Abstract
Main conclusion
P. polyphylla selectively enriches beneficial microorganisms to help their growth.
Abstract
Paris polyphylla (P. polyphylla) is an important perennial plant for Chinese traditional medicine. Uncovering the interaction between P. polyphylla and the related microorganisms would help to utilize and cultivate P. polyphylla. However, studies focusing on P. polyphylla and related microbes are scarce, especially on the assembly mechanisms and dynamics of the P. polyphylla microbiome. High-throughput sequencing of the 16S rRNA genes was implemented to investigate the diversity, community assembly process and molecular ecological network of the bacterial communities in three root compartments (bulk soil, rhizosphere, and root endosphere) across three years. Our results demonstrated that the composition and assembly process of the microbial community in different compartments varied greatly and were strongly affected by planting years. Bacterial diversity was reduced from bulk soils to rhizosphere soils to root endosphere and varied over time. Microorganisms benefit to plants was selectively enriched in P. polyphylla roots as was its core microbiome, including Pseudomonas, Rhizobium, Steroidobacter, Sphingobium and Agrobacterium. The network’s complexity and the proportion of stochasticity in the community assembly process increased. Besides, nitrogen metabolism, carbon metabolism, phosphonate and phosphinate metabolism genes in bulk soils increased over time. These findings suggest that P. polyphylla exerts a selective effect to enrich the beneficial microorganisms and proves the sequential increasing selection pressure with P. polyphylla growth. Our work adds to the understanding of the dynamic processes of plant-associated microbial community assembly, guides the selection and application timing of P. polyphylla-associated microbial inoculants and is vital for sustainable agriculture.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Plants live with numerous microorganisms, which play essential roles in their hosts’ health and productivity and coevolve with hosts (Ling et al. 2022; Martin et al. 2017; Trivedi et al. 2020). Generally, endophytes are considered to be a complement to the host plant’s gene library, helping the hosts adapt to the environment (Vandenkoornhuyse et al. 2015), such as enhancing stress tolerance (e.g., drought and salinity), improving disease resistance (Clarke et al. 2006), aiding mineral uptake (Malinowski et al. 2000) and promoting growth (Schardl et al. 2004). The plant–microbe interaction is diverse. Plant-related microorganisms affect the host's growth and developmental processes but are also regulated by the host metabolite, immune system, and responses to stress (Bai et al. 2022). Studies reported that sesquiterpenes had induced hyphal branching in arbuscular mycorrhizal fungi (Akiyama et al. 2005). Flavonoids are also essential secondary metabolites in improving plant–microbe interactions (Deng et al. 2021; Fu et al. 2022). Dynamic interactions among the environment, microorganisms, and hosts shaped plant-related microbiome assembly and host health, but understanding ecological processes was still shallow (Sessitsch et al. 2019; Fitzpatrick et al. 2020). In addition, researchers discovered that Ginkgo biloba and endophytes were likely to share and compensate for some metabolic processes (Zou et al. 2021). Uncovering the mechanisms of plant microbiome assembly, functions, and networks is vital for applying microbial inoculants in agriculture (Singh et al. 2020; Haskett et al. 2021).
Roots are key organs for plants to obtain nutrients and microorganisms (Chapman et al. 2012; Hirsch and Mauchline 2012). Diverse microbes surround plant roots in the rhizosphere (Edwards et al. 2015; O’Brien and Harrison 2021). However, only some microorganisms are specifically acquired by plants in the root endosphere (Berg and Smalla 2009). The root-associated microbiome, including microorganisms in bulk soil, rhizosphere soil and root endosphere, exerts essential biological and ecological functions in plant health (Mendes et al. 2013; Qu et al. 2020). The rhizosphere is the dominant compartment for plant–microbe interactions that plant root exudates can significantly influence (Bakker et al. 2013; Sun et al. 2021). Low-molecular-weight organic compounds in root exudates, such as organic acids and sugars, shape the structure and function of the root-associated microbial communities (Shi et al. 2011). Benzoxazinoids, the defensive root exudates, were shown to alter root-associated microorganism communities (Saunders and Kohn 2009; Hu et al. 2018). Moreover, root exudates may help the plant enrich specific growth-promoting rhizobacteria (Vacheron et al. 2013; Vives-Peris et al. 2020). Root exudates, including phenolic acids, influence the colonization of plant growth-promoting rhizobacteria in maize and groundnut roots (Ankati and Podile 2019). Furthermore, researchers found that root-associated microbiome assemblage was affected by plant development and planting years (Chaparro et al. 2014). Microbiomes with plants, including bean, maize, rice, cowpea, cabbage, grape, cotton, arabidopsis and tobacco, exhibit age-related variation (Develey-Rivière and Galiana 2007). However, the assembly mechanism of root-associated communities was still unclear. In particular, studies on the dynamic variation in microbial assembly processes across planting years are still lacking.
As an important medicinal plant, P. polyphylla is widely used in the pharmaceutical industry and receiving increasing attention. Researchers isolated endophytes producing multiple antibacterial metabolites from P. polyphylla (Zhao et al. 2010). Some plant growth-promoting bacteria, such as Bacillus megaterium, were also isolated from P. polyphylla (Tao et al. 2021). Previous studies showed that the community composition of both endophytic bacteria and fungi was affected by planting years and related to saponins (Yang et al. 2015). Researchers claimed that there were significant differences in the bacterial community among P. polyphylla organs (Liu et al. 2020), and the bacterial community could be affected by the altitude (Wang et al. 2020). However, the process of P. polyphylla microbial community assembly and the dynamic variation of the community have not been well studied.
In Midu, P. polyphylla was planted in a greenhouse for four years. The bacterial community was examined across 54 samples from soils (rhizosphere and bulk soil) and P. polyphylla root endosphere. This study aimed to clarify how planting years shape microbiome assemblies among compartments and determine the potential sources and keystone taxa of P. polyphylla to guide the sustainable development of P. polyphylla. We hypothesized that the planting years would affect the bacterial community assembly of P. polyphylla from the soils to the root endosphere, and plant growth-promoting bacteria are enriched during planting.
Materials and methods
Sample collection and property
Our study was performed at a greenhouse at Midu (100°29′ E, 25°20‘N), Dali, Yunnan Province in China. Since 2016, the greenhouse has grown P. polyphylla in this area commercially. The weather at Midu was wet and warm, with an annual average temperature of 17.3 °C, annual average precipitation of 824 mm, and annual sunshine hours of 2339.5 h. P. polyphylla was planted in the greenhouse from 2016 to 2020. We maintained the same water and fertilizer management yearly during planting (regular and quantitative watering and no top dressing). Six 3 m × 3 m quadrats and ten P. polyphylla of similar size in each quadrat were randomly selected by five-point sampling. The bulk soils (BS), rhizosphere soils (RS), and root endosphere (RE) of P. polyphylla were collected from quadrats planted for four years, on September 2018, September 2019, and September 2020, respectively. Samples on September 2018, September 2019, and September 2020 were numbered 2, 3 and 4, such as BS2, BS3 and BS4. Bulk soil (BS) samples were collected 5 cm far from the P. polyphylla's root at a depth of 5 ~ 15 cm. The plants were removed from the soil and mildly shooked to remove soil loosely adhering to the roots. After shaking off the loose soil, the roots with still aggregated soil were shaken in 25 mL 0.1 M sterile phosphate buffer (7.1 g Na2HPO4, 4.4 g NaH2PO4·H2O added to 820 mL deionized water, pH 7.0) for 30 min; the roots were then taken out and the suspension centrifuged at 9000 g for 10 min to collect the precipitation, as rhizosphere soil (RS) (Saunders and Kohn 2009). Clean roots were surface disinfected promptly for microbiological analysis. Roots were sterilized in 4% NaClO for 5 min and washed with sterile water (Sorty et al. 2016). After surface sterilizing, the roots were dried in sterile air on a sterile workbench. All soil samples were transported on dry ice, and roots were triturated in liquid nitrogen. All samples were stored at − 80 °C until DNA and polyphyllin extraction.
