Abstract
Key message
HRE1α shows transcriptional activation activity in its C-terminal region via GCC box but not DRE/CRT and plays an important role in root development via root meristem cell division regulation.
Abstract
AtERF73/HRE1 protein, a member of the Arabidopsis AP2/ERF family, contains a conserved AP2/ERF DNA-binding domain. Here, we studied the molecular function of HRE1α, a splicing variant of AtERF73/HRE1, as well as its role in root development. HRE1α-overexpressing transgenic plants (OXs) showed tolerance to submergence. HRE1α showed transcriptional activation activity via GCC box but not DRE/CRT. The 121–211 aa region of HRE1α was responsible for the transcriptional activation activity, and the region was conserved among homologs of other species but was not found in other Arabidopsis proteins. HRE1α OXs showed increased primary root length due to elevated root cell division. Our results suggest that HRE1α functions as a transcription activator in the nucleus, and plays an important role in root development through regulation of root meristem cell division.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
During their life cycle, plants are faced with various environmental stresses. To adjust to such environmental stresses, plants trigger rapid defense responses via a number of signal transduction pathways. A major target of signal transduction is the cell nucleus, where signals lead to the transcriptional activation of numerous genes. Changes in the expression of genes encoding transcriptional regulators can have strong effects on plant stress tolerance (Yi et al. 2004).
AP2/ERF transcription factors, one of the largest plant transcription factor families, contain AP2/ERF DNA-binding domains of 57–66 amino acids in size (Nakano et al. 2006). AP2/ERF genes constitute a large multigene family, including 145 Arabidopsis genes, divided into four subfamilies (AP2, RAV, DREB/CBF, and ERF) according to their sequence similarities and numbers of AP2/ERF domains (Sakuma et al. 2002). DREB/CBF and ERF subfamily proteins contain single AP2/ERF domain. DREB/CBF subfamily genes have a function in the resistance of plants to abiotic stresses by recognizing the DRE/CRT cis-acting element, which has a core motif of (A/G)CCGAC (Thomashow 1999; Yamaguchi-Shinozaki and Shinozaki 1994). ERF subfamily is generally involved in the response to biotic stresses by recognizing the cis-acting element AGCCGCC, known as GCC box (Hao et al. 1998; Xu et al. 2007). Many ERF subfamily members also bind to DRE/CRT (Hsieh et al. 2013; Wu et al. 2007; Zhang et al. 2009; Zhu et al. 2010).
ERF transcription factors have been identified in various plant species, such as Arabidopsis thaliana, tobacco (Nicotiana tabacum), and tomato (Lycopersicon esculentum) (Fischer and Droge-Laser 2004; Onate-Sanchez and Singh 2002; Tournier et al. 2003). ERF proteins involved in defense responses against pathogen infection have also been extensively documented (Gutterson and Reuber 2004; Park et al. 2001; Shin et al. 2002). Recent studies have revealed a role for some ERF proteins in hormone and abiotic stress responses in plants (Shinozaki et al. 2003). Thus, ERF proteins play an important role not only in pathogen defense responses but also in tolerance to abiotic stresses. However, the molecular functions of ERF subfamily genes have been revealed for only a limited number of genes belonging to the subfamily.
Recently, two Arabidopsis ERF transcription factors, AtERF73/HRE1 and AtERF71/HRE2, have been identified to be responsive to hypoxia (Licausi et al. 2010). Expression of both AtERF73/HRE1 and AtERF71/HRE2 is increased under hypoxia, and overexpressing transgenic plants show resistance to hypoxia (Licausi et al. 2010). Although both genes are responsive to hypoxia, the response of AtERF73/HRE1 to hypoxia involves both ethylene-dependent and -independent pathways, while the response of AtERF71/HRE2 only involves ethylene-independent pathways (Hess et al. 2011; Licausi et al. 2010; Yang et al. 2011). It was recently reported that AtERF71/HRE2 is involved in the responses to osmotic stress as well as hypoxia and AtERF71/HRE2 functions as a transcription activator (Park et al. 2011). Single mutants of either AtERF73/HRE1 or AtERF71/HRE2 showed no significant difference compared with wild type (WT), whereas double mutants showed a more sensitive phenotype in response to hypoxic stress conditions (Licausi et al. 2010), suggesting that these two ERF proteins have a partially redundant molecular function in the hypoxic stress response. However, despite these studies, the molecular functions of AtERF73/HRE1 and its role during Arabidopsis development have not been reported.
