Abstract
Claudins are one of the transmembrane proteins found at tight junctions (TJs); they constitute the backbone of TJ strands and comprise a multigene family. Claudins share a YV sequence at the COOH-termini, which is considered to be a ZO-binding motif. Overexpression of claudin-15 (15CL) or claudin-15 tagged with enhanced green fluorescent protein at the NH2-terminus (EGFP-15CL) induced aberrant strands in MDCK II cells, even though claudin-15 has the ZO-binding motif. Morphometric analysis by freeze-fracture electron microscopy revealed that the mean number of apical TJ strands increased by 47% in EGFP-1CL-expressing cells, 21% in EGFP-15CL-expressing cells, and 28% in 15CL-expressing cells. The number of free-ended apical strands increased remarkably in EGFP-15CL- and 15CL-expressing cells, but not in EGFP-1CL-expressing cells. When MDCK cells expressing EGFP-1CL, EGFP-15CL or 15CL were co-cultured with parent MDCK cells, which express claudin-1 but not claudin-15, EGFP-15CL and 15CL could not be concentrated at the apical junctional region between the heterotypic cells, though EGFP-1CL could. These results suggest that not only binding to ZO-1, but also head-to-head compatibility between claudin species, is involved in organizing claudin proteins at the apical junctional region.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
A primary function of epithelial and endothelial monolayer sheets is to form barriers that separate organs into various compartments. Tight junctions (TJs) constitute a junctional complex that controls these barriers by regulating the movement of water, ions and proteins across the monolayer, often referred to as the paracellular permeability pathway (Gumbiner 1993). TJs also maintain cell polarity by acting as a fence that prevents the intermixing of proteins and lipids between the apical and basolateral membrane domains (Rodriguez-Boulan and Nelson 1989). By transmission electron microscopy, TJs appear as a series of very close membrane regions involving the outer leaflets of the plasma membrane of adjacent cells, the so-called “kissing points” (Farquhar and Palade 1963). In freeze-fracture replicas, the TJs appear as a continuous and branching network of particle strands encircling the apical end of the lateral membrane of epithelial cells (Staehelin 1974).
Many scaffolding and integral membrane proteins of TJs have been identified to date. Zonula occludens (ZO)-1, ZO-2 and ZO-3, which belong to a family of membrane-associated guanylate kinase (MAGUK) homologues, MAGI-1, MAGI-2, MAGI-3, MUPP-1, PAR-3, PAR-6, PALS and PATJ are known as scaffolding proteins (Schneeberger and Lynch 2004). Occludin, claudins, JAMs and CAR have been discovered as integral membrane proteins (Schneeberger and Lynch 2004). Among them, the backbones of TJ strands are now believed to be composed of claudins, each with a molecular mass of 20–28 kDa. Claudins have two extracellular loops, one intracellular loop, and NH2- and COOH-terminal cytoplasmic domains (Furuse et al. 1998). Claudins form a multigene family with ∼24 members in humans and mice, and each claudin shows a unique tissue expression pattern. In addition, more than two claudin species are usually expressed in a single cell and their combination and proportion vary among different cell types (Guan et al. 2005; Inai et al. 2005; Kiuchi-Saishin et al. 2002; Li et al. 2004; Morita et al. 1999; Rahner et al. 2001; Wilcox et al. 2001).
It has been proposed that TJ strands may be linear co-polymers composed of occludin and various claudins (Furuse et al. 1999; Tsukita and Furuse 1999) and that they probably present large linear clusters of YV sequences at the COOH-termini of most claudin members toward the cytoplasm. The PDZ1 domains at the NH2-terminal region of ZOs bind in vitro to the COOH-terminal YV sequences of most claudin members (Harris and Lim 2001; Itoh et al. 1999; Pawson and Nash 2003; Sheng and Sala 2001). The COOH-terminal regions of ZO-1 and ZO-2 bind actins (Wittchen et al. 1999). It has been hypothesized from these results that the linkage between claudins-ZOs-actin cytoskeleton at the perijunctional ring may stabilize TJs at the most apical region of the lateral plasma membrane. Consistent with this hypothesis, expression of claudin-1 tagged with a myc-epitope at its COOH-terminus induced aberrant TJ strands along the entire surfaces of lateral plasma membranes, suggesting that the myc-epitope interfered with binding between claudin-1 and ZO-1 and induced aberrant TJ strands (Kobayashi et al. 2002; McCarthy et al. 2000). However, the deletion mutant of claudins that lacked almost all of the COOH-terminal cytoplasmic domains still formed TJ strands in fibroblasts, suggesting that the interaction with peripheral membrane proteins did not seem to be required for the polymerization of claudins into TJ strands (Furuse et al. 1999).
Co-culture experiments of fibroblasts expressing one or two out of claudin-1, -2, and -3 demonstrated interactions within and between individual TJ strands (Furuse et al. 1999). First, different claudin species expressed in a single cell could co-polymerize into TJ strands via homomeric or heteromeric lateral interactions (side-by-side interaction) of claudins. Second, the TJ strands in one cell could interact with those in adjacent cells via homotypic or heterotypic head-to-head interactions, probably via extracellular loop-1 and/or -2 of claudins. However, TJ strands consisting of claudin-1 only could not form paired TJ strands with those consisting of claudin-2 only in fibroblasts (Furuse et al. 1999), suggesting that the extracellular loops of claudin-1 were not compatible with those of claudin-2. Incompatibility between claudin-1 and -5 was also reported (Coyne et al. 2003), but heterologous claudin incompatibility was not perfectly clarified.
When claudin-1 with a myc-epitope at COOH-terminus, referred to as 1CLmyc in this paper, was expressed in MDCK cells under the control of a doxycycline-regulated gene expression system, a high expression level of 1CLmyc induced aberrant TJ strands along the lateral plasma membranes, but low expression levels of 1CLmyc did not (Kobayashi et al. 2002). Although 1CLmyc expressed from a pCI-neo vector did not form aberrant strands (Inai et al. 1999), immunoblot analysis revealed that the expression level of 1CLmyc expressed from the pCI-neo vector was low (Kobayashi et al. 2002). These results suggest that a small amount of 1CLmyc can be localized at the apical junctional region, probably with the help of endogenous claudins via side-by-side and/or head-to-head interactions.
