Abstract
Stromal interaction molecule 1 (STIM1) mediates Ca2+ movements from the extracellular space to the cytosol through a store-operated Ca2+ entry (SOCE) mechanism in various cells including skeletal muscle cells. In the present study, to reveal the unidentified functional role of the STIM1 C terminus from 449 to 671 amino acids in skeletal muscle, binding assays and quadrupole time-of-flight mass spectrometry were used to identify proteins binding in this region along with proteins that mediate skeletal muscle contraction and relaxation. STIM1 binds to sarcoplasmic/endoplasmic reticulum Ca2+-ATPase 1a (SERCA1a) via this region (called STIM1-SBR). The binding was confirmed in endogenous full-length STIM1 in rabbit skeletal muscle and mouse primary skeletal myotubes via co-immunoprecipitation assay and immunocytochemistry. STIM1 knockdown in mouse primary skeletal myotubes decreased Ca2+ uptake from the cytosol to the sarcoplasmic reticulum (SR) through SERCA1a only at micromolar cytosolic Ca2+ concentrations, suggesting that STIM1 could be required for the full activity of SERCA1a possibly during the relaxation of skeletal muscle. Various Ca2+ imaging experiments using myotubes expressing STIM1-SBR suggest that STIM1 is involved in intracellular Ca2+ distributions between the SR and the cytosol via regulating SERCA1a activity without affecting SOCE. Therefore, in skeletal muscle, STIM1 could play an important role in regulating Ca2+ movements between the SR and the cytosol.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The main processes that must occur in skeletal muscle are muscle contraction and relaxation, and Ca2+ from the sarcoplasmic reticulum (SR) is the main source of Ca2+ for these processes [26, 70]. Spatial and temporal distribution of intracellular Ca2+ between the SR and the cytosol is the key factor for the cycling of skeletal muscle contraction and relaxation [26, 70]. Ryanodine receptor 1 (RyR1) and sarcoplasmic/endoplasmic reticulum Ca2+-ATPase 1a (SERCA1a) are the main proteins responsible for the distribution of Ca2+, as follows [26, 27, 50]. During the excitation of skeletal myotubes, RyR1 (an internal Ca2+-releasing channel in the SR membrane) is activated by dihydropyridine receptor (DHPR, a membrane voltage-sensing protein in the transverse (t)-tubule membrane) via physical interaction, which allows Ca2+ releases from the SR to the cytosol through RyR1, and, ultimately, evokes skeletal muscle contraction (excitation–contraction coupling [ECC]). Sub-micromolar Ca2+ acts as an agonist of RyR1 by triggering Ca2+-induced Ca2+ release (CICR) through RyR1, which maximizes skeletal ECC [26, 27, 50]. SERCA1a, the main isoform of adult skeletal muscle (more than 70 % in all types of skeletal muscle fibers), is a Ca2+ pump in the SR membrane and uptakes Ca2+ from the cytosol into the SR to refill the SR with Ca2+ during skeletal muscle relaxation [4, 5].
Store-operated Ca2+ entry (SOCE) is a ubiquitous inside-out signal, and stromal interaction molecule 1 (STIM1) and Orai1 are the main proteins responsible for the SOCE, as follows [24, 54, 55, 59]. STIM1 is a Ca2+ sensor in SR/ER membranes, and Orai1 is a Ca2+ entry channel in t-tubule/plasma membranes. In general, SR/ER depletion induces the dissociation of Ca2+ from STIM1, which allows STIM1 to relocate in the ER membranes near plasma membranes and to interact with Orai1 (called the formation of puncta) followed by extracellular Ca2+ entry through Orai1 [34, 52, 62–64]. In skeletal muscle, the formation of puncta occurs naturally as a part of differentiation that is independent of the SR depletion, which means that SOCE in skeletal myotubes is more rapid than in other types of cells, occurring in a matter of seconds [14, 25, 59].
STIM1 has a single-transmembrane domain, a short intraluminal N terminus that contains a Ca2+-sensing EF hand and a sterile α-motif (SAM) domain [53, 68], and a cytoplasmic C terminus that contains various other domains [35, 53, 58]. The D76, D84, and E87 residues in the EF hand are critical for SR/ER Ca2+-sensing [17, 30, 33, 51]. The EF-SAM domain is responsible for the self-oligomerization and relocalization of STIM1s to form puncta [52, 69]. Either the SOAR (STIM1-Orai1-activating region [66]) or the CAD (Ca2+ release-activated Ca2+ (CRAC)-activating domain [39]) in the C terminus physically activates Orai1. The CAD also participates in the self-oligomerization of STIM1 [10, 37]. The first coiled-coil domain participates in the oligomerization of STIM1 at rest only [10]. The lys-rich domain is responsible for the Orai1-independent plasma membrane targeting of STIM1 [39]. The roles of the other STIM1 domains, or regions, in particular the rear C terminus of STIM1 after CAD/SOAR, either have not been studied or the roles remain a matter of controversy.
STIM1-deficient mice die from perinatal myopathy [32, 54]. Silencing STIM1 in human skeletal myoblasts hampers myotube formation and reduces SOCE [11]. Patients with loss-of-function mutations of STIM1 show congenital myopathies as well as severe combined immunodeficiency due to the lack of SOCE [15, 16, 41]. Muscle fibers from mdx mice, an animal model for Duchenne muscular dystrophy that is characterized by progressive muscle weakness, show increases in STIM1 expression and SOCE [3, 13]. Despite the relevance of STIM1 to skeletal muscle diseases, the direct involvement of STIM1 to Ca2+ movements during skeletal muscle contraction and relaxation is not well understood. Apart from this, the function of the rear C-terminal region of STIM1, which includes 449 to 671 amino acids (called the STIM1-UI region in the present study), has not been identified even in other types of cells. In addition, the STIM1-UI region is a variable region between STIM1 and STIM2 [19]. Therefore, in the present study, proteins binding to the STIM1-UI region, which are among the proteins that mediate skeletal muscle contraction and relaxation, were searched using biochemical and cellular approaches. We found that the STIM1-UI region directly binds to SERCA1a, and we named it the SERCA1a-binding region (STIM1-SBR). The role of STIM1-SBR in skeletal muscle contraction and relaxation was examined by functional approaches using mouse primary skeletal myotubes.
Methods
Cloning and expression of STIM1-UI
Using human STIM1 cDNA (GenBank accession number: NM_003156.3, Addgene plasmid 19755) as a template, the STIM1-UI region was cloned into pGEX-4T-1 (for GST-fused STIM1-UI) or pMO91 mammalian expression vector (for STIM1-SBR), and GST-fused STIM1-UI protein was expressed in Escherichia coli (DH5α), as previously described [29]. PCR primers for the cloning are listed in Fig. S1. The sequences of all constructs were confirmed by sequencing both strands using an ABI Prism 3700 DNA Sequencer (Applied Biosystems, Foster City, CA, USA).
Preparation of a triad vesicle sample and binding assay of GST-fused STIM1-UI protein with triad proteins
The triad vesicles were prepared from rabbit skeletal muscle and were solubilized to make the triad vesicle sample, as previously described [45]. All steps for surgical intervention as well as for pre- and post-surgical animal care were provided in accordance with the Laboratory Animals Welfare Act, the Guide for the Care and Use of Laboratory Animals and the Guidelines and Policies for Rodent Survival Surgery provided by the Institutional Animal Care and Use Committee of the College of Medicine at The Catholic University of Korea. The binding assay of GST-fused STIM1-UI protein with triad proteins was performed as described in Fig. S2a.
