Abstract
Background
Odontogenic tumors (OTs) are rare, with an estimated incidence rate of less than 0.5 cases per 100,000 per year. The causes of OTs remain unclear. Nonetheless, the majority of OTs seem to arise de novo, without an apparent causative factor. Although the etiopathogenesis of most OTs remains unclear, there have been some recent advances in understanding the genetic basis relating to specific histologies and clinical features. Molecular analyses performed by different techniques, including Sanger sequencing, next-generation sequencing, and allele-specific PCR, have uncovered mutations in genes related to the oncogenic MAPK/ERK signaling pathway. Genetic mutations in these pathway genes have been reported in epithelial and mixed OTs, in addition to odontogenic carcinomas and sarcomas. Notably, B‑RAF proto-oncogene serine/threonine kinase (BRAF) and KRAS proto-oncogene GTPase (KRAS) pathogenic mutations have been reported in a high proportion of ameloblastoma and ameloblastoma-related tumors and adenomatoid odontogenic tumors, respectively.
Objective
To discuss how molecular profiling aids in diagnostic classification of odontogenic tumors.
Conclusion
Molecular profiling of odontogenic tumors helps to identify patients for neoadjuvant therapies and reduces postoperative morbidity
Zusammenfassung
Hintergrund
Odontogene Tumoren (OT) sind seltene Tumoren, und die geschätzte Inzidenzrate liegt bei weniger als 0,5 Fällen pro 100.000 pro Jahr. Die Ursachen für OT sind nach wie vor unklar. Die meisten OT scheinen jedoch de novo zu entstehen, ohne dass es einen offensichtlichen ursächlichen Faktor gibt. Obwohl die Ätiopathogenese der meisten odontogenen Tumoren nach wie vor unklar ist, gab es in jüngster Zeit einige Fortschritte beim Verständnis der genetischen Grundlagen bestimmter odontogener Tumoren. Molekulare Analysen mit verschiedenen Techniken, darunter Sanger-Sequenzierung, Next-Generation-Sequenzierung und allelspezifische Polymerasekettenreaktion (PCR), haben Mutationen in Genen aufgedeckt, die mit dem onkogenen MAPK/ERK-Signalweg in odontogenen Tumoren in Verbindung stehen. Genetische Mutationen in diesen Signalweg-Genen wurden bei epithelialen und gemischten odontogenen Tumoren sowie bei odontogenen Karzinomen und Sarkomen festgestellt. Insbesondere die Protoonkogene BRAF, das für die Serin-Threonin-Kinase B‑RAF kodiert, und KRAS, das für die GTPase KRAS kodiert, wurden in einem hohen Anteil von Ameloblastomen und mit Ameloblastomen verwandten Tumoren bzw. adenomatoiden odontogenen Tumoren nachgewiesen.
Ziel
Erörtert wird die Frage, wie die molekulare Typisierung bei der diagnostischen Klassifizierung von odontogenen Tumoren hilft.
Schlussfolgerung
Die molekulare Typisierung bei odontogenen Tumoren hilft bei der Identifizierung von Patienten für neoadjuvante Therapien, präzisiert die histopathologische Klassifikation und vermindert postoperative Morbidität.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Odontogenic tumors (OTs) comprise a group of heterogeneous lesions ranging from hamartomatous lesions to malignant neoplasms with different behavior, histology, and even different geographical distribution [1]. The etiopathogenesis of most OTs remains unclear; however, there have been some recent advances in understanding the genetic basis of specific OTs [2]. Detection of genetic factors that are involved in the molecular pathogenesis of OTs helps us in targeted therapy [3]. Herein, we highlight the molecular profiling of OTs and provide evidence for the clinical utility of targeted therapies.
Odontogenesis and odontogenic tumors
Tooth development (odontogenesis) is initiated by interactions between epithelial and mesenchymal cells derived from the ectoderm of the first branchial arch and the ectomesenchyme of the neural crest. Odontogenesis involves several morphologically distinct stages. Reciprocal signaling between epithelium and ectomesenchyme guides the process of tooth embryonic development, which is fully dependent on Wnt, BMP, FGF, Shh, and Eda signals [4]. The pathogenesis of odontogenic tumors is associated with alterations in components of signaling pathways. For instance, studies in the last decade have described pathogenic mutations in mitogen-activated protein kinases/extracellular signal-regulated kinases (MAPK/ERK) pathway cascade components in benign and malignant odontogenic tumors [2, 5].
