Abstract
Most superficial fungal infections are caused by dermatophytes, a specialized group of filamentous fungi which exclusively infect keratinized host structures such as hair, skin and nails. Since little is known about the molecular basis of pathogenicity and sexual reproduction in dermatophytes, here we functionally addressed two central transcriptional regulators, SteA and StuA. In the zoophilic species Arthroderma benhamiae a strategy for targeted genetic manipulation was recently established, and moreover, the species is teleomorphic and thus allows performing assays based on mating. By comparative genome analysis homologs of the developmental regulators SteA and StuA were identified in A. benhamiae. Knock-out mutants of the corresponding genes as well as complemented strains were generated and phenotypically characterized. In contrast to A. benhamiae wild type and complemented strains, both mutants failed to produce sexual reproductive structures in mating experiments. Analysis of growth on keratin substrates indicated that loss of steA resulted in the inability of ΔsteA mutants to produce hair perforation organs, but did not affect mycelia formation during growth on hair and nails. By contrast, ΔstuA mutants displayed a severe growth defect on these substrates, but were still able to produce hair perforations. Hence, formation of hair perforation organs and fungal growth on hair per se are differentially regulated processes. Our findings on the major role of SteA and StuA during sexual development and keratin degradation in A. benhamiae provide insights into their role in dermatophytes and further enhance our knowledge of basic biology and pathogenicity of these fungi.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The majority of superficial mycoses in humans and animals are caused by dermatophytes, a group of specialized filamentous fungi. Dermatophytes have the ability to infect keratinized host structures (skin, hair, nails) and utilize keratin as a growth substrate, an attribute associated with the pathogenicity of these fungi. Although the disease dermatophytosis affects millions of humans and animals worldwide (Weitzman and Summerbell 1995), little is known about the basic biology and pathogenicity mechanisms of these keratinophilic fungi at the molecular level (White et al. 2008). In the last years, however, full genome sequences of eight dermatophyte species (http://www.broadinstitute.org/annotation/genome/dermatophyte_comparative/MultiHome.html) (Burmester et al. 2011; Martinez et al. 2012) and advances in genetic manipulation have laid the basis for fundamental genetic research on these microorganisms (Grumbt et al. 2011b). In particular, the zoophilic species Arthroderma benhamiae was proven to be an ideal model organism for several reasons. The species grows comparatively fast, produces abundant microconidia and allows efficient targeted genetic manipulation (Grumbt et al. 2011a). Full genome information and global transcriptional profiles are available, and comprehensive in vitro and in vivo infection models have been established (Burmester et al. 2011; Staib et al. 2010; Zaugg et al. 2009). Finally, A. benhamiae is teleomorphic and thus allows studying sexual reproduction in dermatophytes (Ajello and Cheng 1967). Formation of fructifications called cleistothecia occurs when two compatible strains of opposite mating type (mt+ and mt−) meet (Symoens et al. 2011, 2013).
The discovery and functional analysis of major transcriptional regulators and related effector genes have allowed important insights in the basic biology and pathogenicity of various pathogenic fungi, such as Candida albicans (Ernst 2000; Staib et al. 2002) or Aspergillus fumigatus (Dinamarco et al. 2012; Ejzykowicz et al. 2009; Schrettl et al. 2010). Yet, only few transcription factors have been functionally studied in dermatophytes, e.g., PacC in Trichophyton rubrum (Ferreira-Nozawa et al. 2006). In this study, we analyzed the important transcription factors SteA (Ste12-like protein) and StuA of A. benhamiae. In other fungi these transcription factors have been shown to be required for sexual reproduction and pathogenicity (Aramayo et al. 1996; Wong Sak Hoi and Dumas 2010).
Ste12 and Ste12-like proteins are homeodomain transcription factors which are involved in fungal development and pathogenicity (Wong Sak Hoi and Dumas 2010). In Saccharomyces cerevisiae Ste12 combines two important mitogen-activated protein kinase (MAPK) signaling cascades, the pheromone and the filamentous growth pathway, and regulates mating, invasive growth and pseudohyphal growth (Chou et al. 2006; Errede and Ammerer 1989; Gavrias et al. 1996; Madhani and Fink 1997). The term Ste12 derives from gene mutations in S. cerevisiae that lead to sterile mutants designated ste (Hartwell 1980). Homologs of Ste12 have been identified in various yeasts and filamentous fungi and they are characterized by an N-terminal STE motif and a double C2H2 zinc finger domain at the C-terminus, which is absent in yeasts (Wong Sak Hoi and Dumas 2010). SteA of Aspergillus nidulans and pp-1 of Neurospora crassa are required for sexual reproduction (Li et al. 2005; Vallim et al. 2000). Deletion of ste 12 homologs in animal and plant pathogens, e.g., C. albicans and Fusarium oxysporum, demonstrated an essential role of these regulators for virulence (Lo et al. 1997; Rispail and Di Pietro 2009).
The stunted protein StuA belongs to the class of APSES (N. crassa Asm-1, S. cerevisiae Phd1, A. nidulans StuA, C. albicans Efg1 and S. cerevisiae Sok2) transcription factors which are characterized by a conserved basic helix-loop-helix (bHLH) DNA-binding motif (Aramayo et al. 1996; Dutton et al. 1997; Gimeno and Fink 1994; Stoldt et al. 1997; Ward et al. 1995). Members of the APSES protein family are important regulators of morphological processes, i.e., mating, conidiation and dimorphic growth, but they are also involved in the production of secondary metabolites and have been associated with virulence in many plant pathogenic fungi. Asm-1 of N. crassa and StuA of A. nidulans have been shown to be involved in both sexual and asexual development (Aramayo et al. 1996; Clutterbuck 1969; Wu and Miller 1997). In S. cerevisiae, the two APSES proteins Phd1 and Sok2 control opposite effects on dimorphic growth. Phd1 activates pseudohyphal growth, whereas Sok2 represses pseudohyphal differentiation (Gimeno and Fink 1994; Ward et al. 1995). Efg1 of C. albicans is important for hyphal development and chlamydospore formation (Doedt et al. 2004; Sonneborn et al. 1999). Homologs of StuA in the fungi Penicillium chrysogenum and Acremonium chrysogenum have been shown to regulate in part the biosynthesis of the antibiotics penicillin and cephalosporin, respectively (Hu et al. 2015; Sigl et al. 2011). In the plant pathogenic fungi Fusarium culmorum and Leptosphaeria maculans, ΔFcStuA and ΔLmStuA mutants, respectively, showed complete loss of pathogenicity (Pasquali et al. 2013; Soyer et al. 2015).
Given the multifacetted functions of SteA and StuA homologs in fungal biology and pathogenicity, here we set out to investigate the role of these transcriptional regulators in A. benhamiae. Knock-out mutants and complemented strains were constructed and analyzed for their ability to undergo sexual reproduction and keratin degradation. Therefore, mating experiments were performed and in vitro infection of human hair and nails was investigated.
