Abstract
Based on specimens collected from harvested American alligator Alligator mississippiensis Daudin, 1801 in Mississippi, USA, novel molecular data for both nuclear ribosomal genes (18S, ITS1-5.8S, ITS2, and 28S) and mitochondrial genes (cytochrome c oxidase subunit 1 and nicotinamide adenine dinucleotide dehydrogenase subunit 1) are provided for Odhneriotrema incommodum (Leidy, 1856), a trematode of the family Clinostomidae Lühe, 1901 infecting A. mississippiensis and the Florida spotted gar Lepisosteus platyrhincus DeKay, 1842. This represents the first sequencing data available for the genus Odhneriotrema and the subfamily Nephrocephalinae Travassos, 1928. Additionally, the results of phylogenetic analyses, additional morphometric data, a photomicrograph, and a line drawing supporting the present identification of O. incommodum are provided. These data will aid in elucidating the life cycle of O. incommodum through molecular identification of larval stages as well as understanding the evolutionary history of Clinostomidae and its subfamilies. Implications for the currently accepted organization of the Clinostomidae are discussed.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The American alligator Alligator mississippiensis is an abundant top predator and keystone species that can structure aquatic ecosystems throughout their range, spanning the Coastal Plain of the southeastern United States (Mazzotti and Brandt 1994). Acting as an ecosystem engineer, A. mississippiensis can be important predators and prey, mobilizing nutrients, manipulating hydrology, and influencing plant communities within aquatic and adjoining terrestrial ecosystems (Mazzotti et al. 2009; Rosenblatt and Heithaus 2011). A. mississippiensis exist in fresh, brackish (salinity 5–30‰), and marine (salinity > 30‰) systems (Elsey 2005; Rosenblatt and Heithaus 2011). Despite the array of systems occupied by A. mississippiensis, the parasite community associated with this species is considered relatively similar throughout this species’ range (Tellez 2014; Tellez and Nifong 2014). Interest in and taxonomic categorization of these parasites has a long history, riddled with change.
Odhneriotrema incommodum (Leidy 1856) (= Monostomum incommodum; Distoma oricola; Distomum incommodum; Clinostomum incommodum; Homoscaphis incommodum) was first described by Leidy (1856) as M. incommodum based on five specimens reportedly recovered from the feces of A. mississippiensis Daudin, 1801 in Florida, USA. Later, Leidy (1884) described an additional species, D. oricola based on eight worms collected from the mouth of the type host species from Florida. He also noted scarring of the tongue apparently associated with chronic infection. Leidy (1890) later concluded these two trematodes were conspecifics and named the species Distomum incommodum. Pratt (1902) and Ward and Whipple (1918) speculated the species likely belonged in the family Clinostomidae, but indicated insufficient data were available to define its taxonomic relationship within this family. Harrah (1922) moved the species into the genus Clinostomum, thus erecting C. incommodum.
In his description of four archived specimens reportedly collected from the thorax of the type host species from Florida, Canavan (1933) provided the first detailed morphological characterization of the species, although he mistook the pharynx for an oral sucker and thus erroneously concluded the species lacks a pharynx. It must be noted, however, that all subsequently examined adults of the species (McIntosh 1935), as well as observations in the present study, reveal O. incommodum does possess a well-developed, muscular pharynx. In addition to his description of the species, Canavan also erected the novel genus Homoscaphis based on the placement of the genital pore, large body size and dorsal convexity, relatively large acetabulum, placement and arrangement of the testes and ovary, lack of vitelline follicle confluence posterior to the reproductive organs, uterus arrangement, prominent caecal outpocketings, and host species, thus naming the trematode H. incommodum (Canavan, 1933).
Based on similarity of measured characters of three specimens collected from A. mississippiensis in Florida as well as one of Leidy’s original specimens, McIntosh (1935) noted the species was congeneric with Odhneriotrema microcephala, thus establishing the currently accepted O. incommodum. McIntosh (1935) notes the presence of a well-developed pharynx in all four specimens. Thus, prior to the present study, adults of this trematode have been described based on only 20 specimens collected from naturally infected A. mississippiensis, all from Florida.
Subsequent to these examinations, the family Clinostomidae, and thus O. incommodum, has been subject to a number of taxonomic rearrangements. Dollfus (1931) asserted the family Clinostomidae should be raised to the rank of a superfamily, the Clinostomoidea Witenberg, 1925, with the family Clinostomidae containing only the genera Clinostomum Leidy, 1856, Euclinostomum Travassos, 1928, and Ithyclinostomum Witenberg, 1926, while the family Opisthophallidae Travassos, 1926 should contain the genera Odhneriotrema Travassos, 1928, and Opisthophallus Baer, 1921. While agreeing with this organization within the family Clinostomidae, Baer (1933) contended the Clinostomidae should not be raised the rank of a superfamily and the latter two genera did not warrant placement in a distinct family. Skrjabin (1947) maintained the placement of the genus Odhneriotrema in the subfamily Opisthophallinae within the family Clinostomidae based on the presence of a pars prostatica, maturity of miracidia at the point of oviposition, and having crocodilians as definitive hosts. Dollfus (1950) reasserted his prior placement of Odhneriotrema and Opisthophallus into a distinct family, Nephrocephallidae (= Opisthophallidae), within the superfamily Clinostomoidea, distinguished from the Clinostomidae by a number of morphological differences and having reptiles as definitive hosts. Yamaguti (1958) moved Odhneriotrema into the subfamily Clinostominae Pratt, 1902 with Clinostomoides Dollfus, 1950, Clinostomatopsis Dollfus, 1932, and Clinostomum. Conversely, Tavassos et al. (1969) erected an entirely separate subfamily, Odhneriotrematinae Travassos, Freitas, and Kohn 1969 within the family Clinostomidae for Odhneriotrema, distinguishing it from the Clinostominae based on non-confluence of the vitellaria posterior to the gonads, the appearance of the uterus, and intertesticular distance. Yamaguti (1971), considering the work of Tavassos et al. (1969) too narrow in scope, reasserted his placement of Odhneriotrema within the subfamily Clinostominae, distinguished from the Nephrocephalinae by the intertesticular placement of the cirrus sac. Conversely, Feizullaev and Mirzoeva (1983) supported the placement of the genus within its own family within the superfamily Clinostomoidea as proposed by Tavassos et al. (1969). Keys to the Trematoda (Kanev et al. 2002) uses a similar system to that proposed by Yamaguti (1971), but places Odhneriotrema in the subfamily Nephrocephalinae within the family Clinostomidae based on host species, genital pore placement, and oral sucker size.