In this study, we examined whether claudins other than claudin-1 could form aberrant TJ strands when expressed in MDCK II cells that expressed at least claudin-1, -2, -3, -4 and -7 (Hou et al. 2006; Singh and Harris 2004). We selected claudin-15 because it has the ZO-1-binding motif (-YV) at the COOH-terminus (Itoh et al. 1999; Songyang et al. 1997), and the anti-claudin-15 antibody used in this study did not stain MDCK II Tet-Off cells. Claudin-15 was expressed in epithelial cells in the small intestine and colon, endothelial cells of the vasa recta in the kidney, and mesothelial cells of the peritoneum (Fujita et al. 2006; Inai et al. 2005; Kiuchi-Saishin et al. 2002). Expression of claudin-15, which was predicted to create a cation selective pore based on the pattern of its charged extracellular amino acids, increased transepithelial electrical resistance (TER) in MDCK II cells, but decreased TER in LLC-PK1 cells (Van Itallie et al. 2003). By comparing the results obtained from overexpression of EGFP-1CL, EGFP-15CL and 15CL in MDCK II cells, we discuss a possible mechanism of formation of aberrant TJ strands.
Materials and methods
Vector constructions
Human kidney total RNA (Clontech, Palo Alto, CA, USA) was reverse-transcribed into first strand cDNA using ReverTra Ace α Kit (TOYOBO, Osaka, Japan) and human claudin-15 was amplified by PCR using the primers, 5′-GAATTCATGTCGATGGCTGTGGAAACC-3′ and 5′-AAGTCGACCTACACGTAGGCGTTTCTGCCGTA-3′. Underlines indicate the recognition sites of restriction enzymes. Mouse claudin-1 was amplified by PCR using SK-1CLmyc (Inai et al. 1999) as a template and two oligonucleotides, 5′-GCGGCCGCATGGCCAACGCGGGGCTGCAG-3′ and 5′-ACTAGTTCACACATAGTCTTTCCCACT-3′, as primers. All PCR reactions were performed in a 25-μl reaction volume using Pfu Turbo (TOYOBO) DNA polymerase, and 10 μl of each product was examined in a 1.0% agarose gel containing ethidium bromide. Amplifications were performed in a T3 Thermocycler (Biometra, Göttingen, Germany) using the following program: 94°C for 60 s, 30 cycles of 94°C for 60 s, 55°C for 60 s and 72°C for 60 s, and, at last, 72°C for 10 min. As the amplification of claudin-15 was unsuccessful, the RT-PCR product was diluted to 1:100 and re-amplified by the same PCR condition. Although the re-amplified product contained three extra bands, the appropriate band (701 bp) was recovered from agarose gel and subcloned into a pCR-Blunt II-TOPO vector using a Zero Blunt TOPO PCR Cloning Kit (Invitrogen, Carlsbad, CA, USA), producing pCR-15CL. The enhanced green fluorescent protein (EGFP) sequence was amplified by PCR using a pBI-EGFP vector (Clontech) as a template and two oligonucleotides, 5′-GACCGCGGATGGTGAGCAAGGGCGAGGAG-3′ and 5′-ACGCGGCCGC TCTTGTACAGCTCGTCCATGCCG-3′, as primers. The reverse primer has one extra “T” shown in a bold letter between the Not I site and the COOH-terminal sequence of EGFP to prevent a frame shift of a gene to be inserted at the COOH-terminus of EGFP. The amplified product was digested with Sac II and Not I and ligated into SK (−) (Stratagene, La Jolla, CA), producing SK-EGFPC. The EGFP fragment excised by Sac II and Apa I from SK-EGFPC was ligated into pTREM (Kobayashi et al. 2002), producing pTREM-EGFPC, which had a multiple cloning site at the COOH-terminus of the EGFP sequence (239 amino acids) without a stop codon. The amplified claudin-1 fragment digested by Not I and Spe I and the claudin-15 fragment excised from pCR-15CL by Eco RI and Sal I were ligated into pTREM-EGFPC, producing pTREM-EGFP-1CL and pTREM-EGFP-15CL, respectively. When these vectors were introduced into MDCK II Tet-Off cells (Clontech), the stably transfected cells expressed EGFP-1CL (453 amino acids) and EGFP-15CL (482 amino acids) or 15CL (227 amino acids). EGFP-1CL had an additional 3 amino acids (SGR) between the EGFP and claudin-1 sequences, and EGFP-15CL had an additional 15 amino acids (SGRSRTSGSPGLQEF) between the EGFP and claudin-15 sequences. The claudin-15 fragment with a stop codon excised by Eco RI and Sal I from pTREM-EGFP-15CL was ligated into pTREM, producing pTREM-15CL. All sequences were confirmed by dideoxy sequencing.
Cell culture and transfections
Swiss mouse embryonic fibroblast (MEF)/3T3 Tet-Off cells and Madin-Darby canine kidney (MDCK) II Tet-Off cells (Clontech) were maintained in Dulbecco’s modified Eagle medium (DMEM) supplemented with 10% fetal bovine serum (FBS), 100 IU/ml penicillin, 100 μg/ml streptomycin and 100 μg/ml geneticin (Gibco BRL, Rockville, MD, USA). These cells were stably transfected with a pTet-Off regulator plasmid with resistance to geneticin (Jou and Nelson 1998). When Tet-Off cells were further transfected with a pTRE response plasmid containing a gene of interest, the product of the introduced gene is expressed within 12–24 h of removal of doxycycline (Dox) from the culture medium. MEF/3T3 or MDCK II Tet-Off cells plated onto 35 mm dishes were co-transfected by Trans IT-LT2 (PanVera, Madison, WI, USA) in Opti-MEM (Gibco BRL) with a mixture of pTREM-EGFP-1CL, pTREM-EGFP-15CL, or pTREM-15CL and pTK-Hyg providing resistance to hygromycin. After incubation for 4 h, the culture medium was removed and 2 ml of DMEM containing 10% FBS and 2 μg/ml of Dox was added. They were cultured for 36-48 h and then cells were replated onto four 10 cm dishes in the presence of 400 μg/ml hygromycin (Gibco BRL) to select stable transfectants. Induction of EGFP-1CL, EGFP-15CL or 15CL was achieved by the removal of Dox from the culture medium. Colonies were isolated and screened immunohistochemically using anti-claudin-15 antibody or the detection of EGFP 48 h after the removal of Dox. Positive clones were recloned by limiting dilution and stable cells were obtained.