Materials and methods
Strains and culture conditions
The wild-type strain A. benhamiae LAU2354-2 (mt+) was used for the generation of deletion mutants and reconstituted strains. For confrontation assays the wild-type strain A. benhamiae LAU1022 (mt−) was used. Both A. benhamiae wild-type strains have a white phenotype and were isolated from patients at the University of Lausanne (Switzerland) in previous studies. The wild-type A. benhamiae LAU2354-2 = CBS 112371 = IHEM 20161 was isolated by Fumeaux et al. (2004), whereas the wild-type A. benhamiae LAU1022 = IHEM 25063 was described by Symoens et al. (2013). All fungal strains used in this study were stored as frozen glycerol stocks at −80 °C and are listed in Table 1 with their corresponding reference. Sabouraud glucose (SAB) agar (1 % (w/v) peptone, 2 % (w/v) glucose, 1.5 % (w/v) agar) or potato dextrose agar (PDA; Carl Roth, Karlsruhe, Germany) was used for cultivation of the wild-type strains A. benhamiae LAU2354-2 and A. benhamiae LAU1022. Transformants of A. benhamiae LAU2354-2 were grown on SAB supplemented with 200 µg/mL hygromycin (ForMedium, Hunstanton, UK) or G418 (Carl Roth, Karlsruhe, Germany), according to the selectable marker used. MAT agar (0.1 % (w/v) peptone, 0.2 % (w/v) glucose, 0.1 % (w/v) MgSO4, 0.1 % (w/v) KH2PO4; Carl Roth, Karlsruhe, Germany) was used for the production of A. benhamiae microconidia. After 5 days of cultivation at 30 °C the microconidia were harvested with sterile water followed by filtration of the suspension (EASYstrainer™ Cell Strainer, 40 µm pore size, Greiner Bio-One, Frickenhausen, Germany). The number of microconidia was determined with a Thoma counting chamber.
Radial growth of the A. benhamiae LAU2354-2 wild type, ΔsteA and ΔstuA mutants and steA C and stuA C complemented strains was tested on SAB agar inoculated with 1 × 105 microconidia. After incubation at 30 °C for 5 days the diameter of colonies was measured.
For determination of cell dry weight a volume of 50 mL SAB medium was incubated with 5 × 107 microconidia of the respective fungal strain. After 5 days of cultivation at 30 °C and 200 rpm the mycelium was harvested by Miracloth (Calbiochem®, Merck Millipore, Darmstadt, Germany), thoroughly dried at 50 °C and weighed.
For the analysis of the number of conidia produced by A. benhamiae LAU2354-2 wild type, ΔsteA and ΔstuA mutants and steA C and stuA C complemented strains an amount of 107 microconidia of the corresponding strain was plated on MAT agar, incubated, harvested and counted as described above.
For mating/confrontation assays, all strains were directly taken from the frozen glycerol stocks and precultivated on SAB agar at 30 °C for 5–7 days. The selected wild-type strain A. benhamiae LAU1022 (mt−) was co-cultivated on MAT agar with the A. benhamiae LAU2354-2 wild type (mt+), ΔsteA and ΔstuA mutants and steA C and stuA C complemented strains. Small agar plugs (diameter 4 mm) of mycelium from the SAB agar plates were cut and placed on a mating plate 30 mm apart (as illustrated in Fig. 4). The plates were incubated at 25 °C for 4–8 weeks in the dark and regularly examined for the development of cleistothecia using a binocular (Stemi DV4, Zeiss). For visualization of asci and ascospore formation, 6- to 7-week-old cleistothecia/pseudocleistothecia were stained with Calcofluor White (0.1 % (w/v) Calcofluor White solved with 10 % KOH), squashed between a microscope slide and a coverslip, and then analyzed by fluorescence microscopy using an Axio Observer.Z1 microscope (Zeiss, Jena, Germany).
For the analysis of growth on keratin substrates hair and nails were used. Blond human scalp hair from a child and finger nails from a healthy female donor were cut, autoclaved and placed on water agar plates. The hair and nails were inoculated with three plugs of fresh mycelium from SAB agar plates. The cultures were incubated at 25 °C for 30 days (nails) or 40 days (hair) in the dark. In addition, hair perforation of single hair was inspected by light microscopy using an Axiostar plus microscope (Zeiss, Jena, Germany). Images were captured with an AxioCam MRc camera and AxioVision 3.1 software (Zeiss).
Plasmid generation
Sequence information for genes AbenSTEA (locus ARB_04076) and AbenSTUA (locus ARB_07703) were obtained from the annotated A. benhamiae LAU2354-2 genome sequence (http://www.broadinstitute.org/annotation/genome/dermatophyte_comparative/MultiHome.html). Plasmid construction was performed as described before (Grumbt et al. 2011a). In brief, for the generation of deletion mutants, up- and downstream sequences of the AbenSTEA gene were obtained by PCR with the primers AbenSTEA-1/AbenSTEA-2 and AbenSTEA-3/AbenSTEA-4 from genomic DNA of the wild-type A. benhamiae strain LAU2354-2 and cloned successively in the plasmid pHPH1 (Grumbt et al. 2011a) to result in pAbenSTEAM1 and pAbenSTEAM2, respectively. Similarly, the flanking regions of the AbenSTUA gene were amplified with the primers AbenSTUA-1/AbenSTUA-2 as well as AbenSTUA-3/AbenSTUA-4 and used to generate the plasmids pAbenSTUAM1 and pAbenSTUAM2, respectively. The plasmids pAbenSTEAM2 and pAbenSTUAM2 encode the hygromycin resistance gene (hph) flanked by up- and downstream regions of AbenSTEA and AbenSTUA, respectively (Fig. 1a, d). For complementation, the coding region of the AbenSTEA gene and AbenSTEA upstream sequences (amplified with the primers AbenSTEA-1/AbenSTEA-5) as well as AbenSTEA downstream sequences (amplified with the primers AbenSTEA-3/AbenSTEA-4) were successively cloned together with the [CaACT1T] DNA fragment from pJetGFPACT1T1 in the plasmid pNEO1 (Grumbt et al. 2011a) yielding pAbenSTEAK1 and pAbenSTEAK2, respectively.
