Abstract
In the sea urchin embryo, micromeres have two distinct functions: they differentiate cell autonomously into the skeletogenic mesenchyme cells and act as an organizing center that induces endomesoderm formation. We demonstrated that micro1 controls micromere specification as a transcriptional repressor. Because micro1 is a multicopy gene with at least six polymorphic loci, it has been difficult to consistently block micro1 function by morpholino-mediated knockdown. Here, to block micro1 function, we used an active activator of micro1 consisting of a fusion protein of the VP16 activation domain and the micro1 homeodomain. Embryos injected with mRNA encoding the fusion protein exhibited a phenotype similar to that of micromere-less embryos. To evaluate micro1 function in the micromere, we constructed chimeric embryos composed of animal cap mesomeres and a micromere quartet from embryos injected with the fusion protein mRNA. The chimeras developed into dauerblastulae with no vegetal structures, in which the micromere progeny constituted the blastula wall. We also analyzed the phenotype of chimeras composed of an animal cap and a mesomere expressing micro1. These chimeras developed into pluteus larvae, in which the mesomere descendants ingressed as primary mesenchyme cells and formed a complete set of skeletal rods. The hindgut and a part of the midgut were also generated from host mesomeres. However, the foregut and nonskeletogenic mesoderm were not formed in the larvae. From these observations, we conclude that micro1 is necessary and sufficient for both micromere differentiation and mid/hindgut-inducing activity, and we also suggest that micro1 may not fulfill all micromere functions.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The animal–vegetal (A-V) axis of sea urchin embryos is established during oogenesis. A-V polarity is morphologically evident at the 16-cell stage by the formation of a micromere quartet at the vegetal pole by unequal cleavages. At this stage, micromeres are the only autonomously specified blastomeres; when isolated from embryos or transplanted to another location in a host embryo, micromeres give rise to the skeletogenic mesenchyme cells (Hörstadius 1973; Okazaki 1975a). In addition, the micromeres function as an organizing center for endomesoderm development, as indicated by observations showing that the removal of micromeres results in delays and deficiencies in endomesoderm development (Hörstadius 1973; Ransick and Davidson 1995; Sweet et al. 1999). Conversely, micromeres transplanted to the animal pole of a host embryo induce the development of a secondary gut and ectopic mesoderm (Hörstadius 1973; Ransick and Davidson 1993; Sweet et al. 1999). Micromeres also produce endomesodermal structures when combined with animal cap mesomeres; the chimera develops into a larva that metamorphoses into a complete juvenile sea urchin (Hörstadius 1973; Amemiya 1996; Minokawa and Amemiya 1998; Sweet et al. 1999).
Two research groups have provided evidence that a novel paired-like class homeobox gene, designated micro1 or pmar1, participates in micromere specification. Micro1 was isolated from Hemicentrotus pulcherrimus through a subtraction polymerase chain reaction (PCR) survey for micromere-specific genes (Yamaguchi et al. 1994; Kitamura et al. 2002), whereas pmar1 was cloned from a Strongylocentrotus purpuratus library using PlHbox12 cDNA as a probe (Di Bernardo et al. 1995; Oliveri et al. 2002). Although the micro1 A-D and Pmar1 proteins have five to nine amino acid substitutions in their homeodomains (85–92% identity), the two proteins are considered to be products of orthologous genes based on their function and regulation (Nishimura et al. 2004). The injection of pmar1 or micro1 mRNA into eggs converts all blastomeres to the primary mesenchyme cell (PMC) phenotype, and a similar phenotype is obtained by injecting mRNA encoding fusion proteins that combine the engrailed repression domain with the Pmar1 or micro1 homeodomain (Oliveri et al. 2002; Nishimura et al. 2004). These observations indicate that both gene products act as transcriptional repressors to control micromere specification. Pmar1 or micro1 is postulated to repress a ubiquitous repressor that negatively regulates the early micromere specification genes, Ets, Tbr, and Alx1, as well as the signaling genes, including Delta (Oliveri et al. 2003; Kurokawa et al. 1999; Fuchikami et al. 2002; Ettensohn et al. 2003; Sweet et al. 2002).
Nuclear β-catenin is essential for the specification of vegetal cell fates, including those of micromeres (Wikramanayake et al. 1998; Emily-Fenouil et al. 1998; Logan et al. 1999; Vonica et al. 2000). Nishimura et al. (2004) demonstrated that micro1 is a direct target of β-catenin. Oliveri et al. (2003) produced a chimeric embryo in which a micromere quartet was replaced with a mesomere expressing Pmar1. The chimera developed into an almost complete larva except for the skeletal pattern. This phenotype was suggestive of an incomplete oral–aboral axis because skeletal rods are formed according to the oral–aboral axis, the establishment of which requires vegetal signaling (Okazaki 1975b; Angerer and Angerer 2003). On the other hand, Nishimura et al. (2004) showed that morpholino-mediated micro1 knockdown resulted in a phenotype similar to that of micromere-less embryos, suggesting that micro1 is necessary for both micromere differentiation and for inductive signal production. In spite of the previous functional analyses of pmar1 and micro1, it is still uncertain whether these paired-like homeobox genes alone are responsible for all micromere functions.