In this study, we characterized the molecular functions of HRE1α, a splicing variant of AtERF73/HRE1, as a nuclear transcription factor having GCC box-dependent transcriptional activation activity in plants. In addition, we found that HRE1α is involved in root development through regulation of root meristem cell division.
Materials and methods
Plant materials and growth conditions
All Arabidopsis (Arabidopsis thaliana) plants used in this study were Columbia ecotype. Arabidopsis seeds were surface sterilized and germinated on agar plates as previously described (Park et al. 2011). Seedlings were grown under short-day (SD) conditions (cycles of 8-h light/16-h dark) or long-day (LD) conditions (cycles of 16-h light/8-h dark) at 22 °C.
Plasmid construction
To generate a plant expression vector overexpressing HRE1α, pFGL845, the entire open reading frame (ORF) of HRE1α was amplified by PCR and cloned into the SalI-BamHI site of pFGL571 (Park et al. 2011). To construct HRE1 promoter::β-glucuronidase (GUS) vector, the 5′ upstream promoter region (−1,000 to +330) relative to the ATG start codon of HRE1α was amplified by PCR and fused with the GUS gene. For synthetic Green Fluorescence Protein (sGFP)-fused HRE1α construct, the entire ORF of sGFP was amplified by PCR and cloned into pFGL845 in frame with HRE1α. To generate constructs for investigation of the transcriptional activation activity of HRE1α, the entire ORF or truncated HRE1α was amplified by PCR. The PCR products were then cloned into pBDGAL4 (Stratagene) in frame with the GAL4 DNA-binding domain (BD). For the reference plasmid vector expressing Renilla luciferase for transactivation analysis, the entire ORF of Renilla luciferase was amplified and cloned between modified CaMV 35S promoter and nos terminator of pFGL1122. For the effector-reporter plasmid vector for transactivation analysis, the core region of CaMV 35S promoter, the entire ORF of firefly luciferase and nos terminator were amplified and cloned into pBluescriptIIKS, after which four copies of GCC box or DRE/CRT were cloned in front of core CaMV 35S promoter, resulting in pFGL1498 and pFGL1499, respectively. The sequences of four copies of GCC box were 5′-CATAAGAGCCGCCACTCATAAGAGCCGCCACTCATAAGAGCCGCCACTCATAAGAGCCGCCACT-3′ and the sequences of four copies of DRE/CRT were 5′-ATTTCATGGCCGACCTGCTTTCATGGCCGACCTGCTTTCATGGCCGACCTGCTTTCATGGCCGACCTGCTT-3′. Then, CaMV 35S::HRE1α::nos terminator in pFGL845 was amplified and cloned into pFGL1498 or pFGL1499.
Plant transformation
Vectors for plant expression were introduced into Agrobacterium tumefaciens strain GV3101 (pMP90) using the freeze–thaw method (Höfgen and Willmitzer 1988) and Arabidopsis was transformed using the floral-dipping method (Clough and Bent 1998). Transgenic plants were selected on medium containing 25 mg/l of kanamycin.
Stress treatments
For submergence treatment, 4-week-old plants were submerged 5 cm below water level for 10 days, followed by desubmergence for 5 days.
For stress treatments prior to RT-PCR analysis, 10-day-old WT seedlings on MS plates were transferred to filter paper saturated with 300 mM NaCl, 300 mM mannitol, 100 µM ABA, or 10 µM MV, followed by incubation for 0, 1, 2, 4, 8, 12, or 24 h. For hypoxia treatment, mature rosette leaves detached from 4- to 5-week-old WT plants were floated on water and treated with 99.99 % nitrogen gas under dark conditions for 0, 1, 4, 6, 8, 12, 18, or 24 h.
Semi-quantitative reverse-transcription (RT)-PCR and quantitative RT-PCR
Total RNA was isolated using TRIzol reagent (Molecular Research Center). cDNA synthesis, semi-quantitative RT-PCR, and quantitative RT-PCR were performed as previously described (Park et al. 2011). The primers used for the PCR reactions are shown in Supplementary Table S1.
