Abstract
Published data on the association between tumor necrosis factor-alpha (TNF-α) promoter-308 A/G polymorphism and systemic lupus erythematosus (SLE) risk are inconclusive. To derive a more precise estimation of the relationship, a meta-analysis was performed. A total of 28 studies including 2,992 cases and 4,326 controls (5,924 cases and 8,484 controls in A versus G comparison) were involved in this meta-analysis. Meta-analysis was performed for genotypes A/A (recessive effect), A/A+A/G (dominant effect), and A allele in fixed or random effects models. In addition, we also performed a “model-free” analysis by considering the G/G genotype as the reference and estimated the OR for the A/A versus G/G and A/G versus G/G genotype. Overall, an association of TNF-α promoter-308 A/G polymorphism with SLE was found (A versus G: OR = 1.686, 95% CI = 1.400–2.032, P < 0.001; A/A versus A/G+G/G: OR = 3.043, 95% CI = 2.185–4.238, P < 0.001; A/A+A/G versus G/G: OR = 1.822, 95% CI = 1.379–2.407, P < 0.001; A/A versus G/G: OR = 3.686, 95% CI = 2.628–5.172, P < 0.001; A/G versus G/G: OR = 1.691, 95% CI = 1.291–2.215, P < 0.001). However, stratification by ethnicity indicated that the risk A allele was not associated with SLE in Asian (A versus G: OR = 1.207, 95% CI = 0.856–1.702, P = 0.283) and African population (A versus G: OR = 1.225, 95% CI = 0.597–2.516, P = 0.580). In summary, this meta-analysis indicated that TNF-α promoter-308-A/G polymorphism is associated with susceptibility to SLE.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Systemic lupus erythematosus (SLE) is a chronic systemic autoimmune disease, characterized by the production of multiple autoantibodies, complement activation, and immune-complex deposition, resulting in tissue and organ damage. The etiology and pathogenetic mechanisms of SLE are still unclear [1].
Many pro-inflammatory cytokines have been implicated in the pathogenesis of SLE. Tumor necrosis factor-alpha (TNF-α) is a potential pro-inflammatory cytokine that plays an important role in inflammatory and immune responses in SLE [2]. TNF-α stimulates cytokine production, enhancing expression of adhesion molecules and neutrophil activation and acts as a costimulator for T cell activation and antibody production.
TNF-α gene is located on chromosome 6, within the class III region of MHC [3]. Many polymorphisms of the TNF-α gene have been identified. Recently, these polymorphisms have attracted widespread attention, especially the TNF-α promoter-308 A/G polymorphism. A number of case–control studies have been conducted to investigate the association of this polymorphism with autoimmune diseases including systemic lupus erythematosus. However, many studies have shown inconclusive or contradictory results. This inconsistency may be due to studies with inadequate statistical power, racial and ethnic differences, publication bias, or uncorrected multiple hypothesis testing. Although it is unclear whether the TNF-α promoter-308 A/G polymorphism has a functional significance, several evidences suggest that there may be a small but significant effect of the TNF-α promoter-308 A/G polymorphism, with the A allele being associated with slightly greater levels of TNF-α transcription [4].
Meta-analysis is a means of increasing the effective sample size under investigation through the pooling of data from individual association studies, thus enhancing the statistical power of the analysis for the estimation of genetic effects [5]. Using meta-analysis, Lee et al. has demonstrated that the TNF-α promoter-308 A/G polymorphism may confer susceptibility to SLE, especially in European-derived population. In this study, we present an update to Lee’s meta-analysis with recently published studies, to further investigate whether the TNF-α promoter-308 A/G polymorphism is a risk factor to the SLE susceptibility.
