Abstract
We have analyzed brain structure in Macrostomum lignano, a representative of the basal platyhelminth taxon Macrostomida. Using confocal microscopy and digital 3D modeling software on specimens labeled with general markers for neurons (tyrTub), muscles (phalloidin), and nuclei (Sytox), an atlas and digital model of the juvenile Macrostomum brain was generated. The brain forms a ganglion with a central neuropile surrounded by a cortex of neuronal cell bodies. The neuropile contains a stereotypical array of compact axon bundles, as well as branched terminal axons and dendrites. Muscle fibers penetrate the flatworm brain horizontally and vertically at invariant positions. Beside the invariant pattern of neurite bundles, these “cerebral muscles” represent a convenient system of landmarks that help define discrete compartments in the juvenile brain. Commissural axon bundles define a dorsal and ventro-medial neuropile compartment, respectively. Longitudinal axons that enter the neuropile through an invariant set of anterior and posterior nerve roots define a ventro-basal and a central medial compartment in the neuropile. Flanking these “fibrous” compartments are neuropile domains that lack thick axon bundles and are composed of short collaterals and terminal arborizations of neurites. Two populations of neurons, visualized by antibodies against FMRFamide and serotonin, respectively, were mapped relative to compartment boundaries. This study will aid in the documentation and interpretation of patterns of gene expression, as well as functional studies, in the developing Macrostomum brain.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Brain development and morphology has been described in great detail in a few select model systems, ranging from nematodes and insects to vertebrates. Although the brains of these animals are morphologically quite different, recent genetic evidence suggests that most of the genetic mechanisms controlling brain topology are highly conserved (reviewed in Lichtneckert and Reichert 2005). For example, transcriptional regulators controlling cell identity, such as the genes of the Hox complex expressed in a defined sequence along the antero-posterior axis or the Homeobox containing genes vnd, ind, and msh that are expressed along the medio-lateral axis, are found in vertebrates and Drosophila alike (Cornell and Ohlen 2000). This conservation has led developmental biologists to revisit the evolution of the nervous system, searching for clues that would elucidate the structure of the brain of the last common bilaterian (LCB; Holland 2003). As we have no direct access to the ancestral organisms that lived during earlier phases of evolution, taking a comparative route and paying attention to seemingly “primitive” organisms that may have retained more of the original characters of the ancestors appears as the next best approach. In an attempt to shed some light on the origin and structure of the LCB brain, we have begun work on the basal flatworm, Macrostomum lignano.
Flatworms (Platyhelminthes) are traditionally considered to represent the most primitive animals with a true central nervous system (Bullock and Horridge 1965; Hanstroem 1968). Morphological studies at the light and electron microscopic level have concluded that all flatworms exhibit a number of plesiomorphic characters, which could be directly inherited from the bilaterian ancestor (Ax 1996). For example, in the present context, the fact that adult flatworms retain a ciliated epidermis and that movement is largely effected by ciliary beating is significant, as it is likely to have an impact on the morphology of neurons and the type of circuits controlling movement. Similarly, the presence of a diffuse nerve net spread out over the trunk body wall at all dorso-ventral levels, rather than only dorsally or ventrally as in other bilaterians, may represent a primitive character (Lowe et al. 2006). Other less well-discussed traits, such as the absence of morphologically detectable glial cells in basal flatworms (Radojcic and Pentreath 1979; Hartenstein and Ehlers 2000; Morris et al. 2004), should also be taken as a primitive trait. Recent molecular phylogenies are ambiguous as to the position of the phylum Platyhelminthes among other metazoans. It appears adequate for us to follow current reviews (Baguñà and Riutort 2004; Telford et al. 2005) in which platyhelminths, beside other spiralian phyla including annelids and molluscs are grouped side by side within the new superphylum “lophotrochozoa” with no further resolution as to which phylum is more basal.
General aspects of flatworm brain structure, including the shape of neurons and synapses, the distribution of histochemically detectable neuronal cell types, and the overall pattern of nerves connecting the brain to the periphery, have been described for a number of species (Lentz 1967; Keenan et al. 1981; Shaw 1981; Halton et al. 1992; Reuter and Gustafsson 1995; Gustafsson et al. 2002). All flatworms possess a relatively compact anterior brain organized in a way that resembles a typical invertebrate ganglion. Neuronal cell bodies form an external cell layer, the cortex, and processes of the predominantly unipolar neurons form a neuropile in the brain center. A system of nerves, called orthogon (due to its orthogonal arrangement of longitudinal connectives and circular commissures; Reisinger 1925 in Hanstroem 1968) extends from the brain into the trunk. Longitudinal nerves of the orthogon occur at all dorso-ventral levels, although in many species (including those of the genus Macrostomum), the ventral nerves are most prominent. The peripheral nervous system (PNS) of flatworms is formed by one or several subepidermal/submuscular plexus of nerve cell processes formed by the axons/dendrites of neurons whose cell bodies are located in the brain or along the orthogonal nerves. For example, profusely branched neurites of serotonergic neurons located in the brain form a subepidermal network throughout the body of Macrostomum (Ladurner et al. 1997) and other species (Gustafsson et al. 2002). Mechanosensory and chemosensory neurons are located either in the epidermis or the central nervous system (Ehlers 1985). The spatial relationship between motorneurons and muscles in flatworms is different from vertebrates or arthropods. In many cases, long muscle processes approach the neuron, instead of the other way around as found in vertebrates and arthropods (Chien and Koopowitz 1972; Cousin and Dorsey 1991; Biserova et al. 2000), although a similar condition to that in flatworms is found in nematodes. These muscle processes converge upon the longitudinal connectives that carry motor axons; in the head, muscle processes crisscross the brain neuropile.
In this paper, we present a detailed topological analysis of the juvenile brain of M. lignano. Macrostomid flatworms are the most basal flatworms with sexual reproduction (Ehlers 1985). A reliable laboratory culture has been set up for this species, and many molecular markers for the brain have been identified (Morris et al. 2004; Ladurner et al. 2005a, b; Egger et al. 2006). Using antibodies specific to the nervous system we developed a detailed atlas and a digital 3D model of the juvenile brain. A digital 3D model is a convenient way to store and use morphological as well as genetic data (Pereanu and Hartenstein 2004). This study will aid in the interpretation of gene-expression data and future functional studies.
Materials and methods
Macrostomum culture
M. lignano was cultured as described in Morris et al. (2004). Briefly, diatom plates containing Nizschia corvenculate are grown to confluences in modified F/2 media for roughly 10–14 days. One hundred fifty Macrostomum adults were collected and transferred to a confluent algal plate on a monthly basis where they deposited eggs. Freshly deposited eggs were collected on a daily basis and allowed to develop for 5 days, until hatching. Juvenile worms (within 24 h of hatching) were collected and treated as described below.
Immunohistochemistry
Freshly hatched juveniles were then collected and relaxed in an 8% MgCl2 solution, followed by fixation in a freshly prepared solution of 4% paraformaldehyde for 1 hour. The animals were then rinsed five times in phosphate buffered saline pH 7.2 plus 0.1% Tween 20, and the standard immunohistochemistry protocol was subsequently followed (Ashburner 1989). Anti-Tyrosinated tubulin (called tyrTub in the following; Sigma; 1:1,000 dilution) was used to label microtubules of axon tracts and cilia. Anti-FMRF (Sigma; 1:1,000 dilution) was used to label neurons expressing the neuropeptide FMRFamide. Anti-5HT (Sigma; 1:5,000 dilution) was used to label neurons expressing the neuropeptide serotonin. After incubation and washing out of the primary antibody, the appropriate secondary antibody was used. For anti-5HT and anti-FMRF, an anti-rabbit Cy3 (Jackson Labs; 1:100 dilution) was used. For tyrTub, Cy3-conjugated anti-Mouse Ig (Jackson Labs; 1:1,000 dilution) was used. The nuclear marker sytox (Molecular Probes; 1:10,000 dilution) and the actin marker phalloidin (rhodamine-labeled, Molecular Probes; 1:1,000) were used to label the nuclei and muscles, respectively, according to the manufacturer’s recommendations. Preparations were then viewed and recorded using a Biorad Model MRC1024ES Confocal Microscope and Laser Sharp version 3.2 software. The confocal stacks were viewed using the program Image J.