GUS activity analysis and transient gene expression in Arabidopsis protoplasts
GUS activity was histochemically detected using a protocol adapted from a previous report (Park et al. 2011).
To investigate the subcellular localization of HRE1α in Arabidopsis protoplasts, polyethylene glycol (PEG)-mediated protoplast transformations were performed according to the method described by Sheen (2001).
Transcriptional activation activity analysis in yeast
To investigate the transcriptional activation activity of HRE1α in yeast, pBDGAL4-HRE1α constructs were transformed into a yeast strain, YD116, which carries the GAL1pro::URA3 and UASpro::lacZ reporters. Transformants including BD-fusion vectors were selected on Synthetic Dropout (SD)-Trp. Transcriptional activation activities were confirmed by growth assay on SD-Trp-Ura and by quantitative β-galactosidase assay using 2-nitrophenyl-β-d-galacto-pyranoside (ONPG) as a substrate. Quantitative β-galactosidase assay was performed as previously described (Park et al. 2011), and the unit of β-galactosidase activity was calculated using the formula: 1,000 × OD420/(OD600 × assay time in min × assay volume in ml).
Transactivation analysis in Arabidopsis protoplast
For transactivation analysis in Arabidopsis protoplasts, reporter plasmid and effector-reporter plasmid harboring GCC box or DRE/CRT were transformed into Arabidopsis protoplasts. Reference plasmid was co-transformed for normalization. Firefly luciferase and Renilla luciferase activities were determined using the Dual-Luciferase Reporter Assay System (Promega, UK), according to the manufacturer’s instructions.
Measurement of root cell length and root meristem size
To determine root cell length and root meristem size, 8-day-old seedlings were used. The roots were excised from the seedlings and stained with 10 μM propidium iodide (PI) for 10 s, rinsed with distilled water, and mounted with distilled water. The specimens were examined with a confocal laser scanning microscope (LSM510NLO, CarlZeiss, Germany). Excitation wavelength of 536 nm was used to visualize the signals of PI staining. The emission wavelength was 617 nm.
Results and discussion
HRE1α is highly expressed in roots and trichomes and its expression is increased by hypoxia treatment
We isolated a cDNA of AtERF73/HRE1 from seedlings under hypoxia treatment and the cDNA corresponded to gene model 1 (At1g27360.1) of AtERF73/HRE1 on TAIR website (www.arabidopsis.org). Another splicing variant of AtERF73/HRE1, gene model 2 (At1g27360.2), was previously reported (Licausi et al. 2010). We named transcripts of At1g72360.1 and At1g72360.2 HRE1α and HRE1β, respectively (Supplementary Fig. S1). The predicted HRE1α protein, Arabidopsis AP2/ERF transcription factor, contains a conserved 58 aa AP2/ERF DNA-binding domain. Based on sequence homology and its conserved motif, HRE1α belongs to group VII (B-2) (Nakano et al. 2006).
We investigated the spatial and temporal expression patterns of HRE1α using semi-quantitative RT-PCR. Transcription of HRE1α increased as plants developed (Fig. 1a). In mature plants, HRE1α transcripts were highly detected in roots compared with levels in other organs (Fig. 1b). Previously, HRE1β was also reported to be highly expressed in roots under hypoxic conditions (Licausi et al. 2010). These results suggested that HRE1α might play an important role in root development. Expression patterns of HRE1α were also supported by GUS expression analysis using HRE1 promoter::GUS transgenic plants (Fig. 1c, d). Interestingly, high and specific promoter activity of HRE1 was observed in trichomes on leaves (Fig. 1e), suggesting that HRE1α/β might be involved in development and/or function of trichomes. However, in HRE1α-overexpressing transgenic plants (OXs), phenotypes of trichomes did not show any significant difference compared with those in WT plants (data not shown). Further studies will be necessary to elucidate the function of HRE1α/β related to its high expression in trichomes.