To aid in clarification of the long-disputed taxonomy of this species, the present study provides the first molecular description and phylogenetic analyses of O. incommodum in addition to morphometric and ecological data based on adults collected from an alligator processor in Mississippi, USA. Additionally, morphometric data, consisting of previously reported measurements for the species, and ecological data, consisting of the standard parasitological parameters of intensity, abundance, and prevalence are provided.
Materials and methods
Specimen collection
During Mississippi, USA’s 10-day alligator hunting season in August through September of 2016, 11 partial specimens of A. mississippiensis were collected from an alligator processor in Port Gibson, Mississippi. Of these 11 A. mississippiensis, 8 (1.37 to 3.58 m in length) still possessed tongues. Tongues and attached glottal regions of the buccal cavities were separated from hosts and transported in 0.9% saline separately to the Parasitology Laboratory of the Mississippi State University College of Veterinary Medicine, Mississippi State, Mississippi, USA, where they were grossly examined for the presence of parasites. Detected trematodes were removed from the tongues with forceps, relaxed in 0.9% hot saline (~ 95 °C), and stored in 70% molecular grade ethanol for morphological and molecular characterization.
Calculation of intensity, abundance, and prevalence
Intensity, defined as the number of parasites per infected host, abundance, defined as the number of parasite per host in all examined hosts, and prevalence, defined as the percentage of hosts examined found to be infected, were calculated. These definitions are consistent with those provided by Bush et al. (1997) and Rózsa et al. (2000). These parasitological parameters were calculated using Quantitative Parasitology 3.0 (Rózsa et al. 2000). Confidence limits of 95% for intensity, abundance, and prevalence were calculating using the program’s implementation of the Clopper-Pearson method (Clopper and Pearson 1934) with 2000 bootstrap replications.
Morphological identification and characterization
Thirteen ethanol fixed adult O. incommodum excised from three different A. mississippiensis were stained for 1 week in Semichon’s acetocarmine or a concentrated formulation of Van Cleave’s hematoxylin (5 mL of Delafield’s hematoxylin, 5 mL of Ehrlich’s hematoxylin, 6 g potassium aluminum sulfate, and 100 mL of distilled deionized water). After 1 week, specimens were destained in acidic ethanol until the parenchyma was pale and internal organs were distinct. Specimens were then transferred to alkaline ethanol, dehydrated in a series of four ethanol washes of increasing concentrations from 70 to 100%. Specimens were then cleared in Hemo-De (Scientific Safety Solvents, Texas, USA), and mounted on glass slides in Canada balsam (Aldon Corporation, New York, USA). Measurements of stained and mounted trematodes were taken using an Olympus BX50 microscope (Olympus Corporation, Tokyo, Japan) with an Olympus DP72 camera attachment and associated cellSens Standard 1.12 software. A line drawing was made with the aid of a camera lucida and digitized using Adobe Illustrator CC 2017.1 (Adobe Systems, San Jose, CA, USA). A photomicrograph was taken using an Olympus BX41 with an attached Nikon DS-Fi1 camera (Nikon Corporation, Tokyo, Japan).
Molecular analysis
Genomic DNA was extracted from a total of three specimens of O. incommodum, two from one A. mississippiensis, and one from a second using the DNeasy Blood and Tissue Kit (Qiagen, Hilden, Germany). Ribosomal genes (spanning partial 18S rRNA gene, internal transcribed spacer region 1, 5.8S rRNA gene, ITS 2 region, and partial 28S rRNA gene) were amplified using the following primer sets: ERIB1/ERIB10, Dipo1795F/Diplo2549R, BD1/BD2, Diplo2617F/Diplo3170R, and LSU5/1500R. Mitochondrial cytochrome c oxidase subunit 1 (CO1) and nicotinomide adenine dinucleotide dehydrogenase subunit 1 (NAD1) sequences were amplified using the following primer sets: modified versions of Dice1F/Dice14R with T3 and T7 tails removed, cox1_schist5’/modified Dice14R, JB3/JB4.5, and NDJ11/NDJ2a. Primer sequences and references for each primer are listed in Table 1. Each 20-μL reaction consisted of 10 μL of Phusion Green Hot Start II High-Fidelity PCR Master Mix (ThermoFisher, Waltham, MA, USA), 10 pmol/μL of each primer, 1 μL of DNA (> 10 ng/uL), and 7 μL of nuclease free water to volume. Thermal cycling profiles are indicated in Table 2.
To obtain pairwise distances between top BLASTN (Altschul et al. 1990) hits and sequences generated in the present study, sequences were aligned using the MAFFT algorithm (Katoh and Standley 2013) using GUIDANCE2 (Landan and Graur 2008; Sela et al. 2015). The resultant alignments were then trimmed by eye and gaps removed using MEGA7 (Kumar et al. 2016). Finally, pairwise distances were calculated using MEGA7.
To determine intraspecific variability for each gene target between sequences generated in the present study, sequences were aligned using the MUSCLE algorithm (Edgar 2004), trimmed by eye, and pairwise distances between them calculated in MEGA7.
Phylogenetic analysis
Similar to the methods of Caffara et al. (2016), ribosomal (ITS regions) and CO1 sequences belonging to members of the family Clinostomidae, as well as the diplostomids Alaria mustelae Bosma, 1931 and Diplostomum baeri Dubois, 1937 (as outgroups) were downloaded from the NCBI nr/nt database (Supplemental Table 1). Regions of the ITS1, 5.8S, and ITS2 genes were extracted using ITSx 1.0.11 (Bengtsson-Palme et al. 2013), subjected to gap removal, aligned using the MAFFT algorithm as implemented in Geneious 10.2 (Kearse et al. 2012), and concatenated with CO1 sequences in MEGA7 for a final alignment containing 1169 positions. Best-fitting models for each position were selected using the Bayesian Information Criterion: CO1 codon position 1 (TN93 + I; 176 positions), CO1 codon position 2 (HKY; 176 positions), CO1 codon position 3 (TN93 + G; 176 positions), ITS1 region (K2 + G; 402 positions), 5.8S rRNA gene (JC; 157 positions), and ITS2 region (JC + G; 70 positions). Phylogenetic inferences were made with MrBayes 3.2.6 (Ronquist and Huelsenbeck 2003; Altekar et al. 2004) by using Markov chain Monte Carlo searches of two simultaneous runs of four chains with sampling every 100th tree for 1 × 106 generations. This number of generations ensured the value of the standard deviation of split frequencies reached < 0.01. After the first 25% of trees were discarded as burn-in, posterior probability values were calculated from the remaining trees. Maximum likelihood analysis was performed using IQ-Tree (Nguyen et al. 2015) on the IQ-Tree web server (Trifinopoulos et al. 2016) with the concatenated alignment and partition scheme described previously. Branch support was tested using ultrafast bootstrap support (Minh et al. 2013) with 1000 pseudoreplicates. Trees were annotated in FigTree 1.4.3 (Rambaut 2016) and Adobe Illustrator 2017.1.