Antibodies
The following antibodies were used: rabbit anti-claudin-1 polyclonal antibody (pAb), mouse anti-ZO-1 monoclonal antibody (mAb), mouse anti-claudin-15 mAb, rabbit anti-claudin-15 pAb (Zymed, San Francisco, CA, USA), anti-actin pAb (Sigma, Saint Louis, MO, USA), horse radish peroxidase-conjugated anti-mouse or anti-rabbit Ig (GE Healthcare UK Ltd, Buckinghamshire, England), Alexa Fluor 488-conjugated goat anti-mouse or anti-rabbit Ig, and Alexa Fluor 594-conjugated goat anti-mouse or anti-rabbit Ig (Molecular Probes, Eugene, OR, USA).
Gel electrophoresis and immunoblotting
The cells were washed with ice-cold phosphate-buffered saline (PBS), pH 7.4, and treated with 10% trichloroacetic acid (TCA) for 15 min on ice to inactivate intracellular proteases. The cells were lysed in lysis buffer (62.5 mM Tris, pH 6.8, 2% SDS, 10% glycerol) after removal of TCA and the pH was adjusted to 6.8 using 2 M Tris. After determination of the amount of total protein using the BCA protein-assay reagent (Pierce Chemical, Rockford, IL, USA), 2-mercaptoethanol (5%) and bromophenol blue (0.002%) were added to each cell lysate. Proteins were fractionated by sodium dodecyl sulfate (SDS)-polyacrylamide gel electrophoresis and electrophoretically transferred onto PVDF membranes. The membranes were incubated with anti-claudin-1, anti-claudin-15 or anti-actin antibodies and then with horseradish peroxidase-conjugated anti-mouse or anti-rabbit Ig. Antibodies were detected using the ECL Western blotting analysis system (GE Healthcare UK Ltd). Some membranes were reprobed after stripping primary and secondary antibodies with stripping buffer (62.5 mM Tris, pH 6.7, 2% SDS, 100 mM 2-mercaptoethanol) at 50°C for 30 min. Densitometric analysis was done by using the Image J software (http://rsb.info.nih.gov/ij).
Confocal laser scanning microscopy
Cells plated on glass cover slips were rinsed with PBS, fixed with 1.5% paraformaldehyde in PBS for 15 min, washed with PBS, and then permeablized with 0.2% Triton X-100 in PBS for 20 min. After blocking in 1% bovine serum albumin in PBS, they were incubated with primary antibodies for 1 h. They were washed thrice with PBS and incubated for 30 min with Alexa Fluor-conjugated secondary antibodies. After washing thrice with PBS, they were embedded in Vectashield mounting medium (Vector Laboratories, Burlingame, CA, USA). The specimens were examined with a confocal laser scanning microscope LSM 510 meta (Zeiss, Oberkochen, Germany).
Freeze-fracture electron microscopy
For freeze-fracture electron microscopy, confluent cells were fixed with 2.5% glutaraldehyde in 0.1 M cacodylate buffer, pH 7.4, rinsed in cacodylate buffer, cryoprotected in 30% glycerol in cacodylate buffer and then frozen in liquid propane. The frozen samples were fractured at −120°C and platinum-shadowed unidirectionally at an angle of 45° in a JFD-7000 apparatus (JEOL, Tokyo, Japan). Replicas with cells were immersed in household bleach to remove cells, washed in distilled water and then mounted on copper grids. They were examined using a Jeol 2000 EX electron microscope (JEOL).
Morphometrical analysis and statistics
Morphometrical analysis was performed on freeze-fracture replica images of TJs, which were printed at a final magnification of 20,000×. The mean TJ strand number was determined by taking numerous counts along a line drawn perpendicular to the junctional axis at 200 nm intervals (Stevenson et al. 1988). The number of free-ended TJ strands was counted per unit length (1 μm) of TJ strands (Sonoda et al. 1999). Statistical analysis was performed using the two-sided Wilcoxon’s signed rank test for determination of significance. Values of P < 0.05 were considered to be significant.
Results
Inducible expression of EGFP-1CL, EGFP-15CL or 15CL in MEF/3T3 and MDCK II Tet-Off cells
In this study, full-length mouse claudin-1 and human claudin-15 with EGFP at their NH2-terminus and human claudin-15 are referred to as EGFP-1CL, EGFP-15CL and 15CL, respectively. EGFP-1CL, EGFP-15CL or 15CL were expressed in mouse fibroblastic cells (MEF/3T3 Tet-Off cells) and canine epithelial cells (MDCK II Tet-Off cells). The expression of EGFP-1CL, EGFP-15CL or 15CL was induced by the removal of Dox from the culture medium owing to the Tet-Off gene expression system used in this study. Immunoblot analysis showed that bands corresponding to the predicted sizes were detected in the lysates of transfectants in the absence of Dox (Fig. 1, arrowheads), but EGFP-1CL expression in MEF cells was slightly detected in the presence of Dox (Fig. 1A). Densitometric analysis of blots indicated that protein expression of EGFP-1CL (clone 1 and 2), EGFP-15CL (clone 1 and 2), and 15CL (clone 1) were 1.03, 0.97, 0.78, 0.80, and 0.94, respectively, when normalized by actin.