In-gel digestion, protein identification by quadrupole time-of-flight mass spectrometry (qTOF MS), and a database search
Protein bands obtained from the binding assays were subjected to in-gel digestion with trypsin, as previously described [61]. The digested peptide solution was subjected to qTOF MS for protein identification, and the database searches were conducted using the Mascot server (www.matrixscience.com), as described in Fig. S2b.
Cell culture, cDNA transfection, co-immunoprecipitation assay, immunoblot assay, immunocytochemistry, and measurement of the width of myotubes
Mouse primary skeletal myoblasts were derived from the skeletal muscles of mice, then allowed to proliferate and differentiate to myotubes (i.e., myotube formation), as previously described [43, 60]. Reagents and materials for the cell culture were conducted as previously described [59]. For the cDNA transfection of wild-type STIM1 (Addgene plasmid 19755) or STIM1-SBR, after 3 days of culture in the differentiation medium, the immature myotubes were transfected with the cDNA of wild-type STIM1 or STIM1-SBR (with a mixture of 30 μl of FuGENE6 transfection reagent (Promega, Madison, WI, USA) and 20 μg of cDNA per 10-cm dish or the same ratio of the components in the wells of 96-well plates). On differentiation day 5 (at 38 h post-transfection), the fully differentiated myotubes were subjected to further experiments. The successful expression of each protein was confirmed using immunocytochemistry (approximately 40 %). For the immunoblot assay, the fully differentiated myotubes were solubilized as previously described [61]. The experimental procedures for co-immunoprecipitation assay, immunoblot assay, and immunocytochemistry are presented in Fig. S2c. For the width measurement of myotubes that were transfected with either one of the STIM1 constructs or with the #2 siRNA, images of fully differentiated myotubes on D5 were captured using the monochrome camera, and the width of the thickest part in each myotube was measured using the Image J program [8].
STIM1-knockdown and quantitative real-time PCR (qPCR)
Two different sequences of small interference RNAs (siRNAs) for mouse STIM1 (GenBank accession number: NM_009287.4) were selected using siRNA design software (siDirect, Dharmacon RNAi Technologies, Lafayette, CO, USA) (Fig. S3). Transfection of the siRNAs to the immature myotubes on differentiation day 3 and qPCR were performed as previously described [30].
Oxalate-supported 45Ca2+-uptake experiment using STIM1-knockdown myotube homogenate
The STIM1-knockdown myotubes were homogenated in a buffer (50 mM KH2PO4, 10 mM NaF, 1 mM EDTA, 0.3 M sucrose, protease inhibitor cocktail (Roche, Switzerland), and 0.5 mM DTT at pH 7.0) with a homogenizer for 15 s at speed 5 (IKA T10basic Ultra-turrax, Wilmington, NC, USA). Then, 250 μg of the myotube homogenate was subjected to the oxalate-supported 45Ca2+-uptake experiment. Briefly, the reaction buffer was composed of 40 mM imidazole, 100 mM KCl, 5 mM MgCl2, 5 mM NaN3, and 0.5 mM EGTA at pH 7.0. The washing buffer was composed of 100 mM KCl and 20 mM MOPS at pH 7.0. The uptake reaction was begun by the rapid sequential addition of 5 mM MgATP, 5 mM K-oxalate, and either 70 nM or 1 μM of free 45Ca2+ (Perkin-Elmer, Waltham, MA, USA). The rate of 45Ca2+ uptake was calculated from the linear regression of 45Ca2+ uptake at 0, 1, 2, 3, and 4 min.
Single myotube Ca2+ imaging experiment: measurement of resting cytosolic Ca2+ level, caffeine or KCl response, releasable Ca2+ from the SR, and SOCE
Myotubes were loaded with 5 μM of fluo-4, fura-2 (for the measurement of the resting cytosolic Ca2+ level), or fluo-5 N (for the measurement of the releasable Ca2+ from the SR) in an imaging solution (125 mM NaCl, 5 mM KCl, 2 mM KH2PO4, 2 mM CaCl2, 25 mM HEPES, 6 mM glucose, 1.2 mM MgSO4, 0.05 % BSA (fraction V), and pH 7.4.) at 37 °C for 45 min. The loaded myotubes were subjected to single myotube Ca2+ imaging experiments, as previously described [30, 59]. Either KCl or caffeine was dissolved in the imaging solution and applied via an auto-perfusion system (AutoMate Scientific, Berkeley, CA, USA). To measure the releasable Ca2+ from the SR, the SR was depleted by incubating the myotubes in the imaging solution with zero Ca2+ for 5 min followed by treatment with either thapsigargin (TG; dissolved in Me2SO, <0.05 %) or caffeine. Me2SO (0.05 %) alone had no effect on the Ca2+ release from the SR. For the measurement of SOCE, the SR was depleted with TG in the absence of extracellular Ca2+, and once the cytosolic Ca2+ level returned to the baseline, the imaging solution with 2 mM Ca2+ was added to myotubes to measure SOCE. To analyze the Ca2+ release, values for both the peak amplitude and the area under the curve, which exhibited similar increases and decreases, were considered. For relatively long-term release of Ca2+ (more than 10 min of recording), the areas under the curves were analyzed. All reagents for Ca2+ imaging experiments were obtained from Sigma-Aldrich (St. Louis, MO, USA).
In silico approaches
A comparison of amino acid sequences was conducted using Jpred3 [7] and BLAST 2.0 [65]. The secondary structure was predicted using Jpred3 and 3D-JIGSAW 3.0 [1]. The three-dimensional (3D) structure was predicted using RaptorX [20] and was presented using RasMol [48]. The possible phosphorylation sites were predicted using NetPhos [2].
Statistical analysis
The results are presented as the means ± SE for the number of myotubes shown in parentheses in Table 2. The values were normalized to the mean value from the corresponding controls. The significant differences were analyzed using a paired t-test (significant at p < 0.05, GraphPad InStat, v2.04; GraphPad Software, La Jolla, CA, USA). The graphs were prepared using Origin v7 software (Origin Lab, Northampton, MA, USA).
Results
STIM1 binds to SERCA1a via the STIM1-UI region (STIM1-SBR) in skeletal muscle
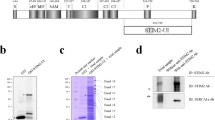
To search for the proteins that could possibly interact with the STIM1-UI (223 amino acids from 449 to 671 of STIM1), the cDNA for a GST-fused STIM1-UI protein was constructed (Fig. 1a and Fig. S1). The GST-fused STIM1-UI protein was expressed in E. coli, the bacterial lysate was separated on a SDS-PAGE gel, and the gel was stained with Coomassie Blue (Fig. 1b). The GST-fused STIM1-UI protein was successfully expressed. For binding assays, immobilized GST-fused STIM1-UI proteins on GST beads (Fig. 1c) were incubated with the solubilized triad vesicle sample from rabbit skeletal muscle. The triad vesicle sample is composed of junctional SRs and t-tubules that are enriched portions with triad proteins mediating the contraction and relaxation of skeletal muscle [26, 27, 61]. The proteins that were bound to the affinity beads were separated on a SDS-PAGE gel and stained with Coomassie Blue in order to evaluate the proteins that were specifically bound to the GST-fused STIM1-UI protein (Fig. 1d). The bands for the proteins bound to the GST itself were excluded from consideration. Four bands appeared as proteins that were specifically bound to the GST-fused STIM1-UI protein, and were subjected to in-gel digestion with trypsin, qTOF MS, and database searches for protein identification. Figure S4 and Table 1 show the results of the qTOF MS and database searches. For band 1, there was no matching signal in the known databases. Band 2 was identified as a SERCA1a that originated from rabbit skeletal muscle, suggesting that the STIM1-UI region could bind to SERCA1a. Bands 3 and 4 were identified as non-specifically bound proteins that originated from the E. coli lysate during the binding assays.
