Abstract
The search for novel pathological and functional amyloids represents one of the most important tasks of contemporary biomedicine. Formation of pathological amyloid fibrils in the aging brain causes incurable neurodegenerative disorders such as Alzheimer’s, Parkinson’s Huntington’s diseases. At the same time, a set of amyloids regulates vital processes in archaea, prokaryotes and eukaryotes. Our knowledge of the prevalence and biological significance of amyloids is limited due to the lack of universal methods for their identification. Here, using our original method of proteomic screening PSIA–LC–MALDI, we identified a number of proteins that form amyloid-like detergent-resistant aggregates in Saccharomyces cerevisiae. We revealed in yeast strains of different origin known yeast prions, prion-associated proteins, and a set of proteins whose amyloid properties were not shown before. A substantial number of the identified proteins are cell wall components, suggesting that amyloids may play important roles in the formation of this extracellular protective sheath. Two proteins identified in our screen, Gas1 and Ygp1, involved in biogenesis of the yeast cell wall, were selected for detailed analysis of amyloid properties. We show that Gas1 and Ygp1 demonstrate amyloid properties both in vivo in yeast cells and using the bacteria-based system C-DAG. Taken together, our data show that this proteomic approach is very useful for identification of novel amyloids.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Amyloids are protein fibrils that are composed of stacked monomers stabilized by intermolecular β-sheets arranged perpendicular to the fibril axis. Functional and pathological amyloids are presented in a broad range of organisms, from archaea and bacteria to mammals. Increased levels of expression or secretion of some proteins as well as specific mutations lead to more than 30 human amyloid-associated disorders (Sipe et al. 2012; Sarto-Jackson and Tomaska 2016). Some amyloids called “prions” are infectious and can be transmitted between organisms of the same or, in some cases, different species. Prion fibrils are fragmented into small oligomers that are infectious and incorporate new monomers, thus growing into the full-size fibrils that are fragmented again providing repeating cycles of prion replication (Kushnirov and Ter-Avanesyan 1998; Chernoff and Kiktev 2016). Formation of prion fibrils causes infectious neurodegenerative diseases in mammals (Prusiner 1982) and protein-based heritable traits in microorganisms such as Saccharomyces cerevisiae (Wickner 1994). The aggregation-prone conformational states of prion proteins are inherited in yeast, since prion aggregates are replicated and passed from mother to daughter cells. Prion conversion, like a mutation, may not only lead to protein inactivation but also to the acquisition of novel functions with specific phenotypic manifestations (Chernoff 2001).
Some proteins form functional amyloid fibrils in physiological conditions. For instance, spider webs and silkworm silk thread consist of amyloid fibrils of spidroin and fibroin proteins, respectively (Kenney et al. 2002; Humenik et al. 2015). Functional amyloids in bacteria contribute to biofilm development and can generate toxic oligomers that cause damage to lipid membranes (Schwartz and Boles 2013). A repeat domain of human Pmel17 protein forms fibrils that are essential for melanin deposition and synthesis in pigment-specific cells (Fowler et al. 2006). Fibrils of Bgl2 stabilize the cell wall of Saccharomyces cerevisiae (Bezsonov et al. 2013). It is likely that the yeast cell wall may contain an ensemble of proteins in amyloid state participating in the maintenance of its structure. Despite extensive study, our knowledge of the prevalence of amyloids in living organisms is very fragmentary. The progress in the discovery of novel amyloids can be provided by recently developed methods of proteomic screening for amyloid-forming proteins (Kryndushkin et al. 2013; Nizhnikov et al. 2014, 2016). All of these methods are based on the universal feature of amyloids to form detergent-resistant aggregates. The SDS-resistant aggregates can be separated from other proteins and identified by mass spectrometry. Two similar methods, TAPI (Kryndushkin et al. 2013) and PSIA (Nizhnikov et al. 2014), include the step of protein extraction from the gel. New gel-free modification of PSIA named PSIA–LC–MALDI is much more sensitive and allows to reveal even minor proteins forming SDS-resistant aggregates (Nizhnikov et al. 2016). Here, we applied this method to identify yeast proteins which form amyloid-like detergent-resistant aggregates in different yeast strains under native conditions. Based on the results of this proteomic screening, we compiled a list of the candidates for amyloid proteins in the yeast proteome. We have shown that two of these proteins, Gas1 and Ygp1, that participate in the maintenance of the cell wall form amyloid aggregates in yeast cells and in the bacterial C-DAG system.
Materials and methods
Yeast and bacterial strains
Strains of S. cerevisiae used in this work are listed in Table 1. The strains have different origin and prion status. GT409 strain is a derivative of the GT81-1C strain that was cured of [PSI +] and [PIN +] by guanidine hydrochloride.
