Abstract
Main conclusion
The 4-coumarate:coenzyme A ligase 4CL4 is involved in enhancing rice P acquisition and use in acid soil by enlarging root growth and boosting functional rhizosphere microbe recruitment.
Abstract
Rice (Oryza sativa L.) cannot easily acquire phosphorus (P) from acid soil, where root growth is inhibited and soil P is fixed. The combination of roots and rhizosphere microbiota is critical for plant P acquisition and soil P mobilization, but the associated molecular mechanism in rice is unclear. 4CL4/RAL1 encodes a 4-coumarate:coenzyme A ligase related to lignin biosynthesis in rice, and its dysfunction results in a small rice root system. In this study, soil culture and hydroponic experiments were conducted to examine the role of RAL1 in regulating rice P acquisition, fertilizer P use, and rhizosphere microbes in acid soil. Disruption of RAL1 markedly decreased root growth. Mutant rice plants exhibited decreased shoot growth, shoot P accumulation, and fertilizer P use efficiency when grown in soil—but not under hydroponic conditions, where all P is soluble and available for plants. Mutant ral1 and wild-type rice rhizospheres had distinct bacterial and fungal community structures, and wild-type rice recruited some genotype-specific microbial taxa associated with P solubilization. Our results highlight the function of 4CL4/RAL1 in enhancing rice P acquisition and use in acid soil, namely by enlarging root growth and boosting functional rhizosphere microbe recruitment. These findings can inform breeding strategies to improve P use efficiency through host genetic manipulation of root growth and rhizosphere microbiota.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Phosphorus (P) is an essential macroelement that limits crop growth in agricultural ecosystems. Although the application of P fertilizer is a routine agronomic practice to meet crop P demand (Withers et al. 2014), applied P is easily bound to soil particle surfaces, resulting in low soil P bioavailability and poor crop P use efficiency (Johnston et al. 2014; Cong et al. 2020). Sustaining high crop yields without large amounts of P fertilization is thus difficult. According to various estimates, rock P reserves will be exhausted in the next 50–100 years (Fixen and Johnston 2012; Johnston et al. 2014), threatening the sustainability of agricultural production. In addition, excess soil P unused by crops poses a huge threat to water environments (Daniel et al. 1998; MacDonald et al. 2011). The improvement of crop P use efficiency is a particularly promising approach for increasing agricultural productivity, while simultaneously decreasing the depletion of P rock resources and the negative environmental effects of P fertilizers.
Given the poor movability of P in soil, plant roots can acquire available P only near root surfaces—a situation that leads to P depletion zones around roots (Hinsinger et al. 2005) and subsequently limits P uptake by roots. To acquire more P for growth, plants must, therefore, explore larger soil volumes and exploit less-soluble P pools (Vance et al. 2003; Lynch and Brown 2008; Strock et al. 2018). Plants can accordingly modify root physiological features and cooperate with rhizosphere microbiota to adapt to P-deficient environments (Richardson and Simpson 2011; Cong et al. 2020). Adaptive modification of plant root system traits is one of the most effective strategies to increase P acquisition efficiency (Shen et al. 2011; Niu et al. 2013; Vejchasarn et al. 2016; Cong et al. 2020). Increases in the root/shoot ratio, root branching, root elongation, root topsoil foraging, and the number of root hairs enable plant roots to explore a given volume of soil more effectively and acquire more P (Lynch and Brown 2008; Shen et al. 2011; Cong et al. 2020). Since root system traits largely determine P acquisition efficiency from soil, the identification of molecular mechanisms behind root system traits for efficient P use by crops is of strong interest.
In addition to the involvement of root systems, microorganisms play a vital role in the regulation of soil P bioavailability. Soil bacteria (e.g., Alcaligenes, Aerobactor, Bacillus, Pseudomonas, and Nguyenibacter) and fungi (e.g., Aspergillus, Chaetomium, Cephalosporium, Fusarium, and Penicillium) can convert inorganic and/or organic P into more plant-bioavailable forms by releasing protons, organic anions, and various phosphatases (Chen et al. 2006; Sharma et al. 2013; Alori et al. 2017; Mohanram and Kumar 2019; Li et al. 2021). Carbon–nutrient exchanges are an important strategy used by plants foraging for resources such as P (Rodriguez et al. 2019; Raymond et al. 2021; Han et al. 2022). Plants deliver the complex carbons produced by photosynthesis into the rhizosphere to nourish microbiota, with microbes in turn facilitating the solubilization and uptake of essential nutrients (e.g., P, N, and Fe) by plants (Rashid et al. 2016; Jacoby et al. 2017). This beneficial effect from rhizosphere microbiota is plant genotype-specific (Rodriguez et al. 2019; Trivedi et al. 2020). Zhang et al. (2019) have reported an association between a nitrate transporter, NRT1.1B, and the recruitment of root microbiota and N use in rice. Although this finding suggests possible breeding strategies to improve crop N use efficiency by modulating rhizosphere microbiota, the molecular mechanisms underlying host genetic regulation of rhizosphere microbiota associated with P utilization are largely unknown.
Rice (Oryza sativa L.), a globally important, staple cereal crop, consumes huge amounts of P fertilizer. Approximately 13% of rice plants worldwide are cultivated in acid soils (pH < 5.5) (von Uexküll and Mutert 1995). In such soils, Al toxicity inhibits root growth (Ma et al. 2001), and the low pH weakens the P biogeochemical cycle driven by soil microbiota (Dai et al. 2020), resulting in decreased P uptake and use by rice plants. Since P deficiency is the main limiting factor for crop production in acid soils (Zhao et al. 2014; Wang et al. 2021), the improvement of rice P use in such areas is vital. The protein kinase gene Pstol1 has been reported to enhance rice root growth and thereby confer tolerance to P deficiency in a nearly neutral (pH 6.1) soil (Gamuyao et al. 2012), but no genes regulating root system traits associated with rice P use efficiency in acid soil have been uncovered thus far. The 4CL4/ral1 gene encoding 4-coumarate:coenzyme A ligase (4CL4), an enzyme putatively involved in lignin biosynthesis, regulates the response of rice root phenotypes to Al toxicity (Liu et al. 2016, 2020). Lignin plays a critical role in cell wall extensibility and root elongation under stressed conditions (Yan et al. 2022). Mutation of the ral1 gene reduces lignin production and results in defective cell elongation of the root mature zone, thereby showing a smaller root system relative to wild-type rice (Liu et al. 2016, 2020). Given the vital role of roots in P acquisition from soil, we used the ral1 mutant and its corresponding wild type, Kasalath, to examine the impact of the ral1 gene on rice P acquisition efficiency and rhizosphere microbiota in acid soil in this study. We hypothesized that (1) the wild-type rice Kasalath, with its larger root system, has a higher P acquisition efficiency than the mutant ral1 in acid soil; (2) mutation of the ral1 gene leads to a rhizosphere microbial pattern that is distinct from that of Kasalath because of the altered root genetic traits; and (3) a link exists between rice P acquisition efficiency and rhizosphere microbiota that is dependent on rice genotype.