Schematic primary sequences of STIM1 and STIM1-UI, and binding assays of GST-fused STIM1-UI protein with triad proteins. a. The position and/or function of each domain in STIM1 is presented according to previous reports: the overall diagram [19], EF [69], EF-SAM [52], OL [10], CAD [39], SOAR [66], L [39], C1 [10], and T [21]. Numbers indicate the sequence of amino acids. b. Bacterial lysate expressing GST-fused STIM1-UI protein was separated on a 10 % SDS-PAGE gel and the gel was stained with Coomassie Blue. GST and GST-fused STIM1-UI proteins are indicated by asterisks. c. Immobilized GST-fused STIM1-UI proteins on GST beads were separated on a 10 % SDS-PAGE gel and the gel was stained with Coomassie Blue. d. The bound proteins obtained from the binding assays of GST-fused STIM1-UI protein with the triad proteins from rabbit skeletal muscle were separated on a 10 % SDS-PAGE gel and the gel was stained with Coomassie Blue. GST was used as a negative control. GST or GST-fused STIM1-UI proteins are indicated by white asterisks. The four specifically bound proteins to the GST-fused STIM1-UI protein compared with the GST control are indicated by white dashes on the right side of lane 4 (SR + STIM1-UI) and were named bands 1 to 4
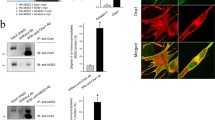
The binding of endogenous full-length STIM1 to SERCA1a was accessed using a triad vesicle sample from rabbit skeletal muscle by co-immunoprecipitation assay with either anti-STIM1 or anti-SERCA1a antibody. SERCA1a was co-immunoprecipitated with STIM1 in both cases (Fig. 2a). Immunocytochemistry using mouse primary skeletal myotubes with anti-STIM1 and anti-SERCA1a antibodies showed the co-localization of STIM1 and SERCA1a near the nucleus (Fig. 2b, indicated by arrowheads in the merged and enlarged image). Therefore, based on the three different approaches, binding assay and qTOP MS, co-immunoprecipitation assay, and immunocytochemistry, the STIM1-UI region participated in the binding to SERCA1a in skeletal muscle and was named STIM1-SBR (SERCA1a-binding region).
Co-immunoprecipitation and co-localization of endogenous full-length STIM1 with SERCA1a. a. Co-immunoprecipitation assay of endogenous full-length STIM1 with SERCA1a was conducted using a triad vesicle sample from rabbit skeletal muscle with either anti-STIM1 (left-hand panel) or anti-SERCA1a antibody (right-hand panel). STIM1 and SERCA1a were co-immunoprecipitated in both cases. The control was the triad vesicle sample alone. Three independent experiments were conducted and a representative result is presented. b. Mouse primary skeletal myotubes were double-stained with anti-STIM1 and anti-SERCA1a antibodies. The boxed area in the merged image is enlarged in the right-hand panel. The merged image shows the co-localization of STIM1 with SERCA1a near the nucleus (indicated by arrowheads in the enlarged image). DIC differential interference contrast microscopy
STIM1 is required for the full activity of SERCA1a in skeletal myotubes
To examine how STIM1 is functionally related to SERCA1a activity in skeletal muscle, STIM1 was knocked down in mouse primary skeletal myotubes by siRNA transfection to the immature myotubes. The results of qPCR using the fully differentiated myotubes showed that the siRNA transfection knocked down STIM1 efficiently (84.5 ± 7.4 % reduction in mRNA, adopted from our previous study [30]; Fig. S3). Immunoblot assay using the lysate of STIM1-knockdown myotubes with anti-STIM1 or anti-SERCA1a antibody showed that STIM1 expression was reduced by up to 95 % by the STIM1-knockdown, and that there was no significant change in the expression level of SERCA1a (Fig. 3a). To rule out the possibility that any changes were due to the considerable differences in the sizes of the myotubes (i.e., the degree of differentiation), the widths of the myotubes were measured. There were no considerable differences in the widths of the myotubes transfected with #2 siRNA compared with those of untransfected myotubes [30]. In addition, the myotube formations were not significantly affected by the STIM1-knockdown compared with either untransfected or scrambled siRNA-transfected controls (Fig. 3b). Therefore, STIM1 was successfully knocked down in mouse primary skeletal myotubes without affecting either SERCA1a expression or myotube formation.
Knockdown of STIM1 in mouse primary skeletal myotubes and 45Ca2+ uptake into the SR in the STIM1-knockdown myotubes. a. The lysate of the STIM1-knockdown myotubes (50 μg of total protein) was subjected to immunoblot assay with anti-STIM1 or anti-SERCA1a antibody. α-actin and gels stained with Coomassie Blue were loading controls. Untransfected or scrambled siRNA-transfected myotubes were used as negative controls. The expression of STIM1 protein was significantly reduced by up to 95 % by the STIM1-knockdown. The expression level of SERCA1a was not significantly changed by the STIM1-knockdown. Three independent experiments were conducted and a representative result was presented. b. The image of STIM1-knockdown myotubes is presented. STIM1-knockdown myotubes showed no significant change in myotube formations compared with controls. Bar represents 150 μm. c. Oxalate-supported 45Ca2+ uptake into the SR using the homogenate of the STIM1-knockdown myotubes was measured at a low or high concentration of free 45Ca2+ (70 nM or 1 μM, respectively). The STIM1-knockdown myotubes showed a significantly decreased 45Ca2+ uptake only at the high concentration of free 45Ca2+. The results are presented as the mean ± SE of three independent experiments. *Significant difference compared with untransfected controls (p < 0.05)
Because it is important for skeletal muscle relaxation, Ca2+ uptake from the cytosol to the SR by SERCA1a was examined in the STIM1-knockdown myotubes using an oxalate-supported 45Ca2+-uptake assay. At a resting cytosolic Ca2+ concentration (70 nM of free 45Ca2+ [59, 60]), the Ca2+-uptake activity of SERCA1a was unchanged (Fig. 3c, left-hand panel). At a higher cytosolic Ca2+ concentration, such as that during the transition from skeletal muscle contraction to relaxation (1 μM of free 45Ca2+ [26, 47]), the Ca2+-uptake activity of SERCA1a was significantly decreased (Fig. 3c, right-hand panel; almost 50 % decrease compared with the untransfected control). Therefore, STIM1 is required for the full activity of SERCA1a by increasing SERCA1a activity during skeletal muscle relaxation but not during the resting state.