The purpose of this study was to further examine the requirement for micro1 in micromere functions. To evaluate the functions of micro1, we constructed chimeric embryos composed of normal blastomeres combined with experimental ones. We used animal cap mesomeres as hosts rather than micromere-less embryos, as the former provide a clearer assay system for testing micromere functions than the latter. Animal caps exclusively develop into dauerblastulae with no vegetal structures; chimeras of animal cap mesomeres and micromeres form archenterons as well as several secondary mesenchyme cell (SMC) types (Minokawa and Amemiya 1998; Sweet et al. 1999). In contrast, micromere-less hosts generate some endomesodermal structures, including the archenteron, skeletogenic cells, and muscle cells, although the formation of these structures is significantly delayed (Sweet et al. 1999). By analyzing the phenotypes of chimeras, composed of animal caps and experimentally manipulated micromeres or mesomeres, we show here that micro1 is necessary and sufficient for both micromere differentiation and mid/hindgut-inducing activity.
Materials and Methods
Animals and embryos
Adult H. pulcherrimus were collected near the Noto Marine Laboratory, Kanazawa University, and near the Tateyama Marine Laboratory, Ochanomizu University. Gametes were obtained by intracoelomic injection of 0.5 M KCl. The embryos were cultured at 15°C in Jamarin U artificial seawater (ASW, Jamarin Laboratory).
Isolation of HpSoxB1 cDNA
Degenerate PCR primers were designed that correspond to the conserved regions of the SRY (Sex-determining region Y) and Sox HMG boxes: 5′-ATGAAYGCNTTYATGGT-3′ coding for MNAFMV and 5′-CCYTTNARRTGYTGRTA-3′ coding for YQHLNG, respectively. PCR was performed under standard conditions with 5 μM each of the primers, using cDNA from the cleavage stage embryo as a template. Products of the expected size (∼0.3 kb) were cloned to the pT7/T vector (Nonagen). The sequence has been deposited as HpSoxB1 in the DNA Data Bank of Japan (DDBJ) database (accession number AB206097), which was 98% identical with nucleotides 245–541 of SpSoxB1 cDNA (accession number NM 214474), and the deduced amino acid sequence of which matched, except one, to residues 71–169 of SpSoxB1.
Reverse transcriptase-polymerase chain reaction
To estimate the expression levels of marker genes in embryos that had been injected with mRNA encoding a VP16 activation domain/micro1 homeodomain fusion protein, semiquantitative reverse transcriptase–PCR (RT-PCR) was carried out using Thermo-Start Taq DNA polymerase (ABgene). Complementary DNA was synthesized from total RNA using ReverTra ACE (Toyobo) and random 9-mer oligonucleotides. The primer sequences for the PCR reactions were as follows: SoxB1-forward, 5′-ACCAGTTCTCCTGTTGC-3′; SoxB1-reverse, 5′-CGTTGAGGTGCTGGTA-3′; Ars-forward, 5′-ATGGTCGGAAAGTGGCA-3′; Ars-reverse, 5′-ACAGGAGAAATCGTCGCT-3′; Endo16-forward, 5′-GCAACTTCCGATCATGTTGT-3′; Endo16-reverse, 5′-GCGATTCTCCTTGTACTC-3′; Delta-forward, 5′-GTACGTGTCGCAATGAAG-3′; Delta-reverse, 5′-AACAGTGGTCACGGATCT-3′; SM50-forward, 5′-GGCTAGTCTTGTAGCCTT-3′; SM50-reverse, 5′-GGCGAATCCGTTAGGATA-3′; Ets-forward, 5′-CCTCCCATGCCATACTT-3′; Ets-reverse, 5′-GGACAGCTTGAATTCCCA-3′; Tbr-forward, 5′-AAGGCGTCGGTTTACCT-3′; Tbr-reverse, 5′-CCTTTGCAAATGGATTGTAGTC-3′; and Mit COI-forward, 5′-AGGCACAGCTATGAGTGT-3′; Mit COI-reverse, 5′-TCATCCAGTCCCTGCTC-3′.
Constructs for in vitro transcription
A modified Bluescript RN3 (Nishimura et al. 2004) was used to make the expression constructs, in which a terminator rrnB sequence had been inserted upstream of the T3 promoter. For the micro1 construct, a full-length micro1 D cDNA, containing the 5′ and 3′ untranslated regions (UTRs), was PCR-amplified and cloned between the EcoRI and NotI sites of the vector. The chimeric construct VP16AD/micro1HD was generated by fusing the sequence encoding the micro1 homeodomain (plus N-terminal 20 and C-terminal 6 residues, accession number AB072733) downstream of that encoding the VP16 activation domain (residues 411–455, accession number U89963). The fragment encoding the VP16 activation domain was excised from the pVP16 plasmid (Clontech) with BglII and EcoRI. The fragment encoding the micro1 homeodomain was PCR-amplified from micro1 D cDNA using the following primers: EcoRI-micro1, 5′-CCGGAATTCAGAATGGCGGATTACACCA-3′, and NotI-micro1-R, 5′-TAAAGCGGCCGCCTAAGCAGAGTTTGGAACAAGA-3′. The two fragments were ligated into the BglII, EcoRI, and NotI sites of the vector.
Synthesis and microinjection of synthetic mRNA and antisense oligonucleotides
Capped RNA was transcribed from linearized constructs using the mMESSAGE mMACHINE kit (Ambion) according to the manufacturer's protocol. The RNA was diluted to 0.1–2.5 pg/pl in 40% glycerol, and ∼3 pl of the solution was injected into each egg as described by Gan et al. (1990). The morpholino antisense oligonucleotides were obtained from Gene Tools. Mmicro1A, B/C, and D were complementary to sequences in the micro1 A, B/C, and D cDNAs, respectively. Their sequences and positions with respect to translational initiation were: Mmicro1A, 5′-TGGTGTAATCCGCCATTCTGATAAA-3′ (−9 to +16); Mmicro1B/C, 5′-GAGTGATCATGGTGTTATCTGCCAT-3′ (+1 to +25); and Mmicro1D, 5′-GAGTGATCATGGTGTAATCTGCCAT-3′ (+1 to +25). Morpholinos were dissolved in 40% glycerol, and 2–3 pl of a solution containing 1 mM of each type was injected individually or in combination into fertilized eggs to give a final concentration in the egg of ∼5 μM each.