3D modeling
A digital 3D model of a representative stack of a Macrostomum juvenile labeled with tyrTub, sytox, and phalloidin was generated using a freeware 3D modeling plug-in developed for Image J (see Cardona et al. 2006). Using the various modeling functions, traces of the head muscles, axonal tracts, and organ outlines (brain, pharynx, midgut) were generated. These traces were rendered in the program to generate a 3D model of the juvenile Macrostomum brain and adjacent structures. The modeling files were exported into the 3D viewing and modeling program Blender (www.blender.org), where color and smoothing changes were made for figure generation.
Results
Overview of Macrostomum juvenile brain
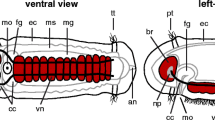
The brain of M. lignano shortly after hatching consists of approximately 1–2 × 103 neurons, including sensory and motor elements and interneurons. Morphologically, the juvenile brain resembles the brain of the adult (Fig. 1), which fills the anterior tip of the worm. One main and two accessory pair of axon tracts project posteriorly. The main tracts, which extend along the ventro-lateral body wall, possess accompanying cell bodies spaced at regular intervals (ventral nerve cord; Ladurner et al. 1997). The anterior part of the ventral nerve cord (anterior ganglion of the ventral nerve cord) is enlarged and contains a substantial central neuropile. Mechanosensory and chemosensory neurons form specialized complexes concentrated at the anterior tip (Xylander et al. 1997; Morris et al. 2004) and the pharynx. The pharynx of Macrostomum is closely attached to the posterior surface of the brain and is intimately connected to the latter by sensory and motor fibers. Conglomerations of neurons (pharyngeal ganglia) flank the pharynx on either side. Simple pigment-cup eyes, consisting of a pair of rhabdomeric photoreceptors ensheathed by pigment cells, are embedded in the dorso-posterior cortex of the brain. In addition to these light-sensing organs, a pair of ciliary receptors with presumed light-sensing function have been described (Sopott-Ehlers et al. 2001) at a position anterior to the brain.
Schematic drawing of the nervous system of Macrostomum (species, hystrinicum marinum; after Ladurner et al. 1997). Shown are the neurites of serotonergic neurons, which outline the pattern of neuropile and peripheral tracts. Left side Dorsal view; right side ventral view
The brain is formed by an outer cortex of neural somata surrounding a fibrous neuropile. The cortex is thick (6–8 cell diameters) anteriorly and laterally but forms only a thin wall (1–2 cell diameters) posteriorly where it demarcates the boundary between brain and pharynx (Fig. 2). Neural somata are relatively uniform in size and packing density, except for the layer of cells that borders the neuropile, and that shows a slightly smaller size and higher packing density (“core cortex”) than the more peripheral cortex (Fig. 2). Peripherally, one cannot easily distinguish without the help of specific markers neurons from muscle cells. Muscle fibers and gland necks, presumably accompanied by cell bodies, also penetrate the brain horizontally and vertically at invariant positions (Fig. 2). Beside the stereotypic pattern of neurite bundles, these cerebral muscles and cerebral glands represent a convenient system of landmarks that help define compartments in the juvenile brain. The pattern of muscles, nerve roots, and compartments is essentially invariant in juvenile specimens. As part of this study, we generated approximately 45 complete stacks of confocal sections. The pattern of nerve roots and muscles was analyzed in eight stacks and turned out to be the same in each specimen. Groups of nerve cells (e.g., FMRFamide-positive neurons, serotonin-positive neurons) appeared slightly more variable in regard to cell number and cell body position, although the neurite projection patterns of these cells, described below, could be verified for each specimen. Considerable variability has been reported for transmitter-defined subpopulations of neurons in adult flatworms (Koopowitz 1986; Joffe and Reuter 1993), and we assume that the variability of nervous and muscular pattern elements described here for Macrostomum will increase during postembryonic development.
Neuropile and cerebral muscles of Macrostomum juvenile brain. All panels show Z projections of horizontal confocal sections of specimens labeled with phalloidin (muscle fibers, green), tyrTub (neurites and cilia, red), and sytox (nuclei, blue). Only head portion of juvenile worms is shown; anterior is to the top. Dorso-ventral levels of Z projections (indicated graphically in Fig. 3) correspond to ventral brain cortex (a), ventral commissural neuropile (b), central compartments at level of central roots (c), central compartments at level of horizontal tracts (d), dorsal commissural compartment (e), dorsal brain cortex (f). In panels of left column (a, c, e), midline is indicated by white hatched line; labeling of nuclei is omitted on right halves of specimens to improve visibility of muscle fibers and neurites. White arrows in d delineate antero-medial domain of body wall that lacks regular layer of epidermal nuclei. Violet circles in e and f indicate position of eyes. agvc Anterior ganglion of ventral nerve cord; amc anterior muscle chiasm; amcd antero-dorsal muscle chiasm; c11, c12a/b, c13, c14, c15 central nerve roots 1–5; CC core cortex; ccn cervical connective; cd centro-dorsal compartment; ci centro-intermediate neuropile compartment; cl centro-lateral neuropile compartment; clf central longitudinal fascicles; clm central longitudinal muscle; cma centro-medial anterior compartment; cmp centro-medial posterior compartment; crp1–2a/b centro-posterior nerve roots 1–2; d11–13 dorsal nerve roots 1–3, dorsal tier; d21a/b, d22a/b, d23 dorsal nerve roots 21–23, ventral tier; db2–6 dorsal cerebral muscles 2–6; dc dorsal compartment; dcom1–6 dorsal commissures 1–6; dcp dorso-posterior compartment; dl1–7 dorsal longitudinal body wall muscles 1–7; dmo dorsal oblique muscle; dpo dorso-posterior oblique muscle; drp1, drp2a/b/c, drp3 dorso-posterior nerve roots 1–3; epi epidermal layer; mg midgut; PC peripheral cortex; ph pharynx; phco postpharyngeal commissure; phg pharyngeal ganglion; phlu lumen of pharynx; pl1–8 longitudinal pharyngeal muscles 1–8; pmna pmnp anterior and posterior pharyngeal muscle net; pt 1–5 transverse pharyngeal muscles; pvtd dorsal pharyngeal vertical muscles; pvtv ventral pharyngeal vertical muscles; sc1–3 central sensory/glandular tracts 1–3; sd11–13 dorsal sensory/glandular tracts 1–3, dorsal tier; sd21–23 dorsal sensory/glandular tracts 1–3, ventral tier; sv21–23 ventral sensory/glandular tracts 1–3, ventral tier; v11–13 ventral nerve roots, ventral tier; v21–23 ventral nerve roots, dorsal tier; vb2, vb3, vb5a, vb5b, vb5c, vb5p, vb6a, vb6b, vb7 ventral cerebral muscles; vc ventral nerve cord; vcb ventro-basal compartment; vcl ventro-lateral compartment; vcm ventral commissural compartment; vcom1–6 ventral commissures 1–6; vl1–7 ventral longitudinal body wall muscles; vpo ventro-posterior oblique muscle; vta1a–d medial row of anterior vertical muscles; vta2–3 lateral anterior vertical muscles; vtf vertical fascicles; vtp2 posterior vertical muscle; vtpd1–4 posterior-dorsal vertical muscles 1–4; vtpv1–4 posterior-ventral vertical muscles 1–4; Scale bar, 10 μm
The pattern of somatic muscles associated with the Macrostomum brain
Muscle fibers are arranged in a pattern of extraordinary regularity underneath the epidermis (Rieger et al. 1991, 1994 in M. hystrinicum; Morris et al. 2004). The inner layer of the body wall muscular plexus is formed by 14 pairs of evenly spaced longitudinal muscles (dorsal longitudinals (dl) 1–7, ventral longitudinals (vl) 1–7 (Figs. 2 and 3). Circular muscles, forming the outer layer of the somatic musculature, are thinner, more closely spaced, and do not appear to display an invariant pattern. In the head of the animal, two systems of preferentially longitudinal fibers diverge from the body wall plexus and extend into the interior of the animal where they form a muscle net around the brain neuropile and the pharynx, respectively. The brain-associated deep muscle plexus forms a group of crescent-shaped cerebral fibers [dorsal cerebrals (db), ventral cerebrals (vb)] that, in part, skirt the outer surface of the brain, and in part, pass through the cortex and neuropile (Figs. 2a and 3d). In addition, several regularly spaced vertical and transverse muscles (vt, tm) branch off the external net and anterior cerebral net. All of these muscles form a highly stereotypical network of fibers that are found in an invariant relationship with compartments and neuronal fiber tracts.
