Abstract
The previous, published data on the association between CYP1B1 polymorphisms and cancer risk remained controversial. To derive a more precise estimation of the association between the CYP1B1 polymorphisms and cancer risk, we performed a meta-analysis to investigate the association between cancer susceptibility and CYP1B1 Leu432Val, Asn453Ser, Arg48Gly, and Ala119Ser polymorphisms. For Asn453Ser and Arg48Gly polymorphisms, significantly decreased endometrial cancer was observed among Caucasians. For Ala119Ser polymorphism, we found that individuals with the minor variant genotypes had a high risk of prostate cancer. For Leu432Val polymorphism, we found that individuals with the minor variant genotypes had a higher risk of endometrial cancer and lung cancer and had a lower risk of ovarian cancer. In summary, this meta-analysis suggests that Leu432Val polymorphism is associated with ovarian cancer, lung cancer, and endometrial cancer risk; Asn453Ser and Arg48Gly polymorphisms are associated with endometrial cancer risk among Caucasians, Ala119Ser polymorphism is associated with prostate cancer risk, and Ala119Ser polymorphism is associated with breast cancer risk in Caucasians. In addition, our work also points out the importance of new studies for Ala119Ser polymorphism in endometrial cancer, because high heterogeneity was observed (I 2 > 75 %).
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Cancer is projected to become the leading cause of death worldwide according to a new edition of the World Cancer Report from the International Agency for Research on Cancer, which has become a major public health challenge (Siegel et al. 2013). Although a recently published paper indicated that cancer prevention and management is moving in the right direction (Karim-Kos et al. 2008). Due to the combination of earlier detection, better access to care and improved treatment, survival is increased and mortality is decreased. Still, cancer prevention efforts have much to attain (Lutz et al. 2003; Soerjomataram et al. 2007). New markers for identifying high-risk populations as well as novel strategies for early detection and preventive care are urgently needed. The mechanism of carcinogenesis is still not fully understood.
CYP1B1 gene is located on chr2p22-p21, which is involved in the metabolic activation of polycyclic aromatic hydrocarbons (PAHs) including benzo(a)pyrene and dimethylbenz(a)anthracene (DMBA), but with a product distribution that is distinct from CYP1A1 (Buters et al. 1999; Shimada et al. 1996). CYP1B1 is commonly overexpressed in human malignancies (Murray et al. 1997) and activates a variety of carcinogens. For example, CYP1B1 catalyzes both the formation of dihydrodiols of specific PAHs and their subsequent oxidation to carcinogenic dihydrodiol epoxides (Kim et al. 1998). CYP1B1 is transcriptionally induced by compounds such as 2,3,7,8-tetrachlorodibenzo-p-dioxin or dioxin, and regulated by several key transcriptional factors including oestrogen receptor and aryl hydrocarbon receptor (Sutter et al. 1994). Apart from its role in xenobiotic metabolism, CYP1B1 is implicated in the bioactivation of pro-carcinogens (Hayes et al. 1996; Martin et al. 2010; Spink et al. 1997). The enzyme also appears to play a role in the metabolism of certain anticancer agents used in the treatment of hormone-induced cancers (Sissung et al. 2008). In humans, CYP1B1 is genetically polymorphic and more than 50 single nucleotide polymorphisms (SNPs) have been reported so far, of which certain deleterious mutations are associated with primary congenital glaucoma (Stoilov et al. 1998). Of the most common SNPs of CYP1B1 gene, four have been reported to result in amino acid substitutions including Arg by Gly at codon 48(rs10012), Ala by Ser at codon 119 (rs1056827), Leu by Val at codon 432 (rs1056836) and Asn by Ser at codon 453 (rs1800440).
In the past decade, a number of molecular epidemiological studies have been done to evaluate the association between CYP1B1 Leu432Val, Asn453Ser, Arg48Gly, and Ala119Ser polymorphisms and different types of cancer risk in diverse populations (Catsburg et al. 2012; Reding et al. 2012; Jang et al. 2012; Salinas-Sánchez et al. 2012a, b; Cerne et al. 2011a,b ; dos Santos et al. 2011; Wang et al. 2011a, 2012; Zhu et al. 2011; Rudolph et al. 2011; Timofeeva et al. 2009; Sainz et al. 2011; Cleary et al. 2010; Hlavata et al. 2010; Trubicka et al. 2010; Ozbek et al. 2010; Soucek et al. 2010; Sachse et al. 2002; Sliwinski et al. 2010; Ashton et al. 2010; Delort et al. 2010; Church et al. 2010; Yadav et al. 2009; Tai et al. 2010; Vrana et al. 2010; Li et al. 2002, 2005, 2013; Kato et al. 2009; Beuten et al. 2009; Rotunno et al. 2009; Fontana et al. 2009; MARIE-GENICA et al. 2010; Sangrajrang et al. 2009; Reding et al. 2009; Kilfoy et al. 2009; Huang et al. 2009; Shimada et al. 2009; Listgarten et al. 2004; Sigurdson et al. 2009; Okobia et al. 2009; Cote et al. 2009; Yuan et al. 2008; Delort et al. 2008; Freedman et al. 2009; Cotterchio et al. 2008; Figueroa et al. 2008; Diergaarde et al. 2008; Shah et al. 2008; Harth et al. 2008; Beuten et al. 2008; Gulyaeva et al. 2008; Van Emburgh et al. 2008; Hirata et al. 2008; Zienolddiny et al. 2008; Singh et al. 2008; Yoon et al. 2008; Justenhoven et al. 2008; Cussenot et al. 2007; Küry et al. 2007; Bethke et al. 2007; Matyjasik et al. 2007; Holt et al. 2007; Berndt et al. 2007; Cote et al. 2007; Tao et al. 2006; Sillanpää et al. 2007; De Roos et al. 2006a, b; Gaudet et al. 2006; Gallicchio et al. 2006; Sobti et al. 2006; Sellers et al. 2005; Huber et al. 2005; Cicek et al. 2005; Le Marchand et al. 2005; Wenzlaff et al. 2005; Landi et al. 2005; Liang et al. 2005; Sørensen et al. 2005; Doherty et al. 2005; Wen et al. 2005; Rylander-Rudqvist et al. 2004; Fukatsu et al. 2004; Zimarina et al. 2004; Cecchin et al. 2004; Dunning et al. 2004; Hung et al. 2004; Ahsan et al. 2004; Wu et al. 2004; Thyagarajan et al. 2004; Sasaki et al. 2003, 2004; McGrath et al. 2004; Chang et al. 2003; Rylander-Rudqvist et al. 2003; Lee et al. 2003; Kocabaş et al. 2002; Tanaka et al. 2002; De Vivo et al. 2002; Ko et al. 2001; Goodman et al. 2001; Watanabe et al. 2000; Zheng et al. 2000; Bailey et al. 1998; Berber et al. 2013; Maurya et al. 2014; Martínez-Ramírez et al. 2013; Rebbeck et al. 2006; Lundin et al. 2012; Tang et al. 2000; Rodrigues et al. 2011; Holt et al. 2013). However, the results were inconsistent or even contradictory. Partially because of the possible small effect of the polymorphism on cancer risk and the relatively small sample size in each of published studies. In addition, some recent meta-analyses analyzed such an association only for single cancer such as breast cancer, lung cancer, prostate cancer, endometrial cancer, and so on (Economopoulos and Sergentanis 2010; Chen et al. 2010; Wang et al. 2011b; Xu et al. 2012; Cui et al. 2012; Yang et al. 2012). Hence, the correlation of these polymorphic genes remains unknown. Every single study may be underpowered to achieve a comprehensive and reliable conclusion. Hence, in order to explore the association we performed a comprehensive meta-analysis by including the most recent and relevant articles to identify statistical evidence of the association between CYP1B1 Leu432Val, Asn453Ser, Arg48Gly, and Ala119Ser polymorphisms and risk of all cancers that have been investigated.
Materials and methods
Identification and eligibility of relevant studies
A comprehensive literature search was performed using the PubMed, Cochrane Library, and EMBASE database for relevant articles published (the last search update was Feb 18, 2014) with the following key words “CYP1B1’’, ‘‘cytochrome P-450 1B1’’, ‘‘cytochrome P450 1B1’’, ‘‘polymorphism’’, “Variant”, or “Mutation”, and “Cancer” or Carcinoma”. In addition, studies were identified by a manual search of the reference lists of reviews and retrieved studies. We included all the case–control studies and cohort studies that investigated the association between CYP1B1 Leu432Val, Asn453Ser, Arg48Gly, and Ala119Ser polymorphisms and cancer risk with genotyping data. All eligible studies were retrieved, and their bibliographies were checked for other relevant publications.
Inclusion and exclusion criteria
The included studies have to meet the following criteria: For inclusion, the study outcome had to be cancer, there had to be at least two comparison groups (case group vs. control group), and the investigation had to provide data on the CYP1B1 genotype distribution. We excluded review articles, editorials, case reports, studies with preliminary results not on CYP1B1 polymorphism or outcome, and investigations of the role of CYP1B1 expression related to disease. When the same sample was used in several publications, only the most complete study was included following careful examination.
Data extraction
Information was carefully extracted from all eligible studies independently by two investigators according to the inclusion criteria listed above. The following data were collected from each study: first author’s name, year of publication, country of origin, ethnicity, source of controls (population-based controls, hospital-based controls, and family-based controls), sample size, and numbers of cases and controls in the CYP1B1 Leu432Val, Asn453Ser, Arg48Gly, and Ala119Ser genotypes whenever possible. Ethnicity was categorized as ‘‘Caucasian’’, ‘‘Asian’’, and “African”. When one study did not state which ethnic group was included or if it was impossible to separate participants according to phenotype, the sample was termed as ‘‘mixed population’’. Meanwhile, studies investigating more than one kind of cancer were counted as individual data set only in subgroup analyses by cancer type. We did not define any minimum number of patients to include in this meta-analysis. Articles that reported different ethnic groups and different countries or locations, we considered them different study samples for each category cited above.