To test the amyloid properties of Ygp1 and Gas1 proteins in the bacterial expression system C-DAG (curli-dependent amyloid generator), we used the Escherichia coli strain VS39 (F −, [araD139] B/r , Δ(argF-lac)169, λ −, e14 −, flhD5301, Δ(fruK-yeiR)725 (fruA25), relA1, rpsL150 (strR), rbsR22, Δ(fimBfimE)632(::IS1), deoC1 and Δ(csgBAC)(::kanR)) kindly provided by A. Hochschild. This strain is a derivative of the strain MC4100 carrying deletions of the curli-encoding genes, csgA, csgB, and csgC, generated by gene replacement with the neo gene providing resistance to kanamycin. VS39 contains a pACYC-derived pVS76 plasmid that directs the synthesis of the outer-membrane curli protein, CsgG, under the control of the IPTG-inducible lacUV5 promoter and carries the cat gene providing resistance to chloramphenicol (Sivanathan and Hochschild 2012).
Plasmids
The plasmids YGPM32106 and YGPM4h01 from the yeast genomic tiling library YSC4613 (Open Biosystems, USA) containing YGP1 and GAS1 genes, respectively, were used for PCR amplification of these genes and their fragments.
The pRS316-based plasmid pU-CUP1-YFP (Antonets et al. 2016) was used as a vector for obtaining plasmids with the hybrid YGP1-YFP and GAS1-YFP genes.
The plasmid pYGP1-YFP contains the chimeric YGP1-YFP gene under the P YGP1 promoter. To construct this plasmid, PCR-generated P YGP1 -YGP1 fragment, amplified using the primers ForPromYgp1 and RevYgp1 (Table 2), was digested with SalI and BamHI and inserted into the SalI–BamHI digested pU-CUP1-YFP plasmid.
pGAS1-YFP plasmid contains chimeric GAS1-YFP gene that encodes the fragment of Gas1 protein (a/a 1–486) fused to YFP, under the P GAS1 promoter. To construct this plasmid, P GAS1 -GAS1 fragment, PCR-generated using the primers ForPromGAS1 and RevGAS1, was digested with HindIII and BamHI and inserted into the HindIII and BamHI-digested pU-CUP1-YFP plasmid.
The plasmids pExport (pVS72) and pVS105 (Sivanathan and Hochschild 2013) were kindly provided by A. Hochshild. These plasmids contain chimeric genes encoding the signal sequence of CsgA protein fused to the M-domain (pVS105) or NM-domain (pVS72) of the Sup35 protein of S. cerevisiae, under the BAD promoter induced by arabinose. The plasmid pVS72 was used as a vector for cloning the selected fragments of YGP1 and GAS1 genes.
The plasmids pVS-YGP172 and pVS-GAS1C contain chimeric genes encoding CsgA signal sequence fused to Ygp1(72–354) and Gas1(291–487) fragments, correspondingly, under the BAD promoter. To obtain these plasmids, the fragments of YGP1 and GAS1 genes generated by PCR with the pairs of primers YGP1F (72) and YGP1R (354), and GAS1F (291) and GAS1R (487), respectively, were digested with NotI and XbaI and substituted for the XbaI-NotI fragment containing the SUP35NM in the pVS72 plasmid.
Proteomic screening and identification of proteins forming amyloid-like aggregates
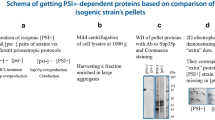
Screening of the S. cerevisiae proteome for potential amyloid-forming proteins was performed using PSIA–LC–MALDI approach (PSIA/Liquid chromatography coupled with mass spectrometry) described recently (Nizhnikov et al. 2016). This method consists of (1) the previously described procedure of isolation of detergent-resistant protein fractions (Nizhnikov et al. 2014), followed by (2) solubilization of proteins with formic acid and by boiling in SDS-PAGE loading buffer, (3) purification of proteins from detergent, trypsinolysis and (4) separation of tryptic peptides by high-performance liquid chromatography coupled with matrix-assisted laser desorption/ionization mass spectrometry (LC-MALDI).
Protein analysis
Preparation of protein lysates was performed as described previously (Newnam et al. 1999). Semi-denaturing detergent agarose gel electrophoresis (SDD-AGE) (Kryndushkin et al. 2003; Bagriantsev et al. 2006) was performed using 1% agarose gel. Before loading onto a gel, protein extracts were treated for 10 min with 1% SDS at room temperature. Then, the extract was subjected to SDD-AGE and transferred onto Immobilon-P PVDF membrane (GE Healthcare, USA). Proteins fused with YFP were reacted with polyclonal chicken primary antibodies against GFP (ab13970) (Abcam, Great Britain). Reactions with the secondary antibodies and chemiluminescent detection were performed using the Amersham ECL Prime Western Blotting Detection Reagent kit (GE Healthcare, USA).
Fluorescent microscopy
Fluorescent assay for YFP fusion proteins was performed with Leica DM6000B microscope (Leica Microsystems GmBH, Germany) and “Leica QWin standart V. 3.2.0.” software. The fluorescence was analyzed using YFP cube (Leica Microsystems GmBH, Germany) equipped with 535-nm barrier and 500-nm excitation filters.
