Abstract
Hrs1/Med3, a component of the Mediator involved in transcription initiation, was previously isolated as a suppressor of hpr1Δ hyper-recombination linked to transcription elongation. Here we show that hrs1Δ-mediated suppression is specific of transcription-associated hyper-recombination (TAR). The decrease in recombination associated with hrs1Δ, either in wild-type or hpr1Δ cells is only observed in DNA repeats constructs in which transcription is Hrs1-dependent. We propose that the suppression of THO mutants by hrs1Δ is due to the specific effect of hrs1Δ on transcription initiation of the recombination system. In parallel we show that the higher the transcription of a gene the more important becomes the THO complex for its expression, implying that the in vivo relevance of this complex is dependent on the frequency of RNAPII transcription initiation. This study furthers the understanding of the importance of THO in transcription and the maintenance of genome stability.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The Mediator of RNA polymerase II (RNAPII) is an essential factor in the regulation of transcription initiation, and is conserved from yeast to humans (Boube et al. 2002; Reeves and Hahn 2005). It is composed of ~20 proteins grouped in different subcomplexes with a putative specialized function each. One of these proteins is Hrs1/Med3, which was isolated as a suppressor of the hyper-recombination phenotype of hpr1Δ (Santos-Rosa and Aguilera 1995). Hpr1 is part of THO/TREX, a conserved eukaryotic complex first identified in yeast (Chavez et al. 2000; Strasser et al. 2002), which functions at the interface between mRNP formation and RNA export during transcription elongation (Aguilera 2005). THO mutations were identified by their strong hyper-recombination phenotype, which is linked to defective transcription elongation of GC-rich and long DNA sequences (Chavez et al. 2001). Different studies have led to the proposal that THO functions during transcription elongation, consistent with the fact that THO immunoprecipitates with transcribed chromatin (Strasser et al. 2002; Kim et al. 2004), and their null mutations impair transcription elongation both in vivo and in vitro (Mason and Struhl 2005; Rondon et al. 2003). The physical and functional interaction of THO with RNA export factors have contributed to the present view that transcription and RNA export are coupled, the THO/TREX complex being a key player in this interconnection. Genome-wide analysis of Nuclear Pore Complex-tethered loci showed an association of highly transcribed genes with the nuclear periphery. This is the case of Saccharomyces cerevisiae INO1 gene and genes induced by galactose or α-factor; they move towards the nuclear periphery after transcriptional activation (Brickner and Walter 2004; Casolari et al. 2004, 2005; Cabal et al. 2006).
The synthetic lethal phenotypes of THO mutants with mutations in transcription elongation factors such as spt4, exosome components such as rrp6, transcription termination/3′-end processing factors, such as rna14 and rna15, or RNA export factors such as mex67 or yra1, and also RNAPII recruitment determined by ChIP analysis (Jimeno et al. 2002; Strasser et al. 2002; Zenklusen et al. 2002; Kim et al. 2004; Luna et al. 2005) support the view that THO functions during elongation and affects later steps of transcription.
Interestingly, different screenings for suppressors of hpr1 have identified several components of the yeast Mediator. One such component is Hrs1/Med3, whose null mutation suppresses the hyper-recombination phenotype of hpr1Δ (Santos-Rosa and Aguilera 1995; Piruat and Aguilera 1996; Santos-Rosa et al. 1996). hrs1Δ also reduces recombination between direct-repeats in wild-type cells (Santos-Rosa et al. 1996). These findings raise the question of how a mutation in a transcription-initiation factor can suppress a phenotype linked to a transcription elongation defect. In order to decipher the molecular basis for these phenotypes, we analysed the effect of hrs1Δ on recombination and found that the decrease in recombination caused by hrs1Δ, both in wild type and in THO mutants, is specific to transcription-associated recombination (TAR) and is only observed in recombination systems in which transcription is dependent on Hrs1. Importantly, we show that Hpr1 becomes critical for transcription driven from a strong promoter but of little importance for transcription driven from weak promoters, implying that the in vivo relevance of THO in the expression of one gene depends on the strength of transcription. Altogether, our analysis shows that suppression of TAR by hrs1Δ/med3Δ is caused by the abolishment of transcription initiation and enhances the understanding of in vivo relevance of THO in transcription, this being dependent on promoter strength.
Materials and methods
Strains
Strains used in this study are listed in Table 1.
Plasmids
Plasmids pRS416GY (pRS416 containing the LYS2 gene under the control of the Gal1 promoter) and pCM184-tet-LYS (pCM184 containing the LYS2 gene under the control of the tet promoter) were generated in this study. Plasmids pRS314LY, pRS314LYΔNS, pRS314LU, pRS314LNA, pRS314LNB, pRS314L-lacZ, pRS314GL-lacZ and pLAUR, used for the studies of recombination and transcription have been previously published (Prado et al. 1997; Jimeno et al. 2002).
Recombination analyses
Recombination frequency was obtained from three different transformants for each genotype. It was determined as the average of 3 median frequencies of 6–12 independent values for each transformant, as previously reported (Santos-Rosa and Aguilera 1994).