Benign odontogenic tumors
The OT classification is mainly divided into two categories, based on biological behavior, as malignant and benign. Benign tumors are classified into three major categories according to their histogenetic origin: epithelial, mesenchymal, and mixed types [2]. OTs may pose both diagnostic and prognostic challenges due to overlapping histology and a high propensity for local recurrence, even though they are considered benign [1].
Epithelial tumors
Ameloblastoma
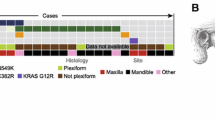
Ameloblastoma (AM) is the most common benign epithelial odontogenic tumor, representing approximately 1% of all oral tumors and about 9 to 11% of all odontogenic tumors [1]. The tumors are classified into four groups: conventional, unicystic, extraosseous/peripheral, and metastasizing variants. In the recent World Health Organization (WHO) classification, adenoid ameloblastoma (AdAM) is a newly recognized entity separate from the AM group of tumors [2]. Although AMs are known as locally aggressive tumors with high recurrence rates, unicystic ameloblastoma (UAM) shows an indolent course different from the other variants. This unpredictable and distinct biological behavior of AMs has lent them priority to trigger molecular studies on understanding their pathogenesis. In the past decade, oncogenic mutations were discovered which constitutively activate signal transduction pathways relating to developmental stages of odontogenesis, including the mitogen-activated protein kinase (MAPK) and hedgehog pathways [6,7,8]. Advanced next-generation sequencing (NGS) analyses identified the high frequency of BRAF V600E and SMO L412F mutations in all types of ameloblastoma [5,6,7,8]. This is followed by KRAS (mostly p.G12R), NRAS, HRAS, FGFR2, and mutations reported in a few BRAF wildtype cases ([7,8,9]; Fig. 1). Additionally, we reported EGFR mutations and the presence of other gene mutations, including somatic mutations in KRAS, PIK3CA, PTEN, FGFR, CDKN2A, and CTNNB1 on the background of either BRAF or SMO mutation-positive ameloblastomas, occurring exclusively in conventional AMs [9].
In line with reports about other neoplasms that harbor a malignant counterpart, the frequency of the BRAF p.V600E mutation is higher in ameloblastoma (64% in conventional, 81% in unicystic, and 63% in peripheral) than in ameloblastic carcinoma (35%) [5]. As both conventional AM and UAM have been found to harbor BRAF p.V600E mutations, aggressive and destructive tumors could be candidates for BRAF-targeted therapy that has the potential to reduce tumor size and ultimately enable a conservative surgical procedure. Preliminary data of biological treatment show effectiveness in selected cases [2, 3].
Adenoid ameloblastoma
Adenoid ameloblastoma (AdAM) is a newly recognized entity separate from the AM group of tumors. AdAM is characterized by an aggressive biological behavior with local infiltration, and the recurrence rate is high (45.5–70%). BRAF p.V600E mutations, usually identified in AM/UAM, are absent in AdAM. Whether AdAM is a unique standalone tumor or a histologic variant of AM requires further investigation [2].
Adenomatoid odontogenic tumor
Adenomatoid odontogenic tumor (AOT) manifests clinically as a slow and self-limiting growth which does not require an aggressive surgical approach [1]. AOTs are characterized by frequent KRAS codon 12 (either p.G12V or p.G12R, and in a single case p.G12D) driver mutations, which occur in approximately 70% of cases [2, 5, 10]. Although they have not been connected to their clinicopathological features, molecular profiling is important for the differential diagnosis of this tumor from other lesions such as AdAM, adenomatoid odontogenic hamartoma, and adenomatoid dentinoma ([2, 5]; Fig. 2).
Calcifying epithelial odontogenic tumor
Calcifying epithelial odontogenic tumor (CEOT) is recognized to have three histopathological subtypes: clear cell, cystic/microcystic, and non-calcified/Langerhans cell rich. Mutations in tumor suppressor genes (PTEN, CDKN2A, PTCH1) and oncogenes (JAK3, MET) have been identified in CEOT; however, so far, these do not contribute to clinical properties or treatment decisions ([2, 5]; Fig. 2).
