Abstract
The previously published data on the association between CYP1A2*1C (rs2069514) and CYP1A2*1F (rs762551) polymorphisms and cancer risk have remained controversial. Hence, we performed a meta-analysis to investigate the association between CYP1A2*1F and CYP1A2*1C polymorphisms and cancer risk under different inheritance models. Overall, significant association was observed between CYP1A2*1F and cancer risk when all the eligible studies were pooled into the meta-analysis (dominant model: OR 1.08, 95 % CI 1.02–1.15; heterozygous model: OR 1.06, 95 % CI 1.01–1.12; additive model: OR 1.07, 95 % CI 1.02–1.13). In the further stratified and sensitivity analyses, for CYP1A2*1F polymorphism, significantly increased lung cancer risk and significantly decreased bladder cancer risk were observed in Caucasians. For CYP1A2*1C polymorphism, no significant association was found in overall and all subgroup analyses. In summary, this meta-analysis suggests that CYP1A2*1F polymorphism is associated with lung cancer and bladder cancer risk in Caucasians.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
As the preservation of genomic integrity is essential in the prevention of tumor initiation and progression, mutations and variations, may play a role in the genetic predisposition to cancer, especially in genes of enzymes in carcinogen metabolism. Phase I enzymes catalyze the activation and detoxification of xenobiotics, drugs, and endogenous compounds. The phase I system is mainly composed of cytochrome P450 (CYP) enzymes, which introduce a reactive group to the exogenous or endogenous compound (Guengerich 2001). The metabolism of xenobiotics and drugs is mainly a detoxification process; however, phase I metabolism has the risk of formation of highly reactive electrophiles that can bind to macromolecules, for example, proteins and DNA, potentially inducing carcinogenesis (Smith et al. 1994; Windmill et al. 1997). Therefore, genetic polymorphisms leading to alteration of activity in phase I enzymes may cause variations in the levels of DNA damage and cancer susceptibility (Brockstedt et al. 2002). CYP1A2 is located on chromosome 15q in opposite orientation and separated by 23.3 kb, a major drug-metabolizing enzyme, with a wide range of substrates (Eaton et al. 1995). CYP1A2 is a key phase I enzyme required for the activation of the major recognized carcinogens [polycyclic aromatic hydrocarbons (PAHs) and heterocyclic amines (HAs)] (Landi et al. 1999). Since both PAHs and HAs are present in food, the activity of the CYP1A2 enzyme may affect the formation of their activated forms after absorption from the large bowel, and thus influencing the risk of cancer. In humans, CYP1A2 is highly polymorphic and several single nucleotide polymorphisms (SNPs) including two common SNPs represented as CYP1A2*1C (rs2069514) and CYP1A2*1F (rs762551) alleles have been identified in different ethnic populations worldwide (Chida et al. 1999). The CYP1A2*1C allele was first reported to be associated with decreased CYP1A2 activity and inducibility in smokers of Japanese ancestry (Nakajima et al. 1999). CYP1A2*1F was reported to influence the inducibility of the enzyme, leading to a higher enzyme activity in the presence of an inducer, such as smoking (Pilgrim et al. 2012, Sachse et al. 1999), omeprazole treatment (Han et al. 2002) or heavy coffee consumption (Djordjevic et al. 2010a, b). However, there is some controversy as to the impact of CYP1A2*1C and CYP1A2*1F polymorphisms on theophylline metabolism in different ethnic populations. Obase et al. (2003) observed that CYP1A2*1C polymorphism was associated with reduced theophylline clearance in asthmatic patients of Japanese ancestry. However, the study of Wang et al. (2013) thought that CYP1A2*1F (rs762551) polymorphism can result in two- to threefold increase in activity/protein and has been associated with increased enzyme inducibility in non-smoking healthy volunteers in male Chinese population. Moreover, another study in Japanese patients reported that there was no influence of CYP1A2*1C and CYP1A2*1F on the theophylline metabolic ratio (2006). As a result, the effect of CYP1A2*1C and CYP1A2*1F polymorphisms on theophylline metabolism remains unconfirmed.
To date, a number of molecular epidemiological studies have been performed to evaluate the association between CYP1A2*1C and CYP1A2*1F polymorphisms and different types of cancer risk in diverse populations (Gervasini et al. 2013; Cui et al. 2013; Pavanello et al. 2012; Wei et al. 2011; Barbieri et al. 2012; Khvostova et al. 2012; Wang et al. 2012a; Rudolph et al. 2011; Jang et al. 2012; Sainz et al. 2011; Goodman et al. 2003; Cleary et al. 2010; Pavanello et al. 2010; Kobayashi et al. 2009a, b; Singh et al. 2010–2011; Zienolddiny et al. 2008; Imaizumi et al. 2009; MARIE-GENICA et al. 2010; Sangrajrang et al. 2009; B’chir et al. 2009; Aldrich et al. 2009; Yeh et al. 2009; Shimada et al. 2009; Kotsopoulos et al. 2009; Saebø et al. 2008; Sachse et al. 2002; Figueroa et al. 2008; Suzuki et al. 2008; Gulyaeva et al. 2008; Hirata et al. 2008; Yoshida et al. 2007; Kiss et al. 2007; Küry et al. 2007; Kotsopoulos et al. 2007; Osawa et al. 2007; Takata et al. 2007; Gemignani et al. 2007; Yeh et al. 2007; Agudo et al. 2006; De Roos et al. 2006; Bae et al. 2006; Long et al. 2006; Chen et al. 2005, 2006; Mikhailova et al. 2006; Le Marchand et al. 2005; Landi et al. 2005; Li et al. 2006; Prawan et al. 2005; Doherty et al. 2005; Tsukino et al. 2004; Goodman et al. 2001; Altayli et al. 2009; Villanueva et al. 2009; Rebbeck et al. 2006; Mochizuki et al. 2005; Chiou et al. 2005; Barbieri et al. 2013; Ghoshal et al. 2014; Lowcock et al. 2013; Lee et al. 2013; Ayari et al. 2013). However, the results were inconsistent or even contradictory. In addition, three recent meta-analyses have confirmed the association between CYP1A2*1C and CYP1A2*1F polymorphisms and risk of cancer. However, some published studies were not included in the three recent meta-analyses (Tian et al. 2013; Zhenzhen et al. 2013; Wang et al. 2012a, b). Therefore, we performed a comprehensive meta-analysis by including the most recent and relevant articles to identify statistical evidence of the association between CYP1A2*1C and CYP1A2*1F polymorphisms and risk of all cancers that have been investigated.
Materials and methods
Identification and eligibility of relevant studies
A comprehensive literature search was performed using the PubMed, ISI, CNKI, and WanFang database for relevant articles published (the last search update was April 15, 2014) with the following keywords: “CYP1A2’’, ‘‘cytochrome P-450 1A2’’, ‘‘cytochrome P450 1A2’’, ‘‘polymorphism’’ and “Cancer” or Carcinoma”. MeSH term: [“Cytochrome P-450 CYP1A2” (mesh) AND “Neoplasms” (mesh)] AND “Polymorphism, Genetic” (mesh). In addition, studies were identified by a manual search of the reference lists of reviews and retrieved studies. We included all the case–control studies and cohort studies that investigated the association between CYP1A2*1C and CYP1A2*1F polymorphisms and cancer risk with genotyping data. All eligible studies were retrieved, and their bibliographies were checked for other relevant publications.