Analysis of amyloid fibril formation in the bacteria-based system C-DAG
All the tests to verify the amyloid properties of Gas1 and Ygp1 proteins using bacterial expression system C-DAG (curli-dependent amyloid generator) were performed as described earlier (Sivanathan and Hochschild 2012, 2013). E. coli strain VS39 was transformed with the plasmids pVS-YGP72 and pVS-GAS1C, encoding the Ygp1(72–354)-and Gas1(291–487) fragments fused to CsgA signal sequence. VS39 transformants with pVS72 and pVS105 encoding the CsgASS-Sup35NM and CsgASS-Sup35M proteins were used as positive and negative controls of amyloid generation, respectively. To perform the tests for colony color phenotype and CR birefringence, spots of the transformants were grown for 5 days at 22 °C on the inducing medium with Congo Red (LB supplemented with 100 mg/l ampicillin, 25 mg/l chloramphenicol, 0.2% w/v L-arabinose, 1 mM IPTG and 10 mg/l Congo Red). To perform CR birefringence analysis, suspensions of the cells grown on CR-containing medium were spotted on a glass slide and analyzed between cross polarizers on the inverted microscope Leica DMI6000 B fitted with cross polarizers. Images were acquired using the Leica Application Suite software. To perform transmission electron microscopy analysis, spots of the transformants were grown for 5 days at 22 °C on the inducing medium without Congo Red (LB supplemented with 100 mg/l ampicillin, 25 mg/l chloramphenicol, 0.2% w/v L-arabinose and 1 mM IPTG). Cell suspensions were then analyzed on the transmission electron microscope Jeol JEM-2100 (“JEOL Ltd”, Japan).
Results
Proteomic screening for proteins forming detergent-resistant aggregates in yeast
To identify the proteins forming amyloid-like SDS-resistant aggregates in yeast, we used the strains of different origin GT81-1C [PSI +][PIN +], BY4742 [psi −][PIN +], GT409 [psi −] [pin −] and DC5 [psi −][pin −]. The isolation of the fraction containing the proteins forming SDS-resistant aggregates was performed as described earlier (Nizhnikov et al. 2014; see also “Materials and methods”). The proteins presented in this fraction were solubilized, treated with trypsin, separated by HPLC and identified by mass spectrometry (Nizhnikov et al. 2016).
Thirty-three proteins identified in each strain with the highest score (see Table S1–S4) were selected for further analysis. The lists of proteins revealed in all strains were very similar. Proteins identified by mass spectrometry in all four analyzed strains are presented in Table 3. We identified Rnq1 protein in both BY4742 [psi −][PIN +] and GT81-1C [PSI +][PIN +] strains and Sup35 protein in the GT81-1C [PSI +][PIN +] strain (Table 3). Identification of Pub1 in [PSI +] strain and Sis1 in both [psi −][PIN +] and [PSI +][PIN +] strains is fully consistent with previously obtained results, according to which the Pub1 protein forms aggregates in the presence of prion polymers of Sup35 (Urakov et al. 2010), whereas Sis1 coaggregates with different Q/N-rich amyloids (Bardill et al. 2009; Kryndushkin et al. 2013; Saarikangas and Barral 2016; Park et al. 2017). Importantly, the Bgl2 protein that is a functional yeast amyloid (Kalebina et al. 2008) was identified in all the strains analyzed (Table 3).
The list of proteins forming SDS-resistant aggregates in different yeast strains includes six cell wall proteins (Bgl2, Toh1, Ygp1, Gas1, Gas3, and Gas5), four RNA-binding proteins (Nop58, Nsr1, Nop1 and Tef1), the ssDNA binding protein Rim1, and also the vacuolar proteins Ape1 and Ape4. Note, all these proteins, except for Sup35 and Rnq1, form SDS-resistant aggregates in all the strains analyzed and do not exhibit prion-like conformational changes.
To determine whether the identified proteins are bona fide amyloids, analysis of their characteristics in vivo and in vitro is required. The ability of the cell wall proteins Gas1 and Ygp1 to form SDS-resistant aggregates in vivo and without overproduction has been shown previously (Nizhnikov et al. 2016). Moreover, Gas1 is functionally related to the Bgl2 amyloid-forming protein (Plotnikova et al. 2006). Only simultaneous deletions of the genes encoding Gas1 and Bgl2 cause significant impairment of the cell wall structure (Plotnikova et al. 2006). Ygp1 production is enhanced in the response to the cell wall damage (Brennan et al. 2013), and this protein may be involved in biofilm formation (Vandenbosch et al. 2013; Moreno-García et al. 2017). Based on these data, we selected Gas1 and Ygp1 proteins as the most promising candidates for amyloid-forming proteins.
