Abstract
Obesity impairs reproductive capacity, and the link between imprinting disorders and obesity has been discussed in many studies. Recent studies indicate that a high-fat diet may cause epigenetic changes in maternal and paternal genes, which may be transmitted to offspring and negatively affect their development. On this basis, our study aims to reveal the changes in DNA methylation and DNA methyltransferase enzymes in the ovaries and testes of C57BL/6 mice fed a high-fat diet and created a model of obesity, by comparing them with the control group. For this purpose, we demonstrated the presence and quantitative differences of DNA methyltransferase 1 and DNA methyltransferase 3a enzymes as well as global DNA methylation in ovaries and testis of C57BL/6 mice fed a high-fat diet by using immunohistochemistry and western blot methods. We found that a high-fat diet induces the levels of Dnmt1 and Dnmt3a proteins (p < 0.05). We observed increased global DNA methylation in testes but, interestingly, decreased global DNA methylation in ovaries. We think that our outcomes have significant value to demonstrate the effects of obesity on ovarian follicle development and testicular spermatogenesis and may bring a new perspective to obesity-induced infertility treatments. Additionally, to the best of our knowledge, this is the first study to document dynamic alteration of Dnmt1 and Dnmt3a as well as global DNA methylation patterns during follicle development in healthy mouse ovaries.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Obesity is a metabolic syndrome that leads to an imbalance in energy expenditure and accumulation of excess fat in the body (Rao et al. 2014; Wang et al. 2019). In the past three decades, the dominance of obesity has steadily increased worldwide (Lainez and Coss 2019). According to the World Health Organization (WHO) 2016 report, 1.9 billion adults are overweight and 650 million people are obese (Maugeri 2020). These figures make overweight and obesity a public health priority (Kroes et al. 2016) and lead to obesity being considered an epidemic in the twenty-first century (Collaboration 2016). With the increase in the prevalence of obesity, the proportion of women and men of childbearing age who are overweight and obese has also increased (Catalano and Shankar 2017).
Recent studies revealed the negative effects of obesity on reproduction (Shah et al. 2011; Atzmon et al. 2017; Kasum et al. 2018). Obesity in men can cause hormonal imbalances related to the hypothalamic–pituitary–gonadal (HPG) axis, leading to chronic inflammation and increasing oxidative stress at the systemic and tissue levels. Testis and epididymis, important organs for normal spermatogenesis and sperm maturation, can be negatively affected by the hormone deficiency, inflammation and oxidative attack caused by obesity. Therefore, these perturbations lead to a decline in sperm quality, including decreased sperm concentration and vitality, insufficient lipid composition, increased DNA damage and abnormal epigenetic changes, which will eventually lead to male infertility and epigenetic health problems (Davidson et al. 2015; Fullston et al. 2015; Liu and Ding 2017). In addition, associated enhanced predisposition for hypoglycaemic agent resistance and metabolic disorders have been identified in youngsters with overweight fathers (Fullston et al. 2015; Potabattula et al. 2019). Similarly, female obesity is considered one of the factors for infertility by negatively affecting the endometrium (Rhee et al. 2016), ovaries (Cheong et al. 2014) and oocyte quality (Snider and Wood 2019). Obesity can alter the levels of follicle-stimulating hormone (FSH) and luteinizing hormone (LH) and leads to irregular menstruation (Mitchell et al. 2005). Moreover, children of obese mothers may also have developmental disorders, insulin resistance and metabolic disorders (Dag and Dilbaz 2015).
The aetiology of obesity is multifactorial and can be caused by a combination of various factors, including genetic predisposition, family history, socio-economic and socio-cultural conditions, sedentary lifestyle and excessive consumption of high-calorie foods that exceed calorie requirements. The current blueprint for the obesity process is that not everyone develops the disease under the same environmental factors, suggesting that genetics may play a role (Sultan et al. 2020). However, the genetic risk locus determined as a result of a genome-wide association study explains only a part of the genetic variability of disease risk, which indicates that more factors need to be considered to elucidate the multifactorial pathology of obesity (Campion et al. 2009). Recently, epigenetics has provided a new explanation for the mechanism by which environmental factors change the expression of genes related to obesity and related diseases (Huang et al. 2018). In addition, the link between epigenetic changes and human metabolic health has been discovered in recent years (van Dijk et al. 2015; Tzika et al. 2018).
