Abstract
Main conclusion
Soil compaction reduces root exploration in chickpea. We found genes related to root architectural traits in chickpea that can help understand and improve root growth in compacted soils.
Abstract
Soil compaction is a major concern for modern agriculture, as it constrains plant root growth, leading to reduced resource acquisition. Phenotypic variation for root system architecture (RSA) traits in compacted soils is present for various crops; however, studies on genetic associations with these traits are lacking. Therefore, we investigated RSA traits in different soil compaction levels and identified significant genomic associations in chickpea. We conducted a Genome-Wide Association Study (GWAS) of 210 chickpea accessions for 13 RSA traits under three bulk densities (BD) (1.1BD, 1.6BD, and 1.8BD). Soil compaction decreases root exploration by reducing 12 RSA traits, except average diameter (AD). Further, AD is negatively correlated with lateral root traits, and this correlation increases in 1.8BD, suggesting the negative effect of AD on lateral root traits. Interestingly, we identified probable candidate genes such as GLP3 and LRX for lateral root traits and CRF1-like for total length (TL) in 1.6BD soil. In heavy soil compaction, DGK2 is associated with lateral root traits. Reduction in laterals during soil compaction is mainly due to delayed seedling establishment, thus making lateral root number a critical trait. Interestingly, we also found a higher contribution of the GxE component of the number of root tips (Tips) to the total variation than the other lateral traits. We also identified a pectin esterase, PPE8B, associated with Tips in high soil compaction and a significantly associated SNP with the relative change in Tips depicting a trade-off between Tips and AD. Identified genes and loci would help develop soil-compaction-resistant chickpea varieties.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Chickpea (Cicer arietinum L.) (2n = 16) is the second most widely grown pulse crop with high nutritional benefits contributing to global food security (Varshney et al. 2013, 2019; Madurapperumage et al. 2021). An alarming increase in the human population in concert with changing environmental conditions threatens crop growth and development (Satterthwaite et al. 2010; Rehman and Khan 2019). To meet the growing demand for food production, agriculture is being modernized. Heavy farm machinery used more frequently negatively impacts soil quality and root growth. It causes soil mechanical impedance or compaction in response to physical consolidation. Soil compaction reduces soil porosity, restricts water and nutrient infiltration, reduces aeration, creates anaerobic conditions, destroys structure, and limits root growth (Schneider et al. 2021). These effects restrict seed germination and root exploration of soil, thus affecting global grain yield production (Ishaq et al. 2001; Lipiec et al. 2012; Cambi et al. 2018).
Roots undergo adaptive growth responses to deal with compacted soils and penetrate cracks. Compact soil significantly impacts root phenotype (Lynch et al. 2014; Vanhees et al. 2020). In such soil, roots tend to become thicker, stunted, and more resistant to buckling and deflection when encountering rugged terrain (Whiteley et al. 1982; Jin et al. 2013). Thicker roots are related to higher penetration pressure (Popova et al. 2016); however, in recent reports, an increase in root diameter is associated with lower penetration (Huang et al. 2022). Nevertheless, this increase in diameter can provide an advantage in terms of higher water uptake, compensating for reduced root length and surface area (Lynch et al. 2014).
Apart from root morphological changes, the plant undergoes various anatomical adaptations. Predicting root penetration and strength in maize is better accomplished by examining cortical cell wall thickness, area, cortical cell count, and stele diameter rather than root diameter (Chimungu et al. 2015). Also, smaller outer cortical cells improve hard soil penetration (Chimungu et al. 2015). Maize plants with deeper roots tend to have thinner layers of cortical cells from the third node regions and increased aerenchyma in the fourth node regions (Vanhees et al. 2022). Similar variations have been observed in other crops such as rice, soybean, wheat, and maize (Chimungu et al. 2015; Vanhees et al. 2020; Schneider et al. 2021; Mondal et al. 2022). Further, the root cortex develops anatomical modifications such as aerenchyma and multiseriate sclerenchyma (MCS) to increase penetration and withstand external pressure (Schneider et al. 2021). These root morphological and anatomical responses in compacted soil are under the strict control of molecular and hormonal regulations (Pandey et al. 2021).
Root growth response is conventionally thought to be the inability of the root to overcome the high external pressure of mechanical impedance (Passioura 2002; Correa et al. 2019), which is recently found to be the complex phytohormonal response (Pandey et al. 2021). Compacted soil restricts ethylene diffusion to the nearby soil, thus accumulating ethylene near plant roots (Pandey et al. 2021). The plant senses this accumulated ethylene as the signal for compacted soil, leading to root extension inhibition and favoring radial swelling (Huang et al. 2022). Ethylene-response-mutants of rice and Arabidopsis were narrow and could penetrate better in compacted soil. Ethylene further deploys auxin and abscisic acid (ABA) as downstream signals to modify rice root cell elongation and radial expansion, causing root tips to swell and reducing their ability to penetrate compacted soil (Huang et al. 2022). ABA causes cortical cell radial expansion. Here, rice mutants of ABA biosynthetic genes have attenuated cortical cell radial expansion in compacted soil. Auxin further aids by altering the cell elongation as osaux1 mutants penetrate compacted soil better than the wild-type roots and do not exhibit cortical cell radial expansion (Huang et al. 2022). Despite the molecular players known in model crops, there is still limited understanding of the root responses in compacted soil for legumes, especially in one of the essential commercial legumes, chickpea.
Harnessing natural variation in root system architecture (RSA) under compaction is an effective strategy for dissecting the root response of chickpeas in compacted soil (Thudi et al. 2014; Varshney et al. 2019; Lynch 2022). Gene banks worldwide have conserved over 90,000 chickpea accessions, yet the full extent of the crop’s diversity remains unexplored due to the limited availability of phenotypic data for crucial adaptive traits (Upadhyaya et al. 2011). To identify genetic variation associated with chickpeas’ ability to penetrate compact soil, we investigated 210 chickpea accessions in normal and two levels of compacted soil. The study aims to find the genetic variants responsible for higher root growth irrespective of compaction. We performed a Genome-Wide Association Study (GWAS) with RSA traits for variant identification. In the present study, we have identified significant associations and underlying genes responsible for root architecture modification in response to soil compaction. This information could benefit future research and breeding programs, as it provides a foundation for understanding the responsiveness and genetic basis of chickpea root architectural traits in compacted soil.
Materials and methods
Plant material and growth conditions
The present study comprises 210 diverse chickpea (Cicer arietinum L.) accessions from the chickpea mini-core collection conserved at gene banks (Suppl. Table S1). These chickpea mini-core accessions are also a part of the 3 K pangenome project containing 3600 genotypes (Varshney et al. 2021). The mini-core panel comprises diverse accessions from more than 25 countries and regions, including commercial varieties and landraces (Suppl. Table S1). The experiment was set up at the National Institute of Plant Genome Research, New Delhi, from November 2022 to January 2023 in a polyhouse with natural conditions. Before sowing, chickpea seeds were overnight soaked in water. The soaked seeds were planted in cylindrical PVC pipes with 5.7, 7.5, and 8.3 kg of sandy loam soil filled up to 26 cm in height. The chickpea association panel was phenotyped for three soil compaction treatments, i.e., zero compacted soil treatment with 5.7 kg soil and 1.1 bulk density g/cm3 (BD), moderately compacted soil stress treatment with 7.5 kg soil and 1.6 BD g/cm3, and heavily compacted soil stress treatment with 8.3 kg soil and 1.8 BD g/cm3 compaction of soil stress in five replications for 9 days (Suppl. Fig. S1 a-j). The field soil was initially screened and filtered using a 2 mm hole-size iron construction sand sieve mesh (1.3 × 2 m). Initially, 15 % soil moisture was adjusted, and throughout the experiment, equal watering was done in all the treatments. The soil compaction was carried out using a hand-held soil compaction proctor.
