Abstract
In yeast, the GCR1 transcription factor is involved in the regulation of glycolysis and its deletion exhibited growth defect, reduced inositol and phosphatidylinositol (PI) levels compared to WT cells. We observed a down regulation of the INO1 and PIS1 expression in gcr1∆ cells under both I− and I+ conditions and the over expression of GCR1 in gcr1∆ cells restored the growth, retrieved the expression of INO1, and PIS1 comparable to WT cells. In the gel shift assay, the Gcr1p binds to its consensus sequence CTTCC in PIS1 promoter and regulates its expression but not in INO1 transcription. The WT cells, under I− significantly reduced the expression of GCR1 and PIS1, but increased the expression of KCS1 and de-repressed INO1. The Kcs1p expression was reduced in gcr1∆ cells; this reduced INO1 expression resulting in abnormal vacuolar structure and reduced autophagy in Saccharomyces cerevisiae.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Yeast has been widely used as a model system to study molecular biology and proliferative response to nutrients. Glycolysis regulation 1 (GCR1) is a transcription factor that tightly controls the expression of glycolytic genes (Willis et al. 2003; Clifton and Fraenkel 1981; Tornow et al. 1993; Barbara et al. 2006; Baker 1986) and serves as a regulator of RNA polymerase II transcription, while being involved in the coordinated regulation of cell cycle progression (Baker 1991; Willis et al. 2003). The Pho85 (cyclin-dependent kinase, CDK) is also involved in cell cycle regulation, responds to nutrient levels and is involved in GCR1 regulation (Lenburg and O’Shea 2001). The loss of PHO85 changes in many cellular events including the repression of INO1 transcription, leading to inositol auxotrophy (Kliewe et al. 2017), and also affects GCR1 transcription (Turkel et al. 2003). The deletion of GCR1 does not affect the glucose 6-phosphate level, but accumulates trehalose and glycogen (Hossain et al. 2016; Seker and Hamamci 2003). The glycolytic intermediates dihydroxyacetone 1-phosphate (DHAP) and glyceraldehyde 3-phosphate (G3P) inhibit inositol 3-phosphate synthase activity in yeast and human (Shi et al. 2005). The inositol 3-P synthase (INO1), is a rate-limiting enzyme that converts glucose 6-P to inositol 3-P which is further dephosphorylated to inositol by inositol monophosphatase (INM1) (Carman and Han 2011; Murray and Greenberg 2000). The glucose 6-phosphate from glycolysis is involved in the de novo synthesis of inositol, but the role of GCR1 in inositol production is still uncertain.
In Saccharomyces cerevisiae, the inositol related molecules modulate diverse cell functions, such as phospholipid metabolism, anchoring of proteins in glycolipids (Carman and Han 2011), cell growth, membrane trafficking (Henry et al. 2012), and apoptosis (York et al. 2001). During inositol limitation (inositol auxotrophy) the Ino1p is highly expressed (Dubois et al. 2002), but the exogenous supplementation of inositol represses the INO1 mRNA expression through the Opi1p (repressor protein). When the phosphatidic acid (PA) level drops, Opi1 is translocated from the ER to nucleus that interacts with Ino2p–Ino4p complex (Murray and Greenberg 2000; Donahue and Henry 1981) and impairs the transcription of genes involved in phospholipid (PL) metabolism (Bachhawat et al. 1995). The phospholipids play an important role in cell size regulation, especially the increased PA levels modulate the cell size (Rao et al. 2017). Recent study shows that INO1 promoter possesses a cis-acting Memory Requirement Sequence (MRS), which is essential for the binding of transcription factor Sfl1p, and SFL1 (Heat shock factor-like DNA-binding domain) was reported to epigenetically control INO1 transcriptional memory, and loss of Sfl1p disrupted INO1 memory under inositol deprivation (D’Urso and Brickner 2017). The increased level of inositol up regulates the phosphatidylinositol synthase 1 (PIS1), and increases the PI levels required for progression of the cell cycle in yeast (Gardocki et al. 2005; Nikawa et al. 1987). The Zap1p transcription factor also controls PIS1 expression under zinc depletion conditions through UASZRE element in the promoter region (Carman and Han 2007; Han et al. 2005). Carbon source has an impact on PIS1 expression (Anderson and Lopes 1996), and the PIS1 promoter region contains a putative binding site for Gcr1p in UAS3 (Gardocki and Lopes 2003).
The inositol pyrophosphate synthase (Kcs1p) incorporates pyrophosphate to inositol hexakisphosphate (IP5) and inositol hexaphosphate (IP6) generating inositol pyrophosphates 5PP-IP4 and 5PP-IP5 (IP6 and IP7) in yeast, and regulates cellular processes like inositol metabolism (Ye et al. 2013), telomere length (Saiardi et al. 2005), vesicular trafficking (Saiardi et al. 2002), vacuolar biogenesis and stress response (Dubois et al. 2002). The phosphatidylinositol-3-kinase-related kinase (PIKK) is also the regulator of cell proliferation and genomic maintenance (Sugimoto 2018). The inositol pyrophosphate 5PP-IP4 (IP6) stabilizes the interaction of Ino2p–Ino4p complex and regulates INO1 transcription (Ye et al. 2013). The 5PP-IP5 (IP7) transfers its pyrophosphate to the serine residues (S515–S518) of Gcr1p, thereby making Gcr1p unable to interact with Gcr2p, thus down regulating the transcription of the glycolytic enzymes (Szijgyarto et al. 2011). The Gcr1p interacts with Gcr2p (GCR1 co activator) and forms a Gcr1p-Gcr2p complex that regulates the transcription of the glycolytic enzymes, which binds to the CTTCC binding motif in UAS sequence of the target glycolytic genes and positively regulates their expression (Willis et al. 2003). The removal of a Gcr1p binding site from the promoter region of Fatty acid synthase (FAS1 and FAS2) genes resulted in a dramatic reduction of FAS1 and FAS2-LacZ expression (Chirala 1992).
The absence of INO1 disturbs the vacuolar structure under inositol limitation (Deranieh et al. 2015). Similarly, anti-epileptic drug Valproate (VPA) inhibits inositol-3-P synthase, and reduces the inositol level (Deranieh et al. 2015). However, the VPA treatment displayed abnormal vacuolar arrangements, also reduced V-ATPase activity and proton pumping under inositol limitation (Deranieh et al. 2015). The vacuole is the counterpart of the mammalian lysosome in yeast, besides being the key cellular site of protein and organelle turnover, and is also involved in proteolysis during micro and macro autophagy process (Stauffer and Powers 2017; Bryant and Stevens 1998; Li and Kane 2009).
Autophagy is maintained at a basal level under normal cell growth conditions and is induced by starvation (Wang and Klionsky 2003). PKA and TOR are two nutrient-responsive signaling components which help to maintain cell growth and autophagy (Kamada et al. 2010). PKA and TOR signaling pathways are regulated by the Rap1-mediated gene response and is based on the nutritional prominence (Santangelo 2006). Indeed, loss of GCR1 reduces transcriptional activation of Rap1, likewise the exposure of rapamycin display no alteration in rapamycin sensitivity or rapamycin-induced down regulation of ribosomal protein gene expression (Menon et al. 2005; Lieb et al. 2001). The earlier reports suggest that the loss of KCS1 displayed abnormal vacuolar structure, membrane abnormalities (Saiardi et al. 2002), defective autophagy flux and reduced autophagosome formation (Taylor et al. 2012).
In the current study, we focused on the functional importance of GCR1 under inositol limitation and autophagy in Saccharomyces cerevisiae. We performed the growth analysis and investigated the gene expression of inositol biosynthesis. The gcr1∆ cells showed reduced intra-cellular inositol levels and the cells became inositol growth defective, which was alleviated by over expression of GCR1 in the gcr1∆ cells. We also show that the deletion of GCR1 decreased Kcs1p expression and our results suggest that GCR1 deletion has an impact on INO1 expression, vacuolar structure and autophagy process through the regulation of Kcs1p.
Materials and methods
Chemicals and media
Yeast extract, peptone, bacteriological agar, yeast nitrogenous bases (YNB) were purchased from Difco (Becton Dickinson and Company, Franklin Lakes, NJ, USA). YNB without inositol, LB media were obtained from HiMedia (Bengaluru, India). Thin-layer silica gel coated aluminum plate 60F254 and solvents were obtained from Merck (Bengaluru, India). Restriction endonuclease and T4 DNA ligase were procured from Fermentas. Glucose (GO) assay kit, ONPG (O-nitrophenyl-β-d-galactopyranoside), PMSF, glass beads, appropriate amino acid mixtures (drop out), anti-His antibody (Cat No: H1029), ALP conjugated secondary antibody (Cat No: A3562) and immunoblot developing substrate NBT/BCIP were from Sigma–Aldrich (Bengaluru, India). Anti-TAP antibody (Cat No: CAB1001), anti-GFP antibody (Cat No: A-11122), anti-PGK antibody (Cat No: PA5-28612), nitrocellulose membrane, FM4-64 dye, Phusion high-fidelity DNA polymerase, dNTPs mix, cDNA synthesis kit and Power SYBR Green PCR Master Mix were obtained from Invitrogen (Bengaluru, India). Myo-inositol assay kit was obtained from Megazyme (New Delhi, India).