Statistical analysis
Crude odds ratios (ORs) together with their corresponding 95 % CIs were used to assess the strength of association between the CYP1B1 Leu432Val, Asn453Ser, Arg48Gly, and Ala119Ser polymorphisms and the risk of cancer. The pooled ORs were performed for co-dominant model (Leu432Val: Val/Val versus Leu/Leu and Leu/Val versus Leu/Leu, Asn453Ser: Ser/Ser versus Asn/Asn and Asn/Ser versus Asn/Asn, Arg48Gly: Gly/Gly versus Arg/Arg and Arg/Gly versus Arg/Arg, and Ala119Ser: Ser/Ser versus Ala/Ala and Ala/Ser versus Ala/Ala); dominant model (Leu432Val: Leu/Val + Val/Val versus Leu/Leu, Asn453Ser: Asn/Ser + Ser/Ser versus Asn/Asn, Arg48Gly: Arg/Gly + Gly/Gly versus Arg/Arg, and Ala119Ser: Ala/Ser + Ser/Ser versus Ala/Ala); recessive model (Leu432Val: Val/Val versus Leu/Val + Leu/Leu, Asn453Ser: Ser/Ser versus Asn/Ser + Asn/Asn, Arg48Gly: Gly/Gly versus Arg/Gly + Arg/Arg, and Ala119Ser: Ser/Ser versus Ala/Ser + Ala/Ala); and additive model (Leu432Val: Val versus Leu, Asn453Ser: Ser versus Asn, Arg48Gly: Gly versus Arg, and Ala119Ser: Ser versus Ala), respectively. Between-study heterogeneity was assessed by calculating Q-statistic (heterogeneity was considered statistically significant if P < 0.10) (Davey and Egger 1997) and quantified using the I 2 value, a value that describes the percentage of variation across studies that are due to heterogeneity rather than chance, where I 2 = 0 % indicates no observed heterogeneity, with 25 % regarded as low, 50 % as moderate, and 75 % as high (Higgins et al. 2003). If results were not heterogeneous, the pooled ORs were calculated by the fixed-effect model (we used the Q-statistic, which represents the magnitude of heterogeneity between-studies) (Mantel and Haenszel 1959). Otherwise, a random-effect model was used (when the heterogeneity between-studies was significant) (DerSimonian and Laird 1986). In addition to the comparison among all subjects, we also performed stratification analyses by cancer type (if one cancer type contained less than three individual studies, it was combined into the ‘‘other cancers’’ group), Moreover, the extent to which the combined risk estimate might be affected by individual studies was assessed by consecutively omitting every study from the meta-analysis (leave-one-out sensitivity analysis). This approach would also capture the effect of the oldest or first positive study (first study effect). In addition, we also ranked studies according to sample size, and then repeated this meta-analysis. Sample size was classified according to a minimum of 200 participants and those with fewer than 200 participants. The cite criteria were previously described (Klug et al. 2009). Last, sensitivity analysis was also performed, excluding studies whose allele frequencies in controls exhibited significant deviation from the Hardy–Weinberg equilibrium (HWE), given that the deviation may denote bias. HWE was calculated by using the goodness-of-fit test, and deviation was considered when P < 0.05. Begg’s funnel plots (Begg and Mazumdar 1994) and Egger’s linear regression test (Egger et al. 1997) were used to assess publication bias. If publication bias existed, the Duval and Tweedie nonparametric “trim and fill” method was used to adjust for it (Dual and Tweedie 2000). A meta-regression analysis was carried out to identify the major sources of between-studies variation in the results, using the log of the ORs from each study as dependent variables, and cancer type, ethnicity, sample size, HWE, and source of controls as the possible sources of heterogeneity. All of the calculations were performed using STATA version 10.0 (STATA Corporation, College Station, TX).
Results
Literature search and meta-analysis databases
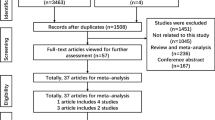
Figure 1 graphically illustrates the trial flow chart. A total of 548 articles regarding CYP1B1 Leu432Val, Asn453Ser, Arg48Gly, and Ala119Ser polymorphisms with respect to cancer were identified. After screening the titles and abstracts, 198 articles were excluded because they were duplicated. In addition, 235 articles were excluded because they were review articles, case reports, or other polymorphisms of CYP1B1. Last, of these published articles, four articles (24, 50, 73, 120) were excluded because their populations overlapped with other four included studies (17, 20, 48, 98). As summarized in Supplemental Table 1, 111 publications with 235 case–control studies were selected among the meta-analyses, including 54,124 cases and 62,932 controls for Leu432Val (115 studies from 107 publications), 30,532 cases and 39,193 controls for Asn453Ser (48 studies from 46 publications), 23,494 cases and 27,083 controls for Arg48Gly (38 studies from 34 publications), and 17,796 cases and 19,891 controls for Ala119Ser (34 studies from 30 publications). Among these studies, for Leu432Val polymorphism, there were 5 bladder cancer studies, 38 breast cancer studies, 12 colorectal cancer studies, 12 endometrial cancer studies, 7 head and neck cancer studies, 13 lung cancer studies, 7 ovarian cancer studies, 11 prostate cancer studies, and 10 studies with the “other cancer”. For Asn453Ser polymorphism, there were 15 breast cancer studies, 8 colorectal cancer studies, 5 endometrial cancer studies, 5 lung cancer studies, 3 ovarian cancer studies, 4 prostate cancer studies, and 8 studies with the “other cancer”. For Arg48Gly polymorphism, there were 12 breast cancer studies, 6 colorectal cancer studies, 5 endometrial cancer studies, 4 ovarian cancer studies, 5 prostate cancer studies, and 6 studies with the “other cancer”. For Ala119Ser polymorphism, there were 11 breast cancer studies, 3 colorectal cancer studies, 3 endometrial cancer studies, 4 lung cancer studies, 4 ovarian cancer studies, 4 prostate cancer studies, and 5 studies with the “other cancer”. All of the cases were pathologically confirmed.
Meta-analysis results
CYP1B1 Leu432Val
The evaluations of the association of CYP1B1 Leu432Val polymorphism with cancer risk are shown in Table 1. Overall, significantly increased cancer risk was found (dominant model: OR = 1.07, 95 % CI = 1.03–1.12, P h < 0.001, I 2 = 54.2 %; homozygous model: OR = 1.09, 95 % CI = 1.02–1.17, P h < 0.001, I 2 = 56.7 %; heterozygous model: OR = 1.06, 95 % CI = 1.01–1.10, P h < 0.001, I 2 = 41.5 %; additive model: OR = 1.05, 95 % CI = 1.02–1.09, P h < 0.001, I 2 = 65.0 %) when all the eligible studies were pooled into the meta-analysis. Then we performed subgroup analysis by cancer type. We found that individuals with the minor variant genotypes had a higher risk of endometrial cancer (dominant model: OR = 1.16, 95 % CI = 1.03–1.31, P h = 0.096, I 2 = 36.9 %), lung cancer (dominant model: OR = 1.25, 95 % CI = 1.07–1.48, P h = 0.061, I 2 = 40.9 %; recessive model: OR = 1.19, 95 % CI = 1.03–1.37, P h = 0.359, I 2 = 8.8 %; homozygous model: OR = 1.36, 95 % CI = 1.15–1.61, P h = 0.124, I 2 = 33.2 %; heterozygous model: OR = 1.19, 95 % CI = 1.06–1.34, P h = 0.153, I 2 = 29.9 %; additive model: OR = 1.19, 95 % CI = 1.05–1.35, P h = 0.015, I 2 = 53.0 %), and prostate cancer (heterozygous model: OR = 1.09, 95 % CI = 1.00–1.19, P h = 0.147, I 2 = 33.9 %) and had a lower risk of ovarian cancer (recessive model: OR = 0.82, 95 % CI = 0.68–1.00, P h = 0.278, I 2 = 19.8 %). We further examined the association of the CYP1B1 Leu432Val polymorphism and cancer risk according to cancer type and ethnicity (Table 1). For samples of Asians, significant association was found among head and neck cancer (recessive model: OR = 1.94, 95 % CI = 1.44–2.61, P h = 0.110, I 2 = 54.7 %; heterozygous model: OR = 1.31, 95 % CI = 1.10–1.57, P h = 0.198, I 2 = 38.2 %), prostate cancer (dominant model: OR = 1.35, 95 % CI = 1.03–1.78, P h = 0.218, I 2 = 34.4 %; homozygous model: OR = 1.85, 95 % CI = 1.03–3.32, P h = 0.401, I 2 = 0.0 %; additive model: OR = 1.36, 95 % CI = 1.07–1.71, P h = 0.160, I 2 = 45.4 %), and other cancer (dominant model: OR = 1.58, 95 % CI = 1.24–2.01, P h = 0.326, I 2 = 10.7 %; recessive model: OR = 2.23, 95 % CI = 1.31–3.79, P h = 0.800, I 2 = 0.0 %; homozygous model: OR = 2.57, 95 % CI = 1.50–4.40, P h = 0.866, I 2 = 0.0 %; heterozygous model: OR = 1.45, 95 % CI = 1.12–1.87, P h = 0.394, I 2 = 0.0 %; additive model: OR = 1.60, 95 % CI = 1.30–1.97, P h = 0.439, I 2 = 0.0 %). For samples of Caucasians, significant association was found among lung cancer (dominant model: OR = 1.37, 95 % CI = 1.04–1.81, P h = 0.018, I 2 = 66.6 %; heterozygous model: OR = 1.26, 95 % CI = 1.08–1.47, P h = 0.106, I 2 = 47.6 %), ovarian cancer (recessive model: OR = 0.77, 95 % CI = 0.62–0.95, P h = 0.575, I 2 = 0.0 %; homozygous model: OR = 0.77, 95 % CI = 0.60–0.97, P h = 0.425, I 2 = 0.0 %), and prostate cancer (dominant model: OR = 1.17, 95 % CI = 1.05–1.30, P h = 0.313, I 2 = 15.8 %; homozygous model: OR = 1.26, 95 % CI = 1.08–1.46, P h = 0.149, I 2 = 47.4 %; heterozygous model: OR = 1.16, 95 % CI = 1.04–1.31, P h = 0.571, I 2 = 0.0 %; additive model: OR = 1.13, 95 % CI = 1.05–1.22, P h = 0.170, I 2 = 43.5 %). We also examined the association of the CYP1B1 Leu432Val polymorphism and cancer risk according to cancer type and source of controls (Table 1). For the population-based studies, significant association was only found among lung cancer (dominant model: OR = 1.26, 95 % CI = 1.00–1.58, P h = 0.017, I 2 = 57.2 %; recessive model: OR = 1.17, 95 % CI = 1.01–1.36, P h = 0.207, I 2 = 26.7 %; homozygous model: OR = 1.40, 95 % CI = 1.05–1.86, P h = 0.050, I 2 = 48.4 %; additive model: OR = 1.17, 95 % CI = 1.00–1.37, P h = 0.006, I 2 = 62.6 %). For the hospital-based studies, significant association was found among breast cancer (homozygous model: OR = 1.32, 95 % CI = 1.13–1.54, P h = 0.182, I 2 = 23.0 %), endometrial cancer (dominant model: OR = 1.33, 95 % CI = 1.03–1.70, P h = 0.030, I 2 = 57.0 %), and prostate cancer (dominant model: OR = 1.14, 95 % CI = 1.00–1.29, P h = 0.177, I 2 = 34.6 %; homozygous model: OR = 1.22, 95 % CI = 1.00–1.50, P h = 0.167, I 2 = 38.1 %).
Significant heterogeneity was observed among these studies for dominant model comparison (P h < 0.001), recessive model comparison (P h < 0.0001), additive model comparison (P h < 0.001), homozygous model comparison (P h < 0.001), and heterozygous model comparison (P h < 0.001). Then, we assessed the source of heterogeneity by ethnicity, cancer type, source of controls, HWE, and sample size. Table 5 lists the results of meta-regression analysis. The results indicated that cancer type (heterozygous model: P = 0.021), ethnicity (heterozygous model: P = 0.044), source of controls (homozygous model: P = 0.013; additive model: P = 0.014; recessive model: P = 0.021; dominant model: P = 0.034), and sample size (heterozygous model: P = 0.004; dominant model: P = 0.010) but not HWE (dominant model: P = 0.262; recessive model: P = 0.607; heterozygous model: P = 0.345; homozygous model: P = 0.273; additive model: P = 0.344) contributed to substantial heterogeneity among the meta-analysis. Examining genotype frequencies in the controls, significant deviation from HWE was detected in the thirteen studies (Catsburg et al. 2012; Wang et al. 2011a,b; Sliwinski et al. 2010; Yadav et al. 2009; Vrana et al. 2010; Shimada et al. 2009; Sellers et al. 2005; Cicek et al. 2005; Le Marchand et al. 2005; Sasaki et al. 2004; Tanaka et al. 2002; Bailey et al. 1998; Maurya et al. 2014). When these studies were excluded, the results were changed among prostate cancer (heterozygous model: OR = 1.09, 95 % CI = 0.93–1.27), Asians of head and neck cancer (recessive model: OR = 1.17, 95 % CI = 0.62–2.24; heterozygous model: OR = 1.09, 95 % CI = 0.81–1.47), and hospital-based studies of other cancer (dominant model: OR = 1.30, 95 % CI = 1.08–1.57; homozygous model: OR = 1.36, 95 % CI = 1.00–1.83; heterozygous model: OR = 1.26, 95 % CI = 1.04–1.54; additive model: OR = 1.20, 95 % CI = 1.05–1.38), as shown in Table 6. In addition, when our meta-analysis was performed excluding studies with small sample sizes, the results were changed among prostate cancer (heterozygous model: OR = 1.08, 95 % CI = 0.99–1.18), Asians of prostate cancer (dominant model: OR = 1.21, 95 % CI = 0.88–1.65; homozygous model: OR = 1.38, 95 % CI = 0.67–2.83; additive model: OR = 1.19, 95 % CI = 0.91–1.57), hospital-based studies of endometrial cancer (dominant model: OR = 1.25, 95 % CI = 0.95–1.63), and hospital-based studies of prostate cancer (dominant model: OR = 1.10, 95 % CI = 0.97–1.26; homozygous model: OR = 1.18, 95 % CI = 0.96–1.44), as shown in Table 7. Last, a single study involved in the meta–analysis was deleted each time to reflect the influence of individual data set to the pooled ORs, the results did not be changed among overall analysis and any subgroup analysis.
