Abstract
We have built an ion-microbeam for studies of the nuclear topography and kinetics of double-strand break repair at the single cell level. Here, we show that a first and a second, delayed single ion exposure at different nuclear sites led to comparable accumulations of phospho-ATM, γ-H2AX and Mdc1 at both earlier (e) and later (l) microirradiated sites. In contrast, accumulations of 53BP1 and the recombination protein Rad51 were strongly reduced at l-sites. This apparent competition effect is accompanied by a reduced amount of 53BP1 in undamaged areas of the irradiated nuclei. We suggest that a critically limited pool size combined with strong binding at irradiated sites leads to the exhaustion of unbound factors freely roaming the nuclear space. The undersupply of these factors at l-sites requires in addition a long-lasting binding at e-sites or a weaker binding at l-sites. The observed effects suggest that DNA damage response at individual nuclear sites depends on the time course of damage load. This may have implications for therapeutic radiation treatments.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
DNA double-strand breaks (DSB) elicit complex response reactions in mammalian cells which lead to cell-cycle arrest, induction of repair, or elimination of damaged cells [1]. The serine/threonine kinase ATM is central in regulating cellular response to DSB by phosphorylating several target proteins [2]. Many of the ATM targets are present in so-called ionizing radiation-induced foci, i.e. large protein accumulations built at DSB sites [3]. Two pathways to ATM activation were proposed: one based on interaction of ATM with a DSB-sensing MRN (Mre11/Rad50/Nbs1) complex [4–7], and one depending on DSB-induced chromatin alterations [8, 9].
53BP1 and Mdc1 are mediator proteins required for efficient ATM activation [10] and ATM-dependent phosphorylation of targets [11–13]. They were proposed to function in parallel pathways towards ATM activation [10]. The Mdc1-dependent branch was suggested to act via the MRN complex, while the 53BP1-dependent branch was proposed to act via chromatin alterations [14]. At low and moderate irradiation doses (from below 1 Gy up to several Gy), efficient activation of ATM appears to depend on the presence of both branches, while at higher doses one branch apparently can compensate for the other [10, 11, 13, 15].
The molecular mechanisms of this and other dose-dependent switches in DNA damage response reactions are presently not understood, but such understanding is required for the assessment of risks associated with radiation exposure. Mechanisms such as exhaustion of limited pools of response factors, saturation of the production of active response factors, or a change in reaction pathways may be involved. Sequential irradiation treatments, where the response to the earlier treatment can develop for some time before the later treatment takes place, have been developed to elucidate this kind of mechanisms. With standard irradiation set-ups, it is difficult to discriminate between the responses to an earlier and a later irradiation event.
In the present investigation, we were able to overcome this problem with the help of the ion microprobe SNAKE [16, 17]. This instrument was applied to irradiate HeLa cells with single carbon or oxygen ions in a line pattern. After irradiation the cells were incubated for a time interval, Δt, before a later irradiation with a line pattern perpendicular to the earlier one was performed. Since SNAKE allows to target the ions with an accuracy better than 0.55 μm full width at half maximum [18], and since foci patterns induced along the paths of the ions turned out to be sufficiently stable over time, we were able to distinguish between the line pattern induced during the earlier and the later microirradiation series. A comparison of foci intensities showed that γ-H2AX, Phospho-ATM (Ser1981) and Mdc1 foci are comparable at sites damaged during the earlier and during the later irradiation. In contrast, 53BP1 and Rad51 focus formation is reduced at sites damaged during the later irradiation.
Materials and methods
Cell cultivation and irradiation
For ion irradiation, HeLa cells were grown on 6 μm micro-structured mylar foil pre-treated with Cell-TAK (BD Bioscience) in stainless steel cell chambers designed as described [16]. Cells were seeded in RPMI 1640 medium with 10% fetal calf serum, 100 U/ml penicillin and 50 μg/ml streptomycin about 24 h before irradiation to obtain sub-confluent cultures. During irradiation at room temperature, the cell chamber is placed vertically in the focal plane of the ion microbeam, i.e. the cells are not covered by medium, but residual medium in a reservoir ensures a saturated atmosphere [16]. Typically, the total time required for sample preparation and irradiation, i.e. the time outside of the CO2 incubator, is less than 15 min, including about 12 min in the upright position. Under these conditions, we did not observe enhanced formation of repair foci in unirradiated cells as compared to samples that remained in the incubator.
