Abstract
In order to determine the different roles of rice (Oryza sativa L.) cytosolic ascorbate peroxidases (OsAPXa and OsAPXb, GenBank accession nos. D45423 and AB053297, respectively) under salt stress, transgenic Arabidopsis plants over-expressing OsAPXa or OsAPXb were generated, and they all exhibited increased tolerance to salt stress compared to wild-type plants. Moreover, transgenic lines over-expressing OsAPXb showed higher salt tolerance than OsAPXa transgenic lines as indicated by root length and total chlorophyll content. In addition to ascorbate peroxidase (APX) activity, antioxidant enzyme activities of catalase (CAT), superoxide dismutase (SOD) and glutathione reductase (GR), which are also involved in the salt tolerance process, and the content of H2O2 were also assayed in both transgenic and wild-type plants. The results showed that the overproduction of OsAPXb enhanced and maintained APX activity to a much higher degree than OsAPXa in transgenic Arabidopsis during treatment with different concentrations of NaCl, enhanced the active oxygen scavenging system, and protected plants from salt stress by equilibrating H2O2 metabolism. Our findings suggest that the rice cytosolic OsAPXb gene has a more functional role than OsAPXa in the improvement of salt tolerance in transgenic plants.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Soil salinity is a major abiotic stress for agriculture practices worldwide, deleteriously affecting the growth and yield of a variety of crops (Zhu 2001). Salinity, mainly caused by NaCl, induces a wide range of responses in plants. In addition to toxic effects, salt stress can also induce oxidative stress with the formation and accumulation of reactive oxygen species (ROS) (Wang et al. 2003). Plants have developed several antioxidant enzymes such as ascorbate peroxidase (APX, EC 1.11.1.11), superoxide dismutase (SOD, EC 1.15.1.1), catalase (CAT, EC 1.11.1.6) and glutathione reductase (GR, EC 1.6.4.2), to detoxify ROS and protect their cells from oxidative injury.
APX is thought to play the most essential role in scavenging ROS and protecting cells against these toxic effects in higher plants, algae, euglena and other organisms (Ishikawa et al. 2003; Panchuk et al. 2005; Sano et al. 2001; Teixeira et al. 2006). APX exists as isoenzymes distributed in distinct cellular compartments such as the cytosol, mitochondria and peroxisomes. Increased activity of different APX isoforms in response to environmental stresses such as salinity and drought has been reported in different plant species, indicating possible functional specialization of the respective isoenzymes in eliminating H2O2 from cells (Sharma and Dubey 2005; Teixeira et al. 2006; Tsai et al. 2005).
Cytosolic APX (cAPX) isoenzymes have been studied extensively. They can be activated by different types of stress and seem to have a more general stress-protective function (Davletova et al. 2005; Fourcroy et al. 2004;). In rice, the APX gene family has eight members that encode two cytosolic and other subcellular isoforms. It has been suggested that individual APX genes are differentially up-regulated during NaCl treatment, and in rice, the accumulation of cytosolic OsAPx2 transcript has been shown to be more remarkable than that of OsAPx1 (Menezes-Benavente et al. 2004; Teixeira et al. 2006). Until now, however, relatively few studies have attempted to characterize the different functions of these two rice cAPX genes using transgenic plants exposed to salinity stress.
The cDNAs encoding two rice APXs in the cytosol have been identified previously (Lu et al. 2005). Further, we observed the different properties of these two cAPX proteins by functional expression in Escherichia coli (Lu et al. 2005). In this study, in an attempt to identify the different potential roles of two rice cAPX genes (OsAPXa and OsAPXb, which correspond to OsAPx1 and OsAPx2, respectively), we generated transgenic plants and explored their stable over-expression in Arabidopsis using northern and western blot analysis. Transgenic Arabidopsis plants over-expressing OsAPXb showed much greater tolerance to NaCl than those over-expressing OsAPXa according to a comprehensive study of root length, total chlorophyll and H2O2 content and the enzymatic activities of APX, CAT, SOD and GR. The results suggest that the function of OsAPXb in relation to salinity tolerance is more important than that of OsAPXa in transgenic plants.