Phenotyping of chickpea accessions for root penetration and root system architecture-related traits under compacted soil
Nine days after germination (DAG), chickpea seedlings were carefully removed from the compacted soil without causing any damage to the roots (Suppl. Fig. S1 f–j). Washed chickpea seedlings were fixed in FAA fixative solution comprising formaldehyde 10 % + ethanol 50 % + acetic acid 5 % + 35 % water (by vol.) (Ruzin 1999). Chickpea seedlings were scanned using EPSON Perfection V850 Pro scanner, and traits were measured using winRHIZHO software (Le Marié et al. 2014). Here, a total of 13 root architecture traits were considered, including total root length (TL_cm), total root surface area (SA_cm2), the average diameter (AD_mm), total root volume (RV_cm3), number of tips, primary root length (PRL_cm), lateral root length (LRL_cm), primary root surface area (PRSA_cm), lateral root surface area (LRSA_cm), primary root projected area (PRPA), lateral root projected area (LRPA), primary root volume (PRVol), and lateral root volume (LRVol). The raw root architecture data were subjected to pre-processing for outlier removal. The clean data was used for ANOVA and correlation analysis using R. Exploratory data analysis of phenotypic traits were performed using R and MS Excel, and boxplots and bar graphs were generated using ggplot2 (Wickham 2011). Correlation plots were generated using the corrplot package (Wei et al. 2017). G x E analysis using a mixed model was performed using statgenGxE and statgenSTA packages (van Rossum et al. 2017). Additionally, plasticity indexes were calculated for all 13 traits by calculating the relative change between compacted (1.6 BD and 1.8 BD) and non-compacted (1.1 BD) soil.
The chickpea whole-genome sequencing (WGS) data retrieval, quality checks, and genome-wide variant detection
The whole-genome sequencing (WGS) data of 210 chickpea accessions were retrieved from the NCBI sequence reads archive (SRA) database (PRJNA657888) (Varshney et al. 2021). Further, SRA data for 210 mini-core chickpea accessions was processed with SRA Toolkit 3.0.5 (Leinonen et al. 2010) and faster-dump to get fastq paired-end reads. Paired-end reads were cleaned with AfterQC 0.9.7, (Chen et al. 2017), and quality was checked using FastQC (version 0.11.9) (Brown et al. 2017). The reads were mapped to the chickpea CDC frontier reference genome (Varshney et al. 2013) using BWA (version 0.7.12) (Li 2013). The SAM files were converted to BAM files using samtools (version 1.18) (Danecek et al. 2021). The variant calling was performed using the Genome Analysis Toolkit (GATK version 4.4.0.0) program’s HaplotypeCaller function (McKenna et al. 2010). The variant calling files (VCFs) were merged using the CombinedGVCF function in GATK. The combined VCF was filtered for missing percentage (keeping sites present in 90 % of genotypes), Minimum mapping quality (mQ = 30), read depth, and minor allele frequency (maf > 0.05) using vcftools (version 4.2) (Danecek et al. 2011; Li 2011). The heterozygous calls were converted to missing using TASSEL software, and sites were filtered for maf > 0.05 and site minimum count of 95 % to identify high-quality SNPs for association analysis (Bradbury et al. 2007).
Phylogenetic diversity and population structure
The population structure of 210 chickpea accession was performed using the maximum likelihood method in Admixture 1.30 (Alexander and Lange 2011). The Admixture K values were set to K = 1 to 10, and each step iterated ten times with 2000 bootstrapping. A phylogenetic tree was constructed for 210 chickpea accessions using the neighbor-joining method (NJ) using TASSEL and R package tree view. Further, genetic principal components analysis was calculated, and a plot was generated using SNPRelate and GAPIT R packages (Lipka et al. 2012; Wang and Zhang 2021).
GWAS to identify significant associations for root architecture traits in compacted soil
The GWAS was performed using the BLINK statistical model in the GAPIT R package (Lipka et al. 2012). Significant loci obtained from the GAPIT (based on Bonferroni correction) were considered for further analysis. Out of all the significant loci, only those showing significant allelic effects were considered for further identification of candidate genes. For the identification of candidate genes, we identified linkage disequilibrium (LD) blocks within the 300 kb region from the significant loci, i.e., 150 kb upstream and downstream of the significant loci, based on the average LD decay of the chickpea genome (Upadhyaya et al. 2016). Further, LD was calculated using the full matrix option in TASSEL (Bradbury et al. 2007) of the LD block containing the significant loci. Within the LD block, only those SNPs in LD (R2 ≥ 0.8) with significant loci were considered further along with significant SNP. Genes adjoining (within 2 kb) of the SNPs were considered as potential candidate genes for further analysis. Only those genes were considered that are in the same LD block as the significant loci. The LD blocks were estimated, and plots were generated using LDBlockShow (version 1.40) (Dong et al. 2021).
Gene expression analysis using qRT-PCR
Chickpea accession ICC4958 was used to study the expression of different putative genes playing a role under soil compaction. Root samples were taken at 9 DAG from three soil compaction treatments (1.1 BD, 1.6 BD, and 1.8 BD). Tissues were frozen in liquid nitrogen and stored at − 80 °C. High-quality RNA was extracted from the chickpea root tissues using the TRIZOl method (Ambion by Life Technologies, Carlsbad, CA, USA). Isolated RNA was subjected to DNase treatment (DNase I, RNase-free (1 U/µL), Thermo Scientific), and cDNA was prepared using a High-Capacity cDNA Reverse Transcription Kit (Applied Biosystems). Primers used in the study were designed from the CDS region of genes using NCBI Primer-BLAST (Ye et al. 2012), preferably from the intron–exon junction, and best-suited ones with minimum self-complementarity index were selected (cf Suppl. Table S7). Gene expression profiling was performed using Applied Biosystems 7900 Fast Real-Time PCR machine. The relative expression of genes (Log2 fold change) in roots under compaction treatments (BD 1.6 and BD1.8) with respect to non-compacted soil (BD1.1) was calculated using the ΔΔCt method (Livak and Schmittgen 2001). The expression of Initiation factor 4 alpha (IF4α) (a housekeeping gene) was taken as the endogenous control as per previous studies for comparing stress samples in chickpea (Garg et al. 2010). The experiment was performed on three biological replicates.
Results
The phenotypic variation of chickpea root system architecture traits response under compacted soil
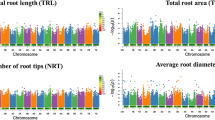
Chickpea accessions were phenotyped for 13 RSA traits under three levels of soil compactions with five replications. Comparing RSA traits between treatments, it was observed that, except for average diameter (AD), all remaining RSA traits showed a decrease in their values with increasing soil compaction (Suppl. Table S2; Fig. 1a–i). In 1.1 BD soil, the population mean of AD was 1.06 mm, which was increased to 1.25 mm in 1.6 BD soil and 1.29 mm in 1.8 BD soil. Contrastingly, the population mean of total length (TL) in 1.1 BD was 27.27 cm, decreasing to 11.33 cm in 1.6 BD soil and 8.61 cm in 1.8 BD soil. Lateral root length was higher in uncompacted soil, but this trend is reversed in compacted soil. Thus, compacted soil delays RSA establishment, resulting in shorter overall root length and increased diameter.