Plasmids, strains and growth conditions
The plasmids and yeast deletion strains used in the study are listed in Table 1. The wild type (BY4741: MATa his3∆1 leu2∆0 met15∆0 ura3∆0) and gcr1∆, gcr2∆, opi1∆, ino2∆, ume6∆ strains were gifted by Prof. Ram Rajasekharan, Central Food Technological Research Institute (CFTRI), Mysore, India, and they obtained it from EUROSCARF. The yeast cells were grown at 30 °C in YPD medium (1% yeast extract, 2% peptone and 2% glucose) and SC (synthetic complete) medium containing 0.67% yeast nitrogenous base, supplemented with the amino acids and 2% glucose. Synthetic nitrogen starvation (SD-N) containing 0.17% yeast nitrogenous base without ammonium sulfate and amino acids and 2% glucose were used to induce autophagy. Cloned plasmids were transformed in Saccharomyces cerevisiae strains by lithium acetate method (Gietz and Schiestl 2007). The yeast transformants harboring the plasmids pYES2-NT/B, YEp357R, pRS316 and pRS315, pRS415 were cultured in SD-Ura and SD-Leu medium containing 2%, 0.2% glucose or 1.8% galactose as required. SC or SD (Synthetic defined) media containing 75 µM inositol was symbolized as I+, whereas media lacking inositol was denoted I−. The cells were pre-cultured until mid-log phase in SC or SD media. Then the cells were collected and washed with fresh inositol-free media and divided into two parts. One part was shifted to I + and another part to I− media at 30 °C for indicated time points.
Analysis of inositol auxotrophy (I−) phenotypes
Yeast cells were grown in SC and SD media lacking leucine (SD-Leu) at 30 °C up to mid-log phase. Cells were harvested, and cell density was adjusted to A600 = 1.0. The cells were serially diluted (1:10, 1:100, 1:1000 and 1:10000) and 3 µl of cells were spotted onto SC-D or SC-Leu agar plates, with or without inositol supplementation (75 µM) and incubated at 30 °C for 3 days. For the growth curve analysis, cell growth was monitored by measuring the cell density (OD at A600) at frequent time intervals until 64 h.
Glucose assay
Wild type (WT), gcr1∆ and opi1∆ strains were grown in synthetic complete (SC), and inositol-free medium at 30 °C with constant shaking at 180 rpm for 12 h. At indicated time points, 1 ml samples were collected and centrifuged and the supernatants were collected to determine the extra-cellular glucose concentrations using the glucose assay kit (Sigma Glucose-GO Assay Kit) according to the manufacturer’s instructions.
Intracellular inositol assay
The WT and deletion strains were grown in SC media and cells were harvested after 12 and 24 h by centrifugation (4 °C) and washed with ice-cold water and resuspended in ice-cold 1 M perchloric acid for deproteinization. The cells were lysed using acid-washed glass beads for 10 min at 30-s interval, alternating with 30-s incubation on ice and pH 7.0 was attained with 1 M KOH solution (ice-cold). The cell extracts were centrifuged at 9000 rpm for 10 min at 4 °C. The supernatant was collected, and intra-cellular inositol was measured using myo-inositol assay kit (K-INOSITOL, Megazyme) according to the manufacturer’s instructions.
RNA isolation and qRT-PCR analysis
Total RNA was extracted using the RNeasy kit from Qiagen according to manufacturer’s instructions. Complementary DNA (cDNA) was synthesized using the high-capacity cDNA reverse transcription kit with 1 × RT buffer, 1 × random primer, 4 mM dNTP mix, 50 U/ml reverse transcriptase and 2 µg of total RNA. The primers were designed with the Primer Express R Software 3.0 (Applied Biosystems), and the sequences are listed in Table 2. For qRT analysis, 1 µl of diluted cDNA (1:20) sample was amplified using Applied Biosystems machine Step One Plus™ Real-Time PCR machine with the Power SYBR Green PCR master mix (Applied Bio systems). Samples were evaluated by the ∆∆Ct method (Livak and Schmittgen 2001). The mRNA expression levels were analyzed in triplicates, and the results were analyzed using relative quantification, with actin (ACT1) as an endogenous control. Data are represented as relative mRNA expression.
Preparation of cell-free lysate
The cells were harvested and centrifuged at 5000 rpm for 5 min, washed and cell extract was prepared by lysing the cells with glass beads and lysis buffer containing 50 mM Tris/HCl (pH 8.0), 1 mM MgCl2, 10% glycerol and 1 mM phenylmethylsulfonyl fluoride (PMSF). Cells were vortexed for 30 s. The cells were kept on ice for 1 min between vortexing for all the 30 cycles. Unbroken cells were removed by centrifugation at 10,000 rpm for 10 min at 4 °C. Total protein was quantified (Bradford 1976) using BSA as standard.
Immunoblot analysis
For Western blotting analysis, proteins (50 µg) from cell extract were separated with 8% or 10% SDS–PAGE and transferred at 120 V for 2 h to the nitrocellulose membrane. The membrane was then incubated with primary anti-TAP antibody to detect Gcr1p (1:2000), anti-His antibody to detect Kcs1p (1:3000), anti-GFP antibody to detect Atg8p (1:2500), anti-Ape1 antibody to detect Ape1p (1:5000) and anti-Pgk1 to identify Pgk1p (1:5000) for 2 h at room temperature, followed by the ALP-conjugated goat anti-rabbit IgG secondary antibody for 1 h. The membrane was washed with PBST (0.1% Tween-20) and PBS. Finally, the blots were developed in the dark using the BCIP/NBT substrate. The intensity of Gcr1p and Kcs1p were normalized to the Pgk1p expression levels, and relative intensity was calculated using ImageJ analysis software (Abramoff et al. 2004).
Cloning, expression and purification of the recombinant His-tagged Gcr1 protein
Yeast genomic DNA was prepared as described previously (Sambrook et al. 1989). GCR1 gene and PIS1 promoter was amplified from yeast genomic DNA using the primers listed in Table 1. The GCR1 gene was cloned into the E. coli expression vector pET28a using BamHI and XhoI restriction sites. PIS1 promoter was cloned into the YEp357 vector using the BamHI and PstI sites. Clones were confirmed by double digestion and DNA sequencing. E. coli BL21 (DE3) cells harboring the pET28a-GCR1 and pET28a vector control plasmids were precultured in 5 ml of LB medium with 50 µg/ml of kanamycin and grown overnight at 37 °C. The precultured cells were inoculated into 200 ml of 2 × LB medium containing 50 µg/ml of kanamycin and grown at 30 °C to an OD 0.4–0.6. The cells were induced with 0.5 mM IPTG (isopropyl β-d-1 thiogalactopyranoside) for 3–4 h at 30 °C. Cells were harvested and the cell pellet was resuspended with lysis buffer (50 mM Tris–HCl, pH 8.0, 300 mM NaCl, 10 mM imidazole, 0.2% Triton-X-100, 10 mM β-mercaptoethanol, 1 mM phenylmethylsulfonyl fluoride (PMSF), and 10% glycerol). The cells were lysed by sonication, and the lysed protein samples were centrifuged. The supernatant containing the recombinant protein having a (His)6 tag was purified using Ni–NTA matrix and allowed to bind to the Ni–NTA beads at 4 °C for 4 h. The protein was eluted with lysis buffer containing 20 mM imidazole followed by a second wash (40 mM and 250 mM) imidazole in lysis buffer. A portion of the collected protein fraction was separated using 10% SDS–PAGE and protein expression was confirmed by immunoblotting using monoclonal His tag antibody as primary antibody (1:5000 dilution), and anti-mouse IgG-alkaline phosphatase as secondary antibody (1:2500 dilution) and developed with BCIP®-NBT substrate. The purified protein was used for EMSA studies.