STIM1-SBR decreases RyR1 activity in skeletal myotubes
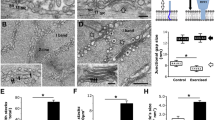
To examine the possible role of STIM1-SBR in skeletal ECC, STIM1-SBR was expressed in mouse primary skeletal myotubes. Myotube formation was not significantly affected by the expression of STIM1-SBR (Fig. 4a). KCl was applied to the myotubes expressing STIM1-SBR. KCl depolarizes t-tubule membranes, activates DHPR, and induces the subsequent Ca2+ release from the SR to the cytosol via RyR1, which ultimately results in skeletal ECC (i.e., the response to KCl reflects ECC) [26, 27, 50]. As expected [30], wild-type STIM1 induced a significant decrease in the Ca2+ release from the SR in response to KCl compared with the control vector (Fig. 4b and Table 2). Similar to wild-type STIM1, STIM1-SBR also induced a decrease in the Ca2+ release from the SR in response to KCl (Fig. 4b and Table 2). There are two possible explanations for the decreased Ca2+ release in response to KCl by STIM1-SBR. STIM1-SBR attenuates either DHPR activity directly, the same as wild-type STIM1 [30], or attenuates RyR1 activity without affecting DHPR activity. To test these possibilities, RyR1 activity was assessed by applying a direct agonist of RyR1, caffeine, to myotubes expressing STIM1-SBR. Unlike wild-type STIM1, STIM1-SBR induced a decrease in Ca2+ release through RyR1 in response to caffeine (Fig. 4c, Table 2), suggesting that RyR1 activity is reduced by STIM1-SBR, and that the decreased Ca2+ release in response to KCl by STIM1-SBR could be primarily due to the reduced RyR1 activity but not due to the changed DHPR activity itself. In addition, considering the facts that STIM1-SBR is a part of STIM1 and that, however, the full-length wild-type STIM1 did not change RyR1 activity, the function of STIM1-SBR in the full-length STIM1 could be masked either by other parts of STIM1 or by other proteins. To rule out the possibility that changes in the responses to either KCl or caffeine were due to considerable differences in the size of the myotubes (i.e., the degree of differentiation), the widths of the myotubes were measured (Fig. 5a). There was no considerable difference between the widths of the myotubes expressing wild-type STIM1 or STIM1-SBR compared with those of myotubes transfected with the control vector. In addition, to rule out the possibility that the functional changes were due to considerable differences in the expression levels of either DHPR or RyR1, the expression levels of DHPR and RyR1 were examined using an immunoblot assay with anti-DHPR or anti-RyR1 antibody, which showed no significant changes in expression levels (Fig. 5b).
The response of myotubes expressing wild-type STIM1 or STIM1-SBR to KCl or caffeine. a. Myotubes expressing wild-type STIM1 or STIM1-SBR were stained with anti-CFP antibodies. Bar represents 100 μm. Each protein was successfully expressed in the myotubes without affecting myotube formation. b. KCl was applied to the myotubes. A representative trace for each group is shown in the left-hand panel. Histograms of the peak amplitude normalized to the mean value of those from the control vectors are shown in the right-hand panel. Wild-type STIM1 or STIM1-SBR significantly decreased Ca2+ release from the SR in response to KCl compared with the control vector. c. Following caffeine administration, only STIM1-SBR significantly decreased Ca2+ release from the SR compared with the control vector. *Significant difference compared with the control vector (p < 0.05, Table 2)
Myotube width, expression levels of RyR1, DHPR, and SERCA1a, resting cytosolic Ca2+ level, and releasable Ca2+ from the SR in myotubes expressing wild-type STIM1 or STIM1-SBR. a. To measure the width of the myotubes, the thickest part of the myotubes expressing either wild-type STIM1 or STIM1-SBR was measured. Width values were normalized to the mean value of those from the control vector and are presented as histograms. The results are presented as the mean ± SE for 50 myotubes per each group. There was no considerable change in the width of myotubes according to the expressions. b. The lysate of the myotubes expressing wild-type STIM1 or STIM1-SBR (50 μg of total protein) was subjected to immunoblot assay with anti-RyR1, anti-DHPR, or anti-SERCA1a antibody. α-Actin was a loading control. The expression levels of RyR1, DHPR, or SERCA1a were not significantly changed by the expression. Three independent experiments were conducted and a representative result is presented. c. Resting cytosolic Ca2+ levels in the myotubes were measured. Resting cytosolic Ca2+ levels were significantly decreased by STIM1-SBR. d. To measure the releasable Ca2+ from the SR to the cytosol, TG was applied to the myotubes in the absence of extracellular Ca2+. A representative trace for each group is shown (left). The releasable Ca2+ from the SR is summarized as histograms in the right-hand panel (the area under the curve was normalized to the mean value of those from the control vectors). The releasable Ca2+ from SR to the cytosol was significantly increased by STIM1-SBR. *Significant difference compared with the control vector (p < 0.05, Table 2)
The reduced RyR1 activity by STIM1-SBR is possibly due to the increased SERCA1a activity
The reduced RyR1 activity in myotubes expressing STIM1-SBR is mediated by either STIM1-SBR directly or via other proteins. Direct mediation is not plausible because STIM1 does not bind to RyR1 [30]. On the other hand, RyR1 is also activated by sub-micromolar Ca2+ through the CICR mechanism although RyR1 is the channel for Ca2+ movement, and cytosolic Ca2+ makes RyR1 more sensitive to RyR1 agonists such as caffeine [26, 27, 70]. To establish how the reduced RyR1 activity by STIM1-SBR might be due to a decrease in Ca2+ levels, first, the cytosolic Ca2+ level at rest was measured in the myotubes. It was noteworthy that STIM1-SBR significantly decreased cytosolic Ca2+ levels at rest compared with the control vector (Fig. 5c, Table 2). Therefore, the decreased cytosolic Ca2+ level would be the cause of the reduced RyR1 activity. In addition, considering that cytosolic Ca2+ levels at rest are decreased, it is possible that SERCA1a activity is increased by STIM1-SBR and that SR-stored Ca2+ levels also are subsequently increased. To test this possibility, SR-stored Ca2+ levels in the myotubes were accessed indirectly by depleting the SR with TG in the absence of extracellular Ca2+ and then measuring the released Ca2+ from the SR (Fig. 5d, Table 2). STIM1-SBR increased the releasable Ca2+ from the SR compared with the control vector. Similar results were obtained from the myotubes, which were loaded with fluo-5 N (which detects higher levels of Ca2+ ranging from μM to mM) and then treated with caffeine in the absence of extracellular Ca2+ (Fig. S5). The possible change in the expression level of SERCA1a was examined using immunoblot assay, which showed no significant change in SERCA1a expression (Fig. 5b). Therefore, STIM1-SBR would reduce RyR1 activity indirectly by increasing SERCA1a activity.
STIM1-SBR is irrelevant to SOCE in skeletal myotubes
The role of STIM1-SBR in SOCE was examined because STIM1 is one of two major SOCE-mediating proteins in skeletal muscle. Myotubes expressing wild-type STIM1 or STIM1-SBR were treated with TG in order to deplete the SR; then, 2 mM of extracellular Ca2+ was applied to the myotubes (Fig. 6). As expected [30], wild-type STIM1 increased SOCE compared with the control vector (approximately 47 %, Table 2). However, STIM1-SBR did not change SOCE, suggesting that STIM1-SBR had no effect on SOCE and is irrelevant to SOCE in skeletal myotubes. Therefore, the effects of STIM1-SBR on SERCA1a activity, and its subsequent effect on RyR1 activity, in skeletal myotubes must be related to the 'intracellular Ca2+ movements' rather than to SOCE.