Muscle fibers and their relationship to the structures of the head of Macrostomum. All panels represent digital 3D models derived from serial confocal horizontal sections of juvenile specimens in which muscles, neurites, and cell nuclei were labeled (see Fig. 2). Models were derived from only the left side of the body; in cases were a right side is shown, this was derived by reflection from its left counterpart. Systems of muscle fibers are shown in separate shades of green (see color key at upper right), with the exception of vl/vb5, vl/vb6, and vl/vb7, which are closely related to the neuropile and for reasons of clarity are accented in yellow, orange, and red, respectively. Surfaces of brain, neuropile, pharynx, and midgut are shaded gray; eyes are shown in black. a Anterior view. b Ventral view. In both panels, left and right half of model are drawn apart; in left halves, brain surface is rendered highly transparent to allow for better visibility of cerebral muscles; in right halves, transparency of brain surface is reduced so that muscles appear at low contrast. Note that only one half was modeled; the complementary half was derived by mirroring the model. Ventral longitudinal muscles vl2 and vl3 in the specimen modeled were split at the level of the head (vl2a/b; vl3a/b). c Dorsal view of half of head. d Medial view. Gray bars with capital letters A–F to the left of model indicate dorso-ventral levels at which the Z projections shown in correspondingly lettered panels of Fig. 2 were taken. For abbreviations, see legend of Fig. 2. Scale bars, 10 μm (a–c); 5 μm (d). Description of muscle trajectories (see also Fig. 2): ventral cerebral muscles (vb group) vb2 and vb3, follow a predominantly longitudinal course, then curve upward and reach the dorsal system of body wall muscles at a right angle (Figs. 2a and 3b). vb2 extends anteriorly next to the pharynx; after traveling through the ventral cortex of the brain, this muscle turns dorsally, grows through the anterior (peripheral cortex), and terminates on the body wall muscle dl2. vb3 branches off vl3 (lateral of vb2) and initially extends parallel to vb2 through the ventral brain cortex. At the point where vb2 makes its sharp dorsal turn, vb3 bends medially and then dorsally. It terminates further medially than vb2. vb5 and vb6, split from vl5 and vl6, respectively. These muscles extend in the horizontal plane, first anteriorly and then medially, which implies that they meet their contra-lateral counterparts in the midline (Figs. 2a and 3b). After originating from vl5/6 behind the level of the pharynx, vb5 and vb6 grow along the inner neuropile surface of the anterior ganglion of the ventral cord. vb5 then turns medially and splits into three branches. The posteriormost branch, vb5a, contacts the ventral surface of the brain neuropile. vb5b and vb5c travel slightly more anteriorly and dorsally through the core cortex. vb5a and 5b converge again near the midline, where they turn antero-dorsally, forming a conspicuous “anterior muscle cross” that reaches from the brain neuropile through the cortex to the apical sensory complex (amc in Fig. 2a,c,e). vb5c follows a dorso-medial trajectory close to the anterior neuropile boundary. Reaching the level of vb7 (see below), vb5c extends parallel to this muscle; shortly before reaching the midline, it turns anteriorly and projects to the anterior sensory field. Extending antero-dorsally of the vb5 system, vb6 also forks and forms three branches (vb6a–c; Figs. 2c and 3b). These fibers travel medially at the boundary between densely packed core-cortex (posterior to vb6 system) and peripheral cortex (anterior to vb6). (iii) vb7 angles steeply from vl7 and follows a medial trajectory (Figs. 2e and 3b). Growing right over the central nerve roots that branch off the brain neuropile, this muscle lies at the anterior-central neuropile boundary. vb7 does not branch. In the midline, it merges with its contralateral counterpart. Dorsal cerebral muscles (db group) Db2 runs close to the dorsal midline, at a level more medial than that of the eye. After passing the cortex, the db2 fibers of either side converge turn ventrally and merge with the vb5 fibers that form the anterior muscle chiasm (see above). db3 branches off the dl3 fiber posterior of the eye (Fig. 3c). This muscle passes right over the pigment cup of the eye. It continues forward and crosses the neuropile right lateral of the dorsal nerve roots exiting the neuropile. Turning ventrally, vb3 meets the horizontal muscle crescent formed by vb6c (Fig. 3d). db5 and db6 extend lateral of db3, penetrating the dorso-lateral cortex of the brain. Like db3, they also meet vb6 at a right angle and appear to merge with this commissural muscle system (Fig. 3c,d). Vertical cerebral muscles (vt group) vta1a–e (anterior verticals) form a paramedian row (Figs. 2a and 3a,b,d) in the anterior cortex. They all branch off the ventral body wall muscle net at the medio-lateral level of muscle vl1. vta1a is the most posterior one of the series; it passes right over the anterior neuropile surface and serves as an external landmark for the boundary between central and lateral neuropile. vta2 and 3 are located laterally adjacent to the vta1 series (Fig. 3b,d). Near the anterior tip of the head, a number of additional, seemingly more irregularly distributed vertical fibers are found (Fig. 2a, not annotated). vtpd1–4 (dorso-posterior verticals), this series of short, vertical muscles branch off the dorsal body wall muscle net (from muscles dl1–4) at a right angle and pass straight ventrally at the surface of the posterior brain cortex (Figs. 2c,e and 3c,d). The fibers converge upon and fuse with the pharyngeal muscle net that surrounds the pharynx wall and forms the muscular boundary between pharynx and posterior brain cortex (Fig. 3d). vtpd3 passes right behind eye, where it crosses the path of db3. vtpv (ventro-posterior verticals), several small, ventrally directed fibers that extend forward from the pharyngeal muscle net and insert at the ventral body wall muscle net (Fig. 3d). vtp2 and clm, the long posterior vertical fiber vtp2 originates at a ventral level in the pharyngeal muscle net and extends dorsally on the posterior neuropile surface; at a central brain level, it gives off the central longitudinal muscle (clm), the only cerebral muscle that actually crosses the neuropile from anterior to posterior (Figs. 2c and 3d). Continuing dorsally, vtp2 branches into 2–4 closely apposed branches that all flank the eye medially (Fig. 3c,d). These fibers converge at a right angle on the dorsal muscle net at around dl2/3. Vertical and transverse pharyngeal muscles. Dpo originates in the medial part of the pharyngeal muscle net. Extending antero-dorsally, it passes the posterior surface of the eye and inserts at cerebral muscle db6 (Figs. 2e and 3c). pt (transverse pharyngeals) connect the pharyngeal muscle net with the lateral subset of body wall muscles (Figs. 2e and 3b,c). pv [vertical pharyngeals (pv)], dorsal vertical pharyngeal muscles (pvtd) and ventral vertical pharyngeal muscles (pvtv) extend between the pharyngeal muscle net and the dorsal and ventral body wall muscles, respectively (Fig. 3d)
Ventral cerebral muscles
One finds on each side of the ventral midline seven body wall longitudinal muscles (vl–vl7; Fig. 3b). With the exception of vl1 and vl4, all ventral longitudinals give off a cerebral branch (vb) at the level of the pharynx. The vb muscles form a meshwork of more or less horizontal fibers within the anterior brain cortex that curve around the anterior neuropile surface. As further detailed below, the muscle crescents vb5, vb6, and vb define the ventral, central, and dorsal compartments of the brain neuropile (Figs. 3a and 4a–c).