Electrophoretic mobility shift assay (EMSA)
To visualize in vitro DNA–protein interactions, EMSA was performed using full-length Gcr1p and the promoters of PIS1, TPI1, KCS1, and INO1 genes (1 kb) that were amplified from WT genomic DNA using the primers listed in Table 2. The gel purified PCR products were used for the gel shift assay. The binding buffer consisted of 10% glycerol, 20 mM Tris, pH-8.0, 50 mM KCl, 5 mM MgCl2, 100 mM ZnSO4, 5 mM β-mercaptoethanol, 1 mM dithiothreitol (DTT) and 10 mM HEPES–NaOH, pH 7.9. Purified Gcr1p was mixed with DNA in the presence of binding buffer and incubated for 25 min at 25 °C. The DNA–protein complex was resolved on non-denaturing polyacrylamide gel 6% with 0.5 × Tris borate–EDTA (TBE) running buffer. The gels were stained with SYBR Green® nucleic acid staining dye (Molecular Probes) and the complex was visualized under UV light at 300 nm using Bio-Rad ChemiDoc™ XRS + Imager (Hellman and Fried 2007; Rajvanshi et al. 2017).
Lipid extraction and analysis
The WT and gcr1∆ cells were grown on SC medium up to mid-log phase at 30 °C. The cells were collected and washed with fresh I− media and divided into two parts. One part was shifted to I+ and another part to I– media containing 2% dextrose and allowed to grow for 4 h. The cells grown up to mid-log phase (0 h and 4 h) with I− and I+ were harvested, and lipids extracted (Bligh and Dyer 1959). Briefly, chloroform and methanol were added to the cell pellet in 2:1 (v/v) ratio and vortexed, and an equal volume of acidified water (2% phosphoric acid) was added and again vigorously vortexed. The lipid containing organic layer was dried, and phospholipids separated by thin-layer chromatography on silica Gel TLC plates using chloroform/ methanol/ acetic acid (65:25:8; v/v) as the solvent system. The TLC plate was exposed to iodine vapor to visualize the lipids, scraped from the TLC plate and quantified (Rouser et al. 1966).
INO1-LacZ promoter reporter assay
The YEp357R-INO1 and YEp357-PIS1 plasmid containing transformants were selected and grown in SD-Ura medium containing 2% glucose. The cells were collected and washed with fresh SD-Ura I− media and resuspended in fresh I− and I+ (SD-Ura) media and incubated at 30 °C for 4 h. Cell-free extract was prepared, and an equal concentration of protein extract was used to measure β-galactosidase activity (Rose and Botstein 1983). The specific activity was expressed as nmol. min−1 mg−1 protein.
Fluorescence microscope
The yeast vacuoles were stained with the lipophilic dye FM4-64 (Molecular Probes) according to the procedure reported earlier (Vida and Emr 1995) with slight modification. The cells were grown in SC medium containing 2% glucose (I− media containing either presence or absence of inositol) at 30 °C with constant shaking. The cells were pelleted and resuspended with 500 µl of fresh medium containing 20 µM FM4-64 for 30 min, followed by washing with fresh medium and incubating with fresh medium for 1 h. The images were acquired on Zeiss LSM 700 confocal laser scanning microscope equipped with a 100X/1.40 oil objective and an AxioCam MRM camera (Zeiss). The yeast Atg8–GFP transformants were grown in an appropriate medium, and the cells were collected and stained with FM 4–64 dye for visualizing the vacuolar membrane. The cells were viewed with an excitation wavelength of 488 nm for GFP and 515 nm for FM 4–64.
Statistical analysis
Experimental quantitative data were analyzed using Student’s t test, and the differences were considered statistically significant when *p< 0.05 and **p < 0.01. Each experiment was repeated at least three independent repeats. Data are presented as the average ± standard deviation (SD). Statistical analysis was performed using the Sigma plot 10.0 software.
Results
Inositol depletion in gcr1∆ cells depicted growth defect
The role of GCR1 in the synthesis of inositol, and growth was studied. Our results revealed that the growth of gcr1∆ cells was significantly reduced during inositol depletion (I− phenotype) condition when compared to the WT I−. Inositol limitation in the WT cells decreased the growth compared to inositol presence (Fig. 1a). However, the gcr1∆ cells under 75 µM inositol supplementation (I+) partially restored the growth defect (Fig. 1a). Further, the growth was studied using plate assay, which depicted growth defect under I − in gcr1∆ cells (Fig. 1b) and the supplementation of choline (100 mM) did not restore the growth and displayed strong inositol growth defect (Fig. S1a). The opi1∆ strain aids inositol overproduction (Fig. 1a, b) and serves as a positive control for inositol auxotrophy (Opi − phenotype) (Greenberg et al. 1982; Ye et al. 2013). Also, we tested diploid strain BY4743 and GCR1/gcr1∆ cell growth in the presence or absence of inositol and choline. The diploid strain, GCR1/gcr1∆ also exhibited a slight growth reduction on the I− C− and I− C + medium compared to wild type (BY4743) cells (Fig. S2). The GCR1/gcr1∆ displayed a semi-dominant growth defect under inositol limitation. However, the growth pattern of gcr2∆ was equivalent to WT (Fig. S1b and c). These results suggest that GCR1 mutant exhibited growth defect under inositol deprivation.
Effect of inositol dependent growth in gcr1Δ cells. The WT, gcr1∆ and opi1∆ cells were grown in YPD medium up to mid-log phase at 30 °C. a Growth curve analysis. The cells were grown in the absence (I−) or presence (I+) of inositol. The growth curve was studied by measuring the OD (A600 nm) of the cells at indicated time points until 64 h. The growth curve is an average of three experimental repeats. b Spot test. The cells were normalized and equal number was serially diluted (tenfold dilution), and 3 µl of cells were spotted onto SC-D (SC-2% dextrose) agar plates, with or without inositol supplementation (75 µM) and incubated at 30 °C for 3 days
GCR1 mutant reduced INO1 expression and decreased intracellular inositol level
The glycolytic genes are regulated by GCR1 transcription factor (Willis et al. 2003), and glucose is essential for inositol production in yeast (Fig. 2a), and the perturbation of glycolysis affects the de novo inositol biosynthesis (Shi et al. 2005). The extracellular glucose and intracellular inositol levels were measured in WT, gcr1∆ and opi1∆ cells. We found that the rate of glucose utilization was reduced in gcr1Δ cells (Fig. 2b) and the intracellular inositol level was also reduced both at 12 (50%) and 24 h (59%) when compared with WT cells (Fig. 2d). The WT and deletion strains were grown up to mid-log phase and shifted to fresh I− media and the glucose measured at indicated time points. We found a rapid consumption of glucose in WT and opi1Δ cells when compared to gcr1Δ cells (Fig. 2c). The consumption of extracellular glucose was increased in the opi1Δ strain, and the intracellular inositol level was higher (Fig. 2b, d). A previous study suggested that the INO1 expression was de-repressed under inositol limitation condition and it is via the Ino2–Ino4 transcriptional activation (Henry et al. 2012). In our study, the absence of inositol (I−) substantially up regulated the INO1 (8.02-fold) and INM1 (4.45-fold) expression in the WT cells (Fig. 2d). However, the gcr1Δ cells exhibited a significant down regulation of INM1 (Fig. 2d), and INO1 transcription both in the absence and presence of inositol compared to the WT. The INO2 mRNA expression was also remarkably decreased in the gcr1Δ cells (Fig. 2e) and was independent of inositol level when compared to the WT cells. We simultaneously performed the in vivo INO1 promoter activity in the gcr1Δ cells using the INO1 promoter fused LacZ reporter assay. We observed a substantial reduction of β-galactosidase activity in the gcr1Δ cells during inositol absence (98.6 ± 6.8 nmol min−1 mg−1), and presence (70.8 ± 4.3 nmol min−1 mg−1) when compared to the WT cells (Fig. 2f). The meiotic transcriptional regulator UME6 positively regulates the CHO1, CHO2, and OPI3 but negatively regulates INO1 as well as the phospholipid biosynthetic genes in yeast (Jackson and Lopes 1996). The INO1 expression was observed both in repressing and derepressing conditions with ume6Δ cells (Jackson and Lopes 1996) and we also observed a substantial increase of β-galactosidase activity in the ume6Δ cells in both I–/I+ conditions. The ino2Δ strain was used as a negative control, whereas opi1Δ and ume6Δ mutant strains served as positive controls for INO1-LacZ reporter assay (Fig. 2f). Together, these results suggested that the growth defect in gcr1∆ cells could be accounted for the decrease in intracellular inositol and INO1 mRNA expression.