Materials and methods
Plant materials and seedling preparation
The rice mutant ral1 and its corresponding wild type, Kasalath, were used in this study. The ral1 mutant, with a short root phenotype, was previously screened from an ethylmethylsulfone-mutagenized population (Liu et al. 2016). Subsequent research revealed that the ral1 gene encodes the 4-coumarate:coenzyme A ligase 4CL4 and impacts Al tolerance and lignin accumulation in rice (Liu et al. 2020). The pathway of 4CL4 involved in lignin synthesis is shown in Fig. S1. Rice seeds were surface-sterilized in 10% H2O2 solution for 30 min to avoid seed-borne diseases, washed with deionized water, soaked in water at 37 °C in the dark for 2 days, and then transferred to a net floating on a 500 μM CaCl2 solution (pH 4.6) for 4 days. After germination, seedlings were cultured in a 3.5 L plastic pot containing half-strength Kimura B solution (Ma et al. 2002) until the three-leaf stage for subsequent soil culture and hydroponic experiments.
Soil culture experiment
The soil used in this study was collected from the surface soil layer (0–20 cm depth) at the Yingtan Red Soil Ecological Experiment Station, Jiangxi Province, China (28° 14′ N, 117° 03′ E). The collected soil was air-dried and then passed through a 2-mm sieve after removal of roots and stones. Soil chemical properties were as follows: total nitrogen (TN), 2.33 g kg−1; pH, 4.4; organic carbon (SOC), 14.42 g kg−1; available P (AP), 2.50 mg kg−1. Since P deficiency and Al toxicity coexist in acidic soils (Chen et al. 2012), lime was used to increase soil pH and decrease soil toxic Al concentrations in this study. Rice seedlings were subjected to three P treatments with or without liming (2 g CaCO3 kg−1). Three P levels were established by applying P at 0, 10, or 50 mg P kg−1 in the form of KH2PO4. All pots were fertilized at the same rate with N (urea, 100 mg N kg−1) and K (100 mg K kg−1), the latter balanced with KCl at the three P levels. Four replicates were carried out per treatment. To ensure the homogeneous distribution of fertilizers in each pot, fertilizers and soil for each pot were weighed accurately and then mixed thoroughly. Each pot (15 cm height, 15 cm top diameter, and 12 cm bottom diameter) contained 2.5 kg of air-dried soil. Three 10-day-old rice plants were planted in each pot and then harvested after 40 days, at the tillering stage. Rhizosphere soil samples were collected immediately after rice plants were harvested. Soil adhering to roots after shaking for 30 s was considered to be rhizosphere soil (Xiao et al. 2022). Apparent P use efficiency was calculated as (total shoot P with P fertilization − total shoot P without P fertilization)/P fertilizer application amount × 100%.
Hydroponic experiment
For the hydroponic experiment, 10-day-old rice seedlings were grown in half-strength Kimura B solution (pH 4.6) containing 5, 20, 90, or 180 μM P. Four rice plants were grown per pot in black plastic pots containing 1.2 L of nutrient solution. P was supplied as NaH2PO4. Three replicates were carried out per P treatment. All pots were maintained in a growth chamber [12-h day (30 ± 1 °C):12-h night (23 ± 1 °C); 65 ± 5% humidity; 900 μmol m−2 s−1 illumination intensity]. The nutrient solution was renewed every 2 days. After 20 days, rice roots were separated from shoots for measurement of dry weight and shoot P concentration.
Short-term P uptake experiment
Roots of 16-day-old seedlings were placed in black bottles containing 100 mL of 10 or 180 μM P in half-strength Kimura B nutrient solution (pH 4.6). After 4 h, the P concentration of the solution in each bottle was determined. We also weighed each bottle before and after the uptake experiment to calculate the amount of solution lost due to transpiration by plants. At the end of the experiment, the roots were harvested, oven-dried, and weighed. The P uptake rate was calculated according to the amount of P depleted from the uptake solution during the experiment.
Plant and soil chemical analyses
For nutrient determinations, harvested plants were dried at 75 °C to a constant weight and then smashed with a grinder. The smashed shoots were digested with 5 mL HNO3 at 140 °C. P concentrations in the digested solution of the soil culture experiment and the uptake solution of the short-term P uptake experiment were determined by the colorimetric molybdenum blue method. Soil total N was quantified on a VarioMAX CNS elemental analyzer (Elementar, Hanau, Germany). Soil pH was measured in a soil–water suspension (1:2.5) using a pH meter (PB-21, Sartorius, Göttingen, Germany). Soil available P was extracted with HCl–NH4F and measured by the molybdenum blue method. Soil exchangeable Al (EAl) was extracted with 1 M KCl (soil:water, 1:25) and determined by inductively coupled plasma–atomic emission spectrophotometry (Optima 8000, PerkinElmer, Waltham, MA, USA). Soil organic carbon was analyzed by the dichromate oxidation method.
Analyses of root system parameters
Harvested roots from the soil culture and hydroponic experiments were scanned on a flatbed scanner (Epson Expression 10000XL, Epson America, San Jose, CA, USA) at a resolution of 400 dpi, and total root lengths and surface areas were measured using WinRHIZO (Regent Instruments, Quebec, QC, Canada).
Expression of Pi starvation response genes
To determine the expression of Pi starvation response genes under various P levels, 10 day-old rice plants were exposed to half-strength Kimura B solution (pH 4.6) containing 5, 20, 90, or 180 μM P for 10 days and then roots (0–3 cm) were excised for RNA isolation. Total RNA was extracted from the excised samples using RNAiso Plus (Takara, Kusatsu, Shiga, Japan) and first-strand cDNA synthesis was prepared from 1 μg of RNA using the HiScript II Q Select RT SuperMix (Vazyme Biotech Co., Ltd., Nanjing, China). Subsequently, cDNA products were diluted tenfold in nuclease-free water and then used for quantitative real-time PCR analysis (qRT-PCR). The qRT-PCR was performed in a 10 μL reaction mixture consisting of 5 μL of SYBR Premix Ex Taq (Vazyme Biotech), 0.5 μL of each of the forward and reverse primers (10 μM), and 4 μL of diluted cDNA. The qRT-PCR analysis was performed on CFX96 Touch real-time PCR detection system (BioRad, Hercules, CA, USA). Histone H3 was used as an internal control. We measured the expression levels of four P starvation response genes including OsPht1;2, OsPht1;3, OsPht1;6, and OsARF12 in roots, and the primers and related function of those genes were shown in Table S1. Relative expression levels were calculated by the comparative Ct method. Three independent biological replicates were made for each treatment.