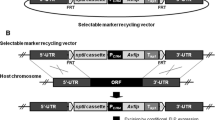
Generation of AbenSTEA and AbenSTUA deletion mutants and reconstituted strains. a For deletion of the AbenSTEA locus (white arrow) in the wild-type strain A. benhamiae LAU2354-2 (bottom) a DNA cassette, containing the hygromycin resistance gene hph (dark gray arrow) under control of the gpd promoter (P gpd , bent arrow) together with the termination sequence fragment T trpC (filled circle) flanked by STEA upstream and downstream regions (STEA up and STEA down, solid lines), was used (top). b For reinsertion of the STEA gene into its original locus in the ΔsteA mutants a DNA cassette, containing the coding region of AbenSTEA and the neomycin resistance gene neo (light gray arrow) under control of the A. benhamiae actin promoter (P ACT1 , bent arrow) together with the Candida albicans actin termination sequence fragment T ACT1 (blank circle) flanked by STEA upstream and downstream regions (STEA up and STEA down, solid lines), was applied. c Southern blot of SalI-digested genomic DNA of the wild-type strain A. benhamiae LAU2354-2, steA deletion mutants and steA C complemented strains with STEA-specific probe 1. d Deletion and e reconstitution of the AbenSTUA locus in the wild-type strain A. benhamiae LAU2354-2 were performed in the same manner as explained for AbenSTEA in a and b. f Southern blot of ClaI-digested genomic DNA of the wild-type strain A. benhamiae LAU2354-2, stuA deletion mutants, and stuA C complemented strains with STUA-specific probe 1. The probes used for Southern analysis of the transformants are indicated by black bars. Only the following relevant restriction sites are given in panels a, b, d and e: A, ApaI; B, BamHI; Bg, BglII; C, ClaI; H, HindIII; S1, SalI; X, XbaI. The sizes of the hybridizing DNA fragments (in kilobases) are given on the left, and their identities on the right
The plasmids pAbenSTUAK1 and pAbenSTUAK2 were constructed as described above using the coding region of the AbenSTUA gene and AbenSTUA upstream sequences amplified with the primers AbenSTUA-1/AbenSTUA-5 as well as AbenSTUA downstream sequences amplified with the primers AbenSTUA-3/AbenSTUA-4. The plasmids pAbenSTEAK2 and pAbenSTUAK2 encode the neomycin resistance gene (neo) flanked by the upstream region plus the coding region of AbenSTEA and AbenSTUA, respectively, under control of the A. benhamiae LAU2354-2 actin promoter (P ACT1 ) followed by the C. albicans actin terminator sequence fragment (T ACT1 ) and the downstream region of AbenSTEA and AbenSTUA, respectively (Fig. 1b, e). All primers used in this study are listed in Table S1.
A. benhamiae LAU2354-2 transformation and Southern hybridization
Transformation of A. benhamiae LAU2354-2 was carried out as previously described (Grumbt et al. 2011a). Hygromycin or neomycin-resistant transformants were selected with either 250 µg/mL hygromycin or G418 depending on the selection marker used. Disruption of the target gene or locus-specific complementation was confirmed by Southern hybridization using the Amersham ECL direct nucleic acid labeling and detection system (GE Healthcare) according to the manufacturer’s instructions (Fig. 1c, f).
Isolation of RNA and cDNA synthesis
For isolation of RNA the wild-type strain A. benhamiae LAU2354-2 was cultivated in SAB medium for 5 days at 30 °C and 200 rpm. The mycelium was harvested, frozen in liquid nitrogen and ground to a fine powder using mortar and pestle. RNA isolation was performed with the RNeasy® Plant Mini Kit (Qiagen) according to the manufacturer’s instructions. For the production of cDNA the RevertAid™ Premium First Strand cDNA Synthesis Kit (Thermo Fisher Scientific) was used. The cDNA was used as a template for amplification of an AbenSTEA gene segment with the primers AbenSTEA-13 and AbenSTEA-14 (Table S1). The resulting PCR product was cloned blunt end in the vector pJET1.2/blunt (CloneJET PCR Clonig Kit, Thermo Fisher Scientific).
Results
Identification of A. benhamiae LAU2354-2 SteA and StuA homologs
Via comparative analysis homologs of the two major transcriptional regulators SteA and StuA from A. nidulans were identified via comparative analysis in the genome sequence of A. benhamiae LAU2354-2 (http://www.broadinstitute.org/annotation/genome/dermatophyte_comparative/MultiHome.html). The analysis of the A. benhamiae AbenSTEA gene in the published data suggested that the gene is encoded by a 2382bp open reading frame interrupted by three introns. The deduced protein of 684 amino acids is characterized by an N-terminal STE homeodomain and a single C2H2 zinc finger domain in the C-terminal region which contrasts with all SteA homologs of filamentous fungi that have been identified so far. To analyze this finding in more detail RNA of A. benhamiae strain LAU2354-2 was isolated and transcribed into cDNA. The downstream region of AbenSTEA was amplified via PCR and the resulting fragment was cloned and sequenced. The identified cDNA, referred to as AbenSTEA-fs, is characterized by a different organization of the exons and introns in the downstream region in comparison to the available genome sequence of AbenSTEA. Thus, AbenSTEA-fs contains 5 exons and 4 introns and encodes a deduced protein of 712 aa with the N-terminal STE homeodomain (residues 64–157) and a double C2H2 zinc finger domain (residues 564–584 and 594–616) at the C-terminus (Fig. 2a). In contrast, the published AbenSTEA gene consists of 4 exons and 3 introns encoding a shorter protein that lacks 28 amino acids and contains only one C2H2 zinc finger domain (Fig. 2b). BLASTP search with the full-sized SteA (AbenSTEA-fs) of A. benhamiae LAU2354-2 revealed high similarities to other fungal Ste12 homologs, such as SteA of Trichophyton tonsurans (99 %), Coccidioides posadasii (70 % identity), A. fumigatus (65 % identity) and A. nidulans (64 % identity) as well as Talaromyces (formerly Penicillium) marneffei (64 % identity).
Two different AbenSTEA transcripts of A. benhamiae. a Localization of exons (E1–E5) and introns (I1–I4) in AbenSTEA-fs as well as the STE homeodomain (gray) and the two C2H2 zinc finger domains (red and green) with their corresponding amino acid sequence are shown. b Localization of exons and introns in the AbenSTEA gene based on the available genome data of A. benhamiae and the deduced amino acid sequence with only one C2H2 zinc finger domain
A. benhamiae LAU2354-2 AbenSTUA is encoded by an open reading frame of 2085 bp interrupted by three introns. AbenSTUA encodes a putative protein of 622 amino acids which is characterized by the highly conserved APSES domain (Fig. S1). A BLASTP search revealed that the amino acid sequence of AbenStuA has high similarity to StuA homologs of A. nidulans (52 % identity), N. crassa (46 % identity), C. albicans (59 % identity) and S. cerevisiae (73 % identity for Sok2 and 69 % identity for Phd1).
Generation of A. benhamiae LAU2354-2 ΔsteA and ΔstuA deletion mutants and reconstituted strains
To assess the functional role of AbenSteA and AbenStuA, ΔsteA and ΔstuA deletion mutants were generated in the wild-type A. benhamiae strain LAU2354-2, following a recently established protocol for gene targeting in A. benhamiae wild-type strain LAU2354-2 (Grumbt et al. 2011a). For further investigation, two independently obtained ΔsteA (AbenSTEAM1A and AbenSTEAM1B) and ΔstuA (AbenSTUAM1B and AbenSTUAM1F) mutants were used. To ensure that the observed phenotypes were a result of the deletion of either AbenSTEA or AbenSTUA, the ΔsteA and ΔstuA knock-out mutants were complemented with a copy of the wild-type AbenSTEA and AbenSTUA gene, respectively. The steA C (AbenSTEAK1A and AbenSTEAK1B) and stuA C (AbenSTUAK1B and AbenSTUAK1F) reconstituted strains were included in the further analysis.
SteA and StuA are not essential for conidiation and vegetative growth
Growth inspection of A. benhamiae LAU2354-2 wild type, mutants and reconstituted strains revealed that SteA and StuA were not essential for conidiation and vegetative growth (Fig. 3), albeit slight differences were noted as follows. In comparison to the A. benhamiae LAU2354-2 wild-type strain, ΔstuA mutants showed slightly impaired radial growth and biomass production on SAB agar (Fig. 3b). Additionally, the colony morphology of the ΔstuA mutants was altered as the mutants produced less aerial hyphae on MAT agar and displayed a folded surface on PDA (Figs. 4, 5). Complementation of the AbenSTUA gene restored the phenotype of the wild-type A. benhamiae LAU2354-2. In contrast, no growth differences were observed for the mutant lacking the AbenSTEA gene (Fig. 3a). Furthermore, conidiation of both mutants ΔsteA and ΔstuA was similar to the wild type (Fig. 3a, b).