We performed Begg’s funnel plot and Egger’s test to assess the publication bias of literatures. Begg’s funnel plots and Egger’s test suggested that there might be publication bias in any genetic model (dominant model: P = 0.004; heterozygous model: P = 0.006; recessive model: P = 0.009; additive model: P = 0.003; homozygous model: P = 0.001). This might be a limitation for the meta-analysis because studies with null findings, especially those with small sample size, are less likely to be published. Adjusting for possible publication bias using the Duval and Tweedie nonparametric ‘‘trim and fill’’ method for overall studies, the results did not change between CYP1B1 Leu432Val polymorphism with the risk of cancer. Figure 2 lists the Duval and Tweedie nonparametric “trim and fill’’ methods funnel plot in any genetic model.
CYP1B1 Asn453Ser
The evaluations of the association of CYP1B1 Asn453Ser polymorphism with cancer risk are shown in Table 2. Overall, no significant association was found among any genetic model (dominant model: OR = 1.01, 95 % CI = 0.96–1.05, P h = 0.012, I 2 = 34.6 %; recessive model: OR = 0.98, 95 % CI = 0.90–1.07, P h = 0.818, I 2 = 0.0 %; homozygous model: OR = 0.98, 95 % CI = 0.90–1.08, P h = 0.787, I 2 = 0.0 %; heterozygous model: OR = 1.00, 95 % CI = 0.95–1.05, P h = 0.016, I 2 = 34.5 %; additive model: OR = 0.99, 95 % CI = 0.95–1.03, P h = 0.050, I 2 = 28.0 %) when all the eligible studies were pooled into the meta-analysis. Then we performed subgroup analysis by cancer type. We found that individuals with the minor variant genotypes had a higher risk of prostate cancer (dominant model: OR = 1.19, 95 % CI = 1.02–1.39, P h = 0.368, I 2 = 0.0 %) and had a lower risk of endometrial cancer (heterozygous model: OR = 0.81, 95 % CI = 0.69–0.95, P h = 0.668, I 2 = 0.0 %; additive model: OR = 0.82, 95 % CI = 0.71–0.94, P h = 0.379, I 2 = 0.0 %). We further examined the association of the CYP1B1 Asn453Ser polymorphism and cancer risk according to cancer type and ethnicity (Table 2). For samples of Caucasians, significant decreased cancer risk was found among endometrial cancer (dominant model: OR = 0.82, 95 % CI = 0.71–0.94, P h = 0.700, I 2 = 0.0 %; heterozygous model: OR = 0.81, 95 % CI = 0.69–0.95, P h = 0.668, I 2 = 0.0 %; additive model: OR = 0.82, 95 % CI = 0.71–0.94, P h = 0.379, I 2 = 0.0 %). We also examined the association of the CYP1B1 Asn453Ser polymorphism and cancer risk according to cancer type and source of controls (Table 2). For the population-based studies, significant association was only found among endometrial cancer (dominant model: OR = 0.80, 95 % CI = 0.68–0.94, P h = 0.564, I 2 = 0.0 %; heterozygous model: OR = 0.81, 95 % CI = 0.69–0.95, P h = 0.668, I 2 = 0.0 %; additive model: OR = 0.82, 95 % CI = 0.71–0.94, P h = 0.379, I 2 = 0.0 %).
Significant heterogeneity was observed among these studies for dominant model comparison (P h = 0.012), additive model comparison (P h = 0.050), and heterozygous model comparison (P h = 0.016). Then, we assessed the source of heterogeneity by ethnicity, cancer type, source of controls, HWE, and sample size. Table 5 lists the results of meta-regression analysis. The results indicated that cancer type (dominant model: P = 0.766; heterozygous model: P = 0.880; additive model: P = 0.717), ethnicity (dominant model: P = 0.847; heterozygous model: P = 0.897; additive model: P = 0.555), source of controls (dominant model: P = 0.635; heterozygous model: P = 0.743; additive model: P = 0.845), sample size (dominant model: P = 0.576; heterozygous model: P = 0.988; additive model: P = 0.291), and HWE (dominant model: P = 0.399; heterozygous model: P = 0.118; additive model: P = 0.618) did not contribute to substantial heterogeneity among the meta-analysis. Examining genotype frequencies in the controls, significant deviation from HWE was detected in the one study (Figueroa et al. 2008). When this study was excluded, the results did not be changed among overall analysis and other subgroup analysis, as shown in Table 6. In addition, when our meta-analysis was performed excluding studies with small sample sizes, the results did not also change in the meta-analysis, as shown in Table 7. Last, a single study involved in the meta-analysis was deleted each time to reflect the influence of individual data set to the pooled ORs, the results were changed among prostate cancer (OR = 1.15, 95 % CI = 0.94–1.40).
Both Begg’s funnel plot and Egger’s test were performed to assess the publication bias of literatures. The Egger’s test results (dominant model: P = 0.497; recessive model: P = 0.078; additive model: P = 0.917; Homozygous model: P = 0.098; Heterozygous model: P = 0.539) and Begg’s funnel plot (Fig. 3) suggested no evidence of publication bias in the meta-analysis.
CYP1B1 Arg48Gly
The evaluations of the association of CYP1B1 Arg48Gly polymorphism with cancer risk are shown in Table 3. Overall, no significant association was found (dominant model: OR = 0.99, 95 % CI = 0.91–1.07, P h < 0.001, I 2 = 71.9 %; recessive model: OR = 0.93, 95 % CI = 0.83–1.04, P h < 0.001, I 2 = 61.7 %; homozygous model: OR = 0.91, 95 % CI = 0.79–1.06, P h < 0.001, I 2 = 74.3 %; heterozygous model: OR = 1.01, 95 % CI = 0.94–1.09, P h < 0.001, I 2 = 65.0 %) when all the eligible studies were pooled into the meta-analysis. In addition, high between-studies heterogeneity was observed among overall analysis (additive model: I 2 = 77.2). Then we performed subgroup analysis by cancer type. We found that individuals with the minor variant genotypes had a lower risk of endometrial cancer (recessive model: OR = 0.55, 95 % CI = 0.42–0.73, P h = 0.176, I 2 = 36.8 %). We further examined the association of the CYP1B1 Arg48Gly polymorphism and cancer risk according to cancer type and ethnicity (Table 3). For samples of Caucasians, significant decreased cancer risk was found among endometrial cancer (recessive model: OR = 0.42, 95 % CI = 0.28–0.61, P h = 0.419, I 2 = 0.0 %; homozygous model: OR = 0.29, 95 % CI = 0.12–0.71, P h = 0.020, I 2 = 74.3 %). We also examined the association of the CYP1B1 Arg48Gly polymorphism and cancer risk according to cancer type and source of controls (Table 3). For the population-based studies, significant association was found among other cancer (heterozygous model: OR = 1.25, 95 % CI = 1.04–1.50, P h = 0.152, I 2 = 46.9 %). For the hospital-based studies, significant decreased cancer risk was found among endometrial cancer (recessive model: OR = 0.46, 95 % CI = 0.31–0.67, P h = 0.114, I 2 = 54.0 %).
Significant heterogeneity was observed among these studies for dominant model comparison (P h < 0.001), recessive model comparison (P h < 0.0001), additive model comparison (P h < 0.001), homozygous model comparison (P h < 0.001), and heterozygous model comparison (P h < 0.001). Then, we assessed the source of heterogeneity by ethnicity, cancer type, source of controls, HWE, and sample size. Table 5 lists the results of meta-regression analysis. The results indicated that source of controls (dominant model: P = 0.005; homozygous model: P = 0.022; heterozygous model: P = 0.009; additive model: P = 0.036) and sample size (recessive model: P = 0.008; homozygous model: P = 0.044) but not cancer type (dominant model: P = 0.137; recessive model: P = 0.102; homozygous model: P = 0.092; heterozygous model: P = 0.164; additive model: P = 0.063), ethnicity (dominant model: P = 0.166; recessive model: P = 0.454; homozygous model: P = 0.303; heterozygous model: P = 0.168; additive model: P = 0.135), and HWE (dominant model: P = 0.703; recessive model: P = 0.759; heterozygous model: P = 0.903; homozygous model: P = 0.505; additive model: P = 0.672) contributed to substantial heterogeneity among the meta-analysis. Examining genotype frequencies in the controls, significant deviation from HWE was detected in the eight studies (Reding et al. 2012; Salinas-Sánchez et al. 2012a; Beuten et al. 2009; Figueroa et al. 2008; Holt et al. 2007; Sasaki et al. 2003, 2004; Tanaka et al. 2002). When these studies were excluded, the results were changed among other cancer (heterozygous model: OR = 1.25, 95 % CI = 1.04–1.50) and hospital-based studies of endometrial cancer (dominant model: OR = 0.33, 95 % CI = 0.16–0.69; homozygous model: OR = 0.19, 95 % CI = 0.10–0.33; heterozygous model: OR = 0.42, 95 % CI = 0.27–0.66; additive model: OR = 0.45, 95 % CI = 0.35–0.59), as shown in Table 6. In addition, when our meta-analysis was performed excluding studies with small sample sizes, the results were changed among hospital-based studies of endometrial cancer (recessive model: OR = 0.53, 95 % CI = 0.22–1.26), as shown in Table 7. Last, a single study involved in the meta–analysis was deleted each time to reflect the influence of individual data set to the pooled ORs, the results did not changed among the meta-analysis.
High between-studies heterogeneity was found among breast cancer, endometrial cancer, and other cancer. When the study of Zimarina et al. (2004) was excluded, the high between-studies heterogeneity were deleted among breast cancer (dominant model: I 2 = 26.2; homozygous model: I 2 = 8.1; heterozygous model: I 2 = 40.9; additive model: I 2 = 16.1), Caucasian of breast cancer (dominant model: I 2 = 0.0; recessive model: I 2 = 17.9; homozygous model: I 2 = 32.1; heterozygous model: I 2 = 0.0; additive model: I 2 = 35.1), hospital-based studies of breast cancer (dominant model: I 2 = 0.0; recessive model: I 2 = 3.0; homozygous model: I 2 = 0.0; heterozygous model: I 2 = 21.3; additive model: I 2 = 0.0), endometrial cancer (dominant model: I 2 = 58.1; homozygous model: I 2 = 42.2; heterozygous model: I 2 = 27.7; additive model: I 2 = 68.8). When the study of Zienolddiny et al. (2008) was excluded, the high between-studies heterogeneity were deleted among other cancer (dominant model: I 2 = 73.5; recessive model: I 2 = 21.1; homozygous model: I 2 = 29.8; heterozygous model: I 2 = 73.2; additive model: I 2 = 68.1).