SOCE in myotubes expressing wild-type STIM1 or STIM1-SBR. After the SR of the myotubes was depleted by the treatment of TG in the absence of extracellular Ca2+, 2 mM of extracellular Ca2+ was applied to the myotubes to induce SOCE. A representative trace for each group is shown in the left-hand panel. Only wild-type STIM1 significantly increased SOCE. The SOCE is summarized in the histograms (the area under the curve was normalized to the mean value of the control vectors). *Significant difference compared with the control vectors (p < 0.05, Table 2)
Discussion
In the present study, to reveal the unidentified functional role of the STIM1 C terminus, from 449 to 671 amino acids in skeletal muscle and the proteins binding to the region were searched along with proteins that mediate skeletal muscle contraction and relaxation. In skeletal muscle, STIM1 binds to SERCA1a via this region (called STIM1-SBR). The binding was confirmed in endogenous full-length STIM1 in the skeletal muscle of rabbits and in the primary skeletal myotubes of mice. STIM1-knockdown in mouse primary skeletal myotubes decreases SERCA1a activity (almost 50 %) only at a micromolar cytosolic Ca2+ concentration, suggesting that STIM1 is required for the full activity of SERCA1a by increasing SERCA1a activity possibly during skeletal muscle relaxation. STIM1-SBR in mouse primary skeletal myotubes reduces RyR1 activity possibly by increasing SERCA1a activity, and STIM1-SBR is irrelevant to SOCE in skeletal muscle.
STIM1 could increase SERCA1a activity in skeletal myotubes at a higher cytosolic Ca2+ concentration
In our previous report [30], STIM1-knockdown in skeletal myotubes did not induce a change in neither the releasable Ca2+ from the SR to cytosol, nor in the resting cytosolic Ca2+ level. Therefore, it was expected that STIM1-knockdown had no effect on SERCA1a activity at resting cytosolic Ca2+ levels (Fig. 3c, left-hand panel). These results suggest that, at resting cytosolic Ca2+ levels, SERCA1a activity does not depend on the presence of STIM1. On the other hand, during the differentiation process of skeletal myoblasts to myotubes, resting cytosolic Ca2+ level is increased via extracellular Ca2+ entry, which is one of the critical requirements for differentiation [12, 42, 49]. Considering that the cDNA of STIM1-SBR was transfected to the immature myotubes in the middle of the differentiation and that STIM1 could increase SERCA1a activity only at micromolar cytosolic Ca2+ concentration (Fig. 3c, right-hand panel), it is possible that STIM1-SBR could increase SERCA1a activity by the increased resting cytosolic Ca2+ level during differentiation and could result in a decrease in resting cytosolic Ca2+ level and/or in an increase in SR-stored Ca2+ in the differentiated myotubes. Indeed, myotubes expressing STIM1-SBR showed a decreased resting cytosolic Ca2+ level and an increase in the releasable Ca2+ from the SR to cytosol (Fig. 5c and d), empathizing again that STIM1 increases SERCA1a activity at a higher cytosolic Ca2+ concentration.
STIM1 could play an ambidextrous role in regulating Ca2+ distributions of skeletal myotubes
The plasma membrane Ca2+-ATPase (PMCA) in Jurkat T-cells plays an important role in cytosolic Ca2+ clearance after they are activated [31], and STIM1 attenuates PMCA-mediated Ca2+ clearance and results in sustained cytosolic Ca2+ elevation without affecting SOCE [23, 44]. On the other hand, STIM1 binds to SERCA2a (cardiac isoform of SRCA1a) either directly [46] or indirectly via the POST in heterologous expression systems [23], but in both cases the functional role of STIM1 in SERCA2 activity has not been addressed. In the present study, we found that STIM1 directly binds to SERCA1a via STIM1-SBR and is required for the full activity of SERCA1a without affecting SOCE possibly during skeletal muscle relaxation. Taken together, it is a possible scenario that the eventual role of STIM1 in skeletal muscle relaxation could be to refill the SR sufficiently both by attenuating PMCA in order not to allow it to extrude too much Ca2+ to the extracellular space, and by maintaining SERCA1a activity at full strength to maximize Ca2+ uptake to the SR. This is a plausible assumption because, in skeletal muscle contraction, Ca2+ from the SR is the main source of Ca2+ for the contraction, and Ca2+ entry from the extracellular space to the cytosol through DHPR is traceable [26, 27].
Myotubes expressing STIM1-SBR show a decrease in resting cytosolic Ca2+ levels and an increase in SR-stored Ca2+ levels (Fig. 5), and STIM1 knockdown myotubes show a decrease in SERCA1a activity (Fig. 3c). Therefore, STIM1 could be involved in the intracellular Ca2+ distribution between the SR and the cytosol in skeletal muscle. Considering that STIM1 is involved in both SOCE via 'its N-terminal EF hand (one hand)' by sensing SR Ca2+ levels [24, 54] and in ECC via 'its C-terminal STIM1-SBR (the other hand)' by regulating SERCA1a activity (in the present study), STIM1 plays an ambidextrous role in regulating Ca2+ movements in skeletal muscle.
STIM1-SBR shares homologies in appearance with other SERCA-binding proteins
The homology of mouse STIM1-SBR with other proteins in an amino acid sequence was searched using Jpred [7] and BLAST 2.0 [65]. Only hypothetical or uncharacterized proteins from chicks, xenopus, zebafish, or puffefish have high homologies with STIM1-SBR. In addition, there is no known domain or motif in STIM1-SBR, suggesting the possibility that STIM1-SBR is an intrinsically disordered region by lacking a stable 3D structure. The secondary structure of mouse STIM1-SBR was predicted using Jpred3 [7] and 3D-JIGSAW 3.0 [1]. Three alpha helixes (I, II, and III) were predicted with a high degree of confidence, and it was predicted that the C-terminal one-third of the STIM1-SBR would be disordered (Fig. 7a). In some cases of intrinsically disordered proteins, they can adopt a fixed 3D structure after binding to other proteins such as binding partners, i.e., a coupled folding [18]. Therefore, it is possible that the STIM1-SBR is a coupled folding region when it binds to a binding partner, such as SERCA1a. Predominance of protein phosphorylation in intrinsically disordered proteins has been reported [9]. Many amino acids of STIM1-SBR were predicted to be possible phosphorylation sites using NetPhos [2] (12.5 %, including 25 serines and three threonines; Fig. S6).
Secondary or 3D structure prediction of mouse STIM1-SBR. a. Secondary structure of mouse STIM1-SBR was predicted using 3D-JIGSAW 3.0 [1] and Jpred3 [7]. H (red), E (blue), or C (black) represents alpha-helix, beta-strand, or random coil, respectively. Confidence scores ranging from 0 to 9 indicate low to high confidence for the prediction, respectively. Numbers indicate the sequence of amino acids. Three dashed boxes indicate three predicted helixes with high confidence scores. D (yellow) or O (green) indicates whether the corresponding amino acid exists in the disordered or ordered state, respectively. The predicted phosphorylation sites in helix I (Fig. S6) are indicated by dots. b. The 3D structure of mouse STIM1-SBR was predicted using RaptorX [20] and presented as a ribbon diagram using RasMol [48]. The alpha-helix, beta-strand, and connecting loop are presented as red, yellow, and white, respectively. N or C means N or C terminus, respectively. The predicted 3D structure of the STIM1-SBR in the right-hand panel resembles a tri-wing boomerang. The left image is the 90° rotation along the vertical axis of the view of the right image. The helix I and helix III in the predicted secondary structure in a. are indicated in the predicted 3D structure
In order to predict the possible 3D structure in the coupled folding state of mouse STIM1-SBR, first, the homologies of the amino acid sequences were searched using RaptorX, which is a prediction server for the 3D structure of proteins and searches the homologies in amino acid sequences with other proteins with the 3D structures that are deposited in the Protein Data Bank (PDB) [20]. A homology between the STIM1-SBR and heme-dependent catalase HPII or one of its mutants (PDB ID: 1gg9, 1gge, 1ggj, 1p80, 1p81 or 1qws) was found (Fig. S7). Based on the 3D structure of HPII and its mutants, the 3D structure of the STIM1-SBR was predicted to be a tri-wing boomerang with four alpha helixes (Fig. 7b). The helix I and helix III in the predicted secondary structure in Fig. 7a was accurately predicted in the 3D structure prediction, although the helix III was predicted to be divided into two shorter helixes.
