Abstract
Plant non-specific lipid transfer proteins (nsLTPs) are encoded by a multigene family and support physiological functions, which remain unclear. We adapted an efficient ligation-mediated polymerase chain reaction (LM-PCR) procedure that enabled isolation of 22 novel Triticum aestivum nsLtp (TaLtp) genes encoding types 1 and 2 nsLTPs. A phylogenetic tree clustered the wheat nsLTPs into ten subfamilies comprising 1–7 members. We also studied the activity of four type 1 and two type 2 TaLtp gene promoters in transgenic rice using the β-Glucuronidase reporter gene. The activities of the six promoters displayed both overlapping and distinct features in rice. In vegetative organs, these promoters were active in leaves and root vascular tissues while no β-Glucuronidase (GUS) activity was detected in stems. In flowers, the GUS activity driven by the TaLtp7.2a, TaLtp9.1a, TaLtp9.2d, and TaLtp9.3e gene promoters was associated with vascular tissues in glumes and in the extremities of anther filaments whereas only the TaLtp9.4a gene promoter was active in anther epidermal cells. In developing grains, GUS activity and GUS immunolocalization data evidenced complex patterns of activity of the TaLtp7.1a, TaLtp9.2d, and TaLtp9.4a gene promoters in embryo scutellum and in the grain epicarp cell layer. In contrast, GUS activity driven by TaLtp7.2a, TaLtp9.1a, and TaLtp9.3e promoters was restricted to the vascular bundle of the embryo scutellum. This diversity of TaLtp gene promoter activity supports the hypothesis that the encoded TaLTPs possess distinct functions in planta.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Non-specific lipid transfer proteins (nsLTPs) are cysteine-rich proteins that are found throughout the plant kingdom. NsLTPs were originally defined by their capacity to transfer various lipid compounds between lipid bilayers in vitro (Kader et al. 1984), but are currently more often defined by sequence homology than on a functional basis. NsLTPs are encoded by a multigene family and can be classified in three types according to their primary structure (Boutrot et al. 2005). The most detailed characterization has been made of types 1 and 2 nsLTPs, which are 9 and 7 kDa basic proteins, respectively. Type 3 nsLTPs also display a 7 kDa molecular mass and all type 3 non-specific lipid transfer protein (nsLtp) genes show anther-specific expression (Lauga et al. 2000). NsLTPs belong to the eight-cysteine motif protein superfamily (José-Estanyol et al. 2004). These cysteine residues are involved in four disulfide bonds that stabilize the nsLTP tertiary structure. This structure is characterized by a hydrophobic cavity whose size plasticity allows the in vitro loading of a great variety of lipid compounds (Sy et al. 2003).
The existence of multiple nsLtp genes in many plant genomes was first revealed by Southern blot and then by analyzing expressed sequence tag (EST) databases. The number of nsLtp genes identified so far ranges from six in pepper (Liu et al. 2006) to 15 in Arabidopsis (Arondel et al. 2000). However, the recent analysis of the complete rice genome indicates the presence of 53 nsLtp genes (F. Boutrot et al. unpublished data).
The different members of several nsLtp gene families exhibit a wide range of expression profiles during plant development. For instance, RT-PCR and northern blot analysis revealed distinct regulations of expression in peach flowers (Botton et al. 2002) and in Arabidopsis and pepper plants (Arondel et al. 2000; Jung et al. 2003). Differences were also found at the protein level since lipid transfer activity differed between nsLTPs from grapevine (Coutos-Thévenot et al. 1993) and from wheat (de Lamotte, INRA, Montpellier, personal communication).
The nsLtp gene transcripts and corresponding proteins are mainly localized in epidermal cell layers such as the leaf and embryo epidermis or the fruit pericarp (for review see Kader 1996). Transcripts are also associated with the vascular system (Buhtz et al. 2004) or localized in anthers, mainly in tapetum cells (Vrinten et al. 1999). At the subcellular level, nsLTPs are mainly found in apoplastic spaces since they are synthesized as preproteins with a signal peptide that directs the mature protein to the secretory pathway (Kader 1996).
The physiological function of nsLTPs is not clear; however, the widely reported localization of nsLTPs in epidermal cell layers and the secretion of these proteins in cell walls support the hypothesis that they are involved in the deposition of cutin monomers that form the cuticle of leaf epidermal cells (Sterk et al. 1991). This hypothesis is further supported by positive correlations established between nsLTP accumulation and conditions that stimulate cuticle synthesis as drought (Cameron et al. 2006b). Several nsLtp genes were also preferentially expressed in the outer epidermal layer of fleshy fruits (Kader 1996; Botton et al. 2002; Liu et al. 2006). Because of the in vitro antibacterial and antifungal activities of certain nsLTPs (Cammue et al. 1995; Dubreil et al. 1998) and the induction of expression of many nsLtp genes in response to biotic infections (Molina and García-Olmedo 1993; Guiderdoni et al. 2002; Jung et al. 2003; Lu et al. 2005), plant nsLTPs are thought to be involved in plant defense mechanisms. NsLTPs could therefore play an indirect role in plant defense by establishing a mechanical barrier of cutin, and a direct role thanks to their intrinsic antibiotic properties and their preferential localization in epidermal cell layers.
Evidence for nsLTP biological activity was provided by Maldonado et al. (2002). Following plant-pathogen interaction, these authors showed that the Arabidopsis dir1 mutant displays a normal local phenotype but fails to develop systemic acquired resistance (SAR). The DIR1 gene encodes a putative nsLTP, which is thought to be involved in long-distance signaling (Maldonado et al. 2002). The localization of the DIR1 transcripts in phloem companion cells (Ivashikina et al. 2003) strongly supports the hypothesis that the SAR-associated molecule signal is transported through the phloem sieve tubes (van Bel and Gaupels 2004). With less than 24% sequence identity to Arabidopsis nsLTPs, the DIR1 protein is not phylogenetically distributed within the three nsLTPs types usually reported (F. Boutrot et al. unpublished data). Moreover, the strong phenotype associated with the disruption of the DIR1 gene indicates that the corresponding protein supports a physiological function that cannot be assigned to other nsLTPs. Biological activity was also demonstrated in a tobacco type 1 nsLTP1 by treating stem sections with recombinant nsLTP1 complexed with jasmonic acid. This treatment enhanced the resistance of tobacco against Phytophthora parasitica, which was not observed with application of nsLTP1 or jasmonic acid alone (Buhot et al. 2004). Several other physiological functions have been proposed for nsLTPs which are widely thought to play a central role in adaptation to abiotic stresses such as desiccation (Cameron et al. 2006b), cold or salinity (Jung et al. 2003). NsLTPs could also be involved in programed cell death since they could protect the living cells from the effects of proteolytic activities triggered during this cell process (Eklund and Edqvist 2003). NsLTPs are also thought to possess a physiological function in male reproductive tissues where they could be involved in the deposition of material in the developing pollen wall (Foster et al. 1992); however their precise physiological function in pollen remains to be elucidated.
In wheat, most knowledge on nsLTPs is related to their protein structure and technological properties (for review see Marion et al. 2003). Transcript accumulation of only a few Triticum aestivum nsLtp genes has been monitored during grain development (Altenbach and Kothari 2004; Boutrot et al. 2005), in crown of wheat subjected to cold (Gaudet et al. 2003) or fungal infection (Lu et al. 2005), and in plants of a wheat-rye translocation line exposed to different biotic and abiotic stress (Jang et al. 2004, 2005).
Taken together these data strongly suggest that different nsLTPs are involved in distinct biological functions. In this context, the identification of multiple wheat nsLtp genes and the characterization of their expression during plant development could provide new insight into nsLTP function. In the present study, we report the isolation and characterization of 22 putative T. aestivum genes belonging to the nsLtp gene family. Six genes belonging to different subfamilies were further analyzed due to the display of diverse patterns of gene transcript accumulation in developing seeds (Boutrot et al. 2005). We compared the tissue-specificity and developmental regulation of these six T. aestivum non-specific lipid transfer protein gene (TaLtp) promoters in transgenic rice using the β-Glucuronidase reporter gene (uidA). β-Glucuronidase (GUS) activity was analyzed by fluorometric and histochemical assays, and by immunolocalization.
Materials and methods
Plant material
Field-grown wheat (T. aestivum L. ‘Apache’; INRA, experimental station Melgueil, 34130 Mauguio, France) leaves were used for genomic DNA extraction. Mature seeds of the japonica rice Oryza sativa L. ‘Nipponbare’ (kindly supplied by Dr. M. Yano, National Institute of Agrobiological Sciences, Tsukuba, Ibaraki, Japan) and O. sativa ‘Zhongzuo321’ (kindly supplied by Dr. Z.L. Chen, Beijing University, Beijing, China) were used for transformation. Histochemical and immunological assays were performed using O. sativa ‘Nipponbare’ and O. sativa ‘Zhongzuo321’ T0 transgenic plants, T1 seeds from progeny, and T1 seedlings. GUS activities were determined using T2 transgenic plants (O. sativa ‘Nipponbare’).
Genomic-library screening and cDNA isolation
The TaLtp9.2c gene was isolated from a genomic library of T. aestivum ‘Chinese spring’, constructed into the λFixII vector (Stratagene, La Jolla, CA, USA), kindly provided by Dr. C. Hartmann (Université Paris VII, Paris, France). Following standard protocols, this library was screened with a nsLtp gene fragment amplified from the pTd6.48 cDNA (Boutrot et al. 2005).
The cDNA clones pTaD2-2, pTa268, pTa232-5, and pTa232-6 were isolated by PCR using plasmid DNA prepared from an aliquot of a 4-day post-anthesis (dpa) T. aestivum seed cDNA library as template. PCRs were done with nsLTP specific antisense-strand primers and the universal T7 promoter as sense-strand primer as described by Boutrot et al. (2005).