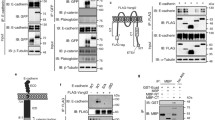
Immunoblot analysis of MEF/3T3 Tet-Off cells expressing EGFP-1CL or EGFP-15CL (a), and MDCK II Tet-Off cells expressing EGFP-1CL, EGFP-15CL or claudin-15 (15CL, b). These cells were cultured for 4 days in the presence or absence of doxycycline (Dox). After fractionation of total cell lysates (20 μg/lane) by SDS-PAGE, immunoblotting was performed using anti-claudin-1 antibody or anti-claudin-15 antibody (a, b). Bands corresponding to the predicted molecular weight (arrowheads) with several degraded forms are detected in the lysates of transfectants in the absence of Dox. Membranes were reprobed with anti-actin antibody after stripping primary and secondary antibodies (b). Arrows indicate non-specific bands (a)
EGFP-1CL and EGFP-15CL were co-localized with ZO-1 at cell-to-cell contacts sites between fibroblasts
Previous studies (Kobayashi et al. 2002; McCarthy et al. 2000) showed that claudin-1 with a myc-tag at its COOH-terminus induced aberrant TJ strands along entire lateral membranes in MDCK cells, because of interference with its binding to ZO-1. In order to examine whether EGFP-1CL or EGFP-15CL is co-localized with ZO-1 at cell-to-cell contact sites, EGFP-1CL or EGFP-15CL was expressed in fibroblasts, which do not have TJs. Confocal microscopy showed that both EGFP-1CL and EGFP-15CL were expressed at cell-to-cell contact sites and co-localized with ZO-1 (Fig. 2), suggesting that EGFP at the NH2-terminus of claudins may not interfere with the binding between the COOH-terminus of claudin and ZO-1.
Confocal laser scanning microscopy of MEF/3T3 Tet-Off cells expressing EGFP-1CL or EGFP-15CL. Cells cultured for 4 days in the absence of Dox were stained with anti-ZO-1 mAb. Green signals for EGFP-1CL (a) or EGFP-15CL (d), red signals for ZO-1 (c, f), and the merged images (b, e) are shown. EGFP-1CL and EGFP-15CL are detected at the cell-to-cell contact sites and they are colocalized with ZO-1. Bars, 10 μm
EGFP-1CL was localized at the apical TJ but EGFP-15CL and 15CL were localized in the lateral membrane by confocal microscopy
The x–y image (Fig. 3a–c) and x–z image (Fig. 3a′–c′) constructed by confocal microscopy confirmed that EGFP-1CL was localized at the most apical region of the lateral membranes corresponding to apical TJs, and co-localized with ZO-1 in EGFP-1CL-expressing MDCK II Tet-Off cells (Fig. 3a–c). EGFP-1CL was also detected in the cytoplasm with a vesicular appearance (Fig. 3a–c). EGFP-1CL was not detected in the presence of Dox (data not shown). We next examined the localization of EGFP-15CL in MDCK II Tet-Off cells. EGFP-15CL was localized at cell-to-cell contact sites and in the cytoplasm with a vesicular appearance (Fig. 3d–f). The x–z image showed that EGFP-15CL was localized along the entire lateral membranes and co-localized with ZO-1 only at the most apical region of the lateral membranes (Fig. 3d′–f′). EGFP-15CL was not detected in the presence of Dox (data not shown). To examine whether EGFP at the NH2-terminus of claudin-15 affected the localization of claudin-15, full-length claudin-15 was expressed in MDCK II Tet-Off cells. The parent MDCK II Tet-Off cells were not stained with the claudin-15 antibodies used in this study (data not shown). 15CL was localized at cell-to-cell contact sites and in the cytoplasm with a vesicular appearance (Fig. 3g–i). The x–z image showed that 15CL was also localized along the entire lateral membranes and co-localized with ZO-1 only at the most apical region of the lateral membranes (Fig. 3g′–i′). 15CL was not detected in the presence of Dox (data not shown).
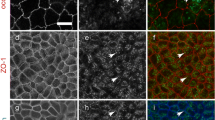
Confocal laser scanning microscopy of MDCK II Tet-Off cells expressing EGFP-1CL (a), EGFP-15CL (b) and 15CL (c). Cells were cultured on glass cover slips for 3 days in the absence of Dox and immunostained with anti-ZO-1 mAb (a, b) and with anti-claudin-15 pAb and anti-ZO-1 mAb (c). Green signals for EGFP-1CL (a, a′), EGFP-15CL (d, d′) and 15CL (g, g′), red signals for ZO-1 (c, c′, f, f′, i, i′), and the merged images (b, b′, e, e′, h, h′) are shown. Fourteen x–y sections were recorded at 0.37 μm intervals from base to apex by confocal microscopy in EGFP-1CL-expressing cells, 23 x–y sections were recorded at 0.37 μm intervals in EGFP-15CL-expressing cells, and 26 x–y sections were recorded at 0.37 μm intervals in 15CL-expressing cells. The x–z images (a′–i′) were reconstructed at white lines drawn in the x–y images (a–i). EGFP-1CL is detected only at the apical junctional regions of cell-to-cell contact sites and colocalized with ZO-1 (a–c, a′–c′). EGFP-15CL and 15CL are detected in the lateral membrane (d′, g′) and colocalized with ZO-1 only at the apical junctional regions (e′, h′). Bars, 10 μm
EGFP-15CL and 15CL, but not EGFP-1CL, formed aberrant TJ strands by freeze-fracture electron microscopy
In parent MDCK II Tet-Off cells and EGFP-1CL-expressing cells, freeze-fracture electron microscopy showed that TJ strands existed only at the apical junctional region, but not in the lateral membranes (Fig. 4a, c). Basal extensions with free ends from apical TJ strands were rarely observed (Fig. 4a–e). In EGFP-15CL-expressing cells and 15CL-expressing cells, freeze-fracture electron microscopy showed that numerous TJ-like structures (aberrant strands) were detected in the lateral membranes in the absence of Dox (Fig. 4f, i). Aberrant strands were never observed in the presence of Dox (data not shown). Most particles of aberrant strands were associated with the P-face (Fig. 4h, k) and the furrows of the E-face were almost free of these particles (Fig.4h, k). Basal extensions with free ends from apical TJ strands were frequently observed (Fig. 4g, j). Confocal microscopy and freeze-fracture electron microscopy showed similar results in EGFP-15CL- and 15CL-expressing cells, suggesting that EGFP at the NH2-terminus of claudin-15 did not affect the localization of claudin-15 and formation of aberrant strands.
