Abstract
MicroRNAs (miRNAs) are a class of small non-coding RNAs that function in transcriptional and post-transcriptional regulation of gene expression. An increasing number of schistosome miRNAs have been identified and are expected possibly involved in differentiation, development, and metabolism. However, limited information is available concerning the target genes of schistosome miRNAs. In the present study, the key target genes of bantam, an abundant miRNA found in paired female Schistosoma japonicum, were predicted by bioinformatics analysis and Solexa technology. Luciferase reporter assay and bantam mimic assay were applied in combination to further verify the targets of bantam. Results showed that ATP synthase (CAX76793.1), one of the three selected predicted targets, was confirmed as the target of bantam; bantam mimic assay results also showed that the two other predicted targets, namely, ataxia telangiectasia mutated (ATM)-related (XP_002571630.1), and ribosomal protein L30 (CAX72575.1), were not confirmed as targets. This research proposed the design and significance of reasonable biological experiments that could be performed to identify miRNA target genes in schistosomes.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Schistosomiasis is a chronic parasitic disease caused by blood flukes of the genus Schistosoma. More than 200 million people are infected, and close to 800 million individuals are at risk of acquiring the parasitic disease (Steinmann et al. 2006). Praziquantel, as the effective schistosomicide, is faced with the drug resistance (Cioli 2000). Thus, novel drug targets should be identified for new therapeutic strategies.
MicroRNAs (miRNAs), generated from endogenous transcripts that form hairpins, are a class of small non-coding RNAs (Kim 2005) that function in transcriptional and post-transcriptional regulation of gene expression (Bartel 2004; Chen and Rajewsky 2007). The binding of miRNA to a target mRNA leads to gene silencing via translational repression and target mRNA degradation (Bartel 2009). MiRNAs have been reportedly involved in translational activation (Filipowicz et al. 2008) and heterochromatin formation (Benetti et al. 2008); in this way, gene expression is regulated during differentiation, development, death, and metabolism of many organisms (Bushati and Cohen 2007). Thus far, numerous schistosome miRNAs have been identified in schistosomes (Cheng and Jin 2012; Hao et al. 2010; Wang et al. 2010; Xue et al. 2008; Huang et al. 2009; Marco et al. 2013; Simões et al. 2011) that provided insights into schistosome development. However, many studies on the target genes of these miRNAs are based on bioinformatics analysis; few targets have been confirmed by biological experiments.
In a previous study, the miRNA expression profiles of 23-day-old female schistosomula from double-sex infections (23DSI) and 23-day-old female schistosomula from single-sex infections (23SSI) were analyzed by Solexa sequencing (Sun et al. 2014a) . During Schistosoma japonicum development, a male and a female begin to pair at approximately 18 days post-infection; the female then lays eggs at approximately 24 days post-infection (He and Yang 1980). Thus, differentially expressed miRNAs of 23DSI can be compared with those of 23SSI to identify key miRNAs that play important roles in female development. The research has revealed that the bantam miRNA possibly functions in the development of female S. japonicum (Sun et al. 2014a). Thus far, limited information is known regarding the targets of the bantam miRNA in S. japonicum. Applying bioinformatics analysis, we obtained numerous target genes of bantam. However, actual biological functional analysis should be based on the genuine targets of bantam. Thus, the present study aimed to identify the target genes of bantam through biological experiments with bioinformatics analysis.
Methods
Ethics statement
This study was carried out in strict accordance with the recommendations of the Regulations for the Administration of Affairs Concerning Experimental Animals of the State Science and Technology Commission. The protocol was approved by the Internal Review Board of Tongji University School of Medicine (China).
Bioinformatics analysis of bantam
We used the algorithms PicTar (Kennell and MacDougald 2005) and TargetScan (Lewis et al. 2003) to predict the target mRNAs of bantam. All of the predicted target genes were evaluated using the scoring system and criteria described by Chi et al. (2011). Sequences with total scores of <3.0 were considered as potential targets. Using Venny (http://bioinfogp.cnb.csic.es/tools/venny/index.html), we obtained the predicted upregulated and downregulated target genes. Kegg pathway analysis of these genes was preformed to investigate the relationship between these genes and important metabolic or regulation pathways.