Results
Prevalence and intensity
Odhneriotrema incommodum was detected in five of eight A. mississippiensis buccal cavities examined, yielding a prevalence of 62.5% (95% bootstrap BCa confidence limit, 24.5–91.5%). The number of parasites in each infected host ranged from 1 to 16. Mean intensity was 5.4 (95% bootstrap BCa confidence limit, 1.8–11.4) O. incommodum per buccal cavity while the median intensity was 4.0 O. incommodum per buccal cavity. Mean abundance of O. incommodum per A. mississippiensis was 3.4 (95% bootstrap BCa confidence limit, 1–9.1).
Morphological characterization
All measurements given in micrometers unless otherwise stated. Measurements are given in the format range (mean ± standard deviation). Measurements for individual worms are also available (Supplementary Table 2). Measurements were consistent with those previously reported for the species while not consistent with those reported for the only congenerous species, Odhneriotrema microcephala (Table 3). Voucher specimens are deposited in the Smithsonian Institution, Museum of Natural History, Washington, District of Columbia, USA (USNM 1457205, 1457207).
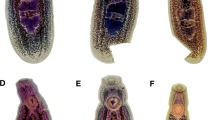
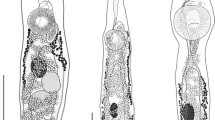
Body elongate, dorsally convex, and ventrally flattened, 8.7–19.2 (14.8 ± 3.6) mm × 1.3–2.3 (1.8 ± 0.3) mm (Figs. 1, 2). Anterior end of body possesses reniform collar surrounding the oral sucker. Collar 0.4–1.4 (0.8 ± 0.3) mm × 0.8–1.4 (1.1 ± 0.2) mm. Oral sucker ovoid to circular, 222.4–559.7 (366.8 ± 94.4) × 286.2–452.9 (381.6 ± 60.5). Pharynx ovoid, immediately posterior to oral sucker, 315.8–562.8 (454.5 ± 72.3) × 181.8–389.0 (299.8 ± 49.5). Caeca lacking distinct diverticula in most specimens, bifurcate just posterior to pharynx, marginal until terminating posterior to anterior testis and anterior to excretory pore. Acetabulum large, muscular 1.0–1.7 (1.4 ± 0.2) mm × 0.6–1.5 (1.2 ± 0.2) mm. Acetabulum muscular ring 238.3–598.5 (413.9 ± 116.2) in thickness. Acetabulum inner orifice 235.0–832.6 (642.2 ± 150.9) in diameter. Distance from anterior end of acetabulum to anterior end of body 2.7–5.2 (3.9 ± 0.7) mm. Acetabulum diameter to pharynx diameter ratio 3.2–4.6 (3.9 ± 0.4). Distance from anterior border of acetabulum to middle of body 0.6–3.3 (2.2 ± 1.1) mm. Testes, two, irregularly pyramidal, unlobed. Anterior testis, 324.6–880.2 (656.6 ± 199.0) × 99.2–545.1 (358.5 ± 126.21). Distance from anterior testis to posterior testis 0.6–1.4 (1.0 ± 0.3) mm. Posterior testis 268.3–956.6 (555.8 ± 196.0) × 218.7–586.9 (426.9 ± 112.2). Cirrus sac large, ovoid, contains pars prostatica, intertesticular, 402.8–1830.9 (1299.4 ± 479.1) × 126.7–364.4 (259.4 ± 84.9). Distance from posterior end of posterior testis to posterior end of the body 0.7–1.6 (1.1 ± 0.3) mm. Vitelline follicles extend from just posterior to point of caecal bifurcation to acetabulum, then extend laterally from posterior to the acetabulum for the length of the ceaca, not convergent posterior to reproductive organs. Ovary round, 97.1–394.8 (295.1 ± 98.8) × 148.1–600.5 (298.5 ± 124.8). Uterus extends anteriorly from ovary, loops posteriorly just before ventral sucker, meets the metraterm at level of anterior testis, contains eggs too numerous to count. Genital pore opening in right margin of body between ovary and posterior testis. Distance from genital pore to posterior testis 40.8–426.6 (147.3 ± 101.5). Distance from center of genital pore to posterior end of body 0.9–2.1 (1.5 ± 0.4) mm. Y-shaped excretory canal at posterior end of body, terminates at excretory pore. Eggs (n = 90, 10 measured from each gravid specimen), elliptical, average 58.4–110.7 (95.9 ± 15.7) × 42.6–59.2 (52.8 ± 5.2), mature eggs containing developed miracidia were located nearest genital pore.
Molecular characterization
NCBI accession numbers for sequences generated from three specimens in the present study are as follows: MF766001-MF766003: cytochrome c oxidase subunit 1; MF765998-MF766000: 18S, ITS1-5.8S, ITS2, and 28S rRNA; and MF766004-MF766006: nicotinamide adenine dinucleotide dehydrogenase subunit 1.