Materials and methods
Plant material and stress treatment
Sterilized seeds of Arabidopsis ecotype Columbia were germinated in MS semi-solid medium for 1 week after 2 days jarovization at 4°C, then grown under a light period of 16 h at 23°C with a humidity of 60%. One-week-old seedlings were subjected to salt stress for 7 days using the following concentrations of NaCl: 50, 100 and 200 mM. The Arabidopsis seedlings were collected and frozen in liquid N2 and stored at −70°C until it is used in the enzyme activity assay and other analyses.
Construct of expressing plasmid and transformation
The full lengths of OsAPXa and OsAPXb (GenBank accession nos: D45423 and AB053297, respectively) amplified by RT-PCR were fused into a T-vector (Lu et al. 2005). Both fragments were then cut with HincII and SacI and ligated into the SmaI–SacI site of the binary vector pBI121 with replacement of the GUS fragment and under the control of the cauliflower mosaic virus 35S (CaMV35S) promoter. The resulting recombinant plasmids (pBI121–OsAPXa and pBI121–OsAPXb) were transformed into Arabidopsis by Agrobacterium-mediated vacuum infiltration.
After harvesting the self-pollinated transformants (T0), the seeds were plated on MS semi-solid medium supplemented with 50 mg/l Kana and transgenic lines (T1) were selected. The integration and expression of each transgene in different homozygous lines (T2) was confirmed by northern and western blot analysis.
Northern and western blot analysis
RNA samples were prepared using the Trizol reagent (Invitrogen) extraction procedure according to the manufacturer’s instructions. Total RNA aliquots of 5 μg were separated in 1.2% denaturation formaldehyde agarose gel and then transferred onto hybond N+ membranes (Amersham). OsAPXa and OsAPXb cDNA labeled with DIG were used as specific probes for the detection of transcripts. Color was developed using NBT/BCIP (Sigma).
OsAPXa and OsAPXb protein were purified as described by Lu et al. (2005), and used to raise polyclonal antibody in rabbits as per standard procedures (Harlow and Lane 1988). For western blotting, extraction of soluble protein from Arabidopsis was essentially carried out as reported previously (Wehmeyer et al. 1996). The amount of protein was estimated according to the Bradford method (Bradford 1976). Briefly, 20 μg of soluble protein was resolved using 12% standard SDS-PAGE and transferred onto a nitrocellulose membrane. Protein blots were probed with rabbit antiserum against OsAPXa and OsAPXb at a dilution of 1:4,000 and then visualized using alkaline phosphatase-conjugated goat anti-rabbit IgG.
Salt tolerance test with transgenic plants
To assess the relative salinity tolerance of various plants, wild-type and T2 homozygous transgenic seeds were germinated on MS semi-solid medium with or without 50 mg/l Kana. After 7 days, surviving seedlings were transferred to MS semi-solid medium supplemented with different concentrations of NaCl and salinity stress was imposed for 7 days. The salt tolerance of the plants was estimated by measuring the length of their roots and their total chlorophyll content.
Preparation of protein extracts
The enzyme activities of the soluble protein were measured after extraction according to the method of Lee and Lee (2000) with some modifications. Total protein extraction was carried out using 500 mg of tissue in extraction buffer (100 mM potassium phosphate buffer, pH 7.8, 1 mM EDTA, 1% (w/v) PVP and 10% glycerol) except for the measurement of APX, in which case the plant tissue was homogenized in 100 mM sodium phosphate (pH 7.0) containing 1 mM EDTA and 5 mM ascorbate.
Enzyme activity assays
APX activity was assayed according to the method of Chen and Asada (1989), and was determined in a reaction mixture consisting of 50 mM phosphate buffer (pH 7.0), 0.5 mM ascorbate and 0.2 mM H2O2 by the change in absorbance at 290 nm (E = 2.8 mM−1 cm−1). The results were calculated in terms of micromole of ascorbate oxidized per minute.
CAT activity was measured by following the decomposition of H2O2 at 240 nm (E = 39.4 mM−1 cm−1) according to the method of Aebi (1974) in a reaction mixture containing the appropriate extract in 50 mM phosphate buffer (pH 7.0). The reaction was initiated by addition of 10 mM H2O2 and 1 U of CAT activity was defined as micromole H2O2 degraded per minute.