Cloning of T. aestivum nsLtp gene promoters by genome walking
A genome walking procedure, the ligation mediated PCR (LM-PCR), was used to clone wheat nsLtp genes. The protocol of Siebert et al. (1995) modified by Dr. E. Bourgeois (personal communication) to allow one-step genomic DNA digestion and adaptor ligation was adapted for wheat. Briefly, 500 ng of wheat genomic DNA were digested overnight with 5 U of restriction enzyme (Promega, Madison, WI, USA) and ligated to adaptors (1.6 μM) with 5 U of T4 DNA ligase (Promega) in a total volume of 10 μl in 1× ligation buffer. Nine blunt-end restriction enzymes BalI, DraI, EcoRV, HindII, NaeI, PvuII, ScaI, SspI, and StuI were used independently. The adaptor consisted of an upper primer (5′-CACTGAATCTTGCTGACTAGGTCTGGGGAGGT-3′) and a lower primer (5′-O4P-ACCTCCCCAGAC-NH2-3′) synthesized by MWG Biotech (Courtaboeuf, France). Two rounds of PCR were then performed on digested-ligated DNAs. The primary PCR reactions contained 1 μM of the adaptor-related primer (5′-CTGAATCTTGCTGACT-3′), 0.2 μM of a nsLtp gene-specific primer, 12.5 μM of each dNTP, 1 mM MgCl2, 1 U Taq DNA polymerase (AmpliTaq Gold, Applied Biosystems, Foster City, CA, USA) and 50 ng digested-ligated DNA in a volume of 20 μl in 1× Taq DNA polymerase buffer. Touchdown PCRs were performed with an initial denaturation stage at 94°C for 3 min, 14 cycles at 94°C for 30 s, 61°C for 45 s (−0.5°C per cycle), 72°C for 2.5 min; and 20 cycles at 94°C for 30 s, 54°C for 45 s, 72°C for 2.5 min; with a final extension at 72°C for 5 min. The second PCRs were performed using 1 μl of the primary PCR product (diluted 1/50) in a reaction mixture containing 0.2 μM of the nested adaptor-related primer (5′-ATCTTGCTGACTAGGT-3′), 0.2 μM of a nested nsLtp gene-specific primer, 12.5 μM of each dNTP, 1.5 mM MgCl2, 2 U Taq DNA polymerase in a total volume of 50 μl in 1× Taq DNA polymerase buffer. PCR cycles were as described above, but with 25 additional cycles instead of 20. All nsLtp gene-specific primers (Eurogentec, Liege, Belgium) were 18–20 nucleotides long with an annealing temperature above 56°C. The secondary PCR products were analyzed on a 1.5% agarose/EtBr gel, cloned into pGEM-T Easy Vector System I (Promega) and recombinant plasmids were introduced into Escherichia coli JM109. Sequencing was performed on an ABI Prism 373 DNA sequencer (Applied Biosystems).
Wheat nsLtp genes obtained from sequence alignment of successive genome walks were confirmed by PCR using wheat genomic DNA as template and primers designed in the most distant 5′ and 3′ sequences of the amplified fragments. All the primers (Eurogentec) were 18–20 nucleotides long with an annealing temperature of above 56°C. Reactions were performed in a final volume of 25 μl containing 100 ng of genomic DNA, 1.6 μM of each primer, 100 μM of each dNTP, 1.9 U of Pfu DNA polymerase (Stratagene), 1× Pfu DNA polymerase reaction buffer and 10% (v/v) glycerol. Initial template denaturation was at 94°C for 2 min, followed by 40 cycles at 94°C for 30 s, annealing temperature for 30 s, 72°C for 2 min, with a final extension at 72°C for 7 min. The PCR products were cloned and then sequenced as described above.
Binary vector constructs and rice transformation
The promoter regions of six wheat nsLtp genes, TaLtp7.1a (−745 to +3), TaLtp7.2a (−1,437 to +3), TaLtp9.1a (−1,390 to +3), TaLtp9.2d (−1,257 to +3), TaLtp9.3e (−824 to +3), and TaLtp9.4a (−843 to +3) were PCR-amplified from genomic DNA and subcloned into the pGEM-T Easy vector (Promega). PCR primers generated an EcoRI or SalI site at their 5′ end and a PstI, SalI or HindIII site at their 3′ end (Table 1). The cloned promoter sequences were digested with EcoRI and PstI (TaLtp7.1a, TaLtp7.2a, TaLtp9.1a, and TaLtp9.2d), EcoRI and SalI (TaLtp9.3e) or SalI and HindIII (TaLtp9.4a) and ligated to the corresponding digested pCAMBIA1381Xa binary vectors (R. Jefferson, CAMBIA, Canberra, Australia). Selectable marker genes allowed hygromycin resistance in plants and kanamycin resistance in bacteria. Each of the resulting binary vectors was then introduced into Agrobacterium tumefaciens EHA105 strain (Hood et al. 1993) by electroporation. DNA digestion, ligation and electroporation were carried out following standard protocols (Sambrook et al. 1989). Agrobacterium-mediated transformation of embryogenic calli derived from mature rice seed embryos and plant regeneration were performed as described by Sallaud et al. (2003). The pCAMBIA1381Xa binary vector containing a promoter-less uidA reporter gene was used as negative control (Xa plants). Transgenic T0 rice plants harboring a CaMV35S::uidA construct (kindly provided by J. Petit, Cirad, Montpellier, France) were used as positive controls.
Molecular analysis of transgenic T0 rice plants
T0 transgenic rice plants presenting a single T-DNA insertion were identified by Southern blot analysis. Genomic DNA was extracted from 40 mg of fresh 3-week-old rice leaves using 320 μl of mixed alkyl trimethyl ammonium bromide (MATAB) buffer [100 mM Tris–HCl, pH 8.0, 1.5 M NaCl, 20 mM EDTA, 2% (w/v) MATAB, 1% (w/v) PEG 6000, 0.5% (w/v) Na2SO2] prewarmed at 72°C. The mix was incubated for 2 h at 72°C, cooled to room temperature, extracted with 360 μl of chloroform: isoamyl alcohol (24:1, v/v) and then, DNA was isopropanol precipitated. Approximately 5 μg of HindIII-digested DNA were separated on a 0.8% agarose gel and blotted onto Hybond-N+ membranes (Amersham Biosciences, Piscataway, NJ, USA). The coding sequences of uidA and hptII were radiolabeled with [α-32P]-dCTP using random primer labeling (Amersham Biosciences) and used as probes. Hybridization signals were detected by autoradiography of X-Ray films exposed to membranes at −80°C.
GUS activity
Two hundred milligrams of 4-week-old leaves from T2 rice plants were ground to powder using liquid nitrogen. Proteins were extracted with 50 mM phosphate buffer, pH 7.0, containing 10 mM EDTA, 0.1% sodium laurylsarcosine and 0.1% Triton X-100. Homogenates were cleared by centrifugation and the resultant supernatants were assayed for total protein (BCA protein assay kit, Pierce, Rockford, IL, USA) and GUS activity (Jefferson et al. 1987). The reaction product 4-methylumbelliferone was measured fluorometrically (Fluoroskan II Ascent, Thermo LabSystems, Helsinki, Finland).
Histochemical GUS assay
Histochemical experiments were performed according to Jefferson et al. (1987). For each construct, expression in the rice root system, flowers, and seeds was carried out in four independent transgenic lines. To monitor expression in leaves, depending on the number of regenerated lines 4–11 independent transgenic lines were analyzed. Rice seeds were longitudinally half-sectioned prior to immersion in X-Gluc reaction buffer and then fixed in 200 mM phosphate buffer (pH 7.0) containing 1% (v/v) acrolein, 2% (w/v) glutaraldehyde, and 50 mM caffeine for 30 min at room temperature. The other tissues were fixed in 200 mM phosphate buffer (pH 7.0) containing 2% (w/v) paraformaldehyde, 1% (w/v) glutaraldehyde, and 50 mM caffeine. Chlorophyll was excluded by soaking rice tissues for several hours in 70% ethanol. Tissue observations were performed with a Leica MZFLIII binocular microscope (Leica Microsystems, Heerbrugg, Switzerland). Roots, stems and leaves were mounted in a 4% (w/v) agar block, and transversal vibratome sections 30–40 μm thick were cut. Flowers and grains were dehydrated and embedded prior to longitudinal microtome sectioning. Dehydration was performed in a graded ethanol series (70–95%, v/v). Grains were finally dehydrated in an additional butanol bath. Flowers were impregnated in a technovit 7100 resin (Kulzer, Wehrheim, Germany)-ethanol (1:1, v/v) mixture for 72 h while a resin-butanol (1:1, v/v) mixture was used for grains. Tissues were then embedded in molds with resin and polymerized for 2 h by adding the polymerization agent to the resin. Sections 3–5 μm thick were cut with a microtome. Bright- or dark-field light microscopic observations were performed with a Leica DM RXA fluorescence microscope (Leica Microsystems).