Top BLASTN hits for the 784–1051-bp CO1 sequences for Clinostomidae were Clinostomum attenuatum Cort, 1913 metacercariae from Lithobates sp. Fitzinger, 1843 (Locke et al. 2015; KP150306), Clinostomum marginatum (Rudolphi, 1819) metacercariae from yellow perch Perca flavescens Mitchill, 1814 (Caffara et al. 2011; JF718610), Clinostomum detruncatum Braun, 1899 metacercariae from marbled swamp eels Synbranchus marmoratus Bloch, 1785 (Locke et al. 2015; KP110519), Clinostomum tataxumui Sereno-Uribe et al., 2013 metacercariae from Pacific sleepers Gobiomorus maculatus Günther, 1859 (Locke et al. 2015; KP110551), and Clinostomum cutaneum Paperna, 1964 metacercariae from Nile tilapia Oreochromis niloticus Linnaeus, 1758 (Locke et al. 2015; KP110516). Pairwise distances between Odhneriotrema incommodum CO1 sequences and top BLASTN hits are shown in Table 4. Intraspecific variability at CO1 was 0–0.13%.
Top BLASTN hits identified to species level for ribosomal sequences for Clinostomidae were Clinostomum marginatum metacercariae from Notropis sp. Rafinesque, 1818 (Sereno-Uribe et al. 2013; JX631101), Clinostomum album Rosser et al., 2017 adults from Great Egrets Ardea alba Linnaeus, 1758 (Rosser et al. 2017; KU708008), Clinosotmum complanatum (Rudolphi, 1814) metacercariae from Nile tilapia Oreochromis niloticus (Gustinelli et al. 2010; FJ609420), Clinostomum phalacrocoracis Dubois, 1931 metacercariae from Nile tilapia Oreochromis niloticus (Gustinelli et al. 2010; FJ609422), and Clinostomum tataxumui Sereno-Uribe et al., 2013 adults from Bare-throated Tiger Herons Tigrisoma mexicanum Swainson, 1834 (Pérez-Ponce de León et al. 2016; JX631050). Pairwise distances between Odhneriotrema incommodum ribosomal sequences and top BLASTN hits are shown in Table 5. Intraspecific variability for ribosomal genes was 0%.
The only Clinostomidae sequence a BLASTN search identified to species level for the NAD1 sequences was that of a complete mitochondrial genome, reportedly from Clinostomum complanatum metacercariae (74.24–74.42% sequence similarity) from goldfish Carassius auratus Linnaeus, 1758 (Chen et al. 2016; KM923964). Comparison of cytochrome c oxidase subunit 1 sequences from the same specimens to this genome showed sequence similarity of 79.35–80.20%. Intraspecific variability at NAD1 was 0–0.46%.
Phylogenetic analysis
Phylogenetic inference based on both conserved (ribosomal) and fast-evolving (mitochondrial) sequencing data placed Odhneriotrema incommodum within the Clinostomidae basal to Euclinostomum, which formed a clade basal to Clinostomum, while the diplostomids Alaria mustelae and Diplostomum baeri were chosen as the outgroup (Fig. 3). While the phylogenetic placement of Odhneriotrema is novel to the present study, placement of Euclinostomum is consistent with analyses carried out in previous studies (Senapin et al. 2014; Caffara et al. 2016). These results are consistent with the systematic arrangement of the family Clinostomidae as reported by Kanev et al. (2002).
Phylogenetic tree constructed using an alignment of concatenated ribosomal and cytochrome c oxidase subunit 1 sequences. Topology based on Bayesian inference analysis. Numbers above branches indicate Bayesian posterior probabilities/bootstrap values. Posterior probabilities less than 0.5 and bootstrap values less than 50 not shown. Redundant species names in each clade are also omitted. Scale bar represents the number of substitutions per site
Discussion
Odhneriotrema incommodum collected in the present study was morphologically consistent with previous accounts of the species (Leidy 1856; McIntosh 1935), including those that provided standard measurements of morphological characters (Leidy 1884; Canavan 1933). However, the measurements of specimens of O. incommodum collected in the present study were not consistent with measurements for the only congenerous species; O. microcephala (Travassos 1922, 1928), collected from the mouth of the spectacled caiman Caiman crocodilus Linnaeus, 1758.
Leidy (1856) described the species as collected from the feces of A. mississippiensis while all subsequent descriptions of adult O. incommodum have been based on specimens collected only from the buccal cavities of A. mississippiensis (Leidy 1884; Canavan 1933; McIntosh 1935). Odhneriotrema incommodum was recovered only from the buccal cavities in the present study despite a full examination of the gastrointestinal tract for the collection of other helminths. Given these findings, coupled with the relationship of O. incommodum to the Clinostomidae, which are typically associated with the mouth, buccal cavity, and esophagus of their various definitive host species (Ukoli 1966), it is probable the specimens Leidy described had died or failed to attach to their host properly and were incidentally ingested. It is also possible that his specimens were simply mislabeled. In his redescription of the species, Canavan (1933) lists their location in the host as “thorax.” Because Canavan’s description is based on archived specimens labeled as having been recovered from the thorax, he likely repeated another’s erroneous labeling. Barring these oddities in reported infection sites, observations presented in the current study are consistent with historical accounts of this species.
While the complete life history of O. incommodum is unknown, encapsulated metacercariae have been briefly described as 6–8 mm orange cysts in the gonads, mesentery, and fatty tissues of Florida spotted gar Lepisosteus platyrhincus DeKay 1842 (Leigh 1960). Subsequently L. platyrhincus was experimentally confirmed as a viable second intermediate host by Leigh (1978) who force-fed metacercariae collected from naturally infected fish to uninfected A. mississippiensis and observed the development of the adult worms in the buccal cavities of the definitive hosts, noting the formation of fibrotic lesions at attachment sites. The finding of numerous fibrotic lesions within the buccal cavities of the A. mississippiensis in the present study supports Leigh’s (1963, 1978) hypothesis that worms relocate to new attachment sites in the buccal cavity after lesions form. Fibrotic lesions on host tongues were often more numerous than the number of O. incommodum detected.