Freeze-fracture electron microscopy of parent MDCK II Tet-Off cells (a), EGFP-1CL-expressing cells (b), EGFP-15CL-expressing cells (c), and 15CL-expressing cells (d). Cells were cultured for 3 days in the absence of Dox. In MDCK II Tet-Off cells and EGFP-1CL-expressing cells, tight junction (TJ) strands existed at the most apical region of the lateral membrane and no strands were detected in the lateral membrane (a, c). Most TJ particles are associated with the P-face and free-ended TJ strands are not frequently observed (b, d, e). In EGFP-15CL and 15CL cells, numerous aberrant strands are widely distributed in the lateral membrane (f, i). Free-ended strands (arrowheads) are frequently observed at the apical junctional regions (g, j). Most aberrant strands are associated with the P-face (P) and the furrow of the E-face (E) of aberrant strands is almost free of particles (h, k). Bars, 200 nm
Statistical analysis of the number of apical TJ strands and free-ended TJ
We performed a morphometrical analysis to examine whether introduced claudins caused an increase in the number of apical TJ strands and free-ended TJs in MDCK II cells. We counted the number of apical TJ strands and free-ended TJs in two clones of EGFP-1CL, two clones of EGFP-15CL and one clone of 15CL in the presence or absence of Dox. The mean number of apical TJ strands increased by 45.7–49.5% in EGFP-1CL cells when the expression of EGFP-1CL was induced (Fig. 5a). Expression of EGFP-15CL increased the mean number of apical TJ strands by 13.7–27.1% (Fig. 5a). Expression of 15CL increased the mean number of apical TJ strands by 27.9% (Fig. 5a). Concerning the increase in the number of apical TJ strands, expression of EGFP-1CL was more effective than that of EGFP-15CL or 15CL. In sharp contrast to the increase in the number of apical TJs, EGFP-1CL did not increase the free-ended TJs (Fig. 5b). However, EGFP-15CL or 15CL dramatically increased free-ended TJs by 195–246% or by 151%, respectively (Fig. 5b).
Statistical analysis of the number of apical TJ strands and free-ended TJ strands. Cells expressing EGFP-1CL (clone 1, 2), EGFP-15CL (clone 1, 2), or 15CL (clone 1) were cultured for 3 days in the presence or absence of Dox, and the mean number of apical TJ strands (a) and the number of free-ended TJ strands per 1 μm of TJ strands (b) were counted in these cells. a The mean numbers of apical TJ strands are significantly increased in all clones by the induction of EGFP-1CL, EGFP-15CL or 15CL, which is performed by the removal of Dox. b Free-ended TJ strands were counted per 1 μm of TJ strands. In EGFP-1CL-expressing cells, the numbers of free-ends do not change following the induction of EGFP-1CL. In EGFP-15CL- or 15CL-expressing cells, however, the numbers of free-ends are significantly increased by the induction of EGFP-15CL or 15CL. * Different from corresponding control (P < 0.05)
EGFP-15CL and 15CL were not concentrated at TJs formed between heterotypic cells
The number of apical TJ strands increased more in EGFP-1CL cells than in EGFP-15CL cells and 15CL cells. By contrast, the number of free-ended TJ strands was dramatically increased in EGFP-15CL cells and 15CL cells compared with EGFP-1CL. From these results, we speculated that claudin-15 may be incompatible with endogenous claudin species expressed in MDCK II Tet-Off cells. Thus, we examined cell-to-cell contact sites between heterotypic cells; that is, exogenous claudin-expressing cells (EGFP-1CL cells, EGFP-15CL cells and 15CL cells) and parent cells (MDCK II Tet-Off cells). For this purpose, EGFP-1CL cells, EGFP-15CL cells or 15CL cells were co-cultured for 24 h with the parent cells at a ratio of 1:10 in the absence of Dox. EGFP-1CL was localized in cell-to-cell contact sites between heterotypic cells (EGFP-1CL-expressing cells and MDCK II Tet-Off cells) as well as between EGFP-1CL-expressing cells, and co-localized with ZO-1 at the apical region (Fig. 6a–c, a′–c′). EGFP-15CL was detected in cell-to-cell contact sites between EGFP-15CL-expressing cells and co-localized with ZO-1 only at the apical junctional region (Fig. 6d–f, d′–f′). However, EGFP-15CL was hardly detected in the cell-to-cell contact sites between heterotypic cells (Fig. 6d–f, d′–f′). Similar to EGFP-15CL, 15CL was only detected in cell-to-cell contact sites between 15CL-expressing cells, and not in those between heterotypic cells (Fig. 6g–i, g′–i′).
Confocal laser scanning microscopy of MDCK II Tet-Off cells expressing EGFP-1CL (a–c, a′–c′), EGFP-15CL (d–f, d′–f′), or 15CL (g–i, g′–i′) cocultured with the parent MDCK II Tet-Off cells. Cells expressing EGFP-1CL, EGFP-15CL, or 15CL were cultured for 3 days in the absence of Dox and then co-cultured for 1 day with the parent cells at a ratio of 1:10. Cells were stained with anti-ZO-1 mAb (a–f, a′–f′) or with anti-claudin-15 pAb and anti-ZO-1 mAb (g–i, g′–i′). Green signals for EGFP-1CL (a, a′), EGFP-15CL (d, d′), or claudin-15 (g, g′), red signals for ZO-1 (c, c′, f, f′, i, i′), and the merged images (b, b′, e, e′, h, h′) are shown. Twelve x–y sections in EGFP-1CL-expressing cells, 17 x–y sections in EGFP-15CL-expressing cells, and 15 x–y sections in 15CL-expressing cells were recorded at 0.37 μm intervals from base to apex by confocal microscopy. The x–z images (a′–i′) were reconstructed at white lines drawn in the x–y images (a–i). Green signals for EGFP-1CL are detected in the heterotypic cell-to-cell junctions (arrowheads) formed by EGFP-1CL-expressing cells and the parent cells in addition to the cell-to-cell junctions between EGFP-1CL-expressing cells (a–c, a′–c′). By contrast, green signals for EGFP-15CL are not detected in the heterotypic cell-to-cell junctions (arrowheads), but are detected in the homotypic cell-to-cell junctions between EGFP-15CL-expressing cells (d–f, d′–f′). Similarly, green signals for 15CL are not detected in the heterotypic cell-to-cell junctions (arrowheads) but are detected in the homotypic cell-to-cell junctions between 15CL-expressing cells (g–i, g′–i′). Bars, 10 μm
Discussion
The detailed process of the formation of TJ strands is still poorly understood. We believe that detailed investigations of the process of aberrant TJ formation may lead us to clues clarifying the mechanisms of normal TJ strand formation. Previous studies have indicated that the hindering of claudin/ZO-1 interactions caused aberrant TJ strands. Overexpression of claudin-1 with a myc-epitope at the COOH-terminus (1CLmyc) in MDCK cells caused the formation of numerous aberrant strands along the lateral plasma membrane, but overexpression of claudin-1 itself did not lead to the formation of aberrant strands (McCarthy et al. 2000). These contradictory results suggest that the myc-epitope at the COOH-terminus of claudin-1 hindered binding to ZO-1 and induced aberrant strands. However, we previously showed using the doxycycline-regulated gene expression system that 1CLmyc was concentrated to apical TJs in MDCK II cells when expressed at low levels, but that it was localized to the lateral membrane and formed aberrant strands when expressed at high levels (Kobayashi et al. 2002). In addition, expression of 1CLmyc from a pCI-neo vector (constitutive expression system) did not lead to the formation of aberrant strands (Inai et al. 1999) and the expression level was found to be low (Kobayashi et al. 2002). Taken together, claudins mutated at the COOH-terminus may induce the formation of aberrant strands when expressed at a high level, but endogenous claudins may partially rescue the mutated claudins to be localized at apical TJs when expressed at low level.