Measurement of soil physical and chemical properties and polyphyllin
We dried bulk soils to constant weight. Because of the lack of rhizosphere soil, only the physicochemical properties of bulk soil were determined. The pH meter measured soil pH using a mixture of air-dried bulk soil and water (1:2.5, w/v). The total organic carbon concentrations (TOC) were measured by dichromate oxidation. The concentrations of total nitrogen (TN) were analyzed by Kjeldahl determination. The total phosphorus (TP) and available phosphorus (AP) were released by NaOH fusion and 0.5 mol/L NaHCO3, respectively, followed by colorimetric analysis. The total potassium (TK) and available potassium (AK) were measured by flame photometry following NaOH fusion extraction and 1 M ammonium acetate. For water-soluble nitrogen (WSN), 3 g soil was mixed with water (30 mL), shaken for 30 min at 100 g, and centrifuged at 6000 g for 20 min. The collected supernatant was filtered with 0.45 µm pore-size syringe filters, and the fractions of organic N are referred to as water-soluble nitrogen. The soil was baked in an oven at 70 °C to a constant weight, and the moisture content (MC) was measured.
Frozen roots were dried in an oven at 60 ℃ until the weight was constant. Afterward, these roots were ground into powder smaller than 40 mesh and extracted in a Soxhlet extractor based on Chinese pharmacopeia (2015). An aliquot of 0.500 g dried powder was homogenized in 25 mL ethanol, followed by 30 min of reflux at 80 ℃. Then, the homogenate was filtrated to remove residue and ethanol was added to 25 mL for HPLC.
HPLC was implemented on a Shimadzu LC-20AD Series HPLC system with an SPD-20A UV–Vis detector, and a connected ACQUITY UPLCTM BEH C18 column (217 mm × 2.1 mm, 1.7 μm) was used for detection at 25 ℃. The mobile phase consisted of acetonitrile (a) and H2O (b). A flow rate of 1.0 mL/min was used. Standard curves were established by a series concentration of the corresponding standard at 203 nm.
DNA extraction, sequencing and sequence analysis
The root fragments were homogenized in a sterilized mortar and pestle with liquid nitrogen. The total genomic DNA of root samples was extracted from the homogenized root material using the OMEGA Plant DNA Kit. The OMEGA Soil DNA Kits were used to extract the total genomic DNA of soil samples. To minimize the interference of hosts' DNA, primer pair fM1 (5′-CCGCGTGNRBGAHGAAGGYYYT-3′) and rC5 (5′-TAATCCTGTTTGCTCCCCAC-3′) were used to amplify the genome of microorganisms (Yu et al. 2013; Sabu et al. 2018). DNA Clean-Up Kit purified the amplicons. High-throughput sequencing of the PCR products was conducted on the Illumina Miseq platform (Miseq PE250).
The raw data were processed by QIIME 2 (version 2020.6) (Bolyen et al. 2019). These steps included: quality filtering, Ribosomal Database Project (RDP) clustering, sequence alignments and community dissimilarities analysis (Wu et al. 2018). Taxonomic assignment of 16S representative sequences was executed with the RDP classifier according to the Greengene database (DeSantis et al. 2006). Resampled 16S reads (10,000 sequences per sample) were to calculate alpha diversity and beta diversity. The raw data have been submitted to the NCBI SRA database. The accession number of all samples is PRJNA 824132.
Statistical analysis
Alpha-diversity indexes, including Shannon index, Simpson index, Simpson_evenness and Pielou_evenness, were calculated on the website (http://mem.rcees.ac.cn:8080/). Dissimilarity tests of operational taxonomic units (OTUs) and predicted function genes (by PICRUSt) had been calculated, and analysis of similarities (ANOSIM) and permutational multivariate analysis of variance (PERMANOVA) were calculated according to the website (http://mem.rcees.ac.cn:8080/). The biomarkers of samples were identified by the linear discriminant analysis (LDA) effect size (LEfSe) (P < 0.05, logarithmic LDA score > 4) (Segata et al. 2011). PICRUSt (Phylogenetic Investigation of Communities by Reconstruction of Unobserved States) was used as a bioinformatics tool to predict the abundance of functional genes in microflora (Langille et al. 2013). By comparison with the sequence data of the 16S Greengene database (Greengenes 13.5), the community functional genes were predicted with reference to the KEGG (Kyoto Encyclopedia of Genes and Genomes) database (DeSantis et al. 2006; Kanehisa and Goto 2000).
The null model test's beta Nearest Taxon Index (βNTI) was calculated to assess the determinism and stochasticity in microbiome assembly. Defined |βNTI|≥ 2 as dominant deterministic processes and |βNTI|< 2 as dominant stochastic processes. Deterministic and stochastic processes were partitioned into five ecological processes based on both βNTI and Bray–Curtis-based Raup-Crick Index (RCBray) values, including heterogeneous selection (βNTI < − 2), homogeneous selection (βNTI > + 2), dispersal limitation (|βNTI|< + 2 and RCBray > 0.95), homogenizing dispersal (|βNTI|< + 2 and RCBray < − 0.95), and undominated (|βNTI|< 2 and |RCBray|< 0.95) [66, 67] (Stegen et al. 2013; Xiong et al. 2021). OTUs with an average abundance of more than 0.03% were chosen to construct correlation networks by calculating Spearman rank correlations with Spearman correlation coefficient (r > 0.6, P < 0.01). Gephi was used to visualize the correlation networks (Team 2008).
Results
Soil property in bulk soil and polyphyllin in roots
Soil property results are listed in Table 1. During the P. polyphylla planting, the pH of the bulk soil decreased from 6.01 to 4.68, suggesting that the planting soil in this area was acidic. TK and AK in the soil decreased significantly across the three years, by almost 50% and 60%, respectively. In contrast, the TOC increased a half from 79.63 g/kg over time. The bulk soils’ AP, TP, and TN increased in 2018–2019 and stayed stable in the last two years. There were irregular changes in the water-soluble nitrogen (WSN) and moisture content (MC) of the bulk soil.
Polyphyllin in roots is presented in Table 2. Polyphyllin VII and polyphyllin VI decreased from 2018 to 2020, while polyphyllin I increased by 3 times across the three years. Diosgenin decreased by 64% from 49.17 mg/kg to 17.77 mg/kg.