Epigenetics describes changes in gene expression that can be inherited without any change in DNA sequence. One of the epigenetic mechanisms is DNA methylation, which plays important roles in the inhibition or activation of transcription, X-chromosome inactivation, cell differentiation and tumourigenesis (Uysal et al. 2015). DNA methylation, which is specifically catalysed by different types of DNA methyltransferases (DNMTs), consists of two different mechanisms as the maintenance process and the de novo process. Maintaining methylation sustains the DNA methylation pattern previously established at the end of each DNA cycle. Conversely, the de novo methylation process generates new DNA methylation signatures in genomic DNA. DNMT, which adds a methyl group to the fifth carbon of the cytosine residue, usually on cytosine-phosphate-guanine (CpG), in rare cases, to non-CpG islands such as cytosine-phosphate-thymine (CpT), cytosine-phosphate-adenine (CpA), and cytosine-phosphate-cytosine (CpC) by using S-adenosyl methionine (AdoMet) as a methyl donor. So far, six different DNMTs have been characterized in structure and function in mammals: DNMT1, DNMT2, DNMT3A, DNMT3B, DNMT3C and DNMT3L (Okano et al. 1999; Uysal et al. 2016; You et al. 2017). The function of DNMT1 is to methylate the hemi-methylated strand of DNA during replication. Additionally, DNMT1 participates in de novo methylation. It was demonstrated that deletion of the Dnmt1 (Dnmt1 −/−) gene can lead to embryonic death, loss of imprinting and overall demethylation of the genome (Okano et al. 1998; Yoder et al. 1997; Kurihara et al. 2008). DNMT3A plays a key role in the de novo methylation of unmethylated DNA strands and creates new repressions. The deletion of the Dnmt3a gene results in embryo lethality and loss of imprints in gametes. DNA methylation is concerned within the management of certain gene expressions throughout oogenesis and spermatogenesis. Global DNA methylation during oogenesis increases (Uysal and Ozturk 2017; Uysal et al. 2015). Dnmt1 and Dnmt3a macromolecules are well known to be expressed in ovaries (Uysal and Ozturk 2017). Moreover, we very recently demonstrated differential alteration of global methylation and the patterns of Dnmt1 and Dnmt3a enzymes in testis focusing on spermatogenesis and Sertoli cells (Uysal et al. 2022). Although it is known that obesity causes epigenetic changes, the effects of obesity on Dnmt expression and therefore global DNA methylation in gonads have not been addressed so far.
In this study, we aimed to evaluate the effects of a high-fat diet (HFD)-induced obesity on Dnmt1 and Dnmt3a expression and global DNA methylation in the ovaries and testes in a mouse model. Briefly, the results demonstrated that HFD differentially alters the levels of Dnmt and global DNA methylation in developing follicles in ovaries and spermatogenesis in testes.
Materials and methods
Animals
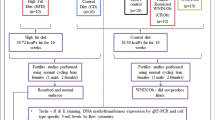
The experimental protocol was approved by the Animal Care and Usage Committee of Ankara University (protocol no. 2020-17-145). C57BL6/J mice (4 weeks old) were housed in cages under a 12-h/12-h light/dark cycle with an ambient temperature of 22–25 °C. After the female (n = 8) and male (n = 8) mice were randomly allocated to the appropriate cages, they were fed either a normal diet (Research Diets, D12450J) consisting of 20% protein, 10% fat and 70% carbohydrate or an HFD (Research Diets, D12492) consisting of 20% protein, 60% fat and 20% carbohydrate for 12 weeks. Body weights were measured weekly. At the end of 12 weeks, animals were euthanized, and the isolated gonads were transferred to buffered solutions.
Immunohistochemistry
The immunoexpression of the Dnmt1 and Dnmt3a proteins, as well as 5-methylcytosine (5mC) staining in the ovaries and testes, was evaluated by using immunohistochemistry. The paraffin sections of the gonads at 5 µm thickness were deparaffinized in fresh xylene and then rehydrated in a series of decreasing ethanol concentrations. After deparaffinization and rehydration processes, citrate buffer (pH 6.0) was used for antigen retrieval using microwave (2 × 5 min at 750 W). The endogenous peroxidase activity in the sections was blocked by 3% hydrogen peroxide for 30 min at room temperature (RT). Following several washes with 1× phosphate-buffered saline (PBS, Sigma-Aldrich, P44417), sections were blocked with blocking buffer (Thermo Scientific, UK) at RT for 7 min to prevent non-specific binding. Then, sections were incubated with the following primary antibodies: Dnmt1 (1:100; catalogue no. ab87654, Abcam), Dnmt3a (1:200; catalogue no. ab23565, Abcam) and 5mC (1:200; catalogue no. 28692S, Cell Signaling) at +4 °C overnight. One of three sections on each slide was incubated with antibody isotype (catalogue no. ab172730, Abcam) for testing primary antibody specificity. For negative control, slides were incubated in PBS without any primary antibody for each group. After that, sections were washed three times in PBS for 15 min each and subsequently incubated with anti-rabbit IgG secondary antibody (1:500; catalogue no. BA-1000, Vector) at RT for 1 h. Finally, sections were incubated with streptavidin–horseradish peroxidase (HRP) complex (catalogue no. TS-125, Thermo Scientific) for 20 min at RT. The immune reactions were visualized using 3,3′-diaminobenzidine (DAB) chromogen (catalogue no. D4168, Sigma-Aldrich) under a light microscope. Sections were washed in tap water and counterstained with Mayer’s haematoxylin (not for 5mC). Slides were examined and images were obtained with a light microscope (ZEISS Scope.A1, Zeiss, Germany) at 20× (Zeiss, A-Plan 20×/0.45 Working Distance (W.D.) = 0.46 M27) and 40× (Zeiss, N-Achroplan 40×/0.65 W.D. = 0.60 M27) magnifications. Images were analysed and captured with AxioCam camera (Zeiss, AxioCam MRc5) and AxioVision image software (Zeiss AxioVision v. 4.8). Their dimensions were at 1292 × 968 (width × height) pixels, and each of them has a depth of 24 bits. Also, all micrographs were evaluated for the related expression using the software and presented a resolution of 300 dpi in the panels. We measured signal intensities of the images for total protein analysis of the whole section, whereas we applied H-score analysis for specific region and cell type evaluations. For intensity analysis, signals obtained from the images were digitized and intensities measured using ImageJ (NIH, USA) software. Relative expression levels were calculated with the ratio of digitized signal intensity of experiment group to that of the negative-control group (as background signal). Ovarian follicles (primordial stage, single-layer primary follicle, multiple-layered primary follicle, early secondary and late secondary, as well as corpus luteum) and testicular cells (spermatogonium, spermatocytes, round spermatids, elongated spermatids and Sertoli cells) were evaluated by H-score. Briefly, sections were evaluated using a numerical score based on the number of positive cells in the section (five different fields in each section). H-score was calculated using the following equation: H-score = ∑Pi (i); where i is the intensity of staining with a value of 1, 2, 3, 4 or 5 (very weak staining, weak but visible, significant staining, intense staining and very intense staining, respectively), Pi is the percentage of stained related cells for each intensity varying from 0% to 100%.