Inclusion criteria
The included studies have to meet the following criteria: (1) only the case–control studies or cohort studies were considered, (2) evaluated the CYP1A2*1C and CYP1A2*1F polymorphisms and the risk of cancer, and (3) the genotype distribution of the CYP1A2*1C and CYP1A2*1F polymorphisms in cases and controls were described in detail and the results were expressed as odds ratio (OR) and corresponding 95 % confidence interval (95 % CI). Major reasons for exclusion of studies were as follows: (1) not for cancer research, (2) only case population, and (3) duplicate of previous publication.
Data extraction
Information was carefully extracted from all eligible studies independently by two investigators according to the inclusion criteria listed above. The following data were collected from each study: first author’s name, year of publication, country of origin, ethnicity, source of controls (population-based controls, hospital-based controls, and family-based controls), source of cases, genotype method, sample size, and numbers of cases and controls in the CYP1A2*1C and CYP1A2*1F genotypes whenever possible. Ethnicity was categorized as ‘‘Caucasian’’, ‘‘Asian’’, and “African”. When one study did not state which ethnic group was included or if it was impossible to separate participants according to phenotype, the sample was termed as ‘‘mixed population’’. Meanwhile, studies investigating more than one kind of cancer were counted as individual data set only in subgroup analyses by cancer type. We did not define any minimum number of patients to include in this meta-analysis. Articles that reported different ethnic groups and countries or locations, we considered them different study samples in this meta-analysis.
Statistical analysis
Crude odds ratios (ORs) together with their corresponding 95 % CIs were used to assess the strength of association between the CYP1A2*1C and CYP1A2*1F polymorphisms and risk of cancer. The pooled ORs were calculated for co-dominant model (AG versus AA or GG versus AA), dominant model (AG + GG versus AA), recessive model (GG versus AA + AG), and additive model (G versus A), respectively. Between-study heterogeneity was assessed by calculating Q-statistic (heterogeneity was considered statistically significant if P < 0.10.) (Davey and Egger 1997) and quantified using the I 2 value, a value that describes the percentage of variation across studies that are due to heterogeneity rather than chance, where I 2 = 0 % indicates no observed heterogeneity, with 25 % regarded as low, 50 % as moderate, and 75 % as high (Higgins et al. 2003). If results were not heterogeneous, the pooled ORs were calculated by the fixed-effect model (we used the Q-statistic, which represents the magnitude of heterogeneity between studies) (Mantel and Haenszel 1959). Otherwise, a random-effect model was used when the heterogeneity between studies was significant (DerSimonian and Laird 1986). Moreover, sensitivity analysis was performed by excluding a single study each time. We also ranked studies according to sample size, and then repeated this meta-analysis. Sample size was classified according to a minimum of 200 participants and those with fewer than 200 participants. The cite criteria were previously described (Klug et al. 2009). HWE was calculated using the goodness-of-fit test, and deviation was considered when P < 0.05. Begg’s funnel plots (Begg and Mazumdar 1994) and Egger’s linear regression test (Egger et al. 1997) were used to assess publication bias. A meta-regression analysis was carried out to identify the major sources of between-studies variation in the results, using the log of the ORs from each study as dependent variables, and cancer type, ethnicity, sample size, and source of controls as the possible sources of heterogeneity. All of the calculations were performed using STATA version 10.0 (STATA Corporation, College Station, TX).
Results
Literature search and meta-analysis databases
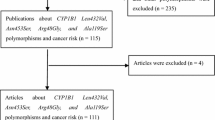
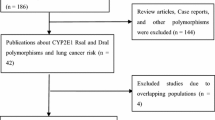
Figure 1 provides a flow chart for this meta-analysis. A total of 64 articles examined the association of CYP1A2*1C and CYP1A2*1F polymorphisms with cancer risk. Of these published articles, 2 (Yeh et al. 2007; Goodman et al. 2001) were excluded because their populations overlapped with another 2 included studies. As summarized in Table 1, 62 publications with 78 case–control studies were included in the meta-analysis, including 27,301 cases and 33,885 controls for CYP1A2*1C polymorphism (57 studies from 54 publications) and 4,722 cases and 6,555 controls for CYP1A2*1F polymorphism (21 studies from 21 publications). Among these studies, 4 were included in the dominant model only because they provided the genotypes of AG + GG ver. AA as a whole and two studies were included in the recessive model only because they provided the genotypes of GG ver. AA + AG. In addition, for CYP1A2*1C polymorphism, there were 4 bladder cancer studies, 13 breast cancer studies, 13 colorectal cancer studies, 4 endometrial cancer studies, 3 gastric cancer studies, 8 lung cancer studies, 3 pancreatic cancer studies, and 9 studies with the “other cancers”. There were 5 colorectal cancer studies, 3 liver cancer studies, 8 lung cancer studies, and 5 studies with the “other cancers” for CYP1A2*1F polymorphism. All of the cases were pathologically confirmed.
Meta-analysis results
CYP1A2*1F
Table 2 lists the main results of the meta-analysis of CYP1A2*1 F polymorphism and cancer risk. Overall, significantly increased cancer risk was observed in any genetic model (dominant model: OR 1.08, 95 % CI 1.02–1.15, P h < 0.001, I 2 = 61.4 %; heterozygous model: OR 1.06, 95 % CI 1.01–1.12, P h < 0.001, I 2 = 50.8 %; additive model: OR 1.07, 95 % CI 1.02–1.13, P h < 0.001, I 2 = 71.4 %) when all the eligible studies were pooled into meta-analysis. Then we performed subgroup analysis by cancer type. Significant association was observed between lung cancer (dominant model: OR 1.21, 95 % CI 1.00–1.46, P h = 0.083, I 2 = 44.3 %; heterozygous model: OR 1.18, 95 % CI 1.02–1.36, P h = 0.157, I 2 = 35.5 %) and bladder cancer (dominant model: OR 0.88, 95 % CI 0.78–0.99, P h = 0.563, I 2 = 0.0 %; recessive model: OR 0.79, 95 % CI 0.66–0.94, P h = 0.849, I 2 = 0.0 %; homozygous model: OR 0.76, 95 % CI 0.63–0.93, P h = 0.702, I 2 = 0.0 %; additive model: OR 0.89, 95 % CI 0.81–0.97, P h = 0.622, I 2 = 0.0 %). We further examined the association between the CYP1A2*1F polymorphism and cancer risk according to cancer type and ethnicity (Table 3). For samples of Caucasians, significant association was observed between CYP1A2*1F polymorphism and bladder cancer (dominant model: OR 0.88, 95 % CI 0.78–0.99, P h = 0.563, I 2 = 0.0 %; recessive model: OR 0.79, 95 % CI 0.66–0.94, P h = 0.849, I 2 = 0.0 %; homozygous model: OR 0.76, 95 % CI 0.63–0.93, P h = 0.702, I 2 = 0.0 %; additive model: OR 0.89, 95 % CI 0.81–0.97, P h = 0.622, I 2 = 0.0 %) and lung cancer risk (dominant model: OR 1.29, 95 % CI 1.11–1.51, P h = 0.948, I 2 = 0.0 %; recessive model: OR 1.33, 95 % CI 1.01–1.75, P h = 0.181, I 2 = 41.4 %; homozygous model: OR 1.49, 95 % CI 1.12–1.98, P h = 0.358, I 2 = 2.7 %; heterozygous model: OR 1.25, 95 % CI 1.06–1.48, P h = 0.540, I 2 = 0.0 %; additive model: OR 1.23, 95 % CI 1.09–1.39, P h = 0.828, I 2 = 0.0 %). For samples of Asians, no significant association was observed between any cancer types. We also examined the association of the CYP1A2*1F polymorphism with cancer risk according to cancer type and source of controls (Table 4). For the population-based studies, no significant association was observed between CYP1A2*1F polymorphism and cancer risk in any cancer type. For the hospital-based studies, significant association was observed only between CYP1A2*1F polymorphism and bladder cancer risk (dominant model: OR 0.88, 95 % CI 0.78–0.99, P h = 0.563, I 2 = 0.0 %; recessive model: OR 0.79, 95 % CI 0.66–0.94, P h = 0.849, I 2 = 0.0 %; homozygous model: OR 0.76, 95 % CI 0.63–0.93, P h = 0.702, I 2 = 0.0 %; additive model: OR 0.89, 95 % CI 0.81–0.97, P h = 0.622, I 2 = 0.0 %).