Microbial composition of P. polyphylla roots and root-related soils
Samples from different compartments of P. polyphylla (including bulk soils, rhizosphere soils, and root endosphere) were collected from the cultivation greenhouse with varying years of planting. After quality filtering, 7,637,512 high-quality sequences were obtained from soil and root bacteria, which were matched to 17,314 operational taxonomic units (OTU) provided in supplementary materials (Supplementary Interactive Plot Data.xlsx) The Good's coverage for the observed_OTUs of bacterial communities was 96.92% ± 1.26%, which showed that the sequencing depth of all samples was adequate to present the microbial community diversity reliably (Fig. S1a). Veen plots showed more total OTUs and unique OTUs in bulk soil. There was less OTU richness in the root endosphere (Fig. S1b and c). All OTUs were classified into 47 phyla and 457 genera, and the relative abundance of 6 phyla and ten genera exceeded 1%.
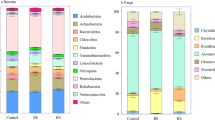
This further study the bacterial community composition in different samples was investigated by LEfSe. Results showed that there were more Proteobacteria, Acidobacteria and Actinomycetes in bulk soil, but Proteobacteria dominated the root endosphere and rhizosphere (Fig. 1a and b). Acidobacteria and Actinomycetes increased with the distance to the root, while Proteobacteria significantly declined (Figs. 1a and 2a). At the genus level, most reads were not assigned in bulk soil. Almost 30% of reads could not be assigned to the rhizosphere and root endosphere (Fig. 1a and b). Excluding these unassigned sequences, sequences were mainly assigned to 20 genera, including Rhizobium, Shingobium, Burkholderia, Pseudomonas and Novosphingobium. Compared with soils, more microorganisms were assigned to Pseudomonas and Novosphingobium in the root endosphere (Fig. 2a and Fig. S2). Besides, microbes with a higher relative abundance in the soil also increased over time, such as Actinomycetes and Acidobacteriales (Fig. 2b and Fig. S3a). Rhizobium and Shingobium gradually enriched in rhizosphere soil (Fig. 2c and Fig. S3b), and Pseudomonas and Burkholderia also increased in the P. polyphylla root endosphere (Fig. 2d and Fig. S3c). The core microbiota was defined as the genus in all samples, with a relative abundance greater than 1% in 95% of samples. It was found that the core microbiota of different compartments was significantly different (Fig. 1b). Rhodoplanes, Candidatus Solibacter and Kaistobacter were the core microbiota of bulk soil. At the same time, the P. polyphylla root endosphere had a core microbiota consisting of Pseudomonas, Rhizobium, Steroidobacter, Sphingobium and Agrobacterium. Pseudomonas in the root endosphere increased about tenfold during the three years (Fig. S4).
Microbial community structure of P. polyphylla roots and root-related soils
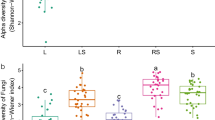
The diversities of bacteria, estimated by phylogenetic distance diversity, Chao1 index, and Shannon index, presented significant increase trends with distance to the root (Fig. 3a). The diversity of bacteria was also affected by planting years, but the effects on different compartments were different. The Shannon index indicated that the alpha diversity of bulk soil and root endosphere significantly decreased during the three years, but the opposite trend was observed in rhizosphere soil (Fig. 3b).
Diversity of bacterial communities in P. polyphylla roots and root-related soils. a Alpha diversity among compartments. Different letters above the boxes indicate a significant difference determined by the Turkey test (SD). b Alpha diversity across three years. Different letters above the boxes indicate a significant difference determined by the Turkey test
In addition, beta diversity showed significant differences among samples at different planting years and in separate compartments. NMDS results showed samples from the same compartments clustered together. The P. polyphylla microbiota shifted with compartment in the first axis and separated by planting years in the second axis, indicating that compartment and planting time were the main factors influencing the root microbiota composition (Fig. 4). ANOSIM and PERMANOVA analysis indicated that the P. polyphylla microbiome assembly was mainly explained by compartment (84.5%) followed by planting years (4.1%) (Table S1). PERMANOVA analysis and NMDS ordinations indicated that planting years explained almost all the bacterial variations (98.6%–100%) in all compartments (BS, RS and RE). Compartment differences explained all the variations in each year. Moreover, microbial community dissimilarity among all samples was much higher in the root endosphere than in soil (Fig. 4). The linear mixed model analysis suggested that planting years had a greater influence on bacterial Shannon index in bulk soil than in root endosphere and rhizosphere (Table S2). The compartment effect on the bacterial Shannon index increased over time (Fig. 4).
Network and community assembly of the root-related microbial community in P. polyphylla across the three planting years
Molecular ecology network (MEN) analyses were performed to reveal the bacterial community interactions in different compartments of P. polyphylla across the three years (Fig. 5). MEN from bulk soil were less densely connected than those from the root endosphere and rhizosphere, with fewer edges and lower density despite more nodes (Fig. 5 and Table 3). In the network of bulk soil, the top 50 core nodes with the highest degree were mainly bacteria belonging to Acidobacteria, Actinobacteria and Proteobacteria, such as Rhodoplanes and Conexibacter. While in rhizosphere soil and root endosphere, the top 50 core nodes with the highest degree were almost all Proteobacteria, such as Rhizobia, Burkholderia and Sphingomonadales (Table S3). OTUs of the five genera (Sphingobium, Pseudomonas, Steroidobacter, Agrobacterium, and Sphingomonas) occupy more network nodes in the root endosphere and rhizosphere (Table S3). In addition, with the prolongation of planting years, fewer nodes were involved in the network construction in the bulk soil, and the network had fewer edges and became more sparse (Fig. 5 and Table 3). Bacterial networks from the root endosphere and rhizosphere shared the different trend of becoming denser over time. Density, average degree and edges of the network from the root endosphere and rhizosphere gradually increased with the planting years. And the total participation of the five core genera in the root endophytic microbial molecular ecology network (MEN) increased with planting, as shown by growing node count and degree (Table S3).
Null model analysis showed that the relative importance of deterministic (|βNTI|≥ 2) and stochastic (|βNTI|< 2) processes in the P. polyphylla microbiome showed a great difference in different compartments. The relative contribution of deterministic processes in microbial community assembly in bulk soil (≥ 43%) was the highest, followed by root endosphere(≤ 43%) and rhizosphere soil (≤ 23%). Stochasticity dominated the community assembly process of the root endosphere, and the relative contribution increased with planting years (57%-78%). The deterministic selection (71%) initially governed community assembly in bulk soil, but stochasticity progressed as the planting age increased (Fig. 6a, b). Stochasticity contributed most to the rhizosphere soil microbial community assembly among the three compartments and dominated their community assembly process. However, the proportion of stochasticity decreased from 91 to 77% with P. polyphylla planting, which showed a significant difference from the other two compartments. Moreover, heterogeneous selection dominated the deterministic process of bacterial community assembly, while the stochastic process was dominated by dispersal limitation (Fig. 6b). Collectively, deterministic processes exerted a greater influence on the bulk soil bacterial community and planting exerted different effects on bacterial community assembly in the three compartments.