We hypothesized that there would be another mechanism of aberrant TJ strand formation, besides the hindering of claudin/ZO-1 interaction. First, we expressed claudin-1 with EGFP at its NH2-terminus (EGFP-1CL) or claudin-15 with EGFP at its NH2-terminus (EGFP-15CL) in fibroblasts that do not form TJs. Both EGFP-1CL and EGFP-15CL were localized at cell-to-cell contact sites and colocalized with ZO-1, suggesting that EGFP-1CL and EGFP-15CL may bind to ZO-1. Next, we expressed EGFP-1CL or EGFP-15CL in TJ-bearing epithelial cells, MDCK II Tet-Off cells. Confocal microscopy showed that EGFP-1CL was concentrated at apical TJs, but EGFP-15CL was localized along the lateral plasma membranes in addition to apical TJs. As seen by freeze-fracture electron microscopy, overexpression of EGFP-15CL, but not EGFP-1CL, induced aberrant TJ strands in lateral plasma membranes. It is interesting that only overexpression of EGFP-15CL induced aberrant TJ strands, because both EGFP-1CL and EGFP-15CL have ZO-1 binding motifs in their COOH-termini and they were colocalized with ZO-1 when expressed in fibroblasts. In order to eliminate another possibility that EGFP at the NH2-terminus of claudin-15 may be related to the formation of aberrant strands, we expressed claudin-15 (15CL) itself in MDCK II Tet-Off cells. 15CL was localized in lateral plasma membranes by confocal microscopy, and aberrant TJ strands were observed in the lateral membranes by freeze-fracture electron microscopy. These results suggest that claudin-15 itself, but not EGFP at the NH2-terminus, induced aberrant strands even though claudin-15 contains the ZO-1-binding motif.
Morphometrical analysis demonstrated that expression of EGFP-1CL, which was colocalized with ZO-1 at apical TJs, resulted in a 47.5% increase in the number of TJ strands. These results indicate that exogenously expressed EGFP-1CL was incorporated into apical TJs and caused the increase in the number of TJ strands. On the other hand, expression of EGFP-15CL or 15CL induced only 20.5 or 28.0% increases in the number of TJ strands, respectively; but expression of EGFP-15CL or 15CL caused approximately 1.5 or 2.5 times greater increases in the number of free-ended TJ strands compared with control cells cultured in the presence of Dox. Expression of EGFP-1CL did not increase the number of free-ended TJ strands. An increase in the number of free-ended TJ strands may represent degradation of preexisted-TJ strands or inhibition of insertion of newly formed TJ strands into apical TJs. Taking the slight increase in the number of apical TJ strands into account, TJ strands newly formed by overexpression of EGFP-15CL or 15CL may be disturbed during insertion into apical TJs by incompatibility between claudin species.
Multiple claudins are generally expressed in single cells (Furuse et al. 1999), but it is still unclear how claudins interact with each other within and between TJ strands, and how claudins are polymerized into TJ strands. It is believed that TJ strands are formed by the linear polymerization of claudins within one plasma membrane, by lateral interaction of claudins, and that paired TJs are formed between adjacent cells by head-to-head interactions between TJ strands (Tsukita et al. 2001). It was demonstrated that claudin-3-based strands could interact with claudin-1- and claudin-2-based strands. However, claudin-1-based strands did not heterotypically interact with claudin-2-based strands (Furuse et al. 1999). These results suggest that distinct species of claudins can interact within and between TJ strands, except in some combinations. The incompatibility of head-to-head interactions between claudin-1 and -5 was also reported (Coyne et al. 2003; Daugherty et al. 2007). Recently, it has been demonstrated that two extracellular loops of claudins are involved in regulating heterotypic claudin compatibility using a series of claudin-3/4 chimeras (Daugherty et al. 2007). Considering the heterotypic compatibility between claudin species, it is possible that the formation of free-ended apical TJ strands and aberrant TJ strands may be due to the incompatibility between claudin-15 and endogenous claudins. To verify this possibility, we co-cultured parent MDCK II Tet-Off cells with either EGFP-1CL-, EGFP-15CL- or 15CL-expressing cells. As expected, EGFP-15CL and 15CL were not localized at heterotypic cell-to-cell contacts between parent cells and EGFP-15CL- or 15CL-expressing cells. These results suggest that claudin-15 could not form paired TJ strands by head-to-head interactions with endogenous claudins in parent MDCK II cells.