There was significant heterogeneity in any genetic model (P h < 0.001). We assessed the source of heterogeneity by ethnicity, cancer type, source of cases, genotype method, source of controls, and sample size. The results indicated that source of controls (dominant model: P = 0.048; heterozygous model: P = 0.013) and cancer type (dominant model: P = 0.024; additive model: P = 0.030; heterozygous model: P = 0.005) but not ethnicity (dominant model: P = 0.089; recessive model: P = 0.522; homozygous model: P = 0.336; heterozygous model: P = 0.091; additive model: P = 0.133), sample size (dominant model: P = 0.874; recessive model: P = 0.259; homozygous model: P = 0.543; heterozygous model: P = 0.643; additive model: P = 0.748), source of cases (dominant model: P = 0.367; recessive model: P = 0.396; homozygous model: P = 0.839; heterozygous model: P = 0.291; additive model: P = 0.686), and genotype method (dominant model: P = 0.777; recessive model: P = 0.155; homozygous model: P = 0.208; heterozygous model: P = 0.881; additive model: P = 0.543) contributed to substantial heterogeneity in the meta-analysis. High between-studies heterogeneity was observed in breast cancer (recessive model: I 2 = 75.1; homozygous model: I 2 = 76.6; heterozygous model: I 2 = 77.0) and other cancer (dominant model: I 2 = 80.0; recessive model: I 2 = 77.2; homozygous model: I 2 = 81.2; additive model: I 2 = 87.9). When we performed subgroup analysis by source of controls and cancer type, high between-studies heterogeneity was also observed in breast cancer (recessive model: I 2 = 54.6; homozygous model: I 2 = 45.5; heterozygous model: I 2 = 0.0) and other cancer (dominant model: I 2 = 0.0; recessive model: I 2 = 0.0; homozygous model: I 2 = 0.0; additive model: I 2 = 0.0) for the population-based studies.
When the meta-analysis was performed excluding studies with small sample sizes, there was no difference in results between overall analysis and subgroup analysis. In addition, a single study involved in the meta-analysis was deleted each time to reflect the influence of individual data set on the pooled ORs, and the corresponding pooled ORs were not essentially altered (data not shown). We performed Begg’s funnel plot and Egger’s test to assess the publication bias of literatures. Begg’s funnel plots and Egger’s test suggested that there was publication bias in any genetic model (dominant model: P = 0.001; heterozygous model: P = 0.001; recessive model: P = 0.025; additive model: P = 0.001; homozygous model: P = 0.004). This might be a limitation for the meta-analysis because studies with null findings, especially those with small sample size, are less likely to be published. Adjusting for possible publication bias using the Duval and Tweedie nonparametric ‘‘trim and fill’’ method for overall studies, the results did not change between CYP1A2*1F polymorphism with cancer risk. Figure 2 lists the Duval and Tweedie nonparametric “trim and fill’’ methods funnel plot.
CYP1A2*1C
Table 2 also lists the main results of the meta-analysis of CYP1A2*1C polymorphism and cancer risk. Overall, no significant association was observed in any genetic model (dominant model: OR 1.02, 95 % CI 0.85–1.23, P h < 0.001, I 2 = 59.9 %; recessive model: OR 1.05, 95 % CI 0.86–1.29, P h = 0.262, I 2 = 18.0 %; homozygous model: OR 1.06, 95 % CI 0.86–1.30, P h = 0.252, I 2 = 18.9 %; heterozygous model: OR 1.01, 95 % CI 0.84–1.22, P h = 0.002, I 2 = 56.1 %; additive model: OR 1.05, 95 % CI 0.88–1.25, P h < 0.001, I 2 = 67.0 %) when all the eligible studies were pooled into meta-analysis. No significant association was observed between CYP1A2*1C polymorphism and any subgroup analysis (Tables 3 and 4).
There was significant heterogeneity in dominant model (P h < 0.001), heterozygous model (P h = 0.002), and additive model (P h < 0.001). Then, we assessed the source of heterogeneity by ethnicity, cancer type, source of cases, genotype method, source of controls, and sample size. The results indicated that ethnicity (dominant model: P = 0.026; additive model: P = 0.004) contributed to substantial heterogeneity in the meta-analysis. High between-studies heterogeneity was also observed in lung cancer (dominant model: I 2 = 76.3; recessive model: I 2 = 88.6; homozygous model: I 2 = 90.6; additive model: I 2 = 85.9). However, when the study of B’chir [36] was excluded, the high between-studies heterogeneity was deleted in lung cancer (dominant model: I 2 = 0.0; heterozygous model: I 2 = 0.0; additive model: I 2 = 0.0).
When the meta-analysis was performed excluding studies with small sample sizes, there was no difference in results between overall analysis and subgroup analysis. In addition, a single study involved in the meta-analysis was deleted each time to reflect the influence of individual data set on the pooled ORs, and the corresponding pooled ORs were not essentially altered (data not shown). Both Begg’s funnel plot and Egger’s test were performed to assess the publication bias of literatures. The Egger’s test results (dominant model: P = 0.463; recessive model: P = 0.216; additive model: P = 0.406; homozygous model: P = 0.326; heterozygous model: P = 0.677) and Begg’s funnel plot (Fig. 3) suggested no evidence of publication bias in the meta-analysis.
