010
Jurinke C, van den Boom D, Cantor CR, Koster H (2002) Automated genotyping using the DNA MassArray technology. Methods Mol Biol 187:179–192
Karanikolou A, Wang G, Pitsiladis Y (2017) Letter to the editor: a genetic-based algorithm for personalized resistance training. Biol Sport 34:31–33
Liang KY, Mintun MA, Fagan AM, Goate AM, Bugg JM, Holtzman DM, Morris JC, Head D (2010) Exercise and Alzheimer's disease biomarkers in cognitively normal older adults. Ann Neurol 68:311–318
Lin Y, Gu SJ, Wu M, Chen Q, Zhou ZY, Yu H, Zhang LJ, Luo WS, Guo ZR (2012) Association between peroxisome proliferator-activated receptors gene polymorphism and essential hypertension. Zhonghua Liu Xing Bing Xue Za Zhi 33:597–601
Liu Y, Berthier-Schaad Y, Fallin MD, Fink NE, Tracy RP, Klag MJ, Smith MW, Coresh J (2006) IL-6 haplotypes, inflammation, and risk for cardiovascular disease in a multiethnic dialysis cohort. J Am Soc Nephrol 17:863–870
McCoy J, Bates M, Eggett C, Siervo M, Cassidy S, Newman J, Moore SA, Gorman G, Trenell MI, Velicki L, Seferovic PM, Cleland JGF, MacGowan GA, Turnbull DM, Jakovljevic DG (2017) Pathophysiology of exercise intolerance in chronic diseases: the role of diminished cardiac performance in mitochondrial and heart failure patients. Open Heart 4:e000632
Montani D, Girerd B, Gunther S, Riant F, Tournier-Lasserve E, Magy L, Maazi N, Guignabert C, Savale L, Sitbon O, Simonneau G, Soubrier F, Humbert M (2014) Pulmonary arterial hypertension in familial hemiplegic migraine with ATP1A2 channelopathy. Eur Respir J 43:641–643
Munoz-Canoves P, Scheele C, Pedersen BK, Serrano AL (2013) Interleukin-6 myokine signaling in skeletal muscle: a double-edged sword? FEBS J 280:4131–4148
Panicker V, Cluett C, Shields B, Murray A, Parnell KS, Perry JR, Weedon MN, Singleton A, Hernandez D, Evans J, Durant C, Ferrucci L, Melzer D, Saravanan P, Visser TJ, Ceresini G, Hattersley AT, Vaidya B, Dayan CM, Frayling TM (2008) A common variation in deiodinase 1 gene DIO1 is associated with the relative levels of free thyroxine and triiodothyronine. J Clin Endocrinol Metab 93:3075–3081
Panoulas VF, Douglas KM, Smith JP, Taffe P, Stavropoulos-Kalinoglou A, Toms TE, Elisaf MS, Nightingale P, Kitas GD (2008) Polymorphisms of the endothelin-1 gene associate with hypertension in patients with rheumatoid arthritis. Endothelium 15:203–212
Pitsiladis YP, Tanaka M, Eynon N, Bouchard C, North KN, Williams AG, Collins M, Moran CN, Britton SL, Fuku N, Ashley EA, Klissouras V, Lucia A, Ahmetov II, de Geus E, Alsayrafi M, Athlome Project C (2016) Athlome Project Consortium: a concerted effort to discover genomic and other “omic” markers of athletic performance. Physiol Genomics 48:183–190
Ramos AV, Bastos-Rodrigues L, Resende BA, Friedman E, Campanha-Versiani L, Miranda DM, Sarquis M, De Marco L (2012) The contribution of FTO and UCP-1 SNPs to extreme obesity, diabetes and cardiovascular risk in Brazilian individuals. BMC Med Genet 13:101
Rankinen T, Perusse L, Rauramaa R, Rivera MA, Wolfarth B, Bouchard C (2004) The human gene map for performance and health-related fitness phenotypes: the 2003 update. Med Sci Sports Exerc 36:1451–1469
Rankinen T, Church T, Rice T, Markward N, Leon AS, Rao DC, Skinner JS, Blair SN, Bouchard C (2007) Effect of endothelin 1 genotype on blood pressure is dependent on physical activity or fitness levels. Hypertension 50:1120–1125
Tejedor MT, Garcia-Sobreviela MP, Ledesma M, Arbones-Mainar JM (2014) The apolipoprotein E polymorphism rs7412 associates with body fatness independently of plasma lipids in middle aged men. PLoS One 9:e108605