The amyloid properties of the Gas1 and Ygp1 proteins in yeast cells
To verify that the Gas1 and Ygp1 proteins demonstrate amyloid properties in yeast cells, we constructed centromeric plasmids carrying the chimeric genes GAS1-YFP and YGP1-YFP under control of the GAS1 and YGP1 promoter, respectively. The BY4742 strain was transformed by these plasmids (see “Materials and methods”) and transformants were selected on the selective medium. Both Gas1-YFP and Ygp1-YFP proteins formed visible fluorescent aggregates (Fig. 1a) in about 38 and 47% of the cells, respectively. The Gas1-YFP protein typically forms from five to ten fluorescent granules per cell, whereas most cells producing the Ygp1-YFP protein contain only one visible aggregate.
Gas1-YFP and Ygp1-YFP proteins form detergent-resistant aggregates in yeast cells. a Localization of Gas1-YFP and Ygp1-YFP aggregates in yeast cells. Transformants bearing the plasmids encoding Gas1-YFP and Ygp1-YFP were grown on -Ura selective media for 48 h prior to fluorescence microscopy. Transformants containing the control pU-CUP1-YFP plasmid were incubated on -Ura selective media supplemented with 150 µM CuSO4. b SDD-AGE of protein lysates extracted from BY4742 cells expressing the proteins YFP (lines 1 and 3), Gas1-YFP (line 2) and Ygp1-YFP (line 4). Protein lysates were treated with 1% SDS at room temperature. The SDS-resistant aggregates of Gas1-YFP and Ygp1-YFP and monomers of YFP were detected using monoclonal rabbit primary antibodies ab32146 (Abcam, Great Britain) and the Amersham ECL Prime Western Blotting Detection Reagent kit (GE Healthcare, USA)
Next, we analyzed Gas1-YFP and Ygp1-YFP aggregation in the BY4742 strain using the semi-denaturing detergent agarose gel electrophoresis (SDD-AGE) assay (see “Materials and methods”). The BY4742 strain was also transformed by he pU-CUP1-YFP plasmid encoding the monomeric YFP protein for control. Both Gas1-YFP and Ygp1-YFP formed SDS-resistant aggregates in yeast cells (Fig. 1b), while the control protein YFP was detected in the monomeric fraction only.
Analysis of the amyloid properties of the Gas1 and Ygp1 proteins in the bacterial expression system C-DAG
To verify the amyloid properties of Gas1 and Ygp1 proteins, we used the bacterial export system for generating extracellular amyloid aggregates, which has been described recently (Sivanathan and Hochschild 2012, 2013). This system called the Curli-dependent amyloid generator (C-DAG) relies on the ability of E. coli cells to generate surface-associated amyloid fibrils (curli) composed of the CsgA and CsgB proteins. These proteins contain the N-terminal bipartite signal sequence (CsgASS) that directs them to the cell surface through the general Sec translocon and curli-specific pore-like structure in the outer membrane formed by the CsgG protein (Chapman et al. 2002; Robinson et al. 2006). Joining of the CsgASS fragment to the heterologous amyloidogenic proteins similarly directs their export to the cell surface where they form amyloid fibrils. The C-DAG approach represents a convenient cell-based alternative to the widely used in vitro methods for the study of protein’s amyloid properties, because it does not require purification of the protein of interest in soluble form, nor optimization of conditions for amyloid fibril assembly in vitro. Extracellular amyloids that are formed when using C-DAG can be easily detected in vivo using a colony color assay and studied by diverse methods including CR birefringence analysis and transmission electron microscopy.
Taking into consideration that C-DAG is used for the study of fragments 150–250 amino acid residues long, we constructed plasmids encoding almost all of the Ygp1 protein (residues 72–354) and central region of Gas1 protein (291–487). Both of these fragments contain the amyloidogenic sequences predicted by “Amylpred” (Table S5) (Tsolis et al. 2013). We studied the amyloid properties of chimeric proteins CsgASS-Ygp1(72–354) and CsgASS-Gas1(291–487) in the E.coli VS39 strain transformed with the plasmids pVS-YGP172 and pVS-GAS1C, respectively. As positive and negative controls for amyloid aggregation, we used VS39 transformants with pVS72 and pVS105 plasmids providing the expression of CsgASS-Sup35NM and CsgASS-Sup35M proteins, respectively. Firstly, we performed colony color phenotype tests and showed that expression of both CsgASS-Ygp1(72–354) and CsgASS-Gas1(291–487) resulted in red colonies of transformants on inducing medium containing Congo Red, although it was not as intense as in transformants expressing CsgASS-Sup35NM. The transformants with the pVS105 plasmid expressing CsgASS-Sup35M protein were pale on this medium (Fig. 2a). These data suggest that the CsgASS-Ygp1(72–354) and CsgASS-Gas1(291–487) proteins probably assemble into extracellular amyloid fibrils, which results in the red color of colonies due to Congo Red binding. To confirm this, we performed the test for CR birefringence by polarization microscopy as well as transmission electron microscopy of the transformants.