Phenotyping of chickpea mini-core GWAS panel for root system architecture (RSA) traits under three treatments (1.1BD, 1.6 BD, and 1.8 BD) of compacted soil. Box plots depict phenotypic variation and the difference between the three treatments for total root length (a, TL) root surface area (b, SA), average root diameter (c, AD), root volume (d, RV), number of root tips (e, Tips), (f, PRL), lateral root length (g, LRL), primary root surface area (h, PRSA), lateral root surface area (i. LRSA). The central line in the box plots depicts the population median for the trait in a particular treatment. Here, except AD, all the remaining traits decrease with increasing soil compaction. Comparison between treatments were analyzed using ANOVA followed by a post-hoc test (P < 0.05) (n = 210). Significant comparisons were depicted using *. The * indicate level of significance as, *P < 0.05, **P < 0.01, ***P < 0.001
All the traits showed significant genotype, environment, and genotype X environment (G × E) variation (Suppl. Table S3). Maximum G × E contribution was shown by root volume (RV) and primary root volume (PRVol), followed by AD. Length traits displayed the least G x E contribution, suggesting low G × E variation in these traits, depicting lower variation in plasticity. By comparing variance with increasing soil compaction, we observed a reduction in variance in all the traits except AD. Higher G × E contribution and increased variance with soil compaction in AD depicted higher genetic variation in response to increasing diameter. Similarly, plasticity indexes of total RV and PRVol showed high variance, followed by PRPA, PRSA, and AD. Interestingly, a high variance was only present in plasticity indexes for 1.8 BD soil, not 1.6 BD soil, depicting higher plasticity variation in 1.8 BD soil compared to 1.6 BD soil (Suppl. Table S4).
In 1.1 BD soil, total length (TL) showed a weak positive correlation for primary root traits and a strong positive correlation for lateral root traits. However, surface area (SA) displayed the opposite trend, depicting laterals contributing more towards total root length and primary root contributing more towards total root surface area. Primary and lateral root traits showed no correlation with each other in 1.1 BD soil. The AD showed a negative correlation between lateral root and branching traits but displayed a positive correlation for primary root traits. In 1.6 BD and 1.8 BD soils, TL and SA showed similar trends in correlation with primary and lateral root traits. However, a weak positive correlation is observed among lateral and primary root traits in 1.6 BD soil, which vanishes in 1.8 BD soil (Suppl. Figs. S2a–c and S3a–f).
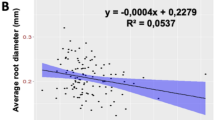
Interestingly, the positive correlation between AD and primary root length (PRL) in 1.1 BD soil vanishes in 1.6 BD and 1.8 BD soils. It suggested that roots with larger diameter are not related to longer primary roots in moderate and heavy soil compaction, which were in uncompacted soil. As 1.1 BD soil, AD negatively correlates with lateral and branching traits in 1.6 BD and 1.8 BD soils. However, the correlation is weaker in 1.8 BD soil. This suggests a trade-off between lateral root development and average diameter in compacted soil. The resources required for increasing diameter in compacted soil could possibly deter resource availability for the lateral root development (Suppl. Figs. S2a–c and S3a–f).
Phylogenetic diversity and population structure
After variant calling and filtering, 238,240 single-nucleotide polymorphisms (SNPs) were extracted to perform GWAS with RSA traits in response to compact soil (Fig. 2a). The extracted SNPs were used to study population structure and genomic variation in the GWAS panel. Population admixture showed an optimum K-value of 9 and was selected based on the minimum cross-validation error, which was fixed to retain pure accessions by eliminating admixtures (Fig. 2b, c). The bar plot designates the distribution of nine chickpea population in six geographical groups. Asia region comprises the highest number of accessions belonging to nine population clusters. Further, phylogenetic diversity analysis and genetic principal component analysis (PCA) obtained a similar clustering pattern, where PC1 and PC2 explained 19.04 % and 10.57 % variation, respectively (Fig. 2d–e).
The phylogenetic variation, population structure, and genetic diversity of chickpea mini-core GWAS panel. a A density plot was created using high-quality SNPs to show the distribution of 200 k SNPs across the chickpea genome. Here, green indicates less SNP density and red indicates higher SNP density. b The population variation was identified using admixture software; the K value varies from 1 to 10 in a 10 iteration, and each iteration was run at 2000 bootstrapping. The bar plot was plotted to identify the distribution of phylogenetic diversity and population STRUCTURE. The bar plot represents the K-9 population among the 210 min-core chickpea accessions. c The bar plot shows the distribution of chickpea accession in nine populations in six groups. d Dendrogram depicts the clustering of chickpea accessions into nine sub-populations. e Principal component analysis (PCA) of 210 chickpea accessions. Here, the PCA biplot (PC1 vs. PC2) depicts diversity in the population, where PC1 explains 19.04 % variation, and PC2 explains 10.57 % variation
Genome-wide association mapping of SNPs linked with root penetration and RSA-responsive traits
The GWAS was performed using the BLINK model to identify significantly associated SNPs for RSA traits under three different bulk densities soil. In control 1.1 BD soil, we have identified two significant SNPs at chromosome 4, i.e., S4_38707673 (P-2.09E–08) and S4_665844 (P-2.81E–06), linked with AD and TL traits (Suppl. Fig. S4a–d; Table 1). Their phenotypic variance explained (PVE %) percentage varies from 38.12 % to 29.86 %. In moderate 1.6 BD soil compaction treatment, we identified four significant SNPs: S7_32834852 (P-6.47E–07), S7_29444770 (P-1.92E–06), S8_3754663 (P-4.03E–06), and S7_29511774 (P-4.04E–06). The most significant SNP, S7_32834852, was associated with lateral root surface area (LRSA) (P-4.01E–06), lateral root volume (LRVol) (P-2.91E–06), LRPA (P-2.58E–06), TL (P-6.47E–07), and their PVE were 28.28 %, 63.04 %, 27.52 %, and 27.17 %, respectively (Fig. 3a–f; Suppl. Figs. S5 a-h; S6 a-f; S7a–f; Table 1). This was followed by S7_29444770, which was associated with lateral root length (LRL) (P-1.92E–06), lateral root projected area (LRPA) (P-2.58E–06), and LRSA (P-4.01E–06), with 61.26 %, 9.79 %, and 8.80 % PVE, respectively. The SNP S8_3754663 linked with primary root volume (PRVol) (P-4.03E–06), and SA (1.10E–06), and the PVE were 17.71 % and 15.14 %. The SNP, S7_29511774, was associated with TL (P-4.04E–06) with a PVE of 7.81 % (Figs. 4a–f; Suppl. Fig. 5a–h; S6a–f; S7a–f; Table 1).