Neuropile compartments and nerve fiber tracts of the Macrostomum brain. All panels represent digital 3D models derived from serial confocal horizontal sections of juvenile specimens in which muscles, neurites, and cell nuclei were labeled (see Fig. 2). The brain surface (gray) is rendered semitransparent. Eyes are rendered black. Neuropile compartments are shown in different shades of green; peripheral nerve roots and central nerve tracts in shades of purple. Muscle fibers closely associated with the central nervous system (vl5–7; vb5–7) are shown in relationship to neural compartments and tracts. a–c Focus on peripheral nerve roots; central commissural tracts rendered in faint color. a Anterior view. b Dorsal view. c Dorso-posterior-medial view of left side of the brain. d–f Focus on central commissural tracts. Peripheral nerve roots are omitted; commissural tracts are rendered purple. d Anterior view of left side of the brain. e Dorsal view of right side. f Dorso-posterior-medial view of the right side of the brain. For abbreviations, see legend of Fig. 2. Scale bar, 10 μm (a–f). Description of nerve roots (see also Fig. 2): ventro-basal nerve roots v11–v13 penetrate the cortex underneath the level of the ventral vb5 muscle crescent (Figs. 2a and 4a). v11 is a thick, short bundle situated in the ventral midline and projecting straight upward into the floor of the neuropile. v12 passes posteriorly underneath the commissures and then turns dorsally; this axon bundle contributes to the system of vertical tracts that dominate the posterior part of the central neuropile compartment (Figs. 2c and 4b,c; see below). In addition, a branch of v12 turns medially and contributes to the anteriormost ventral commissural tract (vcom1). The v13 root connects to the ventral neuropile lateral of v12. Most of the v13 axons seem to contribute to the commissures vcom1/2. Ventro-anterior roots v21–v23 pass through the cortex at a level between the ventral muscle crescent (vb5) and the central crescent (vb6; Figs. 2a and 4a). The medially located v21 consists of at least three individual bundles coming from the medial cortex right in front of the ventral neuropile. Rather than entering the neuropile directly, these bundles turn ventral and posterior, thereby curving around the floor of the neuropile underneath the ventral commissure. At the posterior surface of the neuropile, v21 turns dorsal and contributes to the vertical tracts of the central neuropile. By contrast, axons entering the ventral neuropile through roots v22 and v23 appear mostly to turn medially and contribute to the commissural tracts vcom5/6. Central nerve roots c11–c15 enter the central neuropile from anteriorly (Figs. 2c,d and 4a,b). These roots appear to be formed by clusters of neuronal somata located in the anterior cortex. Many of these somata may constitute sensory neurons, and project dendrites forward into the apical complex (see below). Nerve roots c11–c13 converge and connect to the centro-medial neuropile as one thick stem. Many axons of these roots continue posteriorly as part of the central longitudinal fascicles that form the roof of the centro-medial compartment (Figs. 2c and 4b; see below). Axons forming roots c14 and c15 reach the central neuropile at more lateral levels and cannot be followed any further inside the neuropile. Two roots that connect to the tip of the lateral wing of the central neuropile (cp1/2) carry axons from somata in the lateral and postero-lateral cortex. Dorso-anterior nerve roots d21–23 conduct neurites from somata located in the antero-dorsal cortex into the dorsal neuropile. Like the central roots, the dorsal roots may in part consist of sensory axons whose somata send dendrites into the apical complex (Figs. 2e,f and 4b,f). Dorso-central nerve roots d11–d13 carry vertically directed axons from clusters of somata in the dorsal cortex into the dorsal neuropile (Figs. 2f and 4a). Dorso-posterior nerve roots drp1 and drp2 originate in the postero-medial brain cortex. Their axons extend forward in between the eyes and reach the dorsal cortex, then appear to turn ventrally and merge with the system of vertical fascicles (Figs. 2f and 4b,c)
Dorsal cerebral muscles
The dorsal net of body wall muscles contains seven bilateral pairs of longitudinal fibers (dl1–dl7; Figs. 2e and 3c). dl2, 3, 5, and 6 each gives rise to a cerebral muscle. These dorsal cerebral muscles extend antero-ventrally, thereby approaching the horizontally oriented muscle crescents formed by vb5, 6, and 7 at a right angle (Fig. 3d). All dorsal cerebral muscles extend through the dorsal cortex at a depth corresponding roughly to the boundary between inner core cortex and peripheral cortex (Fig. 2c–e).
Vertical, oblique, and transverse muscles
These form a set of short muscles that interconnect the body wall muscle net with the cerebral muscles. We define a group of anterior vertical fibers that penetrate the anterior cortex of the brain vertically. All anterior verticals are located at a fairly medial level; the position of vta1a, which flanks the anterior surface of the neuropile, defines the boundary between central and lateral neuropile (Figs. 2c and 3d). The posterior verticals are located posterior to the neuropile and can be further divided into a dorsal posterior series (vtpd1–4), several ventral posterior muscles (vtpv), and a prominent fiber reaching from the dorsal body wall to the level of the ventral pharynx (vtp2; Figs. 2c,e and 3c,d). A unique pair of thin fibers, clm (central longitudinals) branch off the vtp2 at a central brain level (Figs. 2c and 3d) and pass anteriorly through the neuropile (they are the only muscles to do that!). The vertical and transverse pharyngeal muscles connect the pharyngeal muscle net with the body wall muscles (Figs. 2e and 3b–d).
Neuropile domains of the Macrostomum brain
The neuropile of the juvenile brain, visualized by global neuronal markers such as tyrTub, forms a crescent-shaped structure in the center of the brain. An invariant set of axon tracts (nerve roots) radiate outward from the neuropile; other tracts (commissures) extend within the neuropile. Based on these tracts, as well as the network of cerebral muscles outlined in the previous section, the neuropile can be subdivided into several discrete compartments (Fig. 4).
The ventral neuropile compartment is defined by a set of ventral commissures (Figs. 4c,f and 5a). Laterally, it connects via a connective (ccn) to the anterior ganglion of the ventral nerve cord (Fig. 4b,c). Nerve roots enter the compartment ventrally and anteriorly. The ventral compartment is bordered by the ventral cerebral muscle crescent formed by vb5a–c (Fig. 4a). The ventral commissures form six bundles of thick, crossing axons. Four of these (vcom1–4) lie next to each other at the same dv level at the ventral floor of the compartment (Figs. 2b, 4f, and 5a); two commissures (vcom5–6) cross dorsal of vcom 1/2 (Fig. 4f). The posterior pair of commissures (vcom3–4) project further lateral, and many of these axons seem to continue through the cerebral connective (ccn) into the anterior ganglion of the ventral nerve cord (agvc; Fig. 4e). Most commissural axons originate from clusters of neurons in the ventral cortex whose axons converge as massive fiber tracts, the “ventral nerve roots,” upon the neuropile. Two sets, one more ventral (ventro-basal roots), the other more dorsal (ventro-anterior roots), are distinguished. The ventro-basal roots (v11–v13) penetrate the cortex ventrally, pass backward underneath the ventral commissures, and then turn dorsally; these axons contribute to the system of vertical tracts that dominate the posterior part of the central neuropile compartment (Figs. 2c and 4b,c; see below), as well as the ventral commissures vcom. The ventro-anterior roots (v21–v23) enter the neuropile more dorsally, at a level between the ventral muscle crescent (vb5) and the central crescent (vb6; Figs. 2a and 4a). These fibers also contribute to the ventral commissures (v22, v23), as well as the centro-medial neuropile compartment (Figs. 2b and 4b,c).