SOD activity was assayed by monitoring the inhibition of the photochemical reduction of nitro blue tetrazolium (NBT) according to the protocol of Giannopolitis and Ries (1977). One unit of SOD activity was defined as the amount of enzyme required to cause 50% inhibition in the reduction of NBT at 560 nm expressed in unit per milligram protein.
GR activity was assayed according to Rao et al. (1996). The reaction was initiated by the addition of GSSG, and was followed by monitoring the oxidation of NADPH at 340 nm. The specific activity of the enzyme was expressed as μmol NADPH oxidized min−1 mg−1 protein.
Measurement of the total chlorophyll and H2O2 content
The total chlorophyll content was determined spectrophotometrically according to the method described by Arnon (1949). The H2O2 content was determined by homogenizing 500-mg leaf samples in 100 mM phosphate buffer (pH 7.0) and then measuring the H2O2 concentration in a reaction mixture 50 mM phosphate buffer (pH 7.0), 40 U/ml HRP and 0.05% guiacol according to the modified method of Bernt and Bergmeyer (1974).
Statistical analysis
All the experiments were carried out in triplicate, and 15 seedlings per transgenic line were examined each time. The values shown in the figures are mean values ± SD. Means were compared by one-way analysis of variance and Duncan’s multiple range test with a 5% level of significance.
Results
Identification of transgenic Arabidopsis
Transgenic Arabidopsis plants over-expressing the rice OsAPXa or OsAPXb gene were produced by introduction of OsAPXa and OsAPXb cassettes from the reconstructed vectors pBI121–OsAPXa and pBI121–OsAPXb, respectively. Respective expression of OsAPXa and OsAPXb in four kanamycin-resistant homozygous T2 transgenic lines was confirmed by northern blot hybridization and APX activity assay (Fig. 1). The results showed different transcriptional levels of OsAPXa or OsAPXb in the transgenic lines compared with the wild-type (Fig. 1a, b) as well as higher APX activity (Fig. 1c). Both results indicated that OsAPXa and OsAPXb were constitutively and functionally expressed in the transgenic Arabidopsis. Further, the transgenic plants did not show any obvious differences in growth at either the vegetative or reproductive stage (data not shown). Subsequently, transgenic plants TA-3 and TB-1 which had a similar APX activity level were chosen for the examination of salt tolerance.
Northern blot analysis and APX activity assay of transgenic T2 generation Arabidopsis. a, b Northern blot of transgenic lines over-expressing OsAPXa (TA-1, TA-3, TA-4 and TA-8) and OsAPXb (TB-1, TB-2, TB-5 and TB-8). c Screening of APX activity in both transgenic lines. WT, wild-type. Data are the mean ± SD of three separate experiments. Different letters in italic (a and b) indicate significant differences (P < 0.05) of individual examined lines as compared to the WT
Salt tolerance in transgenic Arabidopsis over-expressing OsAPXa and OsAPXb
To test whether over-expression of OsAPXa and OsAPXb in Arabidopsis could enhance salt tolerance, wild-type and the T2 generation of the two transgenic lines (TA-3 and TB-1) were treated with 0 mM (control), 50, 100 and 200 mM NaCl. There were no difference between the three lines when grown under control conditions (Fig. 2a); however, during treatment with increasing concentrations of NaCl, wild-type plants displayed progressive chlorosis, growth inhibition, and decreased vigor. In contrast, both transgenic lines showed markedly enhanced salt tolerance, and treatment with 50 mM NaCl had no influence on their growth (Fig. 2a). TB-1 exhibited obviously better growth performance than TA-3 when treated with 100 or 200 mM NaCl; however, both the wild-type and transgenic Arabidopsis were severely damaged when treated with 200 mM NaCl (Fig. 2a).