Discussion
CYP1A2 is an important gene for catalyzing 2- and 4-hydroxylations of estrogens and metabolism of carcinogens. A major reason for the limited number of studies of heterocyclic amine (HCA) and cancer risk is the difficulty in assessing human exposure to HCAs. HCA concentrations depend on cooking methods and the “doneness” level of the meat or fish, hampering the development of a complete and standardized database of concentrations; any estimation of dietary intake from food-frequency questionnaires (FFQs) is thus likely to result in misclassification. Like other environmental chemical carcinogens, HCAs require metabolic activation by host enzymes to become genotoxic. Phase I enzymes, including cytochrome P450 1A2, can metabolically activate carcinogens to form genotoxic electrophilic intermediates (McManus et al. 1990). The relative activity of these metabolizing enzymes, which is in large part genetically determined, is thought to be an important host determinant of cancer incidence. A number of molecular epidemiological studies have been performed to evaluate the association between CYP1A2*1C and CYP1A2*1F polymorphisms and different types of cancer risk in diverse populations. However, the results were inconsistent or even contradictory. In addition, three recent meta-analyses have studied the association between CYP1A2*1C and CYP1A2*1F and risk of cancer. However, some published studies were not included in the three recent meta-analyses (Tian et al. 2013; Zhenzhen et al. 2013; Wang et al. 2012a, b). Hence, we performed a meta-analysis to explore the association between CYP1A2*1C and CYP1A2*1F polymorphisms and cancer risk.
This meta-analysis suggests that CYP1A2*1F polymorphism is associated with increased lung cancer risk and CYP1A2*1F polymorphism is associated with decreased bladder cancer risk, while results from other subgroups remain negative. A possible explanation may be that the biological effect of the genetic mutation is influenced by the variable environmental conditions at different tumor sites, leading to unpredictable physiological characteristics. It may also be attributed to the uncertainty of CYP1A2*1F polymorphism’s function at different tumor positions. Aldrich et al. (2009) reported that CYP1A2 rs762551 polymorphism was associated with an increased risk of lung cancer. Gemignani et al. (2007) reported that CYP1A2 rs762551 was associated with an increased risk of lung cancer in heterozygote carriers (P < 0.05), although not in homozygote. Singh et al. (2010–2011) found that variant genotype of CYP1A2*1F was significantly associated with increased susceptibility to squamous cell carcinoma (SCC) of lung. Pavanello et al. (2012) identified not only increased CYP1A2 metabolic activity but also increased urine mutagenicity in Italian heavy smokers having an ancestral allele of this variant. These findings are consistent with our meta-analysis results being associated with risk of lung cancer. However, at any case, the association between CYP1A2*1F polymorphism and bladder cancer risk remains an open field, as the number of studies (n = 4 for CYP1A2*1F) is considerably smaller than that needed for the achievement of robust conclusions (Higgins and Green 2008).
In the subgroup analysis by ethnicity and cancer type, significantly increased lung cancer risk was found in Caucasians and significantly decreased bladder cancer risk was also found in Caucasians, but not Africans or Asians. The results suggested a possible role of ethnic difference in genetic background and the environment they lived in. The same polymorphisms play different roles in cancer susceptibility in different ethnic populations, because cancer is a complicated multigenetic disease, and different genetic backgrounds may contribute to the discrepancy (Hirschhorn et al. 2002). In the present meta-analysis, between-studies heterogeneity was observed between CYP1A2*1C and CYP1A2*1F polymorphisms and cancer of risk. Meta-regression analysis indicated that source of control and cancer type contributed to substantial heterogeneity between the meta-analyses for CYP1A2*1F polymorphism and ethnicity contributed to substantial heterogeneity between the meta-analyses for CYP1A2*1C. The hospital-based studies may have certain biases for such controls and may only represent a sample of an ill-defined reference population, and may not be representative of the general population or it may be that numerous subjects in the population-based controls were susceptible individuals. The small number of studies hinders the ability to draw more definite conclusions. Therefore, the use of proper and representative population-based control subjects is important to reduce biases in such genetic studies. And this indicates that it may not be appropriate to use an overall estimation of the relationship between CYP1A2*1C and CYP1A2*1F polymorphism and risk of cancer.
We noticed with great interest that 3 previous meta-analyses had been reported on the overall cancer risk with CYP1A2*1C and CYP1A2*1F polymorphisms (Tian et al. 2013; Zhenzhen et al. 2013; Wang et al. 2012a, b). Tian et al. (2013) had 46 case–control studies for CYP1A2*1F polymorphism, in which a total of 22,993 cases and 28,420 controls were included. Their meta-analysis suggested that the allele of CYP1A2*1F polymorphism may be associated with breast and ovarian cancer risk, especially in Caucasian populations. However, two articles (Cui et al. 2013; Tsukino et al. 2004) should be excluded from their meta-analyses, because the two articles explored the CYP1A2*1C polymorphism and cancer risk. In addition, the study subjects should be mixed population, but not Caucasians in the study of Rebbeck et al. (2006). Zhenzhen et al. (2013) included 37 case–control studies for CYP1A2*1F (16,825 cases and 21,513 controls) and 15 studies for CYP1A2*1C (3,677 cases and 5,127 controls). Their meta-analyses suggested that the rs762551 polymorphism of the CYP1A2 gene might be a potential biomarker for the risk of cancer in Caucasians. Wang et al. (2012a, b) had 19 eligible case–control studies for CYP1A2*1F, in which a total of 8,128 cases and 11,165 controls were included. Their meta-analysis suggested that the CYP1A2 rs762551 polymorphism is likely to be associated with susceptibility to cancer in Caucasians. However, some published studies were not included in their meta-analyses (Tian et al. 2013; Zhenzhen et al. 2013; Wang et al. 2012a, b). By analyzing a larger number of studies than the previous meta-analysis (Tian et al. 2013; Zhenzhen et al. 2013; Wang et al. 2012a, b), our meta-analysis included 27,301 cases and 33,885 controls (from 57 studies) for CYP1A2 rs762551 and 4,722 cases and 6,555 controls (from 21 studies) for CYP1A2*1C to perform the two gene polymorphisms and cancer risk. Our meta-analysis suggests that CYP1A2*1F polymorphism is associated with increased lung cancer risk and CYP1A2*1F polymorphism is associated with decreased bladder cancer risk in Caucasians. Our results seem to confirm and establish the trend in the meta-analysis of the CYP1A2*1C and CYP1A2*1F polymorphisms according to the previous meta-analysis (Tian et al. 2013; Zhenzhen et al. 2013; Wang et al. 2012a, b).
There are still some limitations inherited from the published studies. First, our results were based on single-factor estimations without adjustment for other risk factors including alcohol usage, environmental factors and other lifestyles. Second, in the subgroup analysis there may exist insufficient statistical power to check an association. Third, the controls were not uniformly defined. Therefore, non-differential misclassification bias is possible because these studies may have included the control groups who have different risks for developing cancer in the various organs. However, this meta-analysis has also several strengths. First, a systematic review of the association of CYP1A2*1C and CYP1A2*1F polymorphisms with cancer risk is statistically more powerful than any single study. Second, the quality of eligible studies included in current meta-analysis was satisfactory and met our inclusion criteria. Third, we included more published studies than previously published meta-analysis.
In conclusion, this meta-analysis suggests that CYP1A2*1F polymorphism is associated with increased lung cancer risk and CYP1A2*1F polymorphism is associated with decreased bladder cancer risk in Caucasians. However, further studies are still needed to validate the associations between genetic polymorphisms in the CYP1A2 gene and cancer risks.