Temporal and spatial expression patterns of HRE1α. a Semi-quantitative RT-PCR analysis of HRE1α expression in 4-, 7-, 11-, 14-, or 21-day-old WT seedlings grown under SD conditions. b Semi-quantitative RT-PCR analysis of HRE1α expression in organs of 7-week-old WT plants grown under LD conditions. RT, roots; RS, rosette leaves; ST, stems; CA, cauline leaves; FC, floral clusters. In a and b, GAPc was used as an internal control. Similar results were obtained from at least two biological replicates, with one shown here. c Schematic map of HRE1 promoter::GUS for generating transgenic plants. d Histochemical assay of expression in Arabidopsis T2 plants carrying HRE1 promoter::GUS at 7, 11, 14, and 21 DAG under SD conditions. e Histochemical assay of GUS expression in leaves of 21-day-old HRE1 promoter::GUS T2 transgenic plants grown under LD conditions. In d and e, representative staining results are shown
To investigate the expression patterns of HRE1α under abiotic stress conditions, we performed quantitative RT-PCR analysis using HRE1α-specific primers and 10-day-old WT seedlings treated with hypoxia, NaCl, mannitol, ABA, or MV. Transcription of HRE1α increased within 4 h of hypoxia treatment (Fig. 2a), while transcript levels of HRE1α were reduced in response to NaCl, mannitol, ABA, or MV treatment (Supplementary Fig. S2). It has been reported that the expression level of HRE1β is also increased by hypoxia treatment (Licausi et al. 2010).
Submergence response of HRE1α OXs. a Quantitative RT-PCR analysis of HRE1α under 0-, 1-, 4-, 8-, and 12-h hypoxia treatments. GAPc was used as an internal control. Transcript level at 0 h was set to 1. Reactions of each technical replicate were performed in triplicate. Two technical replicates were measured for each biological replicate. Data shown are the mean ± SD (n = 6). Similar results were obtained from at least two biological replicates, with one shown here. b Response of WT and HRE1α OX plants to submergence. c Survival rates of WT and HRE1α OX plants after submergence treatment. Error bars represent standard deviation of three independent experiments. Ten plants were analyzed in each experiment
HRE1α OXs show tolerance to submergence
It was previously reported that HRE1β OXs were more resistant to anoxia than WT (Licausi et al. 2010). It is well known that hypoxic conditions often arise upon water logging or flooding. Since HRE1α expression was highly elevated under hypoxic conditions (Fig. 2a), we investigated the response of HRE1α OXs to submergence. To do this, we generated HRE1α OXs and selected three independent T1 lines for further analysis (Supplementary Fig. S3). HRE1α OXs were more resistant to submergence compared with WT plants (Fig. 2b, c). Both the increase of HRE1α transcript level under hypoxia condition and the submergence resistance of HRE1α OXs suggest that HRE1α is involved in resistance to submergence.
In rice (Oryza sativa), Submergence1 (Sub1) locus contains two or three ERF-like genes whose transcripts are regulated by submergence (Perata and Voesenek 2007). In deepwater rice, a pair of ERFs, SNORKEL1 and 2, also plays a key role in the adaptation of rice to submergence (Hattori et al. 2009). In Arabidopsis, ERF subfamily factors such as RAP2.2, RAP2.12, and AtERF71/HRE2 also function in the hypoxic response (Licausi et al. 2010). Interestingly, all of these genes (Sub1, SK1, SK2, RAP2.2, RAP2.12, AtERF71/HRE2, and AtERF71/HRE2) belong to group VII (B-2) ERFs, suggesting that group VII (B-2) ERFs have a function in the hypoxic response.
HRE1α shows transcriptional activation activity in its C-terminal region
AP2/ERF domain proteins are known to function as transcription factors (Okamuro et al. 1997). However, the molecular functions of HRE1α, including its transcriptional activation activity, have not yet been elucidated. First, we observed that HRE1α was localized in the nucleus using HRE1α-sGFP fusion protein (Fig. 3a) like HRE1β (Licausi et al. 2010).