Genome-wide association mapping of chickpea lateral root traits under moderately compacted soil (1.6 BD) using BLINK GWAS model. Four significant SNPs: S7_32834852, S7_29444770, S8_3754663, and S7_29511774 were identified associated with root traits in moderately compacted 1.6 BD soil. The SNP S7_32834852 is associated with lateral root surface area (LRSA), lateral root projected area (LRPA), lateral root volume (LRVol), and total root length (TL) in chr7 under 1.6 BD soil. a The Manhattan plot represents chromosome-wise SNP distribution and significant SNPs associated with LRSA. b Regional association of SNPs in chromosome 7 with LRSA. c The QQ plot shows the significant SNPs linked with LRSA. d The LD block was plotted around SNP S7_32834852 for the 300 kb region (upstream 150 kb and downstream 150 kb). The LD block plot shows the distribution of candidate genes in the 300 kb region and f genes linked with chickpea root-responsive traits. e The SNP S7_32834852 shows a significant allelic effect (C/T) for LRSA. In the Manhattan plot, a dotted line represents the threshold value calculated using FDR at P < 0.05, and a green line represents the Bonferroni threshold at P < 0.05. Comparison between two alleles is made using Student’s T test, and the P value of the test is depicted in the plot
Genome-wide association mapping of chickpea total root length (TL) under moderately compacted (1.6 BD) soil. The GWAS identified significant SNP, S7_29511774 associated with TL. a The Manhattan plot represents chromosome-wise SNP distribution and significant SNPs associated with TL. b Regional association of SNPs in chromosome 7 with TL. c The QQ plot shows significant SNPs linked with chickpea TL. d The LD block plotted around the SNP, S_29511774 within the 300 kb region. e Significant candidate genes lie in the 300 kb region and the same LD block with the significant SNP. f Box plot depict significant allelic effect (C/T) for TL. In the Manhattan plot, a dotted line represents the threshold value calculated using FDR at P < 0.05, and a green line represents the Bonferroni threshold at P < 0.05. Comparison between two alleles is made using Student’s T test, and the P value of the test is depicted in the plot
We identified seven significant SNPs in 1.8 BD soil (Suppl. Fig. S8 a-f; Suppl. Fig. S9 a-f; Suppl. Fig. S10 a-f; Suppl. Fig. S11a–d). The most significant SNP, S6_34682862, was associated with AD (P-4.21E–11), PRVol (P-6.53E–09), and total root volume RV (P-1.34E–09); their PVE varies between 59.81 %, 42.49 %, and 48.29 %, respectively. Followed by S6_6702892 SNP associated with LRPA (P-5.91E–08), LRSA (P-5.82E–08), and LRVol (P-5.89E–08), their PVE varies between 41.62 %, 42.24 %, and 39.84 %, respectively. The SNP S4_22818995 linked with PRL (P-9.64E–08) showed 39.07 % PVE. S4_22984656 was associated with PRPA (P-1.83E–08) and PRSA (P-1.47E–08) and showed a PVE of 46.5 % and 46.74 %, respectively. The SNP S4_30653480 (P-5.76E–09) was associated with SA (49.60 % PVE), and S1_25029792 (P-1.89E–10) and S4_3178953 (P-1.14E–09) were associated with Tips; their PVE were 13.10 % and 20.89 %, respectively (Suppl. Figs. S8a–f; S9a–f; S10a–f; S11a–d; Table 1).
Similarly, plasticity indexes were used to find significant associations for root plasticity in compacted soil. For plasticity indexes of 1.6 BD, we identified nine significant SNPs. Three SNPs were identified to be associated with volume-related traits. S1_17932204 was associated with RV_RE (P-4.06E–09) and PRVol_RE (P-1.36E–08), showing 8.55 % and 7.86 % PVE. In addition, S4_25935749 was associated with RV_RE (P-5.68E–11) and PRVol_RE (P-5.59E–12), showing 49 % and 54.4 % PVE. For LRVol_RE, S1_46735079 (P- 1.26E-07) showed a significant association with PVE 35.58 %. For area-related traits, S3_27560701 showed the highest significance with PRPA_RE (P-1.50E–12) and PRSA_RE (P-9.70E–13), explaining 19.7 % and 20.2 % PVE. Followed by S7_5017663, which showed significant associations with PRPA_RE (P-1.31E–10) and PRSA_RE (P-1.06E–10), explaining 24.1 % and 24.3 % PVE. For PRL_RE, S3_27560701 (P-1.84E–15) showed the most significant association with PVE, 37.8 %. Further, S3_27904102 explained 34.86 % PVE for PRL_RE (P-1.38E–08) (Suppl. Table S5 and Suppl. Fig. S12a–j).
We identified four significant SNPs for plasticity indexes of 1.8BD, three at chromosome 4 and one at chromosome 6, which were linked with PRL_RE, PRPA_RE, PRSA_RE, PRVol_RE, and Tips_RE, having their PVE varying between 58 % and 11 %. The most significant SNP, S4_25390989, was associated with PRL_RE (P-1.71E–07), showing 31.82 % PVE and followed by S4_30228728, associated with primary root related traits, PRL_RE (P-2.54E–14), PRPA_RE (P-2.95E–18), PRSA_RE (P-3.11E–18) and PRVol_RE (P-1.03E–17), having their PVE 49.39 %, 58.51 %, 58.33 % and 57.39 %, respectively. Further, SNP S4_33340818 was identified, linked with PRPA_RE (P-2.34E–09), PRSA_RE (P-2.17E–09) and PRVol_RE (P-9.207E–10) having PVE of 11.83 %, 12.09 %, and 13.86 % respectively. Finally, the Tips_RE (P-2.06E–10) associated SNP, S6_34682862, was also identified, which had its PVE of 40.91 % (Suppl. Table S5 and Suppl. Fig. S13a, b).
Identification of potential candidate genes
Only those SNPs were retained from the GWAS result for further analyses, which showed a significant allelic effect. In 1.1 BD soil, SNPs associated with AD and TL showed significant allelic effects. For AD-associated SNP, S4_38707673 (G/A), the A allele showed broader AD than the G allele. For S4_38707673, we identified six genes in LD and lie in the same LD block. The significant SNP is localized within a bidirectional sugar transporter SWEET1 (LOC101498274) gene (Suppl. Fig. S14 a). For TL-associated SNP, S4_665844 (G/C), the C allele showed longer roots than the G allele. The significant SNP is localized in the promoter of the low psii accumulation 1 (LOC101508210) gene, and apart from LPA1, no other gene was found to be in LD with the SNP within the 300 kb window (Table 2; Suppl. Fig. S14b).
In moderate soil compaction of 1.6 BD soil, two SNPs, S7_29511774 and S7_32834852, showed significant allelic effect. S7_29511774 (T/C) is associated with TL, and the T allele showed longer roots compared to the C allele. A total of 11 genes were found to be in LD and in the same LD block with significant SNP (Suppl. Fig. S15 a-e). Among the 11 genes, cytokinin response factor-1-like (CRF1-like) (LOC101501547) is an ethylene-responsive transcription factor family protein and could play an important role in regulating root length in compacted soil. SNP S7_32834852 (C/T) was significantly associated with TL, LRSA, LRL, and LRPA. In this, the C allele showed longer roots and a larger surface area compared to the T allele. Within the 300 kb window, 15 genes were found to be in LD and the same LD block with the significant SNP. In upstream of the significant SNP, the nearest gene was pollen-specific leucine-rich repeat extensin-like protein 3 (LRX) (LOC101506805), and in downstream the closest gene was AP2-like ethylene-responsive transcription factor AIL1 (LOC101506248). Both the SNPs were in significant LD with ethylene-responsive genes, suggesting a vital role of ethylene during soil compaction in chickpea (Table 2; Suppl. Fig. S15a–e).
