Abstract
Complex dynamical systems are often governed by equations containing many unknown parameters whose precise values may or may not be important for the system’s dynamics. In particular, for chemical and biochemical systems, there may be some reactions or subsystems that are inessential to understanding the bifurcation structure and consequent behavior of a model, such as oscillations, multistationarity and patterning. Due to the size, complexity and parametric uncertainties of many (bio)chemical models, a dynamics-preserving reduction scheme that is able to isolate the necessary contributors to particular dynamical behaviors would be useful. In this contribution, we describe model reduction methods for mass-action (bio)chemical models based on the preservation of instability-generating subnetworks known as critical fragments. These methods focus on structural conditions for instabilities and so are parameter-independent. We apply these results to an existing model for the control of the synthesis of the NO-detoxifying enzyme Hmp in Escherichia coli that displays bistability.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
(Bio)chemical systems are (often large) dynamical systems that may exhibit complex behaviors such as oscillations, patterning and multistationarity. This dynamical potential of biochemical systems, including coupled gene expression components, is critical to a great deal of biological function, notably in development where, to name just a few examples: multistationarity is the dynamical basis for pluripotency (Delbrück 1949; Laurent and Kellershohn 1999; Thomas 1994; Thomas and Kaufman 2001); Turing patterns have often been invoked as explanations for developmental patterning (Turing 1952; Lacalli and Harrison 1978; Koch and Meinhardt 1994; Nakamura et al. 2006; Müller et al. 2012; Raspopovic et al. 2014); and oscillations underlie the development of somites in vertebrates (Dale and Pourquié 2000; Kærn et al. 2000; Takashima et al. 2011). These nontrivial dynamics arise from bifurcations. In ordinary differential equation (ODE) models, bifurcations are predicted by analyzing a characteristic polynomial with coefficients whose values depend on system parameters. A typical goal of a modeling study is to understand how changes in the parameters, perhaps due to events within a cell, lead to bifurcations, i.e. to changes in the behavior of a cell or, at higher levels of organization, of a tissue. For small models, this kind of analysis can sometimes be carried out by hand, and computer algebra systems facilitate the analysis of models of moderate size. However, purely analytic approaches rapidly exceed typical computer capacities as increasingly complex models are considered. To circumvent these problems and facilitate the discovery of models with given properties as well as the search for parameter ranges in which a given behavior occurs, graphical methods have been developed that transform the problem of analyzing a characteristic polynomial into one of studying the properties of graphs and their subgraphs (Clarke 1974; Ivanova 1979b; Schlosser and Feinberg 1994; Ermakov 2003a; Goldstein et al. 2004; Craciun and Feinberg 2006b; Mincheva and Roussel 2007a; Mincheva 2011; Soliman 2013; Banaji and Pantea 2016; Kaltenbach 2020), or the closely related approach of analyzing matrices derived from the structure of a network (Thomas and Kaufman 2001; Mincheva and Roussel 2007a; Steuer et al. 2006; Blanchini et al. 2014; Culos et al. 2016; Vassena and Stadler 2024). In cases of moderate complexity, the necessary analyses can be carried out with pencil and paper, but they can also be automated (Soranzo and Altafini 2009; Donnell et al. 2014; Walther et al. 2014; Feinberg et al. 2018; Yordanov et al. 2020). Typically, these methods extract the ‘necessary ingredients’ (chemical species and interactions) for a particular behavior to emerge in a model. This opens up the interesting possibility of obtaining reduced models by pruning away unnecessary species and reactions, an idea we explored briefly elsewhere (Roussel and Roussel 2018). We return to this problem in this paper, where we first develop model reduction rules based on the bipartite representation of a chemical reaction mechanism originally introduced for qualitative stability analysis by Ivanova (1979b) and Mincheva and Roussel (2007a). The basic idea underlying these rules is the preservation of critical fragments, which are subnetworks responsible for instabilities arising via bifurcations.
In an age of fast computers allowing for efficient simulation of large models, it may be asked why we would still want model reduction methods. In biochemistry in particular, it is often the case that many aspects of a model are uncertain, including some of the reactions. Even when we know the molecular interactions in a system, we sometimes lack information about the stoichiometry of a particular reaction. The ability first to detect the essential elements of a model and then to offer a reduced model is a way of helping the research community focus on the key interactions in a system, which can then be subjected to greater experimental and theoretical scrutiny.
Model reduction methods for chemical and biochemical systems have recently been reviewed by Snowden and coworkers (2017) and by Gorban et al. (2004a,2018). Timescale-based methods are probably the most commonly applied approaches for reducing models of chemical and biochemical systems (Briggs and Haldane 1925; Schauer and Heinrich 1983; Battelli and Lazzari 1985; Fraser 1988; Segel and Slemrod 1989; Maas and Pope 1992; Lam 1993; Borghans et al. 1996; Stiefenhofer 1998; Gorban et al. 2004b; Lee and Othmer 2010; Noethen and Walcher 2011; Kristiansen et al. 2014; Prescott and Papachristodoulou 2014; Goeke et al. 2017; Eilertsen and Schnell 2020; Golda et al. 2020). Typically in these methods, using the terminology of Haken (1996), ‘fast’ variables that are ‘slaved’ to the slow variables are eliminated. A number of model-reduction methods that do not explicitly invoke timescale separation such as optimization-based methods (Petzold and Zhu 1999), techniques using sensitivity analysis to eliminate chemical species or reactions (Tomlin et al. 1992) and variable aggregation (‘lumping’) (Wei and Kuo 1969; Kuo and Wei 1969; Coxson and Bischoff 1987; Li et al. 1993, 1994; Farkas 1999; Huang et al. 2005; Stagni et al. 2014; Okeke and Roussel 2015) also exist. Although some of the methods listed can be applied symbolically, thus without explicit parameter values, their validity typically depends on the parameter range in which a system operates. Exact lumping (Wei and Kuo 1969; Iwasa et al. 1987; Li and Rabitz 1989; Li et al. 1994; Conzelmann et al. 2008), when it can be applied, is an interesting exception as it generates a reduced model that is exactly equivalent to the original model over some range of timescales using dynamical invariants of the governing equations. In principle, lumping does not require parameter values, although lumping schemes for a given mechanism may not be unique, and different lumping schemes eliminate different sets of timescales (Wei and Kuo 1969).
While graph-theoretical methods only provide necessary conditions for specific dynamical behaviors, they have one great advantage over most conventional approaches: the necessary conditions for bifurcations can be decided in the absence of parameter values. This is particularly helpful in analyzing biochemical systems whose parameter values are typically highly uncertain and often unknown (Radulescu et al. 2015; Lubitz and Liebermeister 2019). If we think of the task of model reduction as one in which we wish to simplify a model while retaining its capacity for particular behaviors, then one can immediately see how parameter-free methods would be especially useful in biochemical modeling. Radulescu and coworkers have also developed methods for model reduction in the face of significant parameter uncertainty, although their method requires at least order-of-magnitude estimates of the parameter values (Radulescu et al. 2008, 2012, 2015).
There has been at least one other attempt to reduce models based on a bipartite graph representation (Gay et al. 2010). This method uses two operations to generate simplified models, namely merging vertices and deleting them. However, it is not clear whether these transformations retain the original model’s dynamics. The idea of preserving dynamics through a model simplification process also has a precedent in the work of Danø et al. (2006). Their method was based on the preservation of a normal form as a model was reduced using a combination of lumping and of fixing the values of a subset of the variables. The implementation of this method requires a time series to which the model outputs are matched. In a related approach, Apri et al. (2012) matched solutions of a model with solutions obtained by perturbing the parameters, eliminating reactions where setting the corresponding rate constant to zero could be accommodated within a desired error by adjusting the other parameters. We also mention the method of Rao et al. (2014), which is based on eliminating Horn–Jackson ‘complexes’ [combinations of chemical species appearing together on one side of a chemical reaction arrow (Horn and Jackson 1972)]. This method simplifies the Laplacian matrix, an alternative description of the structure of a chemical system. In principle, the method of Rao et al. (2014) preserves dynamics, although the examples in their paper displayed only trivial dynamics (unique, stable steady state).
In order to prepare the reader, we present in Sect. 2 some basic ideas associated with bipartite graphs and critical fragments, the central players in the theory. This section is supplemented by appendices containing a glossary (Appendix A) and the analysis of a simple example (Appendix B) for readers unfamiliar with the Ivanova approach to qualitative stability analysis. In Sect. 3, we define terms to be used in our graph-based model reduction methods. Section 4 presents the main ideas of the paper in a non-technical way. Then, we prove the theorems that enable these reductions in Sect. 5.1. We follow this up with a discussion, in Sect. 5.2, of the biochemical interpretation of the reduction operations described in the theorems of Sect. 5.1. The reductions contemplated in Sect. 5 are conservative, in the sense that they provably retain the potential for bifurcations of the original model without introducing the potential for new bifurcations. The conditions on which the reductions are based are necessary but not sufficient for Andronov–Hopf and saddle-node bifurcations, so we cannot guarantee that bifurcations will be preserved, but in practice we find that they almost always are. Since preserving bifurcations also retains the qualitative dynamics to either side of a bifurcation point, as a rule conservative reductions will also be dynamics-preserving.
An example of moderate complexity is treated in detail in Sect. 6. Specifically, we apply our reduction method to a model for the control of the synthesis of the bacterial NO detoxifying enzyme Hmp (Roussel 2019a). This model displays bistability over a wide range of model parameters. We start our analysis by applying all feasible conservative reductions. We then discuss, in Sect. 6.4, aggressive reductions enabled by knowledge of a critical fragment responsible for a particular instability of interest. This class of reductions follows the same general principles as the conservative reductions, but violate some of the conditions of the theorems on which the conservative reductions rely. Knowledge of the critical fragment(s) makes these reductions plausible, if less certain in their application.
The paper concludes with a discussion of the advantages, limitations and possible extensions of this work.
2 Bipartite graphs and critical fragments
In this section, we describe the relationship between the characteristic polynomial, bipartite graphs of mass-action mechanisms and their fragments, and stability analysis. While the necessary terminology is presented in the text below, a glossary is also included in Appendix A for rapid reference.
As noted above, graph-theoretical methods can help identify the key mechanistic contributors to biologically interesting behaviors of a system. The methods we favor are based on a bipartite graph representation of mass-action systems consisting of two distinct sets of vertices, one representing the species and the other the reactions of the system, and of directed edges that point from reactants to reactions and from reactions to products (Vol’pert 1972; Ivanova 1979b; Zeigarnik and Temkin 1994; Mincheva and Roussel 2007a). Examples of bipartite graphs for mass-action models can be seen in Figs. 9 and 19, where ovals represent chemical species and rectangles denote reaction vertices.
The characteristic polynomial of a mass-action system, which governs its linear stability, can be written
where m is the number of species involved in the system and n is the number of independent species, also known as the rank of the model.Footnote 1 For mass-action systems, each of the \(c_k\) can be written in the form
where \({\mathcal {S}}_k\) is the set of all fragments of order k (defined below), \(S_k\) denotes a particular fragment of order k, \(K_{S_k}\) is a numerical coefficient, and \({\mathcal {J}}_{S_k}\) is a monomial. Fragments are subsets of the bipartite graph that graphically represent and are in a 1:1 correspondence with each of the terms (\({\mathcal {J}}_{S_k}\)) of the characteristic polynomial (Ivanova 1979b; Mincheva and Roussel 2007a). A fragment is a union of subgraphs sharing a set of vertices. Each subgraph in turn consists of a union of cycles and edges in which each species vertex is at the origin of exactly one reactant-to-reaction edge or of one path in a cycle. Paths are reactant-reaction-species sequences with their connecting edges, with positive paths encoding reactant-product relationships, and negative paths encoding co-reactant relationships. A cycle is composed of paths and, as the name suggests, can be traversed from a given vertex, returning to the initial vertex after following some number of paths without visiting the same reactant vertex twice. A positive cycle contains an even number of negative paths, while a negative cycle contains an odd number of negative paths.Footnote 2
Each of the monomials appearing in a particular characteristic polynomial coefficient \(c_k\) can be written
where \(A_z\) is the concentration of chemical species z, and \(R_u\) is the rate of elementary reaction u. The concentrations and reaction rates appearing in the products on the right-hand side of Eq. (3) are those corresponding to, respectively, the chemical species and reactions in fragment \(S_k\). Because a chemical species can appear at the origin of only one path or edge in a subgraph, k different chemical species appear in a fragment of order k, and thus k distinct concentrations appear in Eq. (3). However, the k reaction rates in \({\mathcal {J}}_{S_k}\) are not necessarily distinct. When it is necessary to designate a particular fragment, we write
where \(\{z_i\}\) is a set of distinct indices and \(\{u_i\}\) is a set of indices, not necessarily distinct.
The coefficient of a fragment, \(K_{S_k}\), is given by
where
In these equations,Footnote 3g labels subgraphs, C labels cycles, n-paths are negative paths, p-paths are positive paths, and \(\alpha _{jk}\) and \(\beta _{ji}\) refer to stoichiometric coefficients, respectively of reactants and products. Equation (5) gives a fragment’s coefficient as the sum of its subgraphs’ coefficients, \(K_g\), which are given in turn by Eqs. (6). When the coefficient of a fragment is negative, we call it a critical fragment. Because only positive cycles have negative values of \(K_C\) and all other contributors to \(K_{S_k}\) are positive, and taking into account the product over cycles in Eq. (6a), a critical fragment must have at least one subgraph with an odd number of positive cycles (Mincheva and Roussel 2007a).
The appearance of a critical fragment in the bipartite graph of a chemical mechanism, i.e. of a negative term in the characteristic polynomial, is necessary for certain instabilities to appear. In particular, we have the following two theorems (Mincheva and Roussel 2007a; Conradi and Mincheva 2017):
-
A critical fragment of order equal to the number of independent species, n, is necessary for the possibility of a saddle-node bifurcation, which would allow for multistationarity.Footnote 4
-
A critical fragment of order less than n is necessary for oscillations due to positive feedback arising via an Andronov-Hopf bifurcation.Footnote 5 In the simplest case, positive feedback arises from autocatalysis which is well known to be a potential source of oscillatory behavior (Prigogine and Lefever 1968; Tyson 1975). The negative feedback case is more complex (Mincheva 2011).