Uthurralt J, Gordish-Dressman H, Bradbury M, Tesi-Rocha C, Devaney J, Harmon B, Reeves EK, Brandoli C, Hansen BC, Seip RL, Thompson PD, Price TB, Angelopoulos TJ, Clarkson PM, Moyna NM, Pescatello LS, Visich PS, Zoeller RF, Gordon PM, Hoffman EP (2007) PPARalpha L162V underlies variation in serum triglycerides and subcutaneous fat volume in young males. BMC Med Genet 8:55
Voisin S, Jacques M, Lucia A, Bishop DJ, Eynon N (2019) Statistical considerations for exercise protocols aimed at measuring trainability. Exerc Sport Sci Rev 47:37–45
Walsh S, Haddad CJ, Kostek MA, Angelopoulos TJ, Clarkson PM, Gordon PM, Moyna NM, Visich PS, Zoeller RF, Seip RL, Bilbie S, Thompson PD, Devaney J, Gordish-Dressman H, Hoffman EP, Price TB, Pescatello LS (2012) Leptin and leptin receptor genetic variants associate with habitual physical activity and the arm body composition response to resistance training. Gene 510:66–70
Woods D, Hickman M, Jamshidi Y, Brull D, Vassiliou V, Jones A, Humphries S, Montgomery H (2001) Elite swimmers and the D allele of the ACE I/D polymorphism. Hum Genet 108:230–232
Yan X, Eynon N, Papadimitriou ID, Kuang J, Munson F, Tirosh O, O’Keefe L, Griffiths LR, Ashton KJ, Byrne N, Pitsiladis YP, Bishop DJ (2017) The gene SMART study: method, study design, and preliminary findings. BMC Genomics 18:821
Yan X, Dvir N, Jacques M, Cavalcante L, Papadimitriou ID, Munson F, Kuang J, Garnham A, Landen S, Li J, O’Keefe L, Tirosh O, Bishop DJ, Voisin S, Eynon N (2018) ACE I/D gene variant predicts ACE enzyme content in blood but not the ACE, UCP2, and UCP3 protein content in human skeletal muscle in the Gene SMART study. J Appl Physiol 125:923–930
Zervou S, Whittington HJ, Ostrowski PJ, Cao F, Tyler J, Lake HA, Neubauer S, Lygate CA (2017) Increasing creatine kinase activity protects against hypoxia/reoxygenation injury but not against anthracycline toxicity in vitro. PLoS ONE 12:e0182994
Acknowledgements
The authors would like to acknowledge Dr. Cassie Albury for assistance with the molecular methods used in this manuscript.
Funding
This research was chiefly supported by the Commonwealth Collaborative Research Network funding to Bond University CRN-AESS. Mr. Nicholas Harvey was supported by a Ph.D. stipend also provided by Bond University CRN-AESS. This research was also supported by infrastructure purchased with Australian Government EIF Super Science Funds as part of the Therapeutic Innovation Australia—Queensland Node project (LRG). Nir Eynon is supported by the National Health and Medical Research Council, Australia (NHMRC CDF# APP1140644).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflicts of interest
The authors declare that they have no conflict of interest.
Ethical approval
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee (Gene SMART study: VU human research ethics committee: HRE13-233; QUT human research ethics committee: 1600000342; Hawaiian Ironman triathlon study: QUT Human Research Ethics Committee: 1300000499) and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards.
Additional information
Communicated by Stefan Hohmann.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Harvey, N.R., Voisin, S., Dunn, P.J. et al. Genetic variants associated with exercise performance in both moderately trained and highly trained individuals. Mol Genet Genomics 295, 515–523 (2020). https://doi.org/10.1007/s00438-019-01639-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00438-019-01639-8