Deterministic and stochastic processes in microbiome assembly. a The relative contribution of determinism and stochasticity on microbiome assembly along the soil–root continuum based on the β-Nearest Taxon Index (βNTI) values. The βNTI were calculated by the Null model test, and |βNTI|≥ 2 and |βNTI|< 2 represent dominant determinism and stochasticity in driving microbiome assembly, respectively. The percentage above and below the violin plot represents the proportion of the deterministic and stochastic processes in microbiome assembly, respectively. b The relative importance of five ecological processes (heterogeneous selection: βNTI < − 2, homogeneous selection: βNTI > + 2, dispersal limitation: |βNTI|< 2 and RCBray > 0.95, homogenizing dispersal: |βNTI|< 2 and RCBray < – 0.95, and undominated: |βNTI|< 2 and |RCBray|< 0.95) along the soil–root continuum based on the β-Nearest Taxon Index (βNTI) and Bray–Curtis-based Raup-Crick Index (RCBray)
Effects of planting years and compartments on the functions of P. polyphylla root-associated microbial communities
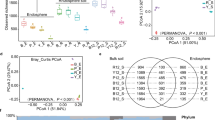
Using PICRUSt as a predictive exploratory tool of function gene, it was found that six orthology groups at the level I (KOs in KEGG) were observed in P. polyphylla roots and root-related soils. The rhizosphere and root endosphere significantly increased cellular processes and environmental information processing associated genes (Fig. 7a). The two genes (cellular processes and environmental information processing associated genes) increased with the planting years in the three compartments. At level III, genes related to nitrogen metabolism, carbon metabolism, phosphonate and phosphinate metabolism were noted to significantly enrich in root endosphere and rhizosphere soil (Fig. 7b). Communities of P. polyphylla root endosphere had more functional gene-encoding protein involved in methyl-accepting chemotaxis (K03406), while bulk soils had the least. And these genes tended to increase in bulk soil over time significantly (Fig. 7a). Different from the microbial community, functional genes' alpha diversity (Simpson_evenness) increased from bulk soil to rhizosphere to root endosphere and increased with age in bulk soils (Fig. 7c). PICRUSt results indicated that the functional composition (i.e., PCA analysis of KEGG Orthology) of P. polyphylla microbiome in different compartments was significantly different. Planting years also significantly affected P. polyphylla root-associated microbiome functions in the three compartments (Fig. 7d). In PCA of Bray–Curtis distance from all samples, bulk soil samples clustered together and were far from the rhizosphere and root endosphere across P. polyphylla planting years (Fig. 7d). Additionally, in the third axis, the P. polyphylla growing years were shown to affect the shift of the functional genes (Fig. 7d). The planting years and compartments significantly affected the richness and diversity of P. polyphylla root-associated microbial community function.
Functional genes of bacterial communities in P. polyphylla roots and root-related soils. a The abundance of genes related to cellular processes, environmental information processing, and K03406 (an encoding protein involved in methyl-accepting chemotaxis). The statistical significance was measured using the Turkey test's SD (n = 6). b Nitrogen metabolism, carbon metabolism and phosphonate and phosphinate metabolism genes’ abundance. “N, C, P” represent genes related to dinitrogen metabolism, carbon metabolism and phosphonate and phosphinate metabolism, respectively. Mean values ± SD (n = 6). c The alpha diversity (Simpson_evenness) of functional genes. Different letters above the boxes indicate a significant difference determined by the Turkey test. d Non-metric multidimensional scaling (NMDS) based on Bray–Curtis distance
Discussion
P. polyphylla selectively enriches its core microbiome with beneficial microorganisms
Microbial differences in different compartments and microbial succession over time have been studied in previous studies (Fan et al. 2017; Zhang et al. 2018; Xiong et al. 2021); this is attributed to significant environmental differences among compartments (Jiang et al. 2017; Xiong et al. 2021). However, there is still a lack of research on the microorganisms related to P. polyphylla. This study sampled different compartments of P. polyphylla for three consecutive years. It was found that the microbial community among P. polyphylla compartments showed significant differences in composition and diversity. Moreover, bacterial alpha diversity in P. polyphylla rhizosphere was generally lower in bulk soil but higher in root endosphere (Fig. 3a). That is common in other plants and was also reported in P. polyphylla (Wang et al. 2020; Ling et al. 2022). The rhizosphere microbiota was generally considered a community subset in bulk soil, ubiquitous in planting soils (Xiong et al. 2021). The microorganisms in the root endosphere are mainly absorbed from the rhizosphere, except from seeds (Wani et al. 2015; Zhang et al. 2018). Bacterial alpha diversity represented by the Shannon index showed a significant reduction in bulk soil and root endosphere but increased in P. polyphylla rhizosphere over time (Fig. 3b). In bulk soil, it may be affected by the enrichment of pathogenic microorganisms, reported in the study of Panax notoginseng (Tan et al. 2017). While in rhizosphere soil, the bacterial diversity might be regulated by P. polyphylla root exudates. Previous research showed root exudates, including phytohormones and defensive compounds, dominated rhizosphere microbial community assembly in some plants (Nannipieri et al. 2008). Slightly different from P. polyphylla, the diversity of ginseng endophytes was highest in the third year and decreased gradually but increased from the second to the third year (Hong et al. 2019). Reasonable speculation is that plant roots extensively absorb microorganisms in the soil to enrich their functional gene pool early in planting and then gradually selectively retain the beneficial microorganisms. P. polyphylla might take less time than ginseng in the first stage. Therefore, the regulation of root microbial assembly by root exudates and signaling molecules also seems to develop in P. polyphylla roots.
Plants exert selective effects on the soil pool to acquire beneficial microorganisms and assemble their core microbiota (Reinhold-Hurek and Hurek 2011). In this study, the bulk soil enriched more Acidobacterial microorganisms because the planting soil was acidic, and the pH decreased with the years of planting (Table 1) (Nguyen et al. 2016). In P. polyphylla root endosphere and rhizosphere soil, Proteobacteria dominate the microbiome. Proteobacteria are used to live in C-rich environments (common in the rhizosphere and root endosphere) for high physiological activity, a common phenomenon in plants (Kuzyakov and Razavi 2019). The core microbiota of P. polyphylla root consisted of Pseudomonas, Rhizobium, Steroidobacter, Sphingobium and Agrobacterium (Figs. 1, 2). These microorganisms were representative plant growth-promoting rhizobacteria (PGPR) (Basu et al. 2021). Pseudomonas and Sphingobium were the core microbiomes of Brassica napus and Salvia miltiorrhiza (Wu et al. 2016; Chen et al. 2018). Pseudomonas, Sphingobium, Rhizobium and Agrobacterium were proven to be vital phosphate solubilizers (Otieno et al. 2015; Li et al. 2017). In addition, Pseudomonas was also found to solubilize potassium and produce indole acetic acid (IAA) and other plant growth regulators (Arruda et al. 2014). It was revealed that Steroidobacter and Sphingobium were beneficial to plants (Li et al. 2017; Zhu et al. 2021). Pseudomonas in the root endosphere increased with polyphyllin I, which indicated that plant metabolites regulated endophytic bacteria (Table 2 and Fig. S4). OTUs belonging to these genera also played important roles in the community network of P. polyphylla rhizosphere soils and root endosphere (Table S3). It further illustrated the importance of these microorganisms in the growth and development of P. polyphylla. P. polyphylla chose to enrich such microbes in favor of their growth. The previous study reported the same conclusion on P. polyphylla (Zhou et al. 2015). Although these core microbes might play a vital role in the growth of P. polyphylla, their relative abundance reached the lowest level in the bulk soil in the fourth year, below 0.1% (Fig. S4). Therefore, supplementing these microorganisms in the bulk soil might benefit P. polyphylla. These findings remind us that supplementing these core microorganisms in bulk soil could improve soil fertility and P. polyphylla growth after P. polyphylla planting.
