Abstract
Purpose
Validated biomarkers of food intake (BFIs) have recently been suggested as a useful tool to assess adherence to dietary guidelines or compliance in human dietary interventions. Although many new candidate biomarkers have emerged in the last decades for different foods from metabolic profiling studies, the number of comprehensively validated biomarkers of food intake is limited. Apples are among the most frequently consumed fruits and a rich source of polyphenols and fibers, an important mediator for their health-protective properties.
Methods
Using an untargeted metabolomics approach, we aimed to identify biomarkers of long-term apple intake and explore how apples impact on the human plasma and urine metabolite profiles. Forty mildly hypercholesterolemic volunteers consumed two whole apples or a sugar and energy-matched control beverage, daily for 8 weeks in a randomized, controlled, crossover intervention study. The metabolome in plasma and urine samples was analyzed via untargeted metabolomics.
Results
We found 61 urine and 9 plasma metabolites being statistically significant after the whole apple intake compared to the control beverage, including several polyphenol metabolites that could be used as BFIs. Furthermore, we identified several endogenous indole and phenylacetyl-glutamine microbial metabolites significantly increasing in urine after apple consumption. The multiomic dataset allowed exploration of the correlations between metabolites modulated significantly by the dietary intervention and fecal microbiota species at genus level, showing interesting interactions between Granulicatella genus and phenyl-acetic acid metabolites. Phloretin glucuronide and phloretin glucuronide sulfate appeared promising biomarkers of apple intake; however, robustness, reliability and stability data are needed for full BFI validation.
Conclusion
The identified apple BFIs can be used in future studies to assess compliance and to explore their health effects after apple intake. Moreover, the identification of polyphenol microbial metabolites suggests that apple consumption mediates significant gut microbial metabolic activity which should be further explored.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Biomarkers of food intake (BFI) are extensively used to assess dietary intake, nutritional status, and to measure biological and health effects of specific diets and dietary components [1]. In particular, BFI are useful tools not only to assess compliance of dietary intake in intervention studies, but also to indicate metabolic pathways induced by the test diet or dietary components. In this framework, different “omics” techniques, including metabolomics and genomics, revolutionized research perspectives of dietary intervention and nutrition. To date, metabolomic techniques have proved to be a valuable tool to assess short- and long-term effects of food intake on the metabolome of biological fluids, although further work is required. Indeed, many new candidate biomarkers have emerged over the last decade from metabolic profiling studies; however, the number of comprehensively validated biomarkers of food intake is limited [2].
Apples are a rich source of polyphenols and fiber and these are considered the principal mediators of the protective health effects reported in epidemiological [3] and dietary intervention studies [4]. The main polyphenolic classes in apples, in increasing order, include dihydrochalcones, flavonols, hydroxycinnamates and flavanols [catechin and proanthocyanidins (PAs)] [5]. The most recognized apple polyphenols are phloridzin and phloretin glycosides. Apples are also rich in insoluble fiber, including cellulose and hemicellulose, while pectin is the major soluble fiber containing homogalacturonans and rhamnogalacturonans. Nonglycosylated polyphenols present in large amounts in apples, such as monomeric and to a lesser extent dimeric flavanols, can be directly absorbed in the small intestine [6]. Prior to passage into the circulation, polyphenols undergo structural modifications in the small intestine and later in the liver through the conjugation process. This involves glucuronidation, methylation and sulfation by phase II enzymes and is part of the metabolic detoxification process common to xenobiotic compounds, which increases their solubility and thus making them easier to excrete by urine or bile [6]. In contrast, polymeric flavanols (proanthocyanidins, PAs), the major polyphenolic class in apples, reach the colon nearly intact where they are metabolized by the colonic microbiota into hydroxyphenylvalerolactones and subsequently to phenolic acids, including phenylpropionic, phenylacetic, and benzoic acid derivatives [4]. These microbial-derived metabolites may have local intestinal effects within the gut or can be further absorbed with systemic effects [4]. A better understanding of the role of the gut microbiota are important, to explain the observed variability (7.5 ×) in the bioavailability of apple procyanidins among subjects, measured by the quantitative excretion of conjugated phenyl-γ-valerolactones and phenyl-γ-valeric acids [7].
Possible biomarkers of apple intake have been identified in animal and human intervention studies and these were extensively discussed in our previous review [8]. Most of these studies focused on the health benefits of apples, where biological fluids were analyzed to determine candidate biomarkers of intake [9,10,11,12,13,14,15]. Three studies aimed explicitly at discovering biomarkers of intake and/or assessing the impact of apple consumption on biological fluid metabolomes in humans [16,17,18] and two in rats [19, 20]; none, however, reported a correlation between two advanced “omics” techniques. Phloretin and its glucuronide and sulfate conjugates are the most frequent biomarkers found in human biofluids after apples or apple product intake. Its plausibility and specificity as BFI after apple consumption were recently discussed [8]. Additionally a dose–response relationship was found by Auclair et al. [15] and Saenger et al. [21] for urinary excretion of phloretin, after different doses of apples and polyphenol intakes. The large inter-individual variability among humans in response to food consumption is an important consideration in terms of understanding how foods and diets impact on human health. It is not clear whether everyone, regardless of age, gender, lifestyle, gut microbiota composition, or genotype, responds similarly to food consumption [22, 23]. These research questions require further investigations in human intervention and large prospective cohort studies in which an accurate assessment of certain food intake can be collected for every subject. For these reasons, the need to systematically and critically assessing the validity of BFI of food intake was highlighted in a series of papers and guidelines published under the Food Biomarker Alliance Project (FoodBAll Project) within the Joint Programming Initiative “A Healthy Diet for a Healthy Life” [24].
Using an untargeted metabolomics approach, we aimed to identify biomarkers of prolonged apple intake (Renetta Canada) and to explore the metabolic changes induced by the dietary intervention that could reflect an impact on host health. Renetta Canada is an apple variety rich in polyphenols which has been previously shown to decrease total and LDL-cholesterol, to improve vascular function in human subjects [25] and beneficially modulate the gut microbiota composition in vitro [26]. Secondarily, we correlated biomarkers found to be statistically significant after apple consumption with fecal microbiota taxa at the genus level. Finally, within the framework of the BFI validation protocol, we combined the information of the biomarkers of apple consumption obtained in this long-term intervention, with the results of a previous postprandial kinetic study [27], to evaluate the plausibility, reproducibility and time–response criteria for phloretin glucuronide and phloretin glucuronide sulfate, which appear promising biomarkers of apple intake.
Materials and methods
Study design
Long-term apple intake study—AVAG
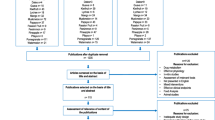
Details of the AVAG apple intervention study have been published previously [25]. Briefly, the trial was a randomized, controlled, crossover, dietary intervention in 40 mildly hypercholesterolemic (total cholesterol levels > 5.2 mmol/L) and otherwise healthy volunteers (23 women, 17 men,), aged 29–69 years, with BMI 19–33 kg/m2. Volunteers were recruited from the general population living in Reading, Berkshire, UK, and the study was conducted at the Hugh Sinclair Unit of Human Nutrition, University of Reading. The study was given a favourable ethical opinion for conduct by the University of Reading Research Ethics Committee [UREC Project No. 13/22 (AVAG study)]. The study was registered at clinicaltrials.gov (NCT01988389) and conducted according to the guidelines laid down in the Declaration of Helsinki, with written informed consent provided by all participants prior to starting the study. Upon informed consent and screening (inclusion/exclusion criteria are described in detail in [25]), subjects were randomly allocated to one of two groups to receive daily either two whole apples (Renetta Canada, 340 g containing 990 mg polyphenols and 8.5 g total fiber) or a sugar and energy-matched control beverage low in apple polyphenols (100 mL concentrated apple beverage containing 2.5 mg polyphenols and no fiber). Clear instructions were given to eat the apple skin and flesh and not to consume the apple core. Detailed description of the nutritional composition and polyphenol analysis of the whole apples and the control beverage is reported in Koutsos et al. [25] Prior to the first intervention period, subjects performed a 2 week run-in period, where they maintained their habitual diet but refrained from eating probiotics, prebiotics and any apples, apple juice or apple-containing foods. The first group (n = 22) consumed two apples daily for 8 weeks, and after a 4-week washout period, a sugar and energy-matched control apple beverage daily for an additional 8 weeks. The second group (n = 18) received the intervention in the reverse order (first the control beverage and then the two apples) (Fig. 1). Subjects were requested to incorporate the intervention products, apples or control beverage, into their normal diet without changing further their habitual intake. Biological samples, including fasted plasma, fecal samples and 24 h urine collections were collected before and after each dietary intervention at week 1, 8, 12 and 20.