References
Aldrich MC, Selvin S, Hansen HM, Barcellos LF, Wrensch MR, Sison JD, Kelsey KT, Buffler PA, Quesenberry CP Jr, Seldin MF, Wiencke JK (2009) CYP1A1/2 haplotypes and lung cancer and assessment of confounding by population stratification. Cancer Res 69:2340–2348
Altayli E, Gunes S, Yilmaz AF, Goktas S, Bek Y (2009) CYP1A2, CYP2D6, GSTM1, GSTP1, and GSTT1 gene polymorphisms in patients with bladder cancer in a Turkish population. Int Urol Nephrol 41:259–266
Ayari I, Fedeli U, Saguem S, Hidar S, Khlifi S, Pavanello S (2013) Role of CYP1A2 polymorphisms in breast cancer risk in women. Mol Med Rep 7:280–286
Bae SY, Choi SK, Kim KR, Park CS, Lee SK, Roh HK, Shin DW, Pie JE, Woo ZH, Kang JH (2006) Effects of genetic polymorphisms of MDR1, FMO3 and CYP1A2 on susceptibility to colorectal cancer in Koreans. Cancer Sci 97:774–779
Barbieri RB, Bufalo NE, Secolin R, Silva AC, Assumpção LV, Maciel RM, Cerutti JM, Ward LS (2012) Evidence that polymorphisms in detoxification genes modulate the susceptibility for sporadic medullary thyroid carcinoma. Eur J Endocrinol 166:241–245
Barbieri RB, Bufalo NE, Cunha LL, Assumpção LV, Maciel RM, Cerutti JM, Ward LS (2013) Genes of detoxification are important modulators of hereditary medullary thyroid carcinoma risk. Clin Endocrinol (Oxf) 79:288–293
B’chir F, Pavanello S, Knani J, Boughattas S, Arnaud MJ, Saguem S (2009) CYP1A2 genetic polymorphisms and adenocarcinoma lung cancer risk in the Tunisian population. Life Sci 84:779–784
Begg CB, Mazumdar M (1994) Operating characteristics of a rank correlation test for publication bias. Biometrics 50:1088–1101
Brockstedt U, Krajinovic M, Richer C, Mathonnet G, Sinnett D, Pfau W et al (2002) Analyses of bulky DNA adduct levels in human breast tissue and genetic polymorphisms of cytochromes P450 (CYPs), myeloperoxidase (MPO), quinone oxidoreductase (NQO1), and glutathione S-transferases (GSTs). Mutat Res 516:41–47
Catsburg C, Joshi AD, Corral R, Lewinger JP, Koo J, John EM, Ingles SA, Stern MC (2012) Polymorphisms in carcinogen metabolism enzymes, fish intake, and risk of prostate cancer. Carcinogenesis 33:1352–1359
Chen K, Jin MJ, Fan CH, Song L, Jiang QT, Yu WP, Ma XY, Yao KY (2005) A case–control study on the association between genetic polymorphisms of metabolic enzymes and the risk of colorectal cancer. Zhonghua Liu Xing Bing Xue Za Zhi 26:659–664
Chen X, Wang H, Xie W, Liang R, Wei Z, Zhi L, Zhang X, Hao B, Zhong S, Zhou G, Zhang L, Gao X, Zhu Y, He F (2006) Association of CYP1A2 genetic polymorphisms with hepatocellular carcinoma susceptibility: a case–control study in a high-risk region of China. Pharmacogenet Genomics 16:219–227
Chida M, Yokoi T, Fukui T, Kinoshita M, Yokota J, Kamataki T (1999) Detection of three genetic polymorphisms in the 5’-flanking region and intron 1 of human CYP1A2 in the Japanese population. Jpn J Cancer Res 90:899–902
Chiou HL, Wu MF, Chien WP, Cheng YW, Wong RH, Chen CY, Lin TS, Lee H (2005) NAT2 fast acetylator genotype is associated with an increased risk of lung cancer among never-smoking women in Taiwan. Cancer Lett 223:93–101
Cleary SP, Cotterchio M, Shi E, Gallinger S, Harper P (2010) Cigarette smoking, genetic variants in carcinogen-metabolizing enzymes, and colorectal cancer risk. Am J Epidemiol 172:1000–1014
Pavanello S, Mastrangelo G, Placidi D, Campagna M, Pulliero A, Carta A, Arici C, Porru S (2010) CYP1A2 polymorphisms, occupational and environmental exposures and risk of bladder cancer. Eur J Epidemiol 25:491–500
Cui X, Lu X, Hiura M, Omori H, Miyazaki W, Katoh T (2013) Association of genotypes of carcinogen-metabolizing enzymes and smoking status with bladder cancer in a Japanese population. Environ Health Prev Med 18:136–142
Davey SG, Egger M (1997) Meta-analyses of randomized controlled trials. Lancet 350:1182
De Roos AJ, Gold LS, Wang S, Hartge P, Cerhan JR, Cozen W, Yeager M, Chanock S, Rothman N, Severson RK (2006) Metabolic gene variants and risk of non-Hodgkin’s lymphoma. Cancer Epidemiol Biomarkers Prev 15:1647–1653
DerSimonian R, Laird N (1986) Meta-analysis in clinical trials. Control Clin Trials 7:177–188
Djordjevic N, Ghotbi R, Jankovic S, Aklillu E (2010a) Induction of CYP1A2 by heavy coffee consumption is associated with the CYP1A2 -163C>A polymorphism. Eur J Clin Pharmacol 66:697–703
Djordjevic N, Carrillo JA, Gervasini G, Jankovic S, Aklillu E (2010b) In vivo evaluation of CYP2A6 and xanthine oxidase enzyme activities in the Serbian population. Eur J Clin Pharmacol 66:571–578
Doherty JA, Weiss NS, Freeman RJ, Dightman DA, Thornton PJ, Houck JR, Voigt LF, Rossing MA, Schwartz SM, Chen C (2005) Genetic factors in catechol estrogen metabolism in relation to the risk of endometrial cancer. Cancer Epidemiol Biomarkers Prev 14:357–366
Eaton DL, Gallagher EP, Bammler TK, Kunze KL (1995) Role of cytochrome P450 1A2 in chemical carcinogenesis: implications for human variability in expression and enzyme activity. Pharmacogenetics 5:259–274
Egger M, Smith DG, Schneider M, Minder C (1997) Bias in meta-analysis detected by a simple, graphical test. Br Med J 315:629–634
Figueroa JD, Malats N, García-Closas M, Real FX, Silverman D, Kogevinas M, Chanock S, Welch R, Dosemeci M, Lan Q, Tardón A, Serra C, Carrato A, García-Closas R, Castaño-Vinyals G, Rothman N (2008) Bladder cancer risk and genetic variation in AKR1C3 and other metabolizing genes. Carcinogenesis 29:1955–1962
Gemignani F, Landi S, Szeszenia-Dabrowska N, Zaridze D, Lissowska J, Rudnai P, Fabianova E, Mates D, Foretova L, Janout V, Bencko V, Gaborieau V, Gioia-Patricola L, Bellini I, Barale R, Canzian F, Hall J, Boffetta P, Hung RJ, Brennan P (2007) Development of lung cancer before the age of 50: the role of xenobiotic metabolizing genes. Carcinogenesis 28:1287–1293