The use of these theorems in qualitative stability analysis is illustrated in an example presented in Appendix B.
The theorems described above provide powerful tools for understanding the dynamics of a chemical network as they enable a determination of the potential for multistationarity or positive-feedback oscillations based on the appearance (or lack thereof) of critical fragments within a mass-action model (Ivanova 1979a, b; Ermakov and Goldstein 2002; Ermakov 2003a, b; Goldstein et al. 2004; Mincheva and Roussel 2007a). It is important to note that these are necessary and not sufficient conditions. Given that these conditions are necessary for the corresponding bifurcations, any model reduction method that would preserve these particular dynamics must preserve the critical fragments of the model. Conversely, a model simplification must not create new critical fragments lest this transformation introduce new dynamical behavior.
Examples of linked cycles. a Linked cycles sharing a single vertex. b Linked cycles sharing multiple vertices. See also Fig. 3
Before we delve into the use of Ivanova’s qualitative stability theory for model reduction, key terms and notations are defined in the next section.
Examples of integrated isolated cycles. a An integrated isolated cycle with a single entry vertex A\(_1\). b Two separate integrated isolated cycles. The cycle on the left has the species vertex A\(_1\) as its entry vertex where the cycle on the right has the species vertex A\(_6\) as its entry vertex. Both of these cycles are considered isolated cycles since none of their vertices are part of other cycles
3 Notation and definitions
This section provides new terminology and notation useful for describing model reduction operations for bipartite graphs.
- Product edge::
-
A directed arrow from a reaction vertex R\(_k\) to a product species vertex A\(_i\), denoted [R\(_k\),A\(_i\)]. Note that in Ivanova’s original terminology, edges only refer to reactant-to-reaction edges (Ivanova 1979b; Mincheva and Roussel 2007a).
- Isolated cycle:
-
A cycle \(C_{i}\) whose vertices are not involved in any other cycle.
- Linked cycle:
-
A cycle \(C_{L}\) where at least one vertex in \(C_{L}\) is part of another cycle. Figure 1 shows examples of linked cycles.
- Unidirectional cycle:
-
A cycle that contains at least one positive path. As a consequence of the positive path, unidirectional cycles can only be traversed in one direction. Note that only unidirectional cycles can be isolated since traversals of a cycle in opposite directions are considered to be different cycles.
- Entry vertex:
-
A vertex within a cycle C that contains at least one arrow connected to it that is not part of C.
- Isolated edge:
-
An edge whose vertices are not entry vertices.
- Integrated cycle:
-
A cycle with one or more entry vertices. See Fig. 2 for examples of isolated integrated cycles. Note that linked cycles are necessarily integrated.
- Unitary edge:
-
An edge (reactant or product) whose weight is one.
- Unitary (positive/negative) path:
-
A path (positive or negative) consisting of unitary edges.
- Unitary cycle:
-
A cycle consisting entirely of unitary paths.
Recall that the weight of an edge is the stoichiometric coefficient of the chemical species participating in that edge. For simplicity, we use the same symbol below for reactant and product stoichiometric coefficients, namely \(a_h\).
-
\([\textrm{A}_k,\textrm{R}_j,\textrm{A}_i]_{a_h, a_{h+1}}\) represents the positive path [A\(_k\),R\(_j\),A\(_i\)] with arrows of weight \(a_h\) leaving A\(_k\) and weight \(a_{h+1}\) leaving R\(_j\).
-
\(\overline{[\text {A}_k,\text {R}_j,\text {A}_i]}_{a_h, a_{h+1}}\) represents the negative path \(\overline{[\text {A}_k,\text {R}_j,\text {A}_i]}\) with arrows of weight \(a_h\) leaving A\(_k\) and weight \(a_{h+1}\) leaving A\(_i\).
4 Reducing (bio)chemical systems
4.1 A graphical approach based on the analysis of critical fragments
As mentioned previously, we will use critical fragment theory as a basis for model reduction. The basic concept is simple: since critical fragments are necessary for behaviors such as oscillations due to positive feedback and multistationarity, then preservation of these fragments is itself necessary (but perhaps not sufficient) to preserve qualitative dynamics, i.e. bifurcations separating parameter regions where distinct dynamical behaviors are observed.
According to Eq. (5), a fragment’s coefficient is a sum of subgraph coefficients. Changing the number of subgraphs in a fragment may thus change the coefficient of a fragment, and could in principle change its sign. As a rule, model reductions should therefore avoid creating or destroying subgraphs. The number of subgraphs that contribute to a fragment depends in large part on the number of cycles that can be built from the fragment.Footnote 6 Thus, it will be essential both to conserve cycles and to avoid creating new ones. Coefficients of subgraphs in turn depend on the number and parities of cycles they include. Again, this highlights the importance of preserving existing cycles and avoiding the creation of new cycles. Moreover, if any of the stoichiometric coefficients differ from unity, these will affect the coefficient of a subgraph and may change the sign of the fragment. Thus, we will need to preserve non-unitary stoichiometric coefficients appropriately in model reduction operations.
4.2 Unidirectional isolated cycles
Reductions can be applied most simply to cycles whose vertices are not shared with other cycles, i.e. to the isolated cycles defined above. Additionally, in order to apply conservative reductions, unidirectionality of these cycles must be established by requiring that each isolated cycle contain at least one positive path. Since a fragment’s coefficient is a sum of subgraph coefficients, transformations that change the number of subgraphs are particularly hazardous unless it can be shown that subgraphs appear or disappear in pairs (or larger groups) whose coefficients sum to zero. This is difficult to guarantee, making reductions involving non-unidirectional cycles tricky since eliminating components of such a cycle will normally change the number of subgraphs in a fragment. In essence, the issue is that non-unidirectional cycles consist of sequences of negative paths, but a negative path and its reverse are considered to be distinct negative paths in the theory (Mincheva and Roussel 2007a). Pruning out an edge that is part of a negative path will therefore generally reduce the number of subgraphs that can be made from a larger non-unidirectional cycle. Preserving the unidirectionality of a cycle eliminates these complications. In addition, since cycles will usually be attached to other vertices in the bipartite graph, the vertex (or vertices) that bridges the cycle to other parts of the bipartite graph, the entry vertex, must be preserved.
With an isolated unidirectional cycle, we do not have to treat vertices in the neighborhood of an entry vertex specially because changes to these vertices will not affect cycles that can be formed with vertices outside of the cycle that is being simplified. For example, removing from an isolated cycle a unitary edge along a positive path when neither of the edge’s vertices is an entry vertex would remove a factor of 1 from the coefficients of any subgraphs in which this cycle appears, and thus leave the coefficients of any fragments containing this cycle invariant. This means that if a particular fragment was (non-)critical, it would remain (non-)critical after simplification. Similarly, since a negative path contributes a negative factor, then removing two unitary negative paths without removing any entry vertices from an isolated cycle would remove a product of \(-1 \cdot -1 = 1\) from the coefficients of subgraphs in which the cycle appears, again leaving the fragment’s coefficient invariant.Footnote 7 The main idea we pursue below is that any number of unitary edges can be removed, provided the cycle remains unidirectional, whereas unitary negative paths may be removable in pairs. We will eventually lift the requirement for paths to be unitary, which will require transformations of the original graph beyond the removal of edges or paths.
4.3 Linked cycles
When an entry vertex is involved in another cycle, reductions are not as straightforward. Cycles of this type are called linked cycles. Note that, by definition, linked cycles involve at least two cycles. When trying to delete vertices near an entry vertex, creation or destruction of cycles may occur even if the vertices involved were connected by a unitary edge along a positive path.
As noted above, changing the number of subgraphs in a fragment can change its coefficient, unless these changes are balanced in the sense that the sum of the coefficients of the subgraphs added or removed is zero, which will in general be difficult to guarantee. Therefore, in simplifications of linked cycles, a conservative approach will leave the number of subgraphs that can be drawn unchanged as well as preserving the coefficients of all cycles in a set of linked cycles. The key to accomplishing this lies in the set of unidirectional cycles within the linked cycle. If we can find isolated edges (with vertices that are not entry vertices) in a unidirectional cycle within a linked cycle, then similar reductions to those discussed in Sect. 4.2 can be applied if any previously unidirectional cycles remain unidirectional. In general, an entry vertex cannot be removed and reductions near the entry vertex must be applied carefully, otherwise unidirectionality of cycles may be lost, or the number of subgraphs may change.
A non-obvious linked cycle. On the surface, this cycle may appear to be isolated. However, if these vertices and edges form a fragment, the negative-path cycle \(\left\{ \overline{\text {[A}_4,\text {R}_4,\text {A}_5]},\overline{\text {[A}_5,\text {R}_4,\text {A}_4]}\right\} \) (among others) appears in one of the subgraphs, breaking the condition for an isolated cycle
Figure 3 illustrates a linked cycle which, on the surface, may appear to be isolated. However, due to the entry vertex being a reaction vertex involved in a negative path with the larger cycle and the external vertex, A\(_5\), it is a linked cycle. The graph in Fig. 3 contains several cycles, each of which will appear in subgraphs of fragments containing these vertices:
-
the “obvious” cycle \(\left\{ \text {[A}_1,\text {R}_1,\text {A}_2],\ldots ,\text {[A}_4,\text {R}_4,\text {A}_1]\right\} \);
-
the negative-path cyclesFootnote 8\(\left\{ \overline{\text {[A}_2,\text {R}_2,\text {A}_3]},\overline{\text {[A}_3,\text {R}_2,\text {A}_2]}\right\} \), \(\left\{ \overline{\text {[A}_3,\text {R}_3,\text {A}_4]},\overline{\text {[A}_4,\text {R}_3,\text {A}_3]}\right\} \) and \(\left\{ \overline{\text {[A}_4,\text {R}_4,\text {A}_5]},\overline{\text {[A}_5,\text {R}_4,\text {A}_4]}\right\} \);
-
and a cycle encompassing the entire set of vertices: \(\Big \{\text {[A}_5,\text {R}_4,\text {A}_1],\ldots ,\overline{\text {[A}_4,\text {R}_4,\text {A}_5]}\Big \}\).
An attempted simplification through pairwise removal of the negative paths \(\overline{\text {[A}_2,\text {R}_2,\text {A}_3]}\) and \(\overline{\text {[A}_3,\text {R}_3,\text {A}_4]}\) by deleting vertices A\(_2\), R\(_2\), A\(_3\) and R\(_3\) would drastically reduce the number of subgraphs of the fragments that contain these cycles, and might thus alter some of their coefficients. However, the profusion of cycles that can be drawn from this portion of a bipartite graph was only possible because of the negative path \(\overline{\text {[A}_4,\text {R}_4,\text {A}_5]}\). Note that if A\(_5\) was a product rather than a reactant of R\(_4\), the cycle becomes an isolated cycle and none of these difficulties arise. This example strikes a note of caution with respect to the removal of negative paths when an entry reaction vertex participates in a negative path with vertices that appear in two distinct cycles. Despite the more delicate nature of linked cycles, the same ideas regarding reductions applied to isolated cycles still hold, but we will need to be more careful with the vertex eliminations we allow.
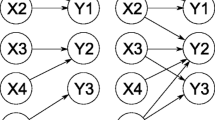
Definition of symbols used in describing the reductions of a Theorem 1, b Theorem 6 and c Theorem 8. The red lines represent arrows that can be oriented in either direction. The dotted black lines are there to show there is an arbitrary number of paths in between the vertices. Note that there must be at least one positive path (pre- and post-reduction) oriented in the clockwise direction in each figure in order for reduction to be successfully undertaken
5 Graph-based reduction rules
5.1 Theorems and Proofs
In this section, we will present the theorems that are the centrepiece of this work. Figure 4 provides a visual aid to the notation used in describing the reductions in the theorems. All of the cycles drawn are assumed unidirectional in the clockwise direction without loss of generality. The weight \(a_h\) refers to an arbitrarily selected edge in a cycle of order \(\nu \). By convention, \(a_0 \equiv a_\nu \) and \(a_{\nu +1} \equiv a_1\). Also, the vertices (and their accompanying arrows) are labeled in the direction of the (unidirectional) cycle.
Refer to Fig. 4a for the interpretation of the symbols in Theorem 1.
Theorem 1
(Positive paths in unidirectional isolated cycles) Assume we have a bipartite graph that contains an integrated unidirectional isolated cycle \(C_{i}\) of order \(\nu \), with \(\nu \) sufficiently large.Footnote 9 Let \(a_1, a_2, \ldots , a_\nu \) be the weights of all \(\nu \) arrows in \(C_{i}\), with \(W_o\) the set containing the weights \(a_{h-1}\), \(a_{h+1}\), and \(W_e\) the set containing the weights \(a_h\), \(a_{h+2}\). Let \([\textrm{A}_k,\textrm{R}_j,\textrm{A}_i]_{a_h, a_{h+1}}\) be a positive path in \(C_{i}\), where neither R\(_j\) nor at least one of the species vertices is an entry vertex. Additionally, let R\(_p\) and R\(_s\) be the reaction vertices immediately preceding and succeeding, respectively, this positive path. We have the following cases:
-
1.
A\(_k\) is not an entry vertex and at most one element in \(W_o\) is non-unitary. Let \(\omega _o =\max (W_o)\). If \(a_h\) is unitary, then construct the resulting cycle \({\bar{C}}_i\) by removing species A\(_k\) and reaction R\(_j\) (and arrows with weights \(a_{h-1}\), \(a_h\), \(a_{h+1}\)) and adding a new arrow that connects R\(_p\) and A\(_i\) with weight \(\omega _o\) oriented in the same direction as the original arrow with weight \(a_{h-1}\).
-
2.
A\(_i\) is not an entry vertex and at most one element in \(W_e\) is non-unitary. Let \(\omega _e =\max (W_e)\). If \(a_{h+1}\) is unitary, then construct the resulting cycle \({\bar{C}}_i\) by removing species A\(_i\) and reaction R\(_j\) (and arrows with weights \(a_h\), \(a_{h+1}\), \(a_{h+2}\)) and adding a new arrow that connects \(A_k\) and \(R_s\) with weight \(\omega _e\) oriented in the direction of the cycle [clockwise as the cycle is drawn in Fig. 4a].