Acute postprandial kinetic apple intake study–AGER
The second dietary intervention, we refer to as postprandial nutrikinetic study, was a randomized postprandial kinetic study involving 12, non-smoking, healthy volunteers (8 men and 4 women), aged 21–42 years; more details have been published elsewhere [27]. A venous blood sample was taken at time 0, and immediately after, participants were provided with a glass of cloudy apple juice (CAJ) (250 mL) or a glass of polyphenol-enriched cloudy apple juice (PAJ) (250 mL)—both prepared from Crispy Pink apples. The order of treatment allocation was randomly assigned: six subjects started with CAJ and the other six with PAJ. Further blood samples were taken at 1, 2, 3 and 5 h after juice consumption. Urine was collected at time 0 and between 0 and 2 h, 2 h and 5 h, 5 h and 8 h, and 8 h and 24 h after juice consumption. Volunteers were instructed to refrain from consuming phenol-rich foods and beverages (wine, coffee, tea, fruits and vegetables), dietary supplements and medications in the 3 days prior to the experiments. To ensure adherence to the dietary instruction, we asked the subjects to keep a 3-day dietary record prior to the study participation [27]. Schemes describing both intervention studies are depicted in Fig. 1.
Chemicals and reagents
HPLC-grade methanol, acetonitrile, 2-propanol and formic acid were obtained from Sigma Aldrich. The ultrapure water was obtained by purifying demineralized water in a Milli-Q system from Millipore (Bedford, MA, USA). Internal standards creatinine-labeled 13C was purchased from Sigma-Aldrich and d5-labeled trans-cinnamic acid, d4-labeled chenodeoxycholic acid and d5-labeled taurocholic acid were obtained from CDN ISOTOPES, Inc. (Pointe-Claire, Quebec, Canada). 96-well plates with PVDF filters were obtained from Millipore, and Ostro protein precipitation plate from Waters (USA). Internal standards were prepared in pure methanol.
Biological sample preparation for metabolomic analysis
Urine and plasma samples were thawed at 4 °C overnight. Extraction methods for serum and urine have been described elsewhere [28]. Briefly, 50 µL of serum was loaded onto 96-well plates Ostro (Waters), and 50 µL of internal standard in methanol was added followed by 150 µL of cold acetonitrile with 1% formic acid. The mixture was shaken for 5 min and samples were filtered using a positive pressure-96 manifold (Waters, USA). Additionally, the filtering plate was further eluted with 150 µL of cold acetonitrile with 1% formic acid, vortexed and filtered again using the same procedure. Samples were evaporated with a gentle stream of nitrogen to dryness using a Techne Dr-block DB 3D heater at room temperature and re-dissolved with 50 µL of water and 50 µL of external standard (trans-cinnamic acid-d5).
100 µL of urine was placed into 96-well plate Millipore PVDF and 100 µL of internal standards in methanol was added. After 5 min of shaking, the mixture was filtered using a positive pressure-96 manifold. Then, 300 µL of external standard (trans-cinnamic acid-d5) was added to the collection plate. The prepared urine and serum samples were submitted to LC–Orbitrap–MS analysis.
Quality control
Quality control (QC) samples were prepared for each biological fluid by mixing equal volumes of each sample. Samples were fully randomized for extraction and injections, and experiments were blinded to the operator. A double QC injection was performed every eight study samples for monitoring of injection performance. A series of ten QC samples was injected at the beginning of the injection queue to equilibrate the LC column.
Chromatographic and mass spectrometry conditions
Samples were analyzed by a hybrid linear ion trap Fourier Transform (LTQ FT) Orbitrap mass spectrometer (Thermo Fisher, Bremen, Germany) interfaced to a Dionex HPLC system, consisting of an auto-sampler and quaternary gradient HPLC pump. Chromatographic separation of compounds was performed using a Kinetex C18 column (150 mm × 2.1 mm I.D., particle size 3.5 µm) with pre-column 4.0 mm × 2.0 mm I.D (Phenomenex Torrance, CA, USA). Details regarding chromatographic conditions for plasma and urine samples have been published elsewhere [27].
The Orbitrap LTQ was equipped with an electrospray ionization (ESI) probe and operated in both positive and negative ionization modes. The conditions have been described elsewhere [28, 29]. Briefly, samples were injected twice. The first injection was dedicated to acquisition of full scan spectra at resolution of 30,000, while the second injection was dedicated to acquisition of high-resolution MS/MS data under data-dependent acquisition (DDA) mode. In DDA mode, the resolving power for MS scan was 7500 and for MS2 scans 7500. The static exclusion list consisted of 300 most abundant ions created by the injection of solvents, which followed the same preparation procedures as the samples. Dynamic exclusion allowed three repeated counts of the same ion in 15 s, while the exclusion duration was 45 s. The sequences were randomized with regard to participants and treatments. For every ten samples, a quality control block was analyzed consisting of solvents and double injection of the quality control sample.
Data treatment and biomarker identification
Raw files were converted to mzXML format with the msconvert utility included in ProteoWizard [30]. Profiling of the data was processed with XCMS [31] using the ‘‘matchedFilter’’ peak picking method. A signal to noise ratio cutoff of 4 was used, full width at half maximum was set to 10 s and a step size of 0.005 Da was used for extracting chromatograms. The resulting feature matrix was annotated using CAMERA [32] to group features corresponding to the same parent ion species.
To identify the features showing a significant difference between the meals, two independent linear mixed models were fitted to each mass feature. In the first model, the concentration of each feature was modeled as a function of meal, the gender of the subject and the individual. Interactions were tested for significance and left out, since they were non-significant for all features. In the second model, the effect of meal was left out. The comparison of the two models allowed us to identify the features significantly affected by the meal factor. The p values were obtained using likelihood ratio tests and the collection of p values was corrected for multiple testing by controlling the false discovery rate (FDR) and the q values were calculated [33, 34]. Post hoc multiple comparison tests (p values uncorrected) were performed to determine the specific effects between means and the collection of p values corrected as above. Urine samples were normalized to the creatinine content. For the plasma samples, feature-wise batch correction was performed such that that the mean intensity for each batch was the same. The R scripts used for pre-processing are available at https://gitlab.com/metabolomics-analyses/avag
Markers contributing to the discrimination between two treatments were identified through a multiple-step procedure. Molecular ions and in-source molecular fragments were assigned as one compound based on a mass accuracy approach and peak shape, as many were statistically significant. Discriminative markers were then compared with the monoisotopic molecular weight, chemical structures and LC–MS/MS spectra of metabolites proposed by freely available databases: m/z Cloud (www.mzcloud.org); Human Metabolome DB [35], METLIN [36], MassBank [37] and LIPID MAPS [38] databases. Mass accuracy was set to 2 mDa while searching online. Additionally, information from MSn experiments were introduced to Met-Fusion to get candidate structures [39]. Final identification was achieved after a combination of LC–HRMS2, LC–HRMS3 experiments, online database information and literature verification. The levels of identification reported in Tables 1 and 2, and in Supplementary Materials Tables 1 and 2 are as follows: level I corresponds to compounds identified by matching masses and retention times with authentic standards in the laboratory using the same analytical method; level II corresponds to compounds identified by LC–(HR)–MS, LC–(HR)MS2 and/or LC–(HR)MSn spectra and matched to spectra from databases and literature. Compounds identified only by spectral similarities to a similar compound class and literature knowledge are reported as level III. Unknown compounds are reported as level IV.