After irradiation, the growth medium was replaced by fresh, pre warmed medium, and the cells were incubated (37°C, 5% CO2) for the time intervals indicated, before either a second irradiation was performed or the cells were fixed for immunofluorescent detection.
Irradiations were performed at the ion-microirradiation facility SNAKE (Superconducting nanoscope for applied nuclear (kern) experiments) [19] situated at the 14 MV tandem accelerator of the Maier–Leibnitz-Laboratorium (http://www.bl.physik.uni-muenchen.de). Cells were irradiated with 100 MeV oxygen or 55 MeV carbon ions in line patterns (lateral distance between ion hits 0.7 μm, distance between lines 6 μm; this results in an average dose for one line wise irradiation of 17 and 12 Gy, for oxygen and carbon ions, respectively). The time intervals, Δt, between two irradiations were measured from the end of the earlier irradiation to the end of the later irradiation. The incubation time intervals, t inc, were measured from the time point when the samples were put back into the incubator. If not indicated otherwise, t inc was 30 min.
Antibodies and immunofluorescent detection of foci
For immunofluorescence, cells were fixed in 2% para-formaldehyde as described [16]. Cells were emerged with vectashield and covered with a cover slip. For mounting, the sandwich consisting of mylar foil, cells and cover slip was cut from the cell chamber using a hot steel needle, placed on a glass slide and sealed with nail polish. Primary mouse antibodies directed against γ-H2AX (Upstate #05-636; 1:350), 53BP1 (Upstate #05-726; 1:400), Phospho-ATM (Ser1981) (Chemicon #24090816; 1:250), and Rad51 (Oncogene # NA71; 1:100), and rabbit antibodies directed against γ-H2AX (Upstate #07-164; 1:200 or Novus Biologicals #NB100-384; 1:200), 53BP1 (Novus Biologicals #NB100-305; 1:500), Mdc1 (Abcam #ab11169; 1:100 or Novus Biologicals #NB100-395; 1:100), and Rad51 (Calbiochem #PC130; 1:100) were used. Secondary antibodies labeled with Cy3 or Alexa488 were purchased from Molecular Probes or Dianova.
Microscopy and image processing
The image acquisition was done with an inverse epifluorescence microscope (Zeiss Axiovert 200 M) using a Zeiss Plan Apocromat 63x/1.4 objective, the filter sets SP-100 for Dapi, SP-101 (Alexa488) and SP-102 (Cy3) from Chroma Technology Corp., and a Spot RT900 CCD camera from DIAGNOSTIC Instruments. Alternatively a Zeiss Axiovert 200 M microscope with a Zeiss LCI Plan Neofluar 63x/1.3 objective, the Zeiss filter set 01 (Dapi), 38 (Alexa488), 20 (Cy3) and a Zeiss AxioCam MRm was used. Z-stacks with slice distances of 0.25 μm were grabbed and deconvoluted with Huygens deconvolution software (Scientific Volume Imaging) using the cmle (classical maximum likelihood estimation) or the qmle (quick maximum likelihood estimation) algorithm with a maximum of 40 iterations and a “quality criterion” of 0.01. Images shown in Figs. 1 and 5 were adjusted using the “auto levels” function in Photoshop.