We performed Begg’s funnel plot and Egger’s test to assess the publication bias of literatures. Begg’s funnel plots and Egger’s test suggested that there might be publication bias in recessive model (P = 0.033) and homozygous model (P = 0.047). Adjusting for possible publication bias using the Duval and Tweedie nonparametric ‘‘trim and fill’’ method for overall studies, the results did not change between CYP1B1 Arg48Gly polymorphism with the risk of cancer.
CYP1B1 Ala119Ser
The evaluations of the association of CYP1B1 Ala119Ser polymorphism with cancer risk are shown in Table 4. Overall, no significant association was found (recessive model: OR = 1.04, 95 % CI = 0.89–1.20, P h < 0.001, I 2 = 66.2 %; homozygous model: OR = 1.03, 95 % CI = 0.87–1.24, P h < 0.001, I 2 = 74.5 %) when all the eligible studies were pooled into the meta-analysis. In addition, high between-studies heterogeneity was observed among overall analysis (dominant model: I 2 = 79.7; heterozygous model: I 2 = 75.3; additive model: I 2 = 81.9). Then we performed subgroup analysis by cancer type. We found that individuals with the minor variant genotypes had a high risk of prostate cancer (recessive model: OR = 1.45, 95 % CI = 1.07–1.97, P h = 0.160, I 2 = 45.4 %; homozygous model: OR = 1.88, 95 % CI = 1.08–3.28, P h = 0.081, I 2 = 60.2 %) and other cancer (recessive model: OR = 1.78, 95 % CI = 1.34–2.37, P h = 0.293, I 2 = 19.2 %). We further examined the association of the CYP1B1 Ala119Ser polymorphism and cancer risk according to cancer type and ethnicity (Table 4). For samples of Caucasians, significant decreased cancer risk was found among other cancer (heterozygous model: OR = 0.62, 95 % CI = 0.38–1.00, P h = 0.045, I 2 = 67.7 %). We also examined the association of the CYP1B1 Arg48Gly polymorphism and cancer risk according to cancer type and source of controls (Table 4). For the population-based studies, significant association was only observed among breast cancer (homozygous model: OR = 1.12, 95 % CI = 1.01–1.25, P h = 0.151, I 2 = 36.3 %). For the hospital-based studies, significant association was found among breast cancer (dominant model: OR = 0.60, 95 % CI = 0.37–0.97, P h = 0.016, I 2 = 70.9 %; heterozygous model: OR = 0.73, 95 % CI = 0.58–0.92, P h = 0.100, I 2 = 52.1 %), prostate cancer (dominant model: OR = 1.45, 95 % CI = 1.07–1.97, P h = 0.160, I 2 = 45.4 %; homozygous model: OR = 1.88, 95 % CI = 1.08–3.28, P h = 0.081, I 2 = 60.2 %), and other cancer (dominant model: OR = 1.78, 95 % CI = 1.34–2.37, P h = 0.293, I 2 = 19.2 %).
Significant heterogeneity was observed among these studies for dominant model comparison (P h < 0.001), recessive model comparison (P h < 0.0001), additive model comparison (P h < 0.001), homozygous model comparison (P h < 0.001), and heterozygous model comparison (P h < 0.001). Table 5 lists the results of meta-regression analysis. Then, we assessed the source of heterogeneity by ethnicity, cancer type, source of controls, HWE, and sample size. The results indicated that cancer type (recessive model: P = 0.008; homozygous model: P = 0.007; additive model: P = 0.038) and ethnicity (dominant model: P = 0.031; heterozygous model: P = 0.043; additive model: P = 0.016) but not source of control (dominant model: P = 0.201; recessive model: P = 0.564; homozygous model: P = 0.269; heterozygous model: P = 0.248; additive model: P = 0.392), sample size (dominant model: P = 0.382; recessive model: P = 0.117; homozygous model: P = 0.084; heterozygous model: P = 0.541; additive model: P = 0.142), and HWE (dominant model: P = 0.676; recessive model: P = 0.586; heterozygous model: P = 0.990; homozygous model: P = 0.644; additive model: P = 0.860) contributed to substantial heterogeneity among the meta-analysis. Examining genotype frequencies in the controls, significant deviation from HWE was detected in the nine studies (Reding et al. 2012; Salinas-Sánchez et al. 2012b; Salinas-Sánchez et al. 2012a; Gulyaeva et al. 2008; Zienolddiny et al. 2008; Sasaki et al. 2003; 2004; Maurya et al. 2014; Rodrigues et al. 2011). When these studies were excluded, the results were changed among other cancer (recessive model: OR = 1.11, 95 % CI = 0.45–2.73) and endometrial cancer (dominant model: OR = 0.38, 95 % CI = 0.24–0.60), as shown in Table 6. In addition, when our meta-analysis was performed excluding studies with small sample sizes, the results did not changed among this meta-analysis, as shown in Table 7. Last, a single study involved in the meta–analysis was deleted each time to reflect the influence of individual data set to the pooled ORs. When the study of Zimarina et al. (2004) was excluded, significant association was found between CYP1B1 Ala119Ser polymorphism and breast cancer susceptibility in Caucasians (homozygous model: OR = 1.15, 95 % CI = 1.03–1.29; recessive model: OR = 1.16, 95 % CI = 1.04–1.29). In addition, when the study of Zimarina et al. (2004) was excluded, the results were also changed among hospital-based studies of breast cancer (dominant model: OR = 0.82, 95 % CI = 0.64–1.03; heterozygous model: OR = 0.82, 95 % CI = 0.63–1.05).
High between-studies heterogeneity was found among breast cancer, endometrial cancer, prostate cancer, and other cancer. When the study of Zimarina et al. (2004) was excluded, the high between-studies heterogeneity were deleted among breast cancer (additive model: I 2 = 51.0 %), Caucasian of breast cancer (dominant model: I 2 = 45.9 %; recessive model: I 2 = 27.3 %; homozygous model: I 2 = 44.4 %; heterozygous model: I 2 = 17.3 %; additive model: I 2 = 58.8 %), hospital-based studies of breast cancer (homozygous model: I 2 = 20.9; additive model: I 2 = 0.0). When the study of Tanaka et al. (2002) was excluded, the high between-studies heterogeneity were deleted among prostate cancer (dominant model: I 2 = 63.7; heterozygous model: I 2 = 41.4; additive model: I 2 = 0.0) and hospital-based studies of prostate cancer (dominant model: I 2 = 63.7; heterozygous model: I 2 = 41.4; additive model: I 2 = 0.0).
Both Begg’s funnel plot and Egger’s test were performed to assess the publication bias of literatures. The Egger’s test results (dominant model: P = 0.949; recessive model: P = 0.271; additive model: P = 0.911; Homozygous model: P = 0.241; Heterozygous model: P = 0.535) and Begg’s funnel plot (Figure not shown) suggested no evidence of publication bias in the meta-analysis.
Discussion
CYP1B1 is commonly over-expressed inhumanmalignancies and activates a variety of carcinogens. For example, CYP1B1 catalyzes both the formation of dihydrodiols of specific PAHs and their subsequent oxidation to carcinogenic dihydrodiol epoxides. The importance of CYP1B1 in chemical carcinogens is well illustrated in animal models in which metabolites of CYP1B1 were shown to induce PCa (Williams et al. 2000; Cavalieri et al. 2002). Furthermore, CYP1B1-null mice, created by targeted gene disruption in embryonic stem cells, were protected from 7,12-dimethylbenz(a)anthracene-induced malignant lymphomas (Buters et al. 1999). Many studies have reported the role of CYP1B1 Leu432Val, Asn453Ser, Arg48Gly, and Ala119Ser with all cancers risk, but the results remained controversial. Some recent meta-analyses analyzed such an association only for single cancer such as breast cancer, lung cancer, prostate cancer, endometrial cancer, and so on. Importantly, several published studies were not included in the previous meta-analysis and additional original studies with larger sample sizes have been published since then. Hence, the correlation of these polymorphic genes remains unknown. In order to derive a more precise estimation of association, we performed the meta-analysis of CYP1B1 Leu432Val (54,124 cases and 62,932 controls), Asn453Ser (30,532 cases and 39,193 controls), Arg48Gly polymorphisms (23,494 cases and 27,083), and Ala119Ser (17,796 cases and 19,891 controls) with cancer risk.
For Asn453Ser and Arg48Gly polymorphisms, significant decreased endometrial cancer was observed among Caucasians. For Ala119Ser polymorphism, we found that individuals with the minor variant genotypes had a high risk of prostate cancer (recessive model: OR = 1.45, 95 % CI = 1.07–1.97; homozygous model: OR = 1.88, 95 % CI = 1.08–3.28) and Caucasians of breast cancer (homozygous model: OR = 1.15, 95 % CI = 1.03–1.29; recessive model: OR = 1.16, 95 % CI = 1.04–1.29). For Leu432Val polymorphisms, We found that individuals with the minor variant genotypes had a higher risk of endometrial cancer (dominant model: OR = 1.16, 95 % CI = 1.03–1.31), lung cancer (dominant model: OR = 1.25, 95 % CI = 1.07–1.48; recessive model: OR = 1.19, 95 % CI = 1.03–1.37; homozygous model: OR = 1.36, 95 % CI = 1.15–1.61; heterozygous model: OR = 1.19, 95 % CI = 1.06–1.34; additive model: OR = 1.19, 95 % CI = 1.05–1.35) and had a lower risk of ovarian cancer (recessive model: OR = 0.82, 95 % CI = 0.68–1.00).
For Asn453Ser and Arg48Gly polymorphisms, significant decreased endometrial cancer risk was observed among Caucasians, but not Asians and Africans. For Ala119Ser polymorphism, significant increased breast cancer risk was observed among Caucasians, but not Asians and Africans. For Leu432Val polymorphisms, significant increased lung cancer among Caucasians, but not Asians and Africans. The results suggested a possible role of ethnic difference in genetic background and the environment they lived in. It should be considered that the apparent inconsistency of these results may underlie differences in ethnicity, lifestyle and disease prevalence as well as possible limitations due to the relatively small sample size. The current knowledge of carcinogenesis indicates a multi-factorial and multistep process that involves various genetic alterations and several biological pathways. Thus, it is unlikely that risk factors of cancer work in isolation from each other. And the same polymorphisms may play different roles in cancer susceptibility, because cancer is a complicated multi-genetic disease, and different genetic backgrounds may contribute to the discrepancy. And even more importantly, the low penetrance genetic effects of single polymorphism may largely depend on interaction with other polymorphisms and/or a particular environmental exposure.
Based on biochemical properties described for CYP1B1 polymorphism, we would expect that the allele would be associated with higher susceptibility for all types of cancer. However, our results showed that such association was observed among ovarian cancer, lung cancer, and endometrial cancer for CYP1B1 Leu432Val, endometrial cancer for Asn453Ser and Arg48Gly, and prostate cancer and breast cancer for CYP1B1 Ala119Ser, suggesting that other factors may be modulating the CYP1B1 polymorphisms functionality. Several previous studies assessed the effect of CYP1B1 Leu432Val, Arg48Gly, and Ala119Ser polymorphisms on these cancers risk, which finding is consistent with our results. However, the exact mechanism for association between different tumor sites and CYP1B1 Leu432Val, Asn453Ser, Arg48Gly, and Ala119Ser polymorphisms was not clear, carcinogenetic mechanism may differ by different tumor sites and the CYP1B1 genetic variants may exert varying effects in different cancers.