Analysis of the amyloid properties of the CsgAss-Gas1 (291–487), CsgAss-Ygp1 (72–385), CsgAss-Sup35NM and CsgAss-Sup35M proteins in the bacterial system C-DAG. a Colony color assay of the cells when plated on agar containing CR. b Micrographs of CsgAss-Gas1 (291–487), CsgAss-Ygp1 (72–385), CsgAss-Sup35NM and CsgAss-Sup35M scraped cell samples harvested from CR-containing agar. Extracellular material binds CR (left) and displays apple-green birefringence when viewed between crossed polarizers (right). The cultures producing the CsgAss-Sup35NM and CsgAss-Sup35M proteins were taken as positive and negative control, respectively
Using polarization microscopy, we showed that the CsgASS-Ygp1(72–354), CsgASS-Gas1(291–487), and CsgASS-Sup35NM proteins, but not CsgASS-Sup35M, bind Congo Red and manifest “apple-green” birefringence when examined between crossed polarizers (Fig. 2b). This property is characteristic of amyloid fibrils (Teng and Eisenberg 2009). Furthermore, we performed transmission electron microscopy analysis of the cell suspension of transformants scraped from inducing medium. The analysis revealed extracellular fibrils of the CsgASS-Ygp1(72–354), CsgASS-Gas1(291–487), and CsgASS-Sup35NM proteins, but not of the negative control protein CsgASS-Sup35M (Fig. 3).
Taken together, these data show that Ygp1 and Gas1 proteins possess amyloid properties in yeast cells, and their fragments form amyloid fibrils in the bacterial-based system C-DAG.
Discussion
Novel proteomic approaches open promising opportunities for identification of the landscapes of amyloids in different organisms. Here, using the PSIA–LC–MALDI proteomic approach, we identified proteins that form SDS-resistant aggregates in yeast and may thus be considered as candidates for amyloid-forming proteins. Comparative analysis was performed in four yeast strains of different origins. One of these strains, BY4742, contained prion [PIN +], while another strain GT81-1C had two prions, [PSI +] and [PIN +]. As expected, the Rnq1 protein was detected in the SDS-resistant fractions of both BY4742 [psi −][PIN +] and GT81-1C [PSI +][PIN +] strains, whereas the Sup35 protein was identified only in the GT81-1C [PSI +][PIN +] strain. Moreover, the Sis1 chaperone that binds to Q/N rich prion polymers (Bardill et al. 2009; Park et al. 2017) was detected only in the BY4742 [psi −][PIN +] and GT81-1C [PSI +][PIN +] strains, but not in the GT409 [psi −] [pin −] and DC5 [psi −][pin −] strains. We also showed that the Pub1 protein, which was previously demonstrated to co-aggregate with the [PSI +] prion (Urakov et al. 2010), was present in the SDS-resistant fraction exclusively in the [PSI +] GT81-1C strain. (see Table 3). All these data show that PSIA–LC–MALDI represents an adequate and very sensitive method for revealing amyloids at the proteome level. The lists of the proteins identified in all the strains were very similar. The proteins forming SDS-resistant aggregates in all four strains are presented in Table 3. We suggest that some of these proteins may function or be stored in yeast cells in an amyloid state. The aggregated state of the Ape1 aspartyl aminopeptidase, detected in our screen, was previously shown to be important for incorporation into the vesicles (Yuga et al. 2011). Based on our data, we may speculate that the Ape1 aggregates are possibly of an amyloid nature. Another interesting candidate for novel amyloid is the mitochondrial single-stranded DNA-binding protein Rim1. It was shown that Rim1 forms homotetramers in a solution (Ramanagoudr-Bhojappa et al. 2013). Our data show that in vivo in yeast cells this protein forms large aggregates that are resistant to SDS treatment like typical amyloids.
Six of the 13 proteins identified in our proteomic screen in all the analyzed strains are cell wall proteins. These data are in agreement with the hypothesis that an ensemble of proteins in an amyloid state participates in the maintenance of cell wall. One of the proteins identified in our work is the 1,3-beta-glucanosyltransferase Bgl2, whose amyloid properties in vivo have been well proven previously (Kalebina et al. 2008). Gas1, Gas3, and Gas5, detected in the proteomic screen are members of the beta-1,3-glucanosyltransferase family required for cell wall assembly (Ragni et al. 2007). Gas1 is also implicated in transcriptional silencing and is found at the nuclear periphery (Burgess et al. 2009; Koch and Pillus 2009). Notably, Gas1 and Bgl2 are functionally related and participate in the incorporation of the GPI-anchored proteins into the cell wall (Plotnikova et al. 2006). Only simultaneous deletion of the genes encoding Gas1 and Bgl2 causes significant disruption of the cell wall structure (Plotnikova et al. 2006). Considering that Gas1 was detected in the SDS-resistant fraction (see Table 3) and is functionally related to the Bgl2 amyloid, we selected this protein as a promising amyloid-forming candidate. Another interesting candidate detected in the proteomic screen was Ygp1, which is a cell wall-related secretory glycoprotein. The expression of YGP1 gene is drastically enhanced in the response to nutrient deprivation-associated growth arrest and cell wall damage (Destruelle et al. 1994; Brennan et al. 2013). Moreover, Ygp1 may be involved in biofilm formation (Vandenbosch et al. 2013; Moreno-García et al. 2017).