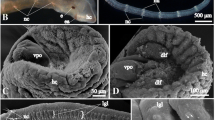
Electron micrographs of cross sections of the Macrostomum juvenile brain. a Cross section at level of rhabdomeric eyes (eye), showing right side of neuropile surrounded by cortex (cx). The cortex contains neuronal cell bodies and nuclei. Muscle fibers are artificially shaded in green, axon bundles in yellow, and gland necks in magenta. Cerebral muscle fibers form a network in the cortex, particularly at the cortex-neuropile boundary. Note the cross-sectioned profile of the central longitudinal muscle (clm), the only muscle that penetrates through the neuropile at the boundary between medial and lateral compartments and the bundle of cerebral rhammite glands (crg) that lies at the center (midline) of the neuropile. Neuropile compartments can be recognized on the basis of location, neurite orientation, and branch morphology: dorsal compartment (dc) with dorsal commissures (dcom), ventral commissural compartment (vcm), centro-medial posterior compartment (cmp), centro-lateral compartment (cl). Among the prominent axon bundles are the central longitudinal fascicle (clf) and the cervical connective (ccn). Boxed areas are shown at higher magnification as b–f. b Eye with pigment cell [pgc, rhabdomere (rha) and adjacent neuronal cell body (ne)]. c Detail of central longitudinal fascicle (clf). d Bundle of cerebral rhammite glands (crg), defined by rod-shaped, electron dense inclusions that lack concentric layering. The centro-medial posterior neuropile (cmp) surrounding the rhammite glands contains highly branched neurites and synapses (f). e Ventral cerebral muscle (ms) and adjacent central roots (vr) passing into the neuropile. g Cross section of brain cortex at an anterior level, showing nerve root (nr; shown at higher magnification in inset) extending alongside rhammite gland (rh). h Cross section of lateral brain showing transition of brain neuropile (np) through cortex (cx) into cervical connective (ccn) that leads to the ventral nerve cord. i Z Projection of confocal sections of head of Macrostomum juvenile showing pharyngeal rhammite glands (prg) and cerebral rhammite glands (crg). The cerebral rhammite glands are formed by cell bodies located posterior to the brain (arrows) and project long necks through the center of the neuropile towards the anterior tip of the head. Other abbreviations, clm central longitudinal muscle; ph pharynx. Scale bars, 5 mm (A, H, I), 2 μm (B, C, D, E), 0.5 μm (f)
The central neuropile
Most domains within this compartment of the Macrostomum brain appear to constitute true “association neuropiles,” based upon the fact that they are formed by branched collaterals of the ventral commissures, as well as branches of dorsal fiber systems (see below). The central neuropile is bounded anteriorly by the vb6 muscle crescent (Fig. 4a,c). TyrTub-positive neurites form short, irregularly oriented fibers that contrast sharply with the commissural and longitudinal systems flanking the central neuropile ventrally and dorsally (Figs. 4c,f and 5a). The central neuropile is divided into an unpaired median domain (anterior and posterior centro-medial compartments) and bilaterally symmetric “wings”; in these wings, a centro-intermediate compartment can be distinguished from a centro-lateral compartment (Figs. 2c and 4a–c). The roof of the central neuropile constitutes the centro-dorsal compartment. The wings of the central neuropile are formed by branches of fibers associated with the ventral commissures vcom 1–6 (see above). The posterior centro-medial compartment stands out by its faint tyrTub immunoreactivity (Fig. 2c). It seems to be formed mainly by thin, vertically oriented axons penetrating into the center of the neuropile from ventral and dorsal. Forming the posterior “wall” of the centro-medial compartment are the Vertical fascicles (vtf; Figs. 2c and 4b), which include dorsally directed axons entering the neuropile through the ventral roots (e.g., v12, 21) and ventrally directed axons stemming from the postero-dorsal roots (see below). The vertical tracts are flanked by a conspicuous pair of muscles, the vtp2 (vertical posterior) muscles (Fig. 2c). The centro-dorsal compartment is formed by and around the central longitudinal fascicles, a system of thick fiber bundles formed by axons entering through the central and postero-dorsal nerve roots (Figs. 2c and 4b; see below). A unique pair of muscles, the central longitudinal muscles (clm), penetrate the central neuropile anteriorly to posteriorly and flank the medial longitudinal fascicles on either side (Figs. 2c,d and 5a,c). Furthermore, the long necks of the rhammite glands, whose cell bodies are located posterior to the brain, pass through the centro-medial neuropile (Fig. 5a,d).
The dorsal neuropile
Like the ventral neuropile described above, the dorsal neuropile compartment is characterized by a system of regularly spaced commissural tracts (dcom1–6; Figs. 2e and 4d,f). The dorsal neuropile is flanked at its anterior surface by the dorsal muscle crescent (vb7; Fig. 4a). The dorsal commissures are shorter and consist of thinner axons than the ventral commissures and, in part, intertwine with the fibers of the central longitudinal fascicles lying right below. Axons of the dorsal commissures cannot be followed very far beyond the midline; laterally, they are lost in a dense mat of fibers that form the lateral wings of the dorsal neuropile (Figs. 2e and 4d–f). Three systems of nerve roots, the dorso-anterior roots (d21–d23), dorso-central roots (d11–d13), and dorso-posterior roots (drp1–2) enter the dorsal neuropile (Figs. 2e,f and 4b,f). A small, unpaired ovoid neuropile compartment (dorso-posterior compartment, dcp; Fig. 4b,c) surrounds the dorso-posterior roots shortly before they enter the main dorsal neuropile. Located in between the eyes (and possibly directly contacted by the somata of photoreceptor neurons), it is reasonable to assume that this compartment may be involved in the processing of visual input to the brain.
The pattern of FMRFamidergic and serotonergic neurons in the Macrostomum brain
The detailed map of neuropile compartments, nerve roots, and cerebral muscles introduced in the previous sections allows one to describe the pattern of specific populations of neurons at a high level of resolution. As an example, we show here the pattern of two groups of neurons that can be visualized with an antibody against FMRF and serotonin. FMRF-positive neurons form six main clusters of 5–10 cells each (see also Ladurner et al. 1997). Cell bodies within these clusters are large, compared to most other neurons, and form part of the central cortex that directly contacts the neuropile. The VA cluster is located adjacent to the ventro-lateral neuropile compartment (Fig. 6a,e), the VP cluster flanks the pharyngeal nerve ring (Fig. 6a), CE surrounds the centro-lateral compartment (Fig. 6b,c,f,g), and DP/DM cap the centro-intermediate and dorso-medial compartment (Fig. 6d,h). Outside the clusters of strong FMRFergic neurons there are additional, small and faintly expressing cells scattered throughout the central cortex. The majority of neurites formed by the FMRF-positive neurons are varicose, scattered fibers that extend within the ventral and dorsal commissural neuropile and the ring-shaped neuropile surrounding the pharynx (Fig. 6a,b,e,f). Some compartments, notably the centro-medial posterior compartment, are conspicuously devoid of FMRF-positive fibers (Fig. 6b,f). Longitudinally oriented fibers extend from the ventral and dorsal neuropile, as well as the pharyngeal nerve ring, and fasciculate with individual longitudinal cerebral and body wall muscles. In most of these cases, a single varicose nerve fiber accompanies a given muscle (Fig. 6d).
Pattern of FMRFamidergic neurons (a–h) and serotonergic neurons (i–l) in the Macrostomum juvenile brain. All panels show Z projections of horizontal confocal sections of specimens. Panels of first row (a, e, i) present views of ventral level (ventral commissural neuropile, cervical connective); second row (b, f, j) represents central level (central neuropile compartments); sections of third row (c, g, k) show the dorsal neuropile (level of eyes), and bottom row (d, h, l) has sections through dorsal cortex and peripheral nerve plexus. Specimen shown in first column (a–d) was labeled with phalloidin (muscle fibers, red) and anti-FMRFamide (green); label in middle column (e–h) was anti-FMRFamide (green), anti-Tryrosinated tubulin (red), and sytox (blue); right column (i–l) illustrates labeling with anti-Serotonin (green), tyrTub (red), and sytox (green). Hatched lines in a–h delineate clusters of FMRFamide-positive neurons (CE centro-anterior cluster; DL dorso-lateral cluster; DM dorso-medial cluster; DP dorso-posterior cluster; VA ventro-anterior cluster; VP ventro-posterior cluster). Clusters of serotonergic neurons shown in i–l include ventro-anterior cluster (AVC) and dorso-medial cluster (DMC). Serotonergic neurites form a conspicuous peripheral nerve plexus (pnpl; i and l). For more details, see text. For other abbreviations, see legend of Fig. 2. Scale bar, 20 μm
Serotonergic neurons are distributed in a pattern that is distinctively different from that of FMRFergic cells (Ladurner et al. 1997). Two clusters of large serotonergic cell bodies, one located in the anterior ganglion of the ventral cord (AVC; 12–15 cells; Fig. 6i) and one capping the dorso-medial neuropile (DMB; 3–4 cells; Fig. 6k,l) can be distinguished. Somata are located in the peripheral cortex, further away from the neuropile of the brain and anterior ganglion. Neurites are loosely scattered throughout the lateral neuropile compartments; many fewer fibers than those of FMRFergic neurons have the tendency to cross the midline (compare Fig. 6e and i; g and k). Peripheral serotonergic fibers emanating from the neuropile of the brain and anterior ganglion form a subepidermal plexus of longitudinally and transversally oriented varicose axons (Fig. 6l).