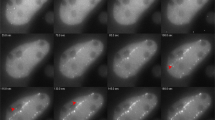
γ-H2AX foci and 53BP1 foci after sequential irradiation. a HeLa cells were irradiated sequentially (Δt = 50 min) with 100 MeV oxygen ions in line patterns perpendicular to each other and fixed after an incubation time t inc = 30 min after the later irradiation. Microscopic images (b, c) show cells that received only in the earlier irradiation (quadrant III), only the later irradiation (I), both irradiations (IV) or none (II). Shown is one microscopic slice from the cell equator, after deconvolution. Crosslike foci pattern are seen after immunostaining for γ-H2AX foci (b) but not for 53BP1 (c) in quadrant IV
Quantification of repair foci at e- and l-sites
A detailed description of data analysis can be found in the accompanying paper [20]. For quantification of the distribution of fluorescence intensity to e- and l-sites we determined for each nucleus I e and I l, the integrated fluorescence intensity per line length of foci at lines of e- and l-sites, after subtracting the mean residual fluorescence intensity, I UD, found in undamaged areas of the investigated nucleus. The normalized difference
describes the competition effect. u is zero if both irradiation have the same fluorescence intensity. If the signal of the earlier irradiation is brighter than the later one, u is negative. u is minus one if I l is zero. Vice versa, u is positive if foci induced by the later irradiation are brighter.
For statistical evaluation of a set of cells the u parameters are entered into a histogram. The fit with a Gaussian results in a mean value \( \bar u \) and the statistical error of the mean. Using the u parameter for statistical evaluation is more robust because singularities are avoided, in contrast to using the ratio of line intensities w = I l/I e By inversion of equation (1) an averaged value for the ratio of line intensities \( \overset{\lower0.5em\hbox{$\smash{\scriptscriptstyle\frown}$}}{w} \) can be calculated:
To compare I UD values from different samples, this value has to be normalized with respect to the mean fluorescence intensity of the whole nucleus \( \bar I_{{{\text{nucl}}}} . \) Thus we define the normalized mean residual fluorescence intensity:
Results
Using the ion microbeam SNAKE, HeLa cells were irrradiated with single carbon or oxygen ions in a line pattern. After irradiation the cells were incubated for a time interval, Δt, before a later irradiation with a line pattern perpendicular to the earlier one was performed (Fig. 1a). In addition to a central irradiation field where cells received both irradiations, neighbouring irradiation fields were generated, where cells received only the earlier or only the later irradiation. After incubation for a second time interval t inc (usually 30 min), cells were fixed and phosphorylated histone H2AX was visualized by indirect immunofluorescence. Phosphorylation of H2AX (to yield γ-H2AX) occurs within minutes after damage induction in megabase-sized regions flanking DSB sites [3]. After sequential irradiation, the patterns of γ–H2AX foci closely reflect the patterns of the irradiation (Fig. 1b).
To compare the signal intensities of foci generated at sites damaged in the earlier irradiation (e-sites) and in the later irradiation (l-sites), a method was used which is described in detail in [20]. The normalized difference \( u = \frac{{I_{{\text{l}}} - I_{{\text{e}}} }}{{I_{{\text{l}}} + I_{{\text{e}}} }} \) between the line fluorescence intensities I l and I e of the later and the earlier irradiations, respectively, was determined in microscopic images. While at short incubation intervals (Δt < 10 min) γ-H2AX foci signals from both irradiation treatments show similar intensity \( (\bar u = 0), \) the intensity ratio of l- and e-sites increases for Δt between 10 and 90 min (Fig. 2). At least up to Δt = 4 h, the signals of the later irradiation were found to be about 50% higher than the signals of the earlier irradiation \( (\bar u = + 0.2). \) This finding is compatible with repair-associated loss of γ-H2AX signals over time.
Normalized averaged difference \( \bar u \) in fluorescence intensity between foci induced by earlier and later irradiation, depending on the time interval Δt between both irradiations. Indicated are the mean value and its standard error of 15–80 cells per data point. Cells were irradiated with 100 MeV oxygen ions (circles) or 55 MeV carbon ions (squares), and γ-H2AX foci (open symbols) and 53BP1 foci (closed symbols) were detected simultaneously by immunofluorescence at t inc = 30 min. For comparison, the right ordinate depicts the corresponding mean quotient \( \hat w = \mathop {\left( {\frac{{I_{{\text{l}}} }}{{I_{{\text{e}}} }}} \right)}\limits^{ \wedge } \) of the line intensities
In sharp contrast to γ-H2AX, the formation of 53BP1 foci was strongly diminished at l-sites in those cells which had experienced an earlier irradiation, but not in cells that had not been pre-irradiated (Fig. 1c). Quantitative evaluation showed that for 53BP1, I l /I e is reduced to about 1/3 \( (\bar u = - 0.5) \) when the time interval, Δt, between both irradiations was between 40 min and at least 4 h (Fig. 2). We conclude that the earlier irradiation treatment induced a process that somehow resulted in a reduction of 53BP1 accumulation at l-sites. We call this effect competition effect. This effect was not observed for incubation times t inc > 150 min (data not shown).