Gene expression analyses
Northern analyses were performed according to standard procedures. 32P-labelled DNA probes used in Northern assays were as follows: lacZ: 3-kb BamHI fragment of pPZ plasmid (Straka and Horz 1991); LEU2: 0.6-kb ClaI EcoRV fragment of pRS415 plasmid; pBR322: pBR322 linearized with EcoRI; 5′ADE2: 1.2-kb HindIII fragment of pHSR1 (ADE2 cloned into BamHI EcoRV of pRS306 plasmid (Sikorski and Hieter 1989); 3′ADE2: 0.5-kb EcoRI-HindIII fragment of pHSR1 plasmid; URA3: 0.9-kb PstI-SmaI fragment of the pRS315-U plasmid (pRS315 with a URA3 HindIII–PstI fragment); YOR129c: 0.55-kb BamHI–HindIII fragment of pHSR1; LYS2: 1.4-kb BstEII fragment of pRS416GY, and the 589-bp internal 25S rDNA fragment obtained from genomic DNA by PCR.
Results
hrs1Δ/med3Δ only suppresses transcription-dependent hyper-recombination mutants
We have previously shown that the absence of Hrs1/Med3 suppresses the transcription-associated hyper-recombination phenotype of hpr1Δ, tho2Δ and thp1Δ mutants in the leuk-2k::ADE2-URA3::leu2-k chromosomal recombination construct, based on two 2.16-kb leu2-k direct repeats (Lk-AU system) (Santos-Rosa and Aguilera 1995; Piruat and Aguilera 1998; Gallardo and Aguilera 2001). We then wondered whether such a suppression was specific to hyper-recombination caused by mutations in the THO and Thp1–Sac3 complexes, or if it could also be observed in other hyper-recombination mutants whose effect was not related to transcription. Consequently, we analysed the effect of hrs1Δ on the hyper-recombination phenotypes of DNA polymerase III (cdc2) and DNA ligase (cdc9) mutants that are presumably caused by their intrinsic defects in DNA replication (Montelone et al. 1981; Kokoska et al. 2000). Recombination was assayed in the Lk-AU system. As can be seen in Fig. 1, cdc2 and cdc9 mutations caused an 800- and 32-fold increase above wild-type levels in the frequency of recombinants, respectively. However, hyper-recombination of neither mutant was suppressed by the strong hrs1-1 mutation, suggesting that the suppression effect of hrs1 is not general for any type of hyper-recombination, but presumably specific for hyper-recombination linked to transcription.
Recombination analyses of cdc2, cdc2 hrs1-1, cdc9 and cdc9 hrs1-1 mutants. Recombination frequencies of cdc2 (MC2-7B), cdc9 (MC9-1D), cdc2 hrs1-1 (MC2-7D), cdc9 hrs1-1 (MC9-8B) and wild type (AYW3-1B) congenic strains carrying the Lk-AU system are shown. A scheme of the system is shown at the top. For recombination analyses, independent colonies were obtained from SC and recombinants were selected in SC medium containing FOA
hrs1 only suppresses hyper-recombination linked to transcription
We next studied the ability of hrs1Δ to suppress hyper-recombination of hpr1Δ in systems such as L-lacZ and LY, both based on two 0.6-kb LEU2 truncated repeats located in CEN (centromeric) plasmids (Chavez et al. 2001). In both systems transcription is driven from the LEU2 promoter and the RNAPII traverses through a long and high G–C DNA fragment as a result of which a severe hyper-recombination phenotype is observed in hpr1Δ mutants. Unexpectedly, hrs1Δ did not suppress the hpr1Δ hyper-recombination phenotype in these systems (Fig. 2a). A slight reduction of recombination in the double mutant hpr1Δ hrs1Δ in the LY system has been reported in a previous study (Santos-Rosa and Aguilera 1995), but we found that this was due to a diminished capacity of the hpr1Δ hrs1Δ transformants to form colonies in the selective minimal medium in which recombinants were scored, independent of recombination (data not shown). Instead, hyper-recombination in the Lk-AU system was completely suppressed. Therefore, suppression of hpr1Δ hyper-recombination seems to be specific of recombination constructs.
Recombination and transcription analyses of hrsΔ, hpr1Δ and hpr1Δ hrs1Δ mutants. a Recombination frequencies of hpr1Δ (UKH-1A), hrs1Δ (UKH-1C), hpr1Δ hrs1Δ (UKH-1B) and wild type (UKH-1D) isogenic strains transformed with L-lacZ and LY systems are shown. The schemes of the systems are shown at the top. Independent colonies of three different transformants were obtained from SC-trp and recombinants were selected in SC-leu-trp. b Effect of hrs1Δ mutant on transcription. Northern analyses of L-lacZ and LY mRNAs driven from the LEU2 promoter in hrs1Δ (UKH-1C) and the isogenic wild type (UKH-1D) are shown. The LEU2, lacZ and rDNA probes used are detailed in “Materials and methods”. RNA levels in arbitrary units (A.U.) were obtained in a Fuji FLA 3000 and were normalized with respect to rRNA levels of each sample
It is important to notice that the specificity of the reduction of TAR by hrs1Δ mutation is not exclusive of THO mutants, but it is also observed in wild-type cells. Whereas hrs1Δ reduces the recombination frequency in the Lk-AU system (Santos-Rosa et al. 1996), this is not the case for the LY and L-lacZ systems, in which we did not observe such a reduction (Fig. 2a) Thus, we can conclude that the effect of hrs1 on recombination is general to TAR.