Immunological assay
Rice seeds were harvested, longitudinally sectioned with a razor blade and immediately fixed for 8 h in 100 mM Pipes buffer, pH 7.0, containing 1.6% (w/v) paraformaldehyde, 10 mM sodium m-periodate, 375 mM lysine, and 1 mM DTT. After several rinses with 100 mM Pipes buffer containing 100 mM glycine and 1 mM DTT, the fixed samples were dehydrated in a graded ethanol series (70–95%, v/v) containing 1 mM DTT, in butanol supplemented with 10 mM DTT and in a butanol-methacrylate series (3:1, 1:1, 1:3, v/v). Tissues were impregnated four times with 100% methacrylate containing 10 mM DTT and 0.4% (v/v) benzoil ethyl ether to enable polymerization. Sections were incubated in the final bath for 12 h at 4°C. Dissolved oxygen was displaced by bubbling gaseous nitrogen through the methacrylate mixture. The samples were then transferred to molds and covered with parafilm to limit contact with oxygen. The molds were placed 28 cm under an 8 W UV light source for 15 h at 4°C. For immunodetection, methacrylate sections (4 μm) were fixed on silanized slides (Dako Cytomation, Carpinteria, CA, USA) and the resin removed with acetone. The sections were re-hydrated with a graded ethanol series from 100 to 50%, soaked in phosphate-buffered saline (PBS) for 10 min, incubated for 8 min in PBS containing 0.1% (w/v) trypsin (type XI, Sigma, St. Louis, MO, USA) and washed for 3 min with PBS. The reaction was stopped by washing with PBS containing 0.05% (w/v) trypsin inhibitor (type II-S, Sigma). The sections were washed two times (5 min each) with PBS, incubated for 1 h in PBS containing 1% (w/v) blocking agent (Roche, Basel, Switzerland), washed three times (10 min each) with PBS and, incubated for 15 h in PBS with 1% blocking agent and a 1:500 dilution of the monoclonal antibody raised against GUS (anti-β-glucuronidase rabbit IgG (H+L) fraction, Molecular Probes, Leiden, The Netherlands). The sections were washed with PBS three times (5 min each) and incubated for 90 min in PBS containing alkaline phosphatase conjugated goat anti-rabbit IgG (Sigma) at a dilution of 1:500. After being washed three times (5 min each) in PBS buffer, the sections were incubated for 15 min with NBT/BCIP (p-nitro blue tetrazolium chloride/5-bromo-4-chloro-3-indolyphosphate-toluidine salt, Chemicon International, Temecula, CA, USA) containing 5 mM levamisole. After incubation with the secondary antibody, the reaction was stopped by washing with distilled water and 95% ethanol and the sections were stained for 2 min in a ruthenium red (0.005%, w/v) solution. Bright-field light microscopic observations were performed with a Leica DM RXA fluorescence microscope (Leica Microsystems).
Sequence alignments and phylogenetic analysis
A systematic search was performed by examinating the EMBL (srs.ebi.ac.uk) and Entrez (www.ncbi.nlm.nih.gov/entrez) databases for wheat nsLtp cDNAs, genes and proteins. Proteins were analyzed for the presence of a signal peptide using SignalP v3.0 (Bendtsen et al. 2004). Deduced primary structures of mature nsLTPs were aligned using the ClustalW v1.83 program (Thompson et al. 1994). The relationship between nsLTPs was investigated with the Phylip v3.6a3 package (Felsenstein 2005) using neighbor-joining analysis. Support for nodes was estimated by the bootstrap procedure, using 1,000 re-samplings of the data. The unrooted phylogenetic tree was graphically displayed using the Treeview v1.6.6 program (Page 1996).
Statistical analysis
The hemizygous or homozygous state of T2 plants was determined by segregation analysis (Chi-square test, P < 0.05) for hygromycin resistance resulting from expression of the hptII selectable gene. For each construct, the GUS activity was measured from two independent T2 transgenic lines analyzed in three replicates. Data are presented as the mean value ± SD of the GUS activity per uidA copy (one for hemizygous or two for homozygous plants). The data were analyzed by ANOVA and means were compared for level of significance (P < 0.05) by Bonferroni’s multiple comparison test using GraphPad Prism v4.00 (GraphPad Software, San Diego, CA, USA).
Results
Identification of Triticum aestivum non-specific lipid transfer protein genes
In this study we isolated 22 T. aestivum genes encoding putative nsLTPs whose belonging to the nsLTP family was based solely on sequence homology (Table 2). The TaLtp9.2c gene was isolated by screening the T. aestivum genomic library whereas the TaLtp9.2d and TaLtp9.5b genes were amplified by PCR using specific primers designed from the TaLtp9.2c gene and the pLTP2 cDNA, respectively. All other wheat nsLtp genes were isolated using a genome walking approach, taking advantage of sequence information from cDNA clones to design specific primers. Thirty-two walks resulted in the amplification of genomic DNA fragments that included part of the coding sequence and/or promoter and led to the identification of 19 wheat nsLtp genes. Since the gene-specific primers were not as specific as expected and revealed complementarities with closely related genes, some DNA fragments belonging to different members of an nsLtp gene subfamily were amplified. In addition, because successive walks were often necessary, the contiguity of fragments resulting from sequential PCRs was confirmed to avoid the creation of chimera genes. To this end, a final PCR was carried out using wheat genomic DNA as template and primers designed in the 5′ and 3′ extremities of the identified sequences, and all the TaLtp genes were fully sequenced. The 22 TaLtp gene sequences contained full-length ORF and the length of 5′ untranslated regions ranged from 9 (TaLtp7.1c) to 2,856 bp (TaLtp9.5a). In the time course of this study we also isolated four new cDNAs from an orientated 4-dpa T. aestivum seed cDNA library whose sequences correspond to TaLtp9.3d, TaLtp9.8a, TaLtp9.7d, and TaLtp9.7e genes. All the nucleotide sequence data were deposited in the EMBL database under accession numbers AJ852536 to AJ852561.
The previously characterized wheat TaLtp9.5a gene, cDNAs encoding nsLTPs and purified proteins were also included in Table 2. Of the 23 wheat nsLtp genes reported, corresponding cDNA clones were identified for nine of them indicating that these genes were transcribed at least in developing seeds (TaLtp7.2a, TaLtp9.3d, TaLtp9.3f, TaLtp9.4a, TaLtp9.4b, TaLtp9.6a, and TaLtp9.7d) or leaves (TaLtp9.5a and TaLtp9.5b). In contrast, no corresponding genes had been identified for seven cDNA clones isolated from developing seeds (pTa4.90, pTd4.90, pTd6.48, pTd6.48, pTaD2, and pTa232) or infected leaves (pLTP3) cDNA libraries and two proteins (TaLTP9.8a and TaLTP9.8b) isolated from wheat embryos.
In the absence of chromosome assignment and given the increasing number of wheat nsLtp genes described, we recently proposed a temporary nomenclature based on the phylogenetic classification of nsLTP sequences (Boutrot et al. 2005). To establish a more rigorous nomenclature, the following criteria were added: nsLTP mature proteins whose sequence identity is over 30% constitute a type and, within a type nsLTPs whose sequence identity is over 75% constitute a protein subfamily. Based on these criteria, the TaLtp9.5a gene (Boutrot et al. 2005) was reclassified as TaLtp9.3f. To date, we have identified two subfamilies within type 2 wheat nsLTPs and eight within type 1. Subfamilies were represented by 1–7 members.
Sequence features of TaLtp genes
In order to characterize the exon/intron organization of the 23 TaLtp genes reported in Table 2, genomic DNA sequences were compared to those of cDNAs when available or deduced from sequence alignments. This analysis indicated that no type 2 TaLtp genes contain introns. This also held true for four type 1 TaLtp genes which are the two members of the TaLtp9.2 gene subfamily and the two members of the TaLtp9.5 gene subfamily. A unique intron whose size ranged from 91 to 444 bp was identified in all the other characterized type 1 TaLtp genes (Table 3). All the identified intron donor sites are located at a highly conserved position 7 bp upstream of the stop codon except the TaLtp9.1a gene whose intron donor site is positioned 4 bp upstream of the stop codon. All introns have canonical splice signals, GU in 5′ and AG in 3′ (Hebsgaard et al. 1996) and are in phase 2 with respect to the ORF.
The TaLtp9.3d gene presents the particularity of harboring a Stowaway foldback element in its intron sequence (Fig. 1). This element belongs to the family of the miniature inverted-repeat transposable elements (MITE) which are less than 400 bp long, flanked by a 2–3 bp target site (TA(A)), terminated by an inverted repeat CTCCCTCC motif and present no coding activity (Petersen and Seberg 2000). The 59 bp long element present in the TaLtp9.3d gene intron is shorter than previously described Stowaway elements (Petersen and Seberg 2000). The terminal inverted motifs are perfectly repeated within the first 6 bp and flanked by the typical TA target site at one end and its duplicated copy at the opposite extremity. The sequence alignment of TaLtp9.3 gene introns at the level of the Stowaway element shows that only the TaLtp9.3d gene harbors this transposable element. However, the five other TaLtp9.3 genes present footprints of the Stowaway element. The presence of the two nucleotides (AG) from the 3′ terminal inverted repeat of the Stowaway element and the duplicated copy of the TA recognition site support the hypothesis that a Stowaway element was excised from these intron sequences.
Sequence alignment of six TaLtp9.3 genes in the region of the Stowaway element identified in the TaLtp9.3d gene intron. The Stowaway terminal inverted motifs are underlined and the flanking TA target site and its duplicated copy are in bold. Highly conserved nucleotides are gray boxed. The nucleotide sequences are numbered with respect to the first base of the initiation codon. Accession numbers of sequences are given in Table 2
The cloned 5′ upstream region of wheat TaLtp genes ranged from 9 to 2,856 bp. Within TaLtp gene subfamilies, alignments of available promoter sequences revealed high homology in the 400 bp upstream of the translation initiation codon (data not shown). The largest sequence deletions were observed within promoters of the TaLtp7.1 subfamily with 32–69 bp deletion polymorphism at different positions. Conversely, with only 31 mutations, the TaLtp9.2c (1,269 bp) and TaLtp9.2d (1,257 bp) promoter regions displayed the highest conservation. Comparison of the other TaLtp promoter regions revealed a few single mutations or small deletions. Nevertheless, some of these modifications could be as crucial as large sequence deletions for the regulation of gene expression.
Phylogeny of TaLTPs
The alignment of the 35 wheat nsLTP sequences described in Table 2 is presented in Fig. 2. NsLTPs are synthesized as preproteins and present an N-terminal sequence (24–36 amino acids) consistent with a secretory signal peptide as defined by SignalP program analysis. Mature type 2 TaLTPs whose molecular mass ranged from 6,971 to 7,046 kDa contain 67 amino acids whereas type 1 TaLTPs contain 90–94 amino acids and have a molecular mass ranging from 8,672 to 9,609 kDa. Wheat nsLTPs displayed the features of plant nsLTPs including a pattern of eight-cysteine in a conserved position and typical amino acids between these cysteine residues.