18- and 23-day-old S. japonicum samples
Oncomelania hupensis snails were obtained from the Jiangsu Institute of Schistosome Diseases, Jiangsu Province, China. Approximately 100 to 150 freshly shed cercariae were used to percutaneously infect each mouse. The mice were then sacrificed at 18 and 23 days post-infection, respectively. S. japonicum worms were recovered by washing with cold saline solution. Eighteen-day-old females were selected with syringe needle under a microscope. The 23-day-old females were carefully separated with syringe needle from the paired worms under a microscope. Some of them were cultured, and others were frozen until further processing.
Solexa analysis
Total RNA samples were extracted respectively from 18- to 23-day-old female schistosomula using Trizol reagent (Invitrogen Life Technologies) according to the manufacturer’s instructions. RNA concentration and purity were evaluated spectrophotometrically at 260 and 280 nm, respectively, by using a NanoDrop ND1000 spectrophotometer (Thermo, USA) and an Agilent 2100 Bioanalyzer (Agilent Technologies, Palo Alto, CA). Tag-seq library was constructed; data were processed; and differentially expressed genes were identified in accordance with previously described methods (Sun et al. 2014b).
In vitro assay of the effect of bantam-mimic on gene expression
RPMI-1640 (Gibco, USA) containing 10 % calf serum, 100 IU/ml penicillin sodium, 100 IU/ml streptomycin, and 0.25 μg/ml amphotericin B (Hyclone, USA) was used to maintain 18- and 23-day-old schistosomes in vitro. The worms were cultured in a 16-well plate containing 2.5 ml of culture medium. A total of 10 females and 12 males were placed in each well. One group comprised four wells. The 5OD of inhibitor control (5′-CAGUACUUUUGUGUAGUACAA-3′), 5OD of bantam mimic (sense: 5′-UGAGAUCGCGAUUAAAGCUGGU-3′; anti-sense: 5′-CAGCUUUAAUCGCGAUCUCAUU-3′), the 5OD of bantam inhibitor (5′-ACCAGCUUUAAUCGCGAUCUCA-3′), and the 5OD of the negative control (sense: 5′-UUCUCCGAACGUGUCACGUTT-3′; anti-sense: 5′-ACGUGACACGUUCGGAGAATT-3′) were added to each well. The plates were incubated at 37 °C in 5 % CO2. The media and agents were replaced with fresh supply daily.
Total RNA was extracted using Trizol reagent according to the manufacturer’s instructions. RNA concentration and purity were respectively evaluated spectrophotometrically at 260 and 280 nm by using a NanoDrop ND1000 spectrophotometer and an Agilent 2100 Bioanalyzer. RNA samples were then stored at −80 °C. Several predicted genes were selected and validated by quantitative RT-PCR in accordance with previously described methods (Sun et al. 2014b). The following primers of the predicted target genes were designed: ATP synthase (CAX76793.1) forward, 5′-GCTACGGCACTGCTCTTTCT-3′, reverse, 5′-TCCACTAAACCCCACGCTTA-3′; major egg antigen (p40; CAX78215.1) forward, 5′-GCGAAGAAGGAGACGACAAC-3′, reverse, 5′-CGATGCAGGAACTGCTTGTA-3′; ataxia-telangiectasia mutated-related gene (ATM; XP_002571630.1) forward, 5′-GGGCTTTAGGTCAATGGGCT-3′, reverse, 5′-AACGAGACTGCCATGCTTCA-3′; and ribosomal protein L30 gene (CAX72575.1) forward, 5′-GCTACGACAAACGCTTCGAC-3′, reverse, 5′-TGAAGTACTTGCCACACGCA-3′. These predicted genes were amplified using SYBR® Premix Ex Taq™ II (Perfect Real Time; Takara Code: DRR081A) in an ABI Prism 7300 sequence detection system (Applied Biosystems). The expression levels of S. japonicum GAPDH were used as endogenous controls in each sample. Relative gene expression levels were calculated using the 2-ΔΔCT method (Livak and Schmittgen 2001).