If the resulting cycle \({\bar{C}}_i\) is unidirectional, then any fragment that contained \(C_i\) will have the same coefficient after replacing the latter cycle by \({\bar{C}}_i\). Moreover, this reduction is conservative.
Proof
Let \(C_{i}\) be an integrated unidirectional isolated cycle of order \(\nu \) in a bipartite graph. Because of its unidirectionality and isolation, the vertices, paths and edges of \(C_i\) can appear in a subgraph in exactly two ways: as the entire cycle, or as a union of reactant edges, in the latter case with all of the edges selected for the subgraph pointing in the direction of the unidirectional cycle. We only allow the removal of non-entry vertices; thus, simplifying the cycle will not affect connectivity to the rest of the bipartite graph. Note in particular that R\(_j\) is a non-entry vertex.
In case 1, we have the non-entry vertex A\(_k\) with \(a_h = 1\). The reactant edge \([\textrm{A}_k,\textrm{R}_j]_{a_h}\) contributes a factor of \(a^2_h=1\) to the coefficient of any subgraph that includes it so removing this edge has no effect on the coefficients of these subgraphs. Furthermore, in any subgraph containing the entire cycle, arrows with weights \(a_{h-1}\), \(a_h\) and \(a_{h+1}\) contribute a magnitude of \((a_{h-1})(a_h)(a_{h+1}) = \omega _o\) to \(K_C\), and thus to the coefficient of any subgraph that contains \(C_i\). Connecting R\(_p\) and A\(_i\) with an arrow of weight \(\omega _o\), \({\bar{C}}_i\) therefore inherits the magnitude of the coefficient of \(C_i\). Having the new arrow drawn in the same direction as the original arrow between R\(_p\) and A\(_k\) preserves the parity of the cycle, and therefore the sign of \(K_C\). Moreover, if unidirectionality was preserved, the cycle resulting from simplification can still appear in a subgraph in only two ways, as a union of reactant edges or as an entire cycle. It follows that the removal of the reactant edge \([\textrm{A}_k,\textrm{R}_j]_{a_h}\) with an appropriate arrow drawn between R\(_p\) and A\(_i\) preserves the number of subgraphs of any fragments in which the cycle appears as well as the coefficients of the subgraphs, and thus the coefficient of the fragment. Case 2 is proved similarly.
The second part of the theorem, that the reduction is conservative, results from the elimination of non-entry vertices from an isolated cycle. Because none of the vertices in the cycle participate in other cycles, removing two non-entry vertices cannot change, destroy or create any other cycles in the bipartite graph. \(\square \)
Figure 5 illustrates the use of Theorem 1.
a An integrated unidirectional isolated cycle that meets the criteria of Theorem 1. We assume that the A\(_4\) to R\(_3\) edge has weight \(a_{h-1} = a\). The vertices in the red box are the vertices that will be removed from the positive path [A\(_4\),R\(_4\),A\(_1\)]. b The result of applying Theorem 1. Note that no further edges could be removed using Theorem 1 because doing so would result in a cycle that was no longer unidirectional, violating one of the conditions of the theorem
Remark 1
Although a reconnection such as the A\(_1\) to R\(_3\) edge in Fig. 5 leaves a reaction labeled R\(_3\) from the original model, this is not the same reaction as in the original model, and it might be best in fact to relabel any reactions affected by a reduction operation, perhaps with an “m" next to the subscript, e.g. R\(_{3\text {m}}\), to indicate a modified reaction. We adopt this convention only when necessary to distinguish a reaction in the original set from a reaction in a transformed bipartite graph. Because we have removed a reaction and consequently may have changed the kinetics of transit through the modeled pathway, the rate constant of R\(_{3\text {m}}\) may have to be different from the rate constant of R\(_3\) in order for the reduced model to remain in the same dynamical regime as the original model. If we consider specifically the case of removing some positive paths from a long chain of positive paths, the appropriate rate constant for the reconnection will be determined by the rate-limiting step of the chain. Given that the chain may contain reactions of different orders, this may not be as simple as picking out the smallest rate constant, which would not be comparable quantities if they have different units.
Corollary 2
(Positive paths in linked cycles) Assume we have a bipartite graph that contains an integrated linked cycle of sufficiently large order, with \(C_{L}\) being the linked cycle. Let \(C_{L_u}\) be the collection of all unidirectional cycles in \(C_{L}\). Let \(C_{L_{u}}[a]\) be an arbitrary cycle from \(C_{L_u}\). Then Theorem 1 can be applied to a positive path in \(C_{L_{u}}[a]\) assuming all conditions in Theorem 1 are fulfilled in addition to maintaining unidirectionality of all cycles in \(C_{L_u}\).
Proof
The proof is essentially the same as for Theorem 1. The key is that the removal of the edge satisfying the appropriate conditions does not create or destroy any cycles or change their coefficients. It only reduces the length of one or more cycles, but that has no effect on the coefficient of a cycle. \(\square \)
While positive paths in cycles can be shortened without changing anything significant about the overall structure of the graph and of its subgraphs, the same cannot be said of negative paths. Part of the problem, as mentioned earlier, is that a negative path by itself is a cycle. Thus, negative paths that appear as part of a large cycle can also show up as cycles in fragments that do not include the entirety of the large cycle. There is also an interpretation issue for the graph obtained following the elimination negative paths, which we discuss in Sect. 5.2. For now though, we focus on the purely mathematical problem of generating reductions that preserve the coefficient of a fragment.
We begin with the following lemmas:
Lemma 3
Fragments that contain a negative-path cycle \(C_i\) that appears without connections to any other vertices of the fragment have a coefficient of zero.
Proof
By hypothesis, we have a fragment
with \(j<k\), the last two species vertices and the repeated reaction vertex being the components of the negative path \([\overline{\textrm{A}_{k-1},\textrm{R}_j,\textrm{A}_{\textrm{k}}}]_{a_h,a_{h+1}}\). If \(C_i\) has no other connections to the vertices in \(S_k\), then due to the combinatorial nature of subgraph construction, the subgraphs will appear in pairs that are identical except that one member of the pair will contain the edge set \(\{[\textrm{A}_{k-1},\textrm{R}_j],[\textrm{A}_k,\textrm{R}_j]\}\) and the other the cycle \(\{[\overline{\textrm{A}_{k-1},\textrm{R}_j,\textrm{A}_{\textrm{k}}}],[\overline{\textrm{A}_k,\textrm{R}_j,\textrm{A}_{\mathrm{k-1}}}]\}\). According to equations (6a) and (6b), the coefficients of these pairs of fragments will differ by factors of \((a_ha_{h-1})^2\) and \(-(a_ha_{h-1})^2\), respectively. Thus, these subgraphs will cancel out in pairs leaving an overall value of \(K_{S_k}\) of zero. \(\square \)
Lemma 4
If a negative-path cycle appears in a subgraph of a fragment of the form (7) and the reaction \(\textrm{R}_j\) only has the reactants \(\textrm{A}_{k-1}\) and \(\textrm{A}_k\), then the two chemical species in that cycle are not at the origin of any other edges in any subgraphs of the fragment.
Proof
For a fragment of the form (7) where \(\textrm{R}_j\) has no other reactants, the edges \([\textrm{A}_{k-1},\textrm{R}_j]\) and \([\textrm{A}_k,\textrm{R}_j]\) must be included in every subgraph of the fragment. This makes it impossible for \(\textrm{A}_{k-1}\) or \(\textrm{A}_k\) to be at the origin of any other edge or path in any of the fragment’s subgraphs. \(\square \)
Lemma 5
A cycle consisting of two consecutive negative paths cannot appear in a subgraph.
Proof
In a subgraph, each species vertex must appear once and only once at the origin of an edge or path. If one attempts to make a return trip to the first species in a pair of consecutive negative paths, one inevitably needs to start a path from the middle species both on the way out and on the way back, which is not allowed. \(\square \)
Refer to Fig. 4b for the interpretation of the symbols in Theorem 6.
Theorem 6
(Consecutive Negative Paths in Unidirectional Isolated Cycles) Assume we have a bipartite graph that contains an integrated unidirectional isolated cycle \(C_{i}\) of sufficiently large order \(\nu \). Let \(a_1, a_2, \ldots , a_\nu \) be the weights of all \(\nu \) arrows oriented in the direction of \(C_{i}\), with \(W_o\) the set containing the weights \(a_{h-1}\), \(a_{h+1}\), \(a_{h+3}\), and \(W_e\) the set containing the weights \(a_h\), \(a_{h+2}\), \(a_{h+4}\). Let \(\overline{[A_k,R_j,A_i]}_{a_h, a_{h+1}}\) and \(\overline{[A_i,R_m,A_p]}_{a_{h+2}, a_{h+3}}\) be two consecutive negative paths in \(C_{i}\), where neither of the reaction vertices nor at least two of the species vertices are entry vertices. Additionally, let \(R_p\) and \(R_s\) be the reaction vertices immediately preceding and succeeding, respectively, this pair of consecutive negative paths. We have the following cases:
-
1.
Neither \(A_k\) nor \(A_i\) are entry vertices, and at most one element in \(W_o\) is non-unitary. Let \(\omega _o=\max (W_o)\). If \(a_h=a_{h+2}=1\), then construct the resulting cycle \({\bar{C}}_i\) as follows: Remove species vertices \(A_k\) and \(A_i\) as well as the reaction vertices \(R_j\) and \(R_m\) along with the consecutive arrows with weights \(a_{h-1}\) to \(a_{h+3}\), then connect \(R_p\) and \(A_p\) with an arrow whose direction is that of the original arrow connecting \(R_p\) to \(A_k\) and weight equal to \(\omega _o\).
-
2.
Neither \(A_k\) nor \(A_p\) are entry vertices, and at most one element in each of the sets \(W_o\) and \(W_e\) is non-unitary. Let \(\omega _o=\max (W_o)\) and \(\omega _e =\max (W_e)\). Construct the resulting cycle \({\bar{C}}_i\) as follows: Remove species vertices \(A_k\) and \(A_p\) and reaction vertices \(R_j\) and \(R_m\) along with the consecutive arrows with weights \(a_{h-1}\) to \(a_{h+4}\), then connect \(R_p\) and \(A_i\) by an arrow with weight \(\omega _o\) and in the same direction as the original arrow connecting \(R_p\) to \(A_k\), and connect \(A_i\) and \(R_s\) with an arrow of weight \(\omega _e\) oriented in the direction of the cycle [clockwise as the cycle is drawn in Fig. 4b].
-
3.
Neither \(A_i\) nor \(A_p\) are entry vertices, and at most one element in \(W_e\) is non-unitary. Let \(\omega _e =\max (W_e)\). If \(a_{h+1}=a_{h+3}=1\), then construct the resulting cycle \({\bar{C}}_i\) as follows: Remove species vertices \(A_i\) and \(A_p\) as well as the reaction vertices \(R_j\) and \(R_m\) along with the consecutive arrows with weights \(a_h\) to \(a_{h+4}\), then connect \(A_k\) and \(R_s\) by an arrow with weight \(\omega _e\) oriented in the direction of the cycle [clockwise as the cycle is drawn in Fig. 4b].
Then any fragment that originally contained \(C_i\) will have the same coefficient after replacing \(C_i\) by \({\bar{C}}_i\). Moreover, the reduction will be conservative.
Proof
Let \(C_i\) be an integrated unidirectional isolated cycle of order \(\nu \) in a bipartite graph. Then, by definition of integrated unidirectional isolated cycles and subgraphs, this cycle contains an equal number of species and reaction vertices and will appear in any subgraph either as the set of reactant edges or as the entire cycle. In each of the cases in Theorem 6, all removed vertices are non-entry vertices. Thus, removal of these vertices will not affect connectivity to any vertices outside of \(C_i\).
In case 1, A\(_k\) and A\(_i\) are non-entry vertices and \(a_h = a_{h+2}= 1\). Thus, the reactant edges [A\(_k\),R\(_j]_{a_h}\) and [A\(_i\),R\(_m]_{a_{h+2}}\) contribute factors of unity to the coefficient of any subgraph that includes the edge subgraph of \(C_i\). Furthermore, in any subgraph containing \(C_{i}\), arrows with weights \(a_{h-1}\), \(a_h\), \(a_{h+1}\), \(a_{h+2}\), and \(a_{h+3}\) contribute a magnitude of \((a_{h-1})(a_h)(a_{h+1})(a_{h+2})(a_{h+3}) = \omega _o\) to \(K_C\), and therefore to the coefficient of any subgraph in which \(C_i\) appears. Connecting R\(_p\) to A\(_p\) with an arrow of weight \(\omega _o\) clearly preserves the magnitude of \(K_C\). Choosing the direction of the new arrow as that of the original R\(_p\) to A\(_k\) arrow preserves the parity of \(K_C\) and thus its sign. Accordingly, the coefficients of all subgraphs containing \(C_i\) will remain unchanged after replacing \(C_i\) by \({\bar{C}}_i\). The number of subgraphs of a fragment containing \(C_i\) or its edge subgraph also does not change because of the unidirectionality of the cycle.
Cases 2 and 3 can be proven similarly, the key idea being that both the number and coefficients of subgraphs in a cycle containing \(C_i\) do not change post-reduction.
Turning to the issue of conservativity of the reduction, we note that the consecutive negative paths cannot appear in any subgraph (Lemma 5). The individual negative-path cycles that have been eliminated cannot contribute to a critical fragment (Lemma 3) and since, by hypothesis, the reaction vertices are not entry vertices, they cannot enter into fragments in more complicated ways (Lemma 4). Thus, no other fragments’ coefficients will be affected by the removal of consecutive negative paths under the conditions of this theorem. \(\square \)
Figure 6 illustrates the use of Theorem 6. It is easily verified that \(K_C=-a\) for both cycles and that their corresponding edge subgraphs both have coefficients of unity.
a An integrated unidirectional isolated cycle that meets the criteria of Theorem 6. For simplicity, we choose only one of the arrows to have an arbitrary weight, \(a_{h-1} = a\). The vertices in the red box are the vertices that will be removed. b The result of applying Theorem 6 to the figure on the left
Corollary 7
Assume we have a bipartite graph that contains an integrated linked cycle of sufficiently large order, with \(C_{L}\) being the linked cycle. Let \(C_{L_u}\) be the collection of all unidirectional cycles in \(C_{L}\). Let \(C_{L_{u}}[a]\) be an arbitrary cycle from \(C_{L_u}\) that does not contain an entry reaction vertex that participates in a negative path with a species vertex in \(C_{L_{u}}[a]\) and a species vertex outside of \(C_{L_{u}}[a]\). Then Theorem 6 can be applied to consecutive negative paths in \(C_{L_{u}}[a]\) assuming all conditions in Theorem 6 are fulfilled in addition to maintaining unidirectionality of all cycles in \(C_{L_u}\).