Lack of GCR1 depletes inositol production and down regulates INO1 transcription. a The schematic diagram represents the inositol formation from glucose. b Analysis of glucose consumption. The WT, gcr1∆ and opi1∆ cells were grown in SC-D (2% dextrose) medium at 30 °C, and the extra-cellular glucose level was measured at indicated time points. c The glucose uptake was measured at indicated time points in WT, gcr1∆ and opi1∆ under inositol deprivation condition. d Intra-cellular inositol quantification. The WT, gcr1∆ and opi1∆ cells were grown in SC-D media and cells were harvested at 12 h or 24 h, and intra-cellular inositol was quantified as described under “Material and methods”. e qRT-PCR analysis of INO1, INM1, INO2, and INO4 gene expression. ACT1 serves as an endogenous control. f β-Galactosidase activity of INO1-LacZ. The WT, gcr1∆, opi1∆, ume6∆ and ino2∆ cells were transformed with the INO1-LacZ fusion gene. The β-Galactosidase activity was measured as described under “Material and methods”. The specific β-Galactosidase activity was expressed as units mg−1 (nmol min−1 mg−1). The data shown are the mean ± SD (*p < 0.05) from three independent experiments
Lack of GCR1 depleted the cellular PI levels
The INO1 expression was down regulated as well as the intra-cellular inositol level decreased in the gcr1∆ cells when compared to the WT cells. Further, we checked the effect of GCR1 deletion on phosphatidylinositol (PI) levels. There was a significant reduction in the PI content (~ 33%) in gcr1∆ cells when compared to the WT cells in SC media (Fig. 3a). The WT and gcr1∆ cells were grown on SC media up to a mid-log phase and equal volume of culture was shifted to I− and I + medium. After 4 h shift to the I + medium, we found an increased PI level in the WT cells relative to WT I− (Fig. 3a). Earlier study also suggested that the exogenous supplementation of inositol increases the rate of PI synthesis, and this could be due to the channelling of PA through CDP-DAG for PI synthesis, which led to a drop in the PA level (Loewen et al. 2004). The increase in PI content also depends on the fatty acids which are obtained from various sources including de novo fatty acid synthesis, TAG hydrolysis and PC turnover (Gardocki et al. 2005; Nikawa et al. 1987; Gaspar et al. 2006, 2011). Likewise, the inositol deprivation reduced the PI level in WT cells (Fig. 3a), and this reduction was accompanied with the accumulation of PA and CDP-DAG (Becker and lester 1977; Gaspar et al. 2006). The gcr1∆ cells showed a decrease in the PI level both in I− (~ 33%) and I+ (~ 36%) compared to respective WT cells (Fig. 3a). The expression study of the PIS1 gene in the gcr1∆ strain was down regulated both in the presence and absence of inositol compared to the WT I (Fig. 3b). The PIS1 mRNA expression was confirmed by the promoter reporter activity using PIS1-LacZ. The β-galactosidase activity was substantially lowered in the gcr1∆ cells under I− and I + compared to WT cells (Fig. 3c). These results suggested that the loss of GCR1 significantly down regulated the PIS1 expression and depleted the cellular PI level.
Deletion of gcr1∆ alters the PI and down regulate PIS1 expression. a Lipid analysis of WT and gcr1∆ cells. The cells were grown until mid-log phase in SC-D media at 30 °C, and cells were washed with I− media and resuspended with fresh I− and I+ media and grown up to 4 h. At this time point, cells were collected for lipid analysis (A600 nm − 100 OD) and resolved on silica TLC plate using solvent systems, chloroform/methanol/acetic acid (65:25:8, v/v). Phosphatidylcholine (PC); phosphatidylinositol (PI); phosphatidylethanolamine (PE). The PI was quantified as described in “Materials and methods”. The inset represents iodine image of TLC plate. (1) Synthetic complete (SC); (2) I−; (3) I+ .The data shown are the mean ± SD (*p < 0.05) from three independent experiments. b Quantitative expression analysis of PIS1 in the wild type and gcr1∆ in the presence or absence of inositol. ACT1 was used as an endogenous control. The data shown is the average of three independent experiments (*p < 0.05). c β-Galactosidase activity of PIS1-LacZ. The WT, gcr1∆ cells were harboring YEp357 containing PIS1-LacZ fusion gene. The β-Galactosidase activity was measured as described under “Material and methods”. The specific β-Galactosidase activity was expressed as units mg −1 (nmol min−1 mg−1). The data shown are the mean ± SD (*p < 0.05) from three independent experiments
Inositol limitation altered the GCR1 expression in WT cells
Further to confirm whether Gcr1 protein responds to inositol, we examined the effect of Gcr1p by shifting cells from SC to fresh I− and I+ medium. Over expression of GCR1 in the WT cells (WT + pRS315-GCR1) decreased the expression of Gcr1p under I− condition when compared to I+ condition in the cells (Fig. 4a). Likewise, the presence of inositol up regulated (1.6-fold) and absence down regulated the mRNA expression of GCR1 compared with WT cells at 0 h (Fig. 4b). The consensus sequence of Gcr1p binding motif is CTTCC (the CT box located in the upstream sequence of the genes (Baker 1991). It was reported that the TPI1 possess two Gcr1p-binding sites (– 385 and − 345) in the UAS region, and Gcr1p is required for its transcriptional activation (Huie et al. 1992). Consistently Gcr1p regulating gene TPI1 (triose phosphate isomerase 1) was up regulated (1.8-fold) under I + in WT cells. The transcription of TPI1 and GCR1 was set as ‘1’ at ‘0’ h and expression was decreased under I− condition when compared to I + in the WT cells (Fig. 4c). The Kcs1 protein expression was increased under I− condition (1.8-fold) when compared to I + in the WT cells (Fig. 5c), but the GCRI expression was increased under I+ condition (Fig. 4a, b). It has already been reported that the KCS1 expression negatively controls the Gcr1p mediated transcriptional regulation in yeast (Szijgyarto et al. 2011), and explains the decreased expression of Gcr1p under I− condition. The above result suggested that under inositol limitation the Kcs1p was increased and reduced Gcr1p expression.
Inositol limitation affects GCR1 expression. a, b The protein expression of GCR1 was measured using western blotting. The overnight culture of WT + pRS315 and WT + pRS315-GCR1 strains were shifted to fresh SD-Leu media and grown up to 0.3 OD. The cells were washed with fresh SD-Leu I− medium and resuspended with I− and I + media, and after 4 h the cells were lysed and subjected to 8% SDS–PAGE. The western blot analysis was performed using anti-TAP tagged antibody to detect Gcr1 protein (109 kDa including TAP tag), and anti-Pgk1 (45 kDa) was the loading control. The Gcr1p expression was normalized by Pgk1p. c The qRT-PCR analysis measured the gene expression of GCR1 and TPI1. The gene expression in wild type cells at 0 h was set as 1 (indicated horizontally as the dotted line) and the expression compared after 4 h both in the presence and absence of inositol. The data showed the mean ± SD of three independent experiments (*p < 0.05)
Role of KCS1 in GCR1 deletion cells under inositol limitation. a Cell growth and INO1- LacZ promoter reporter activity under inositol presence and absence condition in WT and kcs1∆ cells. b The KCS1 mRNA expression of WT and gcr1∆ cells grown with SC media. c The Kcs1p expression was measured using western blotting. The WT and gcr1∆ cells were over expressed with YES2-KCS1 and grown in SD-Ura (0.2% dextrose and 1.8% galactose) medium up to mid-log phase, the cells were washed and resuspended with I− or I+ medium. After 4 h the cells were lysed and subjected to 8% SDS–PAGE and Western blot was analyzed using anti-His tag antibody (to detect Kcs1p of size 126 kDa including His tag), and anti-Pgk1 (45 kDa) was the loading control. Kcs1p expression was normalized by Pgk1p and the data showed the mean ± SD of three independent experiments (*p < 0.05)
GCR1 deletion decreased Kcs1p level
The Kcs1p (inositol pyrophosphate synthase) generates inositol pyrophosphate (5PP-IP4) that stabilizes the interaction of Ino2p–Ino4p complex regulating INO1 transcription (Ye et al. 2013). The promoter reporter assay (INO1-LacZ) revealed a significant increase in the INO1 activity under I− condition with the WT cells. The INO1-LacZ activity was significantly reduced in kcs1∆ under both (I− and I+) conditions (Fig. 5a) and similar results were observed in the gcr1∆ cells (Fig. 2d). We found a growth reduction in gcr1∆ cells during I− condition (Fig. 1) and the kcs1∆ cells also showed a similar pattern (Fig. 5a). The kcs1∆ cells affected cellular inositol production, down regulated INO1 transcription and decreased PI level (Ye et al. 2013) and we observed similar results with gcr1∆ cells. Compared to the WT cells, significant down regulation of KCS1 expression was observed in the gcr1∆ cells (Fig. 5b). The Kcs1p expression was significantly increased in the WT + pYES2-KCS1 cells under I − condition and decreased under I + condition, whereas the Kcs1p expression was significantly decreased (both in the I− and I+) in the gcr1∆+pYES2-KCS1 cells, compared to WT I− cells (Fig. 5c). The INO1 expression was down regulated in both I- and I + in gcr1∆ cells (Fig. 2d). Aforementioned, Kcs1p is important for inositol auxotrophic growth and INO1 transcription (Ye et al. 2013). Thus, the reduced INO1 expression in the gcr1∆ cells might be accounted for the decreased expression of Kcs1p (Fig. 5c).