Luciferase reporter assay
To produce a reporter construct, we synthesized the 3′ untranslated (UTR) of ATP synthase (CAX76793.1) with 322 nucleotides in length (TCTAGATGCATAATATTTATTCCTACATAATACCTTGAGGTTCTTTGACATATTTTCTTTAAGATCAAATGATTAGTACTTCATTCAAGTTGTTGGATTTTTGTAATAGCTTGTATTTCCAATTACAATATCTGATTATCAAAATAACTTGTTCCTTGAAGAATGTATTTCAGCTTATCACTGATTATCATGCATAGTATGGATGAGAACTATTCACATCCGCTTTTTCTTATATAGTTAAGAATTTGTCTTATGTTTAATATTCTTAATCTAGCCGCGTGATTACCGTTTGATTGGTTGGCTCATTGGGTGTTCGTCTAGA). The products were digested with Xbal and then cloned into a GV272 luciferase vector (http://www.genechem.com.cn/Zaiti.aspx?zt=GV272). Sja-bantam precursor was chemically synthesized, digested with Xhol/KpnI, and cloned into a GV268 vector (http://www.genechem.com.cn/Zaiti.aspx?zt=GV268). The two constructs were verified by PCR and sequencing using the two pairs of primers listed respectively as follows: Luc-C-F (GAGGAGTTGTGTTTGTGGAC)/RVprimer4 (GACGATAGTCATGCCCCGCG) and sja-bantam-P1 (TCCGCTCGAGAAGTCGGCTTTTATTGCGCTCTGAGAAAAACCAATTCATTGCTT)/sja-bantam-P2 (ATGGGGTACCAAACCAGCTTTAATCGCGATCTCAGATAAATAAAAAAAGCAATGAATTGGTTTTTC). Afterward, luciferase reporter constructs, together with a bantam construct or a negative control, were transfected into HEK 293T cells. Luciferase activity was detected using a dual luciferase assay system (Promega) at 48 h after transfection in accordance with the manufacturer’s instructions. The system contained 0.2 μg of luciferase plasmid, 0.6 μg of miRNA plasmid (control or bantam), and 0.05 μg of renilla plasmid. The red firefly luciferase signal was used as a normalization control of the green Renilla luciferase signal. The experiments were repeated at least three times.
Statistical analysis
Results were presented as mean ± standard deviation from at least three independent experiments. Statistical analyses were performed using one-way ANOVA and Student’s t test. A P value of <0.05 was considered statistically significant.
Results
Bioinformatics analysis of the target genes of bantam
The differentially expressed genes between 23DSI and 23SSI were analyzed using the Solexa method, and the target genes of bantam were predicted using PicTar and TargetScan algorithms. Then, the differentially expressed target genes of bantam between 23DSI and 23SSI were obtained by analyzing the common genes between the two data sets. Two hundred and thirty-two upregulated genes (Supplementary Table S1) and 1619 downregulated genes (Supplementary Table S2) were predicted as the target genes of bantam in 23DSI compared with 23SSI. Pathway analysis revealed that some pathways, such as protein processing in endoplasmic reticulum, oxidative phosphorylation, mTOR signaling pathway, and ribosome, played a vital role in the development of 23DSI (data not shown). On the basis of these results, we selected three predicted target genes [ATP synthase (CAX76793.1), ATM (XP_002571630.1), and ribosomal protein L30 gene (CAX72575.1)] and one predicted non-target gene [major egg antigen (p40 CAX78215.1)] from these genes which participated in these key pathways, to further investigate the target genes by conducting biological experiments (Table 1). In particular, most of predicted target genes did not contain their 3′ UTR in the database (http://www.chgc.sh.cn/japonicum/Resources.html), which greatly limited our choice in this study. The selected predicted genes come from different groups. ATM, which participated in mTOR signaling pathway, was downregulated in 23DSI, whereas ATP synthase and ribosomal protein L30 gene, which participated in oxidative phosphorylation and ribosome pathway respectively, were upregulated in 23DSI compared with those of 23SSI.