Proof
Assuming the extra condition is fulfilled, then the proof of Theorem 6 applies essentially unchanged. That is, all of the cycles in \(C_{L}\) will retain their parities and \(K_C\) values post-reduction. Accordingly, the number of subgraphs in any fragment and their coefficients also remain unchanged. \(\square \)
Refer to Fig. 4c for the interpretation of the symbols in Theorem 8.
Theorem 8
(Disjoint negative paths in unidirectional isolated cycles) Assume we have a bipartite graph that contains an integrated unidirectional isolated cycle \(C_{i}\) of sufficiently large order \(\nu \). Let \(a_1, a_2, \ldots , a_\nu \) be the weights of all \(\nu \) arrows oriented in the direction of \(C_{i}\). Let \(\overline{[A_k,R_j,A_i]}_{a_h, a_{h+1}}\) and \(\overline{[A_p,R_m,A_q]}_{a_{f}, a_{f+1}}\) be two disjoint negative paths in this isolated cycle where neither of the reaction vertices nor at least one species vertex from each negative path is an entry vertex. Denote by \(W_{o_h}\) the set containing the weights \(a_{h-1}\) and \(a_{h+1}\), by \(W_{e_h}\) the set containing the weights \(a_h\) and \(a_{h+2}\), by \(W_{o_f}\) the set containing the weights \(a_{f-1}\) and \(a_{f+1}\), and by \(W_{e_f}\) the set containing the weights \(a_f\) and \(a_{f+2}\). Additionally, let \(R_{p_h}\), \(R_{s_h}\) be the reaction vertices immediately preceding and succeeding, respectively, \(\overline{[A_k,R_j,A_i]}_{a_h, a_{h+1}}\), and \(R_{p_f}\), \(R_{s_f}\) be the reaction vertices immediately preceding and succeeding, respectively, \(\overline{[A_p,R_m,A_q]}_{a_{f}, a_{f+1}}\).Footnote 10 Consider the following cases:
-
1.
Neither \(A_k\) nor \(A_p\) are entry vertices and at most one element in each of \(W_{o_h}\) and \(W_{o_f}\) is non-unitary. Let \(\omega _{o_h} =\max (W_{o_h})\) and \(\omega _{o_f} =\max (W_{o_f})\). If \(a_h\) and \(a_f\) are unitary, then construct the resulting cycle \({\bar{C}}_i\) as follows: Remove species vertices \(A_k\) and \(A_p\) as well as the reaction vertices \(R_j\) and \(R_m\) along with the arrows with weights \(a_{h-1}\), \(a_h\), \(a_{h+1}\), \(a_{f-1}\), \(a_f\), \(a_{f+1}\), then connect \(R_{p_h}\) and \(A_i\) by an arrow with weight \(\omega _{o_h}\) and in the same direction as the original arrow connecting \(R_{p_h}\) to \(A_k\), and connect \(R_{p_f}\) and \(A_q\) with an arrow of weight \(\omega _{o_f}\) and in the same direction as the original arrow connecting \(R_{p_f}\) and \(A_p\).
-
2.
Neither \(A_k\) nor \(A_q\) are entry vertices and at most one element in each of \(W_{o_h}\) and \(W_{e_f}\) is non-unitary. Let \(\omega _{o_h} =\max (W_{o_h})\) and \(\omega _{e_f} =\max (W_{e_f})\). If \(a_h\) and \(a_{f+1}\) are unitary, then construct the resulting cycle \({\bar{C}}_i\) as follows: Remove species vertices \(A_k\) and \(A_q\) as well as the reaction vertices \(R_j\) and \(R_m\) along with the arrows with weights \(a_{h-1}\), \(a_h\), \(a_{h+1}\), \(a_f\), \(a_{f+1}\), \(a_{f+2}\), then connect \(R_{p_h}\) and \(A_i\) by an arrow with weight \(\omega _{o_h}\) and in the same direction as the original arrow connecting \(R_{p_h}\) to \(A_k\), and connect \(A_p\) and \(R_{s_f}\) with an arrow of weight \(\omega _{e_f}\) oriented in the direction of the cycle [clockwise as the cycle is drawn in Fig. 4c].
-
3.
Neither \(A_i\) nor \(A_p\) are entry vertices and at most one element in each of \(W_{e_h}\) and \(W_{o_f}\) is non-unitary. Let \(\omega _{e_h} =\max (W_{e_h})\) and \(\omega _{o_f} =\max (W_{o_f})\). If \(a_{h+1}\) and \(a_f\) are unitary, then construct the resulting cycle \({\bar{C}}_i\) as follows: Remove species vertices \(A_i\) and \(A_p\) as well as the reaction vertices \(R_j\) and \(R_m\) along with the arrows with weights \(a_h\), \(a_{h+1}\), \(a_{h+2}\), \(a_{f-1}\), \(a_f\), \(a_{f+1}\), then connect \(A_k\) and \(R_{s_h}\) by an arrow with weight \(\omega _{e_h}\) oriented in the direction of the cycle [clockwise as the cycle is drawn in Fig. 4c], and connect \(R_{p_f}\) and \(A_q\) with an arrow of weight \(\omega _{o_f}\) and in the same direction as the original arrow connecting \(R_{p_f}\) and \(A_p\).
-
4.
Neither \(A_i\) nor \(A_q\) are entry vertices and at most one element in each of \(W_{e_h}\) and \(W_{e_f}\) is non-unitary. Let \(\omega _{e_h} =\max (W_{e_h})\) and \(\omega _{e_f} =\max (W_{e_f})\). If \(a_{h+1}\) and \(a_{f+1}\) are unitary, then construct the resulting cycle \({\bar{C}}_i\) as follows: Remove species vertices \(A_i\) and \(A_q\) as well as the reaction vertices \(R_j\) and \(R_m\) along with the arrows with weights \(a_h\), \(a_{h+1}\), \(a_{h+2}\), \(a_f\), \(a_{f+1}\), \(a_{f+2}\), then connect \(A_k\) and \(R_{s_h}\) by an arrow with weight \(\omega _{e_h}\), and connect \(A_p\) and \(R_{s_f}\) by an arrow with weight \(\omega _{e_f}\), both oriented in the direction of the cycle [clockwise as the cycle is drawn in Fig. 4c].
Then any fragment that originally contained \(C_i\) will have the same coefficient after replacing \(C_i\) by \({\bar{C}}_i\). Moreover, reductions using this theorem are conservative.
Proof
By definition of integrated unidirectional isolated cycles, \(C_i\) contains an equal number of species and reaction vertices and will appear in subgraphs of any fragment that includes it either as the set of its reactant edges or as the entire cycle. In each case considered above, all removed vertices are non-entry vertices; thus, removal of these will not affect connectivity to the rest of the vertices outside of \(C_{i}\). By assumption, we have that \(R_j\) and \(R_m\) are non-entry vertices.
In case 1, we have the non-entry vertices A\(_k\) and A\(_p\) and \(a_h = a_f = 1\). The reactant edges [A\(_k\),R\(_j]_{a_h}\) and [A\(_p\),R\(_m]_{a_f}\) thus contribute factors of unity when the cycle’s reactant edge set appears in a subgraph. Furthermore, in any subgraph containing the entire cycle, arrows with weights \(a_{h-1}\), \(a_h\), \(a_{h+1}\), \(a_{f-1}\), \(a_f\), and \(a_{f+1}\) contribute a magnitude of \(\omega _{o_h}\omega _{o_f}\) to the coefficient of the subgraph. Applying these weights to, respectively, the new product edges \([\textrm{R}_{p_h},\textrm{A}_i]\) and \([\textrm{R}_{p_f},\textrm{A}_q]\) means that the coefficient of a subgraph containing the simplified cycle \({\bar{C}}_i\) will have a coefficient of the same magnitude as the original subgraph. A pair of negative paths is removed, and the rule for the directions of the arrows reconnecting the graph ensures that there is no other change in the number of negative paths, i.e. \(C_i\) and \({\bar{C}}_i\) must have the same parity. Thus, the coefficient of a subgraph containing the entire cycle is preserved. Moreover, the number of subgraphs of a fragment containing \(C_i\) or its edge subgraph also does not change after simplification because of the unidirectionality of both \(C_i\) and \({\bar{C}}_i\), which implies that the coefficient of any fragment containing the cycle is preserved.
Cases 2, 3, and 4 can be proven by analogous arguments and are not presented for brevity.
These reductions are conservative because fragments that include the eliminated negative-path cycles are dynamically unimportant according to Lemma 3 and the conditions of the theorem require the eliminated reactions to be non-entry vertices, which means that these cycles cannot appear in a subgraph with connections that would create added complications (Lemma 4). \(\square \)
See Fig. 7 for an example of utilizing this theorem.
a An integrated unitary unidirectional isolated cycle that meets the criteria of Theorem 8. The vertices in the red and blue boxes are the disjoint negative paths \(\overline{[\text {A}_1,\text {R}_1,\text {A}_2]}\) and \(\overline{[\text {A}_3,\text {R}_3,\text {A}_4]}\). The species vertex with the green star \((\text {A}_1)\) will be kept from the blue negative path and the species vertex with the orange star \((\text {A}_3)\) will be kept from the red negative path. b The result of applying Theorem 8 to the figure on the left
Corollary 9
Assume we have a bipartite graph that contains an integrated linked cycle of sufficiently large order, with \(C_{L}\) being the linked cycle. Let \(C_{L_u}\) be the collection of all unidirectional cycles in \(C_{L}\). Let \(C_{L_{u}}[a]\) be an arbitrary cycle from \(C_{L_u}\) that does not contain an entry reaction vertex that participates in a negative path with a species vertex in \(C_{L_{u}}[a]\) and a species vertex outside of \(C_{L_{u}}[a]\). Then Theorem 8 can be applied to disjoint negative paths in \(C_{L_{u}}[a]\) assuming all conditions in Theorem 8 are fulfilled in addition to maintaining unidirectionality of all cycles in \(C_{L_u}\).
Proof
Assuming the extra condition is fulfilled, then the same proof for Theorem 8 applies. That is, all of the cycles in \(C_{L}\) will still be present post-reduction, their contributions to the coefficients of various subgraphs will be the same, and the number of subgraphs in a fragment containing these cycles will be unchanged. \(\square \)
5.2 (Bio)chemical relevance
The theorems and corollaries found in the previous section describe reductions that preserve critical fragments. They do not, however, address the (bio)chemical relevance of the reduced model. For example, it is possible that a removed species or reaction vertex may have been a well-studied component of a mechanism and that there are some observations regarding this component that a model should be able to match. In these cases, it may be desirable to identify species to be preserved prior to any reductions being made, and to avoid eliminating these species in the course of reduction. In many cases, especially if most of the edges are unitary, the theorems presented above offer multiple options for reduction. It will accordingly sometimes be possible to carry out reductions while protecting specific species or reactions.
Unfortunately, reductions involving negative paths are not as straightforward as reductions of positive paths because a negative path encodes a co-reactant relationship, the product not being included in an isolated cycle. In this case, the product species (if any) would have to be outside the cycle. Theorems 6 and 8 remove both reaction vertices involved in the negative paths subject to reduction. Unless the product is formed by another reaction, this therefore eliminates the formation of the product from the model. If the product is dynamically important, then its removal may sabotage the model altogether. However, in order for the product to play an important dynamical role, it would have to be part of a cycle. Thus, the cycle on which we were operating would not be isolated, and the reaction vertex forming the product would in fact be a non-trivial entry vertex, which our theorems explicitly disallow as targets for reduction. The circumstances in which it would be permissible to remove a reaction generating a product would involve products that are part of a pathway that acts as a sink. This includes the trivial case in which the immediate product is a sink species (often represented by an empty-set symbol in reaction mechanisms).
There is also the significant problem of interpreting the bipartite graph resulting from the elimination of negative paths. For instance, consider the example shown in Fig. 7, in which Theorem 8 is applied. The original cycle encoded the reactions (neglecting species not explicitly shown in the cycle) \(\textrm{A}_1+\textrm{A}_2\rightarrow \), \(\textrm{A}_2+\textrm{A}_3\rightarrow \), \(\textrm{A}_3+\textrm{A}_4\rightarrow \) and \(\textrm{A}_4\rightarrow \textrm{A}_1\). In the reduced cycle on the other hand, we have \(\textrm{A}_1+\textrm{A}_3\rightarrow \) and \(\textrm{A}_3\rightarrow \textrm{A}_1\). These are quite different in terms of their chemical meanings, and it may be that, for example, there is no chemical sense to be attached to \(\textrm{A}_1\) reacting with \(\textrm{A}_3\). Even if that is not the case, the reactant-product relationships have been profoundly transformed and, arguably, the reduced model is not a model for the original reaction but for a different reaction. However, an alternative perspective is that we do not care about the chemical interpretation of the reduced bipartite graph, only that it is easier to analyze. The time required for GraTeLPy to check all fragments in a mechanism grows rapidly with the size of the bipartite graph. Thus, it may be useful to simplify a bipartite graph even when an automated search for critical fragments is planned. The discovery of a critical fragment in the reduced model could then be mapped back to a critical fragment of the original model.