The community assembly of P. polyphylla microbes was influenced by compartment and planting years
The community assembly process is an important method for understanding the development dynamics of the community (Singh et al. 2020; Trivedi et al. 2020). Clarifying the ecological processes of plant microbiome assembly is essential to analyze the interaction between P. polyphylla and microbiome and advance the future application of microbiome to P. polyphylla. Our findings demonstrated that compartment and planting years influenced P. polyphylla microbiome assembly (Fig. 6). Niche isolation must be the key factor that causes the difference in community assembly among compartments (Cregger et al. 2018). Microbiomes in bulk soil were more sensitive to the planting years than root endosphere and rhizosphere microbiomes in terms of multiple microbial attributes (i.e., alpha-diversity, beta-diversity and assembly processes), environmental properties may influence this in the bulk soil. The increasing stochastic processes in P. polyphylla root endosphere and bulk soil community assembly might result from decreased alpha diversity and richness over time. The proportion of the homogeneous selection process on the P. polyphylla rhizosphere microbial community increased with the chronosequence advancing, which had also been found in the soybean rhizosphere (Goss-Souza et al. 2020). Root exudates might regulate the unique variation in rhizosphere soil. The relative proportion variation of stochastic processes in rhizosphere soil community assembly was similar to its alpha diversity. Ecological stochasticity was defined as the changes in the community to stochastic processes of birth, death, immigration and emigration, spatiotemporal variation, and historical contingency (e.g., colonization order), which is undoubtedly related to the species richness and alpha diversity of the community (Zhou and Ning 2017). The principal component of stochastic processes in the P. polyphylla root community, dispersal limitation, showed the opposite trend. Previous studies have also revealed that dispersal limitation was an essential factor shaping species richness in agricultural landscapes (Hendrickx et al. 2009). This work shows that species richness and planting years profoundly influence plant microbiome assembly.
Previous studies showed wheat and Avena fatua rhizosphere soil and root endosphere exhibited more complex topology than bulk soil (Fan et al. 2018; Wei et al. 2021). At the same time, the rhizosphere soil community network was revealed over time of Avena fatua growth (Shi et al. 2016) and network analysis was conducted to explore the interaction of microorganisms in different P. polyphylla compartments for three consecutive years. More complex topology was shown in rhizosphere soil and root endosphere than in bulk soil bacterial community, although fewer OTUs were involved in network construction (Fig. 5). High connectance in root endosphere and rhizosphere soil communities could decrease pathogen invasion success (Wei et al. 2015). Therefore, this might be a measure of self-protection by P. polyphylla. In addition, the increase was also observed in the connectivity and complexity of P. polyphylla rhizosphere soil and root endosphere community networks over time, representing significant differences among compartments. Some studies showed that the roots promote the development of dominant taxa, which would concurrently reduce diversity, leading to greater interactions and more complex networks over time (Shi et al. 2016). A consistent conclusion was made in the P. polyphylla root endosphere, different in the rhizosphere. Multiple mechanisms might contribute to the connectivity and complexity of this rhizosphere soil network, such as root exudates (Shi et al. 2011). As reported in previous studies, differences in nutrition, niche, environment properties, etc., might lead to closer associations and more complex networks in root endosphere and rhizosphere soil (Fan et al. 2018). The core microorganisms, including Sphingobium, Pseudomonas, Steroidobacter, Agrobacterium, and Sphingomonas, were more involved in the network’s construction in the root endosphere and rhizosphere. These microorganisms play important roles in the network, so the low connectivity in bulk soil might be due to the lack of these microorganisms. Adding these microorganisms to bulk soils seems to be an excellent method to enhance the complexity and stability of the soil microbial networks, which could enhance the stress resistance of P. polyphylla. These conclusions and further research would offer us the perfect timing for adding microbial inoculants.
P. polyphylla also plays a vital role in shaping the function of the rhizosphere microbial community
In general, the environmental selection of microorganisms is mainly reflected in the collection of functions (Hammesfahr et al. 2011; Yan et al. 2017). In this study, through analysis of function genes predicted by PICRUSTs, the differences in function genes were found among compartments across planting years (Fig. 7). The higher carbon metabolism genes in the root and the rhizosphere indicate that the organic matter secreted by the P. polyphylla root endosphere and rhizosphere attracts numbers of C-loving microorganisms. It was proven by the enriching Proteobacteria in the root endosphere and rhizosphere (Fig. 1) (Kuzyakov and Razavi 2019). The increase of these genes in the bulk soil might be attributed to the rise of TOC in bulk soil across the three years (Table 1). A consensus is that N and P are essential to plant growth and development. Therefore, the increase of genes related to nitrogen metabolism in the bacterial community might be regulated by P. polyphylla for more nutrition. It can be summarized from the enrichment of Rhizobiales in P. polyphylla root and rhizosphere. The reason for the enrichment of phosphonate and phosphonate metabolism genes and nitrogen metabolism-related genes over time in the three compartments may be the combined effects of soil properties and plant selection (Fig. 7b). The increase of TN and TP (Table 1) in the bulk soil encouraged the accumulation of microorganisms with related genes to help P. polyphylla to obtain nutrients (Fig. 7b). The functional gene encoding methyl-accepting chemotaxis (K03406, related to signaling in plant–microbe interactions) was significantly enriched in P. polyphylla rhizosphere and root endosphere (Fig. 7a). It was also a significant evidence that the rhizosphere and root endosphere microbes interacted more with the plant than bulk soil. It was also found that these genes increased over time despite the minor abundance in the bulk soil. It seems to reveal that the selective pressure of plants on bulk soil rises over time. These findings showed that P. polyphylla enriched function genes related to nitrogen and phosphonate metabolism to help their growth, which is consistent with the results of community composition. These results help us uncover the complex plant–microbe interactions and provide an essential guide for applying microorganisms in agriculture.
Conclusion
This study provides comprehensive evidence on the effects of planting years on the P. polyphylla-associative microbes community assembly. Our results show that P. polyphylla-associative microbes were affected by both compartments and the planting years. In addition, it was uncovered that P. polyphylla selection sequentially increased and significantly affected community diversity and network complexity from bulk soils to rhizosphere soils to root endosphere. These findings significantly promote our understanding of the bacterial community assembly in P. polyphylla root and highlight the importance of the host selection effect.
Furthermore, the core microbiome in P. polyphylla roots was enriched over time, including Pseudomonas, Rhizobium, Steroidobacter, Sphingobium and Agrobacterium. Moreover, the core microorganisms of P. polyphylla root reached the lowest level in the bulk soil in the fourth year, with relative abundance below 0.1%. The results indicated that adding phosphate-solubilizing and nitrogen-fixing microorganisms, such as Pseudomonas, could contribute to the stress resistance and growth of P. polyphylla. Our work advances the understanding of dynamic variation host-microbiome assembly.