DNA extraction, PCR amplification and sequencing, and bioinformatics analysis
DNA was extracted using a Fast DNA Spin Kit for Soil (MP Biomedicals, Santa Ana, CA, USA). The quality and concentration of extracted DNA were checked using a NanoDrop ND-1000 spectrophotometer (NanoDrop Technologies, Wilmington, DE, USA).
Bacterial 16S rRNA and fungal ITS gene profiling of rice rhizosphere soil samples were carried out by Illumina sequencing. Thermal cycling protocols and primers for amplification of microbiota are described in Table S2. PCR amplification for sequencing was carried out as described by Xiao et al. (2022). After purification and quantification, amplicons were pooled in equal amounts and sequenced on the Illumina MiSeq PE300 platform (Illumina, San Diego, CA, USA) at Shanghai Personal Biotechnology Co. (Shanghai, China).
Raw 16S rRNA and ITS gene sequences were processed as described by Zhang et al. (2019) and Liu et al. (2021). Briefly, the sequences were quality checked and subjected to barcode and primer trimming and filtering of low-quality sequences. Next, an OTU table was generated by USEARCH, and representative sequences were taxonomically classified using the RDP classifier (Wang et al. 2007). We obtained 1,126,909 and 1,088,148 high-quality sequences, which were, respectively, clustered into 6405 bacterial and 1320 fungal OTUs.
Data analysis
Significant differences at the P < 0.05 level were assessed by Duncan’s test and Student’s t test in SPSS 22.0. To identify treatment differences in the beta diversity of bacterial and fungal community compositions, non-metric multi-dimensional scaling (NMDS) based on Bray–Curtis distances and analysis of similarities (ANOSIM) testing were conducted with the ‘vegan’ package in R v.4.0.5 (Oksanen et al. 2013). And a Mantel test was performed using the same package to examine relationships among edaphic soil properties, root system traits, total shoot P, and rhizosphere microbiota. To explore genotype-specific OTUs, the R package ‘edger’ was used to identify differentially abundant OTUs (relative abundance > 0.01%) between genotypes (Robinson et al. 2010). To construct microbial-OTU co-occurrence networks, we filtered out the OTUs occurring in fewer than four samples and trimmed links according to two criteria: Pearson’s correlation coefficient |r| < 0.7 and P > 0.05. The networks were then visualized using Gephi v.0.9.2, and their topological properties were calculated with the ‘igraph’ package in R.
Results
Rice growth response to different P levels
Application of lime and elevation of P levels increased the shoot and root dry weights of Kasalath and ral1 in the soil culture experiment (Fig. 1a, b, Fig. S2). In the hydroponic experiment, Kasalath and ral1 shoot and root dry weights were markedly increased by boosting P levels from 5 to 20 μM, with these values then remaining constant from 20 to 180 μM P (Fig. 1d, e). The dry weight ratio of roots to shoots decreased with increasing P under both soil culture and hydroponic conditions (Fig. 1c, f). Shoot and root dry weights of Kasalath were both significantly higher than those of ral1 in the soil culture experiment, whereas only the root dry weight of Kasalath was significantly higher than that of ral1 in the hydroponic experiment. In both soil culture and hydroponic experiments, applying lime and boosting P levels increased total root length and surface area—both of which were higher in Kasalath compared with ral1 (Fig. 2).
Dry weights of wild-type Kasalath rice and its mutant ral1 at different phosphorus (P) levels in soil culture (a, b, c) and hydroponic (d, e, f) experiments. a, d Shoot dry weight. b, e Root dry weight. c, f Dry weight ratio of roots to shoots. In the soil culture experiment, 10-day-old seedlings were grown in soil supplemented with 0, 10, or 50 mg kg−1 P under liming (+ Ca) or non-liming (− Ca) conditions for 40 days. In the hydroponic experiment, 10-day-old seedlings were grown in nutrient solution (pH 4.6) supplemented with 5, 20, 90, or 180 μM P for 20 days. Different uppercase and lowercase letters above bars indicate significant differences among different P levels for Kasalath and ral1, respectively (P < 0.05, Duncan’s multiple range test). Asterisks indicate significant differences between Kasalath and ral1 under the same treatment conditions (P < 0.05, independent-sample t test). Data are means ± standard deviation (n = 4 for the soil culture experiment; n = 3 for the hydroponic experiment)
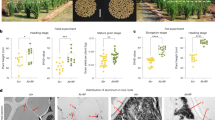
Root parameters of wild-type Kasalath rice and its mutant ral1 at different phosphorus (P) levels in the soil culture (a, b, c) and hydroponic (d, e, f) experiments described in Fig. 1. a, d Root total length. b, e Root surface area. c, f Scanned root images. After plant harvest, rice roots were scanned for measurements of root total length and surface area. Different uppercase and lowercase letters above bars indicate significant differences among different P levels for Kasalath and ral1, respectively (P < 0.05, Duncan’s multiple range test). Asterisks indicate significant differences between Kasalath and ral1 under the same treatment conditions (P < 0.05, independent-sample t test). Data are means ± standard deviation (n = 4 for the soil culture experiment; n = 3 for the hydroponic experiment)
Rice P uptake and use
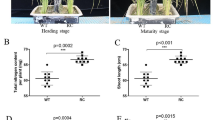
Shoot P concentrations and accumulation increased with rising P levels in both soil culture and hydroponic experiments (Fig. 3a–d). Application of lime increased shoot P accumulation, which was higher in Kasalath compared with ral1 in the soil culture experiment (Fig. 3b) but not significantly different in the hydroponic experiment (Fig. 3d).
Shoot phosphorus (P) accumulation in wild-type Kasalath rice and its mutant ral1 at different P levels in the soil culture (a, b, c) and hydroponic (d, e, f) experiments described in Fig. 1. a, c Shoot P concentration. b, d Total shoot P accumulation. Different uppercase and lowercase letters above bars indicate significant differences among different P levels for Kasalath and ral1, respectively (P < 0.05, Duncan’s multiple range test). Asterisks indicate significant differences between Kasalath and ral1 under the same treatment conditions (P < 0.05, independent-sample t test). Data are means ± standard deviation (n = 4 for the soil culture experiment; n = 3 for the hydroponic experiment)
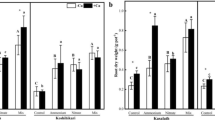
When P levels were increased, the ratio of shoot dry weights of Kasalath to ral1 remained constant in the absence of liming but decreased if lime was applied (Fig. 4a). Similarly, the ratio of shoot P accumulation of Kasalath to ral1 remained constant with increasing P under non-liming conditions and decreased under liming treatment (Fig. 4b). This result suggests that Kasalath is more tolerant to low P conditions compared with ral1 when lime is applied. Addition of lime increased apparent P use efficiency (Fig. 4c). In the soil culture experiment, this parameter was higher in Kasalath compared with ral1, thus indicating that the more highly developed root system of Kasalath under liming conditions was beneficial for root P acquisition. In the short-term P uptake experiment, ral1 had a slightly higher P uptake rate than Kasalath at low (10 μM) and high (180 μM) P levels (Fig. S3). This result indicates that the stronger P acquisition ability of Kasalath relative to ral1 was not due to a higher P uptake rate.