Alignment of wheat nsLTPs. Amino acid sequences were deduced from cDNAs, genes, or indexed in the EMBL protein database. Alignment was performed using the multiple sequence alignment program ClustalW (Thompson et al. 1994). The predicted signal peptides are in italics, the conserved cysteine residues are black boxed, and for each subfamily highly conserved amino acids are gray boxed. Gaps (dashes) were introduced to optimize alignment. Amino acid residues are numbered with respect to the N-terminal of mature nsLTPs. Accession numbers of sequences are given in Table 2
A cladogram of the T. aestivum mature nsLTPs is presented in Fig. 3. The TdLTP7.1b sequence from Triticum durum was also taken into account since no identical mature protein was deduced from the T. aestivum genes characterized to date. Because TaLtp7.1d and TaLtp7.1e genes encoded the same preprotein, and TaLtp9.7b, TaLtp9.7c, and TaLtp9.7d genes the same mature protein, only the TaLTP7.1d and TaLTP9.7b sequences were taken into account in the phylogenetic analysis. The phylogenic tree is presented as an unrooted cladogram since all plant nsLTPs described so far belong to the type 1 or 2 nsLTPs (or with a less extent to the type 3) and could not be used as outgroup to root the paralogous wheat sequences. The robustness of internal branches was estimated by 1,000 bootstrap resamplings. As determined by sequence identity calculations, the phylogenetic representation confirmed that the wheat nsLTPs were distributed within two types, which correspond to the types 1 and 2 nsLTPs usually reported, and ten subfamilies, which were represented by 1–7 proteins. Comparison of all the amino acid sequences revealed at the most 23.4% identity between types 1 (TaLTP9.3e) and 2 (TaLTP7.2a) nsLTPs. As expected from the comparison of the deduced amino acid sequences, the seven TaLTP9.3 sequences (76.3–97.8% identity) appeared to be widely distributed within their subfamily whereas the sequences from other multigene subfamilies were more closely related. Only the TaLTP9.2 and TaLTP9.5 subfamilies appeared to be related in the phylogenetic tree. This subfamily dichotomy was caused by the criteria retained for the wheat nsLTP nomenclature but was also confirmed by noticeable features notably among the unexploited signal peptide sequences.
Neighbor-joining phylogenetic tree of mature wheat nsLTPs. The analysis was based on alignment of 29 amino acid sequences. The evolutionary tree was constructed by the Neighbor-Joining method and drawn by the TreeView program. For each nsLTP subfamily, the bootstrap values are shown on each branch (% of 1,000 re-sampled data set). The scale bar corresponds to 0.1 substitution per amino acid. Accession numbers of sequences are given in Table 2
Transformation and analysis of rice plants
Six wheat nsLtp genes were chosen to compare the spatial and temporal activity of their promoter in transgenic rice using the uidA reporter gene. Their selection was based on their belonging to different subfamilies and displaying diverse patterns of gene transcript accumulation in developing seeds monitored by RT-PCR (Boutrot et al. 2005). Promoter sequences of TaLtp9.1a (1,390 bp), TaLtp9.2d (1,235 bp), TaLtp9.3e (824 bp), and TaLtp9.4a (843 bp) type 1 nsLtp genes, and TaLtp7.1a (745 bp) and TaLtp7.2a (1,437 bp) type 2 nsLtp genes were amplified by PCR and cloned into the pCAMBIA 1381Xa vector resulting in nsLtp promoter::uidA gene transcriptional fusions. These constructs were introduced into O. sativa ‘Zhongzuo321’ and O. sativa ‘Nipponbare’ via Agrobacterium tumefaciens cocultivation. For each construct, 4–20 independent hygromycin-resistant lines were regenerated. Using O. sativa ‘Zhongzuo321’, we obtained nine (TaLtp7.1a::uidA), ten (TaLtp9.1a::uidA and promoterless construct), 11 (TaLtp9.4a::uidA), 15 (TaLtp7.2a::uidA and TaLtp9.3e::uidA), and 20 (TaLtp9.2d::uidA) independent lines. Using O. sativa ‘Nipponbare’, we regenerated four (TaLtp9.4a::uidA and promoterless construct), 11 (TaLtp9.3e::uidA), 13 (TaLtp9.1a::uidA and TaLtp9.2d::uidA), 15 (TaLtp7.2a::uidA), and 20 (TaLtp7.1a::uidA) independent lines. While the biological basis of variations is not understood, we observed that these variations are due to differences in transformation efficiency (18.6–80.0% depending on construct and rice cultivars) as well as differences in ability to develop secondary calli from primary calli (34.5–70.9% depending on rice cultivars). A total of 656 T0 transgenic plants was analyzed to determine the transgene copy number. Southern blot analysis of HindIII-restricted genomic DNA hybridized with the uidA or hptII probe indicated that a mean of 1.98 T-DNA copies was inserted in the genome of O. sativa ‘Zhongzuo321’ and 36.58% of plants contained a single copy of the transgene. For the O. sativa ‘Nipponbare’ these values were 2.03 and 38.14%, respectively. For further analysis, only single copy transgenic lines were propagated and grown in the greenhouse until seed maturity.
Analysis of TaLtp gene promoter activities in transgenic rice plants
In order to gain better insight into the potential role of nsLTPs, we studied the promoter activity of six wheat nsLtp genes in transgenic rice plants by monitoring the expression of the uidA reporter gene. For each TaLtp promoter, we analyzed the localization of GUS activity in all organs throughout the life cycle of the transgenic rice plants. No GUS activity was detected in the transgenic plants harboring the promoterless reporter gene (Xa) or in the untransformed rice plants (wt) (data not shown). No major variation in the uidA expression pattern was observed between plants of the O. sativa ‘Nipponbare’ and O. sativa ‘Zhongzuo321’ cultivars transformed with the same construct.
All six TaLtp gene promoters are active in rice leaves
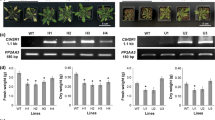
All the TaLtp promoters studied led to expression of the uidA reporter gene in the youngest (F7) leaves as illustrated for the TaLtp9.1a promoter (Fig. 4a). This was corroborated by analysis of GUS activity in the third to seventh leaf blades of 7-leaf seedlings (Fig. 4b). However, we noticed rather wide variability in GUS activity between replicates of the same experiment. Since plants segregate for the TaLtp promoter::uidA constructions, zygosity of the plants was estimated by segregation of hygromycin resistance in their progeny seedlings. Taking into account the ratio of the GUS activity to the transgene copy number rather than raw GUS activity data dramatically reduced the variability between replicates. This analysis highlighted a gene dosage effect, which was relevant for all these single T-DNA copy independent transformation events. As several studies have demonstrated additive transgene expression between homozygous and hemizygous progeny (James et al. 2002), the GUS activity was presented as the ratio of the GUS activity to the transgene copy number (Fig. 4b). While the transgenic plants harboring the 35S::uidA construct (positive control) presented stable GUS activity in the series of leaf blades (16.2 ± 1.3 pmol [4-MU (μg protein)−1 min−1] per transgene copy, data not shown), in untransformed rice plants (wt) and control transgenic plants (Xa) the level of GUS activity was insignificant. The highest and lowest levels of GUS activity were directed by the TaLtp9.1a and TaLtp9.2d promoters, respectively. There was a statistically significant decrease in GUS activity as a function of leaf rank from the youngest to the oldest leaves. This held true for all the TaLtp promoters studied except the TaLtp9.4a, which directed stable GUS activity in blades irrespective of leaf rank (P > 0.05, multiple comparison of all means). The most dramatic decrease was found in plants harboring the TaLtp9.2d::uidA fusion where no GUS activity was observed in the oldest leaves. In adult transgenic rice plants, aging but still growing leaves presented GUS activity that became restricted to the vascular bundles and finally disappeared in the oldest leaves before any external indications of senescence (data not shown).
β-Glucuronidase activity in leaf blades of T2 transgenic rice carrying different TaLtp::uidA constructs. Transgenic rice plants (O. sativa. ‘Nipponbare’) were grown in greenhouse for 4 weeks until they reached the seventh leaf stage. a Histochemical analysis of GUS activity in the F7 leaf blade of TaLtp9.1::uidA plant. b GUS activity driven by six TaLtp promoters and negative control (Xa/wt) measured in the five subsequent younger leaves (F7–F3). GUS activity is expressed as [pmol 4-MU (μg protein)−1 min−1] per transgene copy. Two independent transgenic lines were assayed for each construct and each was performed in triplicate. The values are mean ± SD (n = 6). Values significantly different from the younger leaf (F7) in the same group according to ANOVA test are indicated by asterisks (*P < 0.05; **P < 0.01; ***P < 0.001). Xa Promoterless uidA plants, wt Untransformed rice plants. Bar, 1 mm (a)
All six TaLtp gene promoters are active in rice roots
Six-day-old T1 plantlets displayed GUS activity in the vascular system of the coleoptile (data not shown) and in the root system. All the TaLtp promoters directed uidA gene expression in the root system, in which GUS activity was detected in crown roots (data not shown), lateral roots (Fig. 5a) and the distal part of the seminal root (Fig. 5c) as illustrated for the TaLtp7.1a::uidA construct. At the branching of lateral roots, transversal sections showed GUS activity in the endodermis of the seminal root but not in the central stele (Fig. 5b). In seminal roots the GUS activity was restricted to the central stele cells and mainly associated with vascular tissues (Fig. 5d). In contrast, three distinct profiles were observed in the lateral roots. The most frequent, observed in transgenic rice plants carrying the TaLtp7.1a::uidA, TaLtp7.2a::uidA, TaLtp9.1a::uidA, and TaLtp9.4a::uidA constructs, was characterized by strong GUS activity in the vascular tissues and weaker activity in the parenchyma tissues and in the root apical meristem as illustrated for the TaLtp7.1a::uidA construct (Fig. 5e). The second profile, driven by the TaLtp9.2d promoter, was similar to the latter but GUS staining was weaker and no staining was observed in the root apical meristem (Fig. 5f). Finally, the third profile was observed in TaLtp9.3e::uidA transgenic plants in which GUS staining was restricted to the vascular tissues and decreased from the proximal to the distal part of the lateral roots (Fig. 5g).