Sarcolipin or MG53 binds to and attenuates SERCA1a activity in skeletal muscle, but there has been no report on the counteracting protein against sarcolipin or MG53, although sarcalumenin increases SERCA1a activity [6, 26, 28, 38, 70]. It seems that an alpha-helix in sarcolipin or MG53 participates in the binding to SERCA1a [28, 36]. The predicted helixes in mouse STIM1-SBR in Fig. 7 may play a role in SERCA1a regulation, as does the alpha-helix in sarcolipin or MG53. In addition, there are two phosphorable serines in the predicted helix I of the STIM1-SBR (Fig. 7a, indicated by dots, and Fig. S6). In cardiac and slow-twitch skeletal muscle, phospholamban binds to SERCA2a, which is the major isoform of SERCA in the muscles [6], and SERCA2a activity depends on the phosphorylation state of phospholamban [22]. Therefore, it is possible that the binding of STIM1 to SERCA1a via STIM1-SBR could be mediated by methods of binding that are similar to that of other SERCA-binding proteins, and STIM1 could antagonize the attenuating effects of sarcolipin or MG53 on SERCA1a activity via these similarities in appearance.
SOCE operation could be dependent on the relative degree of SR depletion
SR depletion turns on SOCE [24, 54, 55, 59], but it seems that the size, larger or smaller, of the released Ca2+ from the SR during SR depletion is not a factor on the stimulation of SOCE in skeletal muscle, because myotubes expressing STIM1-SBR show no change in SOCE, although they do show an increase in the releasable Ca2+ from the SR (Figs. 5d and 6). Therefore, rather than the absolute amount of releasable Ca2+ from the SR in skeletal muscle, the relative amount of releasable Ca2+ to the total Ca2+ stored in the SR might be more important for SOCE operation.
Previously, we reported that the cytoplasmic C terminus of STIM1 (from 235 to 685 amino acids) covering STIM1-SBR is involved in the negative regulation of DHPR during ECC in skeletal myotubes [30]. In the present study, STIM1-SBR did not considerably change DHPR activity itself (Fig. 4b and c). These results suggest that STIM1-SBR does not participate in the negative regulation of DHPR by STIM1, and that with the exception of STIM1-SBR, the remaining C terminus, such as coiled-coil 1 or CAD/SOAR, could be responsible for the DHPR regulation in skeletal myotubes. In other words, by the present study, the critical region(s) participating in DHPR regulation was narrowed down to coiled-coil 1 or CAD/SOAR. Indeed, in both vascular smooth muscle cell lines and in rat cortical neurons, CAD/SOAR is known to suppress the activity of Cav1.2 (cardiac isoform of DHPR) [40, 57].
STIM1 could be a novel candidate for the treatment of patients with skeletal muscle diseases
Patients with Brody syndrome, an inherited myopathy due to a reduction in SERCA1a activity with no mutation in the SERCA1a gene, suffer from exercise-induced muscle stiffness and delayed muscle relaxation [56, 67]. Likewise, STIM1-knockdown in skeletal myotubes decreased SERCA1a activity with no mutation in the SERCA1a gene in the present study, and patients with loss-of-function mutations of STIM1 show congenital myopathies [15, 16, 41]. On the other hand, most skeletal muscle diseases involve increases in RyR1 activity and/or cytosolic Ca2+ levels [26, 70]. On the contrary, it is interesting that the myotubes expressing STIM1-SBR in the present study showed decreases in both RyR1 activity and cytosolic Ca2+ levels. Therefore, the regulation of STIM1 expression is a novel candidate for the treatment of patients with skeletal muscle diseases that accompany changes in RyR1 activity, in SERCA1a activity, and/or in cytosolic Ca2+ levels.
Abbreviations
- STIM1:
-
Stromal interaction molecule 1
- STIM1-SBR:
-
SERCA1a-binding region in STIM1
- ECC:
-
Excitation–contraction coupling
- SOCE:
-
Store-operated Ca2+ entry
- SR:
-
Sarcoplasmic reticulum
- ER:
-
Endoplasmic reticulum
- SERCA:
-
Sarcoplasmic/endoplasmic reticulum Ca2+-ATPase
- RyR:
-
Ryanodine receptor
- DHPR:
-
Dihydropyridine receptor
- t-Tubule:
-
Transverse tubule
- SAM:
-
Sterile α-motif
- SOAR:
-
STIM1-Orai1-activating region
- CRAC:
-
Ca2+ release-activated Ca2+
- CAD:
-
CRAC-activating domain
- CICR:
-
Ca2+-induced Ca2+ release
- MG53:
-
Mitsugumin 53
References
Bates PA, Kelley LA, MacCallum RM, Sternberg MJ (2001) Enhancement of protein modeling by human intervention in applying the automatic programs 3D-JIGSAW and 3D-PSSM. Proteins Suppl 5:39–46
Blom N, Gammeltoft S, Brunak S (1999) Sequence and structure-based prediction of eukaryotic protein phosphorylation sites. J Mol Biol 294(5):1351–1362
Boittin FX, Petermann O, Hirn C, Mittaud P, Dorchies OM, Roulet E, Ruegg UT (2006) Ca2+-independent phospholipase A2 enhances store-operated Ca2+ entry in dystrophic skeletal muscle fibers. J Cell Sci 119(Pt 18):3733–3742
Brandl CJ, deLeon S, Martin DR, MacLennan DH (1987) Adult forms of the Ca2+-ATPase of sarcoplasmic reticulum. Expression in developing skeletal muscle. J Biol Chem 262(8):3768–3774
Brandl CJ, Green NM, Korczak B, MacLennan DH (1986) Two Ca2+ ATPase genes: homologies and mechanistic implications of deduced amino acid sequences. Cell 44(4):597–607
Brini M, Carafoli E (2009) Calcium pumps in health and disease. Physiol Rev 89(4):1341–1378
Cole C, Barber JD, Barton GJ (2008) The Jpred 3 secondary structure prediction server. Nucleic Acids Res 36(Web Server issue):W197–W201
Collins TJ (2007) ImageJ for microscopy. Biotechniques 43(1 Suppl):25–30
Collins MO, Yu L, Campuzano I, Grant SG, Choudhary JS (2008) Phosphoproteomic analysis of the mouse brain cytosol reveals a predominance of protein phosphorylation in regions of intrinsic sequence disorder. Mol Cell Proteomics 7(7):1331–1348
Covington ED, Wu MM, Lewis RS (2010) Essential role for the CRAC activation domain in store-dependent oligomerization of STIM1. Mol Biol Cell 21(11):1897–1907
Darbellay B, Arnaudeau S, Konig S, Jousset H, Bader C, Demaurex N, Bernheim L (2009) STIM1- and Orai1-dependent store-operated calcium entry regulates human myoblast differentiation. J Biol Chem 284(8):5370–5380