Discussion
Our analysis shows that neurons and their processes, as wells several nonneuronal elements such as muscles and glands, form a highly reproducible pattern that allows one to subdivide the juvenile Macrostomum brain into numerous compartments. As observed for other invertebrates, the neuropile of the Macrostomum brain exhibits compact bundles of long fibers, next to profusely branched terminal axons and dendrites. Transverse fiber bundles form a dorsal and ventral system of commissures that define a dorsal and ventral medial neuropile compartment, respectively. Neurites that enter the neuropile through stereotypical nerve roots form longitudinal bundles defining a central medial compartment in the neuropile. Other compartments (ventro-lateral and centro-lateral, centro-posterior) are characterized by the relative absence of long neurites. Cerebral muscle fibers form a fine meshed system of longitudinal, transverse, and vertical elements surrounding the neuropile on all sides, providing additional landmarks in the topological map of the Macrostomum brain, a map that will prove its value in future studies analyzing neuronal connectivity and development, as well as the expression of molecules controlling these processes.
Structural elements of the Macrostomum brain
Previous studies in a number of flatworm species had shown that muscles and/or glands penetrate the brain (e.g., Klauser and Tyler 1987; Bedini and Lanfranchi 1991, 1998; Biserova et al. 2000). One of the developmental causes of this interpenetration of neuronal and nonneuronal elements is the absence of a stable glial sheath. In “higher” invertebrates and in vertebrates alike, such glial layers surround the brain on all sides, forming an impermeable blood–brain barrier. Another important aspect of flatworm biology that may explain the mixing of neuronal and nonneuronal cell types is development. Flatworms do not undergo a large scale morphogenetic movement, akin to gastrulation in vertebrates or arthropods, in which endoderm, mesoderm, and ectoderm become established as separate cell layers, which then further differentiate into different tissues and organ such as nervous system (ectodermal) and muscle tissue (from mesoderm). Instead, the early flatworm embryo consists of a conglomerate of progenitor cells (embryonic primordium) from which different cell types appear to differentiate in situ (Thomas 1986; Younossi-Hartenstein et al. 2000, 2001; Younossi-Hartenstein and Hartenstein 2000a, b, 2001; Ramachandra et al. 2002; Morris et al. 2004; Cardona et al. 2005, 2006). As progenitors of muscle cells, nerve cells, glands, and other tissues are intermingled within the embryonic primordium, the corresponding cells, once differentiated, will be intermingled as well. In other words, according to this hypothesis, there is no need to assume that muscle cells or glands “migrate” into the brain primordium at some point in embryonic development; instead, these cells are part of the brain primordium from the very beginning. Studies using cell-specific markers expressed at early stages of embryonic development are urgently needed to reconstruct the specification, origin, and morphogenetic movements of cells in the flatworm embryo. We have generated a collection of ESTs for M. lignano that can act as a resource for such markers (Morris et al. 2006).
Our data indicate that the number and pattern of muscles associated with the Macrostomum brain is highly invariant. Note that this can so far be stated only for the juvenile (the stage within 24 h of hatching) investigated in this study; muscle fibers clearly increase in number during postembryonic development (Rieger et al. 1994; Ladurner et al. 2005a, b). The vertical and longitudinal cerebral muscles branch off the body wall muscle layer at stereotypical positions and enter the brain cortex. The trajectory of three of these muscles, vb5, 6, and 7, is noteworthy because they directly contact the neuropile and thereby may interact with neurons most closely. The other cerebral muscles, vb2–6 and db2–5, traverse the brain cortex but do not come into direct contact with the neuropile. Interestingly, vb5–7, as well as the body wall muscles from which they originate, vl5–7, are the muscles that flank the ventral nerve cord, the most prominent nerve tract extending throughout the trunk of the animal. It therefore appears that vl/vb5–7 form a special subset of muscles that functionally and/or developmentally are most closely related to the nervous system. A close relationship between nerves and musculature had been noted in previous works (e.g., Reiter et al. 1996) and was taken as evidence to suggest developmental interactions between these two tissues.
It can be assumed that synaptic contacts between neurons and cerebral muscles occur in the brain neuropile or even the cortex where muscles are in contact with many neuronal cell bodies. Previous studies had shown in numerous cases that neuromuscular junctions in flatworms are located centrally, whereby muscles send processes with synapses towards the orthogon or brain (Chien and Koopowitz 1972; Cousin and Dorsey 1991; Rieger et al. 1991). Central neuromuscular junctions have not yet been described in Macrostomum, but efforts are under way to analyze ultrastructural details of the brain neuropile compartments, which will reveal the pattern of neuromuscular junctions should they exist. It is possible that central effector synapses could also be formed between brain neurons and apical glands. Thus, as described previously (Rieger 1971; Klauser and Tyler 1987) and shown in this paper, a system of long-necked rhammite gland cells with cell bodies in the dorsal brain cortex extend secretory ducts longitudinally throughout the neuropile and anterior cortex.
In the present paper, we describe the complex pattern of nerve roots connecting to the neuropile. In the absence of further neuro-anatomical studies, such as dye injection or specific antibody labeling experiments, little can be said about the “nature” of these nerve roots, that is, whether they carry afferent or efferent axons or whether they are true nerves at all. It stands to reason that at least some of the nerve roots directed anteriorly towards the apical complex, which contains large numbers of sensory dendrites (Klauser and Tyler 1987; Rieger et al. 1991), are sensory nerves. However, many roots could simply represent bundles of neurites connecting the unipolar neurons located in the more peripheral brain cortex to the central neuropile. In arthropods, clusters of neurons descending from one neuronal progenitors (i.e., lineages of neurons) stay closely together and form a single axon bundle that leads from the brain cortex into the neuropile (Younossi-Hartenstein et al. 2006; Pereanu and Hartenstein 2006). Similar bundles of nascent axons have indeed been described in the embryonic brain of Macrostomum (Morris et al. 2004). We consider it possible that some of the “nerve roots” visualized in the juvenile Macrostomum brain by global axonal markers such as tyrTub are the “descendants” of the fiber bundles formed in the embryonic brain.
Neuropile compartments of the Macrostomum brain
The brain neuropile of all animals is formed by the confluence of multiple sensory systems giving input to central interneurons. In animals with highly evolved sense organs (e.g., image-forming eyes, sensitive olfactory antennae) specialized neuropile domains “dedicated” to the processing of the respective sensory information evolved. These domains, or compartments, can be defined physiologically and morphologically. In the best known cases, the sense organs form a large number of parallel channels of afferent axons (e.g., retinal axons in the insect optic lobe) that impinge upon a matrix of reiterated sets of interneurons (i.e., the “cartridges” in the insect optic lobe; “glomeruli” in the olfactory lobe; Strausfeld 1976). This regular array of afferents and interneurons, called “structured neuropile,” presents an impressive and easily discernible image to the neuroanatomist. Structured neuropile compartments have been described in arthropods, cephalopods, and some annelids (Bullock and Horridge 1965; Hanstroem 1968). Other parts of the neuropile in these derived invertebrates lack this regular structure and, in the absence of careful histochemical or histological investigations, have usually been neglected as “unstructured neuropile.” However, as recently shown for the larval brain of Drosophila (Younossi-Hartenstein et al. 2003, 2006), neuropile compartments can be defined in the unstructured neuropile by using markers that visualize glial cells, density of synapses, or diameters and trajectories of neurites. Compartment boundaries will serve as important landmarks for more in-depth studies of neuronal circuitry and neuronal development.