Given that the current geographic range of L. platyrhincus (IUCN 2013) does not overlap with the host locations for A. mississippiensis in the present study, it is unlikely L. platyrhincus is the only naturally occurring second intermediate host species for O. incommodum. More plausible is that other gar species are capable of serving as second intermediate hosts in the life cycle of O. incommodum. Gar species common to Mississippi include the shortnose gar L. platostomus Rafinesque, 1820, the spotted gar L. oculatus Winchell, 1864, the longnose gar L. osseus Linnaeus, 1758, and the alligator gar Atractosteus spatula Lacepède, 1803 (Ross 2002). Stomach content analyses carried out on A. mississippiensis have shown several of the aforementioned species to be prey items including Atractosteus spatula (Gabrey 2010), unidentified gar species (Overstreet et al. 1985), and Lepisosteus spp. (McNease and Joanen, 1977). Thus, parasitological examinations of potential second intermediate hosts species, particularly of the genera Lepisosteus and Atractosteus, are needed to identify additional hosts, provide a more detailed morphological description of the metacercariae, and to carry out histopathological analyses of infections in second intermediate hosts. Previous surveys of L. oculatus have shown higher prevalence of O. incommodum in females with metacercariae being found encysted chiefly in the ovaries. Additionally, metacercariae were found in the testes of males and the mesenteries of both sexes (Leigh 1960, 1978). Histopathological examinations of fish infected with metacercariae of other species have been shown to cause inflammation in host ovarian tissues (Blazer 2002), which may have implications for host fecundity, as has been suggested for other helminth taxa infecting this site in fish hosts (Clarke et al. 2006).
Although the adult and metacercaria of O. incommodum are both described, the cercaria and first intermediate host species remain unknown. Experimental exposure of snails of the species Planorbella duryi (= Helisoma duryi) Wetherby, 1879 and Physa pumilia Conrad, 1834 to miracidia failed to produce infected snails, possibly indicating these particular snail species are not suitable hosts for O. incommodum (Leigh 1978).
Given that larval stages of Clinostomidae have been previously reported from aquatic planorbid snails of the genera Biomphalaria Preston, 1910 (Pinto et al. 2015; Fernández et al. 2016), Planorbella Halderman, 1842 (Hunter and Hunter 1934), and Bulinus Müller, 1781 (Feizullaev and Mirzoeva 1983) as well as the lymnaeid Radix auricularia Linnaeus, 1758 (Chung et al. 1998), viable snail hosts may be in these genera. Planorbid species susceptible to infection with clinostomids are known to inhabit the range of A. mississippiensis (Hunter and Hunter, 1934). Further malacological surveys, coupled with morphological and molecular analysis of collected cercariae, are needed to identify the natural snail host of O. incommodum to elucidate the complete life cycle. Experimental infections may also reveal additional first intermediate host species. Snails of the Neotropical genus Biomphalaria have recently invaded the range of A. mississippiensis (Pointier et al. 2005) and may be able to serve as hosts for this and other clinostomids, as Biomphalaria spp. have been shown to serve as hosts for Clinostomum sp. (Pinto et al. 2015).
Kanev et al. (2002) present two historical classification systems for the family Clinostomidae, those of Skrjabin (1947) and Yamaguti (1958, 1971). For their part, Kanev et al. employ their own system most similar to that of Yamaguti. Additionally, Feizullaev and Mirzoeva (1983) provide another reclassification. Skrjabin placed Odhneriotrema in the subfamily Ophisthophallinae with Ophisthophallus based on the presence of a prostatic gland in the cirrus sac, whereas Yamaguti placed it within the subfamily Clinostominae with Clinostomum, Clinostomatopsis, and Clinostomoides based on the structure of the excretory system. Finally, Feizullaev and Mirzoeva (1983) placed the Odhneriotrema species in their own family, the Odhneriotremidae, based on definitive host species. Feizullaev and Mirzoeva also state they have based their reassessment of the superfamily on statistical analysis of morphometric data between and within species, but these data are not available for independent analysis, though keys to the families were provided based on said analyses (Feizullaev and Mirzoeva 1986). While significant efforts have been made to better determine these relationships using molecular data within the genus Clinostomum (Locke et al. 2015; Caffara et al. 2017), similar efforts have not been made to determine the systematics of the family as a whole. Molecular data from other taxa within the Clinostomidae are needed to more accurately determine the evolutionary relationships between genera within the Clinostomidae. It is hoped the molecular data provided here will be useful for such a task as it is the first for a member of the Clinostomidae not belonging to Clinostomum, Euclinostomum, or Clinostomoides. It is notable that the present analyses place a member of a Clinostomid subfamily which uses a reptile as a definitive host basal to species which rely on avian definitive hosts. Should future phylogenetic analyses carried out on other members of the Nephrocephalinae place them similarly, this may suggest a parasite evolutionary history that mirrors that of the hosts. However, in the absence of analyses employing calibrated molecular clock techniques, this is highly speculative.
Greater intrafamilial sequence variability at NAD1 than at CO1 suggest this region may be of use for molecular confirmation of species within the Clinostomidae with greater resolution than CO1, as has been shown for other trematode families (Morgan and Blair 1998). To establish this, more sequencing data are needed for these regions from other members of the family Clinostomidae and other closely related members of the superfamily. It should be noted however, that the reported species identity for the sequence of C. complanatum to which sequences from the present study were compared is somewhat doubtful. A BLASTN search using the cytochrome c oxidase subunit 1 barcoding region from this mitochondrial genome sequence reveals a 100% match to be Clinostomum sp. 8 (KP110536; Locke et al., 2010) and only a 96% match to C. complanatum (KU236382; Gaglio et al. 2016). Unfortunately, the sequences of both Chen et al. (2016) and Locke et al. (2015) are based on metacercariae and no morphometric data justifying specific diagnosis is included..
Although Feizullaev and Mirzoeva (1983) did not provide data to justify their reordering of the taxonomy of the Clinostomidae, their observation that many morphometric characters employed have not had their taxonomic utility rigorously assessed is nonetheless compelling. The advent of DNA sequencing provides a useful way to validate the utility of these characters. By using sequencing data to validate ranges of these measurements, which are thought to define particular species, it should be possible to determine which are of more use in distinguishing between taxa. With this in mind, the measurements from individual specimens are provided here (Supplemental Table 2) to better facilitate statistical comparisons of measured characters within and between species in the hopes that the utility of individual morphological characters, diagnostic of species identity and confirmed with molecular data, can be quantitatively assessed. The authors of the present study would encourage other researchers, particularly those working with taxa within the family Clinostomidae, to do the same. Authors who seldom report morphometric data may be more inclined to do so if the utility of those measurements were quantified and not unduly cumbersome to collect.