As an interesting side-note, in the classical theory of kinetic equivalence of mechanisms, two models are kinetically equivalent if they are related by a linear transformation that preserves the entropy production (de Donder 1937; Prigogine 1946; Dutta et al. 1984). Kinetically equivalent models generate the same time series for the chemical species that are common to two mechanisms. This is a considerably more restrictive condition than preserving qualitative dynamics, i.e. the potential for certain bifurcations. However, the reactions in the reduced models generated by our methods will always be linear combinations of reactions in the original model provided we consider only the reactions in the cycle.Footnote 11 The application of Theorem 1 essentially involves “squeezing out” intermediates in a cycle, which can be accomplished by taking a linear superposition of reactions with positive superposition coefficients. The situation with negative paths is more complicated, but this is essentially equivalent to subtracting reactions. For the example of Fig. 7, if we call the resulting reactions R\(_{2\text {m}}\) and R\(_{4\text {m}}\), we have R\(_{2\text {m}}=\text {R}_3-\text {R}_4\) and R\(_{4\text {m}}=\text {R}_2-\text {R}_1\). Consequently, it is possible that the set of transformations that preserve the entropy production is a subset of the transformations allowed by our methods. We leave this question for future study.
6 Application to a gene expression control model
6.1 Model description
A delayed mass-action model (Roussel 1996) for the transcriptional control of Hmp, an NO detoxifying enzyme, by the iron-sulfur protein FNR displays bistability (Roussel 2019a). FNR acts as a transcriptional repressor, preventing the synthesis of the hmp mRNA when bound to the gene’s promoter. Nitric oxide (NO), when present in the cell, reacts with the iron-sulfur cluster of FNR. This causes FNR to lose its ability to bind the hmp promoter, and thus allows this gene to be transcribed. The synthesized Hmp can then remove NO, converting it to nitrate (\(\hbox {NO}^-_3\)), which is much less toxic. A model of this system is complicated by the potential for up to 8 molecules of NO to react with a single iron-sulfur cluster (Crack et al. 2013). Only four of these reactions are kinetically distinguishable, leading to a model with five nitrosylation states of FNR (Roussel 2019a). This model contains 18 species and 32 reactions. Both expression and clearance delays were considered, the latter representing the periods during which the promoter is occluded by an RNA polymerase or the ribosome binding site on an mRNA molecule by a ribosome. Clearance delays turn out to have significant dynamical effects (Trofimenkoff and Roussel 2020; Ünal et al. 2021). Nevertheless, in the present contribution, an ODE model was considered given that the potential for bistability does not depend on the inclusion of delays, thus providing an interesting example to which to apply the theorems presented here. Beyond the neglect of delays, we also considered a simplified version of the original model for two reasons: Perhaps most importantly, an example based on a large model with a complex bipartite graph is poorly suited as an illustration. The original model is also extremely densely connected, largely through NO. As a result, paradoxically, our model simplification methods apply to very few species and reactions in the pre-existing model. It should be noted that this pattern of dense connections is unusual. Most species in metabolic networks, for example, have very few connections to other species via reactions (Jeong et al. 2000). The simplified model presented below is thus, we feel, more representative of models likely to be encountered in practice.
The simplified mechanism consists of the following 15 mass-action reactions connecting 10 species, not counting the sink species \(\hbox {NO}^-_3\):














Reaction (8a) represents a source of nitric oxide, which may be endogeneous or exogeneous, i.e. due to diffusion of NO into the cell from the extracellular medium. Reactions (8b) to (8e) describe the kinetics of the conversion of NO to nitrate (\(\hbox {NO}^-_3\)) catalyzed by Hmp. In reaction (8b), a molecule of oxygen binds into the active site of the enzyme. Because we hold the concentration of oxygen constant, (8b) is represented as a pseudo-first-order reaction with effective rate constant \(k_1=k_{\textrm{O}_{2}}[\hbox {O}_2]\), where \(k_{\textrm{O}_2}\) is the true second-order rate constant for this binding step. Reaction (8f) represents the transcription of the hmp gene, initiated at the gene promoter (\(\hbox {Pro}^{{ hmp}}\)), generating an mRNA. We focus particularly on the ribosome binding site (RBS) of the mRNA because translation [reaction (8g)] can be initiated as soon as the RBS becomes available in bacteria. Moreover, mRNAs are typically degraded starting from the RBS in bacteria (Kennell and Talkad 1976) [reaction (8h)]. This distinction between the RBS and mRNA is admittedly more significant in a model with delays than in an ODE model. Reactions (8i8j) to (8k) are enzyme degradation reactions. FNR is a repressor of hmp, as shown in reaction (8l). Reaction (8m) depicts the inactivation of FNR by nitric oxide, and reaction (8n) represents a recycling pathway. Details of the biochemistry are presented in detail by Roussel (2019a).
The model presented above involves the following simplifications from the original:
-
In the original model, several binding events of substrates with Hmp were assumed to be reversible, specifically reactions (8b), (8c) and (8e). In other words, we treat reactions (8b) and (8c) in the van Slyke-Cullen limit (van Slyke and Cullen 1914), while the formation of the inhibitory complex \(\hbox {Hmp}\cdot \hbox {NO}\) in reaction (8e) now appears in the mechanism as a suicide substrate inhibition reaction (Walsh 1984).
-
We assume that the \(\hbox {Hmp}\cdot \hbox {O}_2\cdot \hbox {NO}\) complex formed in reaction (8c) is stable, i.e. we do not have a sink for this species. This simply reduces the number of reactions to be considered by one without having any significant dynamical effect.
-
The multistep degradation of the iron-sulfur cluster of FNR has been reduced to the single reaction (8m).
Model (8) has two conservation relations, one for the gene promoter, and one for FNR. Accordingly, given the 10 chemical species involved in the dynamics, the model has rank \(n=8\).
The parameters of Table 1 were used throughout this study unless otherwise noted.
6.2 Analysis of the model
Figure 8 shows the bifurcation diagram of the model varying \(k_{\textrm{in}}\). Unlike the original model (Roussel 2019a), the upper branch of steady states diverges near \(k_{\textrm{in}}=0.248\,\mu \text {M s}^{-1}\) because of the suicide substrate inhibition replacing the reversible competitive substrate inhibition in the original model. Between \(k_{\textrm{in}}=0.248\) and the saddle-node bifurcation at \(0.375\,\mu \text {M s}^{-1}\), there is only one stable steady state, the lower one, but that steady state has a relatively small basin of attraction. Many initial conditions result in runaway trajectories in which [NO] increases without bound. For values of \(k_{\textrm{in}}\) greater than the saddle-node value, only runaway trajectories are observed. There is a second saddle-node point at \(k_{\textrm{in}}=2.2\times 10^{-3}\,\mu \text {M s}^{-1}\). Below this value of \(k_{\textrm{in}}\), only the lower steady state exists, and it attracts all trajectories. Between \(k_{\textrm{in}}=2.2\times 10^{-3}\) and \(0.248\,\mu \text {M s}^{-1}\), the system is bistable.
Bifurcation diagram for the Hmp model (8). In this and subsequent bifurcation diagrams, solid lines represent stable steady states while dotted lines represent unstable steady states. The insets magnify the regions in which the two saddle-node bifurcations occur. Note: all bifurcation diagrams were computed using Xppaut version 8 (Ermentrout 2002)
The bipartite graph of the model is illustrated in Fig. 9. This model has five critical fragments of order 8, one of which is shown in Fig. 10. The other four critical fragments are not drawn, but only differ by free-floating edges. For example, in one of the other critical fragments, the edge [\(\hbox {Pro}^{{ hmp}}\),R\(_A\)] is replaced by [FNR,R\(_A\)]. The subgraphs of the critical fragment of Fig. 10 are drawn in Fig. 11. From these subgraphs, we can calculate a fragment coefficient \(K_{S_8} = -1\).
The subgraphs of the critical fragment (from Fig. 10) of the Hmp model
As an aside, consider the structure of the critical fragment in Fig. 10, which includes the edges [\(\hbox {Pro}^{\textrm{hmp}}\),R\(_A\)] and [\(\hbox {RBS}^{\textrm{hmp}}\),R\(_5\)], and [\(\hbox {FNR}_e\),R\(_7\)]. Because these edges just contribute multiplicative factors of 1 to the coefficient of the fragment, a critical fragment of order 5 is obtained by leaving these out. In turn, a critical fragment of order less than the rank implies the possibility of an Andronov-Hopf bifurcation (Mincheva and Roussel 2007a). We have not searched for an oscillatory regime in this model. We point this out because the build-up of larger critical fragments from smaller critical fragments is a common theme in the analysis of bipartite graphs, explaining (in part) why it is not uncommon to find both Andronov-Hopf and saddle-node bifurcations in the same model.
6.3 Conservative reductions
The bipartite graph of the model (Fig. 9) includes a few linked cycles containing a total of three positive paths to which we can apply Corollary 2 (which generalizes Theorem 1). These positive paths are [\(\hbox {FNR}_e\),R\(_r\),FNR], [\(\hbox {Hmp}\cdot \hbox {O}_2\cdot \hbox {NO}\),R\(_{-3}\), Hmp], and [\(\hbox {Hmp}\cdot \hbox {NO}\),R\(_{6d}\),NO]. We can remove the reactant edge of each path (indicated in red in the bipartite graph) by making the appropriate connection from the preceding reaction vertex to the succeeding species vertex of each edge. We rename R\(_2\) to R\(_{2\text {m}}\), R\(_4\) to R\(_{4\text {m}}\), and R\(_7\) to R\(_{7\text {m}}\) to indicate that the modified reaction obtained after each reconnection is different from the original reaction. These connections preserve all of the cycles involved in the linked cycle and change neither the structure nor the coefficients of any of the subgraphs of the system’s critical fragments. However, due to the removal of \(\hbox {FNR}_e\) and R\(_r\), we lose two critical fragments that included the corresponding edge as a free-floating edge. We can think of this kind of loss of critical fragments as the lifting of a degeneracy due to the removal of inessential components that do not participate in the cyclical structures that give rise to criticality. The resulting bipartite graph is shown in Fig. 12. Chemically, the effect of these transformations is to replace reactions (8c) and (8d), (8e) and (8k), and (8m) and (8n) by their respective sums:



“squeezing out” the intermediates \(\hbox {Hmp}\cdot \hbox {O}_2\cdot \hbox {NO}\), \(\hbox {Hmp}\cdot \hbox {NO}\) and \(\hbox {FNR}_e\), respectively.
One of the three critical fragments of order 5 (full rank) of the Hmp model after conservative reduction. This critical fragment corresponds to the critical fragment of the full model found in Fig. 10
The subgraphs of the critical fragment from Fig. 13
The critical fragment of the simplified model and its subgraphs are shown in Figs. 13 and 14, respectively. The elimination of the three species and three reactions has reduced the number of active chemical species to 7 and the number of reactions to 12. There are still two conservation relations, so the rank of the model is now 5. Thus, the critical fragment of order 5 shown in Fig. 13 is a full-rank critical fragment corresponding directly to the critical fragment of the original model shown in Fig. 10. Comparing the subgraphs (Figs. 11 and 14), we see that the reduction has preserved
-
the number of subgraphs of the critical fragment, and
-
the number and parities of cycles in each subgraph.
Thus, the coefficient of each critical fragment in the two models will be the same. Indeed, we can easily calculate \(K_{S_5}=-1\) for this fragment. Since we chose a critical fragment for simplification, the fragment shown in Fig. 13 is still critical. In fact, the bifurcation diagram of the model retains the two saddle-node bifurcations of the original model in essentially identical positions despite the radical simplifications made (Fig. 15; compare Fig. 8). Interestingly however, the reduced model has at least one stable steady state for any value of the control parameter \(k_{\textrm{in}}\), unlike the original model in which a large inflow rate of NO can overwhelm the enzyme and its control system. This is doubtless due to the simplification of Michaelis–Menten kinetics to the simple bimolecular reactions (9) which lack saturation behavior. Indeed, our graph transformations retain the dynamics of the original model only in the sense that they preserve the capacity for bifurcations, but there is no guarantee that the behavior will be identical away from the bifurcation points or for that matter that the bifurcations will occur at similar parameter values, although that turned out to be the case here. It is not obvious, at least to us, that it is possible to predict a priori the effects of changes in the vector field far away from the bifurcating steady states, either in phase space or in parameter space. Thus, the model reductions proposed here are dynamics-preserving only locally. They can have global effects on the vector field, as we saw in Figs. 8 and 15, where the bifurcations were preserved but the geometry of one branch of steady states was dramatically altered, bringing a fixed point at infinity at larger values of \(k_{\textrm{in}}\) in the original model into the phase space of the reduced model.
It is worth pausing to consider the net reaction (9c), and to note that this reduction has eliminated the feedback loop between nitrosylation of FNR and control of the synthesis of Hmp. In other words, this control system is not essential to the bistability of this model. In the reduced model, FNR becomes a second enzyme that eliminates NO. As should perhaps have been clear from the original critical fragment (Fig. 10), the key interactions leading to bistability are located in the catalytic system, including the substrate inhibition by NO. After reduction, the critical fragment (Fig. 13) still contains two competing pathways, one via reactions R\(_1\) and R\(_{2\text {m}}\) that eliminates NO, and the other via reaction R\(_{4\text {m}}\) that eliminates Hmp. An interaction analogous to competitive substrate inhibition has previously been shown to generate oscillations in a model for hydrogen oxidation (Slin’ko and Slin’ko 1978).
The bipartite graph of the aggressively reduced model of Sect. 6.4. This model still exhibits bistability but contains only 6 species and 10 reactions. There are still two conservation relations, one for the promoter and one for FNR, so the rank of the model is now 4
6.4 Aggressive reductions
In Sect. 6.3, we applied the theorems presented in this paper to all paths that met the criteria of our conservative reduction theorems and showed that, upon reduction, the resulting model still exhibits bistability. In our first foray into these methods however, we used knowledge of a critical fragment to propose several dynamics-preserving reductions, validated by numerical computations, some of which involved the removal of entry vertices (Roussel and Roussel 2018). We call these aggressive reductions. The critical fragment of the reduced model, Fig. 13, tells us that we must preserve all of the species in the linked cycles. Anything outside of these linked cycles either doesn’t contribute to critical fragments, or contributes only trivially as free-floating edges (e.g. [\(\hbox {Pro}^{{ hmp}}\),R\(_A\)]). Provided they are not involved in other significant interactions, we could consider deleting some additional species and reactions even if they do not satisfy the conditions of our theorems. Looking at the bipartite graph (Fig. 12), we see for example that \(\hbox {RBS}^{{ hmp}}\) only appears as an intermediate in the Hmp synthesis pathway. It is an entry vertex, and it participates in a cycle of order 1 with Rtl, so it does not satisfy the conditions for conservative reductions. Nevertheless, knowing that it appears in critical fragments only trivially, we can contemplate its removal. On the other hand, it is not clear how the graph could be reconnected in a sensible way if we removed, say, \(\hbox {Pro}^{{ hmp}}\) and Rtc, even though the critical fragment suggests these species are not essential to the bistable behavior. A similar comment could be made about FNR and the reactions with which it shares edges. We therefore attempt to remove only \(\hbox {RBS}^{{ hmp}}\) and the reactions Rtl and R\(_5\), connecting Rtc directly to Hmp. As per our previously established convention, we rename the modified reaction Rtcm. The resulting bipartite graph is shown in Fig. 16. From the graph, the modified reaction Rtcm reads as follows:

We note in passing that models in which mRNA does not explicitly appear and proteins are shown as being synthesized directly from a gene are not uncommon in the literature [e.g. Kepler and Elston (2001); Lipshtat et al. (2006); Sokolowski et al. (2016)].