Bantam is well conserved in different organisms. The two conserved regions of bantam are shown in Fig. 1a. Based on the bantam sequence, the binding regions of bantam were detected in the 3′ UTR of ATP synthase (Fig. 1b). Only short 3′ UTR of ATM gene and ribosomal protein L30 gene were found in the database. The binding regions of bantam were not detected in their 3′ UTR but within the genes (Fig. 1c, d). No similar regions were found in major egg antigen gene or its flanking regions.
Alignment analysis of bantam and its binding sites in ATP synthase, ATM, and ribosomal protein L30 gene or their corresponding 3′ UTRs. a Alignment analysis of bantam using Clustal Omega program (www.ebi.ac.uk/Tools/msa/clustalo/). Analysis of the binding regions of bantam in ATP synthase 3′ UTRs (b), ATM genes (c), and ribosomal protein L30 gene (d)
In vitro assay of the effects of bantam-mimic on gene expression
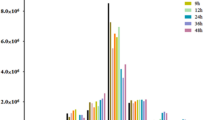
The mimic of bantam is added to cultures of 18- or 23-day-old S. japonicum to investigate the effects of bantam on the expression of the four genes. The results showed that the expression levels of ATP synthase were significantly downregulated, compared with those of the negative control group. Moreover, the bantam mimic inhibited the expression of ATP synthase; the bantam inhibitor simultaneously upregulated the expression of ATP synthase. By contrast, ATM expressions were not significantly changed by the bantam mimic and bantam inhibitor. For the other predicted target genes, the expression of the ribosomal protein L30 was not inhibited by the bantam mimic; by contrast, the addition of the bantam inhibitor could significantly downregulate the expression of ribosomal protein L30(Fig. 2). For the predicted non-target gene (major egg antigen), its expression was not inhibited by bantam mimic, and this result is consistent with our expected.
Effect of bantam mimic on the expressions of predicted target and non-target genes in vitro. a Amount of exclusive and common genes between upregulated and downregulated genes in 23DSI and predicted target genes of bantam in S. japonicum. UP upregulated genes in 23DSI compared with 23SSI, Down downregulated genes in 23DSI compared with 23SSI, Bantam-tar all of the predicted target genes of bantam in S. japonicum. Venn diagram was created using an online Venn diagram maker. Effect of bantam mimic on b ATM, c ATP synthase, d ribosomal protein L30, and e major egg antigen (p40)
Luciferase reporter assay
ATP synthase was selected to further verify whether or not bantam regulates the expression of this gene by binding to the complementary region in the 3′ UTR of ATP synthase. In the luciferase reporter assay, significant changes in luciferase activity were found in bantam-overexpressed plasmid plus luciferase-ATP synthase 3′ UTR plasmid group. By contrast, no control plasmids exhibited the same changes (Fig. 3).
Construct of luciferase reporter vector and effect of bantam on luciferase activity. a Schematic of luciferase reporter construct. 3′ UTR of ATP synthase was cloned into the luciferase vector. b Schematic of bantam-overexpressed construct. c Luciferase activity was detected after the bantam mimic was used in vitro. Test1, miRNA control plasmid + luciferase control plasmid; Test2, bantam-overexpressed plasmid + luciferase control plasmid; Test3, miRNA control plasmid + luciferase-ATP synthase 3′ UTR plasmid; Test4, bantam overexpressed plasmid + luciferase-ATP synthase 3′ UTR plasmid
Discussion
Thus far, 78 mature miRNAs have been identified in S. japonicum genomes and 29 in S. mansoni were deposited in the miRBase database (http://www.mirbase.org/cgi-bin/browse.pl?org=cel) (Hao et al. 2010; Liu et al. 2010; Wang et al. 2010; Xue et al. 2008; Copeland et al. 2009; Huang et al. 2009). MiRNAs are implicated in various biological processes in organisms (Bushati and Cohen 2007). Hence, miRNA target genes should be identified to provide a basis for understanding the function of miRNAs in biological processes, such as growth, development, and sexual maturation of S. japonicum. At present, many biological functions and significance of miRNAs in schistosomes are based on the roles of predicted target genes rather than actual target genes. Some highly conserved miRNAs, such as miR-1, miR-124, and miR-7, have been predicted to interact with phylogenetically conserved mRNA targets in schistosomes based on the analysis of miRNA/mRNA pairs predicted using RNAhybrid (http://bibiserv.techfak.uni-bielefeld.de/rnahybrid/) and PITA (http://genie.weizmann.ac.il/pubs/mir07/mir07_prediction.html). Based on predicted potential targets, signalling pathways, such as Wnt, VEGF, and TGF-β, and other important biological processes, including cell cycle regulation in S. japonicum, are also possibly regulated by these conserved miRNAs (Cheng and Jin 2012). Although predicted target genes may provide relevant data of miRNA functions, the accurate biological significance of miRNA should depend on the understanding of the functions of genuine target genes. In particular, during prediction analysis, a large number of false positive or negative results could provide misleading information and could pose the risk of losing vital information. Thus, identification of actual miRNA targets should be paid more attention in schistosomes.