Methods
Identification of eligible studies and data extraction
We performed an exhaustive search on studies that examined the association of the TNF-α promoter-308 A/G polymorphism with SLE. A search of the literature was made using MEDLINE and PubMed to identify available articles in which the TNF-α promoter-308 A/G polymorphism was determined in patients with SLE and control (most recent one was Nov 2009). References in the studies were reviewed to identify additional studies not indexed by MEDLINE or PubMed. “Tumor necrosis factor”, “TNF-α”, “polymorphism”, “systemic lupus erythematosus” and “SLE” were entered as both medical subject heading (MeSH) terms and text words. No language restrictions were applied. A study was included in the analysis if (1) it was published up to Nov 2009, (2) it was original data (independence among studies), and (3) it provided enough data to calculate odds ratio (OR). When a study reported the results on different subpopulation, we treated them as a separate study in the meta-analysis. We excluded the following: (1) studies that contained overlapping data, (2) studies in which the number of null and wild genotypes could not be ascertained, and (3) studies in which family members had been studied because their analysis is based on linkage considerations. From each study, we extracted the available genotype and allele frequency information from the TNF-α promoter-308 A/G polymorphism.
The literature search identified 81 potentially relevant publications. Among them, 53 studies were excluded because they did not meet the inclusion criteria. They were studies on other TNF polymorphism such as TNF receptor genes (24 publications), studies on other diseases (22 publications) or animal studies (5 publications), or family study (1 publication) or repeat study (1 publications). There were no studies in which the number of null and wild genotypes could not be ascertained. A total of 28 relevant studies with TNF-α promoter-308 A/G polymorphism and SLE were selected for the meta-analysis [6–33]. Among them, two of the eligible studies contained data on two different ethnic groups [10, 14], and we treated them independently. Therefore, a total of 30 separate comparisons were considered in our meta-analysis. These 30 studies consisted of 14 European, 9 Asian, 3 African, 2 South American, and 2 Mexican population samples (Table 1). Although the allele frequency of the TNF-α promoter-308 A/G polymorphism was extracted from 30 studies, the genotype frequency was available from 19 studies. Therefore, the meta-analysis was performed with 19 studies overall and 14 (9 European and 5 Asian) when divided by ethnical origin for genotype-based analysis. We have performed group-specific meta-analysis in European, Asian, South American, Mexican, and African-derived populations. As the genotype data on TNF-α-308 A/G polymorphism was available for one of three studies in African population [10], meta-analysis was performed only on A allele of TNF-α promoter-308 in African population.
Evaluation of publication bias
An estimate of potential publication bias was carried out by the funnel plot, in which the standard error of log (OR) of each study was plotted against its log (OR). An asymmetric plot suggests a possible publication bias. Funnel plot asymmetry was assessed by the method of Egger’s linear regression test, a linear regression approach to measure funnel plot asymmetry on the natural logarithm scale of the OR. The significance of the intercept was determined by the t test suggested by Egger (P < 0.05 was considered representative of statistically significant publication bias).
Evaluation of the statistical association
Allele frequencies at the TNF-α promoter-308 A/G polymorphism from the respective study were determined by the allele counting method. A chi-square test was used to determine whether the observed genotype frequencies in the controls conformed to Hardy–Weinberg (H–W) expectations. We examined the contrast of the allelic effect of A (minor allele) versus G (common allele) and also examined the contrast of A/A versus A/G+G/G genotypes as well as the contrast of A/A+A/G versus G/G genotypes. These contrasts correspond to the recessive and dominant effects of the A allele, respectively. The point estimates of the risk, the OR, and its 95% confidence interval (CI) were estimated for each study. We assessed the within- and between-study variation or heterogeneity by testing Cochran’s Q statistic [34]. This heterogeneity test assessed the null hypothesis that all studies were evaluating the same effect. A significant Q statistic (P < 0.10) indicated heterogeneity across studies, and then the random effect model was used for meta-analysis. Otherwise, the fixed effect model was used. Fixed effect model assumes that all of the studies are estimating the same underlying effect and considers only within-study variation. We also quantified the effect of heterogeneity using a recently developed measure, I 2 = 100% × (Q − df)/Q [35]. The I statistic measures the degree of inconsistency in the studies by calculating what percentage of the total variation across studies is due to heterogeneity rather than by chance [36]. Finally, the overall or pooled estimate of risk (OR) was obtained by a random effects (DerSimonian–Laird) or a fixed effects model (Mantel–Haenszel) in the presence (P ≤ 0.1 or I 2 > 50%) or absence (P > 0.1 and I 2 ≤ 50%) of heterogeneity, respectively. Pooled OR in the meta-analysis was performed weighting individual OR by the inverse of their variance. Statistical manipulations for the meta-analysis were conducted by STATA version 8.0 (Stata Corporation, College Station, TX). The power of each study was computed as the probability of detecting an association between the TNF-α-308 A/G SNP and SLE at the 0.05 level of significance, assuming an OR of 1.5 (small effect size). The power analysis was performed using the statistical program G*Power. (http://www.psycho.uni-duesseldorf.de/aap/projects/gpower).