As expected, the critical fragment of this aggressively reduced version of the model (not shown) is identical to Fig. 13, barring the [\(\hbox {RBS}^{{ hmp}}\),R\(_5\)] edge. This model still contains three critical fragments but now each is of order 4 (full rank). Its bistable regime is depicted in Fig. 17. Because mRNA serves (in part) an amplification role (many mRNAs to one promoter), in order to get similar dynamics in the absence of mRNA, we need to increase the transcription initiation rate constant. If we do, we end up with a similar bifurcation diagram to the one found for the model that includes \(\hbox {RBS}^{{ hmp}}\). (Compare Figs. 15 and 17.)
Bifurcation diagram of the model following removal of \(\hbox {RBS}^{{ hmp}}\) and associated reactions from the model. All parameters are as in Fig. 15 except \(k_{\textrm{tcm}}=50\,\textrm{s}^{-1}\)
As we can see from this example, aggressive reductions may require the injection of some chemical knowledge into the analysis, such as the requirement that an active gene is required to make the protein Hmp.
Up to this point, all model reductions considered have retained a pair of saddle-node bifurcations between which we can observe bistability. We have found that further aggressive reductions destroy at least one of the saddle-node bifurcations, leaving us with a system that does not have a bistable regime. Note that a critical fragment of full rank is a necessary condition for a saddle-node bifurcation to occur. Because the condition is not sufficient, there is no guarantee that there will be parameters where a saddle-node bifurcation occurs, never mind two, just because the network has a critical fragment. Nevertheless, we generally find that conservative reductions preserve the full bifurcation structure, while the aggressive reductions may or may not.
The [R\(_1\),\(\hbox {Hmp}\cdot \hbox {O}_2\)] product edge in Fig. 16 might seem like an attractive target for an aggressive reduction of a slightly different kind. While this edge does appear in the critical fragment (Fig. 13), given that its only other connection is to a sink (R\(_{6b}\)), it is tempting to think that we could remove \(\hbox {Hmp}\cdot \hbox {O}_2\) along with the reaction vertices R\(_1\) and R\(_{6b}\), closing the cycle by connecting Hmp to R\(_{2\text {m}}\). This simplification does not appear at first glance to be very different from that of the [\(\hbox {Hmp}\cdot \hbox {NO}\),R\(_{6d}\)] edge. However, when we do this, we lose unidirectionality of the cycle in \(g_1\) from Fig. 14 because [Hmp,R\(_1\),\(\hbox {Hmp}\cdot \hbox {O}_2\)] is the only positive path in \(g_1\). This causes the number of subgraphs of the corresponding fragment to explode: we gain four subgraphs analogous to \(g_1\) but with different combinations of arrows, and one additional edge subgraph for a total of 10 subgraphs. The corresponding fragment is no longer critical—it has a coefficient of zero—illustrating the importance of maintaining directionality of cycles. Consequently the mechanism no longer has any critical fragments and saddle-node bifurcations are therefore no longer possible.
7 Discussion
7.1 Summary
Models of (bio)chemical networks have been frequent targets for simplification. In this paper, we discussed graph-based dynamics-preserving schemes that can be applied to mass-action (bio)chemical mechanisms represented as bipartite graphs. In this representation, the graph contains two sets of vertices—one for species and one for reactions—and directed arrows (edges) from one type of vertex to the other. The theorems in this paper preserve elements of bipartite graphs known as critical fragments, which are necessary for positive-feedback Andronov-Hopf and saddle-node bifurcations (Ivanova 1979b; Mincheva and Roussel 2007a). Thus, simplifications of models displaying positive-feedback oscillations or multistability should preserve critical fragments in order to preserve their dynamics.
In Sect. 5.1, we presented a set of theorems that will preserve critical fragments, regardless of whether we know ahead of time what critical fragments a model may have, based on the structure of the bipartite graph. We applied these theorems to a model for the control of the synthesis of the NO-detoxifying enzyme Hmp in Sect. 6.3. We then considered how knowledge of the critical fragment may enable more aggressive reductions in Sect. 6.4.
As we saw through the reduction process for the Hmp control model, systematic reduction of a bipartite graph based on our theorems, possibly accompanied by some aggressive reductions that go beyond our theorems, will tend to preserve dynamics. Not all details of the dynamics are preserved. The model used to illustrate the application of our theorems has a limited range within which stable solutions are found (Fig. 8) whereas the simplified models have stable steady states throughout the range of nitric-oxide inflow rates (Figs. 15 and 17). However, all of the correctly reduced models have a pair of saddle-node bifurcations delimiting a range of bistable behavior.
The computational cost of finding critical fragments rises rapidly with the size of the network, so it will not always be possible to identify critical fragments immediately. Simplifying a model using our methods, which can literally be carried out with pen and paper using a printout of the bipartite graph, may in some cases yield a more manageable model that can then be analyzed using GraTeLPy (Walther et al. 2014) or, after converting the bipartite graph back to a set of reactions, other tools (Soranzo and Altafini 2009; Nagy et al. 2012; Donnell et al. 2014; Feinberg et al. 2018; Tóth et al. 2018; Yordanov et al. 2020; Feinberg 2022; Marginean et al. 2023). If critical fragments can be identified in the reduced model, it is easy to backtrack to the original set of reactions in the full model that would correspond to those in the simplified fragment. Thus, these methods may allow us to find critical fragments even in models that are initially too large for direct analysis.
The parameter independence of the reduction methods presented here is a major advantage of this approach over conventional model reduction methods. Numerical model reduction methods are tied to specific parameter values and can be highly demanding of computer resources (Maas 1998; Goussis and Valorani 2006; Golda et al. 2020). In our method, simplifications are applied to the model symbolically, and do not need to be recomputed if parameters change. Although methods based on conditions that are necessary but not sufficient cannot be guaranteed to preserve model behavior, we have seen here that a model can be pared down to a highly simplified core of reactions while preserving major features of the bifurcation diagram. Analytic model reduction methods based on the application of steady-state or equilibrium approximations can also be applied symbolically, but for these methods, substantial analysis or numerical computations are required to determine whether or not they will preserve dynamics (Flach and Schell 2006; Boie et al. 2016).
7.2 Aggressive reductions
The conservative reductions described in the theorems of Sect. 5.1 consist of a set of graph operations that preserve the coefficients of fragments post-reduction. One of the advantages of conservative reductions is that they avoid changing anything in the bipartite graph that would change the number or parities of cycles, and thus can be applied without knowing which fragment, if any, is critical. However, in our first dalliance with these ideas, we carried out reductions that violated the strict conditions of the theorems presented in this contribution while retaining the bistable dynamics of a model for the development of left-right asymmetry (Roussel and Roussel 2018). This observation led us to propose the possibility of critical-fragment-aware aggressive reductions in Sect. 6.4. Aggressive reductions are only possible when we are able to determine the critical fragment leading to instability in a model. In these cases, we can apply a set of reductions that removes species inessential to the criticality of an identified critical fragment using similar rules as in conservative reductions but allowing for, e.g., the elimination of entry vertices. Provided these reductions don’t disconnect a fragment from biochemically essential interactions, they may yield usable reduced models. For example, in Roussel and Roussel (2018), a scaffold protein (Smad4) was eliminated in the reduction process, showing that the key network property required for bistability was the inclusion of two Smad2 molecules in the transcription factor, and not the assembly of the two units of Smad2 on the scaffold. Because aggressive reductions are much more risky in terms of potentially changing the behavior of the model, we would suggest only undertaking them if it is already known prior to their application that the model displays the desired dynamics. It can then be verified that the aggressively reduced model (possibly with some changes in the parameter values) retains these dynamics. The preservation of a critical fragment, even if aspects of the model outside of this fragment are dramatically changed, suggests that aggressive reductions are not only possible but likely to retain at least some aspects of the original dynamics, but again we note that procedures based on necessary but not sufficient conditions can provide no guarantees.
It may be questioned why we would engage in aggressive model reductions if we already know the model’s dynamics. There are at least two answers to this question. The first is that model reduction may yield insights into the dynamical roles of various species, as it did for the Smad2/Smad4 proteins in the left-right asymmetry model we studied previously. The other is that the reduced model will likely be more efficiently simulated, which may facilitate certain compute-heavy tasks such as the numerical solution of reaction-diffusion equations.
It must be noted that even fairly innocent-looking simplifications can alter the system’s dynamics, particularly if they involve species in the critical fragment. For example, it might be tempting to remove the sink R\(_{6\text {b}}\) in the model studied in Sect. 6, reasoning that the sink for the free enzyme, R\(_{6\text {a}}\), might be sufficient. However, setting \(k_{6b}=0\) eliminates bistability. To understand why, we computed the \(k_{\textrm{in}}\times k_{6b}\) phase diagram, shown in Fig. 18a. As \(k_{6b}\rightarrow 0\), the upper saddle-node bifurcation tends towards a limiting value. This does not in itself suggest the loss of bistability. However, if we look at the value of [NO] along the saddle-node curve, we see in Fig. 18b that it tends to zero as \(k_{6b}\rightarrow 0\). Thus, the lower branch of stable steady states drops out of the physically realizable orthant if we eliminate reaction R\(_{6\text {b}}\). This reinforces the potential importance of entry vertices and the risks associated with trying to simplify too aggressively in their vicinity.
7.3 Automation
It should be possible to automate the reduction process. Indeed, the conservative reduction process mostly involves a search for edges or paths within cycles satisfying the conditions of the theorems. The necessary search algorithms are extensions of algorithms that already exist for drawing subgraphs and evaluating their coefficients (Walther et al. 2014). Aggressive reductions are, as noted above, more delicate, but should also be automatable within software that enumerates fragments such as GraTeLPy. The output of a program for carrying out graph-based model reduction could be a hierarchy of models, ranging from models that only use conservative reductions, to models that apply increasingly risky reductions. The search for critical fragments could be carried out after the application of conservative reductions. Indeed, conservative reductions could be built into such a program as a strategy for accelerating the search for critical fragments.
One complication that would have to be addressed in automating the model reduction process is whether the user wants a reduced model, or all reduced models of the same size. Consider the case of a unidirectional cycle containing exactly two edges that enforce the directionality. Assuming both are removable according to our theorems, it would be possible to remove only one of the two edges since the remaining edge is required to maintain the directionality of the cycle. Thus, there would be two possible reduced models, considering only this one cycle, each corresponding to a different edge having been removed. In a sufficiently large model, there would presumably be many such choices, and thus a significant number of possible reduced models. Other methods can also run into issues with non-uniqueness of the reduced model, as in the work of Apri et al. (2012) for example.
This last issue would be somewhat mitigated if the user were able to specify species or reactions that must be retained in all reduced models. For example, if there are observations on the expression of certain proteins, or measurements of some rates of reaction, one would normally want to retain those specific proteins or reactions in the reduced model. These constraints would limit the search space for reduced models, and might even in some cases lead to unique reduced models where there would otherwise have been many.
7.4 Limitations
Graph-theoretical methods, and qualitative stability analysis methods in general, give necessary and not sufficient conditions for oscillations, bistability or other behaviors (Schlosser and Feinberg 1994; Feinberg 1995a; Thomas and Kaufman 2001; Craciun and Feinberg 2005, 2006b, a; Craciun et al. 2006; Mincheva and Roussel 2006, 2007a, b; Mincheva 2011; Mincheva and Craciun 2013; Soliman 2013; Banaji and Pantea 2016; Culos et al. 2016; Kaltenbach 2020; Okada et al. 2021), although sufficient conditions can be obtained for systems satisfying additional criteria (Feinberg 1995b; Pérez Millán and Dickenstein 2018). Preserving a necessary condition cannot of itself guarantee that the reduced model will preserve the original system’s dynamics. While our example shows that these methods are surprisingly robust, one can imagine situations where, for example, the loss of saturable kinetics following the elimination of an enzyme-substrate complex sufficiently alters the dynamics that bistability is lost. Along similar lines, even if the bifurcation structure is retained, the vector field may be changed in ways that change some aspects of the dynamics, as we saw in the reduction of the original model to the model of Fig. 12, which dramatically changed the parametric dependence of the upper steady state (Figs. 8 and 15).
A more serious criticism is that the elimination of pairs of negative paths does not necessarily yield a bipartite graph with a chemical interpretation that is easily related to the original chemistry, unlike the elimination of edges in positive paths. Taking for example the transformation of Fig. 6, the reaction \(\textrm{A}_1\rightarrow a\textrm{A}_4\) that appears in the reduced cycle in panel (b) is the result of the following linear transformation of the reactions: \(R_1+aR_2-aR_3\). In the cycle in panel (a), \(\textrm{A}_1\) is indirectly responsible for preventing the consumption of \(\textrm{A}_4\) by being a precursor of \(\textrm{A}_2\) which in turn participates in a reaction that consumes \(\textrm{A}_3\), the latter being ‘fuel’ in reaction \(\textrm{R}_3\) that also consumes \(\textrm{A}_4\). This is a very different kind of relationship than the direct generation of \(\textrm{A}_4\) from \(\textrm{A}_1\)! However, the point of Theorems 6 and 8 is that the models that result from these transformations are dynamically equivalent, regardless of their chemical interpretation.