During the development of S. japonicum, pairing facilitates female development, thereby leading to female sexual maturation. Previous studies revealed that bantam is significantly upregulated in 23DSI after pairing occurs, suggesting that bantam may play an important role in regulating development and promoting sexual maturation (Sun et al. 2014a). Therefore, bantam targets should be identified to elucidate the mechanisms by which bantam regulates female development. Bioinformatics analysis results have indicated that approximately 5933 genes are predicted as targets of bantam (Sun et al. 2014a). To identify significant targets of bantam in females after pairing occurs, we compared the differentially expressed genes between 23DSI and 23SSI. These differentially expressed genes were considered more responsible for female development than non-differentially expressed genes at this stage. It is obvious that the common genes between differentially expressed genes and all of the predicted bantam target genes can facilitate the identification of targets. Since bantam was expressed in a higher level in 23DSI than in 23SSI, the most possible targets of bantam should be the genes that are common between downregulated genes in 23DSI and predicted bantam target genes. Considering the complex regulatory mechanisms of miRNAs on target genes, we selected the following genes for further analysis: ATM from downregulated predicted targets, ATP synthase, ribosomal protein L30 gene from upregulated predicted targets, and major egg antigen (p40) from non-predicted targets.
The results indicated that the bantam mimic downregulated ATP synthase expression in 18- and 23-day-old schistosome cultures in vitro; by contrast, the expressions of ribosomal protein L30 gene and major egg antigen were inhibited not by the bantam mimic but by the bantam inhibitor. The expression of major egg antigen was not inhibited by the bantam mimic because this gene is not a predicted target of bantam. Although the ribosomal protein L30 gene was predicted as a target of bantam, the expression of this gene was not inhibited by bantam mimic, suggesting that the ribosomal protein L30 gene was not a target of bantam. Likewise, ATM, from the downregulated predicted target group, was not regulated by bantam mimic. On the one hand, these results verified that the prediction of target genes sometimes lead to false positive results. On the other hand, these results indicated that the miRNA mimic assay was an effective experimental verification method to reduce false positives. To further determine whether or not bantam regulates ATP synthase expression by binding to the 3′ UTR, we designed the luciferase reporter assay. Our findings agree with the expected results in which bantam could inhibit ATP synthase gene expression by binding to the 3′ UTR.
In addition, this study provided an effective method to identify target genes in schistosomes. Although bioinformatics analysis may produce false positive results, this method can be used to eliminate a large number of non-target genes. For example, major egg antigen was verified as a non-target gene, as indicated by the results predicted by bioinformatics analysis. Hence, bioinformatics analysis still remains a reliable method. The results of bioinformatics analysis and biological experimentation should be combined to identify actual target genes. Furthermore, rational combination can help us to elucidate the interaction between miRNAs and corresponding potential target genes. For example, gene expression profiles can facilitate the selection of some key genes at a particular development stage. These identified key genes acting as target genes may be more helpful to understand miRNA functions. In particular, a gene often contains various miRNA binding regions, indicating that it is regulated by various miRNAs. Thus, its expression is undoubtedly determined by the effects of miRNAs. However, during identification of target genes, only considering whether or not a miRNA can bind to its binding region of a gene is not enough. Although a miRNA can bind to a corresponding region of a gene and the gene expression can be regulated by it in luciferase reporter assay, the gene may not be the most appropriate “target” if this miRNA does not play a key role in regulating the gene expression at a certain time.