Transcriptional activation activity of HRE1α. a Subcellular localization of HRE1α upon transient expression of HRE1α-sGFP fusion construct in Arabidopsis protoplasts. Left, GFP signal; middle, DAPI (4′,6-diamidino-2-phenylindole) staining; right, light microscope pictures. b Schematic map of GAL4 BD-fusion HRE1α construct along with the entire ORF and truncated HRE1α used in the analysis of transcriptional activation activity. Black box indicates AP2/ERF DNA-binding domain. c Growth assay of yeast transformants on SD-Trp-Ura. d Quantitative β-galactosidase assay in yeast. Data shown are the mean ± SD (n = 4). Similar results were obtained from at least two independent experiments, with one shown here. pBDGAL4 was used as a negative control. NC negative control
We investigated whether or not HRE1α functions as a transcriptional activator using a yeast system. The entire ORF of HRE1α was fused to GAL4 BD (Fig. 3b), after which the fusion construct was transformed into yeast cells. Based on the results of growth assay and quantitative β-galactosidase assay, HRE1α showed transcriptional activation activity in yeast (Fig. 3c, d), suggesting that HRE1α could function as a transcriptional activator in plants.
We next identified the domain of HRE1α responsible for the observed transcriptional activation activity. For this, we constructed a set of GAL4 BD-HRE1α vectors, including N160 (1 to 160 aa), N120 (1 to 120 aa), or N87 (1 to 87 aa) (Fig. 3b). In yeast, when the 161–211 aa region of HRE1α was deleted, transcriptional activation activity was significantly reduced compared with that of the entire ORF (Fig. 3c, d). Moreover, when the 121–211 aa region was deleted, truncated HRE1α showed no transcriptional activation activity (Fig. 3c, d). These results indicate that the region between 121 and 211 aa of HRE1α might be responsible for the observed transcriptional activation activity.
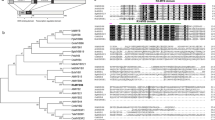
Although the 121–211 aa region of HRE1α had no known functional domain (data not shown), it contained conserved amino acid sequences in 13 ERF family proteins of other plant species, including Gossypium barbadense GbERF1, Gossypium hirsutum GhERF2, Manihot esculenta MeERF, Solanum lycopersicum JERF1, Ipomoea batatas IbERF, Ricinus communis RcERF, Jatropha curcas JcERF, Morus alba MaESF, Fagus sylvatica FsEREBP and FsERF, Capsicum annuum CaEREBP and CaPF1, and Vitis vinifera VvRAP2.12-like (Fig. 4a). These results indicate that the conserved regions might be involved in transcriptional activation activity. In addition, no other Arabidopsis protein except for HRE1α itself contained the conserved sequence in the 121–211 aa region (Fig. 4).
Multiple alignment and phylogenetic tree of HRE1α. a Multiple alignment of 121–211 aa region of HRE1α with corresponding regions of GbERF1, GhERF2, MeERF, JERF, IbERF, RcERF, JcERF, MaESF, FsEREBP, CaEREBP, VvRAP2.12-like, CaPF1, and FsERF. Alignment was made by ClustalW using default parameters. b Phylogenetic tree of 121–211 aa region of HRE1α and corresponding regions of GbERF1, GhERF2, MeERF, JERF, IbERF, RcERF, JcERF, MaESF, FsEREBP, CaEREBP, VvRAP2.12-like, CaPF1, and FsERF. Phylogenetic tree was made by ClustalW2-Phylogeny using default parameters
HRE1α has GCC box-dependent transcriptional activation activity
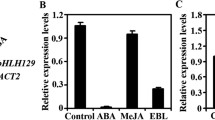
It is well known that the ERF subfamily of AP2/ERF transcription factors activates downstream genes via GCC box and/or DRE/CRT (Lee et al. 2005; Park et al. 2001; Tang et al. 2007; Zhang et al. 2004). To determine GCC box and/or DRE/CRT-dependent transcriptional activation activity of HRE1α in Arabidopsis protoplasts, we generated reporter, effector-reporter, and reference plasmids (Fig. 5a) and then introduced them into Arabidopsis protoplasts. When the GCC box-containing reporter plasmid was introduced together with HRE1α, relative luciferase activity was elevated about fivefold compared to that without HRE1α (Fig. 5b). On the other hand, when the DRE/CRT-containing reporter plasmid was transformed with HRE1α, relative luciferase activity showed no significant differences compared to that without HRE1α (Fig. 5c). These results indicate that HRE1α can activate the firefly luciferase gene via GCC box but not DRE/CRT, suggesting that HRE1α might act as a transcriptional activator via GCC box only. However, we could not confirm specific binding of HRE1α to GCC box in vitro, since host E. coli cells did not produce adequate amount of recombinant HRE1α protein.