Results
Studies included in the meta-analysis
Selected characteristics of 28 case–control studies for TNF-α-308 A/G polymorphism and the risk of SLE are summarized in Table 1. Also, Table 1 shows the expected power of each individual study to demonstrate an association between this polymorphism and SLE. The statistical power of each study ranged from 10.1 to 77.0%. Interestingly, none of the 28 individual studies had more than 80% statistical power to an effect (Table 1). We also calculate the distribution of genotype for H–W equilibrium in control group.
Evaluation of study quality
The distribution of the genotype in the control group of each study (if the genotype data are available in a study) was consistent with H–W equilibrium. Deviation from H–W equilibrium among controls could imply some potential biases in the selection of control or genotyping errors, but excluding the study with the absence of H–W equilibrium in controls did not materially affect the overall results [37]. So, presentation of the meta-analysis is based on a combined data by adding the data which we can only get the alleles distribution.
Evaluation of A/G polymorphism and SLE association
The summary of meta-analysis for the TNF-α-308 A/G polymorphism with SLE is shown (Table 2). The Q test of heterogeneity was almost always significant, and we conducted analyses using random effect models except in one case that was in a subgroup analysis, and the Q test was likely to have been not statistically significant owing to lack of power. So, we also performed the subgroup analysis using random effect model.
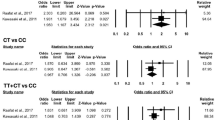
An association between SLE and A/A risk genotype (assuming A allele as recessive allele) was found in the overall population (OR = 3.043, 95% CI = 2.185–4.238, P < 0.001) (Table 2; Fig. 1). However, stratification by ethnicity indicates that the A/A genotype is significantly associated with SLE only in Europeans (OR = 3.720, 95% CI = 2.520–5.491, P < 0.001). Conversely, there was no association detected for the A/A genotype with SLE patients from other population. Genotype data on TNF-α-308 A/G polymorphism were available for one of three studies in African population; therefore, genotype-specific meta-analysis was not performed in African samples. Assuming A allele as dominant allele, the overall OR for the combined A/A+A/G genotypes was 1.822 (95% CI = 1.379–2.407, P < 0.001) (Table 2; Fig. 2). Similarly, using ethnic-specific analysis, OR was increased significantly in the European samples (OR = 2.418, 95% CI = 1.733–3.375, P < 0.001), South American (OR = 2.837, 95% CI = 1.898–4.243, P < 0.001), and Mexican (OR = 2.362, 95% CI = 1.440–3.873, P = 0.001), but not in Asians (OR = 1.021, 95% CI = 0.684–1.522, P = 0.920). The overall OR for the A allele of the TNF-α-308 A/G was 1.686 (95% CI = 1.400–2.032, P < 0.001) (Table 2; Fig. 3). Stratification by ethnicity indicates that the A allele is a risk factor for SLE in European (OR = 1.970, 95% CI = 1.547–2.508, P < 0.001), South American (OR = 2.452, 95% CI = 1.768–3.399, P < 0.001), Mexican (OR = 2.505, 95% CI = 1.548–4.056, P < 0.001), but not in Asian (OR = 1.207, 95% CI = 0.856–1.702, P = 0.283) or African (OR = 1.225, 95% CI = 0.597–2.516, P = 0.580). We also performed a “model-free” analysis by considering the G/G genotype as the reference and estimated the OR for the A/A versus G/G and A/G versus G/G genotype (Table 2; Figs. 4, 5). We found gene dosage effect of the A allele.