Under heavy soil, compaction of 1.8 BD soil, three SNPs, S1_25029792, S6_6702892, and S6_34682862, displayed a significant allelic effect. S1_25029792 (T/A) was associated with the number of tips, and the A allele resulted in more tips than the T allele. Two genes were found in LD with the significant SNP (LOC101495273, LOC101514396). Both of these genes were pectinesterase/pectinesterase inhibitor PPE8B-like. The second SNP, S6_6702892 (G/A), was associated with LRPA and LRSA, and the A allele resulted in a larger surface area than the G allele. The significant SNP lies within the diacylglycerol kinase 2 (DGK2) (LOC101500556) gene, and none of the neighboring genes within the 300 kb window was found to be in LD with the significant SNP (Table 2; Suppl. Fig. S16a–f). The third SNP, S6_34682862 (T/A), was associated with AD, PRVol, and total root volume RV, and the A allele showed more volume and broader diameter compared to the T allele. The SNP lies in an intergenic region, and none of the genes are in LD with the significant SNP. This could be due to the low density of markers in this region. The candidate gene can be identified by increasing marker density in the region; however, the nearest gene from the significant SNP was peroxisomal membrane protein 11B (LOC101501092), around 6.6 kb from the SNP.
Plasticity indexes for moderately compacted soil have four SNPs showing significant allelic effects: S1_17932204, S1_46735079, S3_24492256, and S3_27904102. S1_17932204 (C/A) was associated with PRVol_RE and RV_RE, and the C allele showed a lower volume reduction than the T allele. Four genes were in the same LD block and LD with the significant SNP (Suppl. Table S6 and Suppl. Fig. S17). For LRVol_RE, we found S1_46735079 (C/A) significantly associated and showed a significant allelic effect. Three genes are in the same LD block and LD with the significant SNP. Among these genes, two were uncharacterized, and one was helicase SEN1 (LOC101504865). For PRPA_RE and PRSA_RE, SNP S3_24492256 (G/C) showed a significant association and allelic effect (Suppl. Table S6 and Suppl. Fig. S17). Here, the C allele showed a lower reduction in PRPA_RE and PRSA_RE than the G allele. The SNP lies within the 2 kb of cyclin U4-1-like (LOC101500825), and another gene putative glucan endo-1,3-beta-glucosidase (LOC101493810) was in the same LD block and LD with the significant SNP (Suppl. Tables S6 and S17). Only one SNP (S6_34682862 (T/A)) associated with Tips_RE showed a significant allelic effect for highly compacted soil. Here, the A allele showed a lower reduction in the number of Tips than the T allele (Suppl. Tables S6 and S17).
Differential expression analysis of putative candidate genes for RSA traits in compacted soil
We studied the expression behavior of the 21 putative candidate genes for three different SNPs. In 1.6 BD soil, we studied one SNP, S7_32834852, associated lateral root traits and showed a significant allelic effect. S7_32834852 had 15 genes in the same LD block with it, and among those 15 genes, germin-like protein 3 (GLP3) (LOC101507652) showed the highest fold change in response to compaction of 1.6 BD. In addition, another gene associated with this SNP i.e., LRX (LOC101506805), also showed higher fold change upon soil compaction of 1.6 BD (Fig. 5a). For SNP, S7_29511774, we checked 12 genes, and among them, CRF1-like (LOC101501547) was highly upregulated in 1.6 BD soil (Fig. 5b).
Validation of candidate genes from associated loci with significant allelic effect using real-time qPCR a Expression of 15 putative candidate genes in moderate (1.6 BD) and heavy (1.8 BD) soil compaction for SNP, S7_32834852 associated with lateral root traits in moderate compaction. Out of these 15 genes, GLP3 (LOC101507652) and LRX (LOC101506805) showed significantly elevated expression. b Expression of 12 candidate genes in 1.6 BD and 1.8 BD soil for SNP, S7_29511774. Out of these 12 genes, we found highly elevated expression of CRF-1-like (LOC10501547) in moderately compacted soil. c Expression pattern of candidate genes for two SNPs, S1_25029792 and S6_6702892, associated with root tips and lateral root traits for heavy soil compaction. Of two pectin esterase genes in LD with the first SNP, pectinesterase/pectinesterase inhibitor PPE8B-like (LOC101495273) showed induced expression under both compaction treatments. d For the second SNP, the expression pattern of DGK2 (LOC101500556) shows specifically elevated expression under high compaction, BD1.8. Relative expression plotted to show log twofold change using ΔΔCt method. Comparisons between genes within a treatment were analyzed using ANOVA followed by a Tukey’s Honest Significant Difference (HSD) post-hoc test (P < 0.05)
In heavy compaction of soil 1.8 BD, two SNPs, S1_25029792 and S6_6702892, associated with lateral root traits had significant allelic effects. Among the two pectin esterase genes found in LD with significant SNP (S1_25029792), one of them (LOC101495273) is highly induced (Fig. 5c). It showed remarkably high fold change in both 1.6 BD and 1.8 BD soils in comparison to 1.1 BD soil. The second SNP, S6_6702892, linked to lateral root characteristics, is located in the intronic region of DGK2 (LOC101500556). Interestingly, this gene had high expression in roots grown in 1.8 BD soil, not in 1.6 BD soil (Fig. 5d).
Discussion
Soil compaction has become a significant challenge for agriculture productivity due to the use of heavy machinery in modern farming practices. Addressing this issue is crucial to ensure food security. Roots growth gets hindered in compacted soil as they struggle to penetrate hard soil. This decrease in root growth reduces soil exploration for water and minerals, ultimately causing overall growth retardation and making plants more sensitive to soil compaction (Tracy et al. 2012). However, genetic variations are present in plants, which resuls in varying sensitivity to soil compaction-induced root shortening (Colombi and Walter 2017). Therefore, we evaluated 210 chickpea accessions to understand the important root traits under different degrees of soil compaction. We aimed to identify accessions not deterred by compacted soil and the associated genetic variations.
High-strength soil leads to changes in root system architecture (RSA), including reduced length, surface area, and volume of both primary and lateral roots; contrastingly, during soil compaction, the average diameter increased with an increase in soil compaction. We observed a positive correlation between average root diameter (AD) and primary root length (PRL) in uncompacted soil, which vanishes in compacted soil in chickpea. This suggests neither positive nor negative correlation of AD with penetration ability in compacted soil. However, a broader diameter in maize is positively correlated with higher penetration (Chimungu et al. 2015). Contrastingly, in recent reports, wider diameter is shown to be negatively associated with penetration in rice (Huang et al. 2022). These contrasting results suggest interspecies variation in the ability of thicker roots to penetrate hard soil. In our case, no correlation is observed between AD and PRL, possibly due to the engagement of additional traits influencing root penetration in hard soils.
Further, AD and lateral root traits negatively correlated in uncompacted and compacted soils, suggesting a trade-off between both traits. With increasing root diameter and decreasing lateral length in compacted soil, this trade-off between AD and lateral root traits could have more aggravated consequences for already reduced root exploration in compacted soil. Therefore, the increase in root diameter could reduce root-soil exploration in compacted soil, leading to reduced penetration, as observed in rice (Huang et al. 2022).