The transcriptional regulators SteA and StuA of A. benhamiae are not essential for conidiation and vegetative growth. Conidiation, radial growth and biomass production of A. benhamiae LAU2354-2 wild type, ΔsteA (a) and ΔstuA (b) mutants as well as steA C and stuA C complemented strains were determined after 5 days at 30 °C. Data represent the means ± SDs of three simultaneously cultivated biological replicates. Unpaired t test, two-tailed, **significant at P < 0.05
The transcriptional regulators SteA and StuA of A. benhamiae are required for sexual reproduction. a Confrontation of the wild-type strains A. benhamiae LAU2354-2 (mt+) and A. benhamiae LAU1022 (mt−) (left side of the agar plate) resulted in cleistothecia formation (middle). The squashed cleistothecium shows peridial hyphae and asci with ascospores indicated by arrows (right; Calcofluor White staining and fluorescence microscopy). b Confrontation of ΔsteA mutants with A. benhamiae LAU1022 wild type resulted in the formation of pseudocleistothecia (middle) which consist of hyphae and microconidia (right). d Confrontation of ΔstuA mutants with A. benhamiae LAU1022 led to the formation of pseudocleistothecia (middle) with peridial hyphae and microconidia (right). c and e Confrontation of steA C and stuA C complemented strains with A. benhamiae LAU022, respectively, resulted in the development of cleistothecia (middle) which contain asci with ascospores (right). The two independently generated mutant strains of ΔsteA, ΔstuA, steA C and stuA C behaved identically, and only one of each is shown. Pictures were taken after 8–9 weeks of incubation except for fluorescence microscopy images which were taken after 6–7 weeks. Scale bars: left 10 mm; middle 1 mm; right 10 µm
Both transcriptional regulators SteA and StuA of A. benhamiae are involved in growth on keratin substrates. A. benhamiae LAU2354-2 wild type, ΔsteA and ΔstuA deletion mutants, steA C and stuA C complemented strains were grown on PDA for 6 days at 30 °C, on human nails for 30 days at 25 °C and on human hair for 40 days at 25 °C. Macroscopic inspection revealed a severe growth defect for ΔstuA mutants on human nails and hair (d). Representative microscopic pictures of single hair demonstrate the absence of the characteristic hair perforation organs in ΔsteA mutants (b). The two independently generated mutants of ΔsteA, ΔstuA, steA C and stuA C, respectively, behaved identically; only one of each is shown. White and black scale bars represent 10 mm and 20 μm, respectively
Both SteA and StuA are essential for the formation of cleistothecia
The heterothallic species A. benhamiae is able to undergo sexual reproduction which requires two mating competent strains, i.e., isolates with “+” and “−”mating type. Since fungal homologs of SteA and StuA have been described as important transcriptional regulators of sexual development, the role of SteA and StuA in sexual reproduction was investigated. Hence, the wild-type strain A. benhamiae LAU2354-2 (mt+) as well as the generated ΔsteA and ΔstuA mutants and reconstituted strains steA C and stuA C were analyzed for their ability to mate with the selected wild-type strain A. benhamiae LAU1022 (mt−). Plugs of the A. benhamiae LAU1022 wild type were co-cultivated with A. benhamiae LAU2354-2 wild type and transformant derivatives, repectively. The mating plates were regularly examined regarding cleistothecia formation in the contact zone of the two strains. Mature cleistothecia were visible as small white to beige, globose, firm structures surrounded by interwoven hyphae (peridial hyphae). For further analysis cleistothecia were squashed, stained with Calcofluor White and examined via fluorescence microscopy. Fertile cleistothecia consisted of globose to oval eight-spored asci. The ability of A. benhamiae LAU2354-2 to produce fertile cleistothecia with A. benhamiae LAU1022 has been recently reported (Symoens et al. 2013), and was confirmed in this study (Fig. 4a). By contrast, confrontation of A. benhamiae LAU1022 with ΔsteA mutants led to the development of white, globose, soft structures consisting of hyphae and microconidia which are called pseudocleistothecia; asci were not detected (Fig. 4b). The pseudocleistothecia produced during mating of the ΔstuA mutants with A. benhamiae LAU1022 contained microconidia and peridial hyphae with asymmetrically constricted dumbbell-shaped cells; asci were not detected (Fig. 4d). The reconstituted strains steA C and stuA C behaved like the wild type and developed fertile cleistothecia containing asci with ascospores when mated with A. benhamiae LAU1022 (Fig. 4c, e). The confrontation assays of A. benhamiae LAU1022 (mt−) with either A. benhamiae LAU2354-2 (mt+), ΔsteA and ΔstuA mutants or steA C and stuA C complemented strains were performed twice. No differences in the formation of cleistothecia or pseudocleistothecia were observed between the assays.
StuA but not SteA is required for growth on keratin substrates
The ability of dermatophytes to degrade and utilize keratin as a growth substrate is considered to be highly important for the pathogenicity of these medically important fungi. In order to study a potential role of SteA and StuA for keratin degradation, in vitro growth of A. benhamiae LAU2354-2 wild type, deletion mutants ΔsteA and ΔstuA and reconstituted strains steA C and stuA C on keratin substrates was analyzed. Therefore, human hair and nails were infected with A. benhamiae LAU2354-2 wild type, deletion mutants and reconstituted strains and examined regarding mycelia formation and hair perforation (Fig. 5). The deletion of AbenSTEA resulted in the inability of ΔsteA mutants to produce hair perforation organs but did not affect mycelia formation during growth on human hair and nails (Fig. 5b). In contrast, infection of human hair and nails with ΔstuA mutants demonstrated a severe growth defect on these substrates. Interestingly, however, although fungal multiplication of ΔstuA mutants was not visible, microscopical study of hair still revealed the typical wedge-shaped hair perforation organs (Fig. 5d). Additionally, hair infected with either ΔsteA or ΔstuA mutants was more fragile (analyzed by bending) than non-infected hair. No differences in growth and hair perforation were observed between A. benhamiae LAU2354-2 wild type and reconstituted strains steA C and stuA C during infection of human hair and nails (Fig. 5a, c, e).
Discussion
The transcriptional regulators SteA and StuA have been shown to be important global regulators in yeasts and filamentous fungi (Aramayo et al. 1996; Wong Sak Hoi and Dumas 2010). However, little is known about the function of transcription factors in dermatophytes. The present investigation revealed central roles for major transcriptional regulators in dermatophytes related to both basic biology and pathogenicity. Particularly, A. benhamiae transcription factors SteA and StuA were shown to be required for sexual development and moreover, StuA is essential for the destruction of keratinized host structures.