Mixed tumors
Odontoma
Odontoma is the most common odontogenic tumor and is composed of mesenchymal and epithelial components of the tooth [1]. WNT/beta-catenin pathway activation in embryonic SOX-2-positive dental stem cells can drive odontoma formation [2]. Ameloblastic fibrodentinoma (AFD) and ameloblastic fibroodontoma (AFO) are classified as developing odontomas, although the prevalence of BRAF p.V600E mutations in AFD and AFO is similar to ameloblastic fibroma (AF) but differs from odontoma, which lacks BRAF p.V600E mutations [11].
Ameloblastic fibroma
Ameloblastic fibroma (AF) is a rare benign odontogenic tumor with the potential for recurrence and malignant transformation to ameloblastic fibrosarcoma ([1]; Fig. 2). AFs are characterized by BRAF p.V600E mutations, like other ameloblastic tumors [2, 10, 11]. Early developing stages of odontomas may be comprised of soft tissue closely resembling dental papilla, with prominent epithelial strands and limited or no evidence of dental hard tissue induction. These features overlap with ameloblastic fibroma (AF), sometimes causing a problem differentiating between them. The differentiation between early odontoma and AF is important to avoid unnecessary potentially destructive surgery [1, 2]. Thus, detection of BRAF p.V600E mutations is important for differential diagnosis.
Mesenchymal tumors
Odontogenic myxoma
Odontogenic myxoma (OM) is a rare odontogenic tumor that arises from odontogenic ectomesenchyme. The tumor often behaves in a locally aggressive and infiltrating fashion, with a 25% recurrence rate [1]. Activating mutations in the MAPK/ERK signaling pathway have been identified in this tumor and may serve as targets for pharmacologic therapy.
Cemento-ossifying fibroma
Cemento-ossifying fibroma (COsF) became an integral part of the benign mesenchymal odontogenic tumors in the 2022 WHO classification. A minority of COsFs are linked to inactivating mutations in the tumor suppressor gene CDC73 (HRPT2), especially in those cases that are part of hyperparathyroidism–jaw tumor syndrome. COsF can also be part of gnathodiaphyseal dysplasia, which is characterized by GDD1 gene mutations [2, 11].
Malignant odontogenic tumors
Malignant odontogenic tumors (MOTs) are extremely rare tumors which arise either de novo or from the malignant transformation of benign odontogenic tumors. They can occur as either carcinomas or sarcomas [1, 5]. In recent studies, malignant odontogenic tumors have also been included in the spectrum of MAPK pathway-driven tumors ([5, 11]; Fig. 3).
Entities
Ameloblastic carcinoma
Ameloblastic carcinoma (AMCa) is a highly aggressive, malignant epithelial odontogenic tumor. It is now defined as an entity which is not related to ameloblastoma [2]. However, AMCas harbor BRAF p.V600E mutations like other ameloblastoma-related tumors, with varying prevalence from 25 to 40% [5]. Mutations in other genes which are not related to MAPK/ERK, such as TP53, CTNNB1, and APC, have also been reported inAMCas [2, 5, 11].
Clear cell odontogenic carcinoma
Clear cell odontogenic carcinoma (COdC) is a malignant tumor with high recurrence rate (40%). Its regional lymph node metastases are more common than distant ones and the death rate is about 11%. Differential diagnosis can be critical, which includes jawbone clear cell-containing tumors such as CEOT, amyloid-rich odontogenic fibroma, odontogenic carcinoma with dentinoid, primary or metastatic tumors of salivary glands (e.g., mucoepidermoid carcinoma, clear cell carcinoma, epithelial myoepithelial carcinoma), and metastatic tumors (i.e., clear cell renal carcinoma, melanoma). COdC is characterized by EWSR1 gene rearrangement in about 80% of cases. It has also been shown to harbor BRAF p.V600E in limited cases [2, 5].
Odontogenic sarcoma
Odontogenic sarcoma (OS) is a mixed tumor, histologically characterized by a benign ameloblastic epithelium within a sarcomatous mesenchymal component, with or without dentine and enamel. OS can arise de novo or emerge from a sarcomatous change in AF, and approximately one third of AFS (Ameloblastic Fibrosarcoma) cases stem from a recrudescent AF. Clinically, OS shows locally aggressive behavior. The tumor shows a high recurrence rate of approximately 37% [12]. The BRAF p.V600E mutation has been detected in 67–71% of reported cases. An NRAS mutation has also been reported in one case of OS in a mutually exclusive manner with BRAF p.V600E [5, 11]. Although the rarity of this tumor precludes extensive knowledge about its molecular pathology, the current results could support the role of the MAPK/ERK pathway in its pathogenesis and pave the way for further investigations on targeted therapy [5].