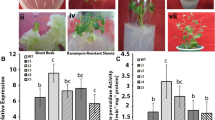
Improvement of salt tolerance with over-expression of OsAPXb compared with over-expression of OsAPXa in transgenic T2 generation Arabidopsis. Seedlings were grown on medium supplemented with 0 (control), 50, 100 or 200 mM NaCl for 7 days (a) then root length and total chlorophyll content were measured (b, c). Values represent the mean ± SD (N = 15). Different italic letters (a–c) indicate significant differences (P < 0.05) between lines. WT, wild-type; TA-3 and TB-1, transgenic plants over-expressing OsAPXa and OsAPXb, respectively
Next, we analyzed the differing salt tolerance of each transgenic line in detail by obtaining measurements of root length and the total chlorophyll content. Without NaCl treatment, there was almost no difference in root length and total chlorophyll content among the wild-type and two transgenic plants (Fig. 2b, c). However, root growth of the transgenic plants was less inhibited by NaCl treatment than that of the wild-type, and TB-1 showed more pronounced root length growth than TA-3 (Fig. 2b). Furthermore, although the total chlorophyll content decreased upon salt stress in both the transgenic and wild-type plants, the extent of this decline in TB-1 was less than that in TA-3 (Fig. 2c).
Salt tolerance and increased APX activity in the transgenic Arabidopsis
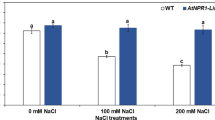
To determine whether the enhanced OsAPXa and OsAPXb activity was correlated with the ability of the plants to survive under salt stress, and to identify the enzyme whose activity is least affected by treatment with NaCl, we investigated APX activity in both the transgenic and wild-type plants under different concentrations of NaCl. As expected, nearly a two-fold increase in enzymatic activity was observed in both transgenic plants under normal conditions (Fig. 3a). When exposed to 50 mM NaCl, APX activity in the transgenic line TB-1 increased more evidently than that in TA-3, but both showed increased activity compared with the wild-type. APX activities in all the tested lines decreased gradually under conditions of 100 and 200 mM NaCl; however, activity in both transgenic plants decreased more slowly than that in the wild-type, and TB-1 seemed more competent than TA-3 in preventing this decline (Fig. 3a).
Higher APX activity in transgenic plants over-expressing OsAPXa or OsAPXb under different concentrations of NaCl. a APX activity in wild-type (WT), OsAPXa (TA-3) and OsAPXb (TB-1) over-expressing transgenic Arabidopsis plants. Data represent the mean ± SD of three independent experiments (N = 15). Means denoted by the same italic letter did not differ significantly (P < 0.05). b, c Northern and western blot analysis of the two transgenic lines treated with 50 mM NaCl
In order to confirm that the increase in APX activity was correlated with the activities of OsAPXa and OsAPXb in the transgenic plants treated with 50 mM NaCl, we further studied the changes in transcript and expression levels of OsAPXa and OsAPXb by northern and western blot analysis. As shown in Fig. 3b, c, when the plants transferred to 50 mM NaCl, both the transcript and expression levels showed evident accumulation of the two genes in the transgenic plants, with OsAPXb being induced at a much higher level than OsAPXa. This suggests that OsAPXb is more sensitive to salt stress than OsAPXa, and higher APX activity caused by this enhanced expression results in the increased tolerance of the transgenic Arabidopsis plants to salt.
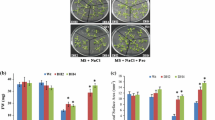
An improvement in salt tolerance was also observed in transgenic Arabidopsis plants TA-1 and TB-5, which exhibited the highest APX activity among the transgenic lines (Fig. 1, 4). When exposed to 100 mM NaCl, TA-1 and TB-5 grew comparatively well, while the growth of TA-3 and TB-1 was influenced to some extent and that of the wild-type plants was severely damaged (Fig. 4a). After treatment with salt, both the OsAPXa and OsAPXb transcripts increased notably in the transgenic plants, with APX activity being higher in TA-1 and TB-5 compared to TA-3 and TB-1 (Fig. 4b, c).