Metabolomic data sharing
The study participants gave written consent to pseudonymized data sharing. All untargeted data in mzXML format and metadata are available for download from the MetaboLights public repository [40]. The AVAG data are deposited in MetaboLights with the persistent unique public identifier MTBLS469, https://www.ebi.ac.uk/metabolights/MTBLS469. A more detailed set of meta-data of the study is also made available in the Dash-in Web-based federated analyses website (https://www.enpadasi.eu/deliverables_final_pdf/D3.2.1.pdf). Metadata in Dash-in are compliant with the need to be extensively annotated in human- and machine-readable form https://dashin.eu/interventionstudies/study/show/38790886.
Fecal sample preparation for DNA extraction and PCR amplification of the V3–V4 region of bacterial 16S rRNA
The fecal DNA extraction, PCR amplification and the Illumina next-generation sequencing of the V3–V4 region of the bacterial 16 rRNA have been described previously by Koutsos et al. [26]. Sequences obtained from Illumina sequencing were processed using Quantitative Insights Into Microbial Ecology (QIIME) software package version 1.9 [41]. Briefly, reads were assigned to each sample according to the unique barcode; forward and reverse Illumina reads (300 bp each) were joined using the fastq-join method [42], quality filtering was performed using 19 as the minimum Phred quality score, chimeric sequences were identified using usearch 6.1 and removed. Then, sequences were assigned to operational taxonomic units (OTU) using the QIIME implementation of UCLUST algorithm at 97% similarity threshold [43]. Representative sequences for each OTU were assigned to different bacterial taxonomic levels—phylum (p.), class (c.), order (o.), family (f.) and genus (g.)—by using Greengenes database release (May 2013).
Correlation analysis between metabolites and microbial taxa
Spearman's rank-order correlation test was applied pairwise to each combination between 69 microbial taxa at genus level and 93 urine and 12 plasma metabolites. FDR adjustment was applied on all tests to correct for multiple testing. Statistical significance cutoff was established at level 0.05; however, results up to 0.07 were also included for further investigation considered as an important trend. The reason for such extended statistical significance consideration derives from previous experience [28] where microbial-derived biomarkers (catabolites, intermediates and final excreted products) belonging to the same metabolic trajectory did not always reach 0.05 cutoff level due to inter-individual variation.
Results
Annotated biomarkers in urine and plasma
The total number of statistically significant m/z features varied between matrices and ionization modes: in plasma: 16 ESI+ and 15 ESI−, and urine: 213 ESI+ and 145 ESI−. The list of discriminating m/z features in urine and plasma after the consumption of whole apples and control beverage is shown in Table 1 and 2, and Supplementary Material Tables 1 and 2. We found 61 putatively annotated biomarkers in 24 h urine and 9 in plasma, which were differentially abundant between the two intervention treatments (whole apples and control beverage). As expected, all the metabolites were found in higher abundance (p < 0.05) after the intake of whole apples compared to the control beverage. The biomarkers were classified into several groups, each containing biochemically related derivatives. Each group was built combining the information on the chemical class of their precursors (metabolites) or their common pathways of origin (for breakdown catabolites) (Tables 1, 2). Two metabolites of phloretin were found to be statistically significant in urine samples after apple intake, but not after the control beverage consumption, namely phloretin glucuronide and phloretin glucuronide sulfate (urine MU1-MU2). Five (epi)catechin metabolites were found in 24 h urine samples, after whole apple consumption (urine MU3–MU7). Fourteen phase II valerolactone metabolites with different hydroxylation patterns and conjugations were found to be statistically significant in urine, and two in plasma after apple intake, but not after intake of control beverage (urine MU8–MU22; plasma MP1–MP2). Feruloylquinic acid (MU23) and hydroxycinnamic acid (MU24–MU26) were found to be statistically significant in urine, but not in plasma, after apple intake. Four metabolites of phenylpropionic and phenylacetic acid with different hydroxylation patterns were found to be statistically significant in urine and two in plasma after apple consumption (urine MU27–MU32; plasma MP3–MP4). Five metabolites belonging to hippurate metabolic pathway were found to be significantly higher in urine and one in plasma after apple intake. Two indole compounds were significantly higher in urine after apple consumption, namely indolylacryloylglycine and indole-acetylaspartic acid, while tryptophan statistically increased in plasma (urine MU55–MU56, plasma MP9) (see Tables 1, 2, Fig. 2a, b). Fourteen fatty acids were found to be statistically significant after apple consumption in urine, and three in plasma (urine MU38–MU54; plasma MP6–MP8). Their accurate masses and fragmentation patterns were consistent with saturated and unsaturated fatty acid glucuronides, including three isomers of (OH)C13:0; four isomers of (OH)C13:2; two isomers of (OH)C15:2; and others (see Table 1 for entire list of fatty acids). Three metabolites of glutamine were found to be significantly higher in urine after apple consumption, namely hydroxyphenylacetylglutamine, hydroxyphenylacetylglutamine sulfate and N-acetylglutamic acid; no glutaminę conjugates were found in plasma (MU57–MU59). Box plots for each of the above-mentioned metabolites are available in Supplementary Material.
Correlations between metabolites and fecal microbiota
The relative abundance of the 69 fecal bacteria taxa identified in the fecal samples was correlated with the concentration of metabolites observed in urine and plasma after consumption of apples. Figure 3 shows a heat map of the correlation coefficients (Spearman’s ρ) of urine metabolites with fecal bacteria taxa related to the consumption of apples. Among the 14 strongest correlations, 6 were statistically significant (FDR < 0.05), while the remaining 8 had an FDR < 0.07; see Table in Fig. 3. Granulicatella was found to be inversely correlated with four metabolites, namely hydroxyphenylacetylglutamine, hydroxyphenylacetylglutamine sulfate, hydroxyphenylacetic acid and hydroxyphenylacetic acid sulfate. Glucuronide conjugated fatty acids C13 with different saturation states were positively correlated with Clostridiales. Finally, two isomers of dihydroxyphenyl propionic acid were negatively correlated with the genus Dehalobacterium and with an unclassified genus of the family Rikenellaceae. Correlations between plasma metabolites and bacterial genera after apple intake did not show any statistical significance after FDR adjustment (see Supplementary Materials Figs. 3–5).
Heat map of pairwise correlations between urine metabolites and gut microbiota at genus level. The correlation coefficients (Spearman’s ρ) are represented by color gradient and size of the circles. The top 14 significant (FDR < 0.07) are listed in the table below. For unclassified genera (“g_”), the bacterial family (“f_”) or order name (“o_”) is reported
Discussion
The most characteristic apple polyphenol is phloridzin which belongs to a family of chalcones. Detection of phloretin metabolites in biological fluids after apple intake is frequently used as indicator of compliance to dietary intervention. Indeed in our study, phloretin glucuronide and phloretin glucuronide sulfate were higher in urine after the whole apple intake compared to the control beverage. However, phloretin metabolites were also detected in eight baseline 24 h urine samples. For this reason, dietary records were manually verified for these participants and revealed the presence of foods that could be potentially adulterated with apple traces such as fruit juice smoothies and muesli with fruits. No phloretin metabolites were found in fasting plasma samples, after apple intake or after control beverage intake. In our previous study we found phloretin glucuronide and sulfates conjugates in plasma with Tmax at 1–2 h postprandial, therefore lack of such metabolites after a 12-h overnight fasting is not surprising [27].