References
Altekar G, Dwarkadas S, Huelsenbeck JP, Ronquist F (2004) Parallel metropolis coupled Markov chain Monte Carlo for Bayesian phylogenetic inference. Bioinform 20:407–415
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410
Baer JG (1933) Note sur un nouveau Trématode, Clinostomum lophophallum sp. nov., avec quelques considerations générales sur la famille des Clinostmoidae. Rev Suisse de Zool 40(23):317–342
Barta JR, Martin DS, Liberator PA, Dashkevicz M, Anderson JAW, Feighner SD, Elbrecht A, Perkins-Barrow A, Jenkins MC, Danforth HD, Ruff MD, Profous-Juchelka H (1997) Phylogenetic relationships among eight Eimeria species infecting domestic fowl inferred using complete small subunit ribosomal DNA sequences. J Parasitol 83:262–271
Bengtsson-Palme J, Ryberg M, Hartmann M, Branco S, Wang Z, Godhe A, De Wit P, Sáchez-García EI, de Sousa F, Amend A, Jumpponen A, Unterseher M, Kristiansson E, Abarenkov K, Bertrand YJK, Sanli K, Eriksson KM, Vik U, Veldre V, Nilsson RH (2013) Improved software detection and extraction of ITS1 and ITS2 from ribosomal ITS sequences of fungi and other eukaryotes from analysis of environmental sequencing data. Methods Ecol Evol 4(10):914–919
Blazer VS (2002) Histopathological assessment of gonadal tissue in wild fishes. Fish Physiol Biochem 26:85–101
Bowles J, Blair D, McManus DP (1995) A molecular phylogeny of the human schistosomes. Mol Phylo Evol 4(2):103–109
Bush AO, Lafferty KD, Lotz JM, Shostak AW (1997) Parasitology meets ecology on its own terms: Margolis et al. revisited. J Parasitol 83(4):575–583
Caffara M, Davidovich N, Falk R, Smirnov M, Ofek T, Cummings D, Gustinelli A, Fioravanti ML (2014) Redescription of Clinostomum phalacrocoracis metacercariae (Digenea: Clinostomidae) in cichlids from Lake Kinneret, Israel. Parasite 21:32
Caffara M, Locke SA, Cristanini C, Nadav D, Perry MP, Fioravanti ML (2016) A combined morphometric and molecular approach to identifying metacercariae of Euclinostomum heterostomum (Digenea: Clinstomidae). J Parasitol 102(2):239–248
Caffara M, Locke SA, Echi PC, Halajian A, Benini D, Luus-Powell WJ, Tavakol S, Fioravanti ML (2017) A morphological and molecular study of clinostomid metacercariae from African fish with a redescription of Clinostomum tilapiae. Parasitol 114(11):1519-1529
Caffarra M, Locke SA, Gustinelli A, Marcogliese DJ, Fioravan ML (2011) Morphological and molecular differentiation of Clinostomum complanatum and Clinostomum marginatum (Digenea: Clinostomidae) metacercariae and adults. J Parasitol 97:884–891
Canavan WPN (1933) A redescription of Distomum incommodum Leidy, from Alligator mississippiensis, and the creation of a new genus (Homoscaphus) for it. Parasitology 25:501–509
Chen L, Feng Y, Chen HM, Wang LX, Feng HL, Yang X, Mughal MN, Fang R (2016) Complete mitochondrial genome analysis of Clinostomum complanatum and its comparison with selected digeneans. Parasitol Res 115:3249–3256
Chung DI, Kong HH, Joo CY (1998) Radix auricularia coreana: natural snail host of Clinostomum complanatum in Korea. Korean J Parasitol 36(1):1–6
Clarke LM, Dove ADM, Conover DO (2006) Prevalence, intensity, and effect of a nematode (Philometra saltatrix) in the ovaries of bluefish (Pomatomus saltatrix). Fish Bull 104(1):118–124
Clopper C, Pearson ES (1934) The use of confidence or fiducial limits illustrated in the case of the binomial. Biometri 26:404–413
Dollfus R (1931) Répertoire des espèces et genres nouveaux. Ann Parasitol Hum Comp 9:492–493
Dollfus R (1950) Trematodes récoltés au Congo Belge par le Professeur Paul Brien:(Mai-août 1937). Ann Mus Congo Belge Zool 1:1–136
Edgar RC (2004) MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 32(5):1792–1797
Elsey RM (2005) Unusual offshore occurrence of an American alligator. Southeast Nat 4:533–536
Feizullaev NA, Mirzoeva SS (1983) Revision of the superfamily Clinostomoidea and analysis of its system. Parazitol 17(1):3–11
Feizullaev NA, Mirzoeva SS (1986) Keys to the taxa of the superfamily Clinostomoidea (Trematoda). Parazitol 20(3):195–201
Fernández MV, Hamann MI, de Núñez MO (2016) New larval trematodes in Biomphalaria species (Planorbidae) from northeastern Argentina. Acta Parasitol 61(3):471–492
Gabrey SW (2010) Demographic and geographic variation in food habits of American alligators (Alligator mississippiensis) in Louisiana. Herpetol Conserv Biol 5(2):241–250
Gaglio G, Reina V, Caffara M, Gjurcevic E, Iaria C, Marino F (2016) Risk of introduction of Clinostomum coplanatum (Digenea: Clinostomidae) to Sicily through use of Cobitis bilineata (Canestrini, 1865) as live baits. Bull Eur Ass Fish Pathol 36(3):105–110
Galazzo DE, Dayanandan S, Marcogliese DJ, McLaughlin JD (2002) Molecular systematics of some North American species of Diplostomum (Digenea) based on rDNA-sequence data and comparisons with European congeners. Can J Zool 80:2207–2217
Gustinelli A, Caffara M, Florio D, Otachi EO, Wathuta EM (2010) First description of the adult stage of Clinostomum cutaneum Paperna, 1964 (Digenea: Clinostomidae) from Grey herons Ardea cinerea L. and a redescription of the metacercaria from the Nile tilapia Oreochromis niloticus niloticus (L.) in Kenya. Syst Parasitol 76(1):39–51
Harrah EC (1922) North American monostomes: primarily from fresh water hosts. Ill Biol Monogr 7(3):66
Hunter GW, Hunter WS (1934) The life cycle of the yellow grub of fish. J Parasitol 20:325
IUCN (2013) Lepisosteus platyrhincus. 10.2305/IUCN.UK.2013-1.RLTS.T202412A18234213.en. Accessed June 27, 2017