Western blotting
Semi-quantitative analysis of Dnmt1 and Dnmt3a was performed with the western blotting (WB) technique. First, we analysed each sample individually, then we pooled the proteins from them and presented the data. For this purpose, tissue samples in lysis buffer [1% sodium dodecyl sulphate (SDS), 1 mM sodium ortho-vanadate, 10 mM Tris pH 7.4] supplemented with 1× protease inhibitor cocktail (Takara, 635672, CA, USA) were mechanically homogenized using a Bandelin Sonopuls homogenizer. The protein concentration was measured using the bicinchoninic acid (BCA) method. Fifty micrograms of protein from each group were loaded on each lane of 10%Tris–HCl gel which was used for protein electrophoresis (Bio-Rad, CA, USA), then electro-transferred to a polyvinylidene difluoride (PVDF) membrane (Bio-Rad, 162-0115, CA, USA) overnight at +4 °C. The membrane was blocked with 5% (w/v) BSA (Sigma-Aldrich, A2153, Saint Louis, USA) prepared in TBS-T (20 mM Tris–HCl and 150 mM NaCl plus 0.05% Tween-20 at pH 7.4) at RT for 1 h, and then were incubated with primary antibodies specific to Dnmt1, Dnmt3a or β-actin (Abcam, ab8227) [1:1000 in 5% (w/v) BSA containing TBS-T] for 2 h at RT. Following a triple-wash in TBS-T for 15 min each, membranes were incubated with IRDye 800CW Goat anti-Rabbit IgG Secondary Antibody (1:2000 in TBS-T) (Licor Biosciences, 926–32211, USA) at RT for 1 h on a shaker. Protein band intensities were measured using a Li-Cor Odyssey CLx infrared detection system (LICOR Biosciences) following the manufacturer’s instructions. Relative band intensities were measured and analysed using ImageJ v.3.91 software. Relative expression levels were calculated as the ratio of digitized signal intensity of experiment group to that of the negative-control group (as background signal).
Statistical analysis
Signal intensities of immunohistochemistry and WB images were digitalized with ImageJ software. Data were transferred to SigmaStat software (version 3.5, Jandel Scientific Corp) for statistical analysis. The Shapiro–Wilk test for normality was used to test whether the data were normally distributed. Logarithmic transformation was applied to non-normally distributed data, when applicable. Normally distributed data were presented as mean ± standard deviation (SD) and analysed by one-way analysis of variance (ANOVA) to examine the differences between the groups, and then a post hoc Tukey’s test was applied. The significance level was set at p < 0.05.
Results
Body weight measurements
C57BL6/J females and males were randomly assigned to the appropriate diet groups, then fed a normal diet (ND) or a high-fat diet (HFD) for 12 weeks. No significant differences were observed in food intake and water consumption between the groups. At the end of 12 weeks, the HFD group showed significant weight gain compared with the ND group (Fig. 1a), and the weight difference between HFD and ND was 14.85 ± 1.79 g for females and 11.4 ± 0.89 g for males (Fig. 1b).
Establishment of an obese mouse model. a Representative pictures of 16-week-old normal diet (ND) and high-fat diet (HFD) mice. b Comparison of time-dependent increase in body weight between ND and HFD males and females. Significant differences in body weight were first noted at the fourth week of diet in males (black arrowhead) and sixth week of diet in females (white arrowhead)
Effect of HFD on testis
Dnmt1 and Dnmt3a expression
Total signals of Dnmt1 and Dnmt3a from each seminiferous tubule section were measured. Significantly increased Dnmt1 and Dnmt3a levels in the HFD group were observed compared with ND (p < 0.05, Fig. 2a–c) when the staining pattern of the whole tubule was taken into consideration. Similarly, WB analyses of total tubules showed a significant increase in Dnmt1 and Dnmt3a levels in the HFD group compared with ND (p < 0.001, Fig. 2d–f).
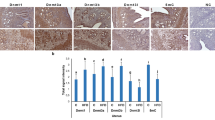
Effect of HFD on Dnmt1 and Dnmt3a expression in mouse testis. Demonstrative images for immunoexpression (a) and relative expression levels in total testis tissues (b, c). Western blotting (WB) (d) and relative signal intensity of Dnmt1 and Dnmt3a to actin from WB (e, f). ND normal diet, HFD high-fat diet, NC negative control (scale bar: 50 µm)
When cellular localization and relative expression levels of Dnmt1 at each stage of spermatogenesis as spermatogonium (Sg), spermatocyte (Sct), round spermatid (RSt), elongated spermatid (ESt) and Sertoli cells (SC) were evaluated, Dnmt1 staining was observed in both cytoplasm and nucleus of all cells but the ESt in both ND and HFD groups. We did not observe any staining in ESt.