Enzymatic mobility shift assay (EMSA) confirms the binding of Gcr1p to promote the expression of PIS1
Transcription factor Gcr1p activates the diverse set of genes involved in glycolysis, RNA polymerase II and cell cycle regulating genes (Willis et al. 2003; Barbara et al. 2006; Baker 1986). We examined the in vitro analysis of DNA–protein interactions, and for this the full-length GCR1 was cloned into pET28a and expressed in BL21 pLys DE3 cells. The recombinant protein was purified using Ni2+ -NTA affinity chromatography and the purity confirmed by immunoblotting with anti-His tag monoclonal antibody (Fig. 6b). The in silico analysis we found Gcr1p binding site 5′CTTCC 3′ (CT Box) in the promoter region (upstream) of the PIS1 (– 885, – 735, – 690 and – 537) and INO1 (– 611, -596 and – 384) genes using YEASTRACT promoter database of S. cerevisiae. A previous study suggested that the Gcr1p positively regulates TPI1 expression in yeast (Huie et al. 1992). The Gcr1p binding site (CTTCC) is present at two positions (– 385 and − 345) on the promoter of TPI1 and serves as a positive control for EMSA. We performed an EMSA with 1000 bp region of the promoter of TPI1, INO1, PIS1 and KCS1 amplicon. The purified Gcr1p strongly binds with TPI1 promoter, which possesses two CT box binding site in the UAS region (Fig. 6c). On the other hand, the Gcr1p did not form a binding complex of INO1 promoter, even with the increasing concentration of Gcr1p (Fig. 6d). The KCS1 promoter does not possess CT box binding site, and so no binding complex was formed (Fig. 6e). Surprisingly, with the 1000 bp from the promoter region of the PIS1 and the purified Gcr1p, an increase in DNA–protein binding complex was observed with increasing concentration of Gcr1p (Fig. 6f). The above result suggested that the Gcr1p is an important for the activation of PIS1 but not for INO1 and KCS1 transcription.
Purification of Gcr1p and EMSA. a Purification of Gcr1p. The bacterially expressed recombinant Gcr1p was purified using nickel affinity chromatography and resolved on a 10% SDS–PAGE and stained with Coomassie brilliant blue (CBB Staining). M marker, FT flow through, W wash, E1–E4 eluted fractions of purified protein. b An immunoblot of Gcr1p using anti-His tag antibody. c EMSA of Gcr1p with TPI1 promoter d EMSA of Gcr1p with INO1 promoter e and f EMSA of Gcr1p with KCS1 and PIS1 promoters
GCR1 expression in gcr1∆ alleviates the inositol growth defect
The gcr1∆ strain depicts inositol dependent growth defect and we checked the impact of GCR1 overexpression in gcr1∆. The over expression of GCR1 in the gcr1∆ cells (pRS415-GCR1) restored the growth as in WT cells (with both I− and I+) unlike the gcr1∆ +pRS415 cells (Fig. 7a). The β-galactosidase activity of INO1 with the gcr1∆+ pRS415-GCR1 cells (in both I− and I + condition) reverted as seen with WT cells (Fig. 7b). Over expression of GCR1 gene in WT cells increased PIS1-LacZ activity under exogenous supplementation of inositol (Fig. 7c), which indicates Gcr1p positively controls PIS1. The INO1 expression in WT + pRS415-GCR1 and WT + pRS415 (in I− or I + condition) were similar (Fig. 7b). The over expression of WT + pRS425-GCR1 did not change INO1 transcription under inositol deprivation condition (Fig. 7b). The IP7 formed by Kcs1p donates the pyrophosphate to the phosphorylation site of Gcr1p, inactivating the Gcr1p-Gcr2p complex formation, as a result disturbing the Gcr1 mediated regulation (Szijgyarto et al. 2011).
Plate assay and promoter reporter assay under GCR1 over expression. a The spot test analysis (agar plates of SD-Leu + 2% dextrose, incubated for 3 days at 30 °C) was performed in wild type and gcr1∆ cells, transformed with pRS415 or pRS415-GCR1 under I− and I + condition. The data is a representative of the experiment repeated thrice. b, c The WT and gcr1∆ cells containing pRS415 and pRS415-GCR1 cells were co-transformed with INO1-LacZ fusion gene or PIS1-LacZ fusion gene and cells were grown up to mid-log phase in SD-Leu-Ura media. The cells were washed with fresh SD-Leu-Ura I− and seeded into the same media, either in the presence or absence of inositol. After 4 h induction, the cells were extracted, and the INO1-promoter activity or PIS1-promoter activity was assayed by measuring the β-Galactosidase activity that was expressed in units mg−1 (nmol min−1 mg−1). The assay represents the mean ± SD of three independent experiments (*p < 0.05)
Lack of GCR1 affects vacuolar morphology
The loss of Kcs1p displayed accumulation of small vacuolar structure and abnormal vacuolar membrane formation (Dubois et al. 2002). We examined the effect of GCR1 on vacuolar morphology, using the lipophilic dye FM 4–64. We visualized the yeast vacuoles, in SC media and the WT cells contain 1–3 vacuoles per cell (Fig. 8a, b), and inositol presence did not effect on the vacuole number or morphology (Fig. S3a–d). Unlike the WT cells, the gcr1∆ cells showed a substantial increase in the fragmented vacuoles (46% of cells display ≥ 4 vacuoles per cell). Normally, Kcs1p deletion affected dynamic cellular process including vacuolar membrane abnormality (Dubois et al. 2002). We observed an increase in the vacuolar number (75% of cells displayed ≥ 4 vacuoles per cell; Fig. 8a, b) with kcs1∆ cells in SC media. The INO1 expression is important for normal vacuolar morphology under inositol deprivation (Deranieh et al. 2015). We also observed an increase in the vacuolar fragmentation in gcr1∆ (67% of cells contain ≥ 4 vacuoles per cell Fig. S3a and b) cells under I − condition compared with WT I−. The addition of inositol altered the vacuolar morphology in the gcr1∆ (45% of cells contain ≥ 4 vacuoles) and kcs1∆ (58% of cells display ≥ 4 vacuoles per cell) cells when compared to WT I + cells (Fig S3c and d). Indeed, the gcr1∆ cells showed an alteration of vacuolar structure similar to kcs1∆ cells (Fig. 8a). The above result suggested that the lack of GCR1 reduced Kcs1p, possibly affecting vacuolar membrane morphology.
Visualization of vacuolar morphology. The WT, gcr1∆ and kcs1∆ cells were grown in SC medium up to mid-log phase and equal volume of culture was collected and vacuoles stained with FM4-64 lipophilic fluorescent dye as described under “Materials and methods”. a Vacuolar morphology of WT, gcr1∆, and kcs1∆ cells. b Quantification of labeled vacuoles per cell (n > 100 cells per stain). Images were observed by laser scanning confocal fluorescence microscope (LSM710-Zeiss). Scale bar, 5 µm
Deletion of GCR1 led to a defect in autophagy degradation
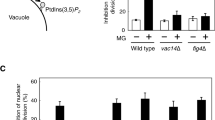
The vacuoles play an important role for cellular pH maintenance, response to osmotic shock, nutrient deprivation, and ion-homeostasis, specifically major site for intracellular proteolysis as well as micro and macroautophagy (Stauffer and Powers 2017; Li and Kane 2009). Normally, Atg8–GFP has been extensively used to observe the process of Cvt pathway and autophagy where Atg8–GFP is embedded in the inner membrane of the completed autophagosome. The synthesis of Atg8 is very low under vegetative growth and is induced upon starvation (Suzuki and Ohsumi 2007) and the autophagic flux was monitored by Atg8–GFP processing (Shintani and Klionsky 2004). The gcr1∆ cells decreased Kcs1 expression, and as a result an increased the number of vacuoles (Figs. 5c and 8). The WT, gcr1∆, atg1∆ and kcs1∆ cells were transformed with pRS316–ATG8–GFP plasmid and the transformants were grown to mid-log phase in SD-Ura with 2% dextrose (0 h), then the cells were pelleted and shifted in to SD-N media for 4 h or 8 h. The immunoblot analysis of Atg8–GFP was monitored using anti-GFP antibody. The WT cells generate free GFP under SD-N after 4 h, which indicates the translocation of Atg8–GFP to intravacuolar vesicles for Atg8–GFP degradation (Fig. 9a). However, the GFP molecule is more resistant to vacuolar protease, so the GFP accumulation is indicative of autophagic flux. Compared with the WT cells, the gcr1∆ cells had no free GFP under SD-N, similar to the atg1∆ and kcs1∆ cells (Fig. 9a). The autophagy defect was monitored by determining the delivery of Atg8–GFP to the vacuoles by fluorescence microscopy. The cells from mid-log phase, were shifted to SD-N media and grown for 4 and 8 h, and cells were collected and stained with FM 4–64 dye and examined under Axio Observed 3 Inverted fluorescence microscope (Zeiss). The WT cells, accumulated Atg8–GFP in the perivacuolar region known as phagophore assembly site (PAS) associated with FM 4–64 vacuolar membrane, while the autophagy deficient atg1∆ failed to accrue PAS and was dispersed in the cytoplasm. The gcr1∆ cells develop single PAS under mid-log phase (Fig. 9b). The WT cells accumulate GFP signal inside the vacuoles under SD-N at 4 and 8 h, indicating the efficient occurrence of autophagy flux (Figs. 9b and S3a). As a negative control, atg1∆ failed to accumulate GFP inside the vacuolar membrane. Similarly, reduced vacuolar GFP accumulation was found gcr1∆ cells both at 4 and 8 h under SD-N grew cells, indicating reduced autophagy than the WT cells (Figs. 9b and S3a). Compared to the WT cells, the co-localization percentage reduced in gcr1∆ (15% and 22% in SD-N 4 and 8 h) cells (Fig. S3b). Similarly, the gcr1∆ cells increased Atg8–GFP punctuate in cytosol under nitrogen starvation condition (Fig. 7c and Fig. S3a and c). Additionally, we also analyzed the function of the Cvt pathway using an anti-Ape1 antibody to detect cytoplasmic protein aminopeptidase 1 (Ape1) expression. Immunoblotting results show that the gcr1∆ cells under nitrogen starvation have a defect in mature Ape1 formation (Fig. 9d). In the atg1∆ (autophagy mutant) cells the mature Ape was not observed in both vegetative and starvation conditions (Fig. 9d). The results strongly suggest that the lack of GCR1 affects autophagosome biogenesis contributing to the deficiency in autophagy flux.