Homologs of SteA have been identified in a variety of fungal species. The C-terminal double C2H2 zinc finger domains are specific for filamentous fungi, but are absent in yeasts (Wong Sak Hoi and Dumas 2010). Although the available genome data of the SteA homolog in A. benhamiae indicate that the protein contains only one C2H2 zinc finger domain, we identified a steA transcript encoding for a protein with a double C-terminal C2H2 zinc finger domain. Interestingly, alternative splicing of SteA homologs of the plant pathogenic fungi Colletotrichum lindemuthianum and Botrytis cinerea resulted in two variants of the protein, one with a double and the other with a single zinc finger domain due to exon skipping (Schamber et al. 2010; Wong Sak Hoi et al. 2007). A similar splicing pattern can also be suggested for A. benhamiae. The full-size SteA protein of A. benhamiae contains two zinc finger domains, whereas the truncated version lacks one zinc finger motif probably as a result of exon skipping. In both fungi C. lindemuthianum and B. cinerea, it has been shown that the truncated transcript played a repressing regulatory role in fungal invasive growth, suggesting that alternative splicing can contribute to gene regulation and virulence (Schamber et al. 2010; Wong Sak Hoi et al. 2007).
A. benhamiae steA knock-out mutants showed no defect in vegetative growth and asexual reproduction, but were not able to form any sexual reproductive structures such as cleistothecia, peridial hyphae, asci or ascospores. Similar results were obtained in studies with various filamentous fungi and yeasts lacking the SteA/Ste12 homolog. The absence of primary sexual structures like peridial hyphae in the ΔsteA mutants of A. benhamiae is in line with results obtained in ΔsteA mutants of A. nidulans where a role for SteA in the early stage of sexual reproduction was suggested (Vallim et al. 2000). StlA, a Ste12 homolog, in the asexual ascomycete Penicillium marneffei, did not affect vegetative growth and asexual reproduction, but was able to complement the defect in sexual development of an A. nidulans ΔsteA mutant (Borneman et al. 2001). The function of Ste12 homologs also appeared to be important in yeast species. For example, in Candida lusitaniae, CLS12 is essential for mating but not for filamentation (Young et al. 2000), whereas disruption of CPH1 in C. albicans led to decreased filamentous growth (Liu et al. 1994; Lo et al. 1997). Both mating and vegetative growth are regulated by Ste12 in S. cerevisiae and its homolog pp-1 in N. crassa (Chou et al. 2006; Hartwell 1980; Li et al. 2005). Similarly, disruption of ste12 in the plant pathogen B. cinerea resulted in a Δste12 mutant with slightly decreased growth and a defect in sexual reproduction (Schamber et al. 2010). Deletion of ste12 in the homothallic ascomycete Sordaria macrospora did not affect fruiting body formation but led to impaired ascus and ascospore development (Nolting and Pöggeler 2006). However, in the plant pathogen Magnaporthe grisea, MST12 is neither involved in vegetative growth, asexual reproduction nor mating (Park et al. 2002). In summary, the function of the transcriptional regulator SteA in vegetative growth, conidiation or sexual reproduction seems to be species-specific, although the role of SteA in sexual reproduction appears to be conserved in some yeasts and filamentous fungi.
A. benhamiae stuA knock-out mutants failed to produce fertile cleistothecia containing asci and ascospores, but were still able to form peridial hyphae. Similarly, stuA deletion mutants of the homothallic fungus A. nidulans failed to form cleistothecia and Hülle cells (nurse cells) which resulted in self-sterility (Clutterbuck 1969; Miller et al. 1991). Abolishment of sexual reproduction has also been demonstrated in Δasm-1 deletion mutants of N. crassa which were unable to form protoperithecia (female organs) (Aramayo et al. 1996). The homolog of StuA is required for mating and sexual organ development in the plant pathogenic fungi Glomerella cingulata and Ustilago maydis (García-Pedrajas et al. 2010; Tong et al. 2007), but it is dispensable for sexual reproduction in M. grisea (Nishimura et al. 2009).
Mutants of A. benhamiae lacking the AbenStuA gene showed retarded growth and produced fewer aerial hyphae during cultivation on solid media. These phenotypes are in line with those observed for inactivation of StuA homologs in the mold N. crassa and in the plant pathogens M. grisea and F. oxysporum (Aramayo et al. 1996; Nishimura et al. 2009; Ohara and Tsuge 2004). Deletion of the stuA gene in P. chrysogenum and A. fumigatus did not affect radial growth (Sheppard et al. 2005; Sigl et al. 2011). A. benhamiae stuA deletion mutants produced microconidia comparable to the wild type, a phenotype also observed in F. oxysporum fostuA deletion mutants (Ohara and Tsuge 2004). Microconidia from A. benhamiae as well as from F. oxysporum are directly released from hyphae. By contrast, inactivation of StuA homologs was found to be deleterious for conidiophore formation and conidiogenesis in other ascomycetes. Deletion of the stuA gene in A. nidulans resulted in the so-called stunted phenotype which is caused by the formation of shortened conidiophores that lack sterigmata, i.e., metulae and phialides. Additionally, stuA deletion mutants of A. nidulans only produced low numbers of conidia which directly bud from the vesicles (Clutterbuck 1969; Miller et al. 1991, 1992). Similar results were obtained in studies with F. oxysporum, A. fumigatus, P. marneffei and A. chrysogenum where deletion of stuA led to the production of abnormal conidiophores (Borneman et al. 2002; Hu et al. 2015; Ohara and Tsuge 2004; Sheppard et al. 2005). A reduced number of conidia was reported for stuA deletion mutants of G. cingulata, M. grisea and A. fumigatus (Nishimura et al. 2009; Sheppard et al. 2005; Tong et al. 2007). Deletion mutants of stuA of P. chrysogenum, Stagonospora nodorum and A. chrysogenum failed to sporulate (Hu et al. 2015; IpCho et al. 2010; Sigl et al. 2011).
The ability of the A. benhamiae ΔsteA mutant to invade and colonize keratinized host structures was analyzed using an in vitro model of human hair and nails. Although ΔsteA mutants were able to grow on both hair and nails, interestingly, deletion of AbenSTEA resulted in loss of hair perforation organs. Vice versa, mutants in AbenSTUA were unable to grow on keratin substrates but formed hair perforations. This observation suggests that perforation organs are dispensable for the destruction and growth on hair, and moreover, the formation of perforation organs and growth per se appear to be differentially regulated processes. Thus, hyphae of the ΔsteA mutants were still able to invade the hair shaft by lifting the cuticle cells and slipping beneath them as it has been described for other dermatophytes which are not able to form perforation organs (Raubitschek and Evron 1963). The ability of dermatophytes to produce hair perforation organs in vitro is a species-specific attribute which was used in the past to distinguish atypical isolates, such as Trichophyton mentagrophytes (perforation positive) from T. rubrum (perforation negative) (Padhye et al. 1980). Due to the development of molecular tools for the identification of dermatophyte species and their phylogenetic relationship, the taxonomy of dermatophytes is subject to constant change (Gräser et al. 2008) and the in vitro hair perforation test became redundant. However, the function of hair perforation organs in dermatophytes remains unknown. Parallels can be drawn between the hair perforation organs produced by A. benhamiae and the development of appressorial penetration pegs by plant pathogens which allow the fungi to enter host cells. Disruption of the Ste12 homologs in the plant pathogens M. grisea, Colletotrichum lagenarium and C. lindemuthianum resulted in the complete loss of penetration peg formation and in the inability to invade host tissue (Park et al. 2002, 2004; Tsuji et al. 2003; Wong Sak Hoi et al. 2007). Decreased virulence was shown for Δste12 mutants in the plant pathogenic fungi Fusarium graminearum, F. oxysporum, B. cinerea, Cryphonectria parasitica and Setosphaeria turcica (Asunción Garcia-Sánchez et al. 2010; Deng et al. 2007; Gu et al. 2014, 2015; Rispail and Di Pietro 2009; Schamber et al. 2010).