Transactivation analysis of HRE1α. a Schematic maps of the effector-reporter, reporter, and reference plasmids used in transactivation analysis. Relative luciferase activity in transactivation analysis for GCC box (b) and DRE/CRT (c). In b and c, firefly luciferase activities of the effector-reporter plasmid and the reporter plasmid were measured in Arabidopsis protoplasts. Transformation efficiency was normalized with Renilla luciferase activity of the reference plasmid. Relative luciferase activity represents the fold-increase in the normalized luciferase activity of the effector-reporter plasmid compared to that of the reporter plasmid alone. Data shown are the mean ± SD (n = 5). Similar results were obtained from at least three independent experiments, with one shown here
HRE1α is involved in root development
HRE1α was highly expressed in roots compared with other organs of mature plants such as leaves, floral clusters, and stems (Fig. 1b). Therefore, we investigated the involvement of HRE1α in root development. Quantitative RT-PCR analysis using HRE1α-specific primers revealed that the expression level of HRE1α was much higher in roots than in shoots in 7-day-old WT seedlings, similar to mature plants (Fig. 6a). Moreover, we found that the primary root length was longer in HRE1α OXs than in WT plants (Fig. 6b, c), suggesting that HRE1α might be involved in root development.
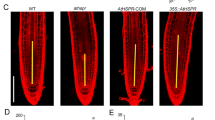
Root development in HRE1α OXs. a Quantitative RT-PCR analysis of HRE1α expression in shoots and roots of 7-day-old WT plants under SD conditions. GAPc was used as an internal control. Transcript level in shoots was set to 1. Reactions of each technical replicate were performed in triplicate. Two technical replicates were measured for each biological replicate. Data shown are the mean ± SD (n = 6). Similar results were obtained from at least two biological replicates, with one shown here. b Root development of 14-day-old WT and HRE1α OX seedlings under SD conditions. c Primary root lengths of WT and HRE1α OXs. Primary root lengths were measured at 14 DAG under SD conditions. Data shown are the mean ± SD (n = 21). The number of root meristem cortex cells (d) and root meristem length (e), and lengths of hair cells and non-hair cells in roots (f) of 8-day-old WT and HRE1α OXs under SD conditions. In d, e, and f, data shown are the mean ± SD (n = 12)
The balance between cell division and expansion is important for the maintenance and the size of root meristem and controls the overall rate of root growth (Beemster and Baskin 1998). To determine the role of HRE1α in the root development, we analyzed root cells of HRE1α OXs. Since cell division activity in roots is reflected by root meristem size, we compared the number of root meristem cortex cells, including cells from the quiescent center to the first elongated cell, as well as root meristem size between WT and HRE1α OXs. The number of cortex cells of HRE1α OXs was almost twofold higher than that of WT (Fig. 6d), and the root meristem size of HRE1α OXs was twofold larger than that of WT (Fig. 6e). These data indicate that cell division activity is higher in root of HRE1α OXs compared with WT. We also analyzed root cell expansion by comparing hair cell length as well as non-hair cell length between HRE1α OXs and WT. The lengths of hair cells and non-hair cells were not significantly different between HRE1α OXs and WT (Fig. 6f), indicating that root cell expansion of HRE1α OXs is similar to that of WT. These results suggest that the increased primary root length in HRE1α OXs can be attributed to increased root cell division.
Taken together, our results suggest that HRE1α might function as a transcription activator via GCC box in the nucleus. In addition, HRE1α is involved in root development through regulation of root meristem cell division.
Author contribution
Most experiments and data analysis were performed by both H.-Y. Seok and V.N. Tarte. S.-Y. Lee contributed to some experimental design and data interpretation. H.-Y Park assisted in the isolation of HRE1α and the generation and analysis of HRE1α OXs. Y.-H. Moon was responsible for the overall conceptualization and supervision of the experiments and worked on data processing and manuscript preparation.