GCR1 deletion affects autophagy process analyzed by western blotting and fluorescent microscopy. The WT and deletion strains (gcr1∆, atg1∆, and kcs1∆) were transformed with the pRS315-Atg8–GFP expression plasmid. The transformants were grown up to mid-log phase in SD-Ura (0 h), followed by induction with nitrogen starvation media (SD-N) for 4 h. a GFP–Atg8 cleavage by immunoblotting analysis. Samples were collected and the cleavage of Atg8–GFP was analyzed by immunoblotting with anti-GFP antibody that detected both Atg8–GFP (40 kDa) and free GFP (26 kDa). The Pgk1 (45 kDa) served as loading control. b and c Fluorescent microscopic analysis of Atg8–GFP localization and FM 4–64 dye (vacuolar morphology) in WT, gcr1∆, kcs1∆, and atg1∆ cells. The cells were collected and the images were captured using Axio Observed 3 Inverted fluorescence microscope (Zeiss) with an excitation wavelength of 488 nm for Green fluorescence and 514 nm for FM 4–64. Scale bar, 5 µm. d Ape1 processing assay by immunoblotting. The WT, gcr1∆, atg1∆ and kcs1∆ cells were grown up to mid-log phase in SC media and starved for 4 h in SD-N media. The cells were collected, and protein extracted and analyzed by immunoblotting with Ape1 antibody (prApe1 ~ 60 kDa; mApe1 ~ 50 kDa size; Asterisk denotes non-specific band). The data are represented as three independent repeats. prApe1, precursor Ape1; mApe1, mature Ape1
Discussion
The Gcr1p is a known transcription factor that regulates glycolysis, ribosomal protein, cell cycle and RNA polymerase II in growing yeast cells (Willis et al. 2003; Clifton and Fraenkel 1981; Tornow et al. 1993; Barbara et al. 2006; Baker 1986). In the present study, we demonstrated that gcr1∆ cells affect inositol production, and cells displayed inositol dependent growth defect. Here we also report that the deletion of GCR1 displays abnormal vacuolar structure and defective autophagy flux.
The transcription factor GCR1 positively regulates the transcription of glycolytic genes (Baker 1986). The mutant TPI1 exhibited a growth defect under inositol limitation, leading to an increase in DHAP (Dihydroxyacetone phosphate) production that inhibited the inositol-P synthase activity (Shi et al. 2005). On the contrary, in gcr1∆ strain the glycolytic intermediate product DHAP was reduced (Willis et al. 2003), and growth defect was observed under I− condition. The growth curve and plate assay revealed that the cells lacking Gcr1p reduced the growth under inositol deprivation (Fig. 1), and the growth was partially restored upon inositol supplementation compared to the WT cells (Fig. 1). However, choline supplementation was not able to restore the growth (Fig. S1). As shown in Fig. 2, deletion of GCR1 delayed glucose consumption and reduced intra-cellular inositol level when compared with the WT cells. The WT and opi1∆ strains rapidly consumed the glucose under I− condition (Fig. 2c). This could be due to the activation of de novo inositol biosynthesis, under inositol limitation. The glucose-6 P is the substrate for inositol-3 P synthase that forms inositol-3 P, and is consequently dephosphorylated to form inositol. The INO1 (inositol-3 P synthase) is the hallmark gene for inositol auxotrophy (Henry et al. 2012; Ye et al. 2013; Shetty and Lopes 2010). In response to inositol depletion, gcr1∆ cells displayed down regulation of INO1 mRNA expression (Fig. 2e) and reduced β-galactosidase reporter (for INO1-LacZ activity) assay compared to WT cells (Fig. 2f) and is associated with inositol dependent growth defect. We conclude that the inositol defect in gcr1∆ cells is caused by defective INO1 transcription. In Saccharomyces cerevisiae, the Ino2p and Ino4p form a complex, and bind to UASINO sequence and activate the transcription of INO1 and other phospholipid biosynthetic genes (Henry et al. 2012). Both the ino2∆ and ino4∆ cells exhibit inositol auxotrophy, and down regulate the INO1 expression (Henry et al. 2012; Bachhawat et al. 1995; Shetty and Lopes 2010). In gcr1∆ cells, the mRNA expression of INO2 was significantly down regulated (in both I− and I+ condition) accounting for the reduction in inositol and INO1 transcription (Fig. 2e).
The PIS1 gene contains a predicted consensus sequence (CTTCC) for the transcription factor GCR1 (Gardocki and Lopes 2003). The expression of PIS1 was down regulated in gcr1∆ strain, resulting in decreased cellular PI level compared to WT (Fig. 3a, b). The PIS1-LacZ was measured by β-galactosidase activity and a reduction was observed in gcr1∆ cells both in the presence and absence of inositol (Fig. 3c). The PI level was reduced under inositol deprivation compared to the inositol presence in WT cells (Fig. 3a), which altered the lipid derivatives including sphingolipid, phosphoinositides (Jesch et al. 2010; Henry et al. 2014), and GPI anchor (Doering and Schekman 1996).
We found a reduced expression of Gcr1 under I− in WT cells (Fig. 4) and this could be attributed to the increased expression of Kcs1p (Fig. 5c) and phosphoinositides metabolism under I− (Jesch et al. 2010; Ye et al. 2013; Henry et al. 2014). The exogenous inositol supplementation increased glycolytic pathway and generated more ATP (Deranieh et al. 2015), and also increased Gcr1p expression in WT-pRS315-GCR1 cells (Fig. 4). The Kcs1p produces 5PP-IP4 that binds with Ino2p–Ino4p complex and activates the INO1 expression (Ye et al. 2013). On the other hand, Kcs1p forms pyrophosphate 5PP-IP5 which binds to Gcr1p and reduces its binding with Gcr2p that affected GCR1 mediated transcriptional regulation (Szijgyarto et al. 2011). The expression of Kcs1p was increased under inositol limitation condition (Fig. 5c), and is important for optimal INO1 transcription and inositol production in yeast (Ye et al. 2013). The Kcs1p expression in gcr1∆ cells, was reduced in both I − and I + condition compared to WT I− (Fig. 5c) validating the reduced INO1 transcription and inositol growth defect in gcr1∆ strain. The INO1 transcription was initiated by epigenetic transcriptional memory under inositol deprivation, through Sfl1p transcription factor (D’Urso and Brickner 2017). The Gcr1p regulates RNA polymerase II transcription (Santangelo 2006; Menon et al. 2005), and its mutation reduced INO1 transcription (Berroteran et al. 1994; Henry et al. 2014). Aforementioned, the loss of Gcr1p exhibited inositol dependent growth defect and it might be due to the misregulation of INO1 transcription (Ye et al. 2013) and the expression of GCR1in the gcr1∆ strain rescued the growth defect and INO1 expression under inositol deprivation (Fig. 7). The over expression of GCR1 in WT cells, did not change the level of INO1 expression and this could be due to the influence of Kcs1p under inositol deprivation (Fig. 7).