Catechins belong to flavanols family, and are common in different classes of fruits and vegetables. Indeed, several studies reported the presence of (epi)catechin conjugates in urine after apple consumption [9, 21], as well as the consumption of cocoa [44], grape juice [45] or tea [46]. However, it has been suggested that (epi)catechin units may be also gradually released by the partial depolymerization of procyanidins during their transit along the intestinal tract [47, 48]. In our previous AGER study [27], we found seven metabolites of (epi)chatechins in postprandial urine samples, and the presence of five of these was confirmed in urine in this study. According to our previous study [27], plasma epi(catechin) metabolites have Tmax at 1–2 h after consumption [27], and as the plasma samples in AVAG study were collected after a 12 h overnight fast, (epi)catechin metabolites were not expected to be present in the plasma samples.
Valerolactone and valeric acid metabolites can arise from procyanidins and/or (epi)catechins which are commonly present in apples. Fourteen phase II valerolactone metabolites were higher in urine after the whole apple intake compared to the control beverage. These metabolites were also found in baseline samples, probably because procyanidins and (epi)catechins are commonly found in several fruits and vegetables, and participants could have consumed these in their habitual diet during the run-in and wash-out periods. The presence of valerolactones and valeric acid metabolites in urine and plasma supports our previous postprandial kinetic study and the literature [27], where a wide range of these compounds were found in biological fluids.
Chlorogenic acids and cinnamic acids are frequently found in biological fluids after apple consumption [8]. Feruloylquinic acid and hydroxycinnamic acid were found to be higher in urine after whole apple intake compared to the control beverage. Stalmach et al. [49] reported an excretion of coumaroylquinic acid and feruloylquinic acid in ileal fluid and urine of ileostomy subjects after coffee consumption, while Kahle et al. [9] and Hagl et al. [11] reported chlorogenic acids in ileal fluids after apple consumption. Similar results have been obtained in our previous study [27], in which chlorogenic and cinnamic acid metabolites were found in urine and in post-prandial plasma after apple juice consumption.
Four metabolites of phenylpropionic and phenylacetic acid were found to be statistically significant in urine and two in plasma after whole apple consumption. The presence of phenylpropionic and phenylacetic acid metabolites in blood and urine confirms that these are formed by the colonic microbiota, then absorbed, conjugated in the liver, transferred in the blood circulation, with potential systemic effects, and then excreted in urine [27]. This is in agreement with our previous study [27] and other studies reporting an increase in the above-mentioned phenolic acids after the intake of coffee and grape juice [50, 51], almonds [52, 53] and apples [14, 36].
Studies of Ulaszewska et al. [28], Trost et al. [27] and Norskov et al. [54] have previously reported several metabolites of hippurate after extensive feeding with fruits and vegetables, and wheat, while Kristensen et al. [19] found hippuric acid and 3-hydroxyhippuric acid in plasma of rats after apple consumption. Similarly, we have found five metabolites belonging to hippurate metabolic pathway in a higher concentration in urine and one in plasma after whole apple intake compared to the control beverage. Hippurate metabolic pathway intersects with the one responsible for metabolism of polyphenols; thus, hippuric acids would be considered as the final end products of several classes of polyphenols. For this reason, hippuric acid per se cannot be considered as a biomarker of a particular food intake; however, its intermediate metabolites such as cyclohexadiene carboxylic acid could result from different parent polyphenols as suggested by Ulaszewska et al. [28]. Moreover, hippuric acid as well as 3- and 4-hydroxyhippuric acids have been also identified as bacterial metabolite of polyphenols [55].
Fatty acids with different hydroxylation patterns are frequently found to be up- or downregulated during dietary interventions [27, 28, 56, 57]. Lack of commercially available standards only allowed annotation at level III of the identified fatty acids in urine and plasma after the whole apple intake. There is scant information explaining the nature of medium and long chain fatty acids with different levels of oxidation and we can only assume they were involved in the energy cycle where free fatty acids play a crucial role.
Indole metabolites deserve further scrutiny, since apple consumption has not been reported to modulate their concentration previously. We found higher abundance of indolylacryloylglycine and indole-acetylaspartic acid in urine after the whole apple intake compared to the control beverage. Indolylacryloylglycine is a regular constituent of human urine, although its origin is not completely understood. It is generally assumed that bacterial activity in the intestine could be an important source of further indolylacryloylglycine in human urine [58]. Tryphophan was in a higher abundance in plasma after the whole apple intake compared to the control beverage. Tryptophan metabolism follows three major pathways in the gastrointestinal tract: (1) direct transformation into indole derivatives, (2) kynurenine pathway through indoleamine 2,3-dioxygenase, (3) serotonin pathway through tryptophan hydroxylase [58]. In the gut, tryptophan can be catabolized, by the resident microbiota, to indole acid derivatives, including indole-3-acetic acid, indole-3-aldehyde, indole-3-lactic acid, and indole-3-acrylic acid (Fig. 2). Different microbiota members, including Bacteroides, certain Clostridia, and Escherichia coli, can catabolize tryptophan to tryptamine and indole pyruvic acid, which are then converted to indole-3-acetic acid, indole propionic acid, and indole lactic acid [59]. Indolyl propionic acid can be further converted to indolyl acrylic acid and combined with glycine to yield indolylacryloyl glycine in the liver or kidney [60]. The effects of several indolic acid derivatives on the gut microbiota and intestinal homeostasis have been reported. Several Peptostreptococcus species are capable of producing indole acrylic acid, which can suppress inflammation by promoting intestinal epithelial barrier function and mitigating inflammatory responses [61]. Other indole metabolites, such as indole-acetylaspartic acid could be a product of conjugation of aspartate with indoleacetic acid, as some bacteria possess this capability. Indole-acetylaspartic acid is better known as the phytohormone auxin, which regulates plant growth and development [62, 63]. To our knowledge, this is the first study where the concentration of these three metabolites was found to be increased in biological fluids after apple consumption. Their presence in urine, origin and biological significance require further investigation.
Hydroxyphenylacetylglutamine, hydroxyphenylacetylglutamine sulfate and N-acetylglutamic acid abundance was higher in urine after whole apple intake compared to the control beverage. However, these were characterized by high variability among individuals. Phenylacetylglutamine and hydroxyphenylacetylglutamine provide a major route of nitrogen excess excretion from the body. Phenylacetylglutamine is formed from the conjugation of phenylacetate (from host or gut microbiome) and glutamine (mainly generated from the Krebs cycle from α-ketoglutarate)—Fig. 2. Phenylacetate is primarily produced by the gut microbiome by the decarboxylation of phenylalanine in unabsorbed protein residue [64], while further phenylacetylglutamine is formed in the liver by glutamination of phenylacetate. In contrast, hydroxylated phenol-substituted acids (such as hydroxyphenylacetic acid) are known to be the main products of tyrosine fermentation in anaerobic bacteria [65]. As it has been shown by Russel et al. [66], hydroxyphenylacetic acid derives from tyrosine, through deamination and chain shortening by different microbiota species, including Bacteroidetes, and to a lesser extent, Firmicutes. Supporting this, apple consumption was associated with the presence of tyrosine metabolites in the study of Kristensen et al. [19], where methoxytyrosine was found in 24 h pooled urine of rats after prolonged consumption of apple pectins. To our knowledge, only one study on grape juice intake reported a higher excretion of phenylacetylglutamine after a dietary intervention in humans [67]. Thus, hydroxyphenylacetylglutamine and hydroxyphenylacetylglutamine sulfate are most likely not metabolites derived from dietary polyphenols, but the result of the modulation of endogenous biological pathways by polyphenols and their metabolites.