Overall, the meta-analysis of the TNF-α-308 A/G genotype (recessive effect), A/A+A/G genotype (dominant effect), and the risk allele A was associated with susceptibility to SLE in Europeans. However, in the subgroup analysis by ethnicity, there were differences in different populations.
Publication bias
Begg’s funnel plot and Egger’s test were performed to access the publication bias of literatures. The shapes of the funnel plots did not reveal any evidence of obvious asymmetry (The figures are not shown). Then, the Egger’s test was used to provide statistical evidence of funnel plot symmetry. The results still did not demonstrate any evidence of publication bias (Table 3).
Discussion
It is well recognized that there is individual susceptibility to SLE even with the same environmental exposure. Host factors, including polymorphisms of genes involved in SLE may have accounted for this difference. Therefore, genetic susceptibility to SLE has been a research focus in scientific community. Recently, genetic variants of the TNF-α gene in the etiology of several autoimmune diseases have drawn increasing attention. Growing number of studies have suggested that -308 A in the promoter region of the TNF-α gene was emerging as a susceptibility allele for SLE. However, the results are inconclusive. To better understanding of the association between this polymorphism and SLE risk, a pooled analysis with a large sample, subgroup analysis performed, and heterogeneity explored is needed.
Overall, our results indicated that TNF-α promoter-308-A/G polymorphism is associated with susceptibility to SLE. These results were similar to that observed by Lee et al. in another previous meta-analysis [38]. However, in the subgroup analysis by ethnicity, there were differences among different populations, suggesting a possible role of ethnic differences in genetic backgrounds and the environment they lived in. In addition, the influence of the TNF-α-308 A allele might be masked by the presence of other as-yet unidentified causal genes involved in SLE development.
Some limitations of this meta-analysis should be acknowledged. Firstly, the controls were not uniformly defined. Although most of the controls were selected mainly from healthy populations, some had benign disease. Therefore, non-differential misclassification bias was possible because these studies may have included the control groups who have different risks of developing SLE. Secondly, in the subgroup analyses, the number of Africans was relatively small, not having enough statistical power to explore the real association. In spite of these limitations, our meta-analysis also had some advantages. First, substantial number of cases and controls were pooled from different studies, which significantly increased the statistical power of the analysis. Second, no publication bias were detected, indicating that the whole pooled results may be unbiased.
In summary, this meta-analysis suggests that the TNF-α promoter-308-A/G polymorphism is associated with SLE susceptibility. However, large sample studies including different ethnic groups with a careful matching between cases and controls should be considered in future association studies to confirm the results from our meta-analysis. Also, further evaluating the effect of gene–gene and gene-environment interactions on the TNF-α promoter-308-A/G polymorphism and SLE risk is necessary.