The integrated 53BP1 fluorescence intensity, normalized with respect to the total nuclear area, was very similar in unirradiated and irradiated nuclei (Fig. 3), supporting the absence of damage-induced protein expression [11]. After irradiation, the residual fluorescence intensity J UD detected in nuclear regions outside the irradiation-induced foci, normalized with respect to the mean intensity of the nucleus, was only about 20% (Fig. 4). The mean value is determined to 0.225 ± 0.007 (SEM), the maximum of the distribution is found at 0.155 ± 0.014. This indicates that the major part of the 53BP1 pool accumulates at DSB sites. Therefore, impaired foci formation after a later irradiation might be explained by strong binding of 53BP1 to chromatin at e-sites and exhaustion of free 53BP1. A rather immobile fraction of 53BP1 at damaged chromatin with a long residence time compatible with this model (at least 30 min) was reported from one group [21], but others have reported much shorter residence times [22].
Constant amount of 53BP1 in cell nuclei, independent of irradiation. Indicated are the mean and the standard error of the mean of the integrated fluorescence signal in each cell nucleus normalised to the area of this nucleus for four sets of cells of one sample (Δt = 124 min; t inc = 30 min): unirradiated cells (cells from region II in Fig. 1), cells irradiated once with the earlier irradiation (region III), once with the later irradiation (region I) and cells irradiated twice (region IV). The data are normalised that the value for unirradiated cell is one. Details about data analysis can be found in [20]
Residual 53BP1 fluorescence intensity. The plot shows the distribution of normalized residual fluorescence intensity found in undamaged regions of about 600 crosswise irradiated cells. Due to the normalization with respect to the mean fluorescence intensity in the nucleus this fluorescence intensity may be interpreted as the fraction of unbound, free available 53BP1. By fitting with a Gaussian the maximum of the distribution is determined to 0.155 ± 0.014, the mean value is 0.225 ± 0.007 (SEM)
At residence times τ much shorter than the incubation time t inc, however, the intensity ratio I l /I e has to be modelled by the ratios of residence times τ l /τ e and of the binding sites n l /n e at l- and e-sites. The unequal distribution of 53BP1 to e- and l-sites can be described with a simple model working on the following assumptions:
-
The release of 53BP1 from a binding site is a statistical process characterized by a release rate 1/τ i, where τ i is the mean residence time of 53BP1 at binding sites on foci of the earlier (i = e) or later (i = l) irradiation. Thus, the total release rates from binding sites are A i /τ i, where is A i the amount of 53BP1 bound to DSB sites induced by the earlier or later irradiation.
-
A released 53BP1 protein binds immediately back to a binding site of earlier or later irradiation with the probabilities p e and p l. The p i’s are given by the ratios \( p_{{\text{i}}} = \frac{{N_{{\text{i}}} }}{{N_{{\text{e}}} + N_{{\text{l}}} }} \), following p e + p l = 1. N i is the number of accessible bindings sites induced by the earlier or later irradiation, which is the product of the number of DSB m i and the number of binding sites per DSB n i: N i = n i m i. Unbound 53BP1 was shown in Fig. 4 to be a negligible amount of total 53BP1. Thus, the number of 53BP1 proteins is much smaller than the number of binding sites, which is a prerequisite of this second assumption.
With these assumptions the amounts A e and A l are described by the following rate equations:
After a period of time much bigger than the residence times τ e and τ l, a steady state is reached and the amounts A e and A l change no longer; then from Eq. (4) follows:
The line intensities I e and I l are defined as the integrated fluorescence intensity around the irradiation lines normalized to the line length. Since the same line pattern is used for both irradiations, the numbers of ion hits per line length are equal and, neglecting repair processes, also the number of DSB per line length are equal. Thus, the ratio w = I l /I e is equal to the ratio of the fluorescence intensities per DSB, which corresponds to the ratio of the amount of 53BP1 bound to one DSB.