David JD, See WM, Higginbotham CA (1981) Fusion of chick embryo skeletal myoblasts: role of calcium influx preceding membrane union. Dev Biol 82(2):297–307
Edwards JN, Friedrich O, Cully TR, von Wegner F, Murphy RM, Launikonis BS (2010) Upregulation of store-operated Ca2+ entry in dystrophic mdx mouse muscle. Am J Physiol Cell Physiol 299(1):C42–C50
Edwards JN, Murphy RM, Cully TR, von Wegner F, Friedrich O, Launikonis BS (2010) Ultra-rapid activation and deactivation of store-operated Ca2+ entry in skeletal muscle. Cell Calcium 47(5):458–467
Feske S, Picard C, Fischer A (2010) Immunodeficiency due to mutations in ORAI1 and STIM1. Clin Immunol 135(2):169–182
Fuchs S, Rensing-Ehl A, Speckmann C, Bengsch B, Schmitt-Graeff A, Bondzio I, Maul-Pavicic A, Bass T, Vraetz T, Strahm B, Ankermann T, Benson M, Caliebe A, Folster-Holst R, Kaiser P, Thimme R, Schamel WW, Schwarz K, Feske S, Ehl S (2012) Antiviral and regulatory T cell immunity in a patient with stromal interaction molecule 1 deficiency. J Immunol 188(3):1523–1533
Grosse J, Braun A, Varga-Szabo D, Beyersdorf N, Schneider B, Zeitlmann L, Hanke P, Schropp P, Muhlstedt S, Zorn C, Huber M, Schmittwolf C, Jagla W, Yu P, Kerkau T, Schulze H, Nehls M, Nieswandt B (2007) An EF hand mutation in Stim1 causes premature platelet activation and bleeding in mice. J Clin Invest 117(11):3540–3550
Gunasekaran K, Tsai CJ, Kumar S, Zanuy D, Nussinov R (2003) Extended disordered proteins: targeting function with less scaffold. Trends Biochem Sci 28(2):81–85
Hogan PG, Lewis RS, Rao A (2010) Molecular basis of calcium signaling in lymphocytes: STIM and ORAI. Annu Rev Immunol 28:491–533
Kallberg M, Wang H, Wang S, Peng J, Wang Z, Lu H, Xu J (2012) Template-based protein structure modeling using the RaptorX web server. Nat Protoc 7(8):1511–1522
Kim JY, Muallem S (2011) Unlocking SOAR releases STIM. EMBO J 30(9):1673–1675
Kim HW, Steenaart NA, Ferguson DG, Kranias EG (1990) Functional reconstitution of the cardiac sarcoplasmic reticulum Ca2+-ATPase with phospholamban in phospholipid vesicles. J Biol Chem 265(3):1702–1709
Krapivinsky G, Krapivinsky L, Stotz SC, Manasian Y, Clapham DE (2011) POST, partner of stromal interaction molecule 1 (STIM1), targets STIM1 to multiple transporters. Proc Natl Acad Sci U S A 108(48):19234–19239
Kurebayashi N, Ogawa Y (2001) Depletion of Ca2+ in the sarcoplasmic reticulum stimulates Ca2+ entry into mouse skeletal muscle fibres. J Physiol 533(Pt 1):185–199
Launikonis BS, Rios E (2007) Store-operated Ca2+ entry during intracellular Ca2+ release in mammalian skeletal muscle. J Physiol 583(Pt 1):81–97
Lee EH (2010) Ca2+ channels and skeletal muscle diseases. Prog Biophys Mol Biol 103(1):35–43
Lee EH, Kim DH, Allen PD (2006) Interplay between intra- and extracellular calcium ions. Mol Cells 21(3):315–329
Lee KJ, Park CS, Woo JS, Kim DH, Ma J, Lee EH (2012) Mitsugumin 53 attenuates the activity of sarcoplasmic reticulum Ca2+-ATPase 1a (SERCA1a) in skeletal muscle. Biochem Biophys Res Commun 428(3):383–388
Lee EH, Rho SH, Kwon SJ, Eom SH, Allen PD, Kim DH (2004) N-terminal region of FKBP12 is essential for binding to the skeletal ryanodine receptor. J Biol Chem 279(25):26481–26488
Lee KJ, Woo JS, Hwang JH, Hyun C, Cho CH, Kim DH, Lee EH (2013) STIM1 negatively regulates Ca2+ release from the sarcoplasmic reticulum in skeletal myotubes. Biochem J 453(2):187–200
Lewis RS (2001) Calcium signaling mechanisms in T lymphocytes. Annu Rev Immunol 19:497–521
Li T, Finch EA, Graham V, Zhang ZS, Ding JD, Burch J, Oh-Hora M, Rosenberg P (2012) STIM1-Ca2+ signaling is required for the hypertrophic growth of skeletal muscle in mice. Mol Cell Biol 32(15):3009–3017
Liou J, Kim ML, Heo WD, Jones JT, Myers JW, Ferrell JE Jr, Meyer T (2005) STIM is a Ca2+ sensor essential for Ca2+-store-depletion-triggered Ca2+ influx. Curr Biol 15(13):1235–1241
Luik RM, Wang B, Prakriya M, Wu MM, Lewis RS (2008) Oligomerization of STIM1 couples ER calcium depletion to CRAC channel activation. Nat Geosci 454(7203):538–542
Manji SS, Parker NJ, Williams RT, van Stekelenburg L, Pearson RB, Dziadek M, Smith PJ (2000) STIM1: a novel phosphoprotein located at the cell surface. Biochim Biophys Acta 1481(1):147–155
Mascioni A, Karim C, Barany G, Thomas DD, Veglia G (2002) Structure and orientation of sarcolipin in lipid environments. Biochemistry 41(2):475–482
Muik M, Fahrner M, Derler I, Schindl R, Bergsmann J, Frischauf I, Groschner K, Romanin C (2009) A cytosolic homomerization and a modulatory domain within STIM1 C terminus determine coupling to ORAI1 channels. J Biol Chem 284(13):8421–8426
Odermatt A, Becker S, Khanna VK, Kurzydlowski K, Leisner E, Pette D, MacLennan DH (1998) Sarcolipin regulates the activity of SERCA1, the fast-twitch skeletal muscle sarcoplasmic reticulum Ca2+-ATPase. J Biol Chem 273(20):12360–12369
Park CY, Hoover PJ, Mullins FM, Bachhawat P, Covington ED, Raunser S, Walz T, Garcia KC, Dolmetsch RE, Lewis RS (2009) STIM1 clusters and activates CRAC channels via direct binding of a cytosolic domain to Orai1. Cell 136(5):876–890
Park CY, Shcheglovitov A, Dolmetsch R (2010) The CRAC channel activator STIM1 binds and inhibits L-type voltage-gated calcium channels. Science 330(6000):101–105
Picard C, McCarl CA, Papolos A, Khalil S, Luthy K, Hivroz C, LeDeist F, Rieux-Laucat F, Rechavi G, Rao A, Fischer A, Feske S (2009) STIM1 mutation associated with a syndrome of immunodeficiency and autoimmunity. N Engl J Med 360(19):1971–1980
Przybylski RJ, Szigeti V, Davidheiser S, Kirby AC (1994) Calcium regulation of skeletal myogenesis: II. Extracellular and cell surface effects. Cell Calcium 15(2):132–142
Rando TA, Blau HM (1997) Methods for myoblast transplantation. Methods Cell Biol 52:261–272
Ritchie MF, Samakai E, Soboloff J (2012) STIM1 is required for attenuation of PMCA-mediated Ca2+ clearance during T-cell activation. EMBO J 31(5):1123–1133
Saito A, Seiler S, Chu A, Fleischer S (1984) Preparation and morphology of sarcoplasmic reticulum terminal cisternae from rabbit skeletal muscle. J Cell Biol 99(3):875–885