The defect in KCS1 depicted a defect in vacuolar membrane morphology (Saiardi et al. 2002; Dubois et al. 2002). The deletion of Ino1p exhibited vacuolar membrane dynamics and affects endocytosis under inositol limitation (Deranieh et al. 2015). The defective vacuolar morphology was increased in gcr1∆ cells, and kcs1∆ cells and both depicted a similar phenotype (Fig. 8). The gcr1∆ cells decreased the Kcs1p expression and thereby accounted for the defective vacuolar membrane. The gcr1∆ cells reduced vacuolar Atg8–GFP co-localization under nitrogen starvation compared to WT cells (Fig. 7c). There is an accumulation of Atg8–GFP punctuate in the cytosol in gcr1∆ cells under SD-N (4 h and 8 h) (Fig. S4). Kcs1p is required for proper localization of PAS for autophagosomal formation under nitrogen starvation (Taylor et al. 2012). The formation of mApe1 is reduced under nitrogen starvation in the gcr1∆ cells (Fig. 9d). Previous, study suggested that gcr1∆ strain is not affected by rapamycin (Menon et al. 2005; Lieb et al. 2001).
In summary, based on our findings we suggest that Gcr1p positively controls PIS1 transcription, and the deletion of GCR1 affects the Kcs1p which declines INO1 expression that resulted in inositol-dependent growth defect, vacuolar abnormality and autophagy defect in yeast. The potential regulatory role of Gcr1p in lipid metabolism is still elusive and yet to be explored.
References
Abramoff MD, Magelhaes PJ, Ram SJ (2004) Image processing with Image J. Biophoton Int 11:36–42
Anderson MS, Lopes JM (1996) Carbon source regulation of PIS1 gene expression in Saccharomyces cerevisiae involves the MCM1 gene and the two-component regulatory gene, SLN1. J Biol Chem 271:26596–26601
Bachhawat N, Ouyang Q, Henry SA (1995) Functional characterization of an inositol-sensitive upstream activation sequence in yeast. A cis-regulatory element responsible for inositol choline-mediated regulation of phospholipid biosynthesis. J Biol Chem 270:25087–25095
Baker HV (1986) Glycolytic gene expression in saccharomyces cerevisiae: nucleotide sequence of GCRI, null mutants, and evidence for expression. Mol Cell Biol 11:3774–3784
Baker HV (1991) GCR1 of Saccharomyces cerevisiae encodes a DNA binding protein whose binding is abolished by mutations in the CTTCC sequence motif. Proc Natl Acad Sci USA 88:9443–9447
Barbara KE, Haley TM, Willis KW, Santangelo GM (2006) The transcription factor Gcr1 stimulates cell growth by participating in nutrient-responsive gene expression on a global level. Mol Genet Genom 277:171–188
Becker GW, Lester RL (1977) Changes in phospholipids of Saccharomyces cerevisiae associated with inositol-less death. J Biol Chem 252:8684–8691
Berroteran RW, Ware DE, Hampsey M (1994) The sua8 suppressors of Saccharomyces cerevisiae encode replacements of conserved residues within the largest subunit of RNA polymerase II and affect transcription start site selection similarly to sua7 (TFIIB) mutations. Mol Cell Biol 14:226–237
Bligh EG, Dyer WJ (1959) A rapid method of total lipid extraction and purification. Can J Biochem Physiol 37:911–917
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254
Bryant NJ, Stevens TH (1998) Vacuole biogenesis in Saccharomyces cerevisiae: Protein transport pathways to the yeast vacuole. Microbiol Mol Biol Rev 62:230–247
Carman GM, Han GS (2007) Regulation of phospholipid synthesis in Saccharomyces cerevisiae by zinc depletion. Biochim Biophys Acta 1771:322–330
Carman GM, Han GS (2011) Regulation of phospholipid synthesis in the yeast Saccharomyces cerevisiae. Annu Rev Biochem 80:859–883
Chirala SS (1992) Coordinated regulation and inositol-mediated and fatty acid-mediated repression of fatty acid synthase genes in Saccharomyces cerevisiae. Proc Natl Acad Sci USA 89:10232–10236
Clifton D, Fraenkel DG (1981) The gcr (glycolysis regulation) mutation of Saccharomyces cerevisiae. J Biol Chem 256:13074–13078
D’Urso A, Brickner JH (2017) Epigenetic transcriptional memory. Curr Genet 63:435–439
Deranieh RM, Shi Y, Tarsio M, Chen Y, McCaffery JM, Kane PM, Greenberg ML (2015) Perturbation of the vacuolar ATPase A Novel Consequence Of Inositol Depletion. J Biol Chem 290:27460–27472
Doering TL, Schekman R (1996) GPI anchor attachment is required for Gas1p transport from the endoplasmic reticulum in COP II vesicles. EMBO J 15:182–191
Donahue TF, Henry SA (1981) myo-Inositol-1-phosphate synthase. Characteristics of the enzyme and identification of its structural gene in yeast. J Biol Chem 256:7077–7085
Dubois E, Scherens B, Vierendeels F, Ho MM, Messenguy F, Shears SB (2002) In Saccharomyces cerevisiae, the inositol polyphosphate kinase activity of Kcs1p is required for resistance to salt stress, cell wall integrity, and vacuolar morphogenesis. J Biol Chem 277:23755–23763
Gardocki ME, Lopes JM (2003) Expression of the yeast PIS1 Gene requires multiple regulatory elements including a Rox1p binding site. J Biol Chem 278:38646–38652
Gardocki ME, Bakewell M, Kamath D, Robinson K, Borovicka K, Lopes JM (2005) Genomic Analysis of PIS1 Gene Expression. Eukaryotic cell 4:604–614
Gaspar ML, Aregullin MA, Jesch SA, Henry SA (2006) Inositol induces a profound alteration in the pattern and rate of synthesis and turnover of membrane lipids in Saccharomyces cerevisiae. J Biol Chem 281:22773–22785
Gaspar ML, Hofbauer HF, Kohlwein SD, Henry SA (2011) Coordination of storage lipid synthesis and membrane biogenesis: evidence for cross-talk between triacylglycerol metabolism and phosphatidylinositol synthesis. J Biol Chem 286:1696–1708
Gietz RD, Schiestl RH (2007) High-efficiency yeast transformation using the LiAc/SS carrier DNA/PEG method. Nat Protoc 2:31–34
Greenberg ML, Reiner B, Henry SA (1982) Regulatory mutations of inositol biosynthesis in yeast: isolation of inositol-excreting mutants. Genetics 100:19–33
Han SH, Han GS, Iwanyshyn WM, Carman GM (2005) Regulation of the PIS1-encoded phosphatidylinositol synthase in Saccharomyces cerevisiae by zinc. J Biol Chem 280:29017–29024
Hellman LM, Fried MG (2007) Electrophoretic mobility shift assay (EMSA) for detecting protein-nucleic acid interactions. Nat Protoc 2:1849–1861
Henry SA, Kohlwein SD, Carman GM (2012) Metabolism and regulation of glycerolipids in the yeast Saccharomyces cerevisiae. Genetics 190:317–349
Henry SA, Gaspar ML, Jesch SA (2014) The response to inositol: Regulation of glycerolipid metabolism and stress response signaling in yeast. Chem Phys Lipids 180:23–43
Hossain MA, Claggett JM, Edwards SR1, Shi A, Pennebaker SL, Cheng MY, Hasty J, Johnson TL (2016) Posttranscriptional Regulation of Gcr1 Expression and Activity Is Crucial for Metabolic Adjustment in Response to Glucose Availability. Mol Cell 62:346–358
Huie MA, Scott EW, Drazinic CM, Lopez MC, Hornstra IK, Yang TP, Baker HV (1992) Characterization of the DNA-Binding Activity of GCR1: In Vivo Evidence for Two GCR1-Binding Sites in the Upstream Activating Sequence of TPI of Saccharomyces cerevisiae. Mol Cell Biol 6:2690–2700
Jackson JC, Lopes JM (1996) The yeast UME6 gene is required for both negative and positive transcriptional regulation of phospholipid biosynthetic gene expression. Nucleic Acids Res 24:1322–1329
Jesch SA, Gaspar ML, Stefan CJ, Aregullin MA, Henry SA (2010) Interruption of inositol sphingolipid synthesis triggers Stt4p-dependent protein kinaseC signaling. J Biol Chem 285:41947–41960
Kamada Y, Yoshino K, Kondo C, Kawamata T, Oshiro N, Yonezawa K, Ohsumi Y (2010) Tor directly controls the Atg1 kinase complex to regulate autophagy. Mol Cell Biol 30:1049–1058