Ratio of shoot dry weight and total shoot phosphorus (P) of wild-type Kasalath rice relative to its mutant ral1 and fertilizer P use efficiency at different P levels in the soil culture experiment described in Fig. 1. a Ratio of shoot dry weight of Kasalath relative to ral1. b Ratio of total shoot P of Kasalath relative to ral1. c Fertilizer P use efficiency. Different lowercase letters above bars indicate significant differences among different P levels under liming (+ Ca) and non-liming (− Ca) conditions (P < 0.05, Duncan’s multiple range test). Asterisks indicate significant differences between Kasalath and ral1 under the same treatment conditions (P < 0.05, independent-sample t test). Data are means ± standard deviation (n = 4)
We further investigated the expression of P starvation response genes including OsPht1;2, OsPht1;3, OsPht1;6, and OsARF12 in the roots of both Kasalath and ral1. Compared with the condition of 90 μM P, the expression levels of OsPht1;2, OsPht1;3, and OsPht1;6 were induced by low P in both Kasalath and ral1 but those of OsARF12 were inhibited (Fig. S4). Moreover, the expression levels of OsPht1;2 and OsPht1;6 in Kasalath were higher than those in ral1 under the 5 μM P condition (Fig. S4).
Rhizosphere microbial diversity
Richness and Shannon indexes were used to estimate the alpha diversities of bacterial and fungal communities. Liming increased the alpha diversity of bacteria but reduced that of fungi (Fig. S5). When soil P levels were increased, the alpha diversity of bacteria showed an upward trend, whereas that of fungi trended downward. Nevertheless, no significant differences in bacterial and fungal alpha diversities were observed between Kasalath and ral1.
NMDS and ANOSIM analyses of Bray–Curtis dissimilarities revealed that bacterial and fungal community structures (beta diversities) differed significantly between liming and non-liming conditions (Fig. S6). To eliminate the influence of this strong liming effect on our investigation of microbial community structural differences between rice genotypes or P levels, we reanalyzed the beta diversities of bacteria and fungi under liming and non-liming conditions separately. In the new analysis, ANOSIM testing uncovered significant effects of rice genotype as well as P level on bacterial and fungal community structures (Table S3), and NMDS divided the rhizosphere microbiota into two distinct groups according to genotype, i.e., Kasalath and ral1 (Fig. 5).
We then compared the composition of bacterial and fungal communities at the phylum level and focused on the variation in community composition between the two rice genotypes (Fig. 6). The main phyla in the bacterial community were Firmicutes, Proteobacteria, and Acidobacteria, which accounted for more than 80% of total bacterial sequences (Fig. 6a). The relative abundance of Firmicutes in the rhizosphere of Kasalath was higher than that of ral1, whereas the opposite was true for Acidobacteria and Chloroflexi (Fig. S7a, b). The main phyla in the fungal community, Ascomycota, Basidiomycota, and Chytridiomycota, accounted for 41.36–82.98% of total fungal sequences (Fig. 6b). The relative abundances of Ascomycota and Basidiomycota were highest in the rhizospheres of Kasalath and ral1, respectively (Fig. S7c, d). According to these results, the mutation of the ral1 gene altered rhizosphere bacterial and fungal community compositions. We further explored relationships among rhizosphere microbiota, several soil properties, root system traits (total root length and surface area), and shoot P accumulation. The changes in bacterial and fungal community structures observed in this study were associated with soil pH and exchangeable Al, total root length and surface area, and total shoot P and soil available P (Table S4). These correlations indicate that alteration in root system size caused by the ral1 mutation affected rhizosphere microbiota by impacting soil chemical properties and rice P uptake.
Genotype-specific rhizosphere microbial taxa
To examine differences in the rhizosphere microbiota of Kasalath and ral1 in further detail, we used Manhattan plots to analyze the enrichment of OTUs according to their taxonomy (Fig. 7; Table S5). Most bacterial OTUs enriched in the rhizosphere of Kasalath belonged to Firmicutes and Proteobacteria. In the ral1 rhizosphere, enriched bacterial OTUs belonged to a wide range of phyla, including Proteobacteria, Acidobacteria, Actinobacteria, and Chloroflexi (Fig. 7a; Table S5a, b). With regard to fungi, only a few genotype-specific OTUs, mostly members of Ascomycota and Basidiomycota, were detected in the rhizospheres of Kasalath and ral1 (Fig. 7b; Table S5c, d). We detected 17 Kasalath-specific and 29 ral1-specific bacterial OTUs overlapping between non-liming and liming conditions (Fig. 7c). In contrast, most genotype-specific enriched fungal OTUs did not overlap between the two liming treatments (Fig. 7d).
Manhattan plot showing bacterial (a) and fungal (b) OTUs enriched in rhizosphere soils of wild-type Kasalath rice and its mutant ral1 under liming (+ Ca) and non-liming (− Ca) conditions. Each dot or triangle represents a single OTU. OTUs enriched in rhizosphere soils of Kasalath or ral1 are represented by filled or empty triangles, respectively. OTUs are colored according to phylum. CPM, counts per million. c, d Overlapping OTUs enriched in rhizosphere soils of Kasalath and ral1 under liming (+ Ca) and non-liming (− Ca) conditions
Rhizosphere microbial co-occurrence networks and keystone taxa
To identify hub species and compare their relative abundances between Kasalath and ral1, we constructed two microbial co-occurrence networks of rhizosphere bacteria and fungi under non-liming and liming conditions (Fig. 8). The network under non-liming conditions included 834 OTUs and 2233 associations, whereas the network under liming conditions consisted of 1085 OTUs and 6971 associations (Fig. 8a, b). As revealed by the higher number of nodes and edges and larger node average degree, the microbial network under liming conditions was more complex than that under non-liming conditions (Table S6). We identified 10 and 13 module hub species under non-liming and liming conditions, respectively (Fig. 8c, d). Most of these species belonged to Acidobacteria, and their relative abundances were higher in the rhizosphere of ral1 than in that of Kasalath (Table S7).