Enhancement of salt tolerance in transgenic Arabidopsis by up-regulated transcript level of OsAPXa (TA-1 and TA-3) and OsAPXb (TB-1 and TB-5). a Transgenic plants were grown on MS medium supplemented with 100 mM NaCl. b Northern blot analysis. c APX activity assay of transgenic plants. Values represent the mean ± SD (N = 15). Different italic letters (a–d) indicate significant differences (P < 0.05) between lines. WT, wild-type
Effects of salt stress on the H2O2 content and activities of CAT, SOD and GR
To clarify the mechanism controlling the metabolic reactions representing cellular salinity damage and protection, we also determined the activities of CAT, SOD and GR. There were almost no differences in the activities of these three enzymes among the two transgenic lines and wild-type under normal conditions. However, when treated with different concentrations of NaCl, the activities of CAT, SOD and GR in wild-type seedlings declined rapidly except for a slight increase in SOD and GR activities under conditions of 50 mM NaCl. At a NaCl concentration of 200 mM almost no enzyme activity was detected in the wild-type plants. In the two transgenic lines, on the other hand, the activities of these three enzymes in response to various concentrations of salt were less affected than those in the wild-type. Further, changes in activities of CAT, SOD and GR were much slower in TB-1 than TA-3 regardless of whether they increased or declined (Fig. 5a–c).
Effect of increasing NaCl concentrations on the activities of CAT (a), SOD (b), GR (c) and levels of H2O2 (d) in transgenic Arabidopsis over-expressing OsAPXa and OsAPXb (TA-3 and TB-1), respectively. Values represent the mean ± SD (N = 15). Different italic letters (a–c) at the top of the error bars indicate statistical different means (P < 0.05). WT, wild-type
To test whether H2O2 was involved in these changes in enzymatic activity, its content was assayed during exposure to different concentrations of NaCl. The results showed that H2O2 content was negatively correlated with changes in the activities of these antioxidant enzymes. Salt stress caused a marked increase in the level of H2O2 in wild-type Arabidopsis with an increase in NaCl concentration. In comparison to the wild-type, the H2O2 content of both transgenic lines increased much more slowly, and in TB-1, there was no obvious change in H2O2 level following treatment with 50 or 100 mM NaCl (Fig. 5d). This slow increase in H2O2 content may have been responsible for the much stronger tolerance of TB-1 grown under different concentrations of NaCl (Fig. 2a).
Discussion
Salt stress is a key environmental stress in agricultural practices around the world, especially in tropical areas and irrigated fields, where salinization affects a large land area. Salt stress can also result in oxidative stress; however, APX, especially cAPX, is thought to play an essential role in protecting plants from such stress (Shigeoka et al. 2002). In Arabidopsis, there are eight APX isoforms, and expression of the cytosolic forms (Apx1 and Apx2) is rapidly induced by elevated levels of light and heat stress. Apx1 is directly involved in H2O2 scavenging during light stress, while the increase in the Apx2 transcript is more profound during heat stress (Davletova et al. 2005; Panchuk et al. 2002). Arabidopsis peroxisomal Apx3 is thought to protect tobacco leaves from oxidative stress damage (Wang et al. 1999). The two rice cAPX gene transcripts have previously been shown to increase after treatment with salt (Menezes-Benavente et al. 2004; Teixeira et al. 2006). However, the different roles of these two cAPXs in plant cells remain elusive. Genetic engineering provides an effective tool for researching gene functions in transgenic plants under salt stress.
In this study, we examined the different potential functions of OsAPXa and OsAPXb in Arabidopsis plants under salt stress. Under normal conditions, over-expression of both OsAPXa and OsAPXb in transgenic lines TA-3 and TB-1 with similar APX activity did not play a crucial role in growth and development (Figs. 1, 2a). Although growth of all the lines tested were influenced to a different degree by salt, transgenic plants over-expressing OsAPXb showed higher tolerance to NaCl than those over-expressing OsAPXa, as indicated by root length and total chlorophyll content (Fig. 2). These results suggest that both leaves and roots were damaged by the salt, and that OsAPXb is more effective in buffering such injury in Arabidopsis. A similar improvement in salt stress tolerance was also observed in transgenic tomato over-expressing pea cAPX and transgenic tobacco over-expressing the Arabidopsis cAPX gene (Badawi et al. 2004; Wang et al. 2005). It has also been reported that, under normal growth conditions, very low expression of OsAPx2 (OsAPXb), which encodes a cytosolic isoform in rice, could be highly induced by NaCl treatment, while the accumulation of the rice cytosolic OsAPx2 transcript was more obvious than that of OsAPx1 (OsAPXa) in the presence of salt (Teixeira et al. 2006). This phenomenon suggests that OsAPXb is more responsive to salt than the OsAPXa gene.
