Abstract
Key message
The glyphosate resistance in Escherichia coli and Arabidopsis was due to d -amino acid oxidase expression.
Abstract
Transgenic glyphosate-resistant crops have a high percentage in the total area devoted to transgenic crops worldwide. d-amino acid oxidase (DAAO) can metabolize glyphosate by oxidative cleavage of the carbon–nitrogen bond on the carboxyl side and yield aminomethyl phosphonic acid and glyoxylate, which are less toxic to plants than glyphosate. To date, reports on the use of DAAO to enhance glyphosate resistance in plants are lacking. In this paper, we report synthesis, and codon usage optimization for plant expression, of the DAAO gene by successive polymerase chain reaction from Bradyrhizobium japonicum. To confirm the glyphosate resistance of the DAAO gene, the recombinant plasmid pYPX251 (GenBank Accession No: AY178046) harboring the wild-type DAAO gene was transformed into DH5α. The positive transformants grew well both on solid and in liquid M9 medium containing 200 mM glyphosate. The optimized DAAO gene was transformed into Arabidopsis and 9 days after application of 10 mM glyphosate, the 4-week-old wild-type plants all died; by contrast, transgenic plants could grow normally. The proline content and peroxidase activity showed that glyphosate could induce proline accumulation and produce reactive oxygen species.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Glyphosate, the most widely used herbicide in the world, inhibits 5-enolpyruvylshikimate-3-phosphate synthase (EPSPS, EC 2.5.1.19). It shows broad-spectrum herbicide activity against various weeds (Duke and Powles 2008; Dill et al. 2008). In transgenic glyphosate-resistant crops, glyphosate can be applied to the crop (post-emergence) to remove emerged weeds without crop damage. Transgenic glyphosate-resistant crops have a high percentage in the total area devoted to transgenic crops worldwide (Duke 2014). Most of the grown transgenic glyphosate-resistant crops presently obtain resistance by overexpressing EPSPS isolated from Agrobacterium sp. CP4 or a variant of EPSPS (T102I/P106S double mutant called TIPS) that is not inhibited by glyphosate, as well as a mutated maize EPSPS. The theoretical disadvantage of this approach is that glyphosate remains in plant and crop yields (Duke et al. 2003).
Glyphosate resistance can also be introduced into crops through detoxification of the glyphosate molecule. Resistance is obtained through glyphosate detoxification by transgene-encoded enzymes, including glyphosate acetyltransferase (GAT) from Bacillus licheniformis (Castle et al. 2004; Siehl et al. 2007), glyphosate oxidoreductase (GOX) from Ochrobactrum anthropi, and glycine oxidase (GO) from Bacillus subtilis (Nicolia et al. 2014). GAT can convert glyphosate into N-acetylglyphosate, which does not inhibit EPSPS. GOX is able to convert glyphosate into aminomethyl phosphonic acid (AMPA) and glyoxylate by cutting the carbon–nitrogen bond on the carboxyl side. GO can also convert glyphosate into AMPA and glyoxylate, but through a different reaction mechanism.
d-amino acid oxidase (DAAO) is a well-known flavoenzyme that catalyzes the oxygen-dependent oxidative deamination of amino acid D-isomers with absolute stereospecificity, which results in α-keto acids, ammonia, and hydrogen peroxide (Pollegioni and Molla 2011). DAAO can metabolize glyphosate by oxidative cleavage of the carbon–nitrogen bond on the carboxyl side, and yield AMPA and glyoxylate. DAAO shows modest sequence similarity to GO; they belong to the same structural family but share different substrate specificities with each other (Pedotti et al. 2009).
As of this writing, no reports have focused on glyphosate-resistant transgenic plants by introducing DAAO genes. In this study, the wild-type DAAO gene (GenBank Accession No: BA000040.2) from Bradyrhizobium japonicum was transformed into glyphosate-susceptible Escherichia coli to detect DAAO-imparted resistance. We also synthesized the DAAO gene according to plant codon preference and transgenic Arabidopsis plants expressing the optimized gene, which were significantly more resistant to glyphosate than untransformed plants. We performed other biological and chemical measurements to assess the function of the DAAO gene in transgenic Arabidopsis.