Apart from AD, all other root traits showed strikingly reduced standard deviation and variance with an increase in the level of soil compaction. Similar results were observed in tomato, where roots showed higher variation in root traits in uncompacted compared to compacted soil (Tracy et al. 2012). This could be due to restricted growth in compacted soil, and maybe the variants responsible for variation in root growth in uncompacted soil are downstream of the pathway, resulting in reduced growth during soil compaction.
In the present GWA study, we have identified several significant SNPs and candidate genes linked with root penetration and RSA-related traits under various levels of soil compaction. Here, we discussed candidate genes for only those SNPs with significant allelic effects.
Two SNPs displayed significant allelic effects in moderate soil compaction of 1.6 BD. Among the two SNPs, S7_32834852 (C/T) is significantly associated with lateral root traits such as LRPA, LRSA, and LRVol. Fifteen genes are in LD and lie in the same LD block with significant SNP; among those 15 genes, GLP3 and LRX showed the highest induction in response to soil compaction of 1.6 BD. GLPs are highly expressed and conserved across different plant species, and several have root-specific expression under various stresses (Zaynab et al. 2022). In Arabidopsis, overexpression of PD-GLP1 and PD-GLP2 leads to a decrease in primary root length and an increase in lateral root length (Ham et al. 2012). Similarly, AtLRX2 and AtLRX6 are involved in lateral root formation, which suggests that LRX could also be involved in lateral root development (Baumberger et al. 2003; Lewis et al. 2013). At this point, we could delimit two genes among the 17, and further characterization of both genes is required to confirm their role in soil compaction.
The other SNP, S7_29511774 (T/C), is associated with TL, and 11 genes are in LD and lie in the same LD block with significant SNP. Among these 11 genes, CRF1-like (LOC101501547) is a transcription factor belonging to the ERF family containing the AP2 domain. Interestingly, CRF1-like also showed increased expression in soil compaction of 1.6 BD and 1.8 BD relative to 1.1 BD. Thus, suggesting a probable role of CRF1-like during soil compaction. AtCRFs are involved in controlling root growth as their quadruple mutant (atcrf1/3/5/6) displays reduced primary root length and lateral number (Raines et al. 2016). In Arabidopsis, CRFs are cytokinin response factors containing the CRF and AP2 domains (Rashotte and Goertzen 2010). A similar protein was reported in Spirogyra, annotated as CRF1, but lacked the CRF domain; however, it displayed steady upregulation upon ethylene treatment (Van de Poel et al. 2016). Ethylene is also reported to induce the expression of SlCRF4 (Shi et al. 2012). Interestingly, changes in ethylene levels within the plant and in the immediate vicinity affect root growth in compacted soil. During soil compaction, ethylene accumulates near the root surface due to less diffusion to the surrounding compacted soil (Pandey et al. 2021). Ethylene employs other downstream signals like ABA and auxin to mediate the soil compaction response, such as cortical radial cell expansion, and reduce primary root length, respectively (Huang et al. 2022). As AtCRFs are known cytokinin response factors, it would be interesting to look into the involvement of cytokinin in regulating root growth along with auxin and ABA in compacted soil. Until now, no study has been related to cytokinin involvement in compacted soil; however, cross-talk of ethylene and cytokinin is well known for regulating primary root growth (Artner and Benkova 2019). Cytokinin induces ethylene biosynthesis, and ethylene is involved in activating cytokinin signal transduction through multi-step phosphorelay resulting in root growth inhibition (Hansen et al. 2009; Zdarska et al. 2019; Yamoune et al. 2023). Thus, it could be possible that CRFs play an essential role in this cross-regulation to cause root growth inhibition during soil compaction.
In heavy soil compaction of 1.8 BD, three SNPs showed significant allelic effects for lateral root traits. S1_25029792 (T/A) is significantly associated with the total number of tips and represents the lateral root number. Two pectin esterase genes were found to be in LD with significant SNP, and among the two, LOC101495273 is highly induced upon heavy soil compaction. It suggests the probable role of pectin esterase in lateral root formation during soil compaction. Pectin esterase acts on cell wall pectin and de-methyl-esterify galacturonic acid residues of homogalacturonan pectin. Desertification by pectin esterase can either lead to cell wall stiffening or loosening, depending on the esterification pattern (Micheli 2001; Willats et al. 2001). Cell wall remodeling by pectin esterases is required for lateral root formation as the ratio between pectin esterification and deesterification is vital for the processes before lateral root primordia formation (Wachsman et al. 2020). In Arabidopsis, single and double mutants of pme2 and pme3 display reduced lateral root numbers, and overexpression of PME5 leads to a reduction in pre-branching site numbers (Wachsman et al. 2020). As discussed, soil compaction delays seedling root architecture establishment (Bassett et al. 2005), resulting in smaller laterals. Thus, the induction of pectin esterase during soil compaction could be an important adaptive strategy by plants to regulate the process required before lateral root primordia formation. Further characterization of the gene and allelic variants will lead to a better understanding of the involvement of pectin esterase in lateral root formation during soil compaction.
The second SNP, S6_6702892 (G/A), is significantly associated with LRPA, LRSA, and LRVol and lies within the intron of DGK2 (LOC101500556). Interestingly, CaDGK2 is induced only during heavy soil compaction and not in moderate soil compaction, suggesting that this gene specifically works in heavy soil compaction. DGKs phosphorylate diacylglycerol to form phosphatidic acid (PA). Diacylglycerol is known to promote lateral root formation, and PA reduces lateral root development. Osdgk1 mutants have more diacylglycerol and lead to higher lateral root density. Upon PA treatment, lateral root density was restored to WT (Yuan et al. 2019). This suggests the critical role of DGK2 in regulating lateral root development in compacted soils.
In terms of plasticity, the contribution of the genotype x environment (G x E) component to the total variation is more for root volume (RV), primary root volume (PRVol), and AD, followed by surface area and least in length traits. Interestingly, we found a significant association (S1_17932204) with plasticity indexes of RV, PRVol, and RV in 1.6 BD soil, showing a significant allelic effect with all three traits. On comparing primary and lateral root traits, primary root traits displayed more G x E contribution than lateral root traits. However, the total number of tips shows comparable G x E contribution as primary root traits. Here, we found a significantly associated SNP with the plasticity index of Tips (S6_34682862) in 1.8 BD soil. Interestingly, the same SNP is associated with AD, RV, and PRVol in 1.8 BD soil; however, it has opposite effects on traits. The T allele leads to a larger diameter and volume in 1.8 BD soil but results in a higher reduction of the number of tips, leading to genotypes with the A allele thinner and less reduction in the number of root tips. Hence, it could be concluded that a larger diameter/volume is traded off with a decrease in the number of laterals and length of laterals during soil compaction, leading to reduced soil exploration. Thus, genotypes with thinner roots and profound laterals could be more beneficial during soil compaction due to higher soil exploration.
Conclusion
Soil compaction in chickpea leads to severe reduction in root system architecture (RSA) traits, except average diameter (AD), which increased with increasing soil compaction. We did not find any direct negative correlation of AD with primary root length (PRL) in compacted soil; however, we identified that the aggravated trade-off between AD and laterals in compacted soil could make increased AD responsible for deteriorated soil exploration in terms of reduced lateral roots. Interestingly, we identified significant associations for various RSA traits and their plasticity indexes for moderate (1.6 BD) and high (1.8 BD) soil compaction, leading to the identification of important probable candidate genes regulating root architecture in compacted soil. These findings will be crucial in developing chickpea varieties resistant to soil compaction.