Kanev I, Radev V, Fried B (2002) Superfamily Clinostomidea Lühe, 1901 In: Gibson, Jones, Bray (eds) Keys to the Trematoda volume 1, 1st edn. CABI Publishing, New York, pp 111–120
Katoh K, Standley DM (2013) MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol Bio Evol 30:772–780
Kearse M, Moir R, Wilson A, Stones-Havas S, Cheung M, Sturrock S, Buxton S, Cooper A, Mrkowitz S, Duran C, Theirer T, Ashton B, Mentjies P, Drummon A (2012) Geneious basic: an integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinform 28:1648–1649
Kostadinova A, Herniou EA, Barret J, Littlewood DTJ (2003) Phylogenetic relationships of Echinostoma Rudolphi, 1809 (Digenea: Echinostomatidae) and related genera re-assessed via DNA and morphological analysis. Syst Parasitol 54(3):159–176
Kudlai O, Kostadinova A, Pulis EE, Tkach VV (2015) A new species of Drepanocephalus Dietz, 1909 (Digenea: Echinostomatidae) from the double-crested cormorant Phalacrocorax auritus (Lesson) (Aves: Phalacrocoracidae) in North America. Syst Parasitol 90(3):221–230
Kumar S, Stecher G, Tamura K (2016) MEGA7: Molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33:1870–1874
Landan G, Graur D (2008) Local reliability measures from sets of co-optimal multipole sequence alignments. Pac Symp Biocomput 13:15–24
Leidy J (1856) A synopsis of entozoa and some other ecto-congeners observed by the author. Proc Acad Nat Sci Phil 8:42–58
Leidy (1884) Distoma and filariae. Proc Acad Nat Sci Philadelphia 36:47–48
Leidy (1890) Notices of entozoa. Proc Acad Nat Sci Philadelphia 42:410–418
Leigh WH (1960) The Florida spotted gar, as the intermediate host for Odhneriotrema incommodum (Leidy, 1856) from Alligator mississippiensis. J Parasitol 46(5 Supplement 2):16
Leigh WH (1963) An unusual host-parasite relationship between Alligator mississippiensis and the trematode Odhneriotrema incommodum (Leidy, 1856). J Parasitol 49(5s2):40
Leigh WH (1978) Studies on Odhneriotrema incommodum (Leidy 1856) (Trematoda: Clinostomidae) from Alligator mississippiensis. J Parasitol 64(5):831–834
Littlewood DTJ, Curini-Galletti M, Herniou EA (2000) The interrelationships of Proseriata (Platyleminthes: Seriata) tested with molecules and morphology. Mol Phylogenet and Evol 16:449–466
Locke SA, McLaughlin JD, Dayanandan S, Marcogliese DJ (2010) Diversity and specificity in Diplostomum spp. metacercariae in freshwater fishes revealed by cytochrome c oxidase I and internal transcribed spacer sequences. Int J Parasitol 40(3):333–343
Locke SA, McLaughlin JD, Lapierre AR, Johnson PT, Marcogliese DJ (2011) Linking larvae and adults of Apharyngostrigea cornu, Hysteromorpha triloba, and Alaria mustelae (Diplostomoidea: Digenea) using molecular data. J Parasitol 97(5):846–851
Locke SA, Caffara M, Marcogliese DJ, Fioravanti ML (2015) A large-scale molecular survey of Clinostomum (Digenea, Clinostomidae). Zool Scr 44(2):203–217
Lockyer AE, Olson PD, Østergaard P, Rollinson D, Johnston DA, Attwood SW, Southgate VR, Horak P, Snyder SD, Le TH, Agatsuma T, McManus DP, Carmichael AC, Naem S, Littlewood DTJ (2002) The phylogeny of the Schistosomatidae based on three genes with emphasis on the interrelationships of Schistosoma Weinland, 1858. Parasitology 126:203–224
Mazotti FJ, Best GR, Brandt LA, Cherkiss MS, Jeffrey BM, Rice KG (2009) Alligators and crocodiles as indicators for restoration of Everglades ecosystems. Ecol Indic 9:S137–S149
Mazzotti FJ, Brandt LA (1994) Ecology of the American alligator in a seasonally fluctuating environment. In: Davis SM, Ogden JC (eds) Everglades: the ecosystem and its restoration, 1st edn. St. Lucie Press, Delray Beach, FL, pp 485–505
McIntosh A (1935) Odhneriotrema incommodum (Leidy, 1856), a trematode from the mouth of Alligator mississippiensis (Daudin). J Parasitol 21:53–55
McNease L, Joanen T (1977) Alligator diets in relation to marsh salinity. Proc Annu Conf S.E Assoc Game and Fish Comm 31:36–40
Minh BQ, Nguyen MAT, von Haeseler A (2013) Ultrafast approximation for phylogenetic bootstrap. Mol Biol Evol 30:1188–1195
Morgan JAT, Blair D (1995) Nuclear rDNA ITS sequence variation in the trematode genus Echinostoma: an aid to establishing relationships within the 37-collar-spine group. Parasitology 111:609–615
Morgan JAT, Blair D (1998) Relative merits of nuclear ribosomal internal transcribed spacers and mitochondrial CO1 and ND1 genes for distinguishing among Echinostoma species (Trematoda). Mol Ecol 9(s1):75–82
Nguyen LT, Schmidt HA, von Haeseler A, Minh BQ (2015) IQ-TREE: a fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol Biol Evol 32(1):268–274
Overstreet RM, Self JT, Vliet KA (1985) The pentastomid Sebekia mississippiensis sp. n. in the American alligator and other hosts. Proc Helminthol Soc Wash 52(2):266–277
Pérez-Ponce de León G, García-Varela M, Pinacho-Pinacho CD, Sereno-Uribe AL, Poulin R (2016) Species delimitation in trematodes using DNA sequences: Middle-American Clinostomum as a case study. Parasitology 143(13):1773–1789
Pinto HA, Caffara M, Fioravanti ML, Melo AL (2015) Experimental and molecular study of cercariae of Clinostomum sp. (Trematoda: Clinostomidae) from Biomphalaria spp. (Mollusca: Planorbidae) in Brazil. J Parasitol 101(1):108–113
Pointier JP, David P, Jarne P (2005) Biological invasions: the case of planorbid snails. J Helminthol 79(3):249–256
Pratt HS (1902) Synopsis of North American invertebrates. XII. The Trematodes, Part 2. Am Nat 36:887–910
Rambaut A (2016) FigTree http://tree.bio.ed.ac.uk/software/figtree/. Accessed 19 June 2017