There are many examples to show that claudins are localized in the lateral plasma membrane in vivo, including non-TJ-bearing cells (Acharya et al. 2004; Brandner et al. 2002; Coyne et al. 2003; Fujita et al. 2006; Furuse et al. 2002; Guan et al. 2005; Inai et al. 2005, 2007; Langbein et al. 2002; Li et al. 2004; Rahner et al. 2001; Troy et al. 2005; Yi et al. 2000). The physiological roles of localization of claudins in the lateral membrane is still obscure, but most claudin molecules in the lateral membrane do not form TJ strands (Inai et al. 2007). The epithelial cell adhesion molecule EpCAM is a Ca2+-independent, homophilic cell-to-cell adhesion molecule, expressed in most epithelial cells and enriched in the basolateral membrane (Balzar et al. 1999; Winter et al. 2003). EpCAM was found to interact directly with claudin-7 (Ladwein et al. 2005), providing a possible mechanism of how claudin-7 is localized in the lateral membrane. Thus, there may be another possibility that claudin-15 could be localized in the lateral membrane in MDCK II cells by binding to EpCAM-like molecule(s) interacting with claudin-15.
If claudin-15 was incompatible with endogenous claudins in MDCK II cells, how could EGFP-15CL or 15CL be localized at the apical TJs containing endogenous claudins in EGFP-15CL- or 15CL-expressing cells? EGFP-15CL or 15CL was probably co-polymerized with endogenous claudins to form TJ strands by side-by-side interactions in one membrane, and incorporated EGFP-15CL or 15CL formed paired TJ strands by homophilic head-to-head coupling between EGFP-15CL or 15CL as shown in “Model C” (Furuse et al. 1999). It was demonstrated by fluorescence recovery after photobleaching in paired TJ strands in fibroblasts expressing GFP-claudin-1 that GFP-claudin-1 molecules, once incorporated into TJ strands, could not move freely within strands (Sasaki et al. 2003). Therefore, head-to-head coupling between compatible claudin species may occur first and then side-by-side interactions follow immediately to elongate paired TJ strands.
In summary, the expression of EGFP-15CL and 15CL in MDCK II cells increased the numbers of apical TJ strands and free-ended strands and induced aberrant TJ strands in the lateral membrane. EGFP-15CL and 15CL could not be localized at TJs between parent cells and EGFP-15CL- or 15CL-expressing cells, suggesting that both EGFP-15CL and 15CL may be incompatible with endogenous claudins expressed in MDCK II cells. Therefore, in addition to the interaction between claudins and ZO-1, the compatibility between different claudin species may play a crucial role in the localization of exogenously expressed claudins at the apical TJs.
References
Acharya P, Beckel J, Ruiz WG, Wang E, Rojas R, Birder L, Apodaca G (2004) Distribution of the tight junction proteins ZO-1, occludin, and claudin-4, -8, and -12 in bladder epithelium. Am J Physiol 287:F305–F318
Balzar M, Winter MJ, de Boer CJ, Litvinov SV (1999) The biology of the 17-1A antigen (Ep-CAM). J Mol Med 77:699–712
Brandner JM, Kief S, Grund C, Rendl M, Houdek P, Kuhn C, Tschachler E, Franke WW, Moll I (2002) Organization and formation of the tight junction system in human epidermis and cultured keratinocytes. Eur J Cell Biol 81:253–263
Coyne CB, Gambling TM, Boucher RC, Carson JL, Johnson LG (2003) Role of claudin interactions in airway tight junctional permeability. Am J Physiol 285:L1166–L1178
Daugherty BL, Ward C, Smith T, Ritzenthaler JD, Koval M (2007) Regulation of heterotypic claudin compatibility. J Biol Chem 282:30005–30013
Farquhar MG, Palade GE (1963) Junctional complexes in various epithelia. J Cell Biol 17:375–412
Fujita H, Chiba H, Yokozaki H, Sakai N, Sugimoto K, Wada T, Kojima T, Yamashita T, Sawada N (2006) Differential expression and subcellular localization of claudin-7, -8, -12, -13, and -15 along the mouse intestine. J Histochem Cytochem 54:933–944
Furuse M, Fujita K, Hiiragi T, Fujimoto K, Tsukita S (1998) Claudin-1 and -2: novel integral membrane proteins localizing at tight junctions with no sequence similarity to occludin. J Cell Biol 141:1539–1550
Furuse M, Hata M, Furuse K, Yoshida Y, Haratake A, Sugitani Y, Noda T, Kubo A, Tsukita S (2002) Claudin-based tight junctions are crucial for the mammalian epidermal barrier: a lesson from claudin-1-deficient mice. J Cell Biol 156:1099–1111
Furuse M, Sasaki H, Tsukita S (1999) Manner of interaction of heterogeneous claudin species within and between tight junction strands. J Cell Biol 147:891–903
Guan X, Inai T, Shibata Y (2005) Segment-specific expression of tight junction proteins, claudin-2 and -10, in the rat epididymal epithelium. Arch Histol Cytol 68:213–225
Gumbiner BM (1993) Breaking through the tight junction barrier. J Cell Biol 123:1631–1633
Harris BZ, Lim WA (2001) Mechanism and role of PDZ domains in signaling complex assembly. J Cell Sci 114:3219–3231
Hou J, Gomes AS, Paul DL, Goodenough DA (2006) Study of claudin function by RNA interference. J Biol Chem 281:36117–36123
Inai T, Kobayashi J, Shibata Y (1999) Claudin-1 contributes to the epithelial barrier function in MDCK cells. Eur J Cell Biol 78:849–855
Inai T, Sengoku A, Guan X, Hirose E, Iida H, Shibata Y (2005) Heterogeneity in expression and subcellular localization of tight junction proteins, claudin-10 and -15, examined by RT-PCR and immunofluorescence microscopy. Arch Histol Cytol 68:349–360
Inai T, Sengoku A, Hirose E, Iida H, Shibata Y (2007) Claudin-7 expressed on lateral membrane of rat epididymal epithelium does not form aberrant tight junction strands. Anat Rec 290:1431–1438