In the present meta-analysis, between-studies heterogeneity was observed between CYP1B1 Leu432Val, Asn453Ser, Arg48Gly, and Ala119Ser polymorphisms and cancer of risk. Meta-regression analysis indicated that cancer type, ethnicity, source of controls, and sample size contributed to substantial heterogeneity among the meta-analysis for Leu432Val polymorphism, source of controls and sample size contributed to substantial heterogeneity among the meta-analysis for Arg48Gly, and cancer type and ethnicity contributed to substantial heterogeneity among the meta-analysis for Ala119Ser. The hospital-based studies may have some biases because such controls may contain certain benign diseases which are prone to develop malignancy and may not be very representative of the general population. The small number studies hinder the ability of drawing more definite conclusions. Thus, the use of a proper and representative cancer-free control subjects and the large sample size studies are very important in reducing biases in such genotype association studies. And this indicates that it may be not appropriate to use an overall estimation of the relationship between CYP1B1 polymorphism and cancer risk.
The current meta-analysis has some strength compared with individual studies and previous meta-analyses. First, differently from previous meta-analyses, we explored the impact of CYP1B1 Leu432Val, Asn453Ser, Arg48Gly, and Ala119Ser on a great diversity of cancer sites, allowing for a general view of its influence on cancer susceptibility. Second, our meta-analysis explores and analyzes the sources of heterogeneity between studies about CYP1B1 in cancer. Third, a systematic review of the association of Leu432Val, Asn453Ser, Arg48Gly, and Ala119Ser polymorphisms with the risk of cancer is statistically more powerful than any single study. Fourth, the quality of eligible studies included in current meta-analysis was satisfactory and met our inclusion criterion.
There are also still some limitations inherited from the published studies. First, our results were based on single-factor estimates without adjustment for other risk factors including alcohol usage, environmental factors and other lifestyle. Second, in the subgroup analysis may have had insufficient statistical power to check an association. Third, the controls were not uniformly defined. Fourth, a potential limitation of our results is the small number of studies for some tumor sites and subgroups, which hinders the ability of drawing more definite conclusions for some results. For these cases, the interpretation of the results should be taken carefully.
In summary, this meta-analysis suggests that Leu432Val polymorphism is associated with ovarian cancer, lung cancer, and endometrial cancer risk, Asn453Ser and Arg48Gly polymorphisms are associated with endometrial cancer risk among Caucasians, and Ala119Ser polymorphism is associated with prostate cancer risk and Caucasians of breast cancer risk. In addition, our work also points out the importance of new studies for Ala119Ser polymorphism in endometrial cancer, because high heterogeneity was observed (I 2 > 75 %).
References
Ahsan H, Chen Y, Whittemore AS, Kibriya MG, Gurvich I, Senie RT, Santella RM (2004) A family-based genetic association study of variants in estrogen-metabolism genes COMT and CYP1B1 and breast cancer risk. Breast Cancer Res Treat 85:121–131
Ashton KA, Proietto A, Otton G, Symonds I, McEvoy M, Attia J, Gilbert M, Hamann U, Scott RJ (2010) Polymorphisms in genes of the steroid hormone biosynthesis and metabolism pathways and endometrial cancer risk. Cancer Epidemiol 34:328–337
Bailey LR, Roodi N, Dupont WD, Parl FF (1998) Association of cytochrome P450 1B1 (CYP1B1) polymorphism with steroid receptor status in breast cancer. Cancer Res 58:5038–5041
Begg CB, Mazumdar M (1994) Operating characteristics of a rank correlation test for publication bias. Biometrics 50:1088–1101
Berber U, Yilmaz I, Yilmaz O, Haholu A, Kucukodaci Z, Ates F, Demirel D (2013) CYP1A1 (Ile462Val), CYP1B1 (Ala119Ser and Val432Leu), GSTM1 (null), and GSTT1 (null) polymorphisms and bladder cancer risk in a Turkish population. Asian Pac J Cancer Prev 14:3925–3929
Berndt SI, Chatterjee N, Huang WY, Chanock SJ, Welch R, Crawford ED, Hayes RB (2007) Variant in sex hormone-binding globulin gene and the risk of prostate cancer. Cancer Epidemiol Biomarkers Prev 16:165–168
Bethke L, Webb E, Sellick G, Rudd M, Penegar S, Withey L, Qureshi M, Houlston R (2007) Polymorphisms in the cytochrome P450 genes CYP1A2, CYP1B1, CYP3A4, CYP3A5, CYP11A1, CYP17A1, CYP19A1 and colorectal cancer risk. BMC Cancer 7:123
Beuten J, Gelfond JA, Byrne JJ, Balic I, Crandall AC, Johnson-Pais TL, Thompson IM, Price DK, Leach RJ (2008) CYP1B1 variants are associated with prostate cancer in non-Hispanic and Hispanic Caucasians. Carcinogenesis 29:1751–1757
Beuten J, Gelfond JA, Franke JL, Weldon KS, Crandall AC, Johnson-Pais TL, Thompson IM, Leach RJ (2009) Single and multigenic analysis of the association between variants in 12 steroid hormone metabolism genes and risk of prostate cancer. Cancer Epidemiol Biomarkers Prev 18:1869–1880
Buters JT, Sakai S, Richter T, Pineau T, Alexander DL, Savas U, Doehmer J, Ward JM, Jefcoate CR, Gonzalez FJ (1999) Cytochrome P450 CYP1B1 determines susceptibility to 7, 12-dimethylbenz (a) Anthraceneinduced lymphomas. Proc Natl Acad Sci 96:1977–1982
Catsburg C, Joshi AD, Corral R, Lewinger JP, Koo J, John EM, Ingles SA, Stern MC (2012) Polymorphisms in carcinogen metabolism enzymes, fish intake, and risk of prostate cancer. Carcinogenesis 33:1352–1359
Cavalieri EL, Devanesan P, Bosland MC, Badawi AF, Rogan EG (2002) Catechol estrogen metabolites and conjugates in different regions of the prostate of Noble rats treated with 4-hydroxyestradiol: implications for estrogen-induced initiation of prostate cancer. Carcinogenesis 23:329–333
Cecchin E, Russo A, Campagnutta E, Martella L, Toffoli G (2004) Lack of association of CYP1 B1*3 polymorphism and ovarian cancer in a Caucasian population. Int J Biol Markers 19:160–163
Cerne JZ, Pohar-Perme M, Novakovic S, Frkovic-Grazio S, Stegel V, Gersak K (2011a) Combined effect of CYP1B1, COMT, GSTP1, and MnSOD genotypes and risk of postmenopausal breast cancer. J Gynecol Oncol. 22:110–119
Cerne JZ, Novakovic S, Frkovic-Grazio S, Pohar-Perme M, Stegel V, Gersak K (2011b) Estrogen metabolism genotypes, use of long-term hormone replacement therapy and risk of postmenopausal breast cancer. Oncol Rep 26:479–485
Chang BL, Zheng SL, Isaacs SD, Turner A, Hawkins GA, Wiley KE, Bleecker ER, Walsh PC, Meyers DA, Isaacs WB, Xu J (2003) Polymorphisms in the CYP1B1 gene are associated with increased risk of prostate cancer. Br J Cancer 89:1524–1529
Chen B, Qiu LX, Li Y, Xu W, Wang XL, Zhao WH, Wu JQ (2010) The CYP1B1 Leu432Val polymorphism contributes to lung cancer risk: evidence from 6501 subjects. Lung Cancer 70:247–252
Church TR, Haznadar M, Geisser MS, Anderson KE, Caporaso NE, Le C, Abdullah SB, Hecht SS, Oken MM, Van Ness B (2010) Interaction of CYP1B1, cigarette-smoke carcinogen metabolism, and lung cancer risk. Int J Mol Epidemiol Genet 1:295–309
Cicek MS, Liu X, Casey G, Witte JS (2005) Role of androgen metabolism genes CYP1B1, PSA/KLK3, and CYP11alpha in prostate cancer risk and aggressiveness. Cancer Epidemiol Biomarkers Prev 14:2173–2177
Cleary SP, Cotterchio M, Shi E, Gallinger S, Harper P (2010) Cigarette smoking, genetic variants in carcinogen-metabolizing enzymes, and colorectal cancer risk. Am J Epidemiol 172:1000–1014
Cote ML, Wenzlaff AS, Bock CH, Land SJ, Santer SK, Schwartz DR, Schwartz AG (2007) Combinations of cytochrome P-450 genotypes and risk of early-onset lung cancer in Caucasians and African Americans: a population-based study. Lung Cancer 55:255–262
Cote ML, Yoo W, Wenzlaff AS, Prysak GM, Santer SK, Claeys GB, Van Dyke AL, Land SJ, Schwartz AG (2009) Tobacco and estrogen metabolic polymorphisms and risk of non-small cell lung cancer in women. Carcinogenesis 30:626–635
Cotterchio M, Boucher BA, Manno M, Gallinger S, Okey AB, Harper PA (2008) Red meat intake, doneness, polymorphisms in genes that encode carcinogen-metabolizing enzymes, and colorectal cancer risk. Cancer Epidemiol Biomarkers Prev 17:3098–3107
Cui L, Dillehay K, Chen W, Shen D, Dong Z, Li W (2012) Association of the CYP1B1 Leu432Val polymorphism with the risk of prostate cancer: a meta-analysis. Mol Biol Rep 39:7465–7471
Cussenot O, Azzouzi AR, Nicolaiew N, Fromont G, Mangin P, Cormier L, Fournier G, Valeri A, Larre S, Thibault F, Giordanella JP, Pouchard M, Zheng Y, Hamdy FC, Cox A, Cancel-Tassin G (2007) Combination of polymorphisms from genes related to estrogen metabolism and risk of prostate cancers: the hidden face of estrogens. J Clin Oncol 25:3596–3602
Davey SG, Egger M (1997) Meta-analyses of randomized controlled trials. Lancet 350:1182
De Roos AJ, Gold LS, Wang S, Hartge P, Cerhan JR, Cozen W, Yeager M, Chanock S, Rothman N, Severson RK (2006a) Metabolic gene variants and risk of non-Hodgkin’s lymphoma. Cancer Epidemiol Biomarkers Prev 15:1647–1653
De Roos AJ, Rothman N, Brown M, Bell DA, Pittman GS, Shapiro WR, Selker RG, Fine HA, Black PM, Inskip PD (2006b) Variation in genes relevant to aromatic hydrocarbon metabolism and the risk of adult brain tumors. Neuro Oncol 8:145–155