Materials and methods
Materials, vectors and chemicals
Escherichia coli strain DH5α, used as a host for heterologous expression of wild-type DAAO, and pYPX251 (p251) vector (GenBank Accession No: AY178046), pCAMBIA1301 (GenBank Accession No: AE234297) vector, and Agrobacterium tumefaciens GV3101, Arabidopsis thaliana (ecotype Columbia L.) were all prepared and maintained in our laboratory. Simple pMD-18 vector and Taq DNA polymerase, T4 DNA ligase and other enzymes were purchased from Takara. RNA isolation kit and Reverse Transcription System kit were purchased from Axygen Co. (China).
Chemical synthesis of DAAO gene from Bradyrhizobium japonicum
According to the sequence (GenBank Accession No: BA000040.2), the DAAO gene from Bradyrhizobium japonicum, was optimized using codon usage bias and synthesized by successive polymerase chain reaction (PCR) method (Xiong et al. 2004; Murray et al. 1989). When optimized the gene sequence, four aspects needed to be considered. (1) all codons were suitable in Arabidopsis, singling out rare codons; (2) replacing the bases in the GC or AT dense distribution, without amino acid change; (3) increasing the A/T content at or near the ribosome binding site, decreasing the possibility of forming stem-loop structure at mRNA 5′; (4) calculating free energy of every domain, to enable the free energy decreased from 5′ to 3′ as possible. PCR was carried out as described in Tian et al. (2011). The primers were in Table S1. The amplified fragment was cloned into Simple pMD-18 and sequenced (Xu et al. 2010).
In vitro glyphosate sensitivity assays
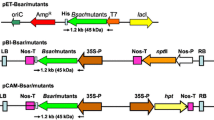
To determine whether the DAAO gene from B. japonicum confers glyphosate resistance to E. coli, recombinant pYPX251 (p251) vector (GenBank Accession No: AY178046) with the wild-type DAAO gene was transformed into DH5α. The recombinant p251-DAAO was generated by inserting the DAAO gene into the BamHI-to-SacI site. Empty p251 without the DAAO gene was transformed into DH5α as a control. The vector p251 carried the moderately strong promoter for aac1 gene encoding gentamicin 3-acetyltransferase and the T1T2 transcription terminator (Tian et al. 2011; Xiong et al. 2007).
The p251-DAAO and p251 (empty) plasmids were transformed into competent E. coli, and applied to solid M9 minimal medium containing 0, 100, 150, and 200 mM glyphosate and 100 mg L−1 Ampicillin. The positive monoclone obtained on medium containing 200 mM glyphosate were grown with shaking at 37 °C in liquid M9 minimal medium for 16 h. To ensure that we had the same amount of bacterial cells in the corresponding cultures prior to glyphosate exposure, we adjusted the amount of overnight suspension culture of the two types of transformants, which were transferred into fresh M9 medium. OD660 values of fresh M9 medium (20 mL) with the two transformants containing different concentrations of glyphosate (0 and 200 mM) were 0.012. Cultures were incubated as above, and OD660 values were determined every 2 h (Sun et al. 2005).
Plant expression vector construction and plant transformation
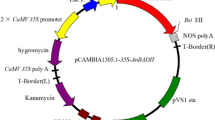
Construction of the plant expression vector was performed as described by Xu et al. (2010) with some modification. We added a chloroplast transit sequence to the constructs to ensure the DAAO gene expression in leaves and stems. The DNA encoding the chloroplast transit peptide (TSP) of Arabidopsis was amplified according to Klee et al. (1987). The primers were: 5′-GTCGACATGGCGCAAGTTAGCAGAATC-3′ and 5′-GGATCCCTCCGCCGT GGAAACACAAGAC-3′. The amplified fragment was digested with BamHI and SalI, inserted into pGEM3Z, and sequenced. Then the double CaMV35S promoter was inserted into pGEM3Z at PstI-SalI site upstream of the TSP. Finally, the aforesaid fused fragment and optimized DAAO gene were all cloned into the plant binary expression vector pCAMBIA1301 upstream of the nopaline synthase terminator (NOS-Ter) at PstI-BamHI and BamHI-SacI restriction sites, respectively. The recombinant plasmid, D35S:TSP:DAAO:Nos, was transformed into Agrobacterium tumefaciens GV3101 by electroporation (Xue et al. 2009), and subsequently transformed into Arabidopsis (ecotype Columbia) by a floral dip method as Zhang et al. described previously (2006) to obtain transgenic plants. Transgenic plants were selected by plating the seeds on solid MS medium containing 30 mg/L hygromycin. Plants were carried for two more generations in order to obtain homozygous transgenic plants.