Despite the capacity of A. benhamiae stuA deletion mutants to penetrate hair by the production of hair perforation organs, their inability to grow on human hair and nails suggest an eminent role of StuA for keratin degradation and consumption, and hence for the pathogenicity of dermatophytes. StuA homologs have been demonstrated to be important for infection in other pathogens before, e.g., GcStuA, Mstu, SnStuA and Ust1 of the plant pathogens G. cingulata, M. grisea, S. nodorum (ascomycetes) and U. maydis (basidiomycete), respectively. Mutants of M. grisea were impaired and mutants of G. cingulata were unable to penetrate intact plant cells (Nishimura et al. 2009; Tong et al. 2007). Ust1 of U. maydis and StuA of S. nodorum were not required for the penetration stage or the initial steps of infection (García-Pedrajas et al. 2010; IpCho et al. 2010). Nevertheless, deletion of S. nodorum StuA resulted in reduced pathogenicity on intact leaves and Ust1 remains a critical virulence factor as it is important for gall induction (García-Pedrajas et al. 2010; IpCho et al. 2010). By contrast, FoStuA of F. oxysporum and StuA of A. fumigatus are dispensable for pathogenicity (Ohara and Tsuge 2004; Sheppard et al. 2005).
The results of the present work on the important global regulators SteA and StuA involved in developmental processes in fungi, give insights into their role in basic biology and pathogenicity of dermatophytes. Both transcriptional regulators of A. benhamiae are essential for sexual reproduction and, additionally, StuA is involved in the degradation of keratin substrates. Future studies, in particular on StuA and its target genes should further enlighten the host–pathogen interaction in dermatophytes.
References
Ajello L, Cheng SL (1967) The perfect state of Trichophyton mentagrophytes. Sabouraudia 5:230–234
Aramayo R, Peleg Y, Addison R, Metzenberg R (1996) Asm-1 +, a Neurospora crassa gene related to transcriptional regulators of fungal development. Genetics 144:991–1003
Asunción Garcia-Sánchez M, Martín-Rodrigues N, Ramos B, de Vega-Bartol JJ, Perlin MH, Díaz-Mínguez JM (2010) Fost12, the Fusarium oxysporum homolog of the transcription factor Ste12, is upregulated during plant infection and required for virulence. Fungal Genet Biol 47:216–225. doi:10.1016/j.fgb.2009.11.006
Borneman AR, Hynes MJ, Andrianopoulos A (2001) An STE12 homolog from the asexual, dimorphic fungus Penicillium marneffei complements the defect in sexual development of an Aspergillus nidulans steA mutant. Genetics 157:1003–1014
Borneman AR, Hynes MJ, Andrianopoulos A (2002) A basic helix-loop-helix protein with similarity to the fungal morphological regulators, Phd1p, Efg1p and StuA, controls conidiation but not dimorphic growth in Penicillium marneffei. Mol Microbiol 44:621–631
Burmester A, Shelest E, Glöckner G, Heddergott C, Schindler S, Staib P, Heidel A, Felder M, Petzold A, Szafranski K, Feuermann M, Pedruzzi I, Priebe S, Groth M, Winkler R, Li W, Kniemeyer O, Schroeckh V, Hertweck C, Hube B, White TC, Platzer M, Guthke R, Heitman J, Wöstemeyer J, Zipfel PF, Monod M, Brakhage AA (2011) Comparative and functional genomics provide insights into the pathogenicity of dermatophytic fungi. Genome Biol 12:R7. doi:10.1186/gb-2011-12-1-r7
Chou S, Lane S, Liu H (2006) Regulation of mating and filamentation genes by two distinct Ste12 complexes in Saccharomyces cerevisiae. Mol Cell Biol 26:4794–4805. doi:10.1128/mcb.02053-05
Clutterbuck AJ (1969) A mutational analysis of conidial development in Aspergillus nidulans. Genetics 63:317–327
Deng F, Allen TD, Nuss DL (2007) Ste12 transcription factor homologue CpST12 is down-regulated by hypovirus infection and required for virulence and female fertility of the chestnut blight fungus Cryphonectria parasitica. Eukaryot Cell 6:235–244. doi:10.1128/ec.00302-06
Dinamarco TM, Almeida RS, de Castro PA, Brown NA, dos Reis TF, Ramalho LN, Savoldi M, Goldman MH, Goldman GH (2012) Molecular characterization of the putative transcription factor SebA involved in virulence in Aspergillus fumigatus. Eukaryot Cell 11:518–531. doi:10.1128/ec.00016-12
Doedt T, Krishnamurthy S, Bockmühl DP, Tebarth B, Stempel C, Russell CL, Brown AJ, Ernst JF (2004) APSES proteins regulate morphogenesis and metabolism in Candida albicans. Mol Biol Cell 15:3167–3180. doi:10.1091/10.1091/mbc.E03-11-0782
Dutton JR, Johns S, Miller BL (1997) StuAp is a sequence-specific transcription factor that regulates developmental complexity in Aspergillus nidulans. EMBO J 16:5710–5721. doi:10.1093/emboj/16.18.5710
Ejzykowicz DE, Cunha MM, Rozental S, Solis NV, Gravelat FN, Sheppard DC, Filler SG (2009) The Aspergillus fumigatus transcription factor Ace2 governs pigment production, conidiation and virulence. Mol Microbiol 72:155–169. doi:10.1111/j.1365-2958.2009.06631.x
Ernst JF (2000) Transcription factors in Candida albicans—environmental control of morphogenesis. Microbiology 146(Pt 8):1763–1774. doi:10.1099/00221287-146-8-1763
Errede B, Ammerer G (1989) STE12, a protein involved in cell-type-specific transcription and signal transduction in yeast, is part of protein-DNA complexes. Genes Dev 3:1349–1361
Ferreira-Nozawa MS, Silveira HC, Ono CJ, Fachin AL, Rossi A, Martinez-Rossi NM (2006) The pH signaling transcription factor PacC mediates the growth of Trichophyton rubrum on human nail in vitro. Med Mycol 44:641–645. doi:10.1080/13693780600876553
Fumeaux J, Mock M, Ninet B, Jan I, Bontems O, Léchenne B, Lew D, Panizzon RG, Jousson O, Monod M (2004) First report of Arthroderma benhamiae in Switzerland. Dermatology 208:244–250. doi:10.1159/000077311
García-Pedrajas MD, Baeza-Montañez L, Gold SE (2010) Regulation of Ustilago maydis dimorphism, sporulation, and pathogenic development by a transcription factor with a highly conserved APSES domain. Mol Plant Microbe Interact 23:211–222. doi:10.1094/mpmi-23-2-0211
Gavrias V, Andrianopoulos A, Gimeno CJ, Timberlake WE (1996) Saccharomyces cerevisiae TEC1 is required for pseudohyphal growth. Mol Microbiol 19:1255–1263