Suppression of TAR by hrs1 is linked to its capacity to reduce transcription initiation
To understand the molecular basis of the difference between the recombination systems in which suppression was observed from those in which it was not, we analysed the effect of hrs1Δ on transcription of the different recombination systems used, as can be seen in Fig. 2b, hrs1Δ was not defective in transcription of either of the LY or L-lacZ systems, in which hyper-recombination was not suppressed. Both wild type and hrs1Δ strains accumulated the same RNA levels. Therefore, since the hpr1Δ hyper-recombination phenotype has only been observed with high levels of transcription, it was possible that the hrs1Δ-mediated suppression of hpr1Δ associated hyper-recombination as well as TAR reduction in the wild-type background could only be seen in recombination systems whose transcription was dependent on Hrs1/Med3. Therefore, we expected that at least some transcripts of the Lk-AU system should be dependent on Hrs1. To test this possibility, we carried out a detailed Northern study of this system. In Fig. 3a we show a scheme of the Lk-AU system, the probes used (marked as lines), and the transcripts obtained (marked as arrows).
Expression analysis of the Lk-AU system in hrs1Δ mutant. a Scheme of 32P-labelled DNA probes used in the study (detailed in “Materials and methods”). b Northern analysis of hrs1Δ (SSAA-12B) and wild type (SSAA-8B) congenic strains carrying the Lk-AU system. Hrs1-dependent transcripts are marked with an asterisk. Probes used are indicated by letters (a–d). RNAs produced by the Lk-AU system are identified by numbers (1–4). E Indicates The RNA of the YOR129c endogenous gene. Other details as in Fig. 2
We first observed that transcription of leu2-k and URA3 was not affected in hrs1Δ mutants (data not shown). However, as can be seen in Fig. 3b, when we hybridized with a probe containing the ADE2 region (probe a) we obtained two bands: a 1.7-kb band (transcript 1) that corresponds to the ADE2 allele and a 3-kb band (transcript 2) present in the wild type and disappeared in hrs1Δ. This latter band corresponded to the fragment we were looking for. To confirm the identity of this band, we carried out a more detailed analysis of transcripts using probes against more specific regions. These were: 5′ ADE2 (probe b), 3′ ADE2 (probe c), 3′ YOR129c (probe d) and pBR322. With probe b (5′ ADE2) we only obtained a 1.7-kb band (transcript 1) that corresponds to the ADE2 region. With probe c (3′ ADE2) we obtained three bands: a 1.7-kb band (transcript 1) that corresponds to ADE2, a 3-kb band (transcript 2) (which was the same hybridizing with probe a), plus a smaller 1.5-kb band (transcript 3) that presumably corresponds to the 3′ end of ADE2 as it does not hybridize with probe b. Transcripts 2 and 3 were Hrs1-dependent. With probe d (YOR129c) we obtained two bands: a 2.8-kb band that corresponds to the endogenous YOR129 gene (transcript E) and a 0.5-kb band that corresponds to the small fragment of this gene present in the Lk-AU system (transcript 4). Hybridization with pBR322 led to a 3-kb band (transcript 2) plus a smaller 1.5-kb band (transcript 3), which are the same obtained with probe c (3′ ADE2) and indeed are Hrs1-dependent. Therefore, we can conclude that there are two Hrs1-dependent transcripts produced by the Lk-AU system. These correspond to the 3′ end of the ADE2 and to pBR322. These data suggest that hyper-recombination in the Lk-AU system is dependent on pBR322-based transcripts, and that the abolishment of transcription through pBR322 by hrs1Δ is the cause of the suppression of hyper-recombination.
Confirmation of this hypothesis required the use of recombination assays lacking the pBR322 region covering transcripts 2 and 3. Since these pBR322 sequences in Lk-AU could only be manipulated in the yeast chromosome and because it implied removal of the unique bacterial ori required for propagation in E. coli, it could not be modified directly without introducing selectable markers that would affect the transcription patterns. For this reason we used the plasmid-borne recombination systems LY, LYΔNS, LU and LA containing also two leu2 repeats flanking different regions of Lk-AU, including pBR322 (Fig. 4a). In contrast to Lk-AU, in these constructs the pBR322 transcripts are originated at the external LEU2 promoter which is independent of Hrs1, as it occurs in the LY system (see Fig. 2b; Prado et al. 1997).