References
Criswell LA, Pfeiffer KA, Lum RF et al (2005) Analysis of families in the multiple autoimmune disease genetics consortium (MADGC) collection: the PTPN22 620 W allele associates with multiple autoimmune phenotypes. Am J Hum Genet 76:561–571
Vassalli P (1992) The pathophysiology of tumor necrosis factors. Annu Rev Immunol 10:411–452
Dunham I, Sargent CA, Trowsdale J et al (1987) Molecular mapping of the human major histocompatibility complex by pulsed-field gel electrophoresis. Proc Natl Acad Sci USA 84:7237–7241
Allen RD (1999) Polymorphism of the human TNF-a promoter—random variation or functional diversity? Mol Immunol 36:1017–1027
Egger M, Davey SG, Schneider M et al (1997) Bias in meta-analysis detected by a simple, graphical test. Br Med J 315:629–634
McHugh NJ, Owen P, Cox B et al (2006) MHC class II, tumour necrosis factor a, and lymphotoxin a gene haplotype associations with serological subsets of systemic lupus erythematosus. Ann Rheum Dis 65:488–494
Hrycek A, Siekiera U, Cieślik P et al (2005) HLA-DRB1 and -DQB1 alleles and gene polymorphisms of selected cytokines in systemic lupus erythematosus. Rheumatol Int 26:1–6
Suárez A, López P, Mozo L et al (2005) Differential effect of IL10 and TNFa genotypes on determining susceptibility to discoid and systemic lupus erythematosus. Ann Rheum Dis 64:1605–1610
Schotte H, Willeke P, Tidow N et al (2005) Extended haplotype analysis reveals an association of TNF polymorphisms with susceptibility to systemic lupus erythematosus beyond HLA-DR3. Scand J Rheumatol 34:114–121
Parks CG, Pandey JP, Dooley MA et al (2004) Genetic polymorphisms in tumor necrosis factor (TNF)-alpha and TNF-beta in a population-based study of systemic lupus erythematosus: associations and interaction with the interleukin-1alpha-889 C/T polymorphism. Hum Immunol 65:622–631
May LA, Huang Q, Morris D et al (2002) Relationship of tumour necrosis factor alpha gene polymorphisms and neuropsychiatric lupus. Lupus 11:114–118
van der Linden MW, van der Slik AR, Zanelli E et al (2001) Six microsatellite markers on the short arm of chromosome 6 in relation to HLA-DR3 and TNF-308A in systemic lupus erythematosus. Genes Immun 2:373–380
Rood MJ, van Krugten MV, Zanelli E et al (2000) TNF-308A and HLA-DR3 alleles contribute independently to susceptibility to systemic lupus erythematosus. Arthritis Rheum 43:129–134
Rudwaleit M, Tikly M, Khamashta M et al (1996) Interethnic differences in the association of tumor necrosis factor promoter polymorphisms with systemic lupus erythematosus. J Rheumatol 23:1725–1728
D’Alfonso S, Colombo G, Della BS et al (1996) Association between polymorphisms in the TNF region and systemic lupus erythematosus in the Italian population. Tissue Antigens 47:551–555
Danis VA, Millington M, Hyland V et al (1995) Increased frequency of the uncommon allele of a tumour necrosis factor alpha gene polymorphism in rheumatoid arthritis and systemic lupus erythematosus. Dis Markers 12:127–133
Wilson AG, Gordon C, di Giovine FS et al (1994) A genetic association between systemic lupus erythematosus and tumor necrosis factor alpha. Eur J Immunol 24:191–195
Goldstein R, Sengar DP (1993) Comparative studies of the major histocompatibility complex in French Canadian and non-French Canadian Caucasians with systemic lupus erythematosus. Arthritis Rheum 36:1121–1127
Fugger L, Morling N, Ryder LP et al (1989) NcoI restriction fragment length polymorphism (RFLP) of the tumor necrosis factor (TNF alpha) region in four autoimmune diseases. Tissue Antigens 34:17–22
Lin YJ, Chen RH, Wan L et al (2009) Association of TNF-α gene polymorphisms with systemic lupus erythematosus in Taiwanese patients. Lupus 18:974–979