The proteomic screen performed in this study revealed amyloid candidates, but we cannot claim that all these proteins really form amyloid fibrils in vivo. For example, we cannot exclude that the pre-rRNA-binding proteins Nop58, Nsr1, and Nop1 presented in Table 3 form heavy non-amyloid ribonucleoprotein RNP complexes that do not completely dissociate even after RNAse and SDS treatment. Alternatively, some yeast RNA-binding proteins identified by PSIA–LC–MALDI may form amyloid-like oligomers, similarly to SDS-resistant RNA-binding oligomers of CPEB in Aplysia californica and its ortholog, Orb2, in Drosophila melanogaster (Si et al. 2003; Majumdar et al. 2012). The conclusion about the amyloid nature of the candidates identified in our screen can be made only after detailed analysis of the amyloid properties of each particular protein. Two proteins, Gas1 and Ygp1, identified by PSIA–LC–MALDI, were selected for such an analysis. Using immunoblotting analysis and fluorescent microscopy we showed that Gas1 and Ygp1 tagged with YFP formed SDS-resistant aggregates in yeast cells under native conditions (Fig. 1). The fragments of Gas1 and Ygp1 in the bacterial C-DAG system form extracellular amyloid fibrils which bind CR and demonstrate apple-green birefringence that is diagnostic of an amyloid state (Fig. 2). All these data allowed us to conclude that Gas1 and Ygp1 exhibit amyloid properties in vivo.
To conclude, in this work we performed proteomic screening and identified the proteins demonstrating amyloid properties in the yeast proteome. Resistance to treatment with ionic detergents is the universal feature of amyloid fibrils, but the levels of this resistance may vary (Nizhnikov et al. 2014). Thus, we cannot claim that our list includes all proteins that form amyloids in yeast. At the same time, PSIA–LC–MALDI yields very reproducible results and allowed us to identify Bgl2 that is the only bona fide amyloid of S. cerevisiae cell wall, well-known prions, and even prion-associated proteins, Sis1 and Pub1. Moreover, we showed that cell wall proteins, Gas1 and Ygp1, identified by PSIA–LC–MALDI, demonstrate amyloid properties in vivo. These data show that the list of proteins revealed in this work includes the most probable candidates for yeast amyloids. We suggest that the new proteomic methodology used here may provide significant progress in the discovery of prions and non-infectious amyloids in various organisms.
References
Allen KD, Wegrzyn RD, Chernova TA, Müller S, Newnam GP, Winslett PA, Wittich KB, Wilkinson KD, Chernoff YO (2005) Hsp70 chaperones as modulators of prion life cycle. Novel effects of Ssa and Ssb on the Saccharomyces cerevisiae Prion [PSI+]. Genetics 169:1227–1242
Antonets KS, Sargsyan HM, Nizhnikov AA (2016) Glutamine/Asparagine-rich fragment of Gln3, but not the full-length protein, aggregates in Saccharomyces cerevisiae. Biochemistry (Mosc) 81:407–413. doi:10.1134/S0006297916040118
Bagriantsev SN, Kushnirov VV, Liebman SW (2006) Analysis of amyloid aggregates using agarose gel electrophoresis. Methods Enzymol 412:33–48. doi:10.1016/S0076-6879(06)12003-0
Bardill JP, Dulle JE, Fisher JR, True HL (2009) Requirements of Hsp104p activity and Sis1p binding for propagation of the [RNQ(+)] prion. Prion 3:151–160
Bezsonov EE, Groenning M, Galzitskaya OV, Gorkovskii AA, Semisotnov GV, Selyakh IO, Ziganshin RH, Rekstina VV, Kudryashova IB, Kuznetsov SA, Kulaev IS, Kalebina TS (2013) Amyloidogenic peptides of yeast cell wall glucantransferase Bgl2p as a model for the investigation of its pH-dependent fibril formation. Prion 7:175–184. doi:10.4161/pri.22992
Brennan TC, Krömer JO, Nielsen LK (2013) Physiological and transcriptional responses of Saccharomyces cerevisiae to d-limonene show changes to the cell wall but not to the plasma membrane. Appl Environ Microbiol 79:3590–3600. doi:10.1128/AEM.00463-13
Broach JR, Strathern JN, Hicks JB (1979) Transformation in yeast: development of a hybrid cloning vector and isolation of the can1 gene. Gene 8:121–133