It may turn out to be difficult to find systems to which Theorems 6 and 8 can be applied, because the reaction vertex of a negative path in a cycle will generally have a product outside the cycle, i.e. the reaction vertex will almost always be an entry vertex. The exception, which might be described as ‘mutual annihilation reactions’, would occur when the reaction forms a sink species. The trivial sink species vertex can be eliminated prior to the analysis since it will have no influence on the reaction. The reaction vertex then no longer appears as an entry vertex. However, having two such reactions in a cycle so that the simplification can proceed would be an even more unusual event. There may be cases where an aggressive simplification can be applied given knowledge of the critical fragment, but even then eliminating entry vertices can cause difficulties reconnecting the graph. For the moment, we do not have a real example where the negative path simplifications could be applied.
7.5 Potential future developments of dynamics-preserving model reduction methods
The basic idea exploited for model reduction in this contribution, the preservation of key properties of a network that determine the dynamic potential of a model, could be applied in the context of other qualitative stability analysis methods. As an example, we consider Chemical Reaction Network Theory (CRNT) (Schlosser and Feinberg 1994; Feinberg 1995a, b; Tóth et al. 2018; Feinberg 2022).
In CRNT, the stoichiometric combination of reactants or products appearing on one side of a reaction arrow is known as a “complex”. In one version of CRNT, a bipartite graph called a Species-Reaction (SR) graph is constructed. Like the Volpert-Ivanova bipartite graphs used in our work (Vol’pert 1972; Ivanova 1979b), SR graphs have distinct species and reaction vertices. The edges, however, are undirected and labeled both with the stoichiometric coefficient (as in Volpert-Ivanova graphs) and with the complex in which a species participates in the corresponding reaction. In an SR graph, a “c-pair” (complex pair) is defined as a pair of edges that meet at a reaction vertex and have the same complex label (the complexes along the edges) (Craciun and Feinberg 2006b). A c-pair could thus denote a pair of edges from co-reactants, i.e. a negative path in our nomenclature, or from two products of a reaction. If a cycle has an even or odd number of c-pairs, it is an “e-cycle” or “o-cycle,” respectively. An “s-cycle” is a cycle for which the result of alternately multiplying and dividing the coefficients along the cycle is 1. Lastly, two cycles have a “species-to-reaction intersection” (S-to-R intersection) if the common edges of the two cycles constitute a path that begins at a species vertex and ends at a reaction vertex. The simplest example of an S-to-R intersection in our terminology would be a single reactant edge shared by two cycles. The following is the main result of Craciun and Feinberg (2006b):
Theorem
(Craciun and Feinberg (2006b)) Consider a reaction network such that in its SR graph
-
i
each cycle is an o-cycle or an s-cycle, and
-
ii
no two e-cycles have an S-to-R intersection.
Then, taken with mass action kinetics, the reaction network does not have the capacity for multiple positive steady states.
Violating at least one of these conditions is therefore a necessary condition for multistability. Accordingly, a dynamics-preserving model reduction method analogous to the work presented here could be developed based on SR graphs in which we would avoid destroying e-cycles or S-to-R intersections. It would be very interesting to pursue this line of thought and to see if the reductions suggested by the two methods are similar or different. Similar comments could be made about other methods of qualitative stability analysis that provide conditions for multistability or oscillations (Thomas and Kaufman 2001; Soliman 2013; Blanchini et al. 2014; Banaji and Pantea 2016; Culos et al. 2016; Kaltenbach 2020; Okada et al. 2021; Vassena and Stadler 2024): each of these could give rise to a dynamics-preserving model-reduction method by ensuring that appropriate properties of the model are retained during the reduction process.
While we have stressed the essential difference between a parameter-free, network-structure-based reduction method that focuses on the dynamical repertoire of a model on the one hand, and intrinsically parameter-dependent methods such as time-scale-based reduction methods on the other, there may be some connections that would be worth exploring. Consider for example, the reduction of reactions (8c) and (8d) to (9a9b). We previously described this reduction as a simple sum of reactions, without giving consideration to any factor other than the placement of these reactions within the bipartite graph. However, we could also arrive at this reduced reaction if we consider reaction (8c) to be rate-limiting, i.e. by applying a time-scale-based reduction method assuming that reaction (8c) is, within the region of parameter space and phase space where this model evolves, much slower than reaction (8d). Thus, there will be cases where the results of a time-scale-based analysis will result in the same simplified model as our network-structure-based methods. In these cases, not only will the reduced models coincide, but the parameters of the reduced model will either be rate constants from the original model or easily computed effective rate constants. However, our method will also apply in cases where the time-scale-based analysis would suggest that reduction is not indicated. Our method seeks to preserve the potential for certain bifurcations, but says nothing in general about the parameter values at which these bifurcations will occur and it may be necessary, as we saw in the model where we reduced gene expression to the single reaction (10), to make large changes to the parameters in order to observe a bifurcation diagram in reasonable agreement with that of the original model. Despite some of the differences between time-scale-based and network-structure-based reduction methods, the relationship between the two would seem like a fertile area for investigation.
The reduction of reactions (8c) and (8d) to (9a9b) raises another issue in terms of the contact between time-scale-based methods and network-structure-based methods: At any given concentration of \(\textrm{Hmp}\cdot \textrm{O}_2\), the reaction pair (8c)–(8d) will saturate if the NO concentration becomes sufficiently high. On the other hand, the bimolecular reaction (9a9b) does not saturate. The loss of saturable enzyme catalysis in favor of bimolecular catalysis in the application of our graph simplification procedure is a common occurrence [this contribution and Roussel and Roussel (2018)]. Time-scale-based methods would tell us that a bimolecular-like rate of catalysis is only recovered at very low substrate concentrations, but our method suggests that, in models where our methods would allow this simplification, its applicability does not depend on the relative concentrations of enzyme and substrate. In other words, in models where the enzyme and enzyme-substrate complex have sufficiently simple connections to the other species of the model, saturability of the catalyst is not dynamically all that important. This is surprising. One would think that limitations in the rates of certain reactions would be important to the qualitative behavior of a model, and not just to quantitative features. We have examined a very limited number of examples, and we point out again that necessary but not sufficient conditions leave a great deal of room for model reductions to go awry. Nevertheless, it would seem an interesting question to ask whether “typical” biochemical oscillator or bistable models require saturable kinetics in order to generate their respective behaviors. Of course, saturation will almost certainly be important for reproducing experimental waveforms or other time-dependent phenomena that depend on the details of the vector field. But, as we have noted above, there are often interesting insights to be gained from studying a reduced model even if it is not fully faithful to the original model’s time evolution.
Even in the context of the bipartite graph methods explored here, we do not believe that we have exhausted the set of graph reductions that could be discovered. Our approach has been to develop provably conservative methods, and then to relax the conditions for their applicability in cases where the critical fragment is known. It is likely that there are more general classes of graph transformation that will retain critical fragments. This approach to model reduction is embryonic and ripe for development.
Data availability
Data sharing not applicable to this article as no datasets were generated or analysed during the current study.
Notes
Linear dependencies between the concentrations often arise in (bio)chemical modeling due to conservation of functional groups such as phosphate groups (Schuster and Höfer 1991). Ultimately, these conservation relations are expressions of element conservation for those elements contained only in chemical species that do not cross the boundaries of the system. Each of the \(m-n\) linear conservation laws contributes a zero eigenvalue, roughly speaking because a change in the total amount of a conserved quantity (e.g. by the external addition of a small amount of one of the chemical species subject to a conservation law) is neither amplified nor annihilated by the chemical reactions.
Positive and negative cycles of Jacobian elements appear in classical methods of qualitative stability analysis, e.g. Kaufman et al. (2007); Mincheva and Roussel (2006). Although there is doubtless a connection, the cycles of classical qualitative stability analysis are defined differently from those that appear in the Ivanova theory.
We write Eqs. (6) slightly differently than Mincheva and Roussel (2007a): rather than including a factor of \((-1)^{t_g}\) in Eq. (6a), where \(t_g\) is the number of cycles in a subgraph, we insert a negative sign in Eq. (6b). This highlights the importance of positive cycles in a manner described below.
Note that a critical fragment of this order would correspond to the constant term in the characteristic polynomial after removing the \(\lambda ^{m-n}\) factor. This means that the constant term must contain at least one negative term. This result is not surprising given that the constant term of the characteristic polynomial must pass through zero in a saddle-node bifurcation (Kuznetsov 1998).
This would correspond to a term in a coefficient \(c_k\) of the characteristic polynomial with \(k<n\). In other words, we require at least one term in some \(c_k\), \(k<n\), to be negative.
The story is a little more complicated for a fragment containing non-unidirectional cycles. These will generally allow multiple edge subgraphs. Focusing on unidirectional cycles therefore guards against some of the subtle issues that arise in non-unidirectional cycles. We note in passing that our definition of fragments, which includes all of the subgraphs that can be made from a set of vertices, differs from that of Walther et al. (2014), who define fragments by their edge graphs. We call the latter ‘WHM fragments’. The disadvantage of the WHM definition of a fragment is that it results in many false critical fragments since it is not uncommon for one edge graph to generate a WHM fragment with a negative coefficient while another edge graph using the same vertices and therefore contributing to the same coefficient of the Jacobian generates a WHM fragment with a positive coefficient that makes the overall Jacobian coefficient non-negative.
There are however some important subtleties in the case of removing negative paths to which we will return later.
Although these satisfy all of the rules for inclusion in subgraphs, we will see later that they do not contribute to the dynamics.
Exactly how large \(\nu \) needs to be depends on what is to either side of the vertices to be eliminated, notably whether the positive path is adjacent to a negative path.
Note that it is possible to have \(R_{p_h} \equiv R_{s_f}\) and/or \(R_{s_h} \equiv R_{p_f}\). In either scenario, this theorem holds.
In Sect. 6.4, we will discuss aggressive reductions, which use the same general principles discussed here along with a knowledge of the critical fragment to carry out reductions that would not be allowed by our conservative theorems. Aggressive reductions thus lack the guarantees associated with theorem-supported reductions. Aggressive reductions may eliminate species outside of the cycle so that the net reaction after reduction may be missing some species. Accordingly, aggressive reductions may change a mechanism in ways that cannot be rationalized as simple superpositions of reactions, precisely because they take no account of species outside a given cycle in a critical fragment.