Thus, a threefold difference between the foci intensities I e and I l may result from a threefold difference either in residence times, τ, or in number of binding sites per DSB, n, at the respective damaged sites, or a combination of both factors.
How can residence times or binding sites differ at e- and l-sites? 53BP1 appears to interact in a bimodal way with chromatin in the vicinity of DSB sites: initial recruitment seems to be mediated by binding to a histone modification, the identity of which is under debate [9, 23], while subsequent retention of 53BP1 was shown to depend on the presence of γ-H2AX [24] and, in certain cell types, Mdc1 [22, 25].
Although in our hands Mdc1 is not required for 53BP1 foci formation in HeLa cells (data not shown), we investigated whether absent 53BP1 foci formation at l-sites was associated with absent Mdc1 foci formation. We found that Mdc1 foci formed readily at l-sites (Fig. 5a), with I l very similar to I e (Fig. 5d). Thus, reduced 53BP1 foci formation cannot be explained by the absence of Mdc1 or γ-H2AX from DSB sites induced in the later irradiation.
Mdc1, phospho-ATM(Ser1981), and Rad51-foci after sequential irradiation. HeLa cells were irradiated sequentially with 100 MeV oxygen ions in a pattern as shown in Fig. 1a, with Δt = 45 min and t inc = 30 min. Cells received only the earlier irradiation (quadrant III), only the later irradiation (I), both irradiations (IV), or none (II). Crosslike foci pattern are seen in quadrant IV after immunostaining for Mdc1 (a) and phospho-ATM (b), but not for Rad51 (c). Note that panels a–c are from different experiments. Shown is one slice from the cell equator, after deconvolution, for panels (a, b), and, because of the limited size of the Rad51 foci, a z-projection of the whole nucleus for panel (c). Panel (d) depicts the mean and its standard error of the normalized averaged difference \( \left( {\bar u} \right) \) obtained from at least 25 cells per data point, with 37 min ≤ Δt ≤ 63 min
It has been proposed that Trrap, a component of several histone acetyl transferase (HAT) complexes, may be involved in regulating chromatin accessibility in regions surrounding DSB sites [26, 27]. In cells lacking Trrap, radiation-induced 53BP1 foci and Rad51 foci were found not to form, while accumulation of Mdc1 and active phospho-ATM (Ser1981) occurred normally [26]. We therefore tested whether accumulation of phospho-ATM (Ser1981) differed between DSB sites induced in earlier and later irradiations and found no difference (Fig. 5b, d). In contrast, Rad51 showed a similar competition effect as did 53BP1: while Rad51 foci built readily at DSB sites formed during the earlier irradiation, they were not visible at DSB sites induced in the same cells at a later irradiation series (with Δt = 45 min), although they were seen in those cells which were only exposed to the second irradiation (Fig. 5c, d).
Discussion
Using an ion microbeam approach, we were able to produce by sequential irradiation line patterns of damaged chromatin in individual nuclei that were clearly distinguishable from each other. We show for the first time a competition effect of factors in the DSB response pathway where these factors act differentially on sites damaged upon sequential irradiation series. With regard to the mechanism, a difference between the foci intensities I e and I l may result from respective differences either in residence times, τ, or in number of binding sites per DSB, n, or a combination of both factors. Published data on the residence times of 53BP1 at damaged chromatin are conflicting [21, 22]; therefore, it is not clear at present whether the residence times at e-sites are long enough to fully explain the observed competition effect.