When the two groups were compared, intensities of Dnmt1 signals in Sg and SC were found to be comparable, whereas a significant increase was noted in Sct and RSt of the HFD group compared with control (p < 0.05) (Fig. 3a and b).
Effect of HFD on Dnmt1 and Dnmt3a cellular localization in mouse testis. Demonstrative images for immunoexpression (a) and relative expression levels during spermatogenesis and Sertoli cells (b, c). Different letters indicate significant differences as p < 0.05. ND normal diet, HFD high-fat diet, NC negative control, Sg spermatogonium, Sct spermatocyte, RSt round spermatid, Est elongated spermatid, SC Sertoli cells (scale bar: 50 µm)
A very similar staining pattern was observed when Dnmt3a was analysed (Fig. 3a, c).
General methylation status
Global DNA methylation indicator 5mC staining showed nuclear localization. Total signal analysis from each seminiferous tubule section revealed that the intensity of 5mC staining significantly decreased (p < 0.001) in the HFD group (Fig. 4a, b). Staining intensity of 5mC was comparable in Sg, Sct and SC, whereas a significant decrease (p < 0.05) was found in RSt and ESt in the HFD group (Fig. 4c). Notably, a remarkable decrease of 5mC signal (p < 0.001) was found in ESt from the HFD group compared with ND (Fig. 4c).
Effect of HFD on global DNA methylation with 5mC staining in mouse testis. Representative images for immunoexpression (a), relative global DNA methylation levels in total tissue sections (b), and during spermatogenesis and Sertoli cells (c). Different letters indicate significant differences as p < 0.05. ND normal diet, HFD high-fat diet, NC negative control, Sg spermatogonium, Sct spermatocyte, RSt round spermatid, Est elongated spermatid, SC Sertoli cells (scale bar: 50 µm)
Effect of HFD on ovaries
Dnmt1 and Dnmt3a expression
Expression levels of Dnmt1 and Dnmt3a signals in total ovary sections of the HFD group were found to be significantly higher than those of the ND group (Fig. 5a–c; p < 0.05). Western blot analysis supported these findings (Fig. 5d–f).
Effect of HFD on Dnmt1 and Dnmt3a expression in the mouse ovary. Demonstrative images for immunoexpression (a) and relative expression levels in total ovary tissues (b, c). Western blotting (WB) (d) and relative signal intensity of Dnmt1 and Dnmt3a to actin from WB (e, f). ND normal diet, HFD high-fat diet, NC negative control (scale bar: 50 µm)
Dnmt1 and Dnmt3a signals from each stage of follicle development, as primordial stage (Pr), single-layered primary follicle (SP), multiple-layered primary follicle (MP), early secondary (ES) and late secondary (LS), as well as corpus luteum (CL) were analysed. Dnmt1 and Dnmt3a positivity was observed in all cells in the follicle. Dnmt1 was found in the cytoplasm and nucleus, whereas Dnmt3a was localized to the cytoplasm. Signal intensities of Dnmt1 and Dnmt3a were significantly lower in the Pr stage, and gradually increased during follicle development in both ND and HFD groups (Fig. 6a, b, d). Signal levels of both enzymes were significantly higher in secondary stages compared with primary ones. We did not find significant differences between SP and MP, or among ES, LS and CL.
Effect of HFD on Dnmt1 and Dnmt3a expression in different stages of follicles in the mouse ovary. Representative images for immunoexpression (a) and relative expression levels in developing follicles and corpus luteum (Pr primordial stage, SP single-layered primary follicle, MP multiple-layered primary follicle, ES early secondary, LS late secondary, CL corpus luteum) (b, d), and follicle cells (O oocyte, GC granulosa cell, SC stromal cell) (c, e). Different letters indicate p < 0.05. ND normal diet, HFD high-fat diet, NC negative control (scale bar: 50 µm)
When ND and HFD groups were compared, Dnmt1 and Dnmt3a levels from each stage of follicle development of the HFD group were significantly higher than those of the ND group (Fig. 6a, b and d; p < 0.05).
Moreover, we analysed Dnmt1 and Dnmt3a signal intensities in follicle cells, oocytes (O), granulosa cells (GC) and stromal cells (SC) individually, in each stage of follicle development (Fig. 6c and e). Dnmt1 signal intensity in oocytes was comparable between ND and HFD during follicle development, but it was significantly higher in GCs of the HFD group at all stages of folliculogenesis. Interestingly, significantly higher levels of Dnmt1 signals from SCs of the HFD group were noted at primary-stage follicles (p < 0.05), while no difference was observed in SCs between ND and HFD at the other stages of follicle development (Fig. 6c).
Dnmt3a levels in oocytes were significantly higher in the HFD group at all stages of folliculogenesis. When GCs were considered, the Dnmt3a signal was comparable in primordial follicles, while a significant increase was observed in subsequent follicle stages in the HFD group. The increase in Dnmt3a levels in stromal cells from the HFD group did not reach significant levels during follicle development, except at ES stage (Fig. 6e; p < 0.05).
General methylation status
Total ovarian assessments revealed that the intensity of 5mC staining significantly increased (p < 0.05) in the HFD group (Fig. 7), in a parallel manner to Dnmt1 and Dnmt3a (Fig. 5a and c).