Histochemical localization of GUS activity under the control of TaLtp promoters in the root system of T1 transgenic rice plants. Six-day-old plantlets (O. sativa ‘Zhongzuo321’) were obtained by germinating T1 transgenic seeds on moist paper. a–d Profile of GUS activity in the seminal root of TaLtp7.1a::uidA transgenic rice plantlets. a Binocular microscope observation and b corresponding transversal vibratome section of the seminal root at the branching of a lateral root. c Binocular microscope observation and d corresponding transversal vibratome section of the distal region of the seminal root. e–g Binocular microscope observation in the distal region of lateral roots of e TaLtp7.1a::uidA, f TaLtp9.2d::uidA and g TaLtp9.3e::uidA plantlets. en Endodermis, lr Lateral root, ram Root apical meristem, sr Seminal root. Bars, 200 μm (a, c), 50 μm (b, d, e–g)
TaLtp gene promoters drive different GUS patterns in rice florets
While no GUS activity was observed in florets carrying the TaLtp7.1a::uidA construct (data not shown), two types of GUS activity profiles were observed for the five other TaLtp::uidA constructs (Fig. 6). The TaLtp7.2a, TaLtp9.1a, TaLtp9.2d, and TaLtp9.3e gene promoters drove similar patterns of uidA gene expression in flowers before anthesis. As illustrated for the TaLtp7.2a::uidA construct, GUS activity was restricted to the extremities of the stamen filaments (Fig. 6a) and to the vascular bundles of the palea and lemma (Fig. 6e, f). Closer examination of glumellas and stamen filaments showed that GUS staining was localized in cells in the immediate vicinity of vessel elements (Fig. 6g, h). The TaLtp9.4a promoter directed uidA gene expression in all glumella tissues and was the only TaLtp gene promoter to drive uidA gene expression in anthers (Fig. 6b). Longitudinal sections revealed that GUS activity was also present in styles and in anthers, where it was restricted to the epidermal cells (Fig. 6c, d).
Histochemical localization of GUS activity in florets of T0 transgenic rice plants harboring TaLtp::uidA fusions. Florets (O. sativa ‘Zhongzuo321’) were collected prior to anthesis in emerged panicles. a, e–h GUS activity in plants carrying the uidA reporter gene under the control of the TaLtp7.2a promoter. a Binocular microscope observation of a floret with the lemma removed. e Transverse section of floret and g detailed observation of a vessel element of a lemma. Under dark-field illumination GUS crystals appear pink and xylem vessels have autofluorescent walls. f Longitudinal section of floret. h Longitudinal section of anthers. b–d TaLtp9.4a::uidA florets. b Binocular microscope observation of open floret. c Longitudinal section of floret and d detailed observation of anthers. fi Filament, l Lemma; p Palea, s Style. Bars, 1 mm (a, b), 250 μm (c, e, f), 100 μm (d, h), 25 μm (g)
TaLtp gene promoters drive complex patterns of GUS activity and GUS localization during rice grain development
The uidA reporter gene expression driven by the wheat nsLtp promoters was studied during grain development from 4 to 30 dpa (Fig. 7). Histochemical analysis of GUS activity was supplemented by immunolocalization of the GUS protein. This analysis was carried out with plant materials fixed immediately after sampling. Depending on the nsLtp promoters, two distinct uidA gene expression profiles were observed. The first was observed in the spikelets of transgenic rice plants carrying the TaLtp7.1a::uidA, TaLtp9.2d::uidA, and TaLtp9.4a::uidA constructs as illustrated for the TaLtp7.1a::uidA plants (Fig. 7a–l). At 4 dpa, GUS staining was detected in the entire immature grain (Fig. 7a), with remarkable intensity in the developing endosperm as shown in the longitudinal section in Fig. 7e. This was confirmed by immunolocalization of GUS in TaLtp9.4a::uidA seeds (data not shown). In contrast, in TaLtp7.1a::uidA and TaLtp9.2d::uidA seeds the GUS protein was only detected in the epicarp layer and to a lesser extent in the cell layers surrounding the testa (aleurone layer, nucellar epidermis, and integument) (Fig. 7i). No immunolocalization data was collected on embryos as these were not included in the samples observed at this stage. At 10 dpa, GUS activity was no longer observed in the central endosperm but was still present in all its surrounding layers and in the embryo (Fig. 7b–f). The immunolocalization signal was restricted to the epicarp, the cell layers surrounding the testa, and the epidermal cells of the embryo coleorhiza (Fig. 7j). GUS proteins were also faintly detected in the embryo vascular system (data not shown). From 20 to 30 dpa, uidA gene expression was restricted to the embryo (Fig. 7c, d, g, h). At 20 dpa, GUS activity and immunolocalization indicated that the TaLtp7.1a, TaLtp9.2d, and TaLtp9.4a promoters drove the reporter gene expression in the embryo epidermal cells (Fig. 7g) and in the embryo vascular system (Fig. 7k). At 30 dpa, GUS activity and immunolocalization signal were almost indiscernible and only faint staining was observed in the epidermal cells located in the basal part of embryo (Fig. 7h, l).
Histochemical localization of GUS activity and GUS immunolocalization in T1 grains of transgenic rice carrying different TaLtp::uidA constructs. a–h GUS activity in grains of TaLtp7.1a::uidA plants. a–d Longitudinal half-sections of grains at 4 dpa (a), 10 dpa (b), 20 dpa (c), and 30 dpa (d). e–h Histological sections of grains under dark-field illumination at 4 dpa (e), 10 dpa (f), 20 dpa (g), and 30 dpa (h). i–l GUS immunolocalization in longitudinal sections of TaLtp7.1a::uidA grains at 4 dpa (i), 10 dpa (j), 20 dpa (k), and 30 dpa (l). m–t GUS activity in grains of TaLtp9.1a::uidA. (m–p) Longitudinal half-sections of rice grains at 4 dpa (m), 10 dpa (n), 20 dpa (o), and 30 dpa (p). q–t Histological sections of rice grains visualized under dark-field illumination at 4 dpa (q), 10 dpa (r), 20 dpa (s), and 30 dpa (t). al Aleurone, cc Cross cells, co Coleoptile, cr Coleorhiza, ep Epicarp, em Embryo, en Endosperm, in Integument, me Mesocarp, ne Nucellar epidermis, ra Radicle, sc Scutellum, tc Tube cells. Bars, 1 mm (a–d, m–p), 300 μm (s, t), 250 μm (e, g, h, l), 200 μm (j, q, r), 100 μm (i, k), 40 μm (f)
The second GUS activity profile was observed in the grains carrying the TaLtp7.2a::uidA, TaLtp9.1a::uidA, and TaLtp9.3e::uidA constructs. TaLtp promoter activity was restricted to the mid-maturation to late-maturation stages of grain development as illustrated for the TaLtp9.1a::uidA plants (Fig. 7m–t). While no GUS activity was detected at 4 dpa (Fig. 7m, q), intense GUS activity restricted to the scutellum vascular bundle and the adjacent endosperm cells appeared at 10 dpa (Fig. 7n, r). This pattern was conserved throughout grain development but GUS activity decreased and was very weak in 20 and 30-dpa-old rice grains (Fig. 7o, p, s, t).
Discussion
Complexity of the wheat nsLtp gene family
The identification of nsLtp genes and the analysis of their expression patterns during plant development are important in understanding nsLTP functions. In the present study, we isolated different members of the wheat nsLtp gene family, and showed that six of them have overlapping but distinct expression patterns in transgenic rice plants. Because we are interested in wheat seed development, we first focused our work on the identification of nsLtp genes expressed in seed. Nineteen of 22 TaLtp genes cloned in this work were isolated by chromosome walking demonstrating that this procedure was particularly productive to clone genes from multigene families and complex genomes as wheat. However, a prerequisite for this technique is to dispose of sequence data from cDNAs or genes to design primers for the LM-PCR. Another important requirement with multigene families composed of closely related members is to avoid the creation of chimera genes by carefully checking the contiguity of fragments amplified from the successive walks.
The 23 TaLtp genes reported here are 420–3,448 bp long and each contains a complete ORF encoding a putative nsLTP. Our phylogenetic analysis indicates that the deduced mature proteins are distributed within distinct clades. Based on this analysis, the proposed wheat nsLtp gene nomenclature (Boutrot et al. 2005) was refined and led to the distribution of 35 identified wheat nsLTPs in two groups representing the types 1 and 2 nsLTPs usually reported, and ten subfamilies characterized by 1–7 members. The nomenclature criteria settled on the basis of amino acid homology allows to discriminate the three nsLTPs types, which display distinct eight-cysteine motifs (data not shown). TaLTP7.1, TaLTP9.3, and TaLTP9.7 subfamilies are represented by more than three homoeologous copies, which suggests that several genes are paralogs resulting from recent gene duplication events. Evolution of wheat nsLtp genes was also revealed by the study of their structure. Type 2 nsLtp genes are intronless whereas the majority of type 1 nsLtp genes examined to date are interrupted by a single intron (Kader 1996). Clearly, within wheat type 1 nsLtp genes only those belonging to the two closely related TaLtp9.2 and TaLtp9.5 subfamilies failed to show an intron that could have been independently lost in the ancestor gene. Intron diversity was also highlighted by the TaLtp9.3d gene that carries a Stowaway miniature inverted-repeat transposable element. Sequence alignments indicated that the five other members of the TaLtp9.3 gene subfamily harbor a footprint that could have resulted from the excision of the Stowaway element. As variable footprints could be generated following excision of the Stowaway element (Petersen and Seberg 2000), no consensus sequence is indicative that a Stowaway foldback element was effectively excised from the five other members of the TaLtp9.3 gene subfamily. Absence of footprint could be evidence of the insertion/excision mechanism but to date no member of the TaLtp9.3 gene subfamily has been shown to harbor this original sequence. With more than 22,000 Stowaway MITEs identified in the rice genome (Feschotte et al. 2003) and a population size which remains to be evaluated in the wheat genome, these repetitive elements present an important disruptive mutation capability. However, the Stowaway element inserted in the TaLtp9.3d intron did not prevent gene transcription since a corresponding cDNA (pTaD2-2) was identified. Because most wheat nsLtp genes were characterized on the basis of sequence information deduced from cDNA clones isolated from developing seed cDNA libraries, it is obvious that we did not identify the full set of wheat nsLtp genes. In higher plants, nsLTPs are encoded by multigene families and 53 nsLtp genes were identified in the rice diploid genome (F. Boutrot et al. unpublished data). Taking into account the synteny between the rice and wheat genomes and the fact that several wheat nsLtp gene subfamilies (TaLtp7.1, TaLtp9.3, and TaLtp9.7) are represented by more than three homoeologous copies, we estimate that the hexaploid wheat T. aestivum contains a minimum of 100–150 nsLtp genes. The LM-PCR procedure that we adapted for walking in the wheat genome proved to be a powerful tool to identify nsLtp genes. An EST data mining strategy has been initiated in order to identify a more complete set of wheat nsLtp genes and appreciate the complexity of this family.