De Vivo I, Hankinson SE, Li L, Colditz GA, Hunter DJ (2002) Association of CYP1B1 polymorphisms and breast cancer risk. Cancer Epidemiol Biomarkers Prev 11:489–492
Delort L, Chalabi N, Satih S, Rabiau N, Kwiatkowski F, Bignon YJ, Bernard-Gallon DJ (2008) Association between genetic polymorphisms and ovarian cancer risk. Anticancer Res 28:3079–3081
Delort L, Satih S, Kwiatkowski F, Bignon YJ, Bernard-Gallon DJ (2010) Evaluation of breast cancer risk in a multigenic model including low penetrance genes involved in xenobiotic and estrogen metabolisms. Nutr Cancer 62:243–251
DerSimonian R, Laird N (1986) Meta-analysis in clinical trials. Control Clin Trials 7:177–188
Diergaarde B, Potter JD, Jupe ER, Manjeshwar S, Shimasaki CD, Pugh TW, Defreese DC, Gramling BA, Evans I, White E (2008) Polymorphisms in genes involved in sex hormone metabolism, estrogen plus progestin hormone therapy use, and risk of postmenopausal breast cancer. Cancer Epidemiol Biomarkers Prev 17:1751–1759
Doherty JA, Weiss NS, Freeman RJ, Dightman DA, Thornton PJ, Houck JR, Voigt LF, Rossing MA, Schwartz SM, Chen C (2005) Genetic factors in catechol estrogen metabolism in relation to the risk of endometrial cancer. Cancer Epidemiol Biomarkers Prev 14:357–366
dos Santos RA, Teixeira AC, Mayorano MB, Carrara HH, de Andrade J, Takahashi CS (2011) Variability in estrogen-metabolizing genes and their association with genomic instability in untreated breast cancer patients and healthy women. J Biomed Biotechnol 2011:571784
Dual S, Tweedie R (2000) A nonparametric “trim and fill” method of accounting for publication bias in meta-analysis. J Am Stat Assoc 95:89–98
Dunning AM, Dowsett M, Healey CS, Tee L, Luben RN, Folkerd E, Novik KL, Kelemen L, Ogata S, Pharoah PD, Easton DF, Day NE, Ponder BA (2004) Polymorphisms associated with circulating sex hormone levels in postmenopausal women. J Natl Cancer Inst 96:936–945
Economopoulos KP, Sergentanis TN (2010) Three polymorphisms in cytochrome P450 1B1 (CYP1B1) gene and breast cancer risk: a meta-analysis. Breast Cancer Res Treat 122:545–551
Egger M, Smith DG, Schneider M, Minder C (1997) Bias in meta-analysis detected by a simple, graphical test. Br Med J 315:629–634
Figueroa JD, Malats N, García-Closas M, Real FX, Silverman D, Kogevinas M, Chanock S, Welch R, Dosemeci M, Lan Q, Tardón A, Serra C, Carrato A, García-Closas R, Castaño-Vinyals G, Rothman N (2008) Bladder cancer risk and genetic variation in AKR1C3 and other metabolizing genes. Carcinogenesis 29:1955–1962
Fontana L, Delort L, Joumard L, Rabiau N, Bosviel R, Satih S, Guy L, Boiteux JP, Bignon YJ, Chamoux A, Bernard-Gallon DJ (2009) Genetic polymorphisms in CYP1A1, CYP1B1, COMT, GSTP1 and NAT2 genes and association with bladder cancer risk in a French cohort. Anticancer Res 29:1631–1635
Freedman ND, Ahn J, Hou L, Lissowska J, Zatonski W, Yeager M, Chanock SJ, Chow WH, Abnet CC (2009) Polymorphisms in estrogen- and androgen-metabolizing genes and the risk of gastric cancer. Carcinogenesis 30:71–77
Fukatsu T, Hirokawa Y, Araki T, Hioki T, Murata T, Suzuki H, Ichikawa T, Tsukino H, Qiu D, Katoh T, Sugimura Y, Yatani R, Shiraishi T, Watanabe M (2004) Genetic polymorphisms of hormone-related genes and prostate cancer risk in the Japanese population. Anticancer Res 24:2431–2437
Gallicchio L, Berndt SI, McSorley MA, Newschaffer CJ, Thuita LW, Argani P, Hoffman SC, Helzlsouer KJ (2006) Polymorphisms in estrogen-metabolizing and estrogen receptor genes and the risk of developing breast cancer among a cohort of women with benign breast disease. BMC Cancer 6:173
Gaudet MM, Chanock S, Lissowska J, Berndt SI, Yang XR, Peplonska B, Brinton LA, Welch R, Yeager M, Bardin-Mikolajczak A, Sherman ME, Sutter TR, Garcia-Closas M (2006) Genetic variation of Cytochrome P450 1B1 (CYP1B1) and risk of breast cancer among Polish women. Pharmacogenet Genomics 16:547–553
Goodman MT, McDuffie K, Kolonel LN, Terada K, Donlon TA, Wilkens LR, Guo C, Le Marchand L (2001) Case–control study of ovarian cancer and polymorphisms in genes involved in catecholestrogen formation and metabolism. Cancer Epidemiol Biomarkers Prev 10:209–216
Gulyaeva LF, Mikhailova ON, PustyInyak VO, Kim IV 4th, Gerasimov AV, Krasilnikov SE, Filipenko ML, Pechkovsky EV (2008) Comparative analysis of SNP in estrogen-metabolizing enzymes for ovarian, endometrial, and breast cancers in Novosibirsk. Russia Adv Exp Med Biol 617:359–366
Harth V, Schafer M, Abel J, Maintz L, Neuhaus T, Besuden M, Primke R, Wilkesmann A, Thier R, Vetter H, Ko YD, Bruning T, Bolt HM, Ickstadt K (2008) Head and neck squamous-cell cancer and its association with polymorphic enzymes of xenobiotic metabolism and repair. J Toxicol Environ Health A 71:887–897
Hayes CL, Spink DC, Spink BC, Cao JQ, Walker NJ et al (1996) 17-b estradiol hydroxylation catalyzed by human cytochrome P450 1B1. Proc Natl Acad Sci USA 93:9776–9781
Higgins JP, Thompson SG, Deeks JJ, Altman DG (2003) Measuring inconsistency in meta-analysis. Br Med J 327:557–560
Hirata H, Hinoda Y, Okayama N, Suehiro Y, Kawamoto K, Kikuno N, Rabban JT, Chen LM, Dahiya R (2008) CYP1A1, SULT1A1, and SULT1E1 polymorphisms are risk factors for endometrial cancer susceptibility. Cancer 112:1964–1973
Hlavata I, Vrana D, Smerhovsky Z, Pardini B, Naccarati A, Vodicka P, Novotny J, Mohelnikova-Duchonova B, Soucek P (2010) Association between exposure-relevant polymorphisms in CYP1B1, EPHX1, NQO1, GSTM1, GSTP1 and GSTT1 and risk of colorectal cancer in a Czech population. Oncol Rep 24:1347–1353
Holt SK, Rossing MA, Malone KE, Schwartz SM, Weiss NS, Chen C (2007) Ovarian cancer risk and polymorphisms involved in estrogen catabolism. Cancer Epidemiol Biomarkers Prev 16:481–489
Holt SK, EM Kwon, Fu R, Kolb S, Feng Z, Ostrander EA, Stanford JL (2013) Association of variants in estrogen-related pathway genes with prostate cancer risk. Prostate 73:1–10
Huang Y, Trentham-Dietz A, García-Closas M, Newcomb PA, Titus-Ernstoff L, Hampton JM, Chanock SJ, Haines JL, Egan KM (2009) Association of CYP1B1 haplotypes and breast cancer risk in Caucasian women. Cancer Epidemiol Biomarkers Prev 18:1321–1323
Huber A, Bentz EK, Schneeberger C, Huber JC, Hefler L, Tempfer C (2005) Ten polymorphisms of estrogen-metabolizing genes and a family history of colon cancer–an association study of multiple gene-gene interactions. J Soc Gynecol Investig 12:e51–e54
Hung RJ, Boffetta P, Brennan P, Malaveille C, Hautefeuille A, Donato F, Gelatti U, Spaliviero M, Placidi D, Carta A, Di Scotto Carlo A, Porru S (2004) GST, NAT, SULT1A1, CYP1B1 genetic polymorphisms, interactions with environmental exposures and bladder cancer risk in a high-risk population. Int J Cancer 110:598–604
Jang JH, Cotterchio M, Borgida A, Gallinger S, Cleary SP (2012) Genetic variants in carcinogen-metabolizing enzymes, cigarette smoking and pancreatic cancer risk. Carcinogenesis 33:818–827
Justenhoven C, Pierl CB, Haas S, Fischer HP, Baisch C, Hamann U, Harth V, Pesch B, Brüning T, Vollmert C, Illig T, Dippon J, Ko YD, Brauch H (2008) The CYP1B1_1358_GG genotype is associated with estrogen receptor-negative breast cancer. Breast Cancer Res Treat 111:171–177
Karim-Kos HE, De Vries E, Soerjomataram I, Lemmens V, Siesling S, Coebergh JW (2008) Recent trends of cancer in Europe: a combined approach of incidence, survival and mortality for 17 cancer sites since the 1990s. Eur J Cancer 44:1345–1389
Kato I, Cichon M, Yee CL, Land S, Korczak JF (2009) African American-preponderant single nucleotide polymorphisms (SNPs) and risk of breast cancer. Cancer Epidemiol 33:24–30
Kilfoy BA, Zheng T, Lan Q, Han X, Qin Q, Rothman N, Holford T, Zhang Y (2009) Genetic polymorphisms in glutathione S-transferases and cytochrome P450 s, tobacco smoking, and risk of non-Hodgkin lymphoma. Am J Hematol 84:279–282
Kim JH, Stansbury KH, Walker NJ, Trush MA, Strickland PT, Sutter TR (1998) Metabolism of benzo(a)pyrene and benzo(a)pyrene-7,8-diol by human cytochrome P450 1B1. Carcinogenesis 19:1847–1853
Klug SJ, Ressing M, Koenig J, Abba MC, Agorastos T, Brenna SM et al (2009) TP53 codon 72 polymorphism and cervical cancer: a pooled analysis of individual data from 49 studies. Lancet Oncol 10:772–784
Ko Y, Abel J, Harth V, Bröde P, Antony C, Donat S, Fischer HP, Ortiz-Pallardo ME, Thier R, Sachinidis A, Vetter H, Bolt HM, Herberhold C, Brüning T (2001) Association of CYP1B1 codon 432 mutant allele in head and neck squamous cell cancer is reflected by somatic mutations of p53 in tumor tissue. Cancer Res 61:4398–4404
Kocabaş NA, Sardaş S, Cholerton S, Daly AK, Karakaya AE (2002) Cytochrome P450 CYP1B1 and catechol O-methyltransferase (COMT) genetic polymorphisms and breast cancer susceptibility in a Turkish population. Arch Toxicol 76:643–649
Küry S, Buecher B, Robiou-du-Pont S, Scoul C, Sébille V, Colman H, Le Houérou C, Le Neel T, Bourdon J, Faroux R, Ollivry J, Lafraise B, Chupin LD, Bézieau S (2007) Combinations of cytochrome P450 gene polymorphisms enhancing the risk for sporadic colorectal cancer related to red meat consumption. Cancer Epidemiol Biomarkers Prev 16:1460–1467
Landi S, Gemignani F, Moreno V, Gioia-Patricola L, Chabrier A, Guino E, Navarro M, de Oca J, Capellà G, Canzian F; Bellvitge Colorectal Cancer Study Group (2005) A comprehensive analysis of phase I and phase II metabolism gene polymorphisms and risk of colorectal cancer. Pharmacogenet Genomics 15:535–546