Co-occurrence network patterns (a and b) and hub species (c and d) under liming (+ Ca) and non-liming (− Ca) conditions. Nodes in the network are subcategorized as follows: peripherals (Zi < 2.5 and Pi < 0.62), connectors (Pi > 0.62), module hubs (Zi > 2.5), and network hubs (Zi > 2.5 and Pi > 0.62)
Discussion
Involvement of the ral1 gene in rice P acquisition via root system enlargement
Mutation of the ral1 gene, which encodes a 4-coumarate:coenzyme A ligase involved in rice lignin biosynthesis, inhibits root cell elongation of the mature zone and impacts rice Al tolerance (Liu et al. 2016, 2020). In the present study, the wild-type rice Kasalath had a larger root system (i.e., root absorbing surface area and total length), higher shoot dry weight, P accumulation, and fertilizer P use efficiency than the mutant ral1. These results demonstrate the function of ral1 in enhancing rice P acquisition by enlarging the root system in acid soil. This finding is similar to the observations of a previous study, namely, that the Pstol1 gene encoding a protein kinase conferred tolerance to P deficiency on rice grown in a nearly neutral (pH 6.1) soil by enhancing root growth (Gamuyao et al. 2012). As deduced from the shoot dry weight and P accumulation of Kasalath relative to that of ral1, the ability of the ral1 gene to improve P acquisition and shoot growth was weakened at high P levels under liming conditions. This result suggests that a large root system is much more important in an acid, P-limited soil than in a high-pH, P-rich one, as toxic Al ions in acid soil have strong inhibitory effects on plant root growth and thereby hinder the acquisition of water and nutrients by roots (Ma et al. 2001). Although wild-type Kasalath is relatively Al-sensitive compared with mutant ral1 (Liu et al. 2016, 2020), the larger root system of Kasalath was still beneficial for rice shoot growth and P uptake in our study. We also found that the dry weight ratio of roots to shoots was increased under low P conditions and that liming enhanced root growth and P acquisition. These results strongly support the importance of a developed root system in rice P acquisition in acid soil.
Owing to the low mobility and availability of P in soil, plants must develop a large root system to exploit a given volume of soil more effectively and maximize P acquisition (Vance et al. 2003; Shen et al. 2011; Gamuyao et al. 2012). In the hydroponic experiment in this study, however, no significant difference was observed in shoot growth, P concentration, or P accumulation between wild-type Kasalath and mutant ral1 even though Kasalath had a larger root system than ral1. This result indicates that a larger root system may not be essential for efficient P uptake under hydroponic conditions, as all P in the nutrient solution was soluble and available to the rice plants. As a result, plants under hydroponic conditions do not necessarily require a large root system to acquire P; this is because P in the solution can easily move to root surfaces for plant uptake. Interestingly, the P uptake rate of ral1 in the nutrient solution was actually slightly higher than that of Kasalath. Consequently, a higher P uptake rate across the root cell membrane may not necessarily result in efficient acquisition of P from soil by rice plants. In other words, the size of the root system, rather than the root P uptake rate, seems to be more important for efficient soil P acquisition by rice plants under the soil culture conditions in this study (Fig. 9). However, the expression levels of P starvation response genes OsPht1;2 and OsPht1;6 were higher in Kasalath than in ral1 under 5 μM P condition, which cannot explain the higher P uptake rate of ral1 relative to Kasalath. Since Kasalath with a larger root system under low P conditions may decrease the P concentrations of the solutions more quickly than ral1, we speculate that the expression of OsPht1;2 and OsPht1;6 was up-regulated much more by P deficiency in Kasalath than in ral1.
Schematic diagram for the role of 4CL4/RAL1 in the improvement of rice P uptake. Briefly, 4CL4/RAL1 catalyzes the 4-coumaric acid and ferulic acid to synthesize lignin and thereby boosts rice root system development. On the one hand, a larger root system (bigger root total length and surface area) enables rice to exploit larger soil volumes for P acquisition. On the other hand, a larger root system solubilizes more P from soils by altering the rhizosphere microbial community and recruiting genotype-specific microbes associated with P solubilization. Consequently, the enlarged root total length and surface area and boosted functional rhizosphere microbe recruitment improve rice P uptake
Involvement of ral1 in the recruitment of rhizosphere microbe taxa associated with P mobilization
According to numerous studies, rhizosphere microbiota differ between host genotypes (Bulgarelli et al. 2013; Turner et al. 2013; Zhang et al. 2019; Xiong et al. 2021; Xiao et al. 2022). Differences in plant nutrient acquisition, stress tolerance, and crop yield between host genotypes are closely associated with distinct rhizosphere microbiota (Rodriguez et al. 2019; Zhang et al. 2019; Trivedi et al. 2020). In the present study, we obtained two pieces of evidence implying that mutation of the ral1 gene changed rhizosphere microbiota patterns. First, rhizosphere bacteria and fungi were well separated into two groups according to rice genotype. Second, Kasalath and ral1 specifically recruited some rhizosphere microbiota. These results suggest that the ral1 gene has a considerable impact on rhizosphere microbiota establishment under fixed environmental conditions. Plants mainly influence their root and rhizosphere microbial communities through the secretion of root exudates, which serves as the mechanistic link between host genetic variation and differences in microbiota composition between different genotypes (Sasse et al. 2018). According to a previous study, the mutation of the ral1 gene reduced lignin production and increased the accumulation of its substrate 4-coumaric and ferulic acids in roots (Liu et al. 2020). Although barley (Hordeum vulgare L.) plants may adjust their lignin contents in response to soil microbial community, neither genotype nor mutations in lignin production affected soil microbial community (Bennett et al. 2015). Also, field-grown transgenic switchgrass (Panicum virgatum L.) with altered lignin did not affect soil microbiology (DeBruyn et al. 2017). Bradley et al. (2007) demonstrated that altered lignin biosynthesis in Populus tremuloides had different effects on soil microbial communities among three distinct soils. Thus, we speculate that the difference in root exudates rather than plant lignin contents due to genetic variation is responsible for the distinct rhizosphere microbial community between the mutant ral1 and its wild-type Kasalath. Our future work will focus on the examination of the difference in root exudates between Kasalath and ral1 to elucidate the mechanism underlying their distinct microbial communities.
Despite the above-mentioned issue, we were interested in exploring whether Kasalath, with its larger root system, recruits a core group of microorganisms that can improve rice growth or serve as an indicator of soil nutrient status. Compared with that of the ral1 mutant, the rhizosphere soil of Kasalath had a higher relative abundance of Firmicutes bacteria. According to a previous report, an increased relative abundance of Firmicutes has a protective effect against bacterial wilt disease (Lee et al. 2021), which would possibly benefit the growth of the wild-type rice Kasalath. By contrast, Kasalath had a lower relative abundance of Acidobacteria compared with the ral1 mutant. Rhizosphere microbiota interacts closely with one another to form a stable ecological network and promote plant growth (Durán et al. 2018). Hub species play a key role in the basic process of microbiota assembly and collaboration, which may help plants adapt to the soil environment (de Vries et al. 2018). In the present study, most module hub species, regardless of liming treatment, were assigned to Acidobacteria (Table S7). Most members of Acidobacteria are acidophilic chemoheterotrophs that exhibit an oligotrophic lifestyle (Kalam et al. 2020). The high abundance of Acidobacteria associated with the ral1 mutant in this study may reflect it is acidic, Al-toxic, infertile rhizosphere environment (Fig. S8).