Expression of the wheat nsLtp genes
The complexity of plant nsLtp gene families and their large spatio-temporal expression patterns raised the question whether the expression of each member is distinctly regulated, and if the corresponding proteins support distinct physiological functions. In wheat, in a preliminary study we showed that nsLtp genes displayed a complex pattern of expression in developing seeds (Boutrot et al. 2005). However, the identification of new wheat nsLtp genes that share high-sequence similarities means that RT-PCR analysis is not really suitable for the evaluation of gene expression. We therefore used a reporter gene strategy and the rice lifecycle to monitor GUS activity driven by the promoter of six TaLtp genes belonging to different subfamilies.
Plant nsLTPs were first isolated from spinach leaves (Kader et al. 1984) and then purified from leaves of different species as broccoli (Pyee et al. 1994). There are also many reports of nsLtp gene expression in leaves of monocots and dicots (Molina and García-Olmedo 1993; Clark and Bohnert 1999). All the six wheat nsLtp gene promoters we studied were active in leaves. However, statistically significant differences in GUS activity were observed in leaves of transgenic lines where the uidA gene is under the control of the TaLtp7.1a, TaLtp7.2a, TaLtp9.1a, TaLtp9.3e, and TaLtp9.2d gene promoters. Our results indicate that the highest GUS activity is observed in the youngest leaf and declines as the leaf ages. Likewise, it was previously reported that the transcript levels of the tobacco Ltp1 gene (Fleming et al. 1992) and broccoli wax9 gene (Pyee et al. 1994) were higher in the youngest leaves. This pattern was correlated with a decrease in nsLTP found in the wax surface (Pyee et al. 1994) and is consistent with an involvement of nsLTP in the secretion of extracellular lipophillic material, including cutin monomers (Sterk et al. 1991). The variability of promoter activities between leaves was also reported for the Arabidopsis AtLtp1 (van Leeuwen et al. 2001) and rice Ltp1 (Guiderdoni et al. 2002) genes. Finally, as described by Guiderdoni et al. (2002), we observed that the decline in GUS staining was progressively limited to the vascular bundles and, then disappeared in the oldest leaves. In immature leaf sheaths of 4-week-old wheat seedlings, TaLtp3 (TaLtp9.4c in our nomenclature) transcripts were only detected in phloem vessels (Jang et al. 2005). This gene is likely a homoeologous copy of the TaLtp9.4a gene considered in this study. Nevertheless, the two genes present differential transcriptional regulation, since the GUS activity driven by the TaLtp9.4a gene promoter was not restricted to immature leaf sheaths. Many studies demonstrated that the expression of nsLtp genes in leaves was mainly associated with epidermal cells and suggested that nsLTPs were involved in the formation of the cuticle (Fleming et al. 1992; Sohal et al. 1999). Nevertheless, we report wheat nsLtp gene promoter activity mainly associated with leaf vascular tissues, suggesting that the physiological function of these wheat nsLTPs in leaves is not be related to cuticle formation, and that their function remains to be elucidated.
All the six wheat nsLtp gene promoters we studied were active in roots while expression of nsLtp genes in root tissues was more rarely reported than in leaves. For example the Arabidopsis Ltp1 gene (Thoma et al. 1994), the tobacco Ltp1 gene (Canevascini et al. 1996), the rape BnLTP gene (Sohal et al. 1999), and the rice Ltp1 gene (Guiderdoni et al. 2002) present a broad pattern of expression that includes roots. Whereas, the tobacco Ltp1 gene promoter drove uidA expression in root hair epidermal cells, the three other gene promoters resulted in GUS staining localized at the emission sites of lateral roots similar to that observed in this study. Since T1 seeds were germinated on a moist paper, the GUS activity in rice seedlings was not related to wound induction. The presence of nsLTPs was also identified in xylem sap (Buhtz et al. 2004), however the GUS activity reported in this study would not support efficient nsLTP translocation in the xylem sap since reporter gene expression under the control of wheat nsLtp promoters was not associated with the seminal root vascular system. On the contrary, the localization of the GUS activity in regions of developing vascular tissues supports the hypothesis of wheat nsLTP involvement in the process of vascular differentiation as reported for a Zinnia nsLTP (Ye and Varner 1994). During this physiological process, nsLTPs might thus be discharged into the xylem sap and travel upward with the flow of sap.
In many plant species, the presence of nsLTPs and nsLtp gene expression were reported in stem tissues, mainly associated with xylem (Eklund and Edqvist 2003) and phloem vascular tissues (Pyee et al. 1994; Horvath et al. 2002) or localized either in epidermal cells (Sossountzov et al. 1991) or subepidermal cells (Clark and Bohnert 1999). In contrast no GUS activity was detected in stems of seedlings or mature transgenic rice plants carrying the TaLtp::uidA constructs. All these findings suggest that if wheat nsLTPs do have a biological function in vascular tissues of leaves and roots as hypothesized above, this function cannot be generalized to the vascular tissues of all vegetative organs.
In rice flowers, three different expression profiles were observed depending on the wheat nsLtp promoter concerned. Under the control of the TaLtp7.1a promoter no uidA gene expression was observed in immature and mature flowers. The GUS activity driven by the TaLtp7.2a, TaLtp9.1a, TaLtp9.2d, and TaLtp9.3e gene promoters was associated with vascular tissues in glumellas and the extremities of anther filaments. As we observed in seedling roots, nsLtp gene expression was not present in all vascular elements since anther filaments and some receptacle autofluorescent xylem vessels were not GUS stained. This supports our hypothesis that in flowers, these wheat nsLTPs may have a physiological function related to particular vascular elements. Unlike in roots, these tissues are not subject to vascular differentiation, so TaLTP7.2a, TaLTP9.1a, TaLTP9.2d, and TaLTP9.3e could be involved either in different physiological processes in roots and flowers or in only one process which remains to be determined. Since GUS staining also appeared in phloem sieve tubes of glumellas, these four nsLTPs could have a function related to these vascular elements. The GUS activity in phloem sieve tubes and companion cells is consistent with reports of the immunodetection of broccoli WAX9 proteins in cell walls of vascular parenchyma cells and sieve elements (Pyee et al. 1994), the highly specific expression of the potato StnsLTP.2 gene promoter in phloem vessels (Horvath et al. 2002) and the RT-PCR detection of Arabidopsis dir1 gene transcripts in phloem companion cells (Ivashikina et al. 2003). However, the predicted apoplastic subcellular localization of plant nsLTPs is inconsistent with their presence in the phloem sieve; thus, like the DIR1 protein, nsLTPs may release a systemic signal responsible for SAR in the vascular system (Maldonado et al. 2002).
Finally, the TaLtp9.4a gene promoter drove a unique GUS activity profile in flowers and is the first nsLtp gene whose broad spatial expression includes the anther epidermal cells. While this profile does not appear to be consistent with a role played by TaLTP9.4a in vascular tissues, GUS activity was not observed in the remaining tapetal cell layer indicating that the protein does not promote pollen development. With GUS staining also present in all glumella cells, the TaLTP9.4a can be assumed to be present in all the flower tissues that protect the pollen. In this way, like several defensins found in flowers displaying antifungal activity (Lay et al. 2003), TaLTP9.4a may support a physiological function by protecting reproductive tissues against potential attack by pathogens.
The six nsLtp gene promoters studied were active in transgenic rice grains and drove two distinct expression profiles. The first profile was observed under the control of the TaLtp7.1a, TaLtp9.2d, and TaLtp9.4a gene promoters and was characterized by GUS activity in all immature grain tissues. Several studies reported induction of nsLtp gene expression following wounding (Guiderdoni et al. 2002; Cameron et al. 2006a). As the transgenic rice grains were half-sectioned prior to GUS histochemical analysis, we performed concomitant immunodetection of GUS proteins which was carried out with plant materials fixed immediately after sampling to determine if the uidA gene expression was or was not wound-induced following longitudinal sectioning of the grain. These combined analyses revealed that at 4 dpa the TaLtp9.4a::uidA construct was expressed in the rice grain. In contrast TaLtp7.1a and TaLtp9.2d genes were restricted to the grain epicarp and the cell layers surrounding the testa while expression was wound-inducible in all other grain tissues. At 10 dpa, the three promoters displayed a similar expression pattern to the GUS protein detected in embryo peripheral cells and vascular tissues, and uidA expression was found to be wound-inducible in grain tissues including embryo, but no longer in endosperm. In subsequent developmental stages, GUS activity and GUS immunolocalization were restricted to the embryo and displayed similar patterns.