Gut microbiota are known to play an important role in the biotransformation of many complex plant polyphenols [68,69,70]. To explore whether the variation in the composition of the gut microbiota could explain, at least partially, the observed inter-individual catabolic differences, we correlated the relative abundances of fecal bacteria taxa with the concentration of metabolites observed in urine and plasma after consumption of apples. Granulicatella was found to be inversely correlated with four metabolites, namely hydroxyphenylacetylglutamine, hydroxyphenylacetylglutamine sulfate, hydroxyphenylacetic acid and hydroxyphenylacetic acid sulfate. The common component for all four metabolites is the hydroxyphenyl acetic moiety, which derives from phenylalanine decarboxylation by the colonic microbiota [64]. Glucuronide conjugated fatty acids C13 with different saturation states were positively correlated with Clostridiales, which is consistent with our previous study, where some saturated and unsaturated fatty acids were correlated with Clostridiales [27].
According to the proposed guidelines for validation of BFIs [1], several fundamental factors should be considered: plausibility, dose–response, time–response, robustness, reliability, stability, analytical performance, and reproducibility.
To approach these factors, in particular plausibility, reproducibility and time–response of BFIs, we compared the untargeted metabolomic data from the current long-term (AVAG) study [25] with a previous single dose postprandial nutrikinetic study [27]. Such strategy was successfully applied in the case of nut consumption [71] and cocoa consumption [72]. An important consideration regarding the participants of the two compared studies is that the long-term dietary intervention (AVAG) involved free living mildly hypercholesterolemic participants, who consumed two apples per day at any time of the day, while the postprandial nutrikinetic study involved healthy participants that consumed polyphenol-enriched cloudy apple juice and cloudy juice in a controlled environment (PAJ and CAJ). Details regarding phloridzin content of apple products used in both intervention studies are given in Fig. 3. We compared biomarkers found in postprandial urine samples: AUC curve from five time points (AGER) with 24 h pool urine (AVAG), while for plasma, postprandial samples from five time points (AGER) with fasting plasma (AVAG). See Fig. 3 for details.
Comparison between putative BFIs found in both studies showed several similarities: some were found at the level of apple-derived metabolites, others at the level of endogenous metabolomics pathways triggered after apple consumption. Twenty-five out of 54 annotated biomarkers in urine from the current study matched with biomarkers of PAJ juice intake from our previous postprandial nutrikinetic study [27]. Figure 4 shows a Venn diagram with metabolites that were common in both studies or unique for only one of them. These metabolites include phloretin glucuronide and phloretin glucuronide sulfate—derived from phlorizin, a characteristic apple constituent [8]; (epi)catechin and valerolactones derivatives—typical metabolites of procyanidins and (epi)catechins common for wide spectra of fruits and vegetables. Two metabolites in plasma were found in both studies: hydroxyhippuric acid and hydroxyphenyl propionic acid sulfate. Similarities were found also between valeric acids metabolites circulating in plasma. Although these metabolites were not identical, they belong to the same chemical class: valeric acid derivatives, arising from microbial activities—see Fig. 4 for details. Furthermore, a wide range of indole-based metabolites were observed in plasma and urine from both dietary interventions, which were derived from indole and tryptophan metabolic pathway, including: tryptophan, acetyltryptophan, indoxyl sulfate, phenol sulfate, dihydroxyindole glucuronide, indolylacryloylglycine, indole acetyl-aspartic acid.
Among all the mentioned metabolites, phloretin metabolites and (epi)catechins fulfill the criterion of plausibility, as phlorizin and (epi)catechins are widely present in apples. However only phloretin metabolites fulfill the criterion of specificity, thus only those were selected for further validation.
Specificity Only minor amounts of phloridzin are present in foods other than apples, although low levels have been found in prune juices (1.70 mg/100 mL [73]), pomegranate juices (0.10 mg/100 mL [74]), or in the peel of immature kumquat fruit (up to 2 g/100 g dry peel of 3′,5′-Di-C-β-glucopyranosylphloretin). However, to our knowledge, there is no report of the presence of the dihydrochalcone phloridzin in any other plant food, in amounts comparable to that reported after apple intake. Accordingly, there is no evidence for the presence of phloretin glucuronide or phloretin glucuronide sulfate in urine or plasma after consumption of the above-mentioned or other foods.
Plausibility Phloretin metabolites, particularly sulfate and glucuronide conjugates are recognized as specific markers of apple consumption [8], as their precursor phlorizin is a typical dihydrochalcone of the Rosacea family, and occurs abundantly in apple cultivars (2.01–15.48 mg/100 g fresh weight) [5]. A large survey investigating 150 apple cultivars (Malus × domestica) highlighted that among the cultivated apples, all Renetta accessions have a high phloridzin content [75]. The selection pressure of domestication and breeding has led to a reduction in the content of phlorizin in the elite cultivars of apple compared with old cultivars. In particular, fruits of several wild Malus genotypes have concentrations of phlorizin well above that of cultivated apples. Therefore, phlorizin is a distinctive apple metabolite, with variability in the concentrations depending on the cultivar.
Time–response Both phloretin glucuronide and phloretin glucuronide sulfate, arising from phase I metabolism, sulfonation and glucuronidation, have relatively short Tmax in biological fluids (1–1.5 h postprandial). However they can still be detected several hours after apple consumption. Phloretin glucuronide was detectable in all urine samples 2–24 h after consumption of both juices. Moreover, phloretin glucuronide sulfate was detected 2–24 h postprandially in all urine samples after PAJ consumption, and 75–91% of the samples after CAJ juice. Both metabolites were found as ubiquitary metabolites in the 24 h urine pool in the long-term study, and thus we can assume that urine pool samples collected over 24 h postprandially is an appropriate sample type to verify apple consumption. However, repeated measurements for both metabolites are needed to provide insights into the reproducibility of biomarker concentrations, and thus the likelihood that the biomarker is a stable estimate of long-term intake. Nevertheless, verification of the postprandial time during which phloretin metabolites return to baseline is also important to elucidate. Kinetic curves from the postprandial kinetic study were over a 5 h period for plasma, while for urine 5–8 h and 8–24 h. In all of these samples, metabolites of phloretin were detectable. Additional time points up to 7–8 h or more postprandially should be included to confirm this.
Dose–response The sum of apple juice-derived phlorizin and phloretin from the postprandial nutrikinetic dietary intervention was 43.2 mg/L and 124.6 mg/L, for the CAJ and PAJ, respectively [27]. This large difference in phlorizin and phloretin content was reflected in the amount of phloretin metabolites found in urine after their consumption. Although analyses of phloretin metabolites were not quantitative, kinetic responses to both interventions were different. The ratio of AUC curves for phloretin glucuronide between enriched apple juice and apple juice was approximately 130 [27]. A similar situation occurred with phloretin glucuronide sulfate; see Fig. 1. Finally, also in the long-term intervention study, differences in phlorizin content in controlled beverage and apples were reflected in detection of phloretin metabolites in biological fluids.
Inter-individual variation Substantial inter-individual variation in the excretion of metabolites was also observed, for example considerable differences in excretions of both phloretin metabolites, especially at 2–5 h postprandially. Considering both metabolites were detected in baseline urine samples in both short-term and long-term interventions, there is a need to evaluate baseline habitual level quantitatively.