Correlation between recombination and expression of pBR322 in different recombination constructs. a Schematic representation of the LY, LYΔNS, LU and LA plasmid-based recombination systems compared with Lk-AU. The fold-increase of the frequency of recombination of hpr1Δ above wild type of each system is shown (data taken from Prado et al. 1997). b Northern analysis of the LY, LYΔNS and LU plasmid-based constructs in wild-type cells using pBR322 as probe. c Northern analysis of LYΔNS system in wild type and hrs1Δ cells using pBR322 as probe. Median frequencies of recombination of hpr1Δ (UKH-1A) and hpr1Δ hrs1Δ (UKH-1B) isogenic strains are shown. pBR322 and rDNA probes used are detailed in “Materials and methods”. Other details as in Fig. 2
As previously reported, whereas the LY and LYΔNS systems, in which the pBR322 sequences corresponding to Lk-AU transcripts 2 and 3 are actively transcribed, are hyper-recombinant in hpr1 strains, this is not the case for LU and LA, which contain, respectively, the URA3 and ADE2 regions present in Lk-AU but not the pBR322 sequences (Fig. 4a, b). This confirms that, as it happens in Lk-AU, transcription of the region of pBR322 covering transcripts 2 and 3 is needed for hpr1Δ hyper-recombination to take place in these systems. hpr1Δ hyper-recombination is only suppressed when such transcripts are removed either by abolishing its transcription in Lk-AU with the hrs1Δ mutation, or by physically removing the DNA sequences, as in LU and LA. Consistently, and as shown with LY (Fig. 2a), hrs1Δ does not suppress hpr1Δ hyper-recombination of the LYΔNS system in agreement with the incapacity of hrs1Δ to abolish transcription through pBR322 that is initiated at the Hrs1-independent LEU2 promoter (Fig. 4c).
Hyper-recombination in hpr1Δ depends on transcription levels
The previous results suggest that suppression of TAR is due to the reduction of transcription caused by hrs1 in the systems tested, which is consistent with the fact that a modulation of transcription affects the levels of recombination in hpr1Δ strains. Thus, as we have previously reported, the increase in recombination of the leu2 repeats under a repressed GAL1 promoter (GL-lacZ system in glucose; low transcription) is much lower (sevenfold above wild type) than under the LEU2 promoter (L-lacZ; medium transcription) (86-fold) (Huertas et al. 2006). However, as can be seen in Fig. 5, the increase in recombination became much higher (224-fold) when transcription from the GAL1 promoter was induced with galactose (GL-lacZ system in galactose; high transcription).
Effect of the level of transcription on the hpr1Δ hyper-recombination phenotype. The increases in recombination of hpr1Δ (UKH-1A) above the isogenic wild type strain (UKH-1D) transformed with L-lacZ or GL-lacZ systems are shown. The data of recombination of the GL-lacZ system in glucose (transcription off; low transcription) and in galactose (transcription on; high transcription) are shown (low and medium transcription data are taken from Huertas et al. 2006). Other details as in Fig. 1
The levels of transcription impairment in hpr1Δ are dependent on the rate of transcription initiation
Another hallmark phenotype of mutants of the THO complex is their gene expression defect. hpr1Δ mutants cannot properly express long and high-GC content genes that are driven from strong promoters or genes containing direct repeats (Chavez et al. 2001; Voynov et al. 2006). This is clearly observed in the pLAUR gene expression system containing a lacZ–URA3 translational fusion under the control of the tet promoter. Whereas wild-type cells express this fusion properly, this is not the case for THO mutants. Thus, hpr1Δ cells carrying the pLAUR system are unable to form colonies on SC-ura and do not show β-galactosidase activity (Jimeno et al. 2002). We wanted to know whether the hrs1-mediated hyper-recombination suppression could be due to the suppression of the gene expression defect. We, therefore, tested whether hrs1Δ suppressed hpr1Δ gene expression defect of pLAUR. As can be seen in Fig. 6, hrs1Δ could not suppress hpr1Δ transcription defect as measured either by its ability to grow on SC-ura (Fig. 6a), or to accumulate LacZ–URA3 mRNA (Fig. 6b). It is important to notice that hrs1Δ mutant is not affected in the expression of the tet promoter as the mutant accumulates similar RNA levels as the wild type (Fig. 6b).
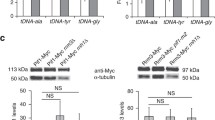
Expression analysis of hrs1Δ and hpr1Δ hrs1Δ mutants in the pLAUR system. a Phenotypic analysis of hpr1Δ (UKH-1A), hrs1Δ (UKH-1C), hpr1Δ hrs1Δ (UKH-1B) and wild type (UKH-1D) isogenic strains transformed with pLAUR system. The scheme of the system is shown at the top. The capacity of each strain to form colonies on synthetic medium lacking tryptophan with or without uracil after four days at 30°C is shown. b Northern analysis of the transformants listed in a. The lacZ and rDNA probes used are detailed in “Materials and methods”. Other details as in Fig. 2
Since the hrs1Δ-mediated suppression of hpr1Δ hyper-recombination is due to the reduction of the transcription through the recombination system, we wanted to know if this was also the case for the gene expression defect of this mutant. To study whether the intensity of transcription could affect the levels of transcription impairment, we determined the effect of hpr1Δ on the transcription of LYS2 as a function of the intensity of the promoter used. For this purpose we constructed a new plasmid-borne gene expression system in which the LYS2 gene was placed under the control of the tet promoter (tetpr::LYS2). We compared the expression of this construct with a similar construct based on the GAL1 promoter (GAL1pr::LYS2), instead of tet. The level of expression of the GAL1 promoter is five times higher than the maximum level of the tet promoter in the wild type (Fig. 7a). As can be seen in Fig. 7a, transcription in the tetpr::LYS2 system is reduced in hpr1Δ mutant (50% of the levels of the wild type). However, the reduction observed in the GAL1pr::LYS2 construct is much stronger given that the mutant only accumulated about 5% of the transcript levels of the wild type (Fig. 7a). Moreover, we can see an actual increase in mRNA levels in hpr1Δ mutant when the promoter strength is decreased, given that the mutant shows a twofold increase in the amount of RNA in the tetpr::LYS2 construct compared to the GAL1pr::LYS2 one, while in the wild type there is a fivefold reduction (Fig. 7a). This result suggests that the strength of transcription could modulate the effect of hpr1Δ in gene expression. To asses this possibility we analysed the effect of hpr1Δ on transcription of the tetpr::LYS2 system.