RT-PCR detection of transgenic Arabidopsis
The homozygous transgenic plants were further confirmed by RT-PCR analysis. The cDNA obtaining of the DAAO gene and RT-PCR were performed as Zhu et al. described previously (2012). The Arabidopsis actin gene, AtAc 2 (GenBank Accession No: NM12764), was used as the internal standard with the primers: 5′-GCACCCTGTTCTTCTTACCGAG-3′ and 5′-AGTAAGGTCACGTCCAGCAAGG-3′. The DAAO gene fragment (300 bp) was amplified by specific primers: 5′-CGCCATAGCGGAAGGCTTGTC-3′ and 5′-CGGATTGCCGAGGATCATGTC-3′.
Glyphosate resistance analysis of plants
For the growth of seedling experiments on MS plates, seeds from wild type (WT) and transgenic T3 plants carrying the DAAO gene (Do3, Do6, Do14) were surface-sterilized and sowed on solid MS medium containing 0, 200, and 500 µM glyphosate, and grown for 12 days.
For glyphosate spraying analysis, the surfaced-sterilized seeds were sowed on solid MS medium and grown for 10 days, and then the seedlings were transferred in pots containing a mixture of vermiculite, peat moss, and perlite (18:6:1, v/v). All the seeds and seedlings were grown in a controlled environmental chamber at 22 °C on a 16 h light/8 h dark cycle. For the leaf spraying experiment, 4-week-old transgenic plants were sprayed with 10 mM herbicide glyphosate (isopropylamine salt of glyphosate as active ingredient), doses of 2 mL per 100 cm−2 every 3 days.
Measurement of physiological indexes
To investigate the stress of glyphosate on WT and transgenic plants, two physiological indexes including peroxidase (POD, EC 1.11.1.7) activity, and proline content were measured. 4-week-old seedlings was sprayed with 10 mM glyphosate 72 h later, fresh tissue of leaves (0.05 g) was harvested and washed with double distilled water. The leaf tissue was crushed using a chilled pestle and mortar kept in an ice bath.
The crushed leaf tissue was homogenized in 200 mM, pH 6.0 phosphate buffer. The homogenate was centrifuged at 12,000 g for 20 min at 4 °C. The supernatant was stored at 4 °C and used for assays within 4 h.
Peroxidase activity was measured using a modified method from MacAdam et al. (1992). 1 U of POD means an absorbance change of 0.01 U/min−1.
Proline was extracted with 3 % sulfosalicylic acid aqueous solution and its content was determined using the ninhydrin assay of Bates et al. (1973).
The total chlorophylls content was measured 1 day after a second spraying with 10 mM glyphosate as Lichtenthaler described previously (1987).
Results
Chemical synthesis of DAAO gene from Bradyrhizobium japonicum
To make sure the gene expression in transgenic plants was stable, the DAAO gene was chemically synthesized using 32 primers (Table S1) according to the original sequence and plant codon usage bias. The optimized sequence derived for the wild-type DAAO gene from B. japonicum is shown in Fig. S1, in which 269 nucleotides of the native sequence were changed. The optimal sequence has 56.8 % GC content and shared 78.75 % similarity to the wild-type gene (Fig. S1). Amino acid sequence was translated using DNAMAN. No differences between the two amino acid sequences were observed (Fig. S1).