Itoh M, Furuse M, Morita K, Kubota K, Saitou M, Tsukita S (1999) Direct binding of three tight junction-associated MAGUKs, ZO-1, ZO-2, and ZO-3, with the COOH termini of claudins. J Cell Biol 147:1351–1363
Jou TS, Nelson WJ (1998) Effects of regulated expression of mutant RhoA and Rac1 small GTPases on the development of epithelial (MDCK) cell polarity. J Cell Biol 142:85–100
Kiuchi-Saishin Y, Gotoh S, Furuse M, Takasuga A, Tano Y, Tsukita S (2002) Differential expression patterns of claudins, tight junction membrane proteins, in mouse nephron segments. J Am Soc Nephrol 13:875–886
Kobayashi J, Inai T, Shibata Y (2002) Formation of tight junction strands by expression of claudin-1 mutants in their ZO-1 binding site in MDCK cells. Histochem Cell Biol 117:29–39
Ladwein M, Pape UF, Schmidt DS, Schnolzer M, Fiedler S, Langbein L, Franke WW, Moldenhauer G, Zoller M (2005) The cell-cell adhesion molecule EpCAM interacts directly with the tight junction protein claudin-7. Exp Cell Res 309:345–357
Langbein L, Grund C, Kuhn C, Praetzel S, Kartenbeck J, Brandner JM, Moll I, Franke WW (2002) Tight junctions and compositionally related junctional structures in mammalian stratified epithelia and cell cultures derived therefrom. Eur J Cell Biol 81:419–435
Li WY, Huey CL, Yu AS (2004) Expression of claudin-7 and -8 along the mouse nephron. Am J Physiol 286:F1063–F1071
McCarthy KM, Francis SA, McCormack JM, Lai J, Rogers RA, Skare IB, Lynch RD, Schneeberger EE (2000) Inducible expression of claudin-1-myc but not occludin-VSV-G results in aberrant tight junction strand formation in MDCK cells. J Cell Sci 113 Pt 19:3387–3398
Morita K, Sasaki H, Fujimoto K, Furuse M, Tsukita S (1999) Claudin-11/OSP-based tight junctions of myelin sheaths in brain and Sertoli cells in testis. J Cell Biol 145:579–588
Pawson T, Nash P (2003) Assembly of cell regulatory systems through protein interaction domains. Science 300:445–452
Rahner C, Mitic LL, Anderson JM (2001) Heterogeneity in expression and subcellular localization of claudins 2, 3, 4, and 5 in the rat liver, pancreas, and gut. Gastroenterology 120:411–422
Rodriguez-Boulan E, Nelson WJ (1989) Morphogenesis of the polarized epithelial cell phenotype. Science 245:718–725
Sasaki H, Matsui C, Furuse K, Mimori-Kiyosue Y, Furuse M, Tsukita S (2003) Dynamic behavior of paired claudin strands within apposing plasma membranes. Proc Natl Acad Sci USA 100:3971–3976
Schneeberger EE, Lynch RD (2004) The tight junction: a multifunctional complex. Am J Physiol 286:C1213–C1228
Sheng M, Sala C (2001) PDZ domains and the organization of supramolecular complexes. Ann Rev Neurosci 24:1–29
Singh AB, Harris RC (2004) Epidermal growth factor receptor activation differentially regulates claudin expression and enhances transepithelial resistance in Madin-Darby canine kidney cells. J Biol Chem 279:3543–3552
Songyang Z, Fanning AS, Fu C, Xu J, Marfatia SM, Chishti AH, Crompton A, Chan AC, Anderson JM, Cantley LC (1997) Recognition of unique carboxyl-terminal motifs by distinct PDZ domains. Science 275:73–77
Sonoda N, Furuse M, Sasaki H, Yonemura S, Katahira J, Horiguchi Y, Tsukita S (1999) Clostridium perfringens enterotoxin fragment removes specific claudins from tight junction strands: Evidence for direct involvement of claudins in tight junction barrier. J Cell Biol 147:195–204
Staehelin LA (1974) Structure and function of intercellular junctions. Int Rev Cytol 39:191–283
Stevenson BR, Anderson JM, Goodenough DA, Mooseker MS (1988) Tight junction structure and ZO-1 content are identical in two strains of Madin-Darby canine kidney cells, which differ in transepithelial resistance. J Cell Biol 107:2401–2408
Troy TC, Rahbar R, Arabzadeh A, Cheung RM, Turksen K (2005) Delayed epidermal permeability barrier formation and hair follicle aberrations in Inv-Cldn6 mice. Mech Dev 122:805–819
Tsukita S, Furuse M (1999) Occludin and claudins in tight-junction strands: leading or supporting players? Trends Cell Biol 9:268–273
Tsukita S, Furuse M, Itoh M (2001) Multifunctional strands in tight junctions. Nat Rev Mol Cell Biol 2:285–293
Van Itallie CM, Fanning AS, Anderson JM (2003) Reversal of charge selectivity in cation or anion-selective epithelial lines by expression of different claudins. Am J Physiol 285:F1078–F1084
Wilcox ER, Burton QL, Naz S, Riazuddin S, Smith TN, Ploplis B, Belyantseva I, Ben-Yosef T, Liburd NA, Morell RJ, Kachar B, Wu DK, Griffith AJ, Riazuddin S, Friedman TB (2001) Mutations in the gene encoding tight junction claudin-14 cause autosomal recessive deafness DFNB29. Cell 104:165–172
Winter MJ, Nagtegaal ID, van Krieken JH, Litvinov SV (2003) The epithelial cell adhesion molecule (Ep-CAM) as a morphoregulatory molecule is a tool in surgical pathology. Am J Pathol 163:2139–2148
Wittchen ES, Haskins J, Stevenson BR (1999) Protein interactions at the tight junction. Actin has multiple binding partners, and ZO-1 forms independent complexes with ZO-2 and ZO-3. J Biol Chem 274:35179–35185
Yi X, Wang Y, Yu FS (2000) Corneal epithelial tight junctions and their response to lipopolysaccharide challenge. Invest Ophthalmol 41:4093–4100
Acknowledgments
This work was supported in part by grants-in-aid for scientific research from the Ministry of Education, Science and Culture, Japan (numbers 11770008, 13670018, 16590146, and 18590187).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Sengoku, A., Inai, T. & Shibata, Y. Formation of aberrant TJ strands by overexpression of claudin-15 in MDCK II cells. Histochem Cell Biol 129, 211–222 (2008). https://doi.org/10.1007/s00418-007-0354-y
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00418-007-0354-y