Gimeno CJ, Fink GR (1994) Induction of pseudohyphal growth by overexpression of PHD1, a Saccharomyces cerevisiae gene related to transcriptional regulators of fungal development. Mol Cell Biol 14:2100–2112
Gräser Y, Scott J, Summerbell R (2008) The new species concept in dermatophytes—a polyphasic approach. Mycopathologia 166:239–256. doi:10.1007/s11046-008-9099-y
Grumbt M, Defaweux V, Mignon B, Monod M, Burmester A, Wöstemeyer J, Staib P (2011a) Targeted gene deletion and in vivo analysis of putative virulence gene function in the pathogenic dermatophyte Arthroderma benhamiae. Eukaryot Cell 10:842–853. doi:10.1128/ec.00273-10
Grumbt M, Monod M, Staib P (2011b) Genetic advances in dermatophytes. FEMS Microbiol Lett 320:79–86. doi:10.1111/j.1574-6968.2011.02276.x
Gu SQ, Li P, Wu M, Hao ZM, Gong XD, Zhang XY, Tian L, Zhang P, Wang Y, Cao ZY, Fan YS, Han JM, Dong JG (2014) StSTE12 is required for the pathogenicity of Setosphaeria turcica by regulating appressorium development and penetration. Microbiol Res 169:817–823. doi:10.1016/j.micres.2014.04.001
Gu Q, Zhang C, Liu X, Ma Z (2015) A transcription factor FgSte12 is required for pathogenicity in Fusarium graminearum. Mol Plant Pathol 16:1–13. doi:10.1111/mpp.12155
Hartwell LH (1980) Mutants of Saccharomyces cerevisiae unresponsive to cell division control by polypeptide mating hormone. J Cell Biol 85:811–822
Hu P, Wang Y, Zhou J, Pan Y, Liu G (2015) AcstuA, which encodes an APSES transcription regulator, is involved in conidiation, cephalosporin biosynthesis and cell wall integrity of Acremonium chrysogenum. Fungal Genet Biol 83:26–40. doi:10.1016/j.fgb.2015.08.003
IpCho SV, Tan KC, Koh G, Gummer J, Oliver RP, Trengove RD, Solomon PS (2010) The transcription factor StuA regulates central carbon metabolism, mycotoxin production, and effector gene expression in the wheat pathogen Stagonospora nodorum. Eukaryot Cell 9:1100–1108. doi:10.1128/ec.00064-10
Li D, Bobrowicz P, Wilkinson HH, Ebbole DJ (2005) A mitogen-activated protein kinase pathway essential for mating and contributing to vegetative growth in Neurospora crassa. Genetics 170:1091–1104. doi:10.1534/genetics.104.036772
Liu H, Köhler J, Fink GR (1994) Suppression of hyphal formation in Candida albicans by mutation of a STE12 homolog. Science 266:1723–1726
Lo HJ, Köhler JR, DiDomenico B, Loebenberg D, Cacciapuoti A, Fink GR (1997) Nonfilamentous C. albicans mutants are avirulent. Cell 90:939–949
Madhani HD, Fink GR (1997) Combinatorial control required for the specificity of yeast MAPK signaling. Science 275:1314–1317
Martinez DA, Oliver BG, Gräser Y, Goldberg JM, Li W, Martinez-Rossi NM, Monod M, Shelest E, Barton RC, Birch E, Brakhage AA, Chen Z, Gurr SJ, Heiman D, Heitman J, Kosti I, Rossi A, Saif S, Samalova M, Saunders CW, Shea T, Summerbell RC, Xu J, Young S, Zeng Q, Birren BW, Cuomo CA, White TC (2012) Comparative genome analysis of Trichophyton rubrum and related dermatophytes reveals candidate genes involved in infection. MBio 3:e00259–12. doi:10.1128/mBio.00259-12
Miller KY, Toennis TM, Adams TH, Miller BL (1991) Isolation and transcriptional characterization of a morphological modifier: the Aspergillus nidulans stunted (stuA) gene. Mol Gen Genet 227:285–292
Miller KY, Wu J, Miller BL (1992) StuA is required for cell pattern formation in Aspergillus. Genes Dev 6:1770–1782
Nishimura M, Fukada J, Moriwaki A, Fujikawa T, Ohashi M, Hibi T, Hayashi N (2009) Mstu1, an APSES transcription factor, is required for appressorium-mediated infection in Magnaporthe grisea. Biosci Biotechnol Biochem 73:1779–1786
Nolting N, Pöggeler S (2006) A STE12 homologue of the homothallic ascomycete Sordaria macrospora interacts with the MADS box protein MCM1 and is required for ascosporogenesis. Mol Microbiol 62:853–868. doi:10.1111/j.1365-2958.2006.05415.x
Ohara T, Tsuge T (2004) FoSTUA, encoding a basic helix-loop-helix protein, differentially regulates development of three kinds of asexual spores, macroconidia, microconidia, and chlamydospores, in the fungal plant pathogen Fusarium oxysporum. Eukaryot Cell 3:1412–1422. doi:10.1128/ec.3.6.1412-1422.2004
Padhye AA, Young CN, Ajello L (1980) Hair perforation as a diagnostic criterion in the identification of Epidermophyton, Microsporum and Trichophyton species. Pan Am Health Org Sci Publ 396:115–120
Park G, Xue C, Zheng L, Lam S, Xu JR (2002) MST12 regulates infectious growth but not appressorium formation in the rice blast fungus Magnaporthe grisea. Mol Plant Microbe Interact 15:183–192. doi:10.1094/mpmi.2002.15.3.183
Park G, Bruno KS, Staiger CJ, Talbot NJ, Xu JR (2004) Independent genetic mechanisms mediate turgor generation and penetration peg formation during plant infection in the rice blast fungus. Mol Microbiol 53:1695–1707. doi:10.1111/j.1365-2958.2004.04220.x
Pasquali M, Spanu F, Scherm B, Balmas V, Hoffmann L, Hammond-Kosack KE, Beyer M, Migheli Q (2013) FcStuA from Fusarium culmorum controls wheat foot and root rot in a toxin dispensable manner. PLoS One 8:e57429. doi:10.1371/journal.pone.0057429
Raubitschek F, Evron R (1963) Experimental invasion of hair by dermatophytes. Arch Dermatol 88:837–845
Rispail N, Di Pietro A (2009) Fusarium oxysporum Ste12 controls invasive growth and virulence downstream of the Fmk1 MAPK cascade. Mol Plant Microbe Interact 22:830–839. doi:10.1094/mpmi-22-7-0830
Schamber A, Leroch M, Diwo J, Mendgen K, Hahn M (2010) The role of mitogen-activated protein (MAP) kinase signalling components and the Ste12 transcription factor in germination and pathogenicity of Botrytis cinerea. Mol Plant Pathol 11:105–119. doi:10.1111/j.1364-3703.2009.00579.x
Schrettl M, Beckmann N, Varga J, Heinekamp T, Jacobsen ID, Jochl C, Moussa TA, Wang S, Gsaller F, Blatzer M, Werner ER, Niermann WC, Brakhage AA, Haas H (2010) HapX-mediated adaption to iron starvation is crucial for virulence of Aspergillus fumigatus. PLoS Pathog 6:e1001124. doi:10.1371/journal.ppat.1001124