Several nsLtp genes were preferentially expressed in the outer epidermal layer of fleshy fruits and the corresponding nsLTPs were thought to be involved in fruit cuticle synthesis (Botton et al. 2002; Liu et al. 2006). Our study reports for the first time nsLtp gene expression in the epicarp of Poaceae caryopsis, and supports the implication of the corresponding TaLTP7.1a, TaLTP9.2d, and TaLTP9.4a proteins in cuticle synthesis. At 2 and 10 dpa, GUS proteins were also detected in the nucellar epidermis and inner integuments. During grain maturation these cells layers will be compressed to form the testa (seed coat). Cutin layers are found in both sides of the testa, which mechanically protect the developing endosperm from water penetration and from microbial attack (for review see Moise et al. 2005). The presence of wheat nsLTPs coincides yet again with the formation of a cuticular layer and strongly supports the hypothesis of nsLTP involvement in cuticle formation. Nevertheless, since induction of TaLtp gene expression following wounding was observed in inner grain tissues, it is tempting to speculate that the three wheat nsLTPs could also be involved in plant defense mechanisms. This presumed physiological function would be limited to immature grains since GUS activity subsequently disappeared from pericarp cell layers and uidA gene expression was no longer wound-inducible in the grain after 10 dpa. Many nsLTPs display intrinsic antimicrobial activity and their involvement in plant defense mechanisms has been widely illustrated (Cammue et al. 1995). In the wheat grain epicarp, TaLTP7.1a, TaLTP9.2d, and TaLTP9.4a are presumed to be constitutively accumulated to become part of antimicrobial and entomotoxic proteins. Such proteins were identified in the epicarp cell layer (Li et al. 2005) and in the testa of different plants (for review see Moise et al. 2005). In the other grain tissues, expression of the corresponding TaLtp7.1a, TaLtp9.2d, and TaLtp9.4a genes is thought to be induced following pathogen development to confer better resistance to biotic stresses.
During grain maturation, GUS activity became restricted to the embryo epidermal cells and embryo vascular tissues, and, in the end, almost no GUS activity was detected in mature transgenic rice grains. Like the maize LTP2 gene (Sossountzov et al. 1991) and the carrot EP2 gene (Sterk et al. 1991), transcripts from several nsLtp genes displayed a similar developmental time course in epidermal cells of developing embryos. These genes are expressed at the early proembryo-stage and confined to the outer cell layer of the proembryo (Sterk et al. 1991; Bommert and Werr 2001). As LTP2 and EP2 were thought to be involved in the secretion or in the deposition of extracellular lipophilic molecules on the protoderm layer, TaLTP7.1a, TaLTP9.2d, and TaLTP9.4a could be involved in transport of lipids to the outer surface. Since the TaLtp7.1a, TaLtp9.2d, and TaLtp9.4a gene promoters drove GUS activity in all rice grain tissues covered by a cuticular layer, our results strongly support the hypothesis of nsLTP involvement in cuticle synthesis. This profile was also illustrated by GUS activity and immunolocalization in embryo vascular tissues. In the scutellum vascular system, the developmental regulation of the uidA reporter gene expression was similar whatever the TaLtp gene promoter. Since the embryo provascular tissue develops from 5 dpa and the vascular bundle system is completed around 10 dpa (Hoshikawa 1993), the appearance of GUS activity between 4 and 10 dpa coincides with the formation of vessel elements. In this way, our results suggest that in grain these six wheat nsLTPs could support a common physiological function related to vessel formation. TaLTP7.2a, TaLTP9.1a, and TaLTP9.3e could be specialized in this function, whereas TaLTP7.1a, TaLTP9.2d, and TaLTP9.4a could be also involved in cuticle formation.
In conclusion, the six wheat nsLtp gene promoters studied directed GUS activity mainly in vascular tissues of leaf, root, flower, and embryo, as well as in grain tissues protected by a cuticle layer. Our study clearly shows that there is no specific expression pattern of type 1 nsLtp genes versus type 2 nsLtp genes. Our observations also revealed that the activity of these promoters presented both overlapping and distinct patterns of transcriptional regulation. The functional significance of the multiple patterns of expression was not elucidated; however, our results suggest that the six wheat nsLTPs could play a role in vascular tissues. Several GUS profiles suggested that TaLTPs could have multiple physiological functions. In this way, TaLTP7.1a, TaLTP9.2d, and TaLTP9.4a could also be involved in cuticle synthesis in grain tissues, while TaLTP9.4a could play a role in anthers. Strong correlative evidence supports the involvement of wheat nsLTPs in plant defense mechanisms. Further experiments are pending to determine the activity of the six wheat nsLtp gene promoters after wounding and microbial infection.
Abbreviations
- dpa:
-
Day post-anthesis
- EST:
-
Expressed sequence tag
- GUS:
-
β-Glucuronidase
- LM-PCR:
-
Ligation mediated PCR
- MATAB:
-
Mixed alkyl trimethyl ammonium bromide
- nsLTP:
-
Non-specific lipid transfer protein
- nsLtp :
-
Non-specific lipid transfer protein gene
- SAR:
-
Systemic acquired resistance
- TaLtp :
-
Triticum aestivum non-specific lipid transfer protein gene
- uidA :
-
β-Glucuronidase gene
References
Altenbach SB, Kothari KM (2004) Transcript profiles of genes expressed in endosperm tissue are altered by high temperature during wheat grain development. J Cereal Sci 40:115–126
Arondel V, Vergnolle C, Cantrel C, Kader J-C (2000) Lipid transfer proteins are encoded by a small multigene family in Arabidopsis thaliana. Plant Sci 157:1–12
Bendtsen JD, Nielsen H, von Heijne G, Brunak S (2004) Improved prediction of signal peptides: SignalP 3.0. J Mol Biol 340:783–795
Bommert P, Werr W (2001) The expression pattern of lipid transfer protein 2 (LTP2) gene indicates regionalisation in the proembryo and confirms the coleoptile to be in lineage with the scutellum. Maize News Lett 75:35–36
Botton A, Begheldo M, Rasori A, Bonghi C, Tonutti P (2002) Differential expression of two lipid transfer protein genes in reproductive organs of peach (Prunus persica L. Batsch). Plant Sci 163:993–1000
Boutrot F, Guirao A, Alary R, Joudrier P, Gautier M-F (2005) Wheat non-specific lipid transfer protein genes display a complex pattern of expression in developing seeds. Biochim Biophys Acta; Gene Struct Exp 1730:114–125
Buhot N, Gomés E, Milat M-L, Ponchet M, Marion D, Lequeu J, Delrot S, Coutos-Thévenot P, Blein J-P (2004) Modulation of the biological activity of a tobacco LTP1 by lipid complexation. Mol Biol Cell 15:5047–5052
Buhtz A, Kolasa A, Arlt K, Walz C, Kehr J (2004) Xylem sap protein composition is conserved among different plant species. Planta 219:610–618
Cameron KD, Moskal WA, Smart LB (2006a) A second member of the Nicotiana glauca lipid transfer protein gene family, NgLTP2, encodes a divergent and differentially expressed protein. Funct Plant Biol 33:141–152
Cameron KD, Teece MA, Smart LB (2006b) Increased accumulation of cuticular wax and expression of lipid transfer protein in response to periodic drying events in leaves of tree tobacco. Plant Physiol 140:176–183
Cammue BPA, Thevissen K, Hendriks M, Eggermont K, Goderis IJ, Proost P, Van Damme J, Osborn RW, Guerbette F, Kader J-C, Broekaert WF (1995) A potent antimicrobial protein from onion seeds showing sequence homology to plant lipid transfer proteins. Plant Physiol 109:445–455
Canevascini S, Caderas D, Mandel T, Fleming AJ, Dupuis I, Kuhlemeier C (1996) Tissue-specific expression and promoter analysis of the tobacco ltp1 gene. Plant Physiol 112:513–524
Clark AM, Bohnert HJ (1999) Cell-specific expression of genes of the lipid transfer protein family from Arabidopsis thaliana. Plant Cell Physiol 40:69–76
Coutos-Thévenot P, Jouenne T, Maes O, Guerbette F, Grosbois M, Le Caer J-P, Boulay M, Deloire A, Kader J-C, Guern J (1993) Four 9-kDa proteins excreted by somatic embryos of grapevine are isoforms of lipid-transfer proteins. Eur J Biochem 217:885–889
Dieryck W, Gautier M-F, Lullien V, Joudrier P (1992) Nucleotide sequence of a cDNA encoding a lipid transfer protein from wheat (Triticum durum Desf.). Plant Mol Biol 19:707–709
Douliez J-P, Jégou S, Pato C, Larré C, Mollé D, Marion D (2001) Identification of a new form of lipid transfer protein (LTP1) in wheat seeds. J Agric Food Chem 49:1805–1808
Dubreil L, Gaborit T, Bouchet B, Gallant DJ, Broekaert WF, Quillien L, Marion D (1998) Spatial and temporal distribution of the major isoforms of puroindolines (puroindoline-a and puroindoline-b) and nonspecific lipid transfer protein (ns-LTP1e1) of Triticum aestivum seeds. Relationships with their in vitro antifungal properties. Plant Sci 138:121–135
Eklund DM, Edqvist J (2003) Localization of nonspecific lipid transfer proteins correlate with programmed cell death responses during endosperm degradation in Euphorbia lagascae seedlings. Plant Physiol 132:1249–1259
Felsenstein J (2005) PHYLIP (Phylogeny Inference Package) version 3.6a3. Distributed by the author. Department of Genome Sciences, University of Washington, Seattle, WA
Feschotte C, Swamy L, Wessler SR (2003) Genome-wide analysis of mariner-like transposable elements in rice reveals complex relationships with Stowaway miniature inverted repeat transposable elements (MITEs). Genetics 163:747–758