Gervasini G, Ghotbi R, Aklillu E, San Jose C, Cabanillas A, Kishikawa J, Benitez J, Carrillo JA (2013) Haplotypes in the 5’-untranslated region of the CYP1A2gene are inversely associated with lung cancer risk but do not correlate with caffeine metabolism. Environ Mol Mutagen 54:124–132
Ghoshal U, Tripathi S, Kumar S, Mittal B, Chourasia D, Kumari N, Krishnani N, Ghoshal UC (2014) Genetic polymorphism of cytochrome P450 (CYP) 1A1, CYP1A2, and CYP2E1 genes modulate susceptibility to gastric cancer in patients with Helicobacter pylori infection. Gastric Cancer 17:226–234
Goodman MT, McDuffie K, Kolonel LN, Terada K, Donlon TA, Wilkens LR, Guo C, Le Marchand L (2001) Case–control study of ovarian cancer and polymorphisms in genes involved in catecholestrogen formation and metabolism. Cancer Epidemiol Biomarkers Prev 10:209–216
Guengerich FP (2001) Forging the links between metabolism and carcinogenesis. Mutat Res 488:195–209
Landi MT, Sinha R, Lang NP, Landi MT, Sinha R, Lang NP, Kadlubar FF (1999) Human cytochrome P4501A2. In: Vineis P, Malats N, Lang M (eds) Metabolic polymorphisms and susceptibility to cancer, vol Chapter 16, 148th edn. IARC Scientific Publications, IARC, Lyon, pp 173–195
Gulyaeva LF, Mikhailova ON, PustyInyak VO, Kim IV 4th, Gerasimov AV, Krasilnikov SE, Filipenko ML, Pechkovsky EV (2008) Comparative analysis of SNP in estrogen-metabolizing enzymes for ovarian, endometrial, and breast cancers in Novosibirsk, Russia. Adv Exp Med Biol 617:359–366
Han XM, Ouyang DS, Chen XP, Shu Y, Jiang CH, Tan ZR et al (2002) Inducibility of CYP1A2 by omeprazole in vivo related to the genetic polymorphism of CYP1A2. Br J Clin Pharmacol 54:540–543
Higgins JPT, Green S (2008) Cochrane handbook for systematic reviews of interventions version 5.0.1. The Cochrane Collaboration, Oxford
Hirschhorn JN, Lohmueller K, Byrne E (2002) A comprehensive review of genetic association studies. Genet Med 4:45–61
Higgins JP, Thompson SG, Deeks JJ, Altman DG (2003) Measuring inconsistency in meta-analysis. Br Med J 327:557–560
Hirata H, Hinoda Y, Okayama N, Suehiro Y, Kawamoto K, Kikuno N, Rabban JT, Chen LM, Dahiya R (2008) CYP1A1, SULT1A1, and SULT1E1 polymorphisms are risk factors for endometrial cancer susceptibility. Cancer 112:1964–1973
Imaizumi T, Higaki Y, Hara M, Sakamoto T, Horita M, Mizuta T, Eguchi Y, Yasutake T, Ozaki I, Yamamoto K, Onohara S, Kawazoe S, Shigematsu H, Koizumi S, Kudo S, Tanaka K (2009) Interaction between cytochrome P450 1A2 genetic polymorphism and cigarette smoking on the risk of hepatocellular carcinoma in a Japanese population. Carcinogenesis 30:1729–1734
Jang JH, Cotterchio M, Borgida A, Gallinger S, Cleary SP (2012) Genetic variants in carcinogen-metabolizing enzymes, cigarette smoking and pancreatic cancer risk. Carcinogenesis 33:818–827
Khvostova EP, Pustylnyak VO, Gulyaeva LF (2012) Genetic polymorphism of estrogen metabolizing enzymes in Siberian women with breast cancer. Genet Test Mol Biomarkers 16:167–173
Kiss I, Orsós Z, Gombos K, Bogner B, Csejtei A, Tibold A, Varga Z, Pázsit E, Magda I, Zölyomi A, Ember I (2007) Association between allelic polymorphisms of metabolizing enzymes (CYP 1A1, CYP 1A2, CYP 2E1, mEH) and occurrence of colorectal cancer in Hungary. Anticancer Res 27:2931–2937
Kobayashi M, Otani T, Iwasaki M, Natsukawa S, Shaura K, Koizumi Y, Kasuga Y, Sakamoto H, Yoshida T, Tsugane S (2009a) Association between dietary heterocyclic amine levels, genetic polymorphisms of NAT2, CYP1A1, and CYP1A2 and risk of stomach cancer: a hospital-based case–control study in Japan. Gastric Cancer 12:198–205
Kobayashi M, Otani T, Iwasaki M, Natsukawa S, Shaura K, Koizumi Y, Kasuga Y, Sakamoto H, Yoshida T, Tsugane S (2009b) Association between dietary heterocyclic amine levels, genetic polymorphisms of NAT2, CYP1A1, and CYP1A2 and risk of colorectal cancer: a hospital-based case–control study in Japan. Scand J Gastroenterol 44:952–959
Kotsopoulos J, Ghadirian P, El-Sohemy A, Lynch HT, Snyder C, Daly M, Domchek S, Randall S, Karlan B, Zhang P, Zhang S, Sun P, Narod SA (2007) The CYP1A2 genotype modifies the association between coffee consumption and breast cancer risk among BRCA1 mutation carriers. Cancer Epidemiol Biomarkers Prev 16:912–916
Kotsopoulos J, Vitonis AF, Terry KL, De Vivo I, Cramer DW, Hankinson SE, Tworoger SS (2009) Coffee intake, variants in genes involved in caffeine metabolism, and the risk of epithelial ovarian cancer. Cancer Causes Control 20:335–344
Küry S, Buecher B, Robiou-du-Pont S, Scoul C, Sébille V, Colman H, Le Houérou C, Le Neel T, Bourdon J, Faroux R, Ollivry J, Lafraise B, Chupin LD, Bézieau S (2007) Combinations of cytochrome P450 gene polymorphisms enhancing the risk for sporadic colorectal cancer related to red meat consumption. Cancer Epidemiol Biomarkers Prev 16:1460–1467
Landi S, Gemignani F, Moreno V, Gioia-Patricola L, Chabrier A, Guino E, Navarro M, de Oca J, Capellà G, Canzian F, Bellvitge Colorectal Cancer Study Group (2005) A comprehensive analysis of phase I and phase II metabolism gene polymorphisms and risk of colorectal cancer. Pharmacogenet Genomics 15:535–546
Le Marchand L, Donlon T, Kolonel LN, Henderson BE, Wilkens LR (2005) Estrogen metabolism-related genes and breast cancer risk: the multiethnic cohort study. Cancer Epidemiol Biomarkers Prev 14:1998–2003
Lee HJ, Wu K, Cox DG, Hunter D, Hankinson SE, Willett WC, Sinha R, Cho E (2013) Polymorphisms in xenobiotic metabolizing genes, intakes of heterocyclic amines and red meat, and postmenopausal breast cancer. Nutr Cancer 65:1122–1131
Li D, Jiao L, Li Y, Doll MA, Hein DW, Bondy ML, Evans DB, Wolff RA, Lenzi R, Pisters PW, Abbruzzese JL, Hassan MM (2006) Polymorphisms of cytochrome P4501A2 and N-acetyltransferase genes, smoking, and risk of pancreatic cancer. Carcinogenesis 27:103–111