Manipulation of embryos
Chimeric embryos were produced according to the method of Amemiya (1996). Transplanted blastomeres were stained with rhodamine B isocyanate (Sigma) to trace the lineage of cells. The chimeras were cultured in ASW containing 100 U/ml penicillin and 50 μg/ml streptomycin sulfate in dishes coated with 1.2% agar.
Immunostaining and histochemistry for alkaline phosphatase
The embryos were fixed for 30 min in ASW containing 1.4% formaldehyde, washed with ASW, and stored in 70% ethanol at 4°C. After washing three times with phosphate-buffered saline containing 0.1% Tween 20 (PBT), the embryos were incubated with P4 (PMC-specific) or Hpoe (oral ectoderm-specific) monoclonal antibody (diluted 1:200 or 1:500 in PBT, respectively) for 3 h at room temperature. The embryos were again washed three times with PBT followed by incubation at 37°C for 30 min, with the secondary antibody diluted 1:100 in PBT (Alexa Fluor 568 goat anti-mouse IgG; Molecular Probes). After a final wash, the embryos were observed with epifluorescence optics. Histochemistry for alkaline phosphatase was performed according to the method of Whittaker and Meedel (1989).
Whole-mount in situ hybridization
Whole-mount in situ hybridization was carried out according to the method of Arenas-Mena et al. (2002), using digoxigenin (DIG)-labeled probes of ∼3.2 kb derived from HpDelta cDNA (accession number AB211538), which was isolated by RT-PCR using an RNA LA PCR kit (TaKaRa). Complementary DNA was synthesized using total RNA from the hatched blastula stage embryos and the oligo(dT) adaptor primer. Three forward gene-specific primers were designed to the conserved regions: 5′-TGGCCGCGTGATTTTTCACTCGCCCTAGATGCC-3′ coding for WPRDFSLALDA between the signal peptide and the DSL domain; 5′-TGTATCCCGAAAGATGACCTCTTTGGGCAT-3′ coding for CIPKDDLFGH in the DSL domain; and 5′-GAAAGAGATCTGAACTACTGCACC-3′ coding for ERDLNYCT in the EGF repeat 3. Three-round nested PCR was performed under standard conditions using the three gene-specific primers and three adaptor primers equipped in the kit. Products of ∼3.2 kb were blunted with T4 DNA polymerase and cloned into the EcoRV site of the pBluescript (Stratagene).
Results
Morpholino-mediated micro1 knockdown
We previously isolated four similar but distinct micro1 cDNAs, designated as micro1 A through D, from a H. pulcherrimus library (Kitamura et al. 2002). To block micro1 function, we previously injected a mixture of morpholino antisense oligonucleotodes complementary to micro1 A, B/C, and D cDNAs (Mmicro1A+B/C+D) to give a final concentration of ∼5 μM each. The injected embryos typically exhibited a phenotype similar to that of micromere-less embryos, characterized by no PMC ingression and a delay in gastrulation (Nishimura et al. 2004). However, the occurrence of this phenotype was inconstant. Table 1 shows the frequency of occurrence of the micromere-less phenotype induced by injection of the morpholinos, individually or in combination, into embryos from a single mating. The morpholino Mmicro1D was more effective than Mmicro1A or Mmicro1B/C in producing the micromere-less phenotype, and the cocktail of all three morpholinos induced the micromere-less phenotype with the highest frequency. We found this tendency (cocktail>Mmicro1D>Mmicro1A, Mmicro1B/C) in ∼10 batches of injected embryos (data not shown). However, even the injection of the cocktail induced the micromere-less phenotype in a maximum of only ∼40% of the injected embryos from a mating of individuals collected on the coast of the Sea of Japan. In contrast, batches of embryos from individuals collected on the Pacific coast of Japan exhibited the micromere-less phenotype in less than 5% of the injected embryos; most of these embryos developed normally. These observations suggest that the fluctuation in the occurrence of the micromere-less phenotype may be attributable to polymorphisms in the multiple loci of micro1 (Nishimura et al. 2004). This fluctuation made it difficult to conduct loss-of-function assays especially in chimeric embryos.
Block of micro1 function with an active activator of micro1, VP16AD/micro1HD
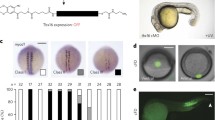
The micro1 encodes a transcription repressor that includes the homeodomain close to the N-terminus and two serine-rich repeats in the C-terminal region (Fig. 1a). The serine-rich repeat includes an octapeptide sequence similar to the eh-1/GEH domain in engrailed/goosecoid (Fig. 1a), which mediates interactive transcriptional repression (Smith and Jaynes 1996; Mailhos et al. 1998). Although a repression domain has not yet been identified in micro1, structural features suggest that one may be present in the C-terminus. Therefore, we designed a construct encoding a fusion protein that replaced this potential micro1 repression domain with the VP16 activation domain, placing it in frame with the N-terminus of micro1 (VP16AD/micro1HD), and used this construct to block micro1 function (Fig. 1a).