Effect of HFD on global DNA methylation with 5mC staining in mouse ovaries. Representative images for total immunoexpression (a) and immunoexpression of each follicle type (b). Relative expression levels in total ovarian tissues (c), in developing follicles and corpus luteum (Pr primordial stage, SP single-layered primary follicle, MP multiple-layered primary follicle, ES early secondary, LS late secondary, CL corpus luteum) (d) and follicle cells (O oocyte, GC granulosa cell, SC stromal cell) (e). Different letters indicate p < 0.05. ND normal diet, HFD high-fat diet, NC negative control (scale bar: 50 µm)
Staining intensities of 5mC during follicle development were analysed. Signal levels of 5mC in primordial and single-layered follicle stages were significantly higher than in the subsequent stages in both ND and HFD groups. Compared with ND, signal intensities of each stage of follicle development in the HFD group were significantly higher (Fig. 7b and d; p < 0.05).
Signal intensities of 5mC were exclusively analysed in oocytes (O), granulosa cells (GC) and stromal cells (SC) in each stage of follicle development, separately (Fig. 7e). 5mC levels in oocytes gradually decreased from the primordial to the late secondary follicle in both ND and HFD groups. Signal intensities of 5mC in oocytes from HFD were significantly higher than those of ND in primordial and SP stages (p < 0.05), whereas they were comparable in the following stages. Considering GCs, we noted that 5mC levels were highest in the Pr stage, and significantly decreased during follicle development in both ND and HFD groups. When two diet regime groups were compared, 5mC levels were higher in GCs of Pr to secondary follicle stages of the HFD group than in those of the ND, while we did not observe a significant difference at secondary stages. 5mC levels in SCs from the HFD group were higher than those of the ND group in all follicle development stages, except SP stage, which showed comparable 5mC levels (Fig. 7e; p < 0.05).
Discussion
Obesity is one of the most common diseases, and negatively impacts male and female reproductive systems (Wang et al. 2019; Barbagallo et al. 2021; Practice Committee of the American Society for Reproductive Medicine. Electronic address and Practice Committee of the American Society for Reproductive 2021). A strong relationship between epigenetic alterations and various disease has been shown (Robertson 2005). Studies focusing on the epigenetic inheritance of obesity have demonstrated that a high-fat diet may cause epigenetic changes in both maternal and paternal genes, and therefore, these changes may be transmitted to the offspring by affecting their development negatively (Ooi and Henikoff 2007; Ozanne 2015).
Gametogenesis is characterized by intense changes in DNA methylation. De novo methylation occurs from globally demethylated primordial germ cells to mature gametes (Robertson 2005). Cellular localization and expression levels of Dnmt1 and Dnmt3a proteins in testis have been demonstrated in previous studies (Marques et al. 2011; Uysal et al. 2019). Omisanjo and colleagues analysed the relationship between Dnmt1 and spermatic arrest or testicular cancer and found that Dnmt1 mRNA and protein were localized in both the nucleus and cytoplasm of spermatogonia and round spermatid. Interestingly, in round spermatid maturation arrest cases, they observed mRNA and protein in spermatocytes but not in spermatogonia (Omisanjo et al. 2007). In our study, Dnmt1 levels of spermatogonia were comparable in ND and HFD groups, but a significant increase was observed at the spermatocyte and round spermatid stage. This alteration may be explained by the effect of obesity on sperm maturation. Although a similar pattern was noted for Dnmt3a, we did not observe any difference in global methylation at spermatogonia and spermatocyte stages, but a significant decrease at round and elongated stages in the HFD group, counter to the expectation that increased Dnmts would induce global methylation. We think that other Dnmts or epigenetic mechanisms such as histone deacetylases (HDACs) or histone methyltransferases (HMTs) may differentially regulate global methylation. Moreover, the finding that there are no Dnmt1 and Dnmt3a proteins but high global methylation in elongated spermatids may support this assumption. Similarly, Fullston and colleagues showed decreased DNA methylation in mouse total testes and elongating spermatids in the HFD group, in line with our findings (Fullston et al. 2013). Recently, Deshpande et al. found a significant decrease in the Dnmt1 transcript levels in the testes of rats fed HFD compared with the ND group, whereas Dnmt3a, Dnmt3b and Dnmt3l were comparable (Deshpande et al. 2021). They also noted a decrease in 5mC levels in spermatids, in line with our outcomes. However, they did not observe any change in 5mC levels, but Dnmt3a, Dnmt3b and Dnmt3l levels were upregulated in genetically obese rats.
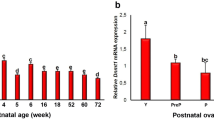
In the current study, we found that both Dnmt1 and Dnmt3a levels and global methylation increased in the ovary in the HFD group. Consistently, the levels of Dnmts and global methylation were high in the HFD group at all stages of follicular development. It is known that developing oocytes, from primordial germ cell (PGC) to mature (MII) oocyte, exhibit unique epigenetic and transcriptional changes (Pan et al. 2012). Genome-wide DNA demethylation occurs in PGCs just after arriving at the genital ridge. Kageyama et al. showed that the level of DNA methylation increased in pre-pubertal oocytes from day 5 to day 15 (Kageyama et al. 2007). During the GV stage to MII maturation, Dnmt1, Dnmt3a, Dnmt3b and Dnmt3l levels increase (Lucifero et al. 2004; Uysal et al. 2017), but interestingly, global DNA methylation decreases (Lucifero et al. 2004) despite some opposite findings (Dean et al. 2003; Uysal and Ozturk 2017). The dynamics of Dnmts and global DNA methylation during follicle development have not been demonstrated before. To the best of our knowledge, this is the first study to document dynamic alteration of Dnmt1 and Dnmt3a as well as global DNA methylation pattern in total follicles and oocytes, granulosa and stromal cells individually during ovarian follicle development not only in an HFD group but also in a ND group.