Effect of the strength of the promoter on hpr1Δ transcription defect of LYS2-based constructs. a Northern analysis of the wild type (HRN1-4A) and hpr1Δ mutant (HRN1-2C) congenic strains transformed either with GAL1pr::LYS2 or with tetpr::LYS2 systems. The LYS2 and rDNA probes used are detailed in “Materials and methods”. Other details as in Fig. 2. b Northern analysis of wild type and hpr1Δ congenic strains carrying the tetpr::LYS2 system using increasing amounts of doxycycline (dox). The percentage of RNA levels respect to the wild type is indicated on the top of each value. Media and the standard deviation of three independent experiments are shown. Other details as in a
Transcription in the tetpr::LYS2 construct was assayed with different doxycycline (dox) concentrations (0, 0.0025 and 0.005 mg/ml). The higher the concentration the weaker the promoter activation. Accordingly, the levels of LYS2 mRNA decreased progressively in correlation with the amount of doxycycline either in wild-type or hpr1Δ cells (Fig. 7b). As expected, hpr1Δ mutant transcript accumulation was reduced with respect to the wild type. However, as the strength of transcription became lower the difference between mutant and wild-type cells became weaker. There was a progressive disappearance of the transcription impairment effect caused by hpr1Δ that correlated with the decreasing frequency of transcription initiation at the tet promoter until reaching a point in which no transcription defect was observed. As can be seen in Fig. 7b, at 0.005 mg/ml doxycycline the difference in transcription levels between wild type and hpr1Δ strains is not significant.
Similar results were obtained using the pLAUR system in hpr1Δ, tho2Δ and mft1Δ mutants. Figure 8 shows that, as for the tetpr::LYS2 construct, the transcriptional defect of tho mutants became weaker when promoter firing was reduced. The hpr1Δ mutant accumulated 9.3% of the wild-type LacZ-URA3 mRNA levels when transcription was high, but reached a 46% value in the presence of 0.0025 mg/ml of doxycycline. Under these conditions, tho2Δ and mft1Δ mutants behaved similarly, the increase going from 2.2 to 25% and from 15 to 41%, respectively (Fig. 8).
Transcription impairment in tho mutants as a function of promoter strength. Northern analyses of wild-type (W303-1A), hpr1Δ (U678-1C), tho2Δ (RK2-6D) and mft1Δ (WMK-2A) isogenic strains harbouring the plasmid-based pLAUR system using increasing amounts of doxycycline (dox) are shown. The percentage of RNA levels respect to the wild type is indicated on the top of each value. Media and the standard deviation of three independent experiments are shown. The lacZ and rDNA probes used are detailed in “Materials and methods”. Other details as in Fig. 2
These results indicate that, together with the G + C content and the length of the DNA sequence, the frequency at which transcription is fired from the promoter influences the physiological relevance of the THO complex in gene expression.
Discussion
Hrs1 (Med3) is a component of the Mediator of RNAPII that was genetically identified as a suppressor of the hyper-recombination phenotype of hpr1Δ mutant. In this study we show that hrs1Δ does not suppress the hyper-recombination that occurs independent of transcription. The hrs1-mediated suppression of hpr1Δ hyper-recombination is not general and is only observed in recombination constructs in which transcription is Hrs1-dependent. In addition we demonstrate that hpr1Δ gene expression defect can be alleviated reducing the frequency of transcription. Altogether, our results agree with the idea that gene expression and recombination phenotypes of THO mutants can be alleviated or suppressed by decreasing the transcription rate.
Suppression of hyper-recombination by hrs1Δ is the result of the abolishment of transcription initiation
We have studied the effect of HRS1 deletion in cdc2 and cdc9 hyper-recombinant mutants, which recombination phenotype is linked to a defect in replication (Montelone et al. 1981; Kokoska et al. 2000). We have observed a small reduction in the recombination frequency in both hrs1Δ cdc2 and hrs1Δ cdc9 double mutants compared to the cdc simple mutants. However, this reduction was due to the decrease in recombination observed in the hrs1Δ simple mutant (Santos-Rosa and Aguilera 1995). hrs1Δ also suppresses the transcription-associated hyper-recombination phenotype observed in other mutants like tho2Δ and thp1Δ (Gallardo and Aguilera 2001). Therefore, hrs1-mediated suppression is not general to all types of hyper-recombination, but specific of transcription-associated hyper-recombination.