Robustness The introduction of two apples per day to habitual diet of free living population in AVAG study was able to change urine profile significantly: phloretin glucuronide was present in all urine samples after apple intervention, while phloretin glucuronide sulfate in 37 samples out of 40. Trace levels (peaks of intensity ca. 2.0E+4 cps, verified manually on chromatogram) of phloretin metabolites were, however, present also in baseline samples (4 samples out of 40). Dietary records were manually verified for these participants and revealed the presence of foods that could be potentially adulterated with apple traces such as fruit juice smoothie, and muesli with fruits, while one person reported the use of apple sauce on one occasion. Apples are largely used for many fruit-based and processed products, thus such unintentional consumption of fruits-based products could result in the presence of phloretin metabolites. Similarly, small peaks of phloretin glucuronide and phloretin glucuronide sulfate were detected in 4 out of 12 postprandial samples at time point 0 h (baselines—before any dietary intervention) in the postprandial nutrikinetic study. Although volunteers were instructed to refrain from consuming polyphenol-rich foods and beverages (wine, coffee, tea, fruits and vegetables) and dietary supplements 3 days prior to the experiments, their dietary records were checked for any food items containing apple residues. None of the consumed foods could explain these residual peaks. In all urine samples collected at 0–2 h, 2–5 h, 5–8 h and 8–24 h after consumption of enriched and unenriched cloudy apple juices, phloretin glucuronide was present; see panel A of Fig. 1. Moreover, phloretin glucuronide sulfate was found in all samples from time points: 0–2 h, 2–5 h, 5–8 h and 8–24 h after enriched apple juice intake, and after cloudy apple juice only in 75–91% of samples from these time points. The lack of presence of phloretin glucuronide sulfate in some samples could be linked to different excretion rates during metabolism. Additional information from observational studies on robustness is needed to clarify impact of eventual confounding factors; however, phloretin glucuronide and phloretin glucuronide sulfate seem potentially robust markers.
Analytical performance and reproducibility In both dietary interventions, the study samples were analyzed by UHPLC Orbitrap mass spectrometer in two different time frames (1 year apart), by two independent operators. The mode of analysis in both cases was untargeted, while data analysis and statistical models were performed independently. Although untargeted analysis is usually recognized as less sensitive than targeted, we were able to detect phloretin metabolites even after 8–24 h in pooled urine samples, though their concentration is expected to be low due to their fast elimination. Similarly, phloretin metabolite was present in pooled 24 h urine (long-term intervention), where dilution factor plays an important role. This confirms the suitability of mass spectrometry in BFI discovery. The capability of phloretin metabolite detection at low concentration is due to the excellent ionization capacities in the ion source and overall sensibility of mass spectrometers. Indeed, mass spectrometry-based analysis of small molecules such as metabolites of polyphenols is particularly effective and robust, and thus frequently employed in nutrimetabolomic investigations [8]. As shown in Trost et al. [27], there is a significant difference in MS response between phloretin glucuronide and phloretin glucuronide sulfate. Although both are visible and detectable by untargeted analysis, phloretin glucuronide was found at higher intensities, while phloretin glucuronide sulfate was found at lower intensities, and sometimes samples lacked signal (see Fig. 1b). This phenomenon can be justified by different excretion quantities, and/or by different LOD for these metabolites. Indeed, formal confirmation of analytical sensitivity and specificity, day-to-day variation and intra-day variation for phloretin glucuronide and glucuronide sulfate is still needed with particular focus on quantitative analysis.
Additional observational studies on robustness, studies with repeated exposure to apples and comparison with other analytical methods are needed to reach full validation according to the criteria established by Dragsted et al. and Gao et al. [1, 24]. Further studies are needed to explore the stability in urine and to evaluate its applicability after intake of cooked or processed products. Moreover, due to the lack of standards, a reliable quantification of phloretin conjugates is a priority.
Conclusions
Differences in the concentration of metabolites in the blood and urine determined by untargeted metabolomics may help in understanding the fate of apple constituents after ingestion. The discovery of validated apple biomarkers could contribute to identifying the relationship between apple consumption and health. Our results suggest that long and sustained consumption of apples triggers the gut microbiota to activate tyrosine, tryptophan and the indole metabolic pathway. The production of endogenous indole metabolites and glutamates and their effect on host and human health require further investigation. Additionally, we approached the validation of phloretin metabolites as BFI, confirming their role as promising biomarkers of apple intake, in light of the evaluation of several criteria such as plausibility, time–response and dose–response. Our results also indicate that 24 h pool urine could be an appropriate biological fluid to identify apple intake.
References
Gao Q, Praticò G, Scalbert A et al (2017) A scheme for a flexible classification of dietary and health biomarkers. Genes Nutr 12:34. https://doi.org/10.1186/s12263-017-0587-x
Garcia-Aloy M, Andres-Lacueva C (2018) Food intake biomarkers for increasing the efficiency of dietary pattern assessment through the use of metabolomics: unforeseen research requirements for addressing current gaps. J Agric Food Chem 66:5–7. https://doi.org/10.1021/acs.jafc.7b05586
Hodgson JM, Prince RL, Woodman RJ et al (2016) Apple intake is inversely associated with all-cause and disease-specific mortality in elderly women. Br J Nutr 115:860–867. https://doi.org/10.1017/S0007114515005231
Koutsos A, Tuohy K, Lovegrove J (2015) Apples and cardiovascular health—is the gut microbiota a core consideration? Nutrients 7:3959–3998. https://doi.org/10.3390/nu7063959
Vrhovsek U, Rigo A, Tonon D, Mattivi F (2004) Quantitation of polyphenols in different apple varieties. J Agric Food Chem 52:6532–6538. https://doi.org/10.1021/jf049317z
D’Archivio M, Filesi C, Varì R et al (2010) Bioavailability of the polyphenols: status and controversies. Int J Mol Sci 11:1321–1342. https://doi.org/10.3390/ijms11041321
Anesi A, Mena P, Bub A et al (2019) Quantification of urinary phenyl-γ-valerolactones and related valeric acids in human urine on consumption of apples. Metabolites 9:254. https://doi.org/10.3390/metabo9110254
Ulaszewska M, Vázquez-Manjarrez N, Garcia-Aloy M et al (2018) Food intake biomarkers for apple, pear, and stone fruit. Genes Nutr 13:29. https://doi.org/10.1186/s12263-018-0620-8
Kahle K, Huemmer W, Kempf M et al (2007) Polyphenols are intensively metabolized in the human gastrointestinal tract after apple juice consumption. J Agric Food Chem 55:10605–10614. https://doi.org/10.1021/jf071942r
Makarova E, Gornas P, Konrade I et al (2014) Acute anti-hyperglycaemic effects of an unripe apple preparation containing phlorizin in healthy volunteers: a preliminary study. J Sci Food Agric 95:560–568. https://doi.org/10.1002/jsfa.6779