Burgess RJ, Guy MP, Zhang Z (2009) Fueling transcriptional silencing with Gas1. Proc Natl Acad Sci USA 106:10879–10880. doi:10.1073/pnas.0905192106
Chapman MR, Robinson LS, Pinkner JS, Roth R, Heuser J, Hammar M, Normark S, Hultgren SJ (2002) Role of Escherichia coli curli operons in directing amyloid fiber formation. Science 295:851–855. doi:10.1126/science.1067484
Chernoff YO (2001) Mutation processes at the protein level: is Lamarck back? Mutat Res 488:39–64
Chernoff YO, Kiktev DA (2016) Dual role of ribosome-associated chaperones in prion formation and propagation. Curr Genet 62:677–685. doi:10.1007/s00294-016-0586-2
Chernoff YO, Newnam GP, Kumar J, Allen K, Zink AD (1999) Evidence for a protein mutator in yeast: role of the Hsp70-related chaperone Ssb in formation, stability, and toxicity of the [PSI] prion. Mol Cell Biol 19:8103–8112
Destruelle M, Holzer H, Klionsky DJ (1994) Identification and characterization of a novel yeast gene: the YGP1 gene product is a highly glycosylated secreted protein that is synthesized in response to nutrient limitation. Mol Cell Biol 14:2740–2754
Fowler DM, Koulov AV, Alory-Jost C, Marks MS, Balch WE, Kelly JW (2006) Functional amyloid formation within mammalian tissue. PLoS Biol 4:e6. doi:10.1371/journal.pbio.0040006
Humenik M, Smith AM, Arndt S, Scheibel T (2015) Ion and seed dependent fibril assembly of a spidroin core domain. J Struct Biol 191:130–138. doi:10.1016/j.jsb.2015.06.021
Kalebina TS, Plotnikova TA, Gorkovskii AA, Selyakh IO, Galzitskaya OV et al (2008) Amyloid-like properties of Saccharomyces cerevisiae cell wall glucantransferase Bgl2p: prediction and experimental evidences. Prion 2:91–96
Kenney JM, Knight D, Wise MJ, Vollrath F (2002) Amyloidogenic nature of spider silk. Eur J Biochem 269:4159–4163
Koch MR, Pillus L (2009) The glucanosyltransferase Gas1 functions in transcriptional silencing. Proc Natl Acad Sci USA 106:11224–11229. doi:10.1073/pnas.0900809106
Kryndushkin DS, Alexandrov IM, Ter-Avanesyan MD, Kushnirov VV (2003) Yeast [PSI+] prion aggregates are formed by small Sup35 polymers fragmented by Hsp104. J Biol Chem 278:49636–49643. doi:10.1074/jbc.M307996200
Kryndushkin D, Pripuzova N, Burnett B, Shewmaker F (2013) Non-targeted identification of prions and amyloid-forming proteins from yeast and mammalian cells. J Biol Chem 288:27100–27111. doi:10.1074/jbc.M113.485359
Kushnirov VV, Ter-Avanesyan MD (1998) Structure and replication of yeast prions. Cell 94:13–16
Majumdar A, Cesario WC, White-Grindley E, Jiang H, Ren F, Khan MR, Li L, Choi EM, Kannan K, Guo F, Unruh J, Slaughter B, Si K (2012) Critical role of amyloid-like oligomers of Drosophila Orb2 in the persistence of memory. Cell 148:515–529. doi:10.1016/j.cell.2012.01.004
Moreno-García J, Mauricio JC, Moreno J, García-Martínez T (2017) Differential proteome analysis of a flor yeast strain under biofilm formation. Int J Mol Sci 18:720. doi:10.3390/ijms18040720
Newnam GP, Wegrzyn RD, Lindquist SL, Chernoff YO (1999) Antagonistic interactions between yeast chaperones Hsp104 and Hsp70 in prion curing. Mol Cell Biol 19:1325–1333. doi:10.1038/nprot.2013.081
Nizhnikov AA, Alexandrov AI, Ryzhova TA, Mitkevich OV, Dergalev AA, Ter-Avanesyan MD, Galkin AP (2014) Proteomic screening for amyloid proteins. PLoS One 9:e116003. doi:10.1371/journal.pone.0116003
Nizhnikov AA, Ryzhova TA, Volkov KV, Zadorsky SP, Sopova JV, Inge-Vechtomov SG, Galkin AP (2016) Interaction of prions causes heritable traits in Saccharomyces cerevisiae. PLoS Genet 12:e1006504. doi:10.1371/journal.pgen.1006504
Park SK, Hong JY, Arslan F, Kanneganti V, Patel B, Tietsort A, Tank EMH, Li X, Barmada SJ, Liebman SW (2017) Overexpression of the essential Sis1 chaperone reduces TDP-43 effects on toxicity and proteolysis. PLoS Genet 13:e1006805. doi:10.1371/journal.pgen.1006805
Plotnikova TA, Selyakh IO, Kalebina TS, Kulaev IS (2006) Bgl2p and Gas1p are the major glucan transferases forming the molecular ensemble of yeast cell wall. Dokl Biochem Biophys 409:244–247