Several transcription mutants have been isolated as hpr1Δ suppressors in different screenings. Although all of them affect transcription they seem to suppress in different ways. Thus, soh1, soh2 and soh4 (mutants in Med31, Rpb2 and TFIIB, respectively) were identified as suppressors of hpr1Δ thermosensitivity phenotype (Fan and Klein 1994). This observation together with the fact that soh mutants suppress the synthetic growth defects of hpr1 Δtop1Δ and hpr1Δ hht1-hhf1Δ double mutants suggests that there is a genetic interaction between these factors. Besides, soh mutants and mutations in other mediator components genes as GAL11/MED15 and SIN4/MED16 have been shown to suppress hpr1Δ hyper-recombination phenotype, although to a lesser extent than hrs1Δ (Fan and Klein 1994; Piruat et al. 1997).
hrs1 and srb2 mutants were identified as the suppressors of hpr1Δ hyper-recombination phenotype. However, while hrs1Δ mutation decreases TAR in the wild type, srb2Δ does not (Santos-Rosa and Aguilera 1995). Here we demonstrated that hrs1Δ-mediated suppression is specific of the recombination assay studied. hrs1Δ reduces TAR both in wild-type and hpr1Δ cells only in the Lk-AU repeat construct, consistent with the idea that in both cases recombination reduction by hrs1Δ occurs by similar mechanisms. These results indicate that the suppression of hyper-recombination by hrs1Δ is not due to a direct genetic interaction between Hrs1 and THO, but is the result of the abolishment of transcription of the DNA repeat construct.
A confirmation that transcription through the pBR322 region covering the amp gene and the ori sequence in Lk-AU is responsible for the hyper-recombination caused by hpr1Δ that is suppressed by hrs1Δ was obtained with recombination assays producing such transcripts from the LEU2 promoter, which is Hrs1-independent. In these systems (LY and LYΔNS) hyper-recombination was not abolished by hrs1Δ. Consistently, identical assays lacking such pBR322 sequences, but containing URA3 and ADE2 (LU and LA) were not hyper-rec in hpr1Δ cells (Fig. 4; Prado et al. 1997). Therefore, we can conclude that the suppression by hrs1Δ is the consequence of the effect of hrs1 on transcription abolishment. Although it is well established the relationship between hyper-recombination observed in hpr1Δ and transcription through the intervening region (Prado et al. 1997) we could still consider the possibility of additional transcription requirements (overlapping transcription, direction of transcription, transcription of the repeats, etc.) that could account for part of the hrs1Δ suppression. However, given the specificity of suppression of hrs1Δ for hpr1Δ and related mutants, we can basically rule out this possibility.
The gene expression defect of hpr1Δ mutants is modulated by the transcription rate
Using the regulatable tet promoter, we see that the transcriptional defect of hpr1Δ mutant decreases with the transcription rate. This decrease is not due to a defect in the repression of the tet promoter in hpr1Δ mutant, because LYS2 mRNA levels increase in this mutant when the strong GAL1 promoter is replaced by the tet promoter. The same effect was observed in other THO mutants (tho2Δ and mft1Δ). Therefore, the stronger the transcription of a gene the more important becomes the THO complex for its transcription. The relevance of the transcription rate in the phenotypes of THO mutants is not restricted to gene expression and hyper-recombination phenotypes. Thus, it has been shown that mutations in the Rad3 component of the TFIIH transcription initiation factor and transcription elongation inhibition suppress the accumulation of the transcript at the site of transcription sites in mft1Δ mutants (Jensen et al. 2004). These observations are not exclusive of THO-complex mutants, since it has also been observed in other mutants defective in mRNA processing such as dbp5 (Estruch and Cole 2003) and ssu72 (Pappas and Hampsey 2000; Dichtl et al. 2002).
One important feature of most of the phenotypes associated to mutants of THO is the dependency on the RNA molecule (Huertas and Aguilera 2003). This, together with the fact that the recruitment of export factors Yra1, Sub2 and Mex67 to chromatin is dependent on the THO/TREX complex (Zenklusen et al. 2002; Gwizdek et al. 2006) opens the possibility that in THO/TREX mutants, at high transcription rates, such export factors are not properly loaded onto the nascent mRNA, while at low transcription rates, there could be an alternative loading mechanism independent of THO/TREX. Moreover, hpr1Δ cells show wild-type levels of transcription when the novel RNA-binding protein Tho1 or the RNA helicase involved in transport Sub2 are overexpressed (Jimeno et al. 2006), suggesting that in hpr1Δ background, the nascent mRNA would need additional RNA-binding factors to alleviate the expression defect. Our results agree with the view that at high transcription rates the demand for RNA packaging factors required for the formation of an export-competent mRNP is higher, in which case THO becomes crucial for the transcription process.