Actually, it is interesting how to define a significant target of a miRNA. A gene usually contains binding regions of various miRNAs or contains multiple binding sites of a miRNA. Each miRNA can potentially contribute to the regulation of gene expression. Some miRNAs may exhibit major implications in gene expression regulation whereas others play minor functions. If a gene is only considered as a target of the miRNAs with critical functions, then the function of these miRNAs should be more clearly understood. However, if a gene is considered as a target of all relevant miRNAs, then this target gene may be not very helpful to reveal the main function of miRNAs. For example, bantam and miR-1 likely regulate opposite developmental processes in paired female S. japonicum (Sun et al. 2014a). Nevertheless, bantam and miR-1 were found to share some same target genes (data not shown), thereby suggesting that different miRNAs exhibit different functions in the same target. In addition, various miRNAs usually exhibit temporal functions in the regulation of gene expression in different tissues, development stages or physiological conditions. Thus, a target containing various miRNA binding sites is not necessarily and simultaneously the target of all these miRNAs. In the present study, ATP synthase was confirmed as a target of bantam, but ATP synthase expression was still upregulated in bantam-abundant paired females (23DSI) from hosts compared with 23SSI. In the bantam mimic assay, the expression of ATP synthase was downregulated. These contradicting results suggested peculiarities in miRNA regulation. Hence, complex relationships among target genes and their corresponding miRNAs should be understood to elucidate the function of miRNAs. In particular, in this research, the pairing provides two different types of development condition in schistosome, which is helpful to recognize the alterable regulation of miRNA on its targets. At present, most studies focus on how to predict miRNA targets (Peterson et al. 2014; Ritchie and Rasko 2014; Zheng et al. 2013) rather than how to understand the alterable relationship between targets and their miRNAs. Furthermore, a target should be given a new understanding, that is, a significant target of a miRNA may belong to a limited space and time.
In summary, the target genes of miRNAs could be determined on the basis of multiple factors. A rational experimental assay design could be used to identify actual target genes from many targets predicted by bioinformatics analysis. In addition, the significant relationship between targets and their miRNAs should be paid an additional attention.
References
Bartel DP (2004) MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116:281–297
Bartel DP (2009) MicroRNAs: target recognition and regulatory functions. Cell 136:215–233
Benetti R, Gonzalo S, Jaco I, Munoz P, Gonzalez S, Schoeftner S, Murchison E, Andl T, Chen T, Klatt P, Li E, Serrano M, Millar S, Hannon G, Blasco MA (2008) A mammalian microRNA cluster controls DNA methylation and telomere recombination via Rbl2-dependent regulation of DNA methyltransferases. Nat Struct Mol Biol 15:998
Bushati N, Cohen SM (2007) MicroRNA functions. Annu Rev Cell Dev Biol 23:175–205
Chen K, Rajewsky N (2007) The evolution of gene regulation by transcription factors and microRNAs. Nat Rev Genet 8:93–103
Cheng G, Jin Y (2012) MicroRNAs: potentially important regulators for schistosome development and therapeutic targets against schistosomiasis. Parasitology 139:669–679
Chi X, Yang Q, Chen X, Wang J, Pan L, Chen M, Yang Z, He Y, Liang X, Yu S (2011) Identification and characterization of microRNAs from peanut (Arachis hypogaea L.) by high-throughput sequencing. PLoS One 6:e27530
Cioli D (2000) Praziquantel: is there real resistance and are there alternatives? Curr Opin Infect Dis 13:659–663