Hagl S, Deusser H, Soyalan B et al (2011) Colonic availability of polyphenols and d-(−)-quinic acid after apple smoothie consumption. Mol Nutr Food Res 55:368–377. https://doi.org/10.1002/mnfr.201000252
Marks SC, Mullen W, Borges G, Crozier A (2009) Absorption, metabolism, and excretion of cider dihydrochalcones in healthy humans and subjects with an ileostomy. J Agric Food Chem 57:2009–2015. https://doi.org/10.1021/jf802757x
Hollands WJ, Hart DJ, Dainty JR et al (2013) Bioavailability of epicatechin and effects on nitric oxide metabolites of an apple flavanol-rich extract supplemented beverage compared to a whole apple puree: a randomized, placebo-controlled, crossover trial. Mol Nutr Food Res 57:1209–1217. https://doi.org/10.1002/mnfr.201200663
Gasper A, Hollands W, Casgrain A et al (2014) Consumption of both low and high (−)-epicatechin apple puree attenuates platelet reactivity and increases plasma concentrations of nitric oxide metabolites: a randomized controlled trial. Arch Biochem Biophys 559:29–37. https://doi.org/10.1016/j.abb.2014.05.026
Auclair S, Chironi G, Milenkovic D et al (2010) The regular consumption of a polyphenol-rich apple does not influence endothelial function: a randomised double-blind trial in hypercholesterolemic adults. Eur J Clin Nutr 64:1158–1165. https://doi.org/10.1038/ejcn.2010.135
Mennen LI, Sapinho D, Ito H et al (2006) Urinary flavonoids and phenolic acids as biomarkers of intake for polyphenol-rich foods. Br J Nutr 96:191. https://doi.org/10.1079/BJN20061808
Rago D, Gurdeniz G, Gitte R-H, Dragsted L (2014) An explorative study of the effect of apple and apple products on the human plasma metabolome investigated by LC–MS profiling. Metabolomics 11:27–39. https://doi.org/10.1007/s11306-014-0668-4
Ravn-Haren G, Dragsted LO, Buch-Andersen T et al (2013) Intake of whole apples or clear apple juice has contrasting effects on plasma lipids in healthy volunteers. Eur J Nutr 52:1875–1889. https://doi.org/10.1007/s00394-012-0489-z
Kristensen M, Engelsen SB, Dragsted LO (2012) LC-MS metabolomics top-down approach reveals new exposure and effect biomarkers of apple and apple-pectin intake. Metabolomics 8:64–73. https://doi.org/10.1007/s11306-011-0282-7
Rago D, Mette K, Gürdeniz G et al (2013) A LC–MS metabolomics approach to investigate the effect of raw apple intake in the rat plasma metabolome. Metabolomics 9:1202–1215. https://doi.org/10.1007/s11306-013-0534-9
Saenger T, Hubner F, Humpf HU (2017) Short-term biomarkers of apple consumption. Mol Nutr Food Res 61:1–10. https://doi.org/10.1002/mnfr.201600629
García-Conesa M-T, Chambers K, Combet E et al (2018) Meta-analysis of the effects of foods and derived products containing ellagitannins and anthocyanins on cardiometabolic biomarkers: analysis of factors influencing variability of the individual responses. Int J Mol Sci 19:694. https://doi.org/10.3390/ijms19030694
Menezes R, Rodriguez-Mateos A, Kaltsatou A et al (2017) Impact of flavonols on cardiometabolic biomarkers: a meta-analysis of randomized controlled human trials to explore the role of inter-individual variability. Nutrients 9:117. https://doi.org/10.3390/nu9020117
Dragsted LO, Gao Q, Scalbert A et al (2018) Validation of biomarkers of food intake—critical assessment of candidate biomarkers. Genes Nutr 13:14. https://doi.org/10.1186/s12263-018-0603-9
Koutsos A, Riccadonna S, Ulaszewska MM et al (2019) Two apples a day lower serum cholesterol and improve cardiometabolic biomarkers in mildly hypercholesterolemic adults: a randomized, controlled, crossover trial. Am J Clin Nutr. https://doi.org/10.1093/ajcn/nqz282
Koutsos A, Lima M, Conterno L et al (2017) Effects of commercial apple varieties on human gut microbiota composition and metabolic output using an in vitro colonic model. Nutrients 9:533. https://doi.org/10.3390/nu9060533
Trošt K, Ulaszewska MM, Stanstrup J et al (2018) Host: microbiome co-metabolic processing of dietary polyphenols—an acute, single blinded, cross-over study with different doses of apple polyphenols in healthy subjects. Food Res Int. https://doi.org/10.1016/j.foodres.2018.06.016
Ulaszewska MM, Trost K, Stanstrup J et al (2016) Urinary metabolomic profiling to identify biomarkers of a flavonoid-rich and flavonoid-poor fruits and vegetables diet in adults: the FLAVURS trial. Metabolomics 12:1–22. https://doi.org/10.1007/s11306-015-0935-z
Ancillotti C, Ulaszewska M, Mattivi F, Del Bubba M (2019) Untargeted metabolomics analytical strategy based on liquid chromatography/electrospray ionization linear ion trap quadrupole/orbitrap mass spectrometry for discovering new polyphenol metabolites in human biofluids after acute ingestion of Vaccinium myrti. J Am Soc Mass Spectrom 30:381–402. https://doi.org/10.1007/s13361-018-2111-y
Chambers MC, Maclean B, Burke R et al (2012) A cross-platform toolkit for mass spectrometry and proteomics. Nat Biotechnol 30:918–920. https://doi.org/10.1038/nbt.2377
Smith CA, Want EJ, O’Maille G et al (2006) XCMS: processing mass spectrometry data for metabolite profiling using nonlinear peak alignment, matching, and identification. Anal Chem 78:779–787. https://doi.org/10.1021/ac051437y
Kuhl C, Tautenhahn R, Böttcher C et al (2012) CAMERA: an integrated strategy for compound spectra extraction and annotation of liquid chromatography/mass spectrometry data sets. Anal Chem 84:283–289. https://doi.org/10.1021/ac202450g
Strimmer K (2008) A unified approach to false discovery rate estimation. BMC Bioinform 9:303. https://doi.org/10.1186/1471-2105-9-303
Strimmer K (2008) fdrtool: a versatile R package for estimating local and tail area-based false discovery rates. Bioinformatics 24:1461–1462. https://doi.org/10.1093/bioinformatics/btn209
Wishart DS, Jewison T, Guo AC et al (2013) HMDB 3.0—the human metabolome database in 2013. Nucleic Acids Res 41:801–807. https://doi.org/10.1093/nar/gks1065
Smith A, O’maille G, Want EJ et al (2005) METLIN a metabolite mass spectral database. Ther Drug Monit 27(6):747–751. https://doi.org/10.1097/01.ftd.0000179845.53213.39
Horai H, Arita M, Kanaya S et al (2010) MassBank: a public repository for sharing mass spectral data for life sciences. J Mass Spectrom. https://doi.org/10.1002/jms.1777
Sud M, Fahy E, Cotter D et al (2007) LMSD: LIPID MAPS structure database. Nucleic Acids Res 35:527–532. https://doi.org/10.1093/nar/gkl838
Gerlich M, Neumann S (2013) MetFusion : integration of compound identification strategies. J Mass Spectrom. https://doi.org/10.1002/jms.3123
Haug K, Salek RM, Conesa P et al (2013) MetaboLights—an open-access general-purpose repository for metabolomics studies and associated meta-data. Nucleic Acids Res 41:D781–D786. https://doi.org/10.1093/nar/gks1004
Caporaso JG, Kuczynski J, Stombaugh J et al (2010) QIIME allows analysis of high-throughput community sequencing data. Nat Methods 7:335–336. https://doi.org/10.1038/nmeth.f.303
Aronesty E (2013) Comparison of sequencing utility programs. Open Bioinforma J 7:1–8. https://doi.org/10.2174/1875036201307010001
Edgar RC (2010) Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26:2460–2461. https://doi.org/10.1093/bioinformatics/btq461
Urpi-Sarda M, Monagas M, Khan N et al (2009) Epicatechin, procyanidins, and phenolic microbial metabolites after cocoa intake in humans and rats. Anal Bioanal Chem 394:1545–1556. https://doi.org/10.1007/s00216-009-2676-1
Stalmach A, Edwards CA, Wightman JD, Crozier A (2011) Identification of (Poly)phenolic compounds in concord grape juice and their metabolites in human plasma and urine after juice consumption. J Agric Food Chem 59:9512–9522. https://doi.org/10.1021/jf2015039
Van Duynhoven J, Vaughan EE, Van Dorsten F et al (2013) Interactions of black tea polyphenols with human gut microbiota: implications for gut and cardiovascular health. Am J Clin Nutr 98:1631–1641. https://doi.org/10.3945/ajcn.113.058263.INTRODUCTION