There are other data that may provide plausible explanations to these observations. Thus, genes containing short direct repeats require the THO complex for its transcription, a phenotype that is suppressed by topoisomerase I overexpression (Voynov et al. 2006). Also, hpr1Δ and top1Δ mutants are synthetically sick (Aguilera and Klein 1990; Sadoff et al. 1995). These data may indicate that the THO complex could be necessary to avoid aberrant DNA structures arising during transcription elongation. In this sense it would be plausible that those types of topological constraints become more frequent as the transcription frequency increases. In the absence of THO, a reduction in the firing of the promoter could allow the action of a topo I-dependent pathway that might remove such structures or directly decrease their formation. It is interesting that, as it occurs with the tetpr::LYS2 assay, THO- dependency of transcription of the TIR1 gene, containing short DNA repeats, is only observed under strong transcription conditions (Voynov et al. 2006).
We have previously reported in vivo and in vitro data that show that THO is important for transcription elongation of strongly transcribed constructs (Chavez et al. 2001; Rondon et al. 2003). Our results confirm that highly transcribed genes require the THO complex for transcription elongation. At low transcription rates, however, THO becomes dispensable.
References
Aguilera A (2005) mRNA processing and genomic instability. Nat Struct Mol Biol 12:737–738
Aguilera A, Klein HL (1990) HPR1, a novel yeast gene that prevents intrachromosomal excision recombination, shows carboxy-terminal homology to the Saccharomyces cerevisiae TOP1 gene. Mol Cell Biol 10:1439–1451
Boube M, Joulia L, Cribbs DL, Bourbon HM (2002) Evidence for a mediator of RNA polymerase II transcriptional regulation conserved from yeast to man. Cell 110:143–151
Brickner JH, Walter P (2004) Gene recruitment of the activated INO1 locus to the nuclear membrane. PLoS Biol 2:e342
Cabal GG et al (2006) SAGA interacting factors confine sub-diffusion of transcribed genes to the nuclear envelope. Nature 441:770–773
Casolari JM, Brown CR, Komili S, West J, Hieronymus H, Silver PA (2004) Genome-wide localization of the nuclear transport machinery couples transcriptional status and nuclear organization. Cell 117:427–439
Casolari JM, Brown CR, Drubin DA, Rando OJ, Silver PA (2005) Developmentally induced changes in transcriptional program alter spatial organization across chromosomes. Genes Dev 19:1188–1198
Chavez S et al (2000) A protein complex containing Tho2, Hpr1, Mft1 and a novel protein, Thp2, connects transcription elongation with mitotic recombination in Saccharomyces cerevisiae. EMBO J 19:5824–5834
Chavez S, Garcia-Rubio M, Prado F, Aguilera A (2001) Hpr1 is preferentially required for transcription of either long or G + C-rich DNA sequences in Saccharomyces cerevisiae. Mol Cell Biol 21:7054–7064
Conrad MN, Newlon CS (1983) Saccharomyces cerevisiae cdc2 mutants fail to replicate approximately one-third of their nuclear genome. Mol Cell Biol 3:1000–1012
Dichtl B et al (2002) A role for SSU72 in balancing RNA polymerase II transcription elongation and termination. Mol Cell 10:1139–1150
Estruch F, Cole CN (2003) An early function during transcription for the yeast mRNA export factor Dbp5p/Rat8p suggested by its genetic and physical interactions with transcription factor IIH components. Mol Biol Cell 14:1664–1676
Fan HY, Klein HL (1994) Characterization of mutations that suppress the temperature-sensitive growth of the hpr1 delta mutant of Saccharomyces cerevisiae. Genetics 137:945–956
Gallardo M, Aguilera A (2001) A new hyperrecombination mutation identifies a novel yeast gene, THP1, connecting transcription elongation with mitotic recombination. Genetics 157:79–89
Gwizdek C et al (2006) Ubiquitin-associated domain of Mex67 synchronizes recruitment of the mRNA export machinery with transcription. Proc Natl Acad Sci USA 103:16376–16381
Hartwell LH, Mortimer RK, Culotti J, Culotti M (1973) Genetic control of the cell division cycle in yeast: V. genetic analysis of cdc mutants. Genetics 74:267–286
Huertas P, Aguilera A (2003) Cotranscriptionally formed DNA:RNA hybrids mediate transcription elongation impairment and transcription-associated recombination. Mol Cell 12:711–721
Huertas P, Garcia-Rubio ML, Wellinger RE, Luna R, Aguilera A (2006) An hpr1 point mutation that impairs transcription and mRNP biogenesis without increasing recombination. Mol Cell Biol 26:7451–7465
Jensen TH, Boulay J, Olesen JR, Colin J, Weyler M, Libri D (2004) Modulation of transcription affects mRNP quality. Mol Cell 16:235–244
Jimeno S, Rondon AG, Luna R, Aguilera A (2002) The yeast THO complex and mRNA export factors link RNA metabolism with transcription and genome instability. EMBO J 21:3526–3535