Copeland CS, Marz M, Rose D, Hertel J, Brindley PJ, Santana CB, Kehr S, Attolini CS, Stadler PF (2009) Homology-based annotation of non-coding RNAs in the genomes of Schistosoma mansoni and Schistosoma japonicum. BMC Genomics 10:464
Filipowicz W, Bhattacharyya SN, Sonenberg N (2008) Mechanisms of post-transcriptional regulation by microRNAs: are the answers in sight? Nat Rev Genet 9:102–114
Hao L, Cai P, Jiang N, Wang H, Chen Q (2010) Identification and characterization of microRNAs and endogenous siRNAs in Schistosoma japonicum. BMC Genomics 11:55
He YX, Yang HZ (1980) Physiological studies on the post-cercarial development of Schistosoma japonicum. Acta Zool Sin 26:32–41
Huang J, Hao P, Chen H, Hu W, Yan Q, Liu F, Han ZG (2009) Genome-wide identification of Schistosoma japonicum microRNAs using a deep-sequencing approach. PLoS One 4:e8206
Kennell JA, MacDougald OA (2005) Wnt signaling inhibits adipogenesis through beta-catenin-dependent and -independent mechanisms. J Biol Chem 280:24004–24010
Kim VN (2005) MicroRNA biogenesis: coordinated cropping and dicing. Nat Rev 6:376–385
Lewis BP, Shih IH, Jones-Rhoades MW, Bartel DP, Burge CB (2003) Prediction of mammalian microRNA targets. Cell 115:787–798
Liu Q, Tuo W, Gao H, Zhu XQ (2010) MicroRNAs of parasites: current status and future perspectives. Parasitol Res 107:501–507
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(−Delta Delta C(T)) Method. Methods 25:402–408
Marco A, Kozomara A, Hui JH, Emery AM, Rollinson D, Griffiths-Jones S, Ronshaugen M (2013) Sex-biased expression of microRNAs in Schistosoma mansoni. PLoS Negl Trop Dis 7:e2402
Peterson SM, Thompson JA, Ufkin ML, Sathyanarayana P, Liaw L, Congdon CB (2014) Common features of microRNA target prediction tools. Front Genet 5:23
Ritchie W, Rasko JE (2014) Refining microRNA target predictions: sorting the wheat from the chaff. Biochem Biophys Res Commun 445:780–4
Simões MC, Lee J, Djikeng A, Cerqueira GC, Zerlotini A, da Silva-Pereira RA, Dalby AR, LoVerde P, El-Sayed NM, Oliveira G (2011) Identification of Schistosoma mansoni microRNAs. BMC Genomics 12:47
Steinmann P, Keiser J, Bos R, Tanner M, Utzinger J (2006) Schistosomiasis and water resources development: systematic review, meta-analysis, and estimates of people at risk. Lancet Infect Dis 6:411–425
Sun J, Wang S, Li C, Ren Y, Wang J (2014a) Novel expression profiles of microRNAs suggest that specific miRNAs regulate gene expression for the sexual maturation of female Schistosoma japonicum after pairing. Parasite Vector 7:177
Sun J, Wang SW, Li C, Hu W, Ren YJ, Wang JQ (2014b) Transcriptome profilings of female Schistosoma japonicum reveal significant differential expression of genes after pairing. Parasitol Res 113:881–892
Wang Z, Xue X, Sun J, Luo R, Xu X, Jiang Y, Zhang Q, Pan W (2010) An "in-depth" description of the small non-coding RNA population of Schistosoma japonicum schistosomulum. PLoS Negl Trop Dis 4:e596
Xue X, Sun J, Zhang Q, Wang Z, Huang Y, Pan W (2008) Identification and characterization of novel microRNAs from Schistosoma japonicum. PLoS One 3:e4034
Zheng H, Fu R, Wang JT, Liu Q, Chen H, Jiang SW (2013) Advances in the Techniques for the Prediction of microRNA Targets. Int J Mol Sci 14:8179–87
Acknowledgments
We would like to thank Pan W. Q. for his valuable advice. We also appreciate the discussions and comments from Xiao S. H., Zhang Q. F., and Xu X. D. We thank Shenzhen Huada Gene Research Institute for Solexa analysis. This research was supported by the National Natural Science Foundation of China (Grant No. 81071383).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Sun, J., Wang, SW. & Li, C. ATP synthase: an identified target gene of bantam in paired female Schistosoma japonicum . Parasitol Res 114, 593–600 (2015). https://doi.org/10.1007/s00436-014-4221-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00436-014-4221-1