It was shown that HFD induces ovarian follicle development and, thus, the rate of follicle loss via different signalling pathways and reduces reproductive capacity (Wang et al. 2014; Sohrabi et al. 2015). DNA methylation also plays an important role in the expression regulation of genes in oocyte and granulosa cells during folliculogenesis as in ovarian cells (Turek-Plewa and Jagodzinski 2005). Our results demonstrated that HFD significantly induces Dnmt1 and Dnmt3a levels and global DNA methylation during folliculogenesis. Here, we think that differential alteration of follicular cells induced by HFD may cause early follicle loss and diminish reproductive capacity.
Conclusion
In this study, the effect of a high-fat diet on Dnmt1 and Dnmt3a expression as well as global DNA methylation in mouse gonads was demonstrated for the first time. Our results showed that the high-fat diet significantly altered Dnmt enzymes and global methylation in mouse gonads, and increased the levels of Dnmt1 and Dnmt3a proteins in the ovary and testis. We observed increased global DNA methylation in testes but, interestingly, decreased global DNA methylation in ovaries. This discrepancy raises new questions to be answered. How do these epigenetic changes, transmitted from either father or mother or both, affect the offspring? What are the molecular pathways that may affect or be affected by Dnmt-enzyme-related DNA methylation altered by HFD?
We think that the current study has a novel role to indicate the necessity of new studies focusing on obesity-induced gene expression patterns and their roles in embryogenesis and further development.
As Dnmt1 and Dnmt3a are considered the most prominent DNA methylating enzymes that we analysed, new studies focusing on the effect of other Dnmts are required. Additionally, this is the first time that dynamic alteration of Dnmt1 and Dnmt3a and global methylation were demonstrated in normal follicle development. Our results indicate the necessity of new studies aiming to document the relationship between DNA methylation and follicle development, in detail.
Code availability
Not applicable.
Data availability
Data and material available upon request.
References
Atzmon Y, Shoshan-Karchovsky E, Michaeli M, Aslih N, Shrem G, Ellenbogen A, Shalom-Paz E (2017) Obesity results with smaller oocyte in in vitro fertilization/intracytoplasmic sperm injection cycles—a prospective study. J Assist Reprod Genet 34(9):1145–1151. https://doi.org/10.1007/s10815-017-0975-6
Barbagallo F, Condorelli RA, Mongioi LM, Cannarella R, Cimino L, Magagnini MC, Crafa A, La Vignera S, Calogero AE (2021) Molecular mechanisms underlying the relationship between obesity and male infertility. Metabolites 11(12):840. https://doi.org/10.3390/metabo11120840
Campion J, Milagro FI, Martinez JA (2009) Individuality and epigenetics in obesity. Obes Rev 10(4):383–392. https://doi.org/10.1111/j.1467-789X.2009.00595.x
Catalano PM, Shankar K (2017) Obesity and pregnancy: mechanisms of short term and long term adverse consequences for mother and child. BMJ 356:j1. https://doi.org/10.1136/bmj.j1
Cheong Y, Sadek KH, Bruce KD, Macklon N, Cagampang FR (2014) Diet-induced maternal obesity alters ovarian morphology and gene expression in the adult mouse offspring. Fertil Steril 102(3):899–907. https://doi.org/10.1016/j.fertnstert.2014.06.015
Collaboration NCDRF (2016) Trends in adult body-mass index in 200 countries from 1975 to 2014: a pooled analysis of 1698 population-based measurement studies with 19.2 million participants. Lancet 387(10026):1377–1396. https://doi.org/10.1016/S0140-6736(16)30054-X
Dag ZO, Dilbaz B (2015) Impact of obesity on infertility in women. J Turk Ger Gynecol Assoc 16(2):111–117. https://doi.org/10.5152/jtgga.2015.15232
Davidson LM, Millar K, Jones C, Fatum M, Coward K (2015) Deleterious effects of obesity upon the hormonal and molecular mechanisms controlling spermatogenesis and male fertility. Hum Fertil (Camb) 18(3):184–193. https://doi.org/10.3109/14647273.2015.1070438
Dean W, Santos F, Reik W (2003) Epigenetic reprogramming in early mammalian development and following somatic nuclear transfer. Semin Cell Dev Biol 14(1):93–100. https://doi.org/10.1016/s1084-9521(02)00141-6
Deshpande SSS, Nemani H, Balasinor NH (2021) High fat diet-induced- and genetically inherited- obesity differential alters DNA demethylation pathways in the germline of adult male rats. Reprod Biol 21(3):100532. https://doi.org/10.1016/j.repbio.2021.100532
Fullston T, Ohlsson Teague EM, Palmer NO, DeBlasio MJ, Mitchell M, Corbett M, Print CG, Owens JA, Lane M (2013) Paternal obesity initiates metabolic disturbances in two generations of mice with incomplete penetrance to the F2 generation and alters the transcriptional profile of testis and sperm microRNA content. FASEB J 27(10):4226–4243. https://doi.org/10.1096/fj.12-224048