Practical conclusion
-
Molecular profiling of BRAF, SMO, KRAS, and bCAT dissects most odontogenic tumors into three groups.
-
Molecular profiling helps to identify patients for neoadjuvant therapies and saves postoperative morbidity.
-
A practical approach could be to stain BRAF p.V600E by immunohistochemistry and apply other markers dependent on morphology.
-
In the rare cases of malignant odontogenic tumors, reference pathology is highly recommended. Be aware of clear cell odontogenic carcinoma.
References
Santosh ARB, Ogle OE (2020) Odontogenic tumors. Dent Clin North Am 64(1):121–138
Vered M, Wright JM (2022) Update from the 5th edition of the World Health Organization classification of head and neck tumors: Odontogenic and maxillofacial bone tumors. Head Neck Pathol 16(1):63–75
González-González R, López-Verdín S, Lavalle-Carrasco J, Molina-Frechero N, Isiordia-Espinoza M, Carreón-Burciaga RG et al (2020) Current concepts in ameloblastoma-targeted therapies in B‑raf proto-oncogene serine/threonine kinase V600E mutation: systematic review. World J Clin Oncol 11:31–42
Hermans F, Hemeryck L, Lambrichts I, Bronckaers A, Vankelecom H (2021) Intertwined signaling pathways governing tooth development: a give-and-take between canonical Wnt and Shh. Front Cell Dev Biol 29(9):758–770
Guimarães LM, Coura BP, Gomez RS, Gomes CC (2021) The molecular pathology of odontogenic tumors: Expanding the spectrum of MAPK pathway driven tumors. Front Oral Health 14(2):740–788
Kurppa KJ, Caton J, Morgan RP, Ristimaki A, Ruhin B, Kelloski J et al (2014) High frequency of BRAFV600E mutations in amelobastoma. J Pathol 2232:492–498
Brown NA, Rolland D, McHugh J, Weigelin HC, Zhao L, Lim M et al (2014) Activating FGFR2-RAS-BRAF mutations in ameloblastoma. Clin Cancer Res 20(21):5517–5552
Sweeney RT, McClary AC, Myers BR, Biscocho J, Neahring L, Kwei KA, Qu K, Gong X, Ng T, Jones CD, Varma S, Odegaard JI, Sugiyama T, Koyota S, Rubin BP, Troxell ML, Pelham RJ, Zehnder JL, Beachy PA, Pollack JR, West RB (2014) Identification of recurrent SMO and BRAF mutations in ameloblastomas. Nat Genet 46(7):722–725
Gültekin SE, Aziz R, Heydt C et al (2018) The landscape of genetic alterations in ameloblastomas relates to clinical features. Virchows Arch 472:807–814
Gültekin SE, Sengüven B, Aziz R, Heydt C, Büttner R (2017) Molecular profiling of odontogenic tumors pilot study. Balkan J Dent Med 21:112–115
Coura BP, Bernardes VF, de Sousa SF, Diniz MG, Moreira RG, de Andrade BAB, Romañach MJ, Pontes HAR, Gomez RS, Odell EW, Gomes CC (2020) Targeted next-generation sequencing and allelespecific quantitative pcr of laser capture microdissected samples uncover molecular differences in mixed odontogenic tumors. J Mol Diagn 22:1393–1399
Ramani P, Krishnan RP, Karunagaran M, Muthusekhar MR (2020) Odontogenic sarcoma: First report after new who nomenclature with systematic review. J Oral Maxillofac Pathol 24(1):157–163
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
R. Buettner and S.E. Gültekin declare that they have no competing interests.
For this article no studies with human participants or animals were performed by any of the authors. All studies mentioned were in accordance with the ethical standards indicated in each case.
The supplement containing this article is not sponsored by industry.
Additional information

Scan QR code & read article online
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Buettner, R., Gültekin, S.E. Molecular diagnostics in odontogenic tumors. Pathologie 43 (Suppl 1), 81–85 (2022). https://doi.org/10.1007/s00292-022-01152-7
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00292-022-01152-7