Long JR, Egan KM, Dunning L, Shu XO, Cai Q, Cai H, Dai Q, Holtzman J, Gao YT, Zheng W (2006) Population-based case–control study of AhR (aryl hydrocarbon receptor) and CYP1A2 polymorphisms and breast cancer risk. Pharmacogenet Genomics 16:237–243
Lowcock EC, Cotterchio M, Anderson LN, Boucher BA, El-Sohemy A (2013) High coffee intake, but not caffeine, is associated with reduced estrogen receptor negative and postmenopausal breast cancer risk with no effect modification by CYP1A2 genotype. Nutr Cancer 65:398–409
Mantel N, Haenszel W (1959) Statistical aspects of the analysis of data from retrospective studies of disease. Natl Cancer Inst 22:719–748
MARIE-GENICA Consortium on Genetic Susceptibility for Menopausal Hormone Therapy Related Breast Cancer Risk (2010) Genetic polymorphisms in phase I and phase II enzymes and breast cancer risk associated with menopausal hormone therapy in postmenopausal women. Breast Cancer Res Treat 119:463–474
McManus ME, Burgess WM, Veronese ME, Huggett A, Quattrochi LC, Tukey RH (1990) Metabolism of 2-acetylaminofluorene and benzo(a)pyrene and activation of food-derived heterocyclic amine mutagens by human cytochromes P-450. Cancer Res 50:3367–3376
Mikhailova ON, Gulyaeva LF, Prudnikov AV, Gerasimov AV, Krasilnikov SE (2006) Estrogen-metabolizing gene polymorphisms in the assessment of female hormone-dependent cancer risk. Pharmacogenomics J 6:189–193
Mochizuki J, Murakami S, Sanjo A, Takagi I, Akizuki S, Ohnishi A (2005) Genetic polymorphisms of cytochrome P450 in patients with hepatitis C virus-associated hepatocellular carcinoma. J Gastroenterol Hepatol 20:1191–1197
Nakajima M, Yokoi T, Mizutani M, Kinoshita M, Funayama M, Kamataki T (1999) Genetic polymorphism in the 5’-flanking region of humanCYP1A2 gene: effect on the CYP1A2 inducibility in humans. J Biochem 125:803–808
Obase Y, Shimoda T, Kawano T, Saeki S, Tomari SY, MitsutaIzaki K et al (2003) Polymorphisms in the CYP1A2 gene and theophylline metabolism in patients with asthma. Clin Pharmacol Ther 73:468–474
Osawa Y, Osawa KK, Miyaishi A, Higuchi M, Tsutou A, Matsumura S, Tabuchi Y, Tsubota N, Takahashi J (2007) NAT2 and CYP1A2 polymorphisms and lung cancer risk in relation to smoking status. Asian Pac J Cancer Prev 8:103–108
Pavanello S, Fedeli U, Mastrangelo G, Rota F, Overvad K, Raaschou-Nielsen O, Tjønneland A, Vogel U (2012) Role of CYP1A2 polymorphisms on lung cancer risk in a prospective study. Cancer Genet 205:278–284
Pilgrim JL, Ruiz Y, Gesteira A, Cruz R, Gerostamoulos D, Carracedo A et al (2012) Characterization of single nucleotide polymorphisms of cytochrome p450 in an Australian deceased sample. Curr Drug Metab 13:679–692
Prawan A, Kukongviriyapan V, Tassaneeyakul W, Pairojkul C, Bhudhisawasdi V (2005) Association between genetic polymorphisms of CYP1A2, arylamine N-acetyltransferase 1 and 2 and susceptibility to cholangiocarcinoma. Eur J Cancer Prev 14:245–250
Rebbeck TR, Troxel AB, Wang Y, Walker AH, Panossian S, Gallagher S, Shatalova EG, Blanchard R, Bunin G, DeMichele A, Rubin SC, Baumgarten M, Berlin M, Schinnar R, Berlin JA, Strom BL (2006) Estrogen sulfation genes, hormone replacement therapy, and endometrial cancer risk. J Natl Cancer Inst 98:1311–1320
Rudolph A, Sainz J, Hein R, Hoffmeister M, Frank B, Försti A, Brenner H, Hemminki K, Chang-Claude J (2011) Modification of menopausal hormone therapy-associated colorectal cancer risk by polymorphisms in sex steroid signaling, metabolism and transport related genes. Endocr Relat Cancer 18:371–384
Sachse C, Smith G, Wilkie MJ, Barrett JH, Waxman R, Sullivan F, Forman D, Bishop DT, Wolf CR, Colorectal Cancer Study Group (2002) A pharmacogenetic study to investigate the role of dietary carcinogens in the etiology of colorectal cancer. Carcinogenesis 23:1839–1849
Sachse C, Brockmoller J, Bauer S, Roots I (1999) Functional significance of a C– > A polymorphism in intron 1 of the cytochrome P450 CYP1A2 gene tested with caffeine. Br J Clin Pharmacol 47:445–449
Saebø M, Skjelbred CF, Brekke Li K, Bowitz Lothe IM, Hagen PC, Johnsen E, Tveit KM, Kure EH (2008) CYP1A2 164 A- > C polymorphism, cigarette smoking, consumption of well-done red meat and risk of developing colorectal adenomas and carcinomas. Anticancer Res 28:2289–2295
Sainz J, Rudolph A, Hein R, Hoffmeister M, Buch S, von Schönfels W, Hampe J, Schafmayer C, Völzke H, Frank B, Brenner H, Försti A, Hemminki K, Chang-Claude J (2011) Association of genetic polymorphisms in ESR2, HSD17B1, ABCB1, and SHBG genes with colorectal cancer risk. Endocr Relat Cancer 18:265–276
Goodman MT, Tung KH, McDuffie K, Wilkens LR, Donlon TA (2003) Association of caffeine intake and CYP1A2 genotype with ovarian cancer. Nutr Cancer 46:23–29
Sangrajrang S, Sato Y, Sakamoto H, Ohnami S, Laird NM, Khuhaprema T, Brennan P, Boffetta P, Yoshida T (2009) Genetic polymorphisms of estrogen metabolizing enzyme and breast cancer risk in Thai women. Int J Cancer 125:837–843
Singh AP, Pant MC, Ruwali M, Shah PP, Prasad R, Mathur N, Parmar D (2010) –2011) Polymorphism in cytochrome P450 1A2 and their interaction with risk factors in determining risk of squamous cell lung carcinoma in men. Cancer 8:351–359
Smith CA, Smith G, Wolf CR (1994) Genetic polymorphisms in xenobiotic metabolism. Eur J Cancer 30A:1921–1935
Suzuki H, Morris JS, Li Y, Doll MA, Hein DW, Liu J, Jiao L, Hassan MM, Day RS, Bondy ML, Abbruzzese JL, Li D (2008) Interaction of the cytochrome P4501A2, SULT1A1 and NAT gene polymorphisms with smoking and dietary mutagen intake in modification of the risk of pancreatic cancer. Carcinogenesis 29:1184–1191
Takata K, Saruwatari J, Nakada N, Nakagawa M, Fukuda K, Tanaka F et al (2006) Phenotype-genotype analysis of CYP1A2 in Japanese patients receiving oral theophylline therapy. Eur J Clin Pharmacol 62:23–28