Fullston T, McPherson NO, Owens JA, Kang WX, Sandeman LY, Lane M (2015) Paternal obesity induces metabolic and sperm disturbances in male offspring that are exacerbated by their exposure to an obesogenic diet. Physiol Rep 3(3):e12336. https://doi.org/10.14814/phy2.12336
Huang Q, Ma C, Chen L, Luo D, Chen R, Liang F (2018) Mechanistic insights into the interaction between transcription factors and epigenetic modifications and the contribution to the development of obesity. Front Endocrinol (Lausanne) 9:370. https://doi.org/10.3389/fendo.2018.00370
Kageyama S, Liu H, Kaneko N, Ooga M, Nagata M, Aoki F (2007) Alterations in epigenetic modifications during oocyte growth in mice. Reproduction 133(1):85–94. https://doi.org/10.1530/REP-06-0025
Kasum M, Oreskovic S, Cehic E, Lila A, Ejubovic E, Soldo D (2018) The role of female obesity on in vitro fertilization outcomes. Gynecol Endocrinol 34(3):184–188. https://doi.org/10.1080/09513590.2017.1391209
Kroes M, Osei-Assibey G, Baker-Searle R, Huang J (2016) Impact of weight change on quality of life in adults with overweight/obesity in the United States: a systematic review. Curr Med Res Opin 32(3):485–508. https://doi.org/10.1185/03007995.2015.1128403
Kurihara Y, Kawamura Y, Uchijima Y, Amamo T, Kobayashi H, Asano T, Kurihara H (2008) Maintenance of genomic methylation patterns during preimplantation development requires the somatic form of DNA methyltransferase 1. Dev Biol 313(1):335–346. https://doi.org/10.1016/j.ydbio.2007.10.033
Lainez NM, Coss D (2019) Obesity, neuroinflammation, and reproductive function. Endocrinology 160(11):2719–2736. https://doi.org/10.1210/en.2019-00487
Liu Y, Ding Z (2017) Obesity, a serious etiologic factor for male subfertility in modern society. Reproduction 154(4):R123–R131. https://doi.org/10.1530/REP-17-0161
Lucifero D, Mann MR, Bartolomei MS, Trasler JM (2004) Gene-specific timing and epigenetic memory in oocyte imprinting. Hum Mol Genet 13(8):839–849. https://doi.org/10.1093/hmg/ddh104
Marques CJ, Joao Pinho M, Carvalho F, Bieche I, Barros A, Sousa M (2011) DNA methylation imprinting marks and DNA methyltransferase expression in human spermatogenic cell stages. Epigenetics 6(11):1354–1361. https://doi.org/10.4161/epi.6.11.17993
Maugeri A (2020) The effects of dietary interventions on DNA methylation: implications for obesity management. Int J Mol Sci 21(22):8670. https://doi.org/10.3390/ijms21228670
Mitchell M, Armstrong DT, Robker RL, Norman RJ (2005) Adipokines: implications for female fertility and obesity. Reproduction 130(5):583–597. https://doi.org/10.1530/rep.1.00521
Okano M, Xie S, Li E (1998) Cloning and characterization of a family of novel mammalian DNA (cytosine-5) methyltransferases. Nat Genet 19(3):219–220. https://doi.org/10.1038/890
Okano M, Bell DW, Haber DA, Li E (1999) DNA methyltransferases Dnmt3a and Dnmt3b are essential for de novo methylation and mammalian development. Cell 99(3):247–257. https://doi.org/10.1016/s0092-8674(00)81656-6
Omisanjo OA, Biermann K, Hartmann S, Heukamp LC, Sonnack V, Hild A, Brehm R, Bergmann M, Weidner W, Steger K (2007) DNMT1 and HDAC1 gene expression in impaired spermatogenesis and testicular cancer. Histochem Cell Biol 127(2):175–181. https://doi.org/10.1007/s00418-006-0234-x
Ooi SL, Henikoff S (2007) Germline histone dynamics and epigenetics. Curr Opin Cell Biol 19(3):257–265. https://doi.org/10.1016/j.ceb.2007.04.015
Ozanne SE (2015) Epigenetic signatures of obesity. N Engl J Med 372(10):973–974. https://doi.org/10.1056/NEJMcibr1414707
Pan Z, Zhang J, Li Q, Li Y, Shi F, Xie Z, Liu H (2012) Current advances in epigenetic modification and alteration during mammalian ovarian folliculogenesis. J Genet Genom 39(3):111–123. https://doi.org/10.1016/j.jgg.2012.02.004
Potabattula R, Dittrich M, Schorsch M, Hahn T, Haaf T, El Hajj N (2019) Male obesity effects on sperm and next-generation cord blood DNA methylation. PLoS ONE 14(6):e0218615. https://doi.org/10.1371/journal.pone.0218615
Practice Committee of the American Society for Reproductive Medicine. Electronic address aao, Practice Committee of the American Society for Reproductive M (2021) Obesity and reproduction: a committee opinion. Fertil Steril 116(5):1266–1285. https://doi.org/10.1016/j.fertnstert.2021.08.018
Rao KR, Lal N, Giridharan NV (2014) Genetic & epigenetic approach to human obesity. Indian J Med Res 140(5):589–603
Rhee JS, Saben JL, Mayer AL, Schulte MB, Asghar Z, Stephens C, Chi MM, Moley KH (2016) Diet-induced obesity impairs endometrial stromal cell decidualization: a potential role for impaired autophagy. Hum Reprod 31(6):1315–1326. https://doi.org/10.1093/humrep/dew048