Block of micro1 function by expression of a fusion protein of the VP16 activation domain and the micro1 homeodomain (VP16AD/micro1HD), an active activator of micro1. a The structures of micro1 and VP16AD/micro1HD. Micro1 includes the homeodomain close to the N-terminus and two serine-rich repeats in the C-terminal region. The serine-rich repeat includes a sequence similar to the eh-1/GEH domain in engrailed/goosecoid that mediates interactive transcriptional repression. b, c Control embryos at 24 and 38 h after fertilization, respectively. e, f Embryos injected with mRNA encoding VP16AD/mciro1HD at 24 and 38 h after fertilization, respectively. PMC ingression and subsequent gastrulation did not occur in these embryos. d, g Control (left) and VP16AD/mciro1HD-injected (right) embryos at 3 days, immunostained using anti-Hpoe monoclonal antibody. Hpoe antigen is restricted to the oral ectoderm in pluteus larvae. The injected embryo remained a hollow blastula, and almost all blastomeres expressed Hpoe antigen. The phenotype is similar to, but more severe than, that produced by morpholino-mediated knockdown of micro1 expression in h. h A 3-day-old embryo injected with morpholinos, Mmicro1A+B/C+D. i A 3-day-old embryo injected with Mmicro1A+B/C+D and j a control larva, respectively, squashed for observation of the pigment cells by the method of Kominami (1998). The Mmicro1-injected embryo has few pigment cells
We injected ∼5 pg of synthetic mRNA encoding VP16AD/micro1HD into fertilized eggs. The injected embryos cleaved normally and developed into blastulae; however, PMC ingression and subsequent gastrulation did not occur (Fig. 1e,f). Three days after fertilization, the injected embryos remained hollow blastulae with practically no vegetal structures (Fig. 1d), whereas some endomesodermal structures were formed in embryos injected with morpholinos, Mmicro1A+B/C+D (Fig. 1i,j). In embryos injected with VP16AD/micro1HD mRNA, almost all of the blastomeres expressed the Hpoe antigen (Fig. 1d,g). The Hpoe antigen is zygotically expressed on the apical surface of all blastomeres during the cleavage stages and is subsequently lost from the PMC, archenteron, and aboral ectoderm; the expression of Hpoe is consequently restricted to the oral ectoderm (Yoshikawa 1997). The presence of Hpoe in the injected embryos indicates that the injection and translation of the RNA for the VP16AD/micro1HD fusion protein did not disrupt early developmental processes, at least those leading to the expression of Hpoe antigen, and suggests that the subsequent loss of Hpoe antigen expression did not occur in the injected embryos.
To further define the phenotype of embryos injected with mRNA encoding the VP16AD/micro1HD, we estimated the transcript levels of marker genes in the injected embryos by semiquantitative RT-PCR. Total RNA was extracted from embryos at 24 h after fertilization (Fig. 1b,e). In injected embryos, vegetal markers Endo16 (Akasaka et al. 1997), SM50 (Katoh-Fukui et al. 1992), Delta (this study), Ets (Kurokawa et al. 1999), and Tbr (Fuchikami et al. 2002) as well as an aboral ectoderm marker, Ars (Akasaka et al. 1990), were downregulated, whereas a nonvegetal/ectodermal marker, SoxB1 (this study), was upregulated (Fig. 2). These findings were consistent with the observed morphologies and the overall expression of the Hpoe antigen observed in the injected embryos.
Vegetal marker genes are downregulated in embryos injected with mRNA encoding VP16AD/micro1HD. The transcripts of marker genes were analyzed by RT-PCR, using total RNA extracted from one embryo at the mesenchyme blastula stage. The numbers below indicate the number of PCR cycles. In the injected embryos, the vegetal markers (Endo16, SM50, Delta, Ets, and Tbr) as well as an aboral ectoderm marker (Ars) were downregulated, whereas a nonvegetal marker (SoxB1) was upregulated
micro1 is necessary for both micromere differentiation and inductive signals
To determine whether micro1 is required for the inductive signals from the micromere, we constructed chimeric embryos composed of animal cap mesomeres from a normal embryo and a micromere quartet from an embryo that had been injected with mRNA encoding the VP16AD/micro1HD (Fig. 3f). The chimeras developed normally into blastulae; however, PMCs and archenterons did not form in the chimeras even 2 days after fertilization (6 of 6 cases, Fig. 3a,b), by which time, a control chimera, composed of an animal cap and normal micromeres, had developed into an early pluteus larva with skeletal structures and an induced gut (Fig. 3c). Three days after fertilization, the experimental chimeras were still hollow blastulae in which the micromere descendants formed a protruding part of the ectodermal wall opposite the apical plate (Fig. 3d). Almost all the blastomeres of the chimera, including the micromere descendants, expressed Hpoe antigen (Fig. 3e). Taken together with the observation of the micromere-less phenotype induced by the injection of the morpholinos, these findings led us to conclude that micro1 is necessary for both micromere differentiation and inductive signals.