Sampieri A, Zepeda A, Asanov A, Vaca L (2009) Visualizing the store-operated channel complex assembly in real time: identification of SERCA2 as a new member. Cell Calcium 45(5):439–446
Sandow A (1965) Excitation–contraction coupling in skeletal muscle. Pharmacol Rev 17(3):265–320
Sayle RA, Milner-White EJ (1995) RASMOL: biomolecular graphics for all. Trends Biochem Sci 20(9):374
Shainberg A, Yagil G, Yaffe D (1969) Control of myogenesis in vitro by Ca2+ concentration in nutritional medium. Exp Cell Res 58(1):163–167
Shamoo AE, MacLennan DH (1974) A Ca++-dependent and -selective ionophore as part of the Ca++ plus Mg++-dependent adenosinetriphosphatase of sarcoplasmic reticulum. Proc Natl Acad Sci U S A 71(9):3522–3526
Spassova MA, Soboloff J, He LP, Xu W, Dziadek MA, Gill DL (2006) STIM1 has a plasma membrane role in the activation of store-operated Ca2+ channels. Proc Natl Acad Sci U S A 103(11):4040–4045
Stathopulos PB, Li GY, Plevin MJ, Ames JB, Ikura M (2006) Stored Ca2+ depletion-induced oligomerization of stromal interaction molecule 1 (STIM1) via the EF-SAM region: an initiation mechanism for capacitive Ca2+ entry. J Biol Chem 281(47):35855–35862
Stathopulos PB, Zheng L, Li GY, Plevin MJ, Ikura M (2008) Structural and mechanistic insights into STIM1-mediated initiation of store-operated calcium entry. Cell 135(1):110–122
Stiber J, Hawkins A, Zhang ZS, Wang S, Burch J, Graham V, Ward CC, Seth M, Finch E, Malouf N, Williams RS, Eu JP, Rosenberg P (2008) STIM1 signalling controls store-operated calcium entry required for development and contractile function in skeletal muscle. Nat Cell Biol 10(6):688–697
Vig M, DeHaven WI, Bird GS, Billingsley JM, Wang H, Rao PE, Hutchings AB, Jouvin MH, Putney JW, Kinet JP (2008) Defective mast cell effector functions in mice lacking the CRACM1 pore subunit of store-operated calcium release-activated calcium channels. Nat Immunol 9(1):89–96
Voermans NC, Laan AE, Oosterhof A, van Kuppevelt TH, Drost G, Lammens M, Kamsteeg EJ, Scotton C, Gualandi F, Guglielmi V, van den Heuvel L, Vattemi G, van Engelen BG (2012) Brody syndrome: a clinically heterogeneous entity distinct from Brody disease: a review of literature and a cross-sectional clinical study in 17 patients. Neuromuscul Disord 22(11):944–954
Wang Y, Deng X, Mancarella S, Hendron E, Eguchi S, Soboloff J, Tang XD, Gill DL (2010) The calcium store sensor, STIM1, reciprocally controls Orai and CaV1.2 channels. Science 330(6000):105–109
Williams RT, Manji SS, Parker NJ, Hancock MS, Van Stekelenburg L, Eid JP, Senior PV, Kazenwadel JS, Shandala T, Saint R, Smith PJ, Dziadek MA (2001) Identification and characterization of the STIM (stromal interaction molecule) gene family: coding for a novel class of transmembrane proteins. Biochem J 357(Pt 3):673–685
Woo JS, Cho CH, Lee KJ, Kim DH, Ma J, Lee EH (2012) Hypertrophy in skeletal myotubes induced by junctophilin-2 mutant, Y141H, involves an increase in store-operated Ca2+ entry via Orai1. J Biol Chem 287(18):14336–14348
Woo JS, Hwang JH, Ko JK, Weisleder N, Kim DH, Ma J, Lee EH (2010) S165F mutation of junctophilin 2 affects Ca2+ signalling in skeletal muscle. Biochem J 427(1):125–134
Woo JS, Kim DH, Allen PD, Lee EH (2008) TRPC3-interacting triadic proteins in skeletal muscle. Biochem J 411(2):399–405
Wu MM, Buchanan J, Luik RM, Lewis RS (2006) Ca2+ store depletion causes STIM1 to accumulate in ER regions closely associated with the plasma membrane. J Cell Biol 174(6):803–813
Wu MM, Luik RM, Lewis RS (2007) Some assembly required: constructing the elementary units of store-operated Ca2+ entry. Cell Calcium 42(2):163–172
Xu P, Lu J, Li Z, Yu X, Chen L, Xu T (2006) Aggregation of STIM1 underneath the plasma membrane induces clustering of Orai1. Biochem Biophys Res Commun 350(4):969–976
Yuan YP, Eulenstein O, Vingron M, Bork P (1998) Towards detection of orthologues in sequence databases. Bioinformatics 14(3):285–289
Yuan JP, Zeng W, Dorwart MR, Choi YJ, Worley PF, Muallem S (2009) SOAR and the polybasic STIM1 domains gate and regulate Orai channels. Nat Cell Biol 11(3):337–343
Zhang Y, Fujii J, Phillips MS, Chen HS, Karpati G, Yee WC, Schrank B, Cornblath DR, Boylan KB, MacLennan DH (1995) Characterization of cDNA and genomic DNA encoding SERCA1, the Ca2+-ATPase of human fast-twitch skeletal muscle sarcoplasmic reticulum, and its elimination as a candidate gene for Brody disease. Genomics 30(3):415–424
Zhang SL, Yu Y, Roos J, Kozak JA, Deerinck TJ, Ellisman MH, Stauderman KA, Cahalan MD (2005) STIM1 is a Ca2+ sensor that activates CRAC channels and migrates from the Ca2+ store to the plasma membrane. Nat Geosci 437(7060):902–905
Zheng L, Stathopulos PB, Schindl R, Li GY, Romanin C, Ikura M (2011) Auto-inhibitory role of the EF-SAM domain of STIM proteins in store-operated calcium entry. Proc Natl Acad Sci U S A 108(4):1337–1342
Zucchi R, Ronca-Testoni S (1997) The sarcoplasmic reticulum Ca2+ channel/ryanodine receptor: modulation by endogenous effectors, drugs and disease states. Pharmacol Rev 49(1):1–51
Acknowledgments
This work was supported by the Basic Science Research Program (2012-0007701 to E.H.L.) and by the Mid-career Researcher Program (2012-0005435 to E.H.L.) through NRF grants funded by the MEST.
Conflict of interest
The authors declare that they have no conflicts of interest.
Author information
Authors and Affiliations
Corresponding author
Additional information
Authors' contributions
Keon Jin Lee and Eun Hui Lee designed the experiments. Keon Jin Lee, Changdo Hyun, Jin Seok Woo, and Chang Sik Park conducted the experiments. Do Han Kim and Eun Hui Lee contributed to the data analysis and to the discussion of the results. Eun Hui Lee wrote the paper.
K.J. Lee and C. Hyun contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(PPTX 5.23 MB)
Rights and permissions
About this article
Cite this article
Lee, K.J., Hyun, C., Woo, J.S. et al. Stromal interaction molecule 1 (STIM1) regulates sarcoplasmic/endoplasmic reticulum Ca2+-ATPase 1a (SERCA1a) in skeletal muscle. Pflugers Arch - Eur J Physiol 466, 987–1001 (2014). https://doi.org/10.1007/s00424-013-1361-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00424-013-1361-6