To further inspect genotype-specific taxa at the OTU level, we carried out a differential OTU abundance analysis. We noticed that most bacterial OTUs enriched in the rhizosphere of Kasalath under non-liming conditions belonged to Desulfosporosinus (20.37% cumulative relative abundance; Table S5a), a genus of acidophilic, Fe (III) and sulfate-reducing bacteria (Bertel et al. 2012; Sánchez-Andrea et al. 2015). Desulfosporosinus species can convert sulfates into sulfides and induce the reduction of Fe (III) to Fe (II) (Fan et al. 2018). Since Fe2+ has a weaker PO43− binding ability relative to Fe3+, we surmise that Desulfosporosinus species release PO43− from FePO4 and improve soil P availability for plants by reducing Fe3+ to Fe2+. We also found that the rhizosphere of Kasalath was enriched in several bacterial OTUs assigned to Rhodocyclaceae, Burkholderiaceae, and Bacillales (Table S5a, b) and some fungal OTUs assigned to Penicillium, Trichoderma, Cladosporium, and Aspergillus (Table S5c, d). These bacteria and fungi are known for their ability to solubilize and mobilize P (Sharma et al. 2013; Alori et al. 2017; Grafe et al. 2018). Our results also suggest that changes in bacterial and fungal community structures are significantly associated with rice shoot P accumulation (Table S4). These results collectively support the potential function of rice genotype-specific bacterial and fungal taxa in impacting rice P nutrition. Similar to the previously reported association of the nitrate transporter NRT1.1B with root microbiota composition and nitrogen use in rice (Zhang et al. 2019), our study has demonstrated that the 4-coumarate:coenzyme A ligase 4CL4/RAL1, an enzyme putatively involved in lignin biosynthesis, is associated with rice P use and rhizosphere microbe recruitment (Fig. 9).
Conclusion
Taken together, our results reveal the new function of the 4-coumarate:coenzyme A ligase 4CL4/ RAL1 in enhancing the fertilizer P use efficiency of rice plants and regulating rhizosphere microbiota in acid soil via root growth enlargement. Our data suggest that the wild-type rice Kasalath with its larger root system can better coordinate its rhizosphere recruitment of genotype-specific microbes to efficiently acquire P from soil than the ral1 mutant. This finding can serve as the basis of an attractive breeding strategy for the improvement of fertilizer P use efficiency in acid soil based on host genetic manipulation of the root microbiome. It is worth exploring the mechanisms for the difference in rhizosphere microbial community between the ral1 mutant and its wild type in the future.
Author contribution statement
XQZ: contributed to the study conception and design; XX: performed the experiments, analyzed the data, prepared the figures and tables, and wrote the manuscript draft; and XQZ, AYH, XYD, and RFS: revised the manuscript. All the authors have read and approved the final manuscript.
Data availability
The raw data for rhizosphere microbiota were submitted to the NCBI BioProject database under accession number PRJNA821392. Further detailed data that support the findings of this study are available from the corresponding author upon reasonable request.
Abbreviations
- 4CL:
-
4-Coumarate:coenzyme A ligase
- RAL1:
-
Resistance to aluminum 1
- OTU:
-
Operational taxonomic unit
- NMDS:
-
Non-metric multi-dimensional scaling
- ANOSIM:
-
Analysis of similarities
References
Alori ET, Glick BR, Babalola OO (2017) Microbial phosphorus solubilization and its potential for use in sustainable agriculture. Front Microbiol 8:971
Bennett AE, Grussu D, Kam J, Caul S, Halpin C (2015) Plant lignin content altered by soil microbial community. New Phytol 206:166–174
Bertel D, Peck J, Quick TJ, Senko JM (2012) Iron transformations induced by an acid-tolerant Desulfosporosinus species. Appl Environ Microb 78:81–88
Bradley KL, Hancock JE, Giardina CP, Pregitzer KS (2007) Soil microbial community responses to altered lignin biosynthesis in Populus tremuloides vary among three distinct soils. Plant Soil 294:185–201
Bulgarelli D, Schlaeppi K, Spaepen S, Van Themaat EVL, Schulze-Lefert P (2013) Structure and functions of the bacterial microbiota of plants. Annu Rev Plant Biol 64:807–838
Chen YP, Rekha PD, Arun AB, Shen FT, Lai WA, Young CC (2006) Phosphate solubilizing bacteria from subtropical soil and their tricalcium phosphate solubilizing abilities. Appl Soil Ecol 34:33–41
Chen RF, Zhang FL, Zhang QM, Sun QB, Dong XY, Shen RF (2012) Aluminium–phosphorus interactions in plants growing on acid soils: does phosphorus always alleviate aluminium toxicity? J Sci Food Agr 92:995–1000
Cong WF, Suriyagoda LDB, Lambers H (2020) Tightening the phosphorus cycle through phosphorus-efficient crop genotypes. Trends Plant Sci 25:967–975
Dai ZM, Liu GF, Chen HH, Chen CR, Wang JK, Ai SY, Wei D, Li DM, Ma B, Tang CX et al (2020) Long-term nutrient inputs shift soil microbial functional profiles of phosphorus cycling in diverse agroecosystems. ISME J 14:757–770
Daniel TC, Sharpley AN, Lemunyon JL (1998) Agricultural phosphorus and eutrophication: a symposium overview. J Environ Qual 27:251–257
de Vries FT, Griffiths RI, Bailey M, Craig H, Girlanda M, Gweon HS, Hallin S, Kaisermann A, Keith AM, Kretzschmar M et al (2018) Soil bacterial networks are less stable under drought than fungal networks. Nat Commun 9:1–12
DeBruyn JM, Bevard DA, Essington ME, McKnight JY, Schaeffer SM, Baxter HL, Mazarei M, Mann DGJ, Dixon RA, Chen F, Zhuo CL, Wang ZY, Stewart CN Jr (2017) Field-grown transgenic switchgrass (Panicum virgatum L.) with altered lignin does not affect soil chemistry, microbiology, and carbon storage potential. GCB Bioenergy 9:1100–1109
Durán P, Thiergart T, Garrido-Oter R, Agler M, Kemen E, Schulze-Lefert P, Hacquard S (2018) Microbial interkingdom interactions in roots promote Arabidopsis survival. Cell 175:973–983
Fan XF, Ding SM, Gong MD, Chen MS, Gao SS, Jin ZF, Tsang DC (2018) Different influences of bacterial communities on Fe (III) reduction and phosphorus availability in sediments of the cyanobacteria- and macrophyte-dominated zones. Front Microbiol 9:2636
Fixen PE, Johnston AM (2012) World fertilizer nutrient reserves: a view to the future. J Sci Food Agr 92:1001–1005
Gamuyao R, Chin JH, Pariasca-Tanaka J, Pesaresi P, Catausan S, Dalid C, Slamet-Loedin I, Tecson-Mendoza EM, Wissuwa M, Heuer S (2012) The protein kinase Pstol1 from traditional rice confers tolerance of phosphorus deficiency. Nature 488:535–539
Grafe M, Goers M, von Tucher S, Baum C, Zimmer D, Leinweber P, Vestergaard G, Kublik S, Schloter M, Schulz S (2018) Bacterial potentials for uptake, solubilization and mineralization of extracellular phosphorus in agricultural soils are highly stable under different fertilization regimes. Env Microbiol Rep 10:320–327