Kliewe F, Kumme J, Grigat M, Hintze S, Schüller HJ (2017) Opi1 mediates repression of phospholipid biosynthesis by phosphate limitation in the yeast Saccharomyces cerevisiae. Yeast 34:67–81
Lenburg ME, O’Shea EK (2001) Genetic evidence for a morphogenetic function of the Saccharomyces cerevisiae Pho85 cyclin dependent kinase. Genetics 157:39–51
Li SC, Kane PM (2009) The yeast lysosome-like vacuole: endpoint and crossroads. Biochim Biophys Acta 1793:650–663
Lieb JD, Liu X, Botstein D, Brown PO (2001) Promoter-specific binding of Rap1 revealed by genome-wide maps of protein-DNA association. Nat Genet 28:327–334
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(–Delta Delta C (T)) method. Methods 25:402–408
Loewen CJ, Gaspar ML, Jesch SA, Delon C, Ktistakis NT, Henry SA, Levine TP (2004) Phospholipid metabolism regulated by a transcription factor sensing phosphatidic acid. Science 304:1644–1647
Menon BB, Sarma NJ, Pasula S, Deminoff SJ, Willis KA, Barbara KE, Andrews B, Santangelo GM (2005) Reverse recruitment: the Nup84 nuclear pore subcomplex mediates Rap1/Gcr1/Gcr2 transcriptional activation. Proc Natl Acad Sci USA 102:5749–5754
Murray M, Greenberg ML (2000) Expression of yeast INM1 encoding inositol monophosphatase is regulated by inositol, carbon source, and growth stage and is decreased by lithium and valproate. Mol Microbiol 36:651–661
Nikawa J, Kodaki T, Yamashita S (1987) Primary structure and disruption of the phosphatidylinositol synthase gene of Saccharomyces cerevisiae. J Biol Chem 262:4876–4881
Rajvanshi PK, Arya M, Rajasekharan R (2017) The stress-regulatory transcription factors Msn2 and Msn4 regulate fatty acid oxidation in budding yeast. J Biol Chem 292:18628–18643
Rao MJ, Srinivasan M, Rajasekharan R (2017) Cell size is regulated by phospholipids and not by storage lipids in Saccharomyces cerevisiae. Curr Genet 64:1071–1087
Rose M, Botstein D (1983) Construction and use of gene fusions to lacZ (β- galactosidase) that are expressed in yeast. Methods Enzymol 101:167–180
Rouser G, Siakotos AN, Fleischer S (1966) Quantitative analysis of phospholipids by thin layer chromatography and phosphorus analysis of spots. Lipids 1:85–86
Saiardi A, Sciambi C, McCaffery JM, Wendland B, Snyder SH (2002) Inositol pyrophosphates regulate endocytic trafficking. Proc Natl Acad Sci 99:14206–14211
Saiardi A, Resnick AC, Snowman AM, Wendland B, Snyder SH (2005) Inositol pyrophosphates regulate cell death and telomere length through phosphoinositide 3-kinase-related protein kinases. Proc Natl Acad Sci 102:1911–1914
Sambrook J, Fritsch EF, Maniatis T (1989) Molecular cloning, a laboratory manual, 2nd edn. Cold Spring Harbor Laboratory, Cold Spring Harbor, pp 6.55–6.56
Santangelo GM (2006) Glucose signaling in Saccharomyces cerevisiae. Microbiol Mol Biol Rev MMBR 70:253–282
Seker T, Hamamci H (2003) Trehalose, Glycogen and Ethanol metabolism in the gcr1 Mutant of Saccharomyces cerevisiae. Folia Microbiol (Praha) 48:193–198
Shetty A, Lopes JM (2010) Derepression of INO1 transcription requires cooperation between the Ino2p-Ino4p heterodimer and Cbf1p and recruitment of the ISW2 chromatin-remodeling complex. Eukaryot cell 12:1845–1855
Shi Y, Vaden DL, Ju S, Ding D, Geiger JH, Greenberg ML (2005) Genetic perturbation of glycolysis results in inhibition of de novo inositol biosynthesis. J Biol Chem 280:41805–41810
Shintani T, Klionsky DJ (2004) Cargo proteins facilitate the formation of transport vesicles in the cytoplasm to vacuole targeting pathway. J Biol Chem 279:29889–29894
Stauffer B, Powers T (2017) Target of rapamycin signaling mediates vacuolar fragmentation. Curr Genet 63:35–42
Sugimoto K (2018) Branching the Tel2 pathway for exact fit on phosphatidylinositol 3-kinase-related kinases. Curr Genet 64:965–970
Suzuki K, Ohsumi Y (2007) Molecular machinery of autophagosome formation in yeast, Saccharomyces cerevisiae. FEBS Lett 581:2156–2161
Szijgyarto Z, Garedew A, Azevedo C, Saiardi A (2011) Influence of inositol pyrophosphates on cellular energy dynamics. Science 334:802–805
Taylor R Jr, Chen PH, Chou CC, Patel J, Jin SV (2012) KCS1 deletion in Saccharomyces cerevisiae leads to a defect in translocation of autophagic proteins and reduces autophagosome formation. Autophagy 8:1300–1311
Tornow J, Zeng X, Gao W, Santangelo GM (1993) GCR1, a transcriptional activator in Saccharomyces cerevisiae, complexes with RAP1 and can function without its DNA binding domain. EMBO J 12:2431–2437
Turkel S, Turgut T, Lopez MC, Uemura H, Baker HV (2003) Mutations in GCR1 affect SUC2 gene expression in Saccharomyces cerevisiae. Mol Gen Genom 268:825–831
Vida TA, Emr SD (1995) A new vital stain for visualizing vacuolar membrane dynamics and endocytosis in yeast. J Cell Biol 128:779–792
Wang CW, Klionsky DJ (2003) The molecular mechanism of autophagy. Mol Med 9:65–76
Willis KA, Barbara KE, Menon BB, MoVat J, Andrews B, Santangelo GM (2003) The global transcriptional activator of Saccharomyces cerevisiae, Gcr1p, mediates the response to glucose by stimulating protein synthesis and CLN-dependent cell cycle progression. Genetics 165:1017–1029
Ye C, Bandara WM, Greenberg ML (2013) Regulation of Inositol metabolism is fine-tuned by inositol pyrophosphates in Saccharomyces cerevisiae. J Biol Chem 288:24898–24908
York JD, Guo S, Odom AR, Spiegelberg BD, Stolz LE (2001) An expanded view of inositol signaling. Adv Enzyme Regul 41:57–71
Acknowledgements
This work was supported by the Science and Engineering Research Board (SERB) Grant No: EMR/2016/001727, New Delhi, under EMR scheme. C. Ravi was supported by a fellowship from SERB, New Delhi. We are grateful to Prof. Ram Rajasekharan (Central Food Technological Research Institute, Mysore, India) for providing yeast strains, over expression plasmids, reagents and instrument facility. We thank Prof. Rashna Bhandari (Centre for DNA Fingerprinting and Diagnostics, India), and John M. Lopes (College of Natural Sciences, University of Massachusetts, Amherst, MA) for providing pYES2-KCS1 and YEp357R-INO1-LacZ plasmids. We thank Prof. Tracy L. Johnson (Molecular Cell and Developmental Biology, University of California, Los Angeles, CA 90095, USA), Prof. Ji-Sook Hahn (School of Chemical and Biological Engineering, Seoul National University, Seoul, Korea), and Prof. Ravi Manjithaya (Molecular Biology and Genetics Unit, Jawaharlal Nehru Centre for Advanced Scientific Research Jakkur, Bangalore 560 064, India) for providing plasmids pRS315-GCR1-TAP tag, pRS415GPD, pRS415GPD-GCR1 and pRS316–ATG8–GFP, respectively. We thank Prof, Yoshinori Oshsumi (Tokyo Institute of Technology, Yokohama, Japan) for providing Ape1 antiserum. We are thankful for the infrastructure facilities of DST-FIST, Department of Biochemistry, Life Sciences and DST-PURSE facilities, of Bharathidasan University.
Author information
Authors and Affiliations
Contributions
V.N. and C.R designed the experiments. C.R and R.G performed the experiments. C.R and V.N. discussed the data and wrote the paper. All authors reviewed the results and approved the final version of the manuscript.
Corresponding author
Additional information
Communicated by M. Kupiec.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Ravi, C., Gowsalya, R. & Nachiappan, V. Impaired GCR1 transcription resulted in defective inositol levels, vacuolar structure and autophagy in Saccharomyces cerevisiae. Curr Genet 65, 995–1014 (2019). https://doi.org/10.1007/s00294-019-00954-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00294-019-00954-2