Data availability
The raw data have been submitted to the NCBI SRA database, which was available. The accession number of all samples is PRJNA 824132. The data of plot construction were provided with supplementary materials.
References
Akiyama K, Matsuzaki K-I, Hayashi H (2005) Plant sesquiterpenes induce hyphal branching in arbuscular mycorrhizal fungi. Nature 435(7043):824–827
Ankati S, Podile AR (2019) Metabolites in the root exudates of groundnut change during interaction with plant growth promoting rhizobacteria in a strain-specific manner. J Plant Physiol 243:153057. https://doi.org/10.1016/j.jplph.2019.153057
Arruda L, Beneduzzi A, Lisboa B, Passaglia L,Vargas LK (2014) Diversity of plant-growth-promoting rhizobacteria associated with maize (Zea mays L.). In: Maheshwari DK (ed) Bacterial diversity in sustainable agriculture. Springer, Cham, pp 167–189. https://doi.org/10.1007/978-3-319-05936-5_7
Bai B, Liu W, Qiu X, Zhang J, Zhang J, Bai Y (2022) The root microbiome: community assembly and its contributions to plant fitness. J Integr Plant Biol 64(2):230–243. https://doi.org/10.1111/jipb.13226
Bakker P, Berendsen R, Doornbos R, Wintermans P, Pieterse C (2013) The rhizosphere revisited: root microbiomics [Mini Review]. Front Plant Sci 4:165. https://doi.org/10.3389/fpls.2013.00165
Basu A, Prasad P, Das SN, Kalam S, Sayyed RZ, Reddy MS, El Enshasy H (2021) Plant growth promoting rhizobacteria (PGPR) as green bioinoculants: recent developments, constraints, and prospects. Sustainability 13(3):1140
Berg G, Smalla K (2009) Plant species and soil type cooperatively shape the structure and function of microbial communities in the rhizosphere. FEMS Microbiol Ecol 68(1):1–13. https://doi.org/10.1111/j.1574-6941.2009.00654.x
Bolyen E, Rideout JR, Dillon MR, Bokulich NA, Abnet CC et al (2019) Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat Biotechnol 37(8):852–857. https://doi.org/10.1038/s41587-019-0209-9
Chaparro JM, Badri DV, Vivanco JM (2014) Rhizosphere microbiome assemblage is affected by plant development. ISME J 8(4):790–803. https://doi.org/10.1038/ismej.2013.196
Chapman N, Miller AJ, Lindsey K, Whalley WR (2012) Roots, water, and nutrient acquisition: let’s get physical. Trends Plant Sci 17(12):701–710. https://doi.org/10.1016/j.tplants.2012.08.001
Chen H, Wu H, Yan B, Zhao H, Liu F, Zhang H, Sheng Q, Miao F, Liang Z (2018) Core microbiome of medicinal plant Salvia miltiorrhiza seed: a rich reservoir of beneficial microbes for secondary metabolism? Int J Mol Sci 19(3):672
Clarke BB, White JF, Hurley RH, Torres MS, Sun S, Huff DR (2006) Endophyte-mediated suppression of dollar spot disease in fine fescues. Plant Dis 90(8):994–998. https://doi.org/10.1094/pd-90-0994
Cregger MA, Veach AM, Yang ZK, Crouch MJ, Vilgalys R, Tuskan GA, Schadt CW (2018) The Populus holobiont: dissecting the effects of plant niches and genotype on the microbiome. Microbiome 6(1):31. https://doi.org/10.1186/s40168-018-0413-8
Deng Y, Huang H, Lei F, Fu S, Zou K, Zhang S, Liu X, Jiang L, Liu H, Miao B, Liang Y (2021) Endophytic bacterial communities of Ginkgo biloba leaves during leaf developmental period. Front Microbiol 12:698703. https://doi.org/10.3389/fmicb.2021.698703
DeSantis TZ, Hugenholtz P, Larsen N, Rojas M, Brodie EL, Keller K, Huber T, Dalevi D, Hu P, Andersen GL (2006) Greengenes, a chimera-checked 16S rRNA gene database and workbench compatible with ARB. Appl Environ Microbiol 72(7):5069–5072. https://doi.org/10.1128/AEM.03006-05
Develey-Rivière MP, Galiana E (2007) Resistance to pathogens and host developmental stage: a multifaceted relationship within the plant kingdom. New Phytol 175(3):405–416
Edwards J, Johnson C, Santos-Medellín C, Lurie E, Podishetty NK, Bhatnagar S, Eisen JA, Sundaresan V (2015) Structure, variation, and assembly of the root-associated microbiomes of rice. Proc Natl Acad Sci USA 112(8):E911–E920. https://doi.org/10.1073/pnas.1414592112
Fan K, Cardona C, Li Y, Shi Y, Xiang X, Shen C, Wang H, Gilbert JA, Chu H (2017) Rhizosphere-associated bacterial network structure and spatial distribution differ significantly from bulk soil in wheat crop fields. Soil Biol Biochem 113:275–284. https://doi.org/10.1016/j.soilbio.2017.06.020
Fan K, Weisenhorn P, Gilbert JA, Chu H (2018) Wheat rhizosphere harbors a less complex and more stable microbial co-occurrence pattern than bulk soil. Soil Biol Biochem 125:251–260. https://doi.org/10.1016/j.soilbio.2018.07.022
Fitzpatrick CR, Salas-González I, Conway JM, Finkel OM, Gilbert S, Russ D, Teixeira PJPL, Dangl JL (2020) The plant microbiome: from ecology to reductionism and beyond. Annu Rev Microbiol 74(1):81–100. https://doi.org/10.1146/annurev-micro-022620-014327
Fu S, Deng Y, Zou K, Zhang S, Liu X, Liang Y (2022) Flavonoids affect the endophytic bacterial community in Ginkgo biloba leaves with increasing altitude. Front Plant Sci 13:982771. https://doi.org/10.3389/fpls.2022.982771
Goss-Souza D, Mendes LW, Rodrigues JLM, Tsai SM (2020) Ecological processes shaping bulk soil and rhizosphere microbiome assembly in a long-term Amazon forest-to-agriculture conversion. Microb Ecol 79(1):110–122. https://doi.org/10.1007/s00248-019-01401-y
Hammesfahr U, Bierl R, Thiele-Bruhn S (2011) Combined effects of the antibiotic sulfadiazine and liquid manure on the soil microbial-community structure and functions. J Plant Nutr Soil Sci 174(4):614–623. https://doi.org/10.1002/jpln.201000322
Haskett TL, Tkacz A, Poole PS (2021) Engineering rhizobacteria for sustainable agriculture. ISME J 15(4):949–964. https://doi.org/10.1038/s41396-020-00835-4
Hendrickx F, Maelfait J-P, Desender K, Aviron S, Bailey D, Diekotter T, Lens L, Liira J, Schweiger O, Speelmans M, Vandomme V, Bugter R (2009) Pervasive effects of dispersal limitation on within- and among-community species richness in agricultural landscapes. Glob Ecol Biogeogr 18(5):607–616. https://doi.org/10.1111/j.1466-8238.2009.00473.x
Hirsch PR, Mauchline TH (2012) Who’s who in the plant root microbiome? Nat Biotechnol 30(10):961–962. https://doi.org/10.1038/nbt.2387