Abbreviations
- ABA:
-
Abscisic acid
- CaMV:
-
Cauliflower mosaic virus
- DAG:
-
Days after germination
- DRE/CRT:
-
Dehydration-responsive element/C-repeat
- GAPc:
-
Glyceraldehyde-3-phosphate dehydrogenase
- MV:
-
Methyl viologen
References
Beemster GT, Baskin TI (1998) Analysis of cell division and elongation underlying the developmental acceleration of root growth in Arabidopsis thaliana. Plant Physiol 116:1515–1526
Clough SJ, Bent AF (1998) Floral dip: a simplified method for Agrobacterium mediated transformation of Arabidopsis thaliana. Plant J 16:735–743
Fischer U, Droge-Laser W (2004) Overexpression of NtERF5, a new member of the tobacco ethylene response transcription factor family enhances resistance to tobacco mosaic virus. Mol Plant Microbe Interact 17:1162–1171
Gutterson N, Reuber TL (2004) Regulation of disease resistance pathways by AP2/ERF transcription factors. Curr Opin Plant Biol 7:465–471
Hao DY, Ohme-Takagi M, Sarai A (1998) Unique mode of GCC box recognition by the DNA-binding domain of ethylene-responsive element-binding factor (ERF domain) in plant. J Biol Chem 273:26857–26861
Hattori Y, Nagai K, Furukawa S, Song XJ, Kawano R, Sakakibara H, Wu J, Matsumoto T, Yoshimura A, Kitano H, Matsuoka M, Mori H, Ashikari M (2009) The ethylene response factors SNORKEL1 and SNORKEL2 allow rice to adapt to deep water. Nature 460:1026–1031
Hess N, Klode M, Anders M, Sauter M (2011) The hypoxia responsive transcription factor genes ERF71/HRE2 and ERF73/HRE1 of Arabidopsis are differentially regulated by ethylene. Physiol Plant 143:41–49
Höfgen R, Willmitzer L (1988) Storage of competent cells for Agrobacterium transformation. Nucleic Acids Res 16:9877
Hsieh EJ, Cheng MC, Lin TP (2013) Functional characterization of an abiotic stress-inducible transcription factor AtERF53 in Arabidopsis thaliana. Plant Mol Biol 82:223–237
Lee JH, Kim DM, Lee JH, Kim J, Bang JW, Kim WT, Pai HS (2005) Functional characterization of NtCEF1, an AP2/EREBP-type transcriptional activator highly expressed in tobacco callus. Planta 222:211–224
Licausi F, van Dongen JT, Giuntoli B, Novi G, Santaniello A, Geigenberger P, Perata P (2010) HRE1 and HRE2, two hypoxia-inducible ethylene response factors, affect anaerobic responses in Arabidopsis thaliana. Plant J 62:302–315
Nakano T, Suzuki K, Fujimura T, Shinshi H (2006) Genome-wide analysis of the ERF gene family in Arabidopsis and rice. Plant Physiol 140:411–432
Okamuro JK, Caster B, Villarroel R, Montagu MV, Jofuku KD (1997) The AP2 domain of APETALA2 defines a large new family of DNA binding proteins in Arabidopsis. Proc Natl Acad Sci USA 94:7076–7081
Onate-Sanchez L, Singh KB (2002) Identification of Arabidopsis ethylene-responsive element binding factors with distinct induction kinetics after pathogen infection. Plant Physiol 128:1313–1322
Park JM, Park CJ, Lee SB, Ham BK, Shin R, Paek KH (2001) Overexpression of the tobacco Tsi1 gene encoding an EREBP/AP2-type transcription factor enhances resistance against pathogen attack and osmotic stress in tobacco. Plant Cell 13:1035–1046
Park HY, Seok HY, Woo DH, Lee SY, Tarte VN, Lee EH, Lee CH, Moon YH (2011) AtERF71/HRE2 transcription factor mediates osmotic stress response as well as hypoxia response in Arabidopsis. Biochem Biophys Res Commun 414:135–141
Perata P, Voesenek LACJ (2007) Submergence tolerance in rice requires Sub1A, an ethylene-response-factor-like gene. Trends Plant Sci 12:43–46
Sakuma Y, Liu Q, Dubouzet JG, Abe H, Shinozaki K, Yamaguchi-Shinozaki K (2002) DNA-binding specificity of the ERF/AP2 domain of Arabidopsis DREBs, transcription factors involved in dehydration- and cold-inducible gene expression. Biochem Biophys Res Commun 290:998–1009