In vitro glyphosate sensitivity assays
Both empty p251 and p251-DAAO plasmids were transformed into DH5α E. coli. The transformations were spread on solid M9 medium plates with 0, 100, 150, and 200 mM glyphosate, and grown for 16 h. The results are shown in Fig. 1. The amount of DH5α cells transformed with empty p251 plasmid was much higher than that of cells transformed with p251-DAAO on control medium. This was an issue with the size of the two plasmids. Size of p251 empty plasmid was smaller than that of p251-DAAO, so the transformation efficiency was higher. DH5α cells transformed with empty p251 plasmid were fully inhibited on M9 plates containing 100 mM glyphosate. However, DH5α cells transformed with p251-DAAO plasmid were much less inhibited and grew well on 200 mM glyphosate plates.
Enhanced glyphosate resistances of E. coli on plates. Both empty p251 and p251-DAAO plasmids were transformed into DH5α. Transformants were spread on solid M9 medium plates with 0, 100, 150, and 200 mM glyphosate, and grown for 16 h. The same results were obtained in three independent experiments and are represented by the effects shown here
The growth curves of E. coli DH5α harboring either p251 or p251-DAAO, in liquid M9 medium containing 0 and 200 mM of glyphosate, are shown in Fig. 2. There is no significant difference at 0 mM concentration once the cell concentrations were standardized. After 24 h of incubation, the growth of cells harboring empty p251 plasmid was strongly inhibited (>95 %) by 200 mM glyphosate, whereas the OD of cultures harboring p251-DAAO plasmid was approximately 37 % of the control values without glyphosate.
Plant transformation and RT-polymerase chain reaction (PCR) detection of transgenic Arabidopsis
Transgenic Arabidopsis was used to evaluate the potential application of the DAAO gene from B. japonicum in developing glyphosate-resistant crops. Expression cassettes comprising a double CaMV35S promoter, Arabidopsis chloroplast transit sequence, and DAAO coding region were cloned into the T-DNA region of a binary vector (Fig. 3a). Transgenic plants were obtained from plantlets grown in medium containing 30 mg/mL hygromycin (Fig. 3b). Fifteen independent lines of transgenic plants were generated. Among them, three overexpression lines (named Do3, Do6, and Do14) were chosen for further experiments.
a The DAAO expression vector for Arabidopsis transformation. Nos-Ter nopaline synthase terminator sequence, SAR scaffold attachment region, TSP chloroplast Arabidopsis transit peptide. b Seeds from wild type (WT) and transgenic T3 plants carrying the DAAO gene (Do3, Do6, Do14), on solid MS medium containing 30 mg/L hygromycin. c RT-PCR analysis of the expression of DAAO (target gene) and actin genes in transgenic lines (Do3, Do6, Do14) and wild type (WT). Each lane contained 5 μL of RT-PCR products obtained using total RNA extracted from 4-week-old plants grown under standard conditions. M markers (DL15000, Takara). Data shown are representative of three independent experiments. Scale bars 1 cm
RT-PCR was performed to examine the transcript levels of the DAAO gene. In the T2 generation, the transcript of the target gene were successfully detected in the three overexpression lines. Furthermore, the transcript was not detected in the WT plants (Fig. 3c). No obvious effects on growth and development were observed in DAAO transgenic plants under normal growth conditions (Fig. 4).
Glyphosate resistance analysis of plants
To analyze glyphosate resistance, Arabidopsis seeds were sown on solid MS medium with 0, 200, and 500 µM glyphosate and then grown for 12 days. Both WT and transgenic plants grew identically on the control medium without glyphosate. On the medium containing 200 µM glyphosate for 12 days, WT plants developed only chlorotic cotyledonary leaves and root growth was inhibited and transgenic plants displayed minimal signs of phytotoxic effects, showing normal development. The 500 µM glyphosate concentration further augment chlorosis and more severely impaired the development of all WT plants. Moreover, transgenic plants gradually developed to later stages and produced longer roots and stronger green leaves than WT plants. These results indicated that transgenic lines (Do3, Do6, and Do14) showed significantly higher resistance than WT plants (Fig. 4).