Ronquist F, Huelsenbeck JP (2003) MRBAYES 3: Bayesian phylogenetic inference under mixed models. Bioinform 19:1572–1574
Rosenblatt AE, Heithaus MR (2011) Does variation in movement tactics and trophic interactions among American alligators create habitat linkages? J Anim Ecol 80:786–798
Ross ST (2002) Inland fishes of Mississippi. University Press of Mississippi, Mississippi
Rosser TG, Alberson NR, Khoo LH, Woodyard ET, Pote LM, Griffin MJ (2016) Characterization of the life cycle of a fish eye fluke, Austrodiplostomum ostrowskiae (Digenea: Diplostomidae), with notes on two other diplostomids infecting Biomphalaria havanensis (Mollusca: Planorbidae) from catfish aquaculture ponds in Mississippi, USA. J Parasitol 102:260–274
Rosser TG, Alberson NR, Woodyard ET, Cunningham FL, Pote LM, Griffin MJ (2017) Clinostomum album n. sp. and Clinostomum marginatum (Rudolphi, 1819), parasites of the great egret Ardea alba L. from Mississippi USA. Syst Parasitol 94(1):35–49
Rózsa L, Reiczigel J, Majoros G (2000) Quantifying parasites in samples of hosts. J Parasitol 86:228–232
Sela I, Ashkenazy H, Katoh K, Pupko T (2015) GUIDANCE2: accurate detection of unreliable alignment regions accounting for the uncertainty of multiple parameters. Nucleic Acids Res 43:W7–W14
Senapin S, Phiwsaiya K, Laosinchai P, Kowasupat C, Ruenwongsa P, Panijpan B (2014) Phylogenetic analysis of parasitic trematodes of the genus Euclinostomum found in Trichopsis and betta fish. J Parasitol 100(3):368–371
Sereno-Uribe AL, Pinacho-Pinacho CD, García-Varela M, de León GP (2013) Using mitochondrial and ribosomal DNA sequences to test the taxonomic validity of Clinostomum complanatum Rudolphi, 1814 in fish-eating birds and freshwater fishes in Mexico, with the description of a new species. Parasitol Res 112(8):2855–2870
Skrjabin KI (1947) Семейство Clinostomatidae Lühe, 1901 In: Трематоды Животных и Человека: Основый Трематодологии Издательство, Tom 1, 1st edn. Академии Наук СССР, Москва [Family Clinostomatidae Lühe, 1901 In: Trematodes of Animals and Man: Basic Trematodology. vol 1, 1st edn. Academy of Sciences of the USSR, Moscow] pp 64-97
Steenkiste NV, Locke SA, Castelin M, Marcogliese DJ, Abbott CL (2015) New primers for DNA barcoding of digeneans and cestodes (Platyhelminthes). Mol Ecol Resour 15:945–952
Tavassos L, de Freitas JFT, Kohn A (1969) Trematódeos do Brasil. Mem Inst Oswaldo Cruz 67:1–886
Tellez CM (2014) Alligator parasitism—the mysterious frontier unfolded: exploration of the ecological interaction between an archaic predator (Alligator mississippiensis) and its parasites. Ph.D. dissertation, University of California, Los Angeles
Tellez CM, Nifong J (2014) Gastric nematode diversity between estuarine and inland freshwater populations of the American alligator (Alligator mississippiensis, Daudin 1802), and the prediction of intermediate hosts. Int J Parasitol Parasites Wildl 3:227–235
Tkach VV, Littlewood DTJ, Olson PD, Kinsella JM, Zdzislaw S (2003) Molecular phylogenetic analysis of the MicrophalloideaWard, 1901 (Trematoda: Digenea). Parasitology 56:1–15
Travassos L (1922) Informações sobre a fauna helmithologica de Matto Grosso. Fòlha Medica 3:187–190
Travassos L (1928) Fauna helmithologica de Matto Grosso. Mem Inst Oswaldo Cruz 21:309–372
Trifinopoulos J, Nguyen LT, von Haeseler A, Minh BQ (2016) W-IQ-TREE: a fast online phylogenetic tool for maximum likelihood analysis. Nucleic Acids Res 44(W1):W232–W235
Ukoli FMA (1966) On Clinostomum tilapiae n. sp., and C. phalacrocoracis Dubois, 1931 from Ghana, and a discussion of the systematics of the genus Clinostomum Leidy, 1856. J Helminthol 40(1/2):187–214
Ward HB, Whipple GC (1918) Freshwater biology. John Wiley and Sons, New York
Yamaguti S (1958) Part IV: Digenea of Birds In: Systema Helmenthum, vol 1. John Wiley and Sons, New York
Yamaguti S (1971) Synopsis of digenetic trematodes of vertebrates, vol 1. Keigaku Publishing Co, Tokyo, Japan
Acknowledgements
This work was supported in part by the Mississippi State University College of Veterinary Medicine and the Mississippi State University College of Forest Resources. The authors are deeply indebted to Mr. Woodrow Cain and Mrs. Vecie Cain for providing us with host specimens. This work would not have been possible without their support. We would also like to acknowledge Dr. Eric Pulis of the Institute for Marine Mammal Studies, Gulfport, Mississippi for his insights regarding the mounting of stained trematode specimens as well as Dr. George Pinchuk of the Department of Sciences & Mathematics, Mississippi University for Women and Dr. Lesya Pinchuk of the Department of Basic Sciences, College of Veterinary Medicine, Mississippi State University, for their translations of Russian texts. The Thad Cochran National Warmwater Aquaculture Center provided material support through use of their microscope and associated imaging equipment.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Supplemental Table 1
(XLSX 49 kb)
Supplemental Table 2
(XLSX 50 kb)
Rights and permissions
About this article
Cite this article
Woodyard, E.T., Rosser, T.G. & Rush, S.A. Alligator wrestling: morphological, molecular, and phylogenetic data on Odhneriotrema incommodum (Leidy, 1856) (Digenea: Clinostomidae) from Alligator mississippiensis Daudin, 1801 in Mississippi, USA. Parasitol Res 116, 2981–2993 (2017). https://doi.org/10.1007/s00436-017-5607-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00436-017-5607-7