The micro1 gene is necessary for micromere differentiation and inductive signal production. The development of chimeras composed of animal cap mesomeres from a normal embryo and a micromere quartet from an embryo injected with mRNA encoding VP16AD/micro1HD (f). a, b Lateral view of a chimera at 2 days. No vegetal structures were formed. The rhodamine-labeled micromere descendants constituted a part of the ectodermal wall opposite the apical plate with apical tuft (at). c Vegetal view of a control chimera composed of animal cap and normal micromeres at 2 days. It developed into a larva with spicules and an induced gut. d, e Lateral view of a chimera at 3 days. The micromere descendants constitute a protruding part of the blastula wall (arrowhead) opposite the apical plate (ap). Almost all blastomeres express Hpoe antigen. Note that chimeras are not polarized along the oral–aboral axis
micro1 is sufficient for micromere differentiation and mid/hindgut-inducing activity
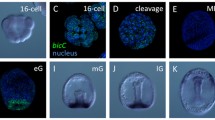
Gain-of-function experiments were conducted by the injection of ∼10 pg of synthetic micro1 D mRNA into fertilized eggs. In situ hybridization demonstrated that Delta mRNA was present in almost all blastomeres of the injected embryos, whereas it was restricted to a single region, probably the presumptive PMCs, in control embryos examined at the swimming blastula stage (Fig. 4a–d). At the time that PMC ingression occurred in the control embryos, almost all blastomeres of the injected embryos had been converted to the PMC phenotype (Fig. 4e–h).
Overexpression of micro1 endows nonvegetal blastomeres with micromere-like properties. a, b, e, f Control embryos. c, d, g, h Embryos injected with micro1 mRNA. a–d Whole mounts showing results of in situ hybridization using Delta probes. a, b Lateral (a) and vegetal (b) views of control swimming blastulae at 20 h. Delta mRNA was restricted to presumptive PMCs. c, d Injected embryos with micro1 at 20 h, hybridized with antisense (c) or sense probes (d). Almost all blastomeres expressed Delta mRNA. e, f Lateral views of a control embryo at 24 h, immunostained with P4 monoclonal antibody. g, h An injected embryo at 24 h. Almost all blastomeres expressed P4, a PMC-specific antigen
Using the same batch of embryos, we constructed chimeric embryos composed of animal cap mesomeres with a mesomere expressing micro1 (Fig. 5k). The volume of each mesomere was almost equivalent to that of four micromeres. At the time that PMC ingression occurred in the control embryos, most of the progeny of the micro1-expressing mesomeres in the chimeric embryos ingressed, like PMCs into the blastocoel (Fig. 5a,b), and subsequently formed a complete set of skeletal rods in the host (Fig. 5c,e). In addition, each chimera formed an archenteron, the tip of which included a minority of the clones from the transplanted mesomere (Fig. 5d,e). A control chimera, composed of an animal cap and a normal mesomere, remained a dauerblastula in which the mesomere progeny formed a part of the ectodermal wall opposite the apical plate (Fig. 5g,h). In the experimental chimera, at 3 days, the archenteron developed into a two-part gut (Fig. 5c): the first part (tip) included clones of the experimental, micro1-expressing mesomere, whereas the second did not (data not shown). Four days after fertilization, the chimeras developed into pluteus larvae with functional mid- and hindguts (Fig. 5i,j). However, the foregut and nonskeletogenic mesoderm, including pigment cells and coelomic pouches, were not formed in the larva (13 of 13 cases). From these observations, we concluded that micro1 is sufficient for micromere differentiation and mid/hindgut-inducing activity.
The micro1 gene is sufficient for micromere differentiation and mid/hindgut-inducing activity. The development of chimeras composed of animal cap mesomeres with a mesomere expressing micro1 (k). a, b Lateral views of a chimera at 24 h. Most of the rhodamine-tagged mesomere descendants have ingressed into the blastocoel. d, e Near-vegetal views of a chimera at 2.5 days. Spicules and archenteron were formed. The descendants of a mesomere lined the spicule and also constituted the tip of the archenteron (arrowhead). g, h Lateral views of a control chimera, composed of an animal cap with a normal mesomere at 2.5 days. The descendants of a mesomere constituted a part of the ectodermal wall opposite the apical plate with apical tuft (at) in a dauerblastula. c, f A chimera at 3 days, showing a two-part archenteron. A complete set of skeletal rods is observed with polarizing microscopy in f; anterolateral rod (alr), body rod (br), horizontal rod (hr), and postoral rod (por). i, j Dorsal (i) and lateral (j) views of a chimera at 4 days. Histochemical staining for alkaline phosphatase indicated that the gut has differentiated to functional stomach (st) and intestine (in). The stomach was in contact with the stomodeum (sd), but the mouth is not open; the esophagus or other mesodermal structures are not observed between the stomach and the stomodeum (arrowhead)
Discussion
Block of micro1 function with an active activator of micro1
Kurokawa et al. (1999) reported that ΔHpEts, which lacks the N-terminal activation domain, acts as a dominant-negative form of the Ets transcription factor in the sea urchin embryo. To conduct a loss-of-function assay for micro1, we first produced constructs encoding a truncated form of micro1A, B, C, or D protein, which lacks the C-terminal potential repression domain, and injected ∼8 pg of mRNA, encoding the truncated form into eggs. Although 10–20% of the injected embryos exhibited phenotypes similar to that of the micromere-less embryo, most of the embryos developed normally regardless of micro1 subtypes (data not shown). Therefore, we blocked micro1 function by injecting mRNA encoding an active activator of micro1 in the present study. The injected embryos remained hollow blastulae with no vegetal structures (Fig. 1). The phenotype was similar to, but more severely compromised than, that of micro1-knockdown embryos produced by the injection of morpholinos or that of micromere-less embryos, both of which exhibited delayed formation of some endomesodermal structures (Fig. 1h). The use of a fusion protein that reverses the function of transcriptional regulators (e.g., from repression to activation in the present case) often causes more severe defects than morpholino-mediated knockdown. For example, SpGsc is a transcriptional repressor that promotes oral ectoderm differentiation and antagonizes Otx, a ubiquitous activator of aboral ectoderm genes. Angerer et al. (2001) examined SpGsc function by using both a morpholino and a fusion protein composed of the VP16 activation domain with the Gsc homeodomain. Although both assays showed a downregulation of an oral ectoderm marker, concomitant with upregulation of an aboral ectoderm marker, the active activator caused more severe defects in ectoderm differentiation than the morpholino. In our study, the more severe developmental defects induced by the active activator of micro1 were unlikely to be attributable to nonspecific toxic effects, as the ubiquitous expression of Hpoe antigen and an upregulation of SoxB1 occurred in the VP16AD/micro1HD-injected embryos (Figs. 1, 2). SoxB1 is an animalizing transcription factor that distributes asymmetrically along the A-V axis and antagonizes nuclear β-catenin (Kenny et al. 1999, 2003; Angerer and Angerer 2003). Our observations indicate that the expression of the active activator of micro1 appears to have resulted in embryos that were more animalized than those formed after the removal of the micromeres or after the morpholino-mediated knockdown of micro1, although the factor(s) antagonizing micro1 has not yet been identified.