The ability of higher plants to scavenge toxic active oxygen seems to be a very important determinant of their tolerance to environmental stress. In active oxygen-scavenging systems, superoxide radicals generated in plant cells are converted to H2O2 by the action of SOD. The accumulation of H2O2 is prevented by either CAT or the ascorbate–glutathione cycle, in which APX and GR reduce it to H2O (Foyer et al. 1994). To obtain more information on the different functions of OsAPXa and OsAPXb, and their relationship with antioxidative systems other than APX activity, we also expanded our analysis to include a comprehensive study of the activity profiles of CAT, SOD and GR as well as examining changes in H2O2 levels in transgenic Arabidopsis under salt stress.
Under normal conditions, higher APX activity was observed in both transgenic plants with over-expression of OsAPXa and OsAPXb, and moreover, this higher activity did not influence the enzyme activities of CAT, SOD and GR (Figs. 3a, 4c, 5a–c). When subjected to salt stress, a rapid decrease in CAT activity was observed in these plants, suggesting that the salt was adversely affecting growth (Fig. 5a). At the same time, the changes in APX, SOD and GR activities in all the tested lines implied that all these enzymes participate in the counteraction of such salt stress, but to differing degrees (Figs. 3a, 4c, 5a–c). When treated with 50 mM NaCl, APX activity increased in all the tested lines, and transgenic plants over-expressing OsAPXb exhibited a much higher APX activity than those over-expressing OsAPXa (Fig. 3). From this, it can be deduced that the increase in APX activity in the transgenic plants was the result of OsAPXa and OsAPXb activity, as confirmed by northern and western blot analysis (Fig. 3). Although APX activity declined under high concentrations of NaCl in all the plant lines tested, over-expression of OsAPXb resulted in maintenance of comparatively high APX activity in the transgenic plants. In the course of salt treatment, changes in CAT, SOD and GR activity in transgenic plants over-expressing OsAPXb were also smaller than those seen in plants over-expressing OsAPXa, regardless of whether an increase or decline was observed (Fig. 5a–c).
The increase in H2O2 levels further confirmed that salt stress produced excessive H2O2, influencing the growth of transgenic and wild-type Arabidopsis. However, there was almost no increase in H2O2 content in transgenic plants over-expressing OsAPXb (Fig. 5d). A concentration of 200 mM NaCl, decreased the activities of APX, CAT, SOD and GR to a minimum, and all lines were severely damaged by a high level of H2O2. When exposed to salt stress, many antioxidant enzyme activities are known to increase or decline in plants to at least some extent (Lee et al. 2001; Orendi et al. 2001; Tsai et al. 2005). In the present study, increased activity of OsAPXa and OsAPXb, which also scavenge H2O2, partly compensated for the increased or decreased activity of SOD, CAT and GR in transgenic lines under salt stress. Severe deactivation of CAT during salt stress may be due to the prevention of new enzyme synthesis or CAT photoinactivation (Feierabend and Dehne 1996; Polle 1997). A similar decline in CAT activity has been reported in rice subjected to salt stress (Lee et al. 2001). At the same time, higher APX activity resulted in enhanced tolerance to salt stress, and OsAPXb played a more important role than OsAPXa in this process.
Interestingly, during salt stress, transcripts of both OsAPXa and OsAPXb increased in the transgenic Arabidopsis lines examined (Figs. 3, 4). Although the 35S promoter was considered to drive genes constitutively, post-transcriptional control caused by salt stress seems to occur in these two cAPXs. It has been reported that cAPX (Apx1) expression in pea is regulated at the post-transcriptional level during recovery from drought stress as indicated by transcript, protein and activity analysis (Mittler and Zilinskas 1994). Post-transcriptional stabilization of SOS1 gene has also been observed in transgenic Arabidopsis over-expressing this Na+/H+ antiporter gene driven by 35S promoter when treated with different concentrations of NaCl (Shi et al. 2002). The present results suggest that stress regulation of rice OsAPXa and OsAPXb is complex, and thus, the many post-transcriptional control processes need to be studied more comprehensively.