Jimeno S, Luna R, Garcia-Rubio M, Aguilera A (2006) Tho1, a novel hnRNP, and Sub2 provide alternative pathways for mRNP biogenesis in yeast THO mutants. Mol Cell Biol 26:4387–4398
Kim M, Ahn SH, Krogan NJ, Greenblatt JF, Buratowski S (2004) Transitions in RNA polymerase II elongation complexes at the 3’ ends of genes. EMBO J 23:354–364
Kokoska RJ, Stefanovic L, DeMai J, Petes TD (2000) Increased rates of genomic deletions generated by mutations in the yeast gene encoding DNA polymerase delta or by decreases in the cellular levels of DNA polymerase delta. Mol Cell Biol 20:7490–7504
Luna R, Jimeno S, Marin M, Huertas P, Garcia-Rubio M, Aguilera A (2005) Interdependence between transcription and mRNP processing and export, and its impact on genetic stability. Mol Cell 18:711–722
Mason PB, Struhl K (2005) Distinction and relationship between elongation rate and processivity of RNA polymerase II in vivo. Mol Cell 17:831–840
Montelone BA, Prakash S, Prakash L (1981) Spontaneous mitotic recombination in mms8–1, an allele of the CDC9 gene of Saccharomyces cerevisiae. J Bacteriol 147:517–525
Pappas DL Jr, Hampsey M (2000) Functional interaction between Ssu72 and the Rpb2 subunit of RNA polymerase II in Saccharomyces cerevisiae. Mol Cell Biol 20:8343–8351
Piruat JI, Aguilera A (1996) Mutations in the yeast SRB2 general transcription factor suppress hpr1-induced recombination and show defects in DNA repair. Genetics 143:1533–1542
Piruat JI, Aguilera A (1998) A novel yeast gene, THO2, is involved in RNA pol II transcription and provides new evidence for transcriptional elongation-associated recombination. EMBO J 17:4859–4872
Piruat JI, Chavez S, Aguilera A (1997) The yeast HRS1 gene is involved in positive and negative regulation of transcription and shows genetic characteristics similar to SIN4 and GAL11. Genetics 147:1585–1594
Prado F, Piruat JI, Aguilera A (1997) Recombination between DNA repeats in yeast hpr1delta cells is linked to transcription elongation. EMBO J 16:2826–2835
Reeves WM, Hahn S (2005) Targets of the Gal4 transcription activator in functional transcription complexes. Mol Cell Biol 25:9092–9102
Rondon AG, Jimeno S, Garcia-Rubio M, Aguilera A (2003) Molecular evidence that the eukaryotic THO/TREX complex is required for efficient transcription elongation. J Biol Chem 278:39037–39043
Sadoff BU, Heath-Pagliuso S, Castano IB, Zhu Y, Kieff FS, Christman MF (1995) Isolation of mutants of Saccharomyces cerevisiae requiring DNA topoisomerase I. Genetics 141:465–479
Santos-Rosa H, Aguilera A (1994) Increase in incidence of chromosome instability and non-conservative recombination between repeats in Saccharomyces cerevisiae hpr1 delta strains. Mol Gen Genet 245:224–236
Santos-Rosa H, Aguilera A (1995) Isolation and genetic analysis of extragenic suppressors of the hyper-deletion phenotype of the Saccharomyces cerevisiae hpr1 delta mutation. Genetics 139:57–66
Santos-Rosa H, Clever B, Heyer WD, Aguilera A (1996) The yeast HRS1 gene encodes a polyglutamine-rich nuclear protein required for spontaneous and hpr1-induced deletions between direct repeats. Genetics 142:705–716
Sikorski RS, Hieter P (1989) A system of shuttle vectors and yeast host strains designed for efficient manipulation of DNA in Saccharomyces cerevisiae. Genetics 122:19–27
Straka C, Horz W (1991) A functional role for nucleosomes in the repression of a yeast promoter. EMBO J 10:361–368
Strasser K et al (2002) TREX is a conserved complex coupling transcription with messenger RNA export. Nature 417:304–308
Thomas BJ, Rothstein R (1989) Elevated recombination rates in transcriptionally active DNA. Cell 56:619–630
Voynov V, Verstrepen KJ, Jansen A, Runner VM, Buratowski S, Fink GR (2006) Genes with internal repeats require the THO complex for transcription. Proc Natl Acad Sci USA 103:14423–14428
Zenklusen D, Vinciguerra P, Wyss JC, Stutz F (2002) Stable mRNP formation and export require cotranscriptional recruitment of the mRNA export factors Yra1p and Sub2p by Hpr1p. Mol Cell Biol 22:8241–8253
Acknowledgments
We would like to thank D. Haun for style supervision. This work was supported by grants from the Ministry of Science and Education of Spain (BMC2006-05260) and Junta de Andalucía (CVI-102 and CVI624).
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by H. Ronne.
Rights and permissions
About this article
Cite this article
Jimeno, S., García-Rubio, M., Luna, R. et al. A reduction in RNA polymerase II initiation rate suppresses hyper-recombination and transcription-elongation impairment of THO mutants. Mol Genet Genomics 280, 327–336 (2008). https://doi.org/10.1007/s00438-008-0368-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00438-008-0368-8