Fleming AJ, Mandel T, Hofmann S, Sterk P, de Vries SC, Kuhlemeier C (1992) Expression pattern of a tobacco lipid transfer protein gene within the shoot apex. Plant J 2:855–862
Foster GD, Robinson SW, Blundell RP, Roberts MR, Hodge R, Draper J, Scott RJ (1992) A Brassica napus mRNA encoding a protein homologous to phospholipid transfer proteins, is expressed specifically in the tapetum and developing microspores. Plant Sci 84:187–192
Gaudet DA, Laroche A, Frick M, Huel R, Puchalski B (2003) Cold induced expression of plant defensin and lipid transfer protein transcripts in winter wheat. Physiol Plant 117:195–205
Guiderdoni E, Cordero MJ, Vignols F, García-Garrido JM, Lescot M, Tharreau D, Meynard D, Ferrière N, Notteghem J-L, Delseny M (2002) Inducibility by pathogen attack and developmental regulation of the rice Ltp1 gene. Plant Mol Biol 49:679–695
Hebsgaard SM, Korning PG, Tolstrup N, Engelbrecht J, Rouze P, Brunak S (1996) Splice site prediction in Arabidopsis thaliana pre-mRNA by combining local and global sequence information. Nucleic Acids Res 24:3439–3452
Hood EE, Gelvin SB, Melchers S, Hoekema A (1993) New Agrobacterium helper plasmids for gene transfer to plant. Transgenic Res 2:208–218
Horvath BM, Bachem CW, Trindade LM, Oortwijn MEP, Visser RGF (2002) Expression analysis of a family of nsLTP genes tissue specifically expressed throughout the plant and during potato tuber life cycle. Plant Physiol 129:1494–1506
Hoshikawa K (1993) Anthesis, fertilization and development of caryopsis. In: Matsuo T, Hosikawa K (eds) Science of the rice plant. 1. Morphology. Food and Agriculture Policy Research Center, Tokyo, pp 339–376
Ivashikina N, Deeken R, Ache P, Kranz E, Pommerrenig B, Sauer N, Hedrich R (2003) Isolation of AtSUC2 promoter-GFP-marked companion cells for patch-clamp studies and expression profiling. Plant J 36:931–945
James VA, Avart C, Worland B, Snape JW, Vain P (2002) The relationship between homozygous and hemizygous transgene expression levels over generations in populations of transgenic rice plants. Theor Appl Genet 104:553–561
Jang CS, Lee HJ, Chang SJ, Seo YW (2004) Expression and promoter analysis of the TaLTP1 gene induced by drought and salt stress in wheat (Triticum aestivum L.). Plant Sci 167:995–1001
Jang CS, Johnson JW, Seo YW (2005) Differential expression of TaLTP3 and TaCOMT1 induced by Hessian fly larval infestation in a wheat line possessing H21 resistance gene. Plant Sci 168:1319–1326
Jefferson RA, Kavanagh TA, Bevan MW (1987) GUS fusions: β-glucuronidase as a sensitive and versatile gene fusion marker in higher plants. EMBO J 6:3901–3907
José-Estanyol M, Gomis-Rüth FX, Puigdomènech P (2004) The eight-cysteine motif, a versatile structure in plant proteins. Plant Physiol Biochem 42:355–365
Jung HW, Kim W, Hwang BK (2003) Three pathogen-inducible genes encoding lipid transfer protein from pepper are differentially activated by pathogens, abiotic, and environmental stresses. Plant Cell Environ 26:915–928
Kader J-C (1996) Lipid-transfer proteins in plants. Annu Rev Plant Physiol Plant Mol Biol 47:627–654
Kader J-C, Julienne M, Vergnolle C (1984) Purification and characterization of a spinach-leaf protein capable of transferring phospholipids from liposomes to mitochondria or chloroplasts. Eur J Biochem 139:411–416
Lauga B, Charbonnel-Campaa L, Combes D (2000) Characterization of MZm3-3, a Zea mays tapetum-specific transcript. Plant Sci 157:65–75
Lay FT, Brugliera F, Anderson MA (2003) Isolation and properties of floral defensins from ornamental tobacco and petunia. Plant Physiol 131:1283–1293
Li YC, Yang YC, Hsu JSF, Wu DJ, Wu HH, Tzen JTC (2005) Cloning and immunolocalization of an antifungal chitinase in jelly fig (Ficus awkeotsang) achenes. Phytochemistry 66:879–886
Liu K, Jiang H, Moore S, Watkins C, Jahn M (2006) Isolation and characterization of a lipid transfer protein expressed in ripening fruit of Capsicum chinense. Planta 223:672–683
Lu ZX, Gaudet DA, Frick M, Puchalski B, Genswein B, Laroche A (2005) Identification and characterization of genes differentially expressed in the resistance reaction in wheat infected with Tilletia tritici, the common bunt pathogen. J Biochem Mol Biol 38:420–431
Maldonado AM, Doerner P, Dixon RA, Lamb CJ, Cameron RK (2002) A putative lipid transfer protein involved in systemic resistance signalling in Arabidopsis. Nature 419:399–403
Marion D, Dubreil L, Douliez J-P (2003) Functionality of lipids and lipid-protein interactions in cereal-derived food products. Ol Corps Gras Li 10:47–56
Moise JA, Han S, Gudynaite-Savitch L, Johnson DA, Miki BLA (2005) Seed coats: structure, development, composition, and biotechnology. In Vitro Cell Dev Biol; Plant 41:620–644
Molina A, García-Olmedo F (1993) Developmental and pathogen-induced expression of three barley genes encoding lipid transfer proteins. Plant J 4:983–991
Monnet F-P (1990) Caractérisation d’une protéine de fixation de lipides du blé dur, purification, séquençage, ADN complémentaire: relations aux protéines végétales de transfert de lipides et aux inhibiteurs d’amylase/trypsine des céréales. PhD Dissertation, Université de Montpellier II, France
Monnet F-P, Dieryck W, Boutrot F, Joudrier P, Gautier M-F (2001) Purification, characterisation and cDNA cloning of a type 2 (7 kDa) lipid transfer protein from Triticum durum. Plant Sci 161:747–755
Neumann GM, Condron R, Thomas I, Polya GM (1994) Purification and sequencing of a family of wheat lipid transfer protein homologues phosphorylated by plant calcium-dependent protein kinase. Biochim Biophys Acta; Prot Struct Mol Enzymol 1209:183–190
Page RDM (1996) TREEVIEW: an application to display phylogenetic trees on personal computers. Comput Appl Biosci 12:357–358
Petersen G, Seberg O (2000) Phylogenetic evidence for excision of Stowaway miniature inverted-repeat transposable elements in triticeae (Poaceae). Mol Biol Evol 17:1589–1596
Pyee J, Yu H, Kolattukudy PE (1994) Identification of a lipid transfer protein as the major protein in the surface wax of broccoli (Brassica oleracea) leaves. Arch Biochem Biophys 311:460–468
Sallaud C, Meynard D, Van Boxtel J, Gay C, Bes M, Brizard JP, Larmande P, Ortega D, Raynal M, Portefaix M, Ouwerkerk PB, Rueb S, Delseny M, Guiderdoni E (2003) Highly efficient production and characterization of T-DNA plants for rice (Oryza sativa L.) functional genomics. Theor Appl Genet 106:1396–408
Sambrook J, Fritsch EF, Maniatis T (1989) Molecular cloning: a laboratory manual, 2nd edn. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY
Siebert PD, Chenchik A, Kellogg DE, Lukyanov KA, Lukyanov SA (1995) An improved PCR method for walking in uncloned genomic DNA. Nucleic Acids Res 23:1087–1088
Sohal AK, Pallas JA, Jenkins GI (1999) The promoter of a Brassica napus lipid transfer protein gene is active in a range of tissues and stimulated by light and viral infection in transgenic Arabidopsis. Plant Mol Biol 41:75–87
Sossountzov L, Ruiz-Avila L, Vignols F, Jolliot A, Arondel V, Tchang F, Grosbois M, Guerbette F, Miginiac E, Delseny M, Puigdomènech P, Kader J-C (1991) Spatial and temporal expression of a maize lipid transfer protein gene. Plant Cell 3:923–933
Sterk P, Booij H, Schellekens GA, van Kammen A, de Vries SC (1991) Cell-specific expression of the carrot EP2 lipid transfer protein gene. Plant Cell 3:907–921
Sy D, Le Gravier Y, Goodfellow J, Vovelle F (2003) Protein stability and plasticity of the hydrophobic cavity in wheat ns-LTP. J Biomol Struct Dyn 21:15–30
Thoma S, Hecht U, Kippers A, Botella J, de Vries SC, Somerville C (1994) Tissue-specific expression of a gene encoding a cell wall-localized lipid transfer protein from Arabidopsis. Plant Physiol 105:35–45
Thompson JD, Higgins DG, Gibson TJ (1994) CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22:4673–4680
van Bel AJE, Gaupels F (2004) Pathogen-induced resistance and alarm signals in the phloem. Mol Plant Pathol 5:495–504
van Leeuwen W, Ruttink T, Borst-Vrenssen AW, van der Plas LH, van der Krol AR (2001) Characterization of position-induced spatial and temporal regulation of transgene promoter activity in plants. J Exp Bot 52:949–959
Vrinten PL, Nakamura T, Kasha KJ (1999) Characterization of cDNAs expressed in the early stages of microspore embryogenesis in barley (Hordeum vulgare) L. Plant Mol Biol 41:455–463
Ye ZH, Varner JE (1994) Expression of an auxin- and cytokinin-regulated gene in cambial region in Zinnia. Proc Natl Acad Sci USA 91:6539–6543
Acknowledgments
The authors wish to thank Caroline Hartman for the wheat genomic library, Julie Petit for the CaMV35S::uidA construct, Jacques Escoute and Geneviève Conejero for helpful advice on histological analysis. We would also like to thank Emmanuelle Bourgeois for help in adapting the LM-PCR protocol to wheat. Freddy Boutrot was the recipient of a fellowship from the French Ministère de l’Education Nationale, de l’Enseignement Supérieur et de la Recherche. The support of the Génopole LR for containment greenhouse infrastructures is also acknowledged.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Boutrot, F., Meynard, D., Guiderdoni, E. et al. The Triticum aestivum non-specific lipid transfer protein (TaLtp) gene family: comparative promoter activity of six TaLtp genes in transgenic rice. Planta 225, 843–862 (2007). https://doi.org/10.1007/s00425-006-0397-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00425-006-0397-7