Mateos-Martín ML, Pérez-Jiménez J, Fuguet E, Torres JL (2012) Non-extractable proanthocyanidins from grapes are a source of bioavailable (epi)catechin and derived metabolites in rats. Br J Nutr 108:290–297. https://doi.org/10.1017/S0007114511005678
Touriño S, Pérez-Jiménez J, Mateos-Martín ML et al (2011) Metabolites in contact with the rat digestive tract after ingestion of a phenolic-rich dietary fiber matrix. J Agric Food Chem 59:5955–5963. https://doi.org/10.1021/jf200159f
Stalmach A, Steiling H, Williamson G, Crozier A (2010) Bioavailability of chlorogenic acids following acute ingestion of coffee by humans with an ileostomy. Arch Biochem Biophys 501:98–105. https://doi.org/10.1016/j.abb.2010.03.005
Stalmach A, Mullen W, Barron D et al (2009) Metabolite profiling of hydroxycinnamate derivatives in plasma and urine after the ingestion of coffee by humans: identification of biomarkers of coffee consumption. Drug Metab Dispos 37:1749–1758. https://doi.org/10.1124/dmd.109.028019
Stalmach A, Edwards CA, Wightman JD, Crozier A (2013) Colonic catabolism of dietary phenolic and polyphenolic compounds from Concord grape juice. Food Funct 4:52–62. https://doi.org/10.1039/C2FO30151B
Urpi-Sarda M, Garrido I, Monagas M et al (2009) Profile of plasma and urine metabolites after the intake of almond [Prunus dulcis (Mill.) D.A. Webb] polyphenols in humans. J Agric Food Chem 57:10134–10142. https://doi.org/10.1021/jf901450z
Redeuil K, Smarrito-Menozzi C, Guy P et al (2011) Identification of novel circulating coffee metabolites in human plasma by liquid chromatography-mass spectrometry. J Chromatogr A 1218:4678–4688. https://doi.org/10.1016/j.chroma.2011.05.050
Nørskov NP, Hedemann MS, Lærke HN, Knudsen KEB (2013) Multicompartmental nontargeted LC–MS metabolomics: explorative study on the metabolic responses of rye fiber versus refined wheat fiber intake in plasma and urine of hypercholesterolemic pigs. J Proteome Res 12:2818–2832. https://doi.org/10.1021/pr400164b
Bergmann H, Triebel S, Kahle K, Richling E (2010) The metabolic fate of apple polyphenols in humans. Curr Nutr Food Sci. https://doi.org/10.2174/157340110790909554
Stanstrup J, Rasmussen JE (2014) Intakes of whey protein hydrolysate and whole whey proteins are discriminated by LC–MS metabolomics. Metabolomics. https://doi.org/10.1007/s11306-013-0607-9
Stanstrup J, Schou SS, Holmer-Jensen J et al (2014) Whey protein delays gastric emptying and suppresses plasma fatty acids and their metabolites compared to casein, gluten, and fish protein. J Proteome Res 13:2396–2408. https://doi.org/10.1021/pr401214w
Agus A, Planchais J, Sokol H (2018) Gut microbiota regulation of tryptophan metabolism in health and disease. Cell Host Microbe 23:716–724. https://doi.org/10.1016/j.chom.2018.05.003
Gao J, Xu K, Liu H et al (2018) Impact of the gut microbiota on intestinal immunity mediated by tryptophan metabolism. Front Cell Infect Microbiol. https://doi.org/10.3389/fcimb.2018.00013
Keszthelyi D, Troost FJ, Masclee AAM (2009) Understanding the role of tryptophan and serotonin metabolism in gastrointestinal function. Neurogastroenterol Motil 21:1239–1249. https://doi.org/10.1111/j.1365-2982.2009.01370.x
Wlodarska M, Luo C, Kolde R et al (2017) Indoleacrylic acid produced by commensal peptostreptococcus species suppresses inflammation. Cell Host Microbe 22:25–37.e6. https://doi.org/10.1016/j.chom.2017.06.007
Bartel B, LeClere S, Magidin M, Zolman BK (2001) Inputs to the active indole-3-acetic acid pool: de novo synthesis, conjugate hydrolysis, and indole-3-butyric acid b-oxidation. J Plant Growth Regul 20:198–216. https://doi.org/10.1007/s003440010025
Chou JC, Kuleck GA, Cohen JD, Mulbry WW (1996) Partial purification and characterization of an inducible indole-3-acetyl-l-aspartic acid hydrolase from Enterobacter agglomerans. Plant Physiol 112:1281–1287. https://doi.org/10.1104/pp.112.3.1281
Semba RD, Trehan I, Li X et al (2017) Environmental enteric dysfunction is associated with carnitine deficiency and altered fatty acid oxidation. EBioMedicine 17:57–66. https://doi.org/10.1016/j.ebiom.2017.01.026
Smith EA, Macfarlane GT (1997) Formation of phenolic and indolic compounds by anaerobic bacteria in the human large intestine. Microb Ecol 33:180–188. https://doi.org/10.1007/s002489900020
Russell WR, Duncan SH, Scobbie L et al (2013) Major phenylpropanoid-derived metabolites in the human gut can arise from microbial fermentation of protein. Mol Nutr Food Res 57:523–535. https://doi.org/10.1002/mnfr.201200594
van Dorsten FA, Grün CH, van Velzen EJJ et al (2009) The metabolic fate of red wine and grape juice polyphenols in humans assessed by metabolomics. Mol Nutr Food Res 54:897–908. https://doi.org/10.1002/mnfr.200900212
Monagas M, Urpi-Sarda M, Sánchez-Patán F et al (2010) Insights into the metabolism and microbial biotransformation of dietary flavan-3-ols and the bioactivity of their metabolites. Food Funct 1:233. https://doi.org/10.1039/c0fo00132e
Scalbert A, Williamson G (2085S) Dietary intake and bioavailability of polyphenols. J Nutr 130:2073S–2085S. https://doi.org/10.1093/jn/130.8.2073S
Del Rio D, Rodriguez-Mateos A, Spencer JPE et al (2013) Dietary (poly)phenolics in human health: structures, bioavailability, and evidence of protective effects against chronic diseases. Antioxid Redox Signal 18:1818–1892. https://doi.org/10.1089/ars.2012.4581
Garcia-Aloy M, Llorach R, Urpi-Sarda M et al (2014) Novel multimetabolite prediction of walnut consumption by a urinary biomarker model in a free-living population: the PREDIMED study. J Proteome Res 13:3476–3483. https://doi.org/10.1021/pr500425r
Garcia-Aloy M, Llorach R, Urpi-Sarda M et al (2015) A metabolomics-driven approach to predict cocoa product consumption by designing a multimetabolite biomarker model in free-living subjects from the PREDIMED study. Mol Nutr Food Res 59:212–220. https://doi.org/10.1002/mnfr.201400434
Van Gorsel H, Li C, Kerbel EL et al (1992) Compositional characterization of prune juice. J Agric Food Chem 40:784–789. https://doi.org/10.1021/jf00017a016
Poyrazoğlu E, Gökmen V, Artιk N (2002) Organic acids and phenolic compounds in pomegranates (Punica granatum L.) grown in Turkey. J Food Compos Anal 15:567–575. https://doi.org/10.1006/jfca.2002.1071
Farneti B, Masuero D, Costa F et al (2015) Is there room for improving the nutraceutical composition of apple? J Agric Food Chem 63:2750–2759. https://doi.org/10.1021/acs.jafc.5b00291
Acknowledgements
We thank Massimo Pindo and the FEM Sequencing Platform for performing the DNA sequencing
Funding
This project was funded in part by the AGER project “Apple fruit quality in the post-genomic era, from breeding new genotypes to post-harvest: nutrition and health”, funded by the AGER (Agribusiness and research) with Grant no. 2010–2119, internal funding of Fondazione Edmund Mach, and the European Union’s Horizon2020 research and innovation grant agreement No 696295—ERA-Net Cofund ERA-HDHL “Biomarkers for Nutrition and Health implementing the JPI HDHL objectives” (https://www.healthydietforhealthylife.eu/), projects FOODBALL (https://foodmetabolome.org/) and CABALA_DIET&HEALTH (https://www.cabalaproject.eu/). The Renetta apples were kindly provided by Consorzio Melinda S.C.A., Cles, Trentino, Italy.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interests
On behalf of all authors, the corresponding author states that there is no conflict of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Ulaszewska, M.M., Koutsos, A., Trošt, K. et al. Two apples a day modulate human:microbiome co-metabolic processing of polyphenols, tyrosine and tryptophan. Eur J Nutr 59, 3691–3714 (2020). https://doi.org/10.1007/s00394-020-02201-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00394-020-02201-8