micro1 is sufficient for mid/hindgut induction
To examine whether micro1 is sufficient for the completion of micromere functions, we constructed chimeras composed of an animal cap host and a mesomere expressing micro1. The transplanted mesomeres gave rise to PMCs and endoderm, suggestive of polarity within the mesomere (see below). A plausible explanation for the formation of the gut in the chimeras is that clones that were converted to a PMC fate had induced the development of ectopic endoderm within the clones of the transplanted mesomere, which gave rise to the midgut, and which further induced the host mesomeres to generate the hindgut and a part of the midgut. Recently, Angerer and Angerer (2003) referred to a region of presumptive ectoderm that gives rise to the apical plate as the “animal pole domain” and described the domain-specific upregulation of NK2.1 (Takacs et al. 2004) and the downregulation of several ectodermal genes. The blastomeres in the animal pole domain are considered to be more resistant to experimentally enhanced canonical Wnt signaling than blastomeres in other regions (Angerer and Angerer 2003). We also observed that blastomeres responded differentially to the overexpression of micro1 along the A-V axis. The injection of micro1 mRNA into eggs increased the number of cells with the PMC phenotype in a dose-dependent manner and resulted in fate conversion from vegetal to animal blastomeres (data not shown). These observations led us to conclude that in transplanted mesomeres, the clones derived from the animal pole domain were converted to an endoderm fate by a signal(s) from the clones that had a PMC fate. Potential signaling pathways include the Delta/Notch pathway, as it has been shown to be sufficient to induce animal cells to form endoderm (Sherwood and McClay 2001; Sweet et al. 2002). However, the possibility is not ruled out that blastomeres in the animal pole domain were autonomously specified to form endoderm.
micro1 may not fulfill all micromere functions
As shown in Fig. 5, chimeras composed of animal cap mesomeres with a mesomere expressing micro1 did not form foreguts or nonskeletogenic mesoderm. On the other hand, a chimera composed of a micromere-less host with a mesomere expressing Pmar1 developed into a larva with a complete gut and a full range of SMCs (Oliveri et al. 2003). Although the difference in phenotypes between the two chimeras may be simply owing to species differences, it is more likely attributable to differences in the hosts, i.e., differences between animal caps and micromere-less hosts. Sweet et al. (1999) have shown that animal caps are less responsive to micromere signals than micromere-less hosts; fewer SMCs are induced in the former than in the latter by the transplantation of micromeres. Two groups have provided a line of strong evidence that the Delta/Notch signaling pathway plays a central role in SMC specification in the sea urchin embryo (Sherwood and McClay 1997, 1999, 2001; Sweet et al. 1999, 2002; McClay et al. 2000). The injection of either Pmar1 or micro1 mRNA into eggs results in the conversion of almost all blastomeres to PMC-phenotype cells that express Delta mRNA (Oliveri et al. 2002; Fig. 4 of this study). Ectopic Delta expression is consistent with the formation of SMCs in a chimera of a micromere-less host with a mesomere expressing Pmar1 (Oliveri et al. 2003). However, no SMC-derived structures were formed in chimeras in the present study.
There are at least two possible explanations for the deficiency of SMC formation in the chimeras in our study. First, the amount of Delta expressed in the clones of the transplanted mesomere might have been insufficient to induce the host cells to generate SMCs, as animal caps are less responsive to micromere signals than micromere-less hosts (see above). Sweet et al. (2002) demonstrated that a mesomere expressing Delta induced mesoderm formation when combined with an animal cap. This induction of mesoderm may be attributable to higher levels of Delta ligands in clones of mesomeres injected with Delta mRNA than in clones of mesomeres injected with micro1 mRNA. However, normal micromeres induce mesoderm in animal caps, and a mesomere injected with micro1 mRNA behaved like normal micromeres in terms of differentiation. These observations suggest the presence of a distinct factor(s) in animal blastomeres that may downregulate micro1-to-Delta production and/or Delta-mediated signaling. Alternatively, micromeres may produce a factor(s) that maintains the gene network and/or cooperatively enhances signaling. It has been shown that Krl and Wnt8, both of which are involved in endomesoderm development, are zygotically activated in vegetal blastomeres, including micromeres, in a β-catenin-dependent manner (Howard et al. 2001; Minokawa et al. 2004; Wikramanayake et al. 2004). Secondly, the lack of mesoderm in the chimeras examined in this study may be explained by the insulation of Delta-mediated signaling from the animal cap hosts. In our chimeras, the transplanted mesomere gave rise to PMCs and endoderm. As we discussed above, endoderm appears to be derived from the animal pole domain. McClay et al. (2000) demonstrated that macromeres require nuclear β-catenin to respond to micromere signals. The animal pole domain is postulated to be a region that lacks maternal mechanisms for the production of nuclear β-catenin (Angerer and Angerer 2003). Combined, these observations suggest that Delta/Notch signaling might be insulated from the host cells within the transplanted mesomere by Delta-insensitive clones derived from the animal pole domain.