Han MG, Chen Y, Li R, Yu M, Fu LC, Li SF, Su JR, Zhu B (2022) Root phosphatase activity aligns with the collaboration gradient of the root economics space. New Phytol 234:837–849
Hinsinger P, Gobran GR, Gregory PJ, Wenzel WW (2005) Rhizosphere geometry and heterogeneity arising from root-mediated physical and chemical processes. New Phytol 168:293–303
Jacoby R, Peukert M, Succurro A, Koprivova A, Kopriva S (2017) The role of soil microorganisms in plant mineral nutrition—current knowledge and future directions. Front Plant Sci 8:1617
Johnston AE, Poulton PR, Fixen PE, Curtin D (2014) Phosphorus: its efficient use in agriculture. Adv Agron 123:177–228
Kalam S, Basu A, Ahmad I, Sayyed R, El-Enshasy HA, Dailin DJ, Suriani NL (2020) Recent understanding of soil acidobacteria and their ecological significance: a critical review. Front Microbiol 11:580024
Lee SM, Kong HG, Song GC, Ryu CM (2021) Disruption of Firmicutes and Actinobacteria abundance in tomato rhizosphere causes the incidence of bacterial wilt disease. ISME J 15:330–347
Li XL, Zhao XQ, Dong XY, Ma JF, Shen RF (2021) Secretion of gluconic acid from Nguyenibacter sp. L1 is responsible for solubilization of aluminum phosphate. Front Microbiol 12:784025
Liu S, Gao H, Wu X, Fang Q, Chen L, Zhao FJ, Huang CF (2016) Isolation and characterization of an aluminum-resistant mutant in rice. Rice 9:1–13
Liu S, Zhao L, Liao YH, Luo ZL, Wang H, Wang P, Zhao H, Xia JX, Huang CF (2020) Dysfunction of the 4-coumarate: coenzyme A ligase 4CL4 impacts aluminum resistance and lignin accumulation in rice. Plant J 104:1233–1250
Liu YX, Qin Y, Chen T, Lu M, Qian X, Guo X, Bai Y (2021) A practical guide to amplicon and metagenomic analysis of microbiome data. Protein Cell 12:315–330
Lynch JP, Brown KM (2008) Root strategies for phosphorus acquisition. In: White PJ, Hammond JP (eds) The ecophysiology of plant-phosphorus interactions. Springer, pp 83–116
Ma JF, Ryan PR, Delhaize E (2001) Aluminium tolerance in plants and the complexing role of organic acids. Trends Plant Sci 6:273–278
Ma JF, Shen RF, Zhao ZQ, Wissuwa M, Takeuchi Y, Ebitani T, Yano M (2002) Response of rice to Al stress and identification of quantitative trait loci for Al tolerance. Plant Cell Physiol 43:652–659
MacDonald GK, Bennett EM, Potter PA, Ramankutty N (2011) Agronomic phosphorus imbalances across the world’s croplands. Proc Natl Acad Sci USA 108:3086–3091
Mohanram S, Kumar P (2019) Rhizosphere microbiome: revisiting the synergy of plant-microbe interactions. Ann Microbiol 69:307–320
Niu YF, Chai RS, Jin GL, Wang H, Tang CX, Zhang YS (2013) Responses of root architecture development to low phosphorus availability: a review. Ann Bot 112:391–408
Oksanen J, Blanchet FG, Kindt R, Legendre P, Minchin P, O’hara R, Simpson G, Solymos P, Stevens MHH, Wagner H (2013) Community ecology package. R Pack Ver 2:321–326
Rashid MI, Mujawar LH, Shahzad T, Almeelbi T, Ismail IM, Oves M (2016) Bacteria and fungi can contribute to nutrients bioavailability and aggregate formation in degraded soils. Microbiol Res 183:26–41
Raymond NS, Gómez-Muñoz B, van der Bom FJT, Nybroe O, Jensen LS, Müller-Stöver DS, Oberson A, Richardson AE (2021) Phosphate-solubilising microorganisms for improved crop productivity: a critical assessment. New Phytol 229:1268–1277
Richardson AE, Simpson RJ (2011) Soil microorganisms mediating phosphorus availability update on microbial phosphorus. Plant Physiol 156:989–996
Robinson MD, McCarthy DJ, Smyth GK (2010) edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26:139–140
Rodriguez PA, Rothballer M, Chowdhury SP, Nussbaumer T, Gutjahr C, Falter-Braun P (2019) Systems biology of plant-microbiome interactions. Mol Plant 12:804–821
Sánchez-Andrea I, Stams AJ, Hedrich S, Ňancucheo I, Johnson DB (2015) Desulfosporosinus acididurans sp. nov.: an acidophilic sulfate-reducing bacterium isolated from acidic sediments. Extremophiles 19:39–47
Sasse J, Martinoia E, Northen T (2018) Feed your friends: do plant exudates shape the root microbiome? Trends in Plant Sci 23:25–41
Sharma SB, Sayyed RZ, Trivedi MH, Gobi TA (2013) Phosphate solubilizing microbes: sustainable approach for managing phosphorus deficiency in agricultural soils. Springerplus 2:587
Shen JB, Yuan LX, Zhang JL, Li HG, Bai ZH, Chen XP, Zhang WF, Zhang FS (2011) Phosphorus dynamics: from soil to plant. Plant Physiol 156:997–1005
Strock CF, Morrow de la Riva L, Lynch JP (2018) Reduction in root secondary growth as a strategy for phosphorus acquisition. Plant Physiol 176:691–703
Trivedi P, Leach JE, Tringe SG, Sa T, Singh BK (2020) Plant–microbiome interactions: from community assembly to plant health. Nat Rev Microbiol 18:607–621
Turner TR, James EK, Poole PS (2013) The plant microbiome. Genome Biol 14:1–10
Vance CP, Uhde-Stone C, Allan DL (2003) Phosphorus acquisition and use: critical adaptations by plants for securing a nonrenewable resource. New Phytol 157:423–447
Vejchasarn P, Lynch JP, Brown KM (2016) Genetic variability in phosphorus responses of rice root phenotypes. Rice 9:1–16
von Uexküll HR, Mutert E (1995) Global extent, development and economic impact of acid soils. Plant Soil 171:1–15
Wang Q, Garrity GM, Tiedje JM, Cole JR (2007) Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microb 73:5261–5267
Wang JL, Liu KL, Zhao XQ, Zhang HQ, Li D, Li JJ, Shen RF (2021) Balanced fertilization over four decades has sustained soil microbial communities and improved soil fertility and rice productivity in red paddy soil. Sci Total Environ 793:148664
Withers PJA, Sylvester-Bradley R, Jones DL, Healey JR, Talboys PJ (2014) Feed the crop not the soil: rethinking phosphorus management in the food chain. Environ Sci Technol 48:6523–6530
Xiao X, Wang JL, Li JJ, Li XL, Dai XJ, Shen RF, Zhao XQ (2022) Distinct patterns of rhizosphere microbiota associated with rice genotypes differing in aluminum tolerance in an acid sulfate soil. Front Microbiol 13:933722
Xiong JB, Lu J, Li X, Qiu Q, Chen J, Yan C (2021) Effect of rice (Oryza sativa L.) genotype on yield: Evidence from recruiting spatially consistent rhizosphere microbiome. Soil Biol Biochem 161:108395
Yan L, Li S, Cheng J, Zhang YR, Jiang CC (2022) Boron-mediated lignin metabolism in response to aluminum toxicity in citrus (Poncirus trifoliata (L.) Raf.) root. Plant Physiol Biochem 185:1–12
Zhang JY, Liu YX, Zhang N, Hu B, Jin T, Xu HR, Qin Y, Yan PX, Zhang XN, Guo XX et al (2019) NRT1.1B is associated with root microbiota composition and nitrogen use in field-grown rice. Nat Biotechnol 37:676–684
Zhao XQ, Chen RF, Shen RF (2014) Coadaptation of plants to multiple stresses in acidic soils. Soil Sci 179:503–513
Acknowledgements
We are grateful to C-F Huang for providing rice seed materials and nice comments on this study. We thank HZ, JL, XL, and YW: for their assistance with rice cultivation and soil sampling. We thank LB (Edanz) (www.liwenbianji.cn/ac) for editing the English text of a draft of this manuscript.