Takata Y, Maskarinec G, Le Marchand L (2007) Breast density and polymorphisms in genes coding for CYP1A2 and COMT: the multiethnic cohort. BMC Cancer 7:30
Tian Z, Li YL, Zhao L, Zhang CL (2013) Role of CYP1A2 1F polymorphism in cancer risk: evidence from a meta-analysis of 46 case–control studies. Gene 524:168–174
Tsukino H, Kuroda Y, Nakao H, Imai H, Inatomi H, Osada Y, Katoh T (2004) Cytochrome P450 (CYP) 1A2, sulfotransferase (SULT) 1A1, and N-acetyltransferase (NAT) 2 polymorphisms and susceptibility to urothelial cancer. J Cancer Res Clin Oncol 130:99–106
Villanueva CM, Silverman DT, Murta-Nascimento C, Malats N, Garcia-Closas M, Castro F, Tardon A, Garcia-Closas R, Serra C, Carrato A, Rothman N, Real FX, Dosemeci M, Kogevinas M (2009) Coffee consumption, genetic susceptibility and bladder cancer risk. Cancer Causes Control 20:121–127
Wang J, Joshi AD, Corral R, Siegmund KD, Marchand LL, Martinez ME, Haile RW, Ahnen DJ, Sandler RS, Lance P, Stern MC (2012a) Carcinogen metabolism genes, red meat and poultry intake, and colorectal cancer risk. Int J Cancer 130:1898–1907
Wang HG, Zhang Z, Han SG, Lu YJ, Feng FM, Yuan JX (2012b) CYP1A2 rs762551 polymorphism contributes to cancer susceptibility: a meta-analysis from 19 case–control studies. BMC Cancer 12:528
Wang L, Hu Z, Deng X, Wang Y, Zhang Z, Cheng ZN (2013) Association between common CYP1A2 polymorphisms and theophylline metabolism in non-smoking healthy volunteers. Basic Clin Pharmacol Toxicol 112:257–263
Wei W, Ge JP, Dong J, Gao JP, Zhang ZY, Gong J (2011) Single nucleotide polymorphisms of CYP1A2 and their correlation with prostate cancer. Zhonghua Nan Ke Xue 17:998–1001
Windmill KF, McKinnon RA, Zhu X, Gaedigk A, Grant DM, McManus ME (1997) The role of xenobiotic metabolizing enzymes in arylamine toxicity and carcinogenesis: functional and localization studies. Mutat Res 376:153–160
Klug SJ, Ressing M, Koenig J, Abba MC, Agorastos T, Brenna SM, Ciotti M, Das BR, Del Mistro A, Dybikowska A, Giuliano AR, Gudleviciene Z, Gyllensten U, Haws AL, Helland A, Herrington CS, Hildesheim A, Humbey O, Jee SH, Kim JW, Madeleine MM, Menczer J, Ngan HY, Nishikawa A, Niwa Y, Pegoraro R, Pillai MR, Ranzani G, Rezza G, Rosenthal AN, Roychoudhury S, Saranath D, Schmitt VM, Sengupta S, Settheetham-Ishida W, Shirasawa H, Snijders PJ, Stoler MH, Suárez-Rincón AE, Szarka K, Tachezy R, Ueda M, van der Zee AG, von Knebel Doeberitz M, Wu MT, Yamashita T, Zehbe I, Blettner M (2009) TP53 codon 72 polymorphism and cervical cancer: a pooled analysis of individual data from 49 studies. Lancet Oncol 10:772–784
Yeh CC, Sung FC, Tang R, Chang-Chieh CR, Hsieh LL (2007) Association between polymorphisms of biotransformation and DNA-repair genes and risk of colorectal cancer in Taiwan. J Biomed Sci 14:183–193
Agudo A, Sala N, Pera G, Capellá G, Berenguer A, García N, Palli D, Boeing H, Del Giudice G, Saieva C, Carneiro F, Berrino F, Sacerdote C, Tumino R, Panico S, Berglund G, Simán H, Stenling R, Hallmans G, Martínez C, Bilbao R, Barricarte A, Navarro C, Quirós JR, Allen N, Key T, Bingham S, Khaw KT, Linseisen J, Nagel G, Overvad K, Tjonneland A, Olsen A, Bueno-de-Mesquita HB, Boshuizen HC, Peeters PH, Numans ME, Clavel-Chapelon F, Boutron-Ruault MC, Trichopoulou A, Lund E, Offerhaus J, Jenab M, Ferrari P, Norat T, Riboli E, González CA (2006) Polymorphisms in metabolic genes related to tobacco smoke and the risk of gastric cancer in the European prospective investigation into cancer and nutrition. Cancer Epidemiol Biomarkers Prev 15:2427–2434
Yeh CC, Sung FC, Tang R, Chang-Chieh CR, Hsieh LL (2009) Polymorphisms of cytochrome P450 1A2 and N-acetyltransferase genes, meat consumption, and risk of colorectal cancer. Dis Colon Rectum 52:104–111
Shimada N, Iwasaki M, Kasuga Y, Yokoyama S, Onuma H, Nishimura H, Kusama R, Hamada GS, Nishimoto IN, Iyeyasu H, Motola J Jr, Laginha FM, Kurahashi N, Tsugane S (2009) Genetic polymorphisms in estrogen metabolism and breast cancer risk in case–control studies in Japanese, Japanese Brazilians and non-Japanese Brazilians. J Hum Genet 54:209–215
Yoshida K, Osawa K, Kasahara M, Miyaishi A, Nakanishi K, Hayamizu S, Osawa Y, Tsutou A, Tabuchi Y, Shimada E, Tanaka K, Yamamoto M, Takahashi J (2007) Association of CYP1A1, CYP1A2, GSTM1 and NAT2 gene polymorphisms with colorectal cancer and smoking. Asian Pac J Cancer Prev 8:438–444
Zhenzhen L, Xianghua L, Ning S, Zhan G, Chuanchuan R, Jie L (2013) Current evidence on the relationship between three polymorphisms in the CYP1A2 gene and the risk of cancer. Eur J Cancer Prev 22:607–619
Zienolddiny S, Campa D, Lind H, Ryberg D, Skaug V, Stangeland LB, Canzian F, Haugen A (2008) Acomprehensive analysis of phase I and phase II metabolism gene polymorphisms and risk of non-small cell lung cancer in smokers. Carcinogenesis 29:1164–1169
Acknowledgments
Wen-Xia Zhang and Ying-Hua Chen contributed equally to this study and should be considered as co-first authors. This study was supported by the Natural Science Foundation of Shanxi Province (2013011056-3), The Science and Technology Development Projects of Colleges and Universities of Shanxi Province (20121013), and The Scientific Research Subject of the Health Department of Shanxi Province (201202008).
Conflict of interest
None.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Communicated by S. Hohmann.
Rights and permissions
About this article
Cite this article
Sun, WX., Chen, YH., Liu, ZZ. et al. Association between the CYP1A2 polymorphisms and risk of cancer: a meta-analysis. Mol Genet Genomics 290, 709–725 (2015). https://doi.org/10.1007/s00438-014-0956-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00438-014-0956-8