In the present study, we showed that micro1 is necessary and sufficient for both micromere differentiation and partial endoderm-inducing activity. However, the ectopic expression of micro1 did not endow transplanted mesomeres with all micromere functions, suggesting that micro1 alone may not be sufficient for the full inducing activity. To understand the molecular mechanisms of patterning along the A-V axis, including micromere specification, we consider it important to identify the other factor(s) in micromeres whose existence is suggested by our experiments and to illuminate the molecular nature that subdivides mesomeres into the animal pole and preectoderm domains.
References
Akasaka K, Ueda T, Higashinakagawa T, Yamada K, Shimada H (1990) Spatial pattern of arylsulfatase mRNA expression in sea urchin embryo. Dev Growth Differ 32:9–13
Akasaka K, Uemoto H, Wilt F, Mitsunaga-Nakatsubo K, Shimada H (1997) Oral–aboral ectoderm differentiation of the sea urchin embryos is disrupted in response to calcium ionophore. Dev Growth Differ 39:373–379
Amemiya S (1996) Complete regulation of development throughout metamorphosis of sea urchin embryos devoid of macromeres. Dev Growth Differ 38:465–476
Angerer LM, Angerer RC (2003) Patterning the sea urchin embryos: gene regulatory networks, signaling pathways, and cellular interactions. Curr Top Dev Biol 53:159–198
Angerer LM, Oleksyn DW, Levine AM, Li X, Klein WH, Angerer RC (2001) Sea urchin goosecoid function links fate specification along the animal–vegetal and oral–aboral embryonic axes. Development 128:4393–4404
Arenas-Mena C, Cameron RA, Davidson EH (2000) Spatial expression of Hox cluster genes in the ontogeny of a sea urchin. Development 127:4631–4643
Di Bernardo M, Russo R, Oliveri P, Melfi R, Spinelli G (1995) Homeobox-containing gene transiently expressed in a spatially restricted pattern in the early sea urchin embryo. Proc Natl Acad Sci U S A 92:8180–8184
Emily-Fenouil F, Ghiglione C, Lhomond G, Lepage T, Gache C (1998) GSK3beta/shaggy mediates patterning along the animal–vegetal axis of the sea urchin embryo. Development 125:2489–2498
Ettensohn CA, Illies MR, Oliveri P, De Jong DL (2003) Alx1, a member of the Cart1/Alx3/Alx4 subfamily of paired-class homeodomain proteins, is an essential component of the gene network controlling skeletogenic fate specification in the sea urchin embryo. Development 130:2917–2928
Fuchikami T, Mitsunaga-Nakatsubo K, Amemiya S, Hosomi T, Watanabe T, Kurokawa D, Karaoka M, Harada Y, Satoh N, Kusunoki S, Takata K, Shimotori T, Yamamoto T, Sakamoto N, Shimada H, Akasaka K (2002) T-brain homologue (HpTb) is involved in the archenteron induction signals of micromere descendant cells in the sea urchin embryo. Development 129:5205–5216
Gan L, Wessel GM, Klein WH (1990) Regulatory elements from the Spec genes of Strongylocentrotus purpuratus yield different spatial patterns with a lacZ reporter gene. Dev Biol 142:346–359
Hörstadius S (1973) Experimental embryology of echinoderms. Clarendon, Oxford
Howard EW, Newman LA, Oleksyn DW, Angerer RC, Angerer LM (2001) SpKrl: a direct target of beta-catenin regulation required for endoderm differentiation in sea urchin embryos. Development 128:365–375
Katoh-Fukui Y, Noce T, Ueda T, Fujiwara Y, Hashimoto N, Tanaka S, Higashinakagawa T (1992) Isolation and characterization of cDNA encoding a spicule matrix protein in Hemicentrotus pulcherrimus micromeres. Int J Dev Biol 36:353–361
Kenny AP, Kozlowski DJ, Oleksyn DW, Angerer LM, Angerer RC (1999) SpSoxB1, a maternally encoded transcription factor asymmetrically distributed among early sea urchin blastomeres. Development 126:5473–5483
Kenny AP, Oleksyn DW, Newman LA, Angerer RC, Angerer LM (2003) Tight regulation of SoxB factors is required for patterning and morphogenesis in sea urchin embryos. Dev Biol 261:412–425
Kitamura K, Nishimura Y, Kubotera N, Higuchi Y, Yamaguchi M (2002) Transient activation of the micro1 homeobox gene family in the sea urchin (Hemicentrotus pulcherrimus) micromere. Dev Genes Evol 212:1–10
Kominami T (1998) Role of cell adhesion in the specification of pigment cell lineage in embryos of the sea urchin, Hemicentrotus pulcherrimus. Dev Growth Differ 40:609–618
Kurokawa D, Kitajima T, Mitsunaga-Nakatsubo K, Amemiya S, Shimada H, Akasaka K (1999) HpEts, an ets-related transcription factor implicated in primary mesenchyme cell differentiation in the sea urchin embryo. Mech Dev 80:41–52
Logan CY, Miller JR, Fercowicz MJ, McClay DR (1999) Nuclear β-catenin is required to specify vegetal cell fates in the sea urchin embryo. Development 126:345–357
Mailhos C, André S, Mollereau B, Goriely A, Hemmati-Brivanlou A, Desplan C (1998) Drosophila Goosecoid requires a conserved heptapeptide for repression of Paired-class homeoprotein activators. Development 125:937–947