Le Marchand L, Donlon T, Kolonel LN, Henderson BE, Wilkens LR (2005) Estrogen metabolism-related genes and breast cancer risk: the multiethnic cohort study. Cancer Epidemiol Biomarkers Prev 14:1998–2003
Lee KM, Abel J, Ko Y, Harth V, Park WY, Seo JS, Yoo KY, Choi JY, Shin A, Ahn SH, Noh DY, Hirvonen A, Kang D (2003) Genetic polymorphisms of cytochrome P450 19 and 1B1, alcohol use, and breast cancer risk in Korean women. Br J Cancer 88:675–678
Li D, Walcott FL, Chang P, Zhang W, Zhu J, Petrulis E, Singletary SE, Sahin AA, Bondy ML (2002) Genetic and environmental determinants on tissue response to in vitro carcinogen exposure and risk of breast cancer. Cancer Res 62:4566–4570
Li G, Liu Z, Sturgis EM, Chamberlain RM, Spitz MR, Wei Q (2005) CYP2E1 G1532C, NQO1 Pro187Ser, and CYP1B1 Val432Leu polymorphisms are not associated with risk of squamous cell carcinoma of the head and neck. Cancer Epidemiol Biomarkers Prev 14:1034–1036
Li Y, Tan SQ, Ma QH, Li L, Huang ZY, Wang Y, Li SW (2013) CYP1B1 C4326G polymorphism and susceptibility to cervical cancer in Chinese Han women. Tumour Biol 34:3561–3567
Liang G, Pu Y, Yin L (2005) Rapid detection of single nucleotide polymorphisms related with lung cancer susceptibility of Chinese population. Cancer Lett 223:265–274
Listgarten J, Damaraju S, Poulin B et al (2004) Predictive models for breast cancer susceptibility from multiple single nucleotide polymorphisms. Clin Cancer Res 10:2725–2737
Lundin E, Wirgin I, Lukanova A, Afanasyeva Y, Krogh V, Axelsson T, Hemminki K, Clendenen TV, Arslan AA, Ohlson N, Sieri S, Roy N, Koenig KL, Idahl A, Berrino F, Toniolo P, Hallmans G, Försti A, Muti P, Lenner P, Shore RE, Zeleniuch-Jacquotte A (2012) Selected polymorphisms in sex hormone-related genes, circulating sex hormones and risk of endometrial cancer. Cancer Epidemiol 36:445–452
Lutz JM, Francisci S, Mugno E, Usel M, Pompe-Kirn V, Coebergh JW (2003) Cancer prevalence in Central Europe: the EUROPREVAL Study. Ann Oncol 14:313–322
Mantel N, Haenszel W (1959) Statistical aspects of the analysis of data from retrospective studies of disease. Natl Cancer Inst 22:719–748
MARIE-GENICA Consortium on Genetic Susceptibility for Menopausal Hormone Therapy Related Breast Cancer Risk (2010) Genetic polymorphisms in phase I and phase II enzymes and breast cancer risk associated with menopausal hormone therapy in postmenopausal women. Breast Cancer Res Treat 119:463–474
Martin FL, Patel II, Sozeri O, Singh PB, Ragavan N et al (2010) Constitutive expression of bioactivating enzymes in normal human prostate suggests a capability to activate pro-carcinogens to DNA-damaging metabolites. Prostate 70:1586–1599
Martínez-Ramírez OC, Pérez-Morales R, Castro C, Flores-Díaz A, Soto-Cruz KE, Astorga-Ramos A, Gonsebatt ME, Casas L, Valdés-Flores M, Rubio J (2013) Polymorphisms of catechol estrogens metabolism pathway genes and breast cancer risk in Mexican women. Breast 22:335–343
Matyjasik J, Cybulski C, Masojć B, Jakubowska A, Serrano-Fernandez P, Górski B, Debniak T, Huzarski T, Byrski T, Gronwald J, Złowocka E, Narod SA, Scott R, Lubinski J (2007) CYP1B1 and predisposition to breast cancer in Poland. Breast Cancer Res Treat 106:383–388
Maurya SS, Anand G, Dhawan A, Khan AJ, Jain SK, Pant MC, Parmar D (2014) Polymorphisms in drug-metabolizing enzymes and risk to head and neck cancer: evidence for gene-gene and gene-environment interaction. Environ Mol Mutagen 55:134–144
McGrath M, Hankinson SE, Arbeitman L, Colditz GA, Hunter DJ, De Vivo I (2004) Cytochrome P450 1B1 and catechol-O-methyltransferase polymorphisms and endometrial cancer susceptibility. Carcinogenesis 25:559–565
Murray GI, Taylor MC, McFadyen MC, McKay JA, Greenlee WF, Burke MD, Melvin WT (1997) Tumor-specific expression of cytochrome P450 CYP1B1. Cancer Res 57:3026–3031
Okobia MN, Bunker CH, Garte SJ, Zmuda JM, Ezeome ER, Anyanwu SN, Uche EE, Osime U, Ojukwu J, Kuller LH, Ferrell RE, Taioli E (2009) Cytochrome P450 1B1 Val432Leu polymorphism and breast cancer risk in Nigerian women: a case–control study. Infect Agent Cancer 4(Suppl 1):S12
Ozbek YK, Oztürk T, Tüzüner BM, Calay Z, Ilvan S, Seyhan FM, Kisakesen HI, Oztürk O, Isbir T (2010) Combined effect of CYP1B1 codon 432 polymorphism and N-acetyltransferase 2 slow acetylator phenotypes in relation to breast cancer in the Turkish population. Anticancer Res 30:2885–2889
Rebbeck TR, Troxel AB, Wang Y, Walker AH, Panossian S, Gallagher S, Shatalova EG, Blanchard R, Bunin G, DeMichele A, Rubin SC, Baumgarten M, Berlin M, Schinnar R, Berlin JA, Strom BL (2006) Estrogen sulfation genes, hormone replacement therapy, and endometrial cancer risk. J Natl Cancer Inst 98:1311–1320
Reding KW, Weiss NS, Chen C, Li CI, Carlson CS, Wilkerson HW, Farin FM, Thummel KE, Daling JR, Malone KE (2009) Genetic polymorphisms in the catechol estrogen metabolism pathway and breast cancer risk. Cancer Epidemiol Biomarkers Prev 18:1461–1467
Reding KW, Chen C, Lowe K, Doody DR, Carlson CS, Chen CT, Houck J, Weiss LK, Marchbanks PA, Bernstein L, Spirtas R, McDonald JA, Strom BL, Burkman RT, Simon MS, Liff JM, Daling JR, Malone KE (2012) Estrogen-related genes and their contribution to racial differences in breast cancer risk. Cancer Causes Control 23:671–681
Rodrigues IS, Kuasne H, Losi-Guembarovski R, Fuganti PE, Gregório EP, Kishima MO, Ito K, de Freitas Rodrigues MA, de Syllos Cólus IM (2011) Evaluation of the influence of polymorphic variants CYP1A1 2B, CYP1B1 2, CYP3A4 1B, GSTM1 0, and GSTT1 0 in prostate cancer. Urol Oncol 29:654–663
Rotunno M, Yu K, Lubin JH, Consonni D, Pesatori AC, Goldstein AM, Goldin LR, Wacholder S, Welch R, Burdette L, Chanock SJ, Bertazzi PA, Tucker MA, Caporaso NE, Chatterjee N, Bergen AW, Landi MT (2009) Phase I metabolic genes and risk of lung cancer: multiple polymorphisms and mRNA expression. PLoS One 4:e5652
Rudolph A, Sainz J, Hein R, Hoffmeister M, Frank B, Försti A, Brenner H, Hemminki K, Chang-Claude J (2011) Modification of menopausal hormone therapy-associated colorectal cancer risk by polymorphisms in sex steroid signaling, metabolism and transport related genes. Endocr Relat Cancer 18:371–384
Rylander-Rudqvist T, Wedren S, Granath F, Humphreys K, Ahlberg S, Weiderpass E, Oscarson M, Ingelman-Sundberg M, Persson I (2003) Cytochrome P450 1B1 gene polymorphisms and postmenopausal breast cancer risk. Carcinogenesis 24:1533–1539
Rylander-Rudqvist T, Wedrén S, Jonasdottir G, Ahlberg S, Weiderpass E, Persson I, Ingelman-Sundberg M (2004) Cytochrome P450 1B1 gene polymorphisms and postmenopausal endometrial cancer risk. Cancer Epidemiol Biomarkers Prev 13:1515–1520
Sachse C, Smith G, Wilkie MJ, Barrett JH, Waxman R, Sullivan F, Forman D, Bishop DT, Wolf CR, Colorectal Cancer Study Group (2002) A pharmacogenetic study to investigate the role of dietary carcinogens in the etiology of colorectal cancer. Carcinogenesis 23(11):1839–1849
Sainz J, Rudolph A, Hein R, Hoffmeister M, Buch S, von Schönfels W, Hampe J, Schafmayer C, Völzke H, Frank B, Brenner H, Försti A, Hemminki K, Chang-Claude J (2011) Association of genetic polymorphisms in ESR2, HSD17B1, ABCB1, and SHBG genes with colorectal cancer risk. Endocr Relat Cancer 18:265–276
Salinas-Sánchez AS, Sánchez-Sánchez F, Donate-Moreno MJ, Rubio-Del-Campo A, Serrano-Oviedo L, Gimenez-Bachs JM, Martínez-Sanchiz C, Segura-Martín M, Escribano J (2012a) GSTT1, GSTM1, and CYP1B1 gene polymorphisms and susceptibility to sporadic renal cell cancer. Urol Oncol 30:864–870
Salinas-Sánchez AS, Donate-Moreno MJ, López-Garrido MP, Giménez-Bachs JM, Escribano J (2012b) Role of CYP1B1 gene polymorphisms in bladder cancer susceptibility. J Urol 187:700–706
Sangrajrang S, Sato Y, Sakamoto H, Ohnami S, Laird NM, Khuhaprema T, Brennan P, Boffetta P, Yoshida T (2009) Genetic polymorphisms of estrogen metabolizing enzyme and breast cancer risk in Thai women. Int J Cancer 125:837–843
Sasaki M, Tanaka Y, Kaneuchi M, Sakuragi N, Dahiya R (2003) CYP1B1 gene polymorphisms have higher risk for endometrial cancer, and positive correlations with estrogen receptor alpha and estrogen receptor beta expressions. Cancer Res 63:3913–3918
Sasaki M, Tanaka Y, Okino ST, Nomoto M, Yonezawa S, Nakagawa M, Fujimoto S, Sakuragi N, Dahiya R (2004) Polymorphisms of the CYP1B1 gene as risk factors for human renal cell cancer. Clin Cancer Res 10:2015–2019
Sellers TA, Schildkraut JM, Pankratz VS, Vierkant RA, Fredericksen ZS, Olson JE, Cunningham J, Taylor W, Liebow M, McPherson C, Hartmann LC, Pal T, Adjei AA (2005) Estrogen bioactivation, genetic polymorphisms, and ovarian cancer. Cancer Epidemiol Biomarkers Prev 14:2536–2543
Shah PP, Singh AP, Singh M, Mathur N, Mishra BN, Pant MC, Parmar D (2008) Association of functionally important polymorphisms in cytochrome P4501B1 with lung cancer. Mutat Res 643:4–10
Shimada T, Hayes CL, Yamazaki H, Amin S, Hecht SS, Guengerich FP, Sutter TR (1996) Activation of chemically diverse procarcinogens by human cytochrome P-450 1B1. Cancer Res 56:2979–2984
Shimada N, Iwasaki M, Kasuga Y, Yokoyama S, Onuma H, Nishimura H, Kusama R, Hamada GS, Nishimoto IN, Iyeyasu H, Motola J Jr, Laginha FM, Kurahashi N, Tsugane S (2009) Genetic polymorphisms in estrogen metabolism and breast cancer risk in case-control studies in Japanese, Japanese Brazilians and non-Japanese Brazilians. J Hum Genet 54:209–215