Prusiner SB (1982) Novel proteinaceous infections particles cause scrapie. Science 216:136–144
Ragni E, Fontaine T, Gissi C, Latgè JP, Popolo L (2007) The Gas family of proteins of Saccharomyces cerevisiae: characterization and evolutionary analysis. Yeast 24:297–308. doi:10.1002/yea.1473
Ramanagoudr-Bhojappa R, Blair LP, Tackett AJ, Raney KD (2013) Physical and functional interaction between yeast Pif1 helicase and Rim1 single-stranded DNA binding protein. Nucleic Acids Res 41:1029–1046. doi:10.1093/nar/gks1088
Robinson LS, Ashman EM, Hultgren SJ, Chapman MR (2006) Secretion of curli fibre subunits is mediated by the outer membrane-localized CsgG protein. Mol Microbiol 59:870–881. doi:10.1111/j.1365-2958.2005.04997
Saarikangas J, Barral Y (2016) Protein aggregation as a mechanism of adaptive cellular responses. Curr Genet 62:711–724. doi:10.1007/s00294-016-0596-0
Sarto-Jackson I, Tomaska L (2016) How to bake a brain: yeast as a model neuron. Curr Genet 62:347–370. doi:10.1007/s00294-015-0554-2
Schwartz K, Boles BR (2013) Microbial amyloids-functions and interactions within the host. Curr Opin Microbiol 16:93–99. doi:10.1016/j.mib.2012.12.001
Si K, Lindquist S, Kandel ER (2003) A neuronal isoform of the aplysia CPEB has prion-like properties. Cell 115:879–891
Sipe JD, Benson MD, Buxbaum JN, Ikeda S, Merlini G et al (2012) Amyloid fibril protein nomenclature: 2012 recommendations from the Nomenclature Committee of the International Society of Amyloidosis. Amyloid 19:167–170. doi:10.3109/13506129.2012.734345
Sivanathan V, Hochschild A (2012) Generating extracellular amyloid aggregates using E. coli cells. Genes Dev 26:2659–2667. doi:10.1101/gad.205310.112
Sivanathan V, Hochschild A (2013) A bacterial export system for generating extracellular amyloid aggregates. Nat Protoc 8:1381–1390. doi:10.1038/nprot.2013.081
Teng PK, Eisenberg D (2009) Short protein segments can drive a non-fibrillizing protein into the amyloid state. Protein Eng Des Sel 22:531–536. doi:10.1093/protein/gzp037
Tsolis AC, Papandreou NC, Iconomidou VA, Hamodrakas SJ (2013) A consensus method for the prediction of “aggregation-prone” peptides in globular proteins. PLoS One 8(1):e54175
Urakov VN, Vishnevskaya AB, Alexandrov IM, Kushnirov VV, Smirnov VN, Ter-Avanesyan MD (2010) Interdependence of amyloid formation in yeast: implications for polyglutamine disorders and biological functions. Prion 4:45–52
Vandenbosch D, Canck E, Dhondt I, Rigole P, Nelis H, Coenye T (2013) Genomewide screening for genes involved in biofilm formation and miconazole susceptibility in Saccharomyces cerevisiae. FEMS Yeast Res 13:720–730. doi:10.1111/1567-1364.12071
Wickner RB (1994) [URE3] as an altered Ure2 protein: evidence for a prion analog in Saccharomyces cerevisiae. Science 264:566–569
Yuga M, Gomi K, Klionsky DJ, Shintani T (2011) Aspartyl aminopeptidase is imported from the cytoplasm to the vacuole by selective autophagy in Saccharomyces cerevisiae. J Biol Chem 286:13704–13713. doi:10.1074/jbc.M110.173906
Acknowledgements
The authors acknowledge Dr. A.A. Aleksandrov for critical reading of the manuscript. Special thanks go to Dr. A. Hochschild for providing the bacterial C-DAG system. The authors acknowledge St. Petersburg State University for opportunity to use facilities of the Research Resource Center for Molecular and Cell Technologies and the Resource Centers “CHROMAS” of SPbSU. This work was partially supported by the grant of SPbSU to A.P.G. and by the Russian Foundation for Basic Research (14-04-01463to A.P.G. and 16-34-60153 to A.A.N). The experiments on proteomic screening were supported by the Russian Science Foundation 14-50-00069 to SPbSU.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by M. Kupiec.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Ryzhova, T.A., Sopova, J.V., Zadorsky, S.P. et al. Screening for amyloid proteins in the yeast proteome. Curr Genet 64, 469–478 (2018). https://doi.org/10.1007/s00294-017-0759-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00294-017-0759-7