Funding
This work was supported by the Strategic Priority Research Program of the Chinese Academy of Sciences (nos XDA24020104 and XDA24040203) and the National Natural Science Foundation of China (nos. 42077101 and 31672229).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
425_2023_4158_MOESM1_ESM.docx
Supplementary file 1: Table S1 Primers used for detection of Pi starvation response gene expression. Table S2 Primers and thermal cycling protocols used for high-throughput sequencing. Table S3 Analysis of similarities (ANOSIM) test of the effects of P level and genotype on the beta diversity of bacterial and fungal community structures based on Bray–Curtis distance matrixes. Table S4 Mantel test of the correlation between selected soil and rice properties based on Euclidean distances and rhizosphere microbiological structure based on Bray–Curtis distances. Table S5 Bacterial and fungal OTUs enriched in the rhizosphere of Kasalath and ral1 under non-liming (-Ca) and liming (+ Ca) conditions. Table S6 Key topological features of co-occurrence network patterns under non-liming (-Ca) and liming (+ Ca) conditions. Table S7 Relative abundance and taxonomy of hub species in co-occurrence networks. Fig. S1 The general biosynthesis pathway of lignin in higher plants modified from Liu et al. (2018). Green words indicate the specific step of 4CL4 involved in lignin biosynthesis. PAL, phenylalanine ammonia-lyase; TAL, tyrosine ammonia-lyase; C4H, cinnamate 4-hydroxylase; 4CL, 4-coumarate: CoA ligase; CCR, cinnamoyl-CoA reductase; HCT, hydroxycinnamoyl-CoA shikimate/Quinatehydroxycinnamoyltransferase; C3H, p-coumarate 3-hydroxylase; CCoAOMT, caffeoyl-CoA O-methyltransferase; F5H, ferulate 5-hydroxylase; CSE, caffeoyl shikimate esterase; COMT, caffeic acid O-methyltransferase; CAD, cinnamyl alcohol dehydrogenase; LAC, laccase; POD, peroxidase. Fig. S2 Shoot growth picture of Kasalath and ral1 under the soil culture experiment. 10-day-old seedlings were grown in soil supplemented with 0, 10, or 50 mg kg−1 P under liming (+ Ca) or non-liming (-Ca) conditions for 40 days. Fig. S3 Short-term P uptake rate of wild-type Kasalath rice and its mutant ral1. Sixteen-day-old seedlings were exposed to 10 or 180 μM P for 4 h. Asterisks indicate significant differences between Kasalath and ral1 at the same P level (P < 0.05, independent-sample t-test). Data are means ± standard deviation (n = 4). Fig. S4 Gene expression related to rice P starvation responses. 10-day-old rice plants were exposed to half-strength Kimura B solution (pH 4.6) containing 5, 20, 90, or 180 μM P for 10 days, and then roots were sampled for the expression analysis. Expression level relative to Kasalath with 90 μM P is shown. Different uppercase and lowercase letters above bars indicate significant differences among different P levels for Kasalath and ral1, respectively (P < 0.05, Duncan’s multiple range test). Asterisks indicate significant differences between Kasalath and ral1 under the same treatment conditions (P < 0.05, independent-sample t-test). Data are means ± standard deviation (n = 3). Fig. S5 Richness and Shannon diversity indexes of bacteria (a, c) and fungi (b, d) in rhizosphere soils of wild-type Kasalath rice and its mutant ral1 at different P levels under liming (+ Ca) and non-liming (-Ca) conditions. Different uppercase and lowercase letters above bars indicate significant differences among different P levels for Kasalath and ral1, respectively (P < 0.05, Duncan’s multiple range test). Data are means (n = 4). Fig. S6 Non-metric multi-dimensional scaling (NMDS) analysis of bacterial (a) and fungal (b) community structure based on a Bray–Curtis dissimilarity matrix. Fig. S7 Stamp analysis of the relative abundance of bacterial (a, b) and fungal (c, d) phyla at the 95% confidence interval level between Kasalath and ral1 under liming (+ Ca) and non-liming (-Ca) conditions. Fig. S8 Soil chemical properties of rhizospheres of Kasalath and ral1 at different P levels under liming (+ Ca) and non-liming (-Ca) conditions. a Soil pH. b Bray P. c Exchangeable Al. Different uppercase and lowercase letters above bars indicate significant differences among different P levels for Kasalath and ral1, respectively (P < 0.05, Duncan’s multiple range test). Asterisks indicate significant differences between Kasalath and ral1 under the same treatment conditions (P < 0.05, independent-sample t-test). Data are means ± standard deviation (n = 4) (DOCX 8032 KB)
425_2023_4158_MOESM2_ESM.xlsx
Supplementary file 2: Table S5 Bacterial and fungal OTUs enriched in the rhizosphere of Kasalath and ral1 under non-liming (-Ca) and liming (+ Ca) conditions (XLSX 55 KB)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Xiao, X., Hu, A.Y., Dong, X.Y. et al. Involvement of the 4-coumarate:coenzyme A ligase 4CL4 in rice phosphorus acquisition and rhizosphere microbe recruitment via root growth enlargement. Planta 258, 7 (2023). https://doi.org/10.1007/s00425-023-04158-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00425-023-04158-4