Sheen J (2001) Signal transduction in maize and Arabidopsis mesophyll protoplasts. Plant Physiol 127:1466–1475
Shin R, Park JM, An JM, Paek KH (2002) Ectopic expression of Tsi1 in transgenic hot pepper plants enhances host resistance to viral, bacterial, and oomycete pathogens. Mol Plant Microbe Interact 15:983–989
Shinozaki K, Yamaguchi-Shinozaki K, Seki M (2003) Regulatory network of gene expression in the drought and cold stress responses. Curr Opin Plant Biol 6:410–417
Tang M, Sun J, Liu Y, Chen F, Shen S (2007) Isolation and functional characterization of the JcERF gene, a putative AP2/EREBP domain-containing transcription factor, in the woody oil plant Jatropha curcas. Plant Mol Biol 63:419–428
Thomashow MF (1999) Plant cold acclimation: freezing tolerance genes and regulatory mechanisms. Annu Rev Plant Physiol Plant Mol Biol 50:571–599
Tournier B, Sanchez-Ballesta MT, Jones B, Pesquet E, Regad F, Latche A, Pech JC, Bouzayen M (2003) New members of the tomato ERF family show specific expression pattern and diverse DNA binding capacity to the GCC box element. FEBS Lett 550:149–154
Wu L, Chen X, Ren H, Zhang Z, Zhang H, Wang J, Wang XC, Huang R (2007) ERF protein JERF1 that transcriptionally modulates the expression of abscisic acid biosynthesis-related gene enhances the tolerance under salinity and cold in tobacco. Planta 226:815–825
Xu ZS, Xia LQ, Chen M, Cheng XG, Zhang RY, Li LC, Zhao YX, Lu Y, Ni YZ, Liu L, Qiu ZG, Ma YZ (2007) Isolation and molecular characterization of the Triticum aestivum L. ethylene-responsive factor 1 (TaERF1) that increases multiple stress tolerance. Plant Mol Biol 65:719–732
Yamaguchi-Shinozaki K, Shinozaki K (1994) A novel cis-acting element in an Arabidopsis gene is involved in responsiveness to drought, low-temperature, or high-salt stress. Plant Cell 6:251–264
Yang CY, Hsu FC, Li JP, Wang NN, Shih MC (2011) The AP2/ERF transcription factor AtERF73/HRE1 modulates ethylene responses during hypoxia in Arabidopsis. Plant Physiol 156:202–212
Yi SY, Kim JH, Joung YH, Lee S, Kim WT, Yu SH, Choi D (2004) The pepper transcription factor CaPF1 confers pathogen and freezing tolerance in Arabidopsis. Plant Physiol 136:2862–2874
Zhang H, Huang Z, Xie B, Chen Q, Tian X, Zhang X, Zhang H, Lu X, Huang D, Huang R (2004) The ethylene-, jasmonate-, abscisic acid-, and NaCl-responsive tomato transcription factor JERF1 modulates expression of GCC box-containing genes and salt tolerance in tobacco. Planta 220:262–270
Zhang G, Chen M, Li L, Xu Z, Chen X, Guo J, Ma Y (2009) Overexpression of the soybean GmERF3 gene, an AP2/ERF type transcription factor for increased tolerances to salt, drought, and diseases in transgenic tobacco. J Exp Bot 60:3781–3796
Zhu Q, Zhang J, Gao X, Tong J, Xiao L, Li W, Zhang H (2010) The Arabidopsis AP2/ERF transcription factor RAP2.6 participates in ABA, salt and osmotic stress responses. Gene 457:1–12
Acknowledgments
This work was supported by a 2-Year Research Grant of Pusan National University.
Conflict of interest
The authors have no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by Y.-I. Park.
H.-Y. Seok and V.N. Tarte equally contributed to the paper.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Seok, HY., Tarte, V.N., Lee, SY. et al. Arabidopsis HRE1α, a splicing variant of AtERF73/HRE1, functions as a nuclear transcription activator in hypoxia response and root development. Plant Cell Rep 33, 1255–1262 (2014). https://doi.org/10.1007/s00299-014-1613-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00299-014-1613-8