Data Availability
Data sharing not applicable to this article as no datasets were generated or analyzed during the current study.
Abbreviations
- AD:
-
Average diameter (mm)
- BD:
-
Bulk density
- GWAS:
-
Genome-Wide Association Study
- LR:
-
Lateral root
- PA:
-
Projected area
- PR:
-
Primary root
- PVE:
-
Phenotypic variation explained
- RSA:
-
Root system architecture
- SA:
-
Surface area (cm2)
- Tips:
-
Root tips
- Tips_RE:
-
Relative change in total number of tips
- TL:
-
Total length (cm)
- RE:
-
Relative change in
- RV:
-
Total volume (cm3)
References
Alexander DH, Lange K (2011) Enhancements to the ADMIXTURE algorithm for individual ancestry estimation. BMC Bioinform 12:246. https://doi.org/10.1186/1471-2105-12-246
Artner C, Benkova E (2019) Ethylene and cytokinin: partners in root growth regulation. Mol Plant 12:1312–1314. https://doi.org/10.1016/j.molp.2019.09.003
Bassett IE, Simcock RC, Mitchell ND (2005) Consequences of soil compaction for seedling establishment: implications for natural regeneration and restoration. Austral Ecol 30:827–833. https://doi.org/10.1111/j.1442-9993.2005.01525.x
Baumberger N, Doesseger B, Guyot R et al (2003) Whole-genome comparison of leucine-rich repeat extensins in Arabidopsis and rice. A conserved family of cell wall proteins form a vegetative and a reproductive clade. Plant Physiol 131:1313–1326. https://doi.org/10.1104/pp.102.014928
Bradbury PJ, Zhang Z, Kroon DE et al (2007) TASSEL: software for association mapping of complex traits in diverse samples. Bioinformatics 23:2633–2635. https://doi.org/10.1093/bioinformatics/btm308
Brown J, Pirrung M, McCue LA (2017) FQC Dashboard: integrates FastQC results into a web-based, interactive, and extensible FASTQ quality control tool. Bioinformatics 33:3137–3139. https://doi.org/10.1093/bioinformatics/btx373
Cambi M, Mariotti B, Fabiano F et al (2018) Early response of Quercus robur seedlings to soil compaction following germination. Land Degrad Dev 29:916–925
Chen S, Huang T, Zhou Y et al (2017) AfterQC: automatic filtering, trimming, error removing and quality control for fastq data. BMC Bioinform 18:80. https://doi.org/10.1186/s12859-017-1469-3
Chimungu JG, Loades KW, Lynch JP (2015) Root anatomical phenes predict root penetration ability and biomechanical properties in maize (Zea mays). J Exp Bot 66:3151–3162. https://doi.org/10.1093/jxb/erv121
Colombi T, Walter A (2017) Genetic diversity under soil compaction in wheat: root number as a promising trait for early plant vigor. Front Plant Sci 8:420. https://doi.org/10.3389/fpls.2017.00420
Correa J, Postma JA, Watt M, Wojciechowski T (2019) Soil compaction and the architectural plasticity of root systems. J Exp Bot 70:6019–6034
Danecek P, Auton A, Abecasis G et al (2011) The variant call format and VCFtools. Bioinformatics 27:2156–2158
Danecek P, Bonfield JK, Liddle J et al (2021) Twelve years of SAMtools and BCFtools. Gigascience 10:giab008
Dong S-S, He W-M, Ji J-J et al (2021) LDBlockShow: a fast and convenient tool for visualizing linkage disequilibrium and haplotype blocks based on variant call format files. Briefi Bioinformatics 22:bbaa227. https://doi.org/10.1093/bib/bbaa227
Hansen M, Chae HS, Kieber JJ (2009) Regulation of ACS protein stability by cytokinin and brassinosteroid. Plant J 57:606–614. https://doi.org/10.1111/j.1365-313X.2008.03711.x
Huang G, Kilic A, Karady M et al (2022) Ethylene inhibits rice root elongation in compacted soil via ABA- and auxin-mediated mechanisms. Proc Natl Acad Sci USA 119:e2201072119. https://doi.org/10.1073/pnas.2201072119
Ishaq M, Ibrahim M, Hassan A et al (2001) Subsoil compaction effects on crops in Punjab, Pakistan. Soil Tillage Res 60:153–161. https://doi.org/10.1016/S0167-1987(01)00177-5
Jin K, Shen J, Ashton RW et al (2013) How do roots elongate in a structured soil? J Exp Bot 64:4761–4777. https://doi.org/10.1093/jxb/ert286
Le Marié C, Kirchgessner N, Marschall D et al (2014) Rhizoslides: paper-based growth system for non-destructive, high throughput phenotyping of root development by means of image analysis. Plant Methods 10:13. https://doi.org/10.1186/1746-4811-10-13
Leinonen R, Sugawara H, Shumway M, Collaboration INSD (2010) The sequence read archive. Nucl Acids Res 39:D19–D21
Lewis DR, Olex AL, Lundy SR et al (2013) A kinetic analysis of the auxin transcriptome reveals cell wall remodeling proteins that modulate lateral root development in Arabidopsis. Plant Cell 25:3329–3346. https://doi.org/10.1105/tpc.113.114868
Li H (2011) A statistical framework for SNP calling, mutation discovery, association mapping and population genetical parameter estimation from sequencing data. Bioinformatics 27:2987–2993. https://doi.org/10.1093/bioinformatics/btr509
Li H (2013) Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. arXiv preprint arXiv:13033997
Lipiec J, Horn R, Pietrusiewicz J, Siczek A (2012) Effects of soil compaction on root elongation and anatomy of different cereal plant species. Soil Tillage Res 121:74–81. https://doi.org/10.1016/j.still.2012.01.013
Lipka AE, Tian F, Wang Q et al (2012) GAPIT: genome association and prediction integrated tool. Bioinformatics 28:2397–2399
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2- ΔΔCT method. Methods 25:402–408
Lynch JP (2022) Harnessing root architecture to address global challenges. Plant J 109:415–431. https://doi.org/10.1111/tpj.15560
Lynch JP, Chimungu JG, Brown KM (2014) Root anatomical phenes associated with water acquisition from drying soil: targets for crop improvement. J Exp Bot 65:6155–6166. https://doi.org/10.1093/jxb/eru162
Madurapperumage A, Tang L, Thavarajah P et al (2021) Chickpea (Cicer arietinum L.) as a source of essential fatty acids—a biofortification approach. Front Plant Sci 12:734980. https://doi.org/10.3389/fpls.2021.831140
McKenna A, Hanna M, Banks E et al (2010) The genome analysis toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res 20:1297–1303
Micheli F (2001) Pectin methylesterases: cell wall enzymes with important roles in plant physiology. Trends Plant Sci 6:414–419. https://doi.org/10.1016/S1360-1385(01)02045-3
Mondal S, Christopher S, Chakraborty D, Mandal PK (2022) Soil compaction affects root growth and gene expression of major N-assimilating enzymes in wheat. J Soil Sci Plant Nutr 22:3958–3967. https://doi.org/10.1007/s42729-022-00945-2
Pandey BK, Huang G, Bhosale R et al (2021) Plant roots sense soil compaction through restricted ethylene diffusion. Science 371:276–280. https://doi.org/10.1126/science.abf3013
Passioura JB (2002) Soil conditions and plant growth. Plant Cell Environ 25:311–318. https://doi.org/10.1046/j.0016-8025.2001.00802.x