Robertson KD (2005) DNA methylation and human disease. Nat Rev Genet 6(8):597–610. https://doi.org/10.1038/nrg1655
Shah DK, Missmer SA, Berry KF, Racowsky C, Ginsburg ES (2011) Effect of obesity on oocyte and embryo quality in women undergoing in vitro fertilization. Obstet Gynecol 118(1):63–70. https://doi.org/10.1097/AOG.0b013e31821fd360
Snider AP, Wood JR (2019) Obesity induces ovarian inflammation and reduces oocyte quality. Reproduction 158(3):R79–R90. https://doi.org/10.1530/REP-18-0583
Sohrabi M, Roushandeh AM, Alizadeh Z, Vahidinia A, Vahabian M, Hosseini M (2015) Effect of a high fat diet on ovary morphology, in vitro development, in vitro fertilisation rate and oocyte quality in mice. Singap Med J 56(10):573–579. https://doi.org/10.11622/smedj.2015085
Sultan S, Patel AG, El-Hassani S, Whitelaw B, Leca BM, Vincent RP, le Roux CW, Rubino F, Aywlin SJB, Dimitriadis GK (2020) Male obesity associated gonadal dysfunction and the role of bariatric surgery. Front Endocrinol (Lausanne) 11:408. https://doi.org/10.3389/fendo.2020.00408
Turek-Plewa J, Jagodzinski PP (2005) The role of mammalian DNA methyltransferases in the regulation of gene expression. Cell Mol Biol Lett 10(4):631–647
Tzika E, Dreker T, Imhof A (2018) Epigenetics and metabolism in health and disease. Front Genet 9:361. https://doi.org/10.3389/fgene.2018.00361
Uysal F, Ozturk S (2017) DNA methyltransferases in mammalian oocytes. Results Probl Cell Differ 63:211–222. https://doi.org/10.1007/978-3-319-60855-6_10
Uysal F, Akkoyunlu G, Ozturk S (2015) Dynamic expression of DNA methyltransferases (DNMTs) in oocytes and early embryos. Biochimie 116:103–113. https://doi.org/10.1016/j.biochi.2015.06.019
Uysal F, Akkoyunlu G, Ozturk S (2016) DNA methyltransferases exhibit dynamic expression during spermatogenesis. Reprod Biomed Online 33(6):690–702. https://doi.org/10.1016/j.rbmo.2016.08.022
Uysal F, Ozturk S, Akkoyunlu G (2017) DNMT1, DNMT3A and DNMT3B proteins are differently expressed in mouse oocytes and early embryos. J Mol Histol 48(5–6):417–426. https://doi.org/10.1007/s10735-017-9739-y
Uysal F, Akkoyunlu G, Ozturk S (2019) Decreased expression of DNA methyltransferases in the testes of patients with non-obstructive azoospermia leads to changes in global DNA methylation levels. Reprod Fertil Dev. https://doi.org/10.1071/RD18246
Uysal F, Sukur G, Cinar O (2022) DNMT enzymes differentially alter global DNA methylation in a stage-dependent manner during spermatogenesis. Andrologia. https://doi.org/10.1111/and.14357
van Dijk SJ, Tellam RL, Morrison JL, Muhlhausler BS, Molloy PL (2015) Recent developments on the role of epigenetics in obesity and metabolic disease. Clin Epigenetics 7:66. https://doi.org/10.1186/s13148-015-0101-5
Wang N, Luo LL, Xu JJ, Xu MY, Zhang XM, Zhou XL, Liu WJ, Fu YC (2014) Obesity accelerates ovarian follicle development and follicle loss in rats. Metabolism 63(1):94–103. https://doi.org/10.1016/j.metabol.2013.09.001
Wang S, Pan MH, Hung WL, Tung YC, Ho CT (2019) From white to beige adipocytes: therapeutic potential of dietary molecules against obesity and their molecular mechanisms. Food Funct 10(3):1263–1279. https://doi.org/10.1039/c8fo02154f
Yoder JA, Soman NS, Verdine GL, Bestor TH (1997) DNA (cytosine-5)-methyltransferases in mouse cells and tissues. Studies with a mechanism-based probe. J Mol Biol 270(3):385–395. https://doi.org/10.1006/jmbi.1997.1125
You D, Nilsson E, Tenen DE, Lyubetskaya A, Lo JC, Jiang R, Deng J, Dawes BA, Vaag A, Ling C, Rosen ED, Kang S (2017) Dnmt3a is an epigenetic mediator of adipose insulin resistance. eLife. https://doi.org/10.7554/eLife.30766
Acknowledgements
We thank Prof. Erkan Yilmaz, who helped with animal diet regime, from Ankara University Biotechnology Institute.
Funding
This study was financially supported by TUBITAK 220S736.
Author information
Authors and Affiliations
Contributions
All authors qualify for authorship by contributing substantially to this article. G.S., F.U. and O.C. developed the original concept of this study. G.S. and F.U. collected data and performed statistical analyses, and G.S. and O.C. wrote the manuscript. All authors have contributed to critical discussion, reviewed the final version of the article and approved it for publication.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflicts of interest.
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Sukur, G., Uysal, F. & Cinar, O. High-fat diet induced obesity alters Dnmt1 and Dnmt3a levels and global DNA methylation in mouse ovary and testis. Histochem Cell Biol 159, 339–352 (2023). https://doi.org/10.1007/s00418-022-02173-2
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00418-022-02173-2