Hong CE, Kim JU, Lee JW, Bang KH, Jo IH (2019) Metagenomic analysis of bacterial endophyte community structure and functions in Panax ginseng at different ages. 3 Biotech 9(8):300. https://doi.org/10.1007/s13205-019-1838-x
Hu L, Robert CA, Cadot S, Zhang X, Ye M, Li B, Manzo D, Chervet N, Steinger T, Van Der Heijden MG (2018) Root exudate metabolites drive plant-soil feedbacks on growth and defense by shaping the rhizosphere microbiota. Nat Commun 9(1):1–13
Jiang Y, Li S, Li R, Zhang J, Liu Y, Lv L, Zhu H, Wu W, Li W (2017) Plant cultivars imprint the rhizosphere bacterial community composition and association networks. Soil Biol Biochem 109:145–155. https://doi.org/10.1016/j.soilbio.2017.02.010
Kanehisa M, Goto S (2000) KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res 28(1):27–30. https://doi.org/10.1093/nar/28.1.27
Kuzyakov Y, Razavi BS (2019) Rhizosphere size and shape: temporal dynamics and spatial stationarity. Soil Biol Biochem 135:343–360. https://doi.org/10.1016/j.soilbio.2019.05.011
Langille MG, Zaneveld J, Caporaso JG, McDonald D, Knights D, Reyes JA, Clemente JC, Burkepile DE, Vega Thurber RL, Knight R, Beiko RG, Huttenhower C (2013) Predictive functional profiling of microbial communities using 16S rRNA marker gene sequences. Nat Biotechnol 31(9):814–821. https://doi.org/10.1038/nbt.2676
Li Y, Liu X, Hao T, Chen S (2017) Colonization and maize growth promotion induced by phosphate solubilizing bacterial isolates. Int J Mol Sci 18(7):1253
Ling N, Wang T, Kuzyakov Y (2022) Rhizosphere bacteriome structure and functions. Nat Commun 13(1):836. https://doi.org/10.1038/s41467-022-28448-9
Liu TH, Zhou Y, Tao WC, Liu Y, Zhang XM, Tian SZ (2020) bacterial diversity in roots, stems, and leaves of Chinese medicinal plant Paris polyphylla var. yunnanensis. Pol J Microbiol 69(1):91–97. https://doi.org/10.33073/pjm-2020-012
Malinowski DP, Alloush GA, Belesky DP (2000) Leaf endophyte Neotyphodium coenophialum modifies mineral uptake in tall fescue. Plant Soil 227(1):115–126. https://doi.org/10.1023/A:1026518828237
Martin FM, Uroz S, Barker DG (2017) Ancestral alliances: plant mutualistic symbioses with fungi and bacteria. Science 356(6340):eaad4501. https://doi.org/10.1126/science.aad4501
Mendes R, Garbeva P, Raaijmakers JM (2013) The rhizosphere microbiome: Significance of plant beneficial, plant pathogenic, and human pathogenic microorganisms. FEMS Microbiol Rev 37(5):634–663. https://doi.org/10.1111/1574-6976.12028
Nannipieri P, Ascher J, Ceccherini MT, Landi L, Pietramellara G, Renella G, Valori F (2008) Effects of root exudates in microbial diversity and activity in rhizosphere soils. In: Nautiyal CS, Dion P (eds) Molecular mechanisms of plant and microbe coexistence. Springer, Berlin, pp 339–365. https://doi.org/10.1007/978-3-540-75575-3_14
Nguyen NL, Kim YJ, Hoang VA, Subramaniyam S, Kang JP, Kang CH, Yang DC (2016) Bacterial diversity and community structure in Korean ginseng field soil are shifted by cultivation time. PLoS One 11(5):e0155055. https://doi.org/10.1371/journal.pone.0155055
O’Brien AM, Harrison TL (2021) Host match improves root microbiome growth. Nat Microbiol 6(9):1103–1104. https://doi.org/10.1038/s41564-021-00957-1
Otieno N, Lally R, Kiwanuka S, Lloyd A, Ryan D, Germaine K, Dowling D (2015) Plant growth promotion induced by phosphate solubilizing endophytic Pseudomonas isolates. Front Microbiol 6:745. https://doi.org/10.3389/fmicb.2015.00745
Qu Q, Zhang Z, Peijnenburg WJGM, Liu W, Lu T, Hu B, Chen J, Chen J, Lin Z, Qian H (2020) Rhizosphere microbiome assembly and its impact on plant growth. J Agric Food Chem 68(18):5024–5038. https://doi.org/10.1021/acs.jafc.0c00073
Reinhold-Hurek B, Hurek T (2011) Living inside plants: bacterial endophytes. Curr Opin Plant Biol 14(4):435–443. https://doi.org/10.1016/j.pbi.2011.04.004
Saunders M, Kohn LM (2009) Evidence for alteration of fungal endophyte community assembly by host defense compounds. New Phytol 182(1):229–238. https://doi.org/10.1111/j.1469-8137.2008.02746.x
Schardl CL, Leuchtmann A, Spiering MJ (2004) Symbioses of grasses with seedborne fungal endophytes. Annu Rev Plant Biol 55(1):315–340. https://doi.org/10.1146/annurev.arplant.55.031903.141735
Segata N, Izard J, Waldron L, Gevers D, Miropolsky L, Garrett WS, Huttenhower C (2011) Metagenomic biomarker discovery and explanation. Genome Biol 12(6):R60. https://doi.org/10.1186/gb-2011-12-6-r60
Sessitsch A, Pfaffenbichler N, Mitter B (2019) Microbiome applications from lab to field: facing complexity. Trends Plant Sci 24(3):194–198. https://doi.org/10.1016/j.tplants.2018.12.004
Shi S, Richardson AE, O’Callaghan M, DeAngelis KM, Jones EE, Stewart A, Firestone MK, Condron LM (2011) Effects of selected root exudate components on soil bacterial communities. FEMS Microbiol Ecol 77(3):600–610. https://doi.org/10.1111/j.1574-6941.2011.01150.x
Shi S, Nuccio EE, Shi ZJ, He Z, Zhou J, Firestone MK (2016) The interconnected rhizosphere: high network complexity dominates rhizosphere assemblages. Ecol Lett 19(8):926–936. https://doi.org/10.1111/ele.12630
Singh BK, Trivedi P, Egidi E, Macdonald CA, Delgado-Baquerizo M (2020) Crop microbiome and sustainable agriculture. Nat Rev Microbiol 18(11):601–602. https://doi.org/10.1038/s41579-020-00446-y
Sorty AM, Meena KK, Choudhary K, Bitla UM, Minhas PS, Krishnani KK (2016) Effect of plant growth promoting bacteria associated with halophytic weed (Psoralea corylifolia L) on germination and seedling growth of wheat under saline conditions. Appl Biochem Biotechnol 180(5):872–882. https://doi.org/10.1007/s12010-016-2139-z
Stegen JC, Lin X, Fredrickson JK, Chen X, Kennedy DW, Murray CJ, Rockhold ML, Konopka A (2013) Quantifying community assembly processes and identifying features that impose them. ISME J 7(11):2069–2079. https://doi.org/10.1038/ismej.2013.93
Sun H, Jiang S, Jiang C, Wu C, Gao M, Wang Q (2021) A review of root exudates and rhizosphere microbiome for crop production. Environ Sci Poll Res 28(39):54497–54510. https://doi.org/10.1007/s11356-021-15838-7