Four-week-old seedlings were sprayed with 10 mM glyphosate once every 3 days. To evaluate the response of the WT plants and transgenic plants to glyphosate, the proline content and POD activity were detected 3 days after one spray. Proline and POD are involved in the response to a number of environmental stresses. The proline content and POD activity were not significantly different in the WT and transgenic overexpression lines before stress was induced. After glyphosate spraying, the proline content and POD activity of transgenic lines both increased. The increments of proline content for transgenic lines were 2.0, 3.07 and 7.1 µM/g FW, respectively, which were much lower than that of WT plants (25.3 µM/g FW) (Fig. 5a). So as the POD activity. The increments of POD activity for transgenic lines were 1739, 3226, 4164 U/g FW, respectively, which were also much lower than that of WT plants (6511 U/g FW) (Fig. 5b).
Changes in the content of proline, POD enzyme activities and chlorophylls content in transgenic plants carrying the DAAO gene (Do3, Do6, Do14) and wild type (WT) treated with 10 mM glyphosate. Proline content and POD activity were measured 3 days after first treatment. The total chlorophylls content was measured 1 day after a second spraying with 10 mM glyphosate. Values are means of three replications ±SD
A previous study revealed that sublethal concentrations of glyphosate cause chlorotic symptoms in the leaves of transgenic tobacco expressing OsEPSPS gene (Zhou et al. 2006). Therefore, the total chlorophyll content was also measured at 1 d after a second spray (leaves of control plants showed slight yellowing). The chlorophyll contents of transgenic lines were 0.71, 0.66, 0.67 mg/g FW, respectively, which were higher than that of WT plants (0.45 mg/g FW) (Fig. 5c).
At 3 days after the third spray, all leaves on WT plants showed severe yellowing and wilting, whereas transgenic plants grew well with normal morphology (Fig. 6). These results indicated that DAAO transgenic plants were more resistant to glyphosate exposure than WT plants.
Wild type (WT) and transgenic plants carrying the DAAO gene (Do3, Do6, Do14) were sprayed with 10 mM glyphosate. Control Photograph taken before the first spray treatment. Glyphosate treatment Photograph taken 3 days after the third spray treatment. The same results were obtained in three independent experiments and are represented by the effects shown here
Discussion
Two basic strategies have been successful in introducing glyphosate resistance into plants, namely, expression of an insensitive form of the target enzyme and detoxification of the glyphosate molecule. The former strategy is used in existing commercial glyphosate-resistant crops, employing a microbial (Agrobacterium sp. CP4) or a mutated (TIPS) form of EPSPS that is not inhibited by glyphosate. In contrast to the existing commercial glyphosate-resistant crops, resistance to the herbicide in transgenic plants using the second strategy is achieved through degradation to non-toxic or less toxic compounds. DAAO can metabolize glyphosate via oxidative cleavage of the carbon–nitrogen bonds on the carboxyl side, resulting in the formation of AMPA and glyoxylate. These mechanisms have not been shown to occur in higher plants to a significant degree (Duke 2011).
Previous studies have reported increasing instances of evolved glyphosate resistance in weed species following wide planting of glyphosate-resistant crops, based mainly on EPSPS being insensitive to the herbicide (Dill et al. 2008; Waltz 2010). In several cases, moderate resistance is imparted by mutations of the target enzyme (Powles and Preston 2006), but no documented case of a plant species with native or evolved resistance to glyphosate by virtue of a metabolic enzyme has been found (Pollegioni et al. 2011; Trovato et al. 2008). Thus, new genes should be discovered and utilized for the development of glyphosate-resistant crops.
The main objective of our research was to identify a DAAO gene with properties appropriate for the development of transgenic glyphosate-resistant plants. The wild-type DAAO gene was successfully transformed into DH 5α to detect its resistance to glyphosate. Compared with the control, we found that DH 5α with p251-DAAO plasmid could tolerate glyphosate at concentrations of 200 mM on M9 plates or liquid M9 medium. By contrast, the growth of cultures harboring p251-aroA H. Orenii (generated by inserting the EPSPS gene aroA H. Orenii into the BamHI-to-SacI site in pYPX251) was completely inhibited at 120 mM glyphosate (Tian et al. 2011).