Hirankarn N, Avihingsanon Y, Wongpiyabovorn J (2007) Genetic susceptibility to SLE is associated with TNF-alpha gene polymorphism -863, but not -308 and -238, in Thai population. Int J Immunogenet 34:425–430
Azizah MR, Kuak SH, Ainol SS et al (2004) Association of the tumor necrosis factor alpha gene polymorphism with susceptibility and clinical-immunological findings of systemic lupus erythematosus. Asian Pac J Allergy Immunol 22:159–163
Wang M, Dong Y, Huang S (1999) Study on the association between tumor necrosis factor alpha gene polymorphism and systemic lupus erythematosus. Zhonghua Nei Ke Za Zhi 38:393–396
Wang Y, Zhang Y, Zhu S (1998) The association of susceptibility of SLE and the gene polymorphism of TNF. Zhonghua Yi Xue Za Zhi 78:111–114
Chen CJ, Yen JH, Tsai WC et al (1997) The TNF2 allele does not contribute towards susceptibility to systemic lupus erythematosus. Immunol Lett 55:1–3
Fong KY, Howe HS, Tin SK et al (1996) Polymorphism of the regulatory region of tumour necrosis factor alpha gene in patients with systemic lupus erythematosus. Ann Acad Med Singapore 25:90–93
Tomita Y, Hashimoto S, Yamagami K et al (1993) Restriction fragment length polymorphism (RFLP) analysis in the TNF genes of patients with systemic lupus erythematosus (SLE). Clin Exp Rheumatol 11:533–536
Atsumi T (1992) Tumor necrosis factor alpha in systemic lupus erythematosus: evaluation by restriction fragment length polymorphism and production by peripheral blood mononuclear cells. Hokkaido Igaku Zasshi 67:408–419
Sullivan KE, Wooten C, Schmeckpeper BJ et al (1997) A promoter polymorphism of tumor necrosis factor alpha associated with systemic lupus erythematosus in African–Americans. Arthritis Rheum 40:2207–2211
Guarnizo-Zuccardi P, Lopez Y, Giraldo M et al (2007) Cytokine gene polymorphisms in Colombian patients with systemic lupus erythematosus. Tissue Antigens 70:376–382
Correa P, Gomez ML, Cadena J et al (2005) Autoimmunity and tuberculosis. Opposite association with TNF polymorphism. J Rheumatol 32:219–224
Jiménez-Morales S, Velázquez-Cruz R, Ramírez-Bello J et al (2009) Tumor necrosis factor-alpha is a common genetic risk factor for asthma, juvenile rheumatoid arthritis, and systemic lupus erythematosus in a Mexican pediatric population. Hum Immunol 70:251–256
Zuniga J, Vargas-Alarcon G, Hernandez-Pacheco G et al (2001) Tumor necrosis factoralpha promoter polymorphisms in Mexican patients with systemic lupus erythematosus (SLE). Genes Immun 2:363–366
Cochran WG (1954) The combination of estimates from different experiments. Biometrics 10:101–129
Higgins JP, Thompson SG (2002) Quantifying heterogeneity in a metaanalysis. Stat Med 21:1539–1558
Higgins JP, Thompson SG, Deeks JJ et al (2003) Measuring inconsistency in meta-analyses. BMJ 327:557–560
Lee YH, Rho YH, Choi SJ et al (2007) The PTPN22 C1858T functional polymorphism and autoimmune diseases—a meta-analysis. Rheumatology 46:49–56
Lee YH, Harley JB, Nath SK (2006) Meta-analysis of TNF-alpha promoter-308 A/G polymorphism and SLE susceptibility. Eur J Hum Genet 14:364–371
Acknowledgments
This work was supported by grants from the key program of National Natural Science Foundation of China (30830089, 30771848) and the Specialized Research Fund for the Doctoral Program of Higher Education of China (20070366002).
Author information
Authors and Affiliations
Corresponding author
Additional information
Hai-Feng Pan, and Rui-Xue Leng contributed equally to this work and should be considered co-first authors.
Rights and permissions
About this article
Cite this article
Pan, HF., Leng, RX., Wang, C. et al. Association of TNF-α promoter-308 A/G polymorphism with susceptibility to systemic lupus erythematosus: a meta-analysis. Rheumatol Int 32, 2083–2092 (2012). https://doi.org/10.1007/s00296-011-1924-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00296-011-1924-9