Concerning the possibility that the number of binding sites for 53BP1 at l-sites is smaller than at e-sites, the very similar effects conferred by Trrap deficiency and pre-irradiation of cells raise the possibility that a step in the Trrap-mediated pathway is blocked upon pre-irradiation. It is conceivable that binding sites for 53BP1 (e.g. histone modifications H3K79me2 or H4K20me2 [9, 23]) become exposed upon Trrap-mediated histone acetylation in the vicinity of DSB sites under normal irradiation conditions, but not under the conditions of a later irradiation as applied here. Our data are thus compatible with the existence of a chromatin-dependent damage response pathway [8, 26] in which 53BP1 plays a central role [10, 14]. To what extent 53BP1 and the recombination factor Rad51 are functionally related remains to be established. 53BP1 foci formation precedes foci formation of Rad51 (which depends on prior single-strand processing) and may be involved in ensuring right conditions for Rad51 binding. Alternatively, insufficient chromatin accessibility may inhibit processing of DNA ends and exposure of single-strands.
Concerning the biological significance of a differential response in earlier and subsequent irradiations, proficient formation of phospho-ATM(Ser1981) foci at DSB induced in the later irradiation suggests that general signalling is not affected. Reduced Rad51 foci formation, however, may suggest that this differential response affects the choice of DSB repair pathways. The consequence of Rad51 inhibition may be that DSB repair has to occur via other and potentially more error-prone repair pathways. Our findings may, therefore, have implications for therapeutic radiation treatment with respect to fractionation and repetitive irradiation schemes.
References
Khanna KK, Jackson SP (2001) DNA double-strand breaks: signaling, repair and the cancer connection. Nat Genet 27:247–254
Lavin MF, Birrell G, Chen P, Kozlov S, Scott S, Gueveen N (2005) ATM signaling and genomic stability in response to DNA damage. Mutat Res 569:123–132
Sedelnikova OA, Pilch DR, Redon C, Bonner WM (2003) Histone H2AX in DNA damage and repair. Cancer Biol Ther 2:233–235
Carson CT, Schwartz RA, Stracker TH, Lilley CE, Lee DV, Weitzmann M (2003) The Mre11 complex is required for ATM activation and the G2/M checkpoint. EMBO J 22:6610–6620
Horejsi Z, Falck J, Bakkenist CJ, Kastan MB, Lukas J, Bartek J (2004) Distinct functional domains of Nbs1 modulate the timing and magnitude of ATM activation after low doses of ionizing radiation. Oncogene 23:3122–3127
Lee JH, Paull TT (2004) Direct activation of the ATM protein kinase by the Mre11/Rad50/Nbs1 complex. Science 304:93–96
Lee JH, Paull TT (2005) ATM activation by DNA double-strand breaks through the Mre11-Rad50-Nbs1 complex. Science 308:551–554
Bakkenist CJ, Kastan MB (2004) DNA damage activates ATM through intermolecular autophosphorylation and dimer dissociation. Nature 421:499–506
Huyen Y, Zgheib O, DiTullio RA, Gorgoulis VG, Zacharatos P, Petty TJ, Sheston EA, Mellert HS, Stavridi ES, Halazonetis TD (2004) Methylated lysine 79 of histone H3 targets 53BP1 to DNA double-strand breaks. Nature 432:406–411
Mochan TA, Venere M, DiTullio RA, Halazonetis TD (2003) 53BP1 and NFBD1/MDC1-Nbs1 function in parallel interacting pathways activating ataxia-telangiectasia mutated (ATM) in response to DNA damage. Cancer Res 63:8586–8591
Wang B, Matsuoka S, Carpenter PB, Elledge SJ (2002) 53BP1, a mediator of the DNA damage checkpoint. Science 298:1435–1438
Stewart GS, Wang B, Bignell CR, Taylor AMR, Elledge SJ (2003) MDC1 is a mediator of the mammalian DNA damage checkpoint. Nature 421:961–966
Peng A, Chen PL (2003) NFBD1, like 53BP1, is an early and redundant transducer mediating Chk2 phosphorylation in response to DNA damage. J Biol Chem 278:8873–8876