Our PCR results confirmed that the DAAO gene was successfully inserted into the Arabidopsis genome and expressed (Fig. 3b, c). DAAO expression could significantly improve glyphosate resistance in plants. In our study, the growth of WT plants was inhibited on medium containing 200 and 500 µM glyphosate, whereas the selected transgenic lines developed normally (Fig. 4). The 4-week-old WT plants all died within 9 days after application of 10 mM glyphosate, whereas transgenic plants showed no significant effects to the herbicide after being sprayed with the same dose (Fig. 6).
Proline has been known to be involved in the response to a number of environmental stresses, and it accumulates upon osmotic stress in a large number of different organisms. When stress is relieved, the proline concentration rapidly drops (Trovato et al. 2008). In our study, following glyphosate spraying, the proline content and POD activity increased in both WT and transgenic plants. This finding showed that glyphosate could induce proline accumulation and produce ROS. The proline content and POD activity in transgenic lines only slightly increased compared with those in WT plants (Fig. 5). Plant cells don’t possess endogenous DAAO activity (Pollegioni et al. 2007), so proline content in transgenic lines could be affected not only by a reduced stress due to glyphosate degradation but also by the direct activity of DAAO on proline.This result indicated that the effects of stress on WT plants were much more serious than those on transgenic plants under the same glyphosate concentration. The expression of DAAO in plants could convert glyphosate into AMPA and glyoxylate, which are less toxic than glyphosate (Pedotti et al. 2009); however, the time course of glyphosate degradation and the evaluation of residues were not carried out in our study, therefore more biochemical experiments need to be done in future. To date, no studies have reported the use of DAAO to enhance glyphosate resistance in plants. However, researchers have paid much attention on GO, an enzyme that belongs to the same structural family as DAAO (Mortl 2004). GO catalyzes the oxidation of several amines and d-amino acids (e.g., d-proline) using the same reaction mechanism as DAAO (Molla et al. 2003).
GO and d-amino acid oxidase catalyze the oxidative deamination of amino acids to yield the corresponding d-amino acids and, after hydrolysis, α-keto acids, ammonia (or primary amines), and hydrogen peroxide, but they differ in substrate specificity. In addition to neutral d-amino acids, GO catalyzes the oxidation of primary and secondary amines (e.g., glycine and sarcosine) partially sharing substrate specificity with monomeric sarcosine oxidase (Job et al. 2002). Glyphosate is a poor substrate of the flavoprotein GO. A number of GO variants with improved activity on glyphosate were recognized after site saturation mutagenesis on the positions of the active site (Nicolia et al. 2014; Pollegioni et al. 2011; Pedotti et al. 2009; Zhan et al. 2013).
GOX was isolated by Monsanto Co. in 1995 from Ochrobactrum anthropi, and it has actually been used in combination with resistant EPSP synthase to generate commercial varieties of glyphosate-resistant canola (Barry and Kishore 1995). Though the activity of wild-type GOX with glyphosate as substrate is quite low, the researchers improved the activity significantly by inserting genetic variability in the gene sequence (Pollegioni et al. 2011). Both GOX and DAAO are able to convert AMPA and glyoxylate by cutting the carbon–nitrogen bond on the carboxyl side. But the result of nucleotide sequence alignment showed there was no sequence identity between the two. Up to now, only one GOX gene has been found and used to achieve glyphosate-resistant plants, whereas the DAAO gene is present in many organisms, and thus potentially characterized by high variability (Pollegioni and Molla 2011). We propose that we can obtain a DAAO enzyme with high catalytic efficiency on the herbicide glyphosate through molecular evolution and DNA shuffling.
In conclusion, evolved DAAO may be a novel system for the development of glyphosate-resistant transgenic plants. It may represent an effective supplement or alternative to the transgene currently being used, which is based on EPSP being insensitive to inhibition by glyphosate.
Author contribution statement
RP, HH and QY conceived and designed the research project, analyzed the data and wrote the manuscript. BZ, XF, SY, BW, and JX conducted experiments and helped with data analysis. The manuscript was approved by all other authors.