References
Apri M, de Gee M, Molenaar J (2012) Complexity reduction preserving dynamical behavior of biochemical networks. J Theor Biol 304:16–26. https://doi.org/10.1016/j.jtbi.2012.03.019
Banaji M, Pantea C (2016) Some results on injectivity and multistationarity in chemical reaction networks. SIAM J Appl Dyn Syst 15:807–869. https://doi.org/10.1137/15M1034441
Battelli F, Lazzari C (1985) On the pseudo-steady-state approximation and Tikhonov theorem for general enzyme systems. Math Biosci 75:229–246. https://doi.org/10.1016/0025-5564(85)90039-2
Blanchini F, Franco E, Giordano G (2014) A structural classification of candidate oscillatory and multistationary biochemical systems. Bull Math Biol 76:2542–2569. https://doi.org/10.1007/s11538-014-0023-y
Boie S, Kirk V, Sneyd J, Wechselberger M (2016) Effects of quasi-steady-state reduction on biophysical models with oscillations. J Theor Biol 393:16–31. https://doi.org/10.1016/j.jtbi.2015.12.011
Borghans JAM, De Boer RJ, Segel LA (1996) Extending the quasi-steady state approximation by changing variables. Bull Math Biol 58:43–63. https://doi.org/10.1007/BF02458281
Briggs GE, Haldane JBS (1925) A note on the kinetics of enzyme action. Biochem J 19:338–339. https://doi.org/10.1042/bj0190338
Clarke BL (1974) Graph theoretic approach to the stability analysis of steady state chemical reaction networks. J Chem Phys 60:1481–1492. https://doi.org/10.1063/1.1681221
Conradi C, Mincheva M (2017) Graph-theoretic analysis of multistationarity using degree theory. Math Comput Simul 133:76–90. https://doi.org/10.1016/j.matcom.2015.08.010
Conzelmann H, Fey D, Gilles ED (2008) Exact model reduction of combinatorial reaction networks. BMC Syst Biol 2:78. https://doi.org/10.1186/1752-0509-2-78
Coxson PG, Bischoff KB (1987) Lumping strategy. 1. Introductory techniques and applications of cluster analysis. Ind Eng Chem Res 26:1239–1248. https://doi.org/10.1021/ie00066a031
Craciun G, Feinberg M (2005) Multiple equilibria in complex chemical reaction networks: I. The injectivity property. SIAM J Appl Math 65:1526–1546. https://doi.org/10.1137/S0036139904440278
Craciun G, Feinberg M (2006) Multiple equilibria in complex chemical reaction networks: extensions to entrapped species models. IEE Proc Syst Biol 153:179–186. https://doi.org/10.1049/ip-syb:20050093
Craciun G, Feinberg M (2006) Multiple equilibria in complex chemical reaction networks: II. The species-reaction graph. SIAM J Appl Math 66:1321–1338. https://doi.org/10.1137/050634177
Craciun G, Tang Y, Feinberg M (2006) Understanding bistability in complex enzyme-driven reaction networks. Proc Natl Acad Sci USA 103:8697–8702. https://doi.org/10.1073/pnas.0602767103
Crack JC, Stapleton MR, Green J et al (2013) Mechanism of [4Fe-4S](Cys)\(_4\) cluster nitrosylation is conserved among NO-responsive regulators. J Biol Chem 288:11492–11502. https://doi.org/10.1074/jbc.M112.439901
Culos GJ, Olesky DD, van den Driessche P (2016) Using sign patterns to detect the possibility of periodicity in biological systems. J Math Biol 72:1281–1300. https://doi.org/10.1007/s00285-015-0906-z
Dale KJ, Pourquié O (2000) A clock-work somite. BioEssays 22:72–83. https://doi.org/10.1002/(SICI)1521-1878(200001)22:1<72::AID-BIES12>3.0.CO;2-S
Danø S, Madsen MF, Schmidt H, Cedersund G (2006) Reduction of a biochemical model with preservation of its basic dynamical properties. FEBS J 273:4862–4877. https://doi.org/10.1111/j.1742-4658.2006.05485.x
de Donder T (1937) Sur la vitesse réactionelle. Bull Classe Sci Acad Roy Belg 23:936–941
Delbrück M (1949) In: Unités biologiques douées de continuité génétique, Colloques internationaux du Centre national de la recherche scientifique, vol VIII. Centre national de la recherche scientifique, Paris, pp 33–34
Donnell P, Banaji M, Marginean A, Pantea C (2014) CoNtRol: an open source framework for the analysis of chemical reaction networks. Bioinformatics 30:1633–1634. https://doi.org/10.1093/bioinformatics/btu063
Dutta C, Dasgupta A, Das J (1984) On thermodynamic and kinetic equivalence of chemical systems. Proc Indian Acad Sci (Chem Sci) 93:817–830. https://doi.org/10.1007/BF02866343
Eilertsen J, Schnell S (2020) The quasi-steady-state approximations revisited: timescales, small parameters, singularities, and normal forms in enzyme kinetics. Math Biosci 325:108339. https://doi.org/10.1016/j.mbs.2020.108339
Ermakov GL (2003) A theoretical graph method for search and analysis of critical phenomena in biochemical systems. I. Graphical rules for detecting oscillators. Biochemistry (Moscow) 68:1109–1120. https://doi.org/10.1023/A:1026358628659
Ermakov GL (2003) A theoretical graph method for search and analysis of critical phenomena in biochemical systems. II. Kinetic models of biochemical oscillators including two and three substances. Biochemistry (Moscow) 68:1121–1131. https://doi.org/10.1023/A:1026310712729
Ermakov GL, Goldstein BN (2002) Simplest kinetic schemes for biochemical oscillators. Biochemistry (Moscow) 67:473–484. https://doi.org/10.1023/A:1015294208979
Ermentrout B (2002) Simulating, analyzing, and animating dynamical systems. SIAM, Philadelphia. https://doi.org/10.1137/1.9780898718195
Farkas G (1999) Kinetic lumping schemes. Chem Eng Sci 54:3909–3915. https://doi.org/10.1016/S0009-2509(99)00028-7
Feinberg M (1995) The existence and uniqueness of steady states for a class of chemical reaction networks. Arch Rational Mech Anal 132:311–370. https://doi.org/10.1007/BF00375614
Feinberg M (1995) Multiple steady states for chemical reaction networks of deficiency one. Arch Rational Mech Anal 132:371–406. https://doi.org/10.1007/BF00375615
Feinberg M (2022) Foundations of chemical reaction network theory. Applied Mathematical Sciences, vol 202. Springer, Cham. https://doi.org/10.1007/978-3-030-03858-8
Feinberg M, Ellison P, Ji H, Knight D (2018) Chemical reaction network toolbox, version 2.35. https://doi.org/10.5281/zenodo.5149265
Flach EH, Schell S (2006) Use and abuse of the quasi-steady-state approximation. IEE Proc Syst Biol 153:187–191. https://doi.org/10.1049/ip-syb:20050104
Fraser SJ (1988) The steady state and equilibrium approximations: a geometrical picture. J Chem Phys 88:4732–4738. https://doi.org/10.1063/1.454686
Gay S, Soliman S, Fages F (2010) A graphical method for reducing and relating models in systems biology. Bioinformatics 26:i575–i581. https://doi.org/10.1093/bioinformatics/btq388
Goeke A, Walcher S, Zerz E (2017) Classical quasi-steady state reduction—a mathematical characterization. Physica D 345:11–26. https://doi.org/10.1016/j.physd.2016.12.002
Golda P, Blattmann A, Neagos A et al (2020) Implementation problems of manifolds-based model reduction and their generic solution. Combust Theory Modell 24:377–406. https://doi.org/10.1080/13647830.2019.1682198
Goldstein BN, Ermakov G, Centelles JJ et al (2004) What makes biochemical networks tick? A graphical tool for the identification of oscillophores. Eur J Biochem 271:3877–3887. https://doi.org/10.1111/j.1432-1033.2004.04324.x
Gorban AN (2018) Model reduction in chemical dynamics: slow invariant manifolds, singular perturbations, thermodynamic estimates, and analysis of reaction graph. Curr Opin Chem Eng 21:48–59. https://doi.org/10.1016/j.coche.2018.02.009
Gorban AN, Karlin IV, Zinovyev AYu (2004) Constructive methods of invariant manifolds for kinetic problems. Phys Rep 396:197–403. https://doi.org/10.1016/j.physrep.2004.03.006
Gorban AN, Karlin IV, Zinovyev AYu (2004) Invariant grids for reaction kinetics. Physica A 333:106–154. https://doi.org/10.1016/j.physa.2003.10.043
Goussis DA, Valorani M (2006) An efficient iterative algorithm for the approximation of the fast and slow dynamics of stiff systems. J Comput Phys 214:316–346. https://doi.org/10.1016/j.jcp.2005.09.019
Haken H (1996) Slaving principle revisited. Physica D 97:95–103. https://doi.org/10.1016/0167-2789(96)00080-2
Horn F, Jackson R (1972) General mass action kinetics. Arch Rational Mech Anal 47:81–116. https://doi.org/10.1007/BF00251225
Huang H, Fairweather M, Griffiths JF et al (2005) A systematic lumping approach for the reduction of comprehensive kinetic models. Proc Combust Inst 30:1309–1316. https://doi.org/10.1016/j.proci.2004.08.001
Ivanova AN (1979) Conditions for the uniqueness of the stationary states of kinetic systems, connected with the structures of their reaction mechanisms. II. Open systems. Kinet Catal 20:837–840
Ivanova AN (1979) Conditions for uniqueness of the stationary states of kinetic systems, connected with the structures of their reaction mechanisms. I. Kinet Catal 20:833–837
Iwasa Y, Andreasen V, Levin S (1987) Aggregation in model ecosystems. I. Perfect aggregation. Ecol Modelling 37:287–302. https://doi.org/10.1016/0304-3800(87)90030-5
Jeong H, Tombor B, Albert R et al (2000) The large-scale organization of metabolic networks. Nature 407:651–654. https://doi.org/10.1038/35036627
Kærn M, Menzinger M, Hunding A (2000) Segmentation and somitogenesis derived from phase dynamics in growing oscillatory media. J Theor Biol 207:473–493. https://doi.org/10.1006/jtbi.2000.2183
Kaltenbach HM (2020) A unified view on bipartite species-reaction graphs and their relation to interaction graphs and qualitative dynamics of chemical reaction networks. Electron Notes Theor Comput Sci 350:73–90. https://doi.org/10.1016/j.entcs.2020.06.005
Kaufman M, Soulé C, Thomas R (2007) A new necessary condition on interaction graphs for multistationarity. J Theor Biol 248:675–685. https://doi.org/10.1016/j.jtbi.2007.06.016
Kennell D, Talkad V (1976) Messenger RNA potential and the delay before exponential decay of messages. J Mol Biol 104:285–298. https://doi.org/10.1016/0022-2836(76)90014-0
Kepler TB, Elston TC (2001) Stochasticity in transcriptional regulation: origins, consequences, and mathematical representations. Biophys J 81:3116–3136. https://doi.org/10.1016/S0006-3495(01)75949-8
Koch AJ, Meinhardt H (1994) Biological pattern formation: from basic mechanisms to complex structures. Rev Mod Phys 66:1481–1507. https://doi.org/10.1103/RevModPhys.66.1481
Kristiansen KU, Brøns M, Starke J (2014) An iterative method for the approximation of fibers in slow-fast systems. SIAM J Appl Dyn Syst 13:861–900. https://doi.org/10.1137/120889666
Kuo JCW, Wei J (1969) A lumping analysis in monomolecular reaction systems. Analysis of approximately lumpable systems. Ind Eng Chem Fundam 8:124–133. https://doi.org/10.1021/i160029a020
Kuznetsov YA (1998) Elements of applied bifurcation theory, 2nd edn. Springer, New York
Lacalli TC, Harrison LG (1978) The regulatory capacity of Turing’s model for morphogenesis, with application to slime moulds. J Theor Biol 70:273–295. https://doi.org/10.1016/0022-5193(78)90377-6
Lam SH (1993) Using CSP to understand complex chemical kinetics. Combust Sci Technol 89:375–404. https://doi.org/10.1080/00102209308924120
Laurent M, Kellershohn N (1999) Multistability: a major means of differentiation and evolution in biological systems. Trends Biochem Sci 24:418–422. https://doi.org/10.1016/S0968-0004(99)01473-5
Lee CH, Othmer HG (2010) A multi-time-scale analysis of chemical reaction networks: I. Deterministic systems. J Math Biol 60:387–450. https://doi.org/10.1007/s00285-009-0269-4
Li G, Rabitz H (1989) A general analysis of exact lumping in chemical kinetics. Chem Eng Sci 44:1413–1430. https://doi.org/10.1016/0009-2509(89)85014-6
Li G, Tomlin AS, Rabitz H, Tóth J (1993) Determination of approximate lumping schemes by a singular perturbation method. J Chem Phys 99:3562–3574. https://doi.org/10.1063/1.466153
Li G, Rabitz H, Tóth J (1994) A general analysis of exact nonlinear lumping in chemical kinetics. Chem Eng Sci 49:343–361. https://doi.org/10.1016/0009-2509(94)87006-3
Lipshtat A, Loinger A, Balaban NQ, Biham O (2006) Genetic toggle switch without cooperative binding. Phys Rev Lett 96:188101. https://doi.org/10.1103/PhysRevLett.96.188101
Lubitz T, Liebermeister W (2019) Parameter balancing: consistent parameter sets for kinetic metabolic models. Bioinformatics 35:3857–3858. https://doi.org/10.1093/bioinformatics/btz129
Maas U (1998) Efficient calculation of intrinsic low-dimensional manifolds for the simplification of chemical kinetics. Comput Visual Sci 1:69–81. https://doi.org/10.1007/s007910050007
Maas U, Pope SB (1992) Simplifying chemical kinetics: intrinsic low-dimensional manifolds in composition space. Combust Flame 88:239–264. https://doi.org/10.1016/0010-2180(92)90034-M
Marginean A, Pantea C, Banaji M et al (2023) CoNtRol, version 1.5.3. URL https://control.math.wvu.edu
Mincheva M (2011) Oscillations in biochemical reaction networks arising from pairs of subnetworks. Bull Math Biol 73:2277–2304. https://doi.org/10.1007/s11538-010-9620-6
Mincheva M, Craciun G (2013) Graph-theoretic conditions for zero-eigenvalue Turing instability in general chemical reaction networks. Math Biosci Eng 10:1207–1226. https://doi.org/10.3934/mbe.2013.10.1207
Mincheva M, Roussel MR (2006) A graph-theoretic method for detecting potential Turing bifurcations. J Chem Phys 125:204102. https://doi.org/10.1063/1.2397073
Mincheva M, Roussel MR (2007) Graph-theoretic methods for the analysis of chemical and biochemical networks. I. Multistability and oscillations in ordinary differential equation models. J Math Biol 55:61–86. https://doi.org/10.1007/s00285-007-0099-1
Mincheva M, Roussel MR (2007) Graph-theoretic methods for the analysis of chemical and biochemical networks. II. Oscillations in networks with delays. J Math Biol 55:87–104. https://doi.org/10.1007/s00285-007-0098-2
Müller P, Rogers KW, Jordan BM et al (2012) Differential diffusivity of Nodal and Lefty underlies a reaction-diffusion patterning system. Science 336:721–724. https://doi.org/10.1126/science.1221920
Nagy AL, Papp D, Tóth J (2012) ReactionKinetics—a Mathematica package with applications. Chem Eng Sci 83:12–23. https://doi.org/10.1016/j.ces.2012.01.039
Nakamura T, Mine N, Nakaguchi E et al (2006) Generation of robust left-right asymmetry in the mouse embryo requires a self-enhancement and lateral-inhibition system. Dev Cell 11:495–504. https://doi.org/10.1016/j.devcel.2006.08.002
Noethen L, Walcher S (2011) Tikhonov’s theorem and quasi-steady state. Discrete Contin Dyn Syst B 16:945–961. https://doi.org/10.3934/dcdsb.2011.16.945
Okada T, Mochizuki A, Furuta M, Tsai JC (2021) Flux-augmented bifurcation analysis in chemical reaction network systems. Phys Rev E 103:062212. https://doi.org/10.1103/PhysRevE.103.062212
Okeke BE, Roussel MR (2015) An invariant-manifold approach to lumping. Math Model Nat Phenom 10:149–167. https://doi.org/10.1051/mmnp/201510312
Pérez Millán M, Dickenstein A (2018) The structure of MESSI biological systems. SIAM J Appl Dyn Syst 17:1650–1682. https://doi.org/10.1137/17M1113722
Petzold L, Zhu W (1999) Model reduction for chemical kinetics: an optimization approach. AIChE J 45:869–886. https://doi.org/10.1002/aic.690450418
Prescott TP, Papachristodoulou A (2014) Layered decomposition for the model order reduction of timescale separated biochemical reaction networks. J Theor Biol 356:113–122. https://doi.org/10.1016/j.jtbi.2014.04.007
Prigogine I (1946) Remarque sur le principe de réciprocité d’Onsager et le couplage des réactions chimiques. Bull Classe Sci Acad Roy Belg 32:30–35
Prigogine I, Lefever R (1968) Symmetry breaking instabilities in dissipative systems. II. J Chem Phys 48:1695–1700. https://doi.org/10.1063/1.1668896
Radulescu O, Gorban AN, Zinovyev A, Lilienbaum A (2008) Robust simplifications of multiscale biochemical networks. BMC Syst Biol 2:86. https://doi.org/10.1186/1752-0509-2-86
Radulescu O, Gorban AN, Zinovyev A, Noel V (2012) Reduction of dynamical biochemical reactions networks in computational biology. Front Genet 3:131. https://doi.org/10.3389/fgene.2012.00131
Radulescu O, Vakulenko S, Grigoriev D (2015) Model reduction of biochemical reactions networks by tropical analysis methods. Math Model Nat Phenom 10:124–138. https://doi.org/10.1051/mmnp/201510310
Rao S, van der Schaft A, van Eunen K et al (2014) A model reduction method for biochemical reaction networks. BMC Syst Biol 8:52. https://doi.org/10.1186/1752-0509-8-52
Raspopovic J, Marcon L, Russo L, Sharpe J (