Zgheib O, Huyen Y, DiTullio RA, Snyder A, Venere M, Stavridi ES, Halazonetis TD (2005) ATM signaling and 53BP1. Radiother Oncol 76:119–122
DiTullio RA, Mochan TA, Venere M, Bartkova J, Sehested M, Bartek J, Halazonetis TD (2002) 53BP1 functions in an ATM-dependent checkpoint pathway that is constitutively activated in human cancer. Nat Cell Biol 4:998–1002
Hauptner A, Dietzel S, Drexler GA, Reichart P, Krücken R, Cremer T, Friedl AA, Dollinger G (2004) Microirradiation of cells with energetic heavy ions. Radiat Environ Biophy 42:237–245
Reichart P, Datzmann G, Hauptner A, Hertenberger R, Wild C, Dollinger G (2004) Three-dimensional hydrogen microscopy in diamond. Science 306:1537–1540
Dollinger G, Hable V, Hauptner A, Krücken R, Reichart P, Friedl AA, Drexler GA, Cremer T, Dietzel S (2005) Microirradiation of cells with energetic heavy ions. Nucl Instr Meth B 231:195–201
Datzmann G, Dollinger G, Goeden C, Hauptner A, Körner HJ, Reichart P, Schmelmer O (2001) The Munich microprobe SNAKE: first results using 20 MeV protons and 90 MeV sulfur ions. Nucl Instr Meth B 181:20–26
Greubel C, Hable V, Drexler GA, Hauptner A, Dietzel S, Strickfaden H, Baur I, Krücken R, Cremer T, Friedl AA, Dollinger G (2008) Quantitative analysis of DNA-damage response factors after sequential ion microirradiation. Radiat Environ Biophys (submitted)
Pryde F, Khalili S, Robertson K, Selfridge J, Ritchie AM, Melton DW, Jullien D, Adachi Y (2005) 53BP1 exchanges slowly at the sites of DNA damage and appears to require RNA for its association with chromatin. J Cell Biol 118:2043–2055
Bekker-Jensen S, Lukas C, Melander F, Bartek J, Lukas J (2005) Dynamic assembly and sustained retention of 53BP1 at the sites of DNA damage are controlled by Mdc1/NFBD1. J Cell Biol 170:201–211
Botuyan MV, Lee J, Ward IM, Kim JE, Thompson JR, Chen J, Mer G (2003) Structural basis for the methylation state-specific recognition of histone H4-K20 by 53BP1 and Crb2 in DNA repair. Cell 127:1361–1373
Celeste A, Fernandez-Capetillo O, Kruhlak MJ, Pilch DR, Staudt DW, Lee A, Bonner RF, Bonner WM, Nussenzweig A (2003) Histone H2AX phosphorylation is dispensable for the initial recognition of DNA breaks. Nat Cell Biol 5:675–679
Bekker-Jensen S, Lukas C, Kitagawa R, Melander F, Kastan MB, Bartek J, Lukas J (2006) Spatial organization of the mammalian genome surveillance machinery in response to DNA strand breaks. J Cell Biol 173:195–206
Murr R, Loizou JI, Yang YG, Cuenin C, Li H, Wang ZQ, Herceg Z (2006) Histone acetylation by Trrap-Tip60 modulates loading of repair proteins and repair of DNA double-strand breaks. Nat Cell Biol 8:91–99
Downey M, Durocher D (2006) Chromatin and DNA repair: the benefits of relaxation. Nat Cell Biol 8:9–10
Acknowledgments
We thank the staff of the Maier–Leibnitz-Laboratorium for operating the accelerator and F. Eckardt-Schupp, GSF Research Centre Neuherberg, for providing access to the α-irradiation facility. This work was supported by grants of the European Science Foundation (ESF) under the EUROCORES Programme EuroDYNA (G.D., T.C., A.A.F.), the DFG Cluster of Excellence Munich Centre for Advanced Photonics (G.D., A.A.F.), the Bundesamt für Strahlenschutz (A.A.F.) and the Bundesministerium für Bildung und Forschung (A.A.F.).
Author information
Authors and Affiliations
Corresponding author
Additional information
Christoph Greubel, Volker Hable and Guido A. Drexler contributed equally to this work.
Rights and permissions
About this article
Cite this article
Greubel, C., Hable, V., Drexler, G.A. et al. Competition effect in DNA damage response. Radiat Environ Biophys 47, 423–429 (2008). https://doi.org/10.1007/s00411-008-0182-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00411-008-0182-z