Abbreviations
- AMPA:
-
Aminomethyl phosphonic acid
- DAAO:
-
d-amino acid oxidase
- EPSPS:
-
5-Enolpyruvylshikimate-3-phosphate synthase
- GAT:
-
Glyphosate acetyl-transferase
- GO:
-
Glycine oxidase
- GOX:
-
Glyphosate oxidoreductase
- PCR:
-
Polymerase chain reaction
- POD:
-
Peroxidase
- p251:
-
pYPX251
- ROS:
-
Reactive oxygen species
References
Barry GF, Kishore GM (1995) Glyphosate tolerant plants. US Patent 5,462,175
Bates LS, Waldren RP, Teare ID (1973) Rapid determination of free proline in water-stress studies. Plant Soil 39:205–207
Castle LA, Siehl DL, Gorton R, Lassner MW et al (2004) Discovery and directed evolution of a glyphosate tolerance gene. Science 304:1151–1154
Dill GM, Cajacob CA, Padgette SR (2008) Glyphosate-resistant crops: adoption, use and future considerations. Pest Manag Sci 64:326–331
Duan BQ, Wang XJ, Lu W, Lin M et al (2014) Development of highly glyphosate-tolerant tobacco by co-expression of glyphosate acetyltransferase gat and EPSPS G2-aroA genes. Crop J 2:164–169
Duke SO (2011) Glyphosate degradation in glyphosate-resistant and –susceptible crops and weeds. J Agric Food Chem 59:5835–5841
Duke SO (2014) Biotechnology: herbicide-resistant crops. In: Van Alfen N (ed) Encyclopedia of agriculture and food systems, vol 2. Elsevier, San Diego, pp 94–116
Duke SO, Powles SB (2008) Glyphosate: a once-in-a-century herbicide. Pest Manag Sci 64:319–325
Duke SO, Rimando AM, Pace PF, Reddy KN, Smeda RJ (2003) Isoflavone, glyphosate, and aminomethylphosphonic acid levels in seeds of glyphosate-treated, glyphosate-resistant soybean. J Agric Food Chem 51:340–344
Job V, Marcone GL, Pilone MS, Pollegioni L (2002) Glycine oxidase from Bacillus subtilis: characterization of a new flavoprotein. J Biol Chem 277:6985–6993
Klee HJ, Muskopf YM, Gasser CS (1987) Cloning of an Arabidopsis thaliana gene encoding 5-enolpyruvylshikimate-3-phosphate synthase: sequence analysis and manipulation to obtain glyphosate-tolerant plants. Mol Gen Genet 210:437–442
Lichtenthaler HK (1987) Chlorophylls and carotenoids: pigments of photosynthetic biomembranes. Methods Enzymol 148:350–382
Macadam JW, Nelson CJ, Sharp RE (1992) Peroxidase activity in the leaf elongation zone of tall fescue I. Spatial distribution of ionically bound peroxidase activity in genotypes differing in length of the elongation zone. Plant Physiol 99:872–878
Molla G et al (2003) Kinetic mechanisms of glycine oxidase from Bacillus subtilis. Eur J Biochem 270:1474–1482
Mortl M (2004) Structure–function correlation in glycine oxidase from Bacillus subtilis. J Biol Chem 279:29718–29727
Murray EE, Lotzer J, Eberle M (1989) Codon usage in plant genes. Nucleic Acids Res 17:477–498
Nicolia N, Ferradini G, Molla E, Veronesi D et al (2014) Expression of an evolved engineered variant of a bacterial glycine oxidase leads to glyphosate resistance in alfalfa. J Biotechnol 184:201–208
Pedotti M, Rosini E, Molla G, Savino C et al (2009) Glyphosate resistance by engineering the flavoenzyme glycine oxidase. J Biol Chem 284:36415–36423
Pollegioni L, Molla G (2011) New biotech applications from evolved d-amino acid oxidases. Trends Biotechnol 29:276–283
Pollegioni L, Piubelli L, Sacchi S, Pilone MS, Molla G (2007) Physiological functions of d-amino acid oxidase from yeast to humans. Cell Mol Llfe Sci 64:1373–1394
Pollegioni L, Schonbrunn E, Siehl D (2011) Molecular basis of glyphosate resistance-different approaches through protein engineering. FEBS J 278:2753–2766
Powles SB, Preston C (2006) Evolved glyphosate resistance in plants: biochemical and genetic basis of resistance. Weed Technol 20:282–289