Tan Y, Cui Y, Li H, Kuang A, Li X, Wei Y, Ji X (2017) Diversity and composition of rhizospheric soil and root endogenous bacteria in Panax notoginseng during continuous cropping practices. J Basic Microbiol 57(4):337–344. https://doi.org/10.1002/jobm.201600464
Tao L, Qiuhong L, Fuqiang Y, Shuhui Z, Suohui T, Linyuan F (2021) Plant growth-promoting activities of bacterial endophytes isolated from the medicinal plant Pairs polyphylla var. yunnanensis. World J Microbiol Biotechnol 38(1):15. https://doi.org/10.1007/s11274-021-03194-0
Team TG (2008) Gephi, an open source graph visualization and manipulation software
Trivedi P, Leach JE, Tringe SG, Sa T, Singh BK (2020) Plant-microbiome interactions: from community assembly to plant health. Nat Rev Microbiol 18(11):607–621. https://doi.org/10.1038/s41579-020-0412-1
Vacheron J, Desbrosses G, Bouffaud M-L, Touraine B, Moënne-Loccoz Y, Muller D, Legendre L, Wisniewski-Dyé F, Prigent-Combaret C (2013) Plant growth-promoting rhizobacteria and root system functioning. Front Plant Sci 4:356. https://doi.org/10.3389/fpls.2013.00356
Vandenkoornhuyse P, Quaiser A, Duhamel M, Le Van A, Dufresne A (2015) The importance of the microbiome of the plant holobiont. New Phytol 206(4):1196–1206. https://doi.org/10.1111/nph.13312
Vives-Peris V, de Ollas C, Gómez-Cadenas A, Pérez-Clemente RM (2020) Root exudates: from plant to rhizosphere and beyond. Plant Cell Rep 39(1):3–17. https://doi.org/10.1007/s00299-019-02447-5
Wang Y, Wang H, Cheng H, Chang F, Wan Y, She X (2020) Niche differentiation in the rhizosphere and endosphere fungal microbiome of wild Paris polyphylla Sm. PeerJ 8:e8510. https://doi.org/10.7717/peerj.8510
Wani ZA, Ashraf N, Mohiuddin T, Riyaz-Ul-Hassan S (2015) Plant-endophyte symbiosis, an ecological perspective. Appl Microbiol Biotechnol 99(7):2955–2965. https://doi.org/10.1007/s00253-015-6487-3
Wei Z, Yang T, Friman VP, Xu Y, Shen Q, Jousset A (2015) Trophic network architecture of root-associated bacterial communities determines pathogen invasion and plant health. Nat Commun 6:8413. https://doi.org/10.1038/ncomms9413
Wei G, Ning K, Zhang G, Yu H, Yang S, Dai F, Dong L, Chen S (2021) Compartment niche shapes the assembly and network of Cannabis sativa-associated microbiome. Front Microbiol 12:714993. https://doi.org/10.3389/fmicb.2021.714993
Wu L, Liu R, Niu Y, Lin H, Ye W, Guo L, Hu X (2016) Whole genome sequence of Pantoea ananatis R100, an antagonistic bacterium isolated from rice seed. J Biotechnol 225:1–2
Wu Y, Yang Y, Cao L, Yin H, Xu M, Wang Z, Liu Y, Wang X, Deng Y (2018) Habitat environments impacted the gut microbiome of long-distance migratory swan geese but central species conserved. Sci Rep 8(1):13314. https://doi.org/10.1038/s41598-018-31731-9
Xiong C, Singh BK, He JZ, Han YL, Li PP, Wan LH, Meng GZ, Liu SY, Wang JT, Wu CF, Ge AH, Zhang LM (2021) Plant developmental stage drives the differentiation in ecological role of the maize microbiome. Microbiome 9(1):171. https://doi.org/10.1186/s40168-021-01118-6
Yan Y, Kuramae EE, de Hollander M, Klinkhamer PG, van Veen JA (2017) Functional traits dominate the diversity-related selection of bacterial communities in the rhizosphere. ISME J 11(1):56–66. https://doi.org/10.1038/ismej.2016.108
Yang Y, Yang SC, Zhao J, Udikeri S, Liu T (2015) Microbial diversity in Paris polyphylla var. yunnanensis rhizomes of varying ages. Genet Mol Res 14(4):17612–17621. https://doi.org/10.4238/2015.December.21.34
Zhang J, Zhang N, Liu YX, Zhang X, Hu B, Qin Y, Xu H, Wang H, Guo X, Qian J, Wang W, Zhang P, Jin T, Chu C, Bai Y (2018) Root microbiota shift in rice correlates with resident time in the field and developmental stage. Sci China Life Sci 61(6):613–621. https://doi.org/10.1007/s11427-018-9284-4
Zhao J, Mou Y, Shan T, Li Y, Zhou L, Wang M, Wang J (2010) Antimicrobial metabolites from the endophytic fungus Pichia guilliermondii isolated from Paris polyphylla var. yunnanensis. Molecules 15(11):7961–7970. https://doi.org/10.3390/molecules15117961
Zhou N, Qi W, Xiao G, Ding B, Zhang H, Guo D, Wei S (2015) [Correlation between distribution of rhizospheric microorganisms and contents of steroidal saponins of Paris polyphylla var. yunnanensis]. China J Chin Materia Medica 40(6):1055
Zhou J, Ning D (2017) Stochastic community assembly: Does it matter in microbial ecology? Microbiol Mol Biol Rev 81(4):e0002–e00017. https://doi.org/10.1128/MMBR.00002-17
Zhu J, Cao A, Wu J, Fang W, Huang B, Yan D, Wang Q, Li Y (2021) Effects of chloropicrin fumigation combined with biochar on soil bacterial and fungal communities and Fusarium oxysporum. Ecotoxicol Environ Saf 220:112414. https://doi.org/10.1016/j.ecoenv.2021.112414
Zou K, Liu X, Hu Q, Zhang D, Fu S, Zhang S, Huang H, Lei F, Zhang G, Miao B, Meng D, Jiang L, Liu H, Yin H, Liang Y (2021) Root endophytes and Ginkgo biloba are likely to share and compensate secondary metabolic processes, and potentially exchange genetic information by LTR-RTs. Front Plant Sci 12:704985. https://doi.org/10.3389/fpls.2021.704985
Funding
This work was supported by the following grants: the National Natural Science Foundation of China (31570113).
Author information
Authors and Affiliations
Contributions
Shaodong Fu and Yili Liang designed the experiments, analyzed the data, and wrote the manuscript. Yan Deng, Kai Zou and Shuangfei Zhang helped to perform the experiments. Yili Liang revised this article. All authors read and approved of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that the research was conducted without any commercial or financial relationships construed as a potential conflict of interest.
Additional information
Communicated by Dorothea Bartels.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Fu, S., Deng, Y., Zou, K. et al. Dynamic variation of Paris polyphylla root-associated microbiome assembly with planting years. Planta 257, 61 (2023). https://doi.org/10.1007/s00425-023-04074-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00425-023-04074-7