The neuropile of flatworm brains is small and “unstructured.” Therefore, despite the wealth of ultrastructural data mostly pertaining to synaptic morphology, no compartments have been described up until now. In the present study, the global axonal marker tyrTub served as a useful tool to visualize axonal trajectories, which revealed a subdivision of the neuropile into several easily identifiable compartments. Several compartments, including the dorsal compartment, the ventro-medial compartment, and the centro-dorsal compartment, are characterized by their wealth of parallel long axon bundles, whereas other compartments lack such long fibers and may constitute centers where highly branched terminal axons and dendrites interact. Systematic electron microscopic analysis of serially sectioned brains, in conjunction with visualization of entire neurons by dye injection or specific antibody markers, will help to progress in the understanding of brain compartments. Thanks to the abundance of landmarks that can be recognized light and electron microscopically, this analysis is under way. For example, as exemplified in Fig. 5, the commissural and longitudinal fiber bundles are readily distinguishable in ultrathin cross sections of the juvenile Macrostomum brain. Bundles consist of 20–40 axons each, with axon diameter ranging from 0.2 to 0.4 μm. Distinctively different from these fibrous compartments are other neuropile domains, such as the centro-medial posterior compartments, which are composed of abundant small diameter, electron dense profiles (terminal axonal arborizations) surrounding large, translucent structures, which most likely correspond to dendrites.
The labeling of individual neurons by dye injection or specific antibodies will be another prerequisite for elucidation of brain circuitry. A large number of studies employing antibodies against specific neurotransmitters have revealed the distribution of neurons expressing these chemicals in the central and peripheral nervous system of many different flatworm taxa (reviewed in Reuter and Gustafsson 1995; Shaw et al. 1996; Reuter et al. 2001). In conjunction with specific landmarks forming a fine-meshed neuroanatomical framework, such immuno-histochemical studies will be useful to add functional information to the brain map. For example, as shown in Fig. 6, FMRFamidergic neurons provide a heavy innervation to the ventro-lateral and centro-lateral “association” compartments, whereas the projection of serotonergic neurons is dense in the pharyngeal nerve ring but relatively sparse in the brain neuropile. However, the use of markers such as anti-Serotonin and anti-FMRFamide is limited when it comes to the reconstruction of individual neurons, because neurites of multiple closely spaced cells fasciculate and cannot be distinguished light microscopically. Antibodies and molecular probes that recognize individual neurons located far apart start to become available in planarian (triclad) flatworms (Bueno et al. 1997; Cebria et al. 2002); these will make it possible to reconstruct neuronal shapes. In addition, the injection of fluorescent dyes label neurons in their entirety. Studies utilizing this approach in flatworms are rare (Elvin and Koopowitz 1994; Koopowitz et al. 1996; Okamoto et al. 2005) but have revealed a number of interesting facts. For example, in both polyclads (representative of the more primitive archoophoran clade) and rhabdocoels (derived neoophorans), the brain contains a large fraction of multipolar neurons. Most neurons visualized by dye injection form several long processes with relatively sparse arborizations along their shafts. Such long fibers (when bundled together) would account for the commissural and longitudinal tracts described in the present study for the Macrostomum brain. In addition, many dye-filled neurons exhibited small but dense terminal arbors, which represent the sites of synaptic interactions with other cells. Dye injections in the Macrostomum brain, carried out against the backdrop of the topological information provided by the brain map described in this paper, will help significantly to unravel neuronal circuitry in a primitive invertebrate organism.
High resolution maps are indispensable for the documentation and interpretation of gene expression patterns. Probes for genes expressed in the nervous system have become available in large numbers in triclads (Sanchez Alvarado et al. 2002; Reddien et al. 2005) and Macrostomum (Morris et al. 2006). To study the function of these genes in brain development or function, the first question that needs to be addressed is the precise expression patterns. Does the expression of a given gene in a population of neurons signify that these cells are related by ancestry from one progenitor? Or that they produce a common neurotransmitter, receptor, or ion channel? Or that they innervate a discrete compartment or contribute to a specific circuit? The first step in answering these questions is the precise mapping of gene expression in relationship to neuroanatomical landmarks. The map of compartments and cerebral muscles outlined in the present study will provide landmarks that will improve the resolution of future analyses of gene expression in the Macrostomum brain.
References
Ashburner M (1989) Drosophila, a laboratory manual. Cold Springs Harbor Laboratory, New York
Ax P (1996) Multicellular animals: a new approach to the phylogenetic order in nature. Springer, Berlin Heidelberg New York
Baguñà J, Riutort M (2004) The dawn of bilaterian animals: the case of acoelomorph flatworms. BioEssays 26:1046–1057
Bedini C, Lanfranchi A (1991) The central and peripheral nervous system of Acoela (Plathelminthes)—an electron-microscopic study. Acta Zool 72:101–106
Bedini C, Lanfranchi A (1998) Ultrastructural study of the brain of a typhloplanid flatworm. Acta Zool 79:243–249
Biserova NM, Dudicheva VA, Terenina NB, Reuter M, Halton DW, Maule AG, Gustafsson MK (2000) The nervous system of Amphilina foliacea (Platyhelminthes, Amphilinidea). an immunocytochemical, ultrastructural and spectrofluorometrical study. Parasitology 121:441–453
Bueno D, Baguna J, Romero R (1997) Cell-, tissue-, and position-specific monoclonal antibodies against the planarian Dugesia (Girardia) tigrina. Histochem Cell Biol 107:139–149
Bullock TH, Horridge GA (1965) Structure and function in the nervous system of invertebrates. Two volumes. Freeman, San Francisco, CA
Cardona A, Hartenstein V, Romero R (2005) The embryonic development of the triclad Schmidtea polychroa. Dev Genes Evol 215:109–131
Cardona A, Hartenstein V, Romero R (2006) Early embryogenesis of planaria: a cryptic larva feeding on maternal resources. Dev Genes Evol 216:667–681
Cebria F, Nakazawa M, Mineta K, Ikeo K, Gojobori T, Agata K (2002) Dissecting planarian central nervous system regeneration by the expression of neural-specific genes. Dev Growth Differ 44:135–146
Chien P, Koopowitz H (1972) The ultrastructure of neuromuscular systems in Notoplana acticola,a free-living polyclad flatworm. Z Zellforsch 133:277–288
Cornell RA, Ohlen TV (2000) Vnd/nkx, ind/gsh, and msh/msx: conserved regulators of dorsoventral neural patterning? Curr Opin Neurobiol 10:63–71
Cousin CE, Dorsey CH (1991) Nervous system of Schistosoma mansoni cercaria: organization and fine structure. Parasitol Res 77:132–141
Egger B, Ladurner P, Nimeth K, Gschwentner R, Rieger R (2006) The regeneration capacity of the flatworm Macrostomum lignano-on repeated regeneration, rejuvenation, and the minimal size needed for regeneration. Dev Genes Evol 216:565–577
Ehlers U (1985) Das phylogenetische System der Plathelminthes. Fischer, Jena
Elvin M, Koopowitz H (1994) Neuroanatomy of the rhabdocoel flatworm Mesostoma ehrenbergii (Focke, 1836). I. Neuronal diversity in the brain. J Comp Neurol 343:319–331
Gustafsson MK, Halton DW, Kreshchenko ND, Movsessian SO, Raikova OI, Reuter M, Terenina NB (2002) Neuropeptides in flatworms. Peptides 23:2053–2061