It can be concluded that, under normal conditions, salt stress seriously damages the growth of wild-type Arabidopsis; however, the expression of the OsAPXa and OsAPXb transgenes enhances salt tolerance, especially in transgenic plants over-expressing OsAPXb. The results suggest that over-expression of OsAPXb in Arabidopsis plays more pivotal role in preventing the over-accumulation of ROS and protecting cells against ROS caused by salt stress, thereby enhancing salt tolerance in transgenic plants. This study will help to explain the differential and essential roles of rice cAPXs in the adaptive responses of plant cells to environmental stresses, while suggesting that OsAPXb is more functional than OsAPXa in transgenic Arabidopsis exposed to salinity stress.
Abbreviations
- APX:
-
Ascorbate peroxidase
- cAPX:
-
Cytosolic ascorbate peroxidase
- CAT:
-
Catalase
- GR:
-
Glutathione reductase
- ROS:
-
Reactive oxygen species
- SOD:
-
Superoxide dismutase
References
Abei H (1974) Catalases. In: Bergmeyer HU (eds) Methods of enzymatic analysis. vol. 2, Academic, New York, pp 673–684
Arnon DI (1949) Copper enzymes in isolated chloroplasts in Beta vulgaris. Plant Physiol 24:1–15
Badawi GH, Kawano N, Yamauchi Y, Shimada E, Sasaki R, Kubo A, Tanaka K (2004) Over-expression of ascorbate peroxidase in tobacco chloroplasts enhances the tolerance to salt stress and water deficit. Physiol Plant 121:231–238
Bernt E, Bergmeyer HU (1974) Inorganic peroxides. In: Bergmeyer HU (ed) Methods of enzymatic analysis, vol. 4, Academic, New York, pp 2246–2248
Bradford M (1976) A rapid and sensitive method for the quantification of microgram quantities of proteins utilizing the principal of protein–dye binding. Anal Biochem 72:248–254
Chen GX, Asada K (1989) Ascorbate peroxidase in tea leaves: occurrence of two isozymes and the differences in their enzymatic and molecular properties. Plant Cell Physiol 30:987–998
Davletova S, Rizhsky L, Liang H, Shengqiang Z, Oliver DJ, Coutu J, Shulaev V, Schlauch K, Mittler R (2005) Cytosolic ascorbate peroxidase 1 is a central component of the reactive oxygen gene network of Arabidopsis. Plant Cell 17:268–281
Feierabend J, Dehne S (1996) Fate of the porphyrin cofactors during the light-dependent turnover of catalase and of the photosystem II reaction-center protein D1 in mature rye leaves. Planta 198:413–422
Fourcroy P, Vansuyt G, Kushnir S, Inzé D, Briat J (2004) Iron-regulated expression of a cytosolic ascorbate peroxidase encoded by the APX1 gene in Arabidopsis seedlings. Plant Physiol 134:605–613
Foyer CH, Lelandais M, Kenert KJ (1994) Photooxidative stress in plants. Physiol Plant 92:696–717
Giannopolitis CN, Ries SK (1977) Superoxide dismutases. I. Occurrence in higher plants. Plant Physiol 59:309–314
Harlow E, Lane D (1988) Antibodies: a laboratory manual. Cold Spring Harbor Lab Press, Plainview
Ishikawa T, Madhusudhan R, Shigeoka S (2003) Effects of iron on the expression of ascorbate peroxidase in Euglena gracilis. Plant Sci 165:1363–1367
Lee DH, Lee CB (2000) Chilling stress-induced changes of antioxidant enzymes in the leaves of cucumber: in gel enzyme activity assay. Plant Sci 159:75–85
Lee DH, Kim YS, Lee CB (2001) The inductive responses of the antioxidant enzymes by salt stress in the rice (Oryza sativa L.). J Plant Physiol 158:737–745
Lu ZQ, Takano T, Liu SK (2005) Purification and characterization of two ascorbate peroxidases of rice (Oryza sativa L.) expressed in Escherichia coli. Biotechnol Lett 27:63–67
Menezes-Benavente L, Teixeira FK, Kamei CLA, Margis-Pinheiro M (2004) Salt stress induces altered expression of genes encoding antioxidant enzymes in seedlings of a Brazilian indica rice (Oryza sativa L.). Plant Sci 166:323–331