Popova L, van Dusschoten D, Nagel KA et al (2016) Plant root tortuosity: an indicator of root path formation in soil with different composition and density. Ann Bot 118:685–698
Raines T, Shanks C, Cheng C-Y et al (2016) The cytokinin response factors modulate root and shoot growth and promote leaf senescence in Arabidopsis. Plant J 85:134–147
Rashotte AM, Goertzen LR (2010) The CRF domain defines cytokinin response factor proteins in plants. BMC Plant Biol 10:74. https://doi.org/10.1186/1471-2229-10-74
Rehman T, Khan MU (2019) Early trends, current status and future prospects of farm mechanization in Asia. Agric Eng Int CIGR J 21:76–87
Ruzin SE (1999) Plant microtechnique and microscopy. Oxford University Press, New York
Satterthwaite D, McGranahan G, Tacoli C (2010) Urbanization and its implications for food and farming. Philos Trans R Soc Lond B Biol Sci 365:2809–2820. https://doi.org/10.1098/rstb.2010.0136
Schneider HM, Strock CF, Hanlon MT et al (2021) Multiseriate cortical sclerenchyma enhance root penetration in compacted soils. Proc Natl Acad Sci USA 118:e2012087118. https://doi.org/10.1073/pnas.2012087118
Shi X, Gupta S, Rashotte AM (2012) Solanum lycopersicum cytokinin response factor (SlCRF) genes: characterization of CRF domain-containing ERF genes in tomato. J Exp Bot 63:973–982. https://doi.org/10.1093/jxb/err325
Thudi M, Upadhyaya HD, Rathore A et al (2014) Understanding the genetic architecture of drought and heat tolerance in chickpea through genome-wide and candidate gene-based association mapping. PLoS One 9:e96758
Tracy SR, Black CR, Roberts JA et al (2012) Quantifying the impact of soil compaction on root system architecture in tomato (Solanum lycopersicum) by X-ray micro-computed tomography. Ann Bot 110:511–519. https://doi.org/10.1093/aob/mcs031
Upadhyaya HD, Thudi M, Dronavalli N et al (2011) Genomic tools and germplasm diversity for chickpea improvement. Plant Genetic Resour 9:45–58
Upadhyaya HD, Bajaj D, Narnoliya L et al (2016) Genome-wide scans for delineation of candidate genes regulating seed-protein content in chickpea. Front Plant Sci 7:302. https://doi.org/10.3389/fpls.2016.00302
Van de Poel B, Cooper ED, Van Der Straeten D et al (2016) Transcriptome profiling of the green alga Spirogyra pratensis (Charophyta) suggests an ancestral role for ethylene in cell wall metabolism, photosynthesis, and abiotic stress responses. Plant Physiol 172:533–545
van Rossum B-J, van Eeuwijk F, Boer M (2017) Package ‘statgenSTA.’ Statistician 56:24
Vanhees DJ, Loades KW, Bengough AG et al (2020) Root anatomical traits contribute to deeper rooting of maize under compacted field conditions. J Exp Bot 71:4243–4257. https://doi.org/10.1093/jxb/eraa165
Vanhees DJ, Schneider HM, Sidhu JS et al (2022) Soil penetration by maize roots is negatively related to ethylene-induced thickening. Plant Cell Environ 45:789–804. https://doi.org/10.1111/pce.14175
Varshney RK, Song C, Saxena RK et al (2013) Draft genome sequence of chickpea (Cicer arietinum) provides a resource for trait improvement. Nat Biotechnol 31:240–246. https://doi.org/10.1038/nbt.2491
Varshney RK, Thudi M, Roorkiwal M et al (2019) Resequencing of 429 chickpea accessions from 45 countries provides insights into genome diversity, domestication and agronomic traits. Nat Genet 51:857–864. https://doi.org/10.1038/s41588-019-0401-3
Varshney RK, Roorkiwal M, Sun S et al (2021) A chickpea genetic variation map based on the sequencing of 3366 genomes. Nature 599:622–627. https://doi.org/10.1038/s41586-021-04066-1
Wachsman G, Zhang J, Moreno-Risueno MA et al (2020) Cell wall remodeling and vesicle trafficking mediate the root clock in Arabidopsis. Science 370:819–823. https://doi.org/10.1126/science.abb7250
Wang J, Zhang Z (2021) GAPIT version 3: boosting power and accuracy for genomic association and prediction. Genom Proteom Bioinform 19:629–640. https://doi.org/10.1016/j.gpb.2021.08.005
Wei T, Simko V, Levy M et al (2017) Package ‘corrplot.’ Statistician 56:e24
Whiteley GM, Hewitt JS, Dexter AR (1982) The buckling of plant roots. Physiol Plant 54:333–342. https://doi.org/10.1111/j.1399-3054.1982.tb00268.x
Wickham H (2011) ggplot2. Wiley Interdisciplinary Reviews. Comput Stat 3:180–185
Willats WG, Orfila C, Limberg G et al (2001) Modulation of the degree and pattern of methyl-esterification of pectic homogalacturonan in plant cell walls. Implications for pectin methyl esterase action, matrix properties, and cell adhesion. J Biol Chem 276:19404–19413. https://doi.org/10.1074/jbc.M011242200
Yamoune A, Zdarska M, Depaepe T, et al (2023) Cytokinins regulate spatially-specific ethylene production to control root growth in Arabidopsis. http://biorxiv.org/lookup/doi/10.1101/2023.01.07.522790
Ye J, Coulouris G, Zaretskaya I et al (2012) Primer-BLAST: a tool to design target-specific primers for polymerase chain reaction. BMC Bioinformatics 13:1–11
Yuan S, Kim S, Deng X et al (2019) Diacylglycerol kinase and associated lipid mediators modulate rice root architecture. New Phytol 223:261–276. https://doi.org/10.1111/nph.15801
Zaynab M, Peng J, Sharif Y et al (2022) Genome-wide identification and expression profiling of germin-like proteins reveal their role in regulating abiotic stress response in potato. Front Plant Sci 12:831140. https://doi.org/10.3389/fpls.2021.831140
Zdarska M, Cuyacot AR, Tarr PT et al (2019) ETR1 Integrates response to ethylene and cytokinins into a single multistep phosphorelay pathway to control root growth. Mol Plant 12:1338–1352. https://doi.org/10.1016/j.molp.2019.05.012
Acknowledgements
RD and US acknowledge the research fellowship from DBT funded project (BT/PR31665/AGIII/103/1172/2019). PSK and MP acknowledge fellowship from DBT, India. BS acknowledge a fellowship from Indo-Swiss joint research project (BT/IN/Swiss/46/JG/2018 by DBT, India. JG was supported by NIPGR core grant. We thank Swarup K Parida for sharing chickpea seeds.
Author information
Authors and Affiliations
Contributions
RD, PSK, MP conducted experiments, analyzed data and wrote the manuscript. US and BS helped with the experiments. JG designed the project, supervised experiments, analyzed data and wrote the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Additional information
Communicated by Dorothea Bartels.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Donde, R., Kohli, P.S., Pandey, M. et al. Dissecting chickpea genomic loci associated with the root penetration responsive traits in compacted soil. Planta 259, 17 (2024). https://doi.org/10.1007/s00425-023-04294-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00425-023-04294-x