Siegel R, Naishadham D, Jemal A (2013) Cancer statistics, 2013. CA Cancer J Clin 63:11–30
Sigurdson AJ, Bhatti P, Chang SC, Rajaraman P, Doody MM, Bowen L, Simon SL, Weinstock RM, Linet MS, Rosenstein M, Stovall M, Alexander BH, Preston DL, Struewing JP (2009) Polymorphisms in estrogen biosynthesis and metabolism-related genes, ionizing radiation exposure, and risk of breast cancer among US radiologic technologists. Breast Cancer Res Treat 118:177–184
Sillanpää P, Heikinheimo L, Kataja V, Eskelinen M, Kosma VM, Uusitupa M, Vainio H, Metsola K, Hirvonen A (2007) CYP1A1 and CYP1B1 genetic polymorphisms, smoking and breast cancer risk in a Finnish Caucasian population. Breast Cancer Res Treat 104:287–297
Singh AP, Shah PP, Mathur N, Buters JT, Pant MC, Parmar D (2008) Genetic polymorphisms in cytochrome P4501B1 and susceptibility to head and neck cancer. Mutat Res 639:11–19
Sissung TM, Danesi R, Price DK, Steinberg SM, de Wit R et al (2008) Association of the CYP1B1*3 allele with survival in patients with prostate cancer receiving docetaxel. Mol Cancer Ther 7:19–26
Sliwinski T, Sitarek P, Stetkiewicz T, Sobczuk A, Blasiak J (2010) Polymorphism of the ERalpha and CYP1B1 genes in endometrial cancer in a Polish subpopulation. J Obstet Gynaecol Res 36:311–317
Sobti RC, Onsory K, Al-Badran AI, Kaur P, Watanabe M, Krishan A, Mohan H (2006) CYP17, SRD5A2, CYP1B1, and CYP2D6 gene polymorphisms with prostate cancer risk in North Indian population. DNA Cell Biol 25:287–294
Soerjomataram I, de Vries E, Pukkala E, Coebergh JW (2007) Excess of cancers in Europe: a study of eleven major cancers amenable to lifestyle change. Int J Cancer 120:1336–1343
Sørensen M, Autrup H, Tjønneland A, Overvad K, Raaschou-Nielsen O (2005) Genetic polymorphisms CYP1B1, GSTA1, NQO1 and NAT2 and the risk of lung cancer. Cancer Lett 221:185–190
Soucek P, Susova S, Mohelnikova-Duchonova B, Gromadzinska J, Moraviec-Sztandera A, Vodicka P, Vodickova L (2010) Polymorphisms in metabolizing enzymes and the risk of head and neck squamous cell carcinoma in the Slavic population of the central Europe. Neoplasma 57:415–421
Spink DC, Spink BC, Cao JQ, Gierthy JF, Hayes CL et al (1997) Induction of cytochrome P450 1B1 and catechol estrogen metabolism in ACHN human renal adenocarcinoma cells. J Steroid Biochem Mol Biol 62:223–232
Stoilov I, Akarsu AN, Alozie I, Child A, Barsoum-Homsy M, Turacli ME, Or M, Lewis RA, Ozdemir N, Brice G, Aktan SG, Chevrette L, Coca-Prados M, Sarfarazi M (1998) Sequence analysis and homology modeling suggest that primary congenital glaucoma on 2p21 results from mutations disrupting either the hinge region or the conserved core structures of Cytochrome P4501B1. Am J Hum Genet 62:573–584
Sutter TR, Tang YM, Hayes CL, Wo YY, Jabs EW et al (1994) Complete cDNA sequence of a human dioxin-inducible Mrna identifies a new gene subfamily of cytochrome P450 that maps to chromosome 2. J Biol Chem 269:13092–13099
Tai J, Yang M, Ni X, Yu D, Fang J, Tan W, Huang Z, Wu C, Chen X, Wang G, Zhou W, Chen X, Zhang W, Ma L, Lin D, Han D (2010) Genetic polymorphisms in cytochrome P450 genes are associated with an increased risk of squamous cell carcinoma of the larynx and hypopharynx in a Chinese population. Cancer Genet Cytogenet 196:76–82
Tanaka Y, Sasaki M, Kaneuchi M, Shiina H, Igawa M, Dahiya R (2002) Polymorphisms of the CYP1B1 gene have higher risk for prostate cancer. Biochem Biophys Res Commun 296:820–826
Tang YM, Green BL, Chen GF, Thompson PA, Lang NP, Shinde A, Lin DX, Tan W, Lyn-Cook BD, Hammons GJ, Kadlubar FF (2000) Human CYP1B1 Leu432Val gene polymorphism: ethnic distribution in African-Americans, Caucasians and Chinese; oestradiol hydroxylase activity; and distribution in prostate cancer cases and controls. Pharmacogenetics 10:761–766
Tao MH, Cai Q, Xu WH, Kataoka N, Wen W, Zheng W, Xiang YB, Zhang ZF, Shu XO (2006) Cytochrome P450 1B1 and catechol-O-methyltransferase genetic polymorphisms and endometrial cancer risk in Chinese women. Cancer Epidemiol Biomarkers Prev 15:2570–2573
Thyagarajan B, Brott M, Mink P, Folsom AR, Anderson KE, Oetting WS, Gross M (2004) CYP1B1 and CYP19 gene polymorphisms and breast cancer incidence: no association in the ARIC study. Cancer Lett 207:183–189
Timofeeva MN, Kropp S, Sauter W, Beckmann L, Rosenberger A, Illig T, Jager B, Mittelstrass K, Dienemann H, Bartsch H, Bickeboller H, Chang-Claude JC, Risch A, Wichmann HE, LUCY-Consortium (2009) CYP450 polymorphisms as risk factors for early-onset lung cancer: gender-specific differences. Carcinogenesis 30:1161–1169
Trubicka J, Grabowska-Kłujszo E, Suchy J, Masojć B, Serrano-Fernandez P, Kurzawski G, Cybulski C, Górski B, Huzarski T, Byrski T, Gronwald J, Złowocka E, Kładny J, Banaszkiewicz Z, Wiśniowski R, Kowalska E, Lubinski J, Scott RJ (2010) Variant alleles of the CYP1B1 gene are associated with colorectal cancer susceptibility. BMC Cancer 10:420
Van Emburgh BO, Hu JJ, Levine EA, Mosley LJ, Perrier ND, Freimanis RI, Allen GO, Rubin P, Sherrill GB, Shaw CS, Carey LA, Sawyer LR, Miller MS (2008) Polymorphisms in CYP1B1, GSTM1, GSTT1 and GSTP1, and susceptibility to breast cancer. Oncol Rep 19:1311–1321
Vrana D, Novotny J, Holcatova I, Hlavata I, Soucek P (2010) CYP1B1 gene polymorphism modifies pancreatic cancer risk but not survival. Neoplasma 57:15–19
Wang Q, Li H, Tao P, Wang YP, Yuan P, Yang CX, Li JY, Yang F, Lee H, Huang Y (2011a) Soy isoflavones, CYP1A1, CYP1B1, and COMT polymorphisms, and breast cancer: a case–control study in southwestern China. DNA Cell Biol 30:585–595
Wang F, Zou YF, Sun GP, Su H, Huang F (2011b) Association of CYP1B1 gene polymorphisms with susceptibility to endometrial cancer: a meta-analysis. Eur J Cancer Prev 20:112–120
Wang J, Joshi AD, Corral R, Siegmund KD, Marchand LL, Martinez ME, Haile RW, Ahnen DJ, Sandler RS, Lance P, Stern MC (2012) Carcinogen metabolism genes, red meat and poultry intake, and colorectal cancer risk. Int J Cancer 130:1898–1907
Watanabe J, Shimada T, Gillam EM, Ikuta T, Suemasu K, Higashi Y, Gotoh O, Kawajiri K (2000) Association of CYP1B1 genetic polymorphism with incidence to breast and lung cancer. Pharmacogenetics 10:25–33
Wen W, Cai Q, Shu XO, Cheng JR, Parl F, Pierce L, Gao YT, Zheng W (2005) Cytochrome P450 1B1 and catechol-O-methyltransferase genetic polymorphisms and breast cancer risk in Chinese women: results from the shanghai breast cancer study and a meta-analysis. Cancer Epidemiol Biomarkers Prev 14:329–335
Wenzlaff AS, Cote ML, Bock CH, Land SJ, Santer SK, Schwartz DR, Schwartz AG (2005) CYP1A1 and CYP1B1 polymorphisms and risk of lung cancer among never smokers: a population-based study. Carcinogenesis 26:2207–2212
Williams JA, Martin FL, Muir GH, Hewer A, Grover PL, Phillips DH (2000) Metabolic activation of carcinogens and expression of various cytochromes P450 in human prostate tissue. Carcinogenesis 21:1683–1689
Wu MF, Wu WJ, Chang GC, Chen CY, Hu SW, Tsai WT, Lee H, Lin P (2004) Increased expression of cytochrome P4501B1 in peripheral leukocytes from lung cancer patients. Toxicol Lett 150:211–219
Xu W, Zhou Y, Hang X, Shen D (2012) Current evidence on the relationship between CYP1B1 polymorphisms and lung cancer risk: a meta-analysis. Mol Biol Rep 39:2821–2829
Yadav S, Singhal NK, Singh V, Rastogi N, Srivastava PK, Singh MP (2009) Association of single nucleotide polymorphisms in CYP1B1 and COMT genes with breast cancer susceptibility in Indian women. Dis Markers 27:203–210
Yang J, Xu DL, Lu Q, Han ZJ, Tao J, Lu P, Wang C, Di XK, Gu M (2012) Prostate cancer risk and aggressiveness associated with the CYP1B1 4326C/G (Leu432Val) polymorphism: a meta-analysis of 2788 cases and 2968 controls. Asian J Androl 14:560–565
Yoon KA, Kim JH, Gil HJ, Hwang H, Hwangbo B, Lee JS (2008) CYP1B1, CYP1A1, MPO, and GSTP1 polymorphisms and lung cancer risk in never-smoking Korean women. Lung Cancer 60:40–46
Yuan X, Zhou G, Zhai Y, Xie W, Cui Y, Cao J, Zhi L, Zhang H, Yang H, Zhang X, Qiu W, Peng Y, Zhang X, Yu L, Xia X, He F (2008) Lack of association between the functional polymorphisms in the estrogen-metabolizing genes and risk for hepatocellular carcinoma. Cancer Epidemiol Biomarkers Prev 17:3621–3627
Zheng W, Xie DW, Jin F, Cheng JR, Dai Q, Wen WQ, Shu XO, Gao YT (2000) Genetic polymorphism of cytochrome P450-1B1 and risk of breast cancer. Cancer Epidemiol Biomarkers Prev 9:147–150
Zhu ZY, Mu YQ, Fu XM, Li SM, Zhao FX (2011) Association of CYP1B1 gene polymorphisms and the positive expression of estrogen alpha and estrogen beta with endometrial cancer risk. Eur J Gynaecol Oncol 32:188–191
Zienolddiny S, Campa D, Lind H, Ryberg D, Skaug V, Stangeland LB, Canzian F, Haugen A (2008) A comprehensive analysis of phase I and phase II metabolism gene polymorphisms and risk of non-small cell lung cancer in smokers. Carcinogenesis 29:1164–1169
Zimarina TC, Kristensen VN, Imianitov EN, Bershteĭn LM (2004) Polymorphisms of CYP1B1 and COMT in breast and endometrial cancer. Mol Biol (Mosk) 38:386–393
Conflict of interest
None.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by S. Hohmann.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Liu, JY., Yang, Y., Liu, ZZ. et al. Association between the CYP1B1 polymorphisms and risk of cancer: a meta-analysis. Mol Genet Genomics 290, 739–765 (2015). https://doi.org/10.1007/s00438-014-0946-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00438-014-0946-x