McClay DR, Peterson RE, Range RC, Winter-Vann AM, Ferkowicz MJ (2000) A micromere induction signal is activated by β-catenin and acts through Notch to initiate specification of secondary mesenchyme cells in the sea urchin embryo. Development 127:5113–5122
Minokawa T, Amemiya S (1998) Mesodermal cell differentiation in echinoid embryos derived from the animal cap recombined with a quartet of micromeres. Zoolog Sci 15:541–545
Minokawa T, Rast JP, Arenas-Mena C, Franco CB, Davidson EH (2004) Expression patterns of four different regulatory genes that function during sea urchin development. Gene Expr Patterns 4:449–456
Nishimura Y, Sato T, Morita Y, Yamazaki A, Akasaka K, Yamaguchi M (2004) Structure, regulation, and function of micro1 in the sea urchin Hemicentrotus pulcherrimus. Dev Genes Evol 214:525–536
Okazaki K (1975a) Spicule formation by isolated micromeres of the sea urchin embryo. Am Zool 15:567–581
Okazaki K (1975b) Normal development to metamorphosis. In: Czihak G (ed) The sea urchin embryo. Springer, Berlin Heidelberg New York, pp 177–233
Oliveri P, Carric DM, Davidson EH (2002) A regulatory network that directs micromere specification in the sea urchin embryo. Dev Biol 246:209–228
Oliveri O, Davidson EH, McClay DR (2003) Activation of pmar1 controls specification of micromeres in the sea urchin embryo. Dev Biol 258:32–43
Ransick A, Davidson EH (1993) A complete second gut induced by transplanted micromeres in the sea urchin embryo. Science 259:1134–1138
Ransick A, Davidson EH (1995) Micromeres are required for normal vegetal plate specification in sea urchin embryos. Development 121:3215–3222
Sherwood DR, McClay DR (1997) Identification and localization of a sea urchin Notch homologue: insights into vegetal plate regionalization and Notch receptor regulation. Development 124:3363–3374
Sherwood DR, McClay DR (1999) LvNotch signaling mediates secondary mesenchyme specification in the sea urchin embryo. Development 126:1703–1713
Sherwood DR, McClay DR (2001) LvNotch signaling plays a dual role in regulating the position of the ectoderm–endoderm boundary in the sea urchin embryo. Development 128:2221–2232
Smith SJ, Jaynes JB (1996) A conserved region of engrailed, shared among all en-, gsc-, Nk1-, Nk2- and msh-class homeoproteins, mediates active transcriptional repression in vivo. Development 122:3141–3150
Sweet HC, Hodor PG, Ettensohn CA (1999) The role of micromere signaling in Notch activation and mesoderm specification during sea urchin embryogenesis. Development 126:5155–5265
Sweet HC, Gehring M, Ettensohn CA (2002) LvDelta is a mesoderm-inducing signal in the sea urchin embryo and can endow blastomeres with organizer-like properties. Development 129:1945–1955
Takacs CM, Amore G, Oliveri P, Poustka AJ, Wang D, Burke RD, Peterson KJ (2004) Expression of an NK2 homeodomain gene in the apical ectoderm defines a new territory in the early sea urchin embryo. Dev Biol 269:152–164
Vonica A, Weng W, Gunbiner BM, Venuti JM (2000) TCF is the nuclear effector of the beta-catenin signal that patterns the sea urchin animal–vegetal axis. Dev Biol 217:230–243
Whittaker JR, Meedel TH (1989) Two histospecific enzyme expressions in the same cleavage-arrested one-celled ascidian embryos. J Exp Zool 250:168–175
Wikramanayake AH, Huang L, Klein WH (1998) β-Catenin is essential for patterning the maternally specified animal–vegetal axis in the sea urchin embryo. Proc Natl Acad Sci U S A 95:9343–9348
Wikramanayake AH, Peterson R, Chen J, Huang L, Bince JM, McClay DR, Klein WH (2004) Nuclear β-catenin-dependent Wnt8 signaling in vegetal cells of the early sea urchin embryo regulates gastrulation and differentiation of endoderm and mesoderm cell lineages. Genesis 39:194–205
Yamaguchi M, Kinoshita T, Ohba Y (1994) Fractionation of micromeres, mesomeres, and macromeres of 16-cell stage sea urchin embryos by elutriation. Dev Growth Differ 36:381–387
Yoshikawa S (1997) Oral/aboral ectoderm differentiation of the sea urchin embryo depends on a planar or secretory signal from the vegetal half. Dev Growth Differ 39:319–327
Acknowledgements
We thank Hiroshi Kubota for the Hpoe monoclonal antibody, Hideki Katow for the P4 monoclonal antibody, and Masato Kiyomoto for supplying the mature adult H. pulcherrimus.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by N. Satoh.
A. Yamazaki and R. Kawabata contributed equally to this study
Rights and permissions
About this article
Cite this article
Yamazaki, A., Kawabata, R., Shiomi, K. et al. The micro1 gene is necessary and sufficient for micromere differentiation and mid/hindgut-inducing activity in the sea urchin embryo. Dev Genes Evol 215, 450–459 (2005). https://doi.org/10.1007/s00427-005-0006-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00427-005-0006-y