Sheppard DC, Doedt T, Chiang LY, Kim HS, Chen D, Nierman WC, Filler SG (2005) The Aspergillus fumigatus StuA protein governs the up-regulation of a discrete transcriptional program during the acquisition of developmental competence. Mol Biol Cell 16:5866–5879. doi:10.1091/mbc.E05-07-0617
Sigl C, Haas H, Specht T, Pfaller K, Kürnsteiner H, Zadra I (2011) Among developmental regulators, StuA but not BrlA is essential for penicillin V production in Penicillium chrysogenum. Appl Environ Microbiol 77:972–982. doi:10.1128/aem.01557-10
Sonneborn A, Bockmühl DP, Ernst JF (1999) Chlamydospore formation in Candida albicans requires the Efg1p morphogenetic regulator. Infect Immun 67:5514–5517
Soyer JL, Hamiot A, Ollivier B, Balesdent MH, Rouxel T, Fudal I (2015) The APSES transcription factor LmStuA is required for sporulation, pathogenic development and effector gene expression in Leptosphaeria maculans. Mol Plant Pathol 16:1000–1005. doi:10.1111/mpp.12249
Staib P, Kretschmar M, Nichterlein T, Hof H, Morschhauser J (2002) Transcriptional regulators Cph1p and Efg1p mediate activation of the Candida albicans virulence gene SAP5 during infection. Infect Immun 70:921–927
Staib P, Zaugg C, Mignon B, Weber J, Grumbt M, Pradervand S, Harshman K, Monod M (2010) Differential gene expression in the pathogenic dermatophyte Arthroderma benhamiae in vitro versus during infection. Microbiology 156:884–895. doi:10.1099/mic.0.033464-0
Stoldt VR, Sonneborn A, Leuker CE, Ernst JF (1997) Efg1p, an essential regulator of morphogenesis of the human pathogen Candida albicans, is a member of a conserved class of bHLH proteins regulating morphogenetic processes in fungi. EMBO J 16:1982–1991. doi:10.1093/emboj/16.8.1982
Symoens F, Jousson O, Planard C, Fratti M, Staib P, Mignon B, Monod M (2011) Molecular analysis and mating behaviour of the Trichophyton mentagrophytes species complex. Int J Med Microbiol 301:260–266. doi:10.1016/j.ijmm.2010.06.001
Symoens F, Jousson O, Packeu A, Fratti M, Staib P, Mignon B, Monod M (2013) The dermatophyte species Arthroderma benhamiae: intraspecies variability and mating behaviour. J Med Microbiol 62:377–385. doi:10.1099/jmm.0.053223-0
Tong X, Zhang X, Plummer KM, Stowell KM, Sullivan PA, Farley PC (2007) GcSTUA, an APSES transcription factor, is required for generation of appressorial turgor pressure and full pathogenicity of Glomerella cingulata. Mol Plant Microbe Interact 20:1102–1111. doi:10.1094/mpmi-20-9-1102
Tsuji G, Fujii S, Tsuge S, Shiraishi T, Kubo Y (2003) The Colletotrichum lagenariu Ste12-like gene CST1 is essential for appressorium penetration. Mol Plant Microbe Interact 16:315–325. doi:10.1094/mpmi.2003.16.4.315
Vallim MA, Miller KY, Miller BL (2000) Aspergillus SteA (sterile12-like) is a homeodomain-C2/H2-Zn+2 finger transcription factor required for sexual reproduction. Mol Microbiol 36:290–301
Ward MP, Gimeno CJ, Fink GR, Garrett S (1995) SOK2 may regulate cyclic AMP-dependent protein kinase-stimulated growth and pseudohyphal development by repressing transcription. Mol Cell Biol 15:6854–6863
Weitzman I, Summerbell RC (1995) The dermatophytes. Clin Microbiol Rev 8:240–259
White TC, Oliver BG, Gräser Y, Henn MR (2008) Generating and testing molecular hypotheses in the dermatophytes. Eukaryot Cell 7:1238–1245. doi:10.1128/ec.00100-08
Wong Sak Hoi J, Dumas B (2010) Ste12 and Ste12-like proteins, fungal transcription factors regulating development and pathogenicity. Eukaryot Cell 9:480–485. doi:10.1128/ec.00333-09
Wong Sak Hoi J, Herbert C, Bacha N, O’Connell R, Lafitte C, Borderies G, Rossignol M, Rougé P, Dumas B (2007) Regulation and role of a STE12-like transcription factor from the plant pathogen Colletotrichum lindemuthianum. Mol Microbiol 64:68–82. doi:10.1111/j.1365-2958.2007.05639.x
Wu J, Miller BL (1997) Aspergillus asexual reproduction and sexual reproduction are differentially affected by transcriptional and translational mechanisms regulating stunted gene expression. Mol Cell Biol 17:6191–6201
Young LY, Lorenz MC, Heitman J (2000) A STE12 homolog is required for mating but dispensable for filamentation in Candida lusitaniae. Genetics 155:17–29
Zaugg C, Monod M, Weber J, Harshman K, Pradervand S, Thomas J, Bueno M, Giddey K, Staib P (2009) Gene expression profiling in the human pathogenic dermatophyte Trichophyton rubrum during growth on proteins. Eukaryot Cell 8:241–250. doi:10.1128/ec.00208-08
Acknowledgments
This work was supported by the DFG funded excellence graduate school Jena School for Microbial Communication (JSMC; GSC 214; www.jsmc.uni-jena.de) and the Leibniz Institute for Natural Product Research and Infection Biology (HKI) (www.leibniz-hki.de).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
The study did not include any diagnostic procedure or therapeutic method. Furthermore, the sample collection was non-invasive (the physical integrity of the donor was maintained) and did not intrude into the privacy of the donor. Based on the regulations of the ethics commission at the Jena University Hospital, Jena (Germany), an approval of the study was not necessary in this case.
Informed consent
Informed consent was obtained from all individual participants included in the study.
Additional information
Communicated by M. Kupiec.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary Table S1: Primers used in this study
294_2016_608_MOESM1_ESM.docx
Supplementary Fig. S1: Multiple sequence alignments of StuA homologs of N. crassa (Asm-1, XP_960837.1), A. nidulans (StuA, AN5836), A. benhamiae (AbenStuA, ARB_07703); C. albicans (Efg1, CR_07890W_A) and S. cerevisiae (Phd1, CAA81878.1; Sok2, AAB35749.1) using EBI Clustal Omega. The APSES domain is shaded in grey (DOCX 17 kb)
Rights and permissions
About this article
Cite this article
Kröber, A., Etzrodt, S., Bach, M. et al. The transcriptional regulators SteA and StuA contribute to keratin degradation and sexual reproduction of the dermatophyte Arthroderma benhamiae . Curr Genet 63, 103–116 (2017). https://doi.org/10.1007/s00294-016-0608-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00294-016-0608-0