Siehl DL, Castle LA, Gorton R, Keenan RJ (2007) The molecular basis of glyphosate resistance by an optimized microbial acetyltransferase. J Biol Chem 15:11446–11455
Sun YC, Chen YC, Tian ZX, Wang YP et al (2005) Novel AroA with high tolerance to glyphosate, encoded by a gene of Pseudomonas putida 4G-1 isolated from an extremely polluted environment in China. Appl Environ Microbiol 71:4771–4776
Tian YS, Xu J, Xiong AS, Yao QH et al (2011) Functional characterization of Class II 5-enopyruvylshikimate-3-phosphate synthase from Halothermothrix orenii H168 in Escherichia coli and transgenic Arabidopsis. Appl Microbiol Biotechnol 93:241–250
Trovato M, Mattioli R, Paolo C (2008) Multiple roles of proline in plant stress tolerance and development. Rendiconti Lincei 19:325–346
Waltz E (2010) Glyphosate resistance threatens Roundup hegemony. Nat Biotechnol 28:537–538
Xiong AS, Yao QH, Peng RH, Cheng ZM et al (2004) A simple, rapid, high fidelity and cost-effective PCR based two-step DNA synthesis (PTDS) method for long gene sequences. Nucleic Acids Res 32:e98
Xiong AS, Peng RH, Liu JG, Yao QH et al (2007) High efficiency and throughput system in directed evolution in vitro of reporter gene. Appl Microbiol Biotechnol 74:160–168
Xu J, Tian YS, Peng RH, Yao QH et al (2010) AtCPK6, a functionally redundant and positive regulator involved in salt/drought stress tolerance in Arabidopsis. Planta 231:1251–1260
Xue Y, Peng R, Xiong A, Li X, Zhang D, Yao Q (2009) Yeast heat-shock protein gene HSP26 enhances freezing tolerance in Arabidopsis. J Plant Physiol 166:844–850
Zhan T, Zhang K, Liu ZD et al (2013) Improving glyphosate oxidation activity of glycine oxidase from Bacillus cereus by derected evolution. PLoS One 8:e79175
Zhang X, Henriques R, Lin SS, Niu QW, Chua NH (2006) Agrobacterium-mediated transformation of Arabidopsis thaliana using the floral dip method. Nat Protoc 1:641–646
Zhou M, Xu H, Wei X, Zhu Z et al (2006) Identification of a glyphosate-resistant mutant of rice 5-enolpyruvylshikimate 3-phosphate synthase using a directed evolution strategy. Plant Physiol 140:184–195
Zhu B, Peng RH, Fu XY, Yao QH et al (2012) Enhanced transformation of TNT by Arabidopsis plants expressing an old yellow enzyme. PLoS One 7:e39861
Acknowledgments
The research was supported by the International Scientific and Technological Cooperation (13440701700), Agriculture Science Technology Achievement Transformation Fund (133919N1300), Young Foundation of Shanghai Academy of Agricultural Science (2010-14), National Natural Science Foundation (31200075, 31200076, 31200212). The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
We declare that we have no conflict of interest.
Additional information
Communicated by L. Peña.
Electronic supplementary material
Below is the link to the electronic supplementary material.
299_2015_1850_MOESM1_ESM.docx
Supplemental Table 1 Sequences for all primer sets used for chemical synthesis of DAAO gene according to plant codon usage bias. Letters in boxes are restriction endonuclease sites (DOCX 12 kb)
299_2015_1850_MOESM3_ESM.jpg
Fig. S2 RT-PCR analysis of the expression of DAAO (target gene) in other transgenic lines (Do1-15 except for Do3, Do6 and Do14) (JPEG 43 kb)
Rights and permissions
About this article
Cite this article
Han, H., Zhu, B., Fu, X. et al. Overexpression of d-amino acid oxidase from Bradyrhizobium japonicum, enhances resistance to glyphosate in Arabidopsis thaliana . Plant Cell Rep 34, 2043–2051 (2015). https://doi.org/10.1007/s00299-015-1850-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00299-015-1850-5