Halton DW, Shaw C, Maule AG, Johnston CF, Fairweather I (1992) Peptidergic messengers: a new perspective of the nervous system of parasitic platyhelminths. J Parasitol 78:179–193
Hanstroem B (1968) Vergleichende Anatomie des Nervensystems der Wirbellosen Tiere. A. Asher, Amsterdam
Hartenstein V, Ehlers U (2000) The embryonic development of the rhabdocoel flatworm Mesostoma lingua. Dev Genes Evol 210:399–415
Holland ND (2003) Early central nervous system evolution: an era of skin brains? Nat Rev Neurosci 4:617–627
Joffe B, Reuter M (1993) The nervous system of Bothriomolud balticus (Proseriata)—a contribution to the knowledge of the orthogon in Plathelminthes. Zoomorphology 113:113–127
Keenan CL, Coss R, Koopowitz H (1981) Cytoarchitecture of primitive brains: Golgi studies in flatworms. J Comp Neurol 195:697–716
Klauser MD, Tyler S (1987) Frontal glands and sensory structures in the Macrostomida (Turbellaria). Zool Scr 16:95–110
Koopowitz H (1986) On the evolution of central nervous systems: Implications from polyclad turbellarian neurobiology. Hydrobiologia 132:79–87
Koopowitz H, Elvin M, Keenan L (1996) In vivo visualization of living flatworm neurons using Lucifer yellow intracellular injections. J Neurosci Methods 69:83–89
Ladurner P, Mair GR, Reiter D, Salvenmoser W, Rieger R (1997) Serotonergic nervous system of two macrostomid species: recent or ancient divergence. Invertebr Biol 116:178–191
Ladurner P, Pfister D, Seifarth C, Scharer L, Mahlknecht M, Salvenmoser W, Gerth R, Marx F, Rieger R (2005a) Production and characterisation of cell- and tissue-specific monoclonal antibodies for the flatworm Macrostomum sp. Histochem Cell Biol 123:89–104
Ladurner P, Schärer L, Salvenmoser W, Rieger R (2005b) A new model organism among the lower Bilateria and the use of digital microscopy in taxonomy of meiobenthic Platyhelminthes: Macrostomum lignano, n. sp. (Rhabditophora, Macrostomorpha). J Zoolog Syst Evol Res 43:114–126
Lentz TL (1967) Fine structure of nerve cells in a planarian. J Morphol 121:323–337
Lichtneckert R, Reichert H (2005) Insights into the urbilaterian brain: conserved genetic patterning mechanisms in insect and vertebrate brain development. Heredity 94:465–477
Lowe CJ, Terasaki M, Wu M, Freeman RM Jr, Runft L, Kwan K, Haigo S, Aronowicz J, Lander E, Gruber C, Smith M, Kirschner M, Gerhart J (2006) Dorsoventral patterning in hemichordates: insights into early chordate evolution. PLoS Biol 4:e291
Morris J, Nallur R, Ladurner P, Egger B, Rieger R, Hartenstein V (2004) The embryonic development of the flatworm Macrostomum sp. Dev Genes Evol 214:220–239
Morris J, Ladurner P, Rieger R, Pfister D, De Miguel-Bonet M, Jacobs D, Hartenstein V (2006) The Macrostomum lignano EST database as a molecular resource for studying platyhelminth development and phylogeny. Dev Genes Evol 216:695–707
Okamoto K, Takeuchi K, Agata K (2005) Neural projections in planarian brain revealed by fluorescent dye tracing. Zoolog Sci 22:535–546
Pereanu W, Hartenstein V (2004) Digital three-dimensional models of Drosophila development. Curr Opin Genet Dev 14:382–391
Pereanu W, Hartenstein V (2006) Neural lineages of the Drosophila brain: a three-dimensional digital atlas of the pattern of lineage location and projection at the late larval stage. J Neurosci 26:5534–5553
Radojcic T, Pentreath VW (1979) Invertebrate glia. Prog Neurobiol 12:115–179
Ramachandra NB, Gates R, Ladurner P, Jacobs D, Hartenstein V (2002) Neurogenesis in the primitive bilaterian Neochildia I. Normal development and isolation of genes controlling neural fate. Dev Genes Evol 212:55–69
Reddien PW, Bermange AL, Murfitt KJ, Jennings JR, Sanchez Alvarado A (2005) Identification of genes needed for regeneration, stem cell function, and tissue homeostasis by systematic gene perturbation in planaria. Dev Cell 8:635–649
Reiter D, Boyer B, Ladurner P, Mair G, Salvenmoser W, Rieger R (1996) Differentiation of the bodywall musculature in Macrostomum hystrinicum marinum and Hoploplana inquilina (Plathelminthes), as models for muscle development in lower spiralia. Roux’s Arch Dev Biol 205:410–423
Reuter M, Gustafsson MKS (1995) The flatworm nervous system: pattern and phylogeny. In: Breidbach O, Kutsch W (eds) The nervous system of invertebrates. Birkhäuser Verlag Basel
Reuter M, Raikova OI, Jondelius U, Gustafsson MKS, Maule AG, Halton DW (2001) Organisation of the nervous system in the Acoela: an immunocytochemical study. Tissue Cell 33:119–128
Rieger R (1971) Die Turbellarienfamilie Dolichmacrostomidae Rieger. II. Teil: Dolichomacrostominae. Zool Jahrb Syst 98:569–703
Rieger RM, Tyler S, Smith JPS III, Rieger GE (1991) Platyhelminthes: turbellarida. In: Harrison FW, Bogitsh BJ (eds) Microscopic anatomy of invertebrates, vol.3. Wiley-Liss, New York
Rieger RM, Salvenmoser W, Legniti A, Tyler S (1994) Phalloidin-rhodamine preparations of Macrostomum hystrinicum marinum (Plathelminthes)-morphology and postembryonic development of the musculature. Zoomorphology 114:133–147
Sanchez Alvarado A, Newmark PA, Robb SM, Juste R (2002) The Schmidtea mediterranea database as a molecular resource for studying Platyhelminthes, stem cells and regeneration. Development 129:5659–5665
Shaw MK (1981) The ultrastructure of synapses in the brain of Gastrocotyle trachuri (Monogenea, Platyhelminthes). Cell Tissue Res 220:181–189
Shaw C, Maule AG, Halton DW (1996) Platyhelminth FMRFamide-related peptides. Int J Parasitol 26:335–345
Sopott-Ehlers B, Salvenmoser W, Reiter D, Rieger R, Ehlers U (2001) Photoreceptors in species of the Macrostomida (Plathelminthes): ultrastructural findings and phylogenetic implications. Zoomorphology 121:1–12
Strausfeld NJ (1976) Atlas of an insect brain. Springer, Berlin Heidelberg New York
Telford MJ, Wise MJ, Gowri-Shankar V (2005) Consideration of RNA secondary structure significantly improves likelihood-based estimates of phylogeny: Examples from the Bilateria. Mol Biol Evol 22:1129–1136
Thomas MB (1986) Embryology of the turbellaria and its phylogenetic significance. Hydrobiologia 132:105–115
Xylander WER, Rohde K, Watson NA (1997) Ultrastructural investigations of the sensory receptors of Macrostomum cf bulbostylum (Plathelminthes, Macrostomida). Zool Anz 236:1–12
Younossi-Hartenstein A, Hartenstein V (2000a) The embryonic development of the polyclad flatworm Imgogine mcgrathi. Dev Genes Evol 210:383–398
Younossi-Hartenstein A, Hartenstein V (2000b) Comparative approach to developmental analysis: the case of the dalyellid flatworm, Gieysztoria superba. Int J Dev Biol 44:499–506
Younossi-Hartenstein A, Hartenstein V (2001) The embryonic development of the temnocephalid flatworms Craspedella pedum and Diceratocephala sp. Cell Tissue Res 304:295–310
Younossi-Hartenstein A, Ehlers U, Hartenstein V (2000) Embryonic development of the nervous system of the rhabdocoel flatworm Mesostoma lingua (Abildgaard, 1789). J Comp Neurol 416:461–476
Younossi-Hartenstein A, Jones M, Hartenstein V (2001) The embryonic development of the nervous system of the temnocephalid flatworm Craspedella pedum. J Comp Neurol 434:56–68
Younossi-Hartenstein A, Salvaterra P, Hartenstein V (2003) Early development of the Drosophila brain IV. Larval neuropile compartments defined by glial septa. J Comp Neurol 45:435–450
Younossi-Hartenstein A, Nguyen B, Shy D, Hartenstein V (2006) Embryonic origin of the Drosophila brain neuropile. J Comp Neurol 497:981–998
Acknowledgements
This work was supported by NSF Grant IBN-0110718 to V.H. and the Graduate Student Training Fellowship GM0718 to J.M.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by D.A. Weisblat
Rights and permissions
About this article
Cite this article
Morris, J., Cardona, A., De Miguel-Bonet, M.D.M. et al. Neurobiology of the basal platyhelminth Macrostomum lignano: map and digital 3D model of the juvenile brain neuropile. Dev Genes Evol 217, 569–584 (2007). https://doi.org/10.1007/s00427-007-0166-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00427-007-0166-z