Mittler R, Zilinskas BA (1994) Regulation of pea cytosolic ascorbate peroxidase and other antioxidant enzymes during the progression of drought stress and following recovery from drought. Plant J 5(3):397–405
Orendi G, Zimmermann P, Baar C, Zentgraf U (2001) Loss of stress-induced expression of catalases 3 during leaf senescence in Arabidopsis thaliana is restricted to oxidative stress. Plant Sci 161:301–314
Panchuk II, Volkov RA, Schöffl F (2002) Heat stress- and heat shock transcription factor-dependent expression and activity of ascorbate peroxidase in Arabidopsis. Plant Physiol 129:838–853
Panchuk II, Zentgraf U, Volkov RA (2005) Expression of the Apx gene family during leaf senescence of Arabidopsis thaliana. Planta 222:926–932
Polle A (1997) Defense against photooxidative damage in plants. In: Scandalios J (ed) Oxidative stress and molecular biology of antioxidant defenses. Cold Spring Harbor Laboratory Press. Woodbury, pp 785–813
Rao MV, Paliyath G, Ormrod DP (1996) Ultraviolet-B- and ozone-induced biochemical changes in antioxidant enzymes of Arabidopsis thaliana. Plant Physiol 110:125–136
Sano S, Ueda M, Kitajima S, Takeda T, Shigeoka S, Kurano N, Miyachi S, Miyake C, Yokota A (2001) Characterization of ascorbate peroxidases from unicellular red alga Galdieria partita. Plant Cell Physiol 42:433–440
Sharma P, Dubey RS (2005) Drought induces oxidative stress and enhances the activities of antioxidant enzymes in growing rice seedlings. Plant Growth Regul 46:209–211
Shigeoka S, Ishikawa T, Tamoi M, Miyagawa Y, Takeda T, Yabuta Y, Yoshimura K (2002) Regulation and function of ascorbate peroxidase isoenzymes. J Exp Bot 53:1305–1319
Shi H, Lee B, Wu SJ, Zhu JK (2002) Overexpression of a plasma membrane Na+/H+ antiporter gene improves salt tolerance in Arabidopsis thaliana. Nat Biotechnol 21:81–85
Teixeira FK, Menezes-Benavente L, Galvão VC, Margis R, Margis-Pinheiro M (2006) Rice ascorbate peroxidase gene family encodes functionally diverse isoforms localized in different subcellular compartments. Planta 224:300–314
Tsai YC, Hong CY, Liu LF, Kao CH (2005) Expression of ascorbate peroxidase and glutathione reductase in roots of rice seedlings in response to NaCl and H2O2. J Plant Physiol 162:291–299
Wang J, Zhang H, Allen RD (1999) Overexpression of an Arabidopsis peroxisomal ascorbate peroxidase gene in tobacco increases protection against oxidative stress. Plant Cell Physiol 40:725–732
Wang WX, Vinocur B, Altman A (2003) A plant responses to drought, salinity and extreme temperatures: towards genetic engineering for stress tolerance. Planta 218:1–14
Wang Y, Wisniewski M, Meilan R, Cui M, Webb R, Fuchigami L (2005) Overexpression of cytosolic ascorbate peroxidase in tomato confers tolerance to chilling and salt stress. J Am Soc Hortic Sci 130:167–173
Wehmeyer N, Hernandez LD, Finkelstein RR, Vierling E (1996) Synthesis of small heat-shock proteins is part of the developmental program of late seed maturation. Plant Physiol 112:747–757
Zhu JK (2001) Plant salt tolerance. Trends Plant Sci 6:66–71
Acknowledgments
This work was supported by the Excellent Teachers Program Foundation of Helongjiang University and the National High Technology Research and Development Program (863 Program) from the People’s Republic of China.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by W.-H. Wu.
Zhenqiang Lu and Dali Liu contributed equally.
Rights and permissions
About this article
Cite this article
Lu, Z., Liu, D. & Liu, S. Two rice cytosolic ascorbate peroxidases differentially improve salt tolerance in transgenic Arabidopsis . Plant Cell Rep 26, 1909–1917 (2007). https://doi.org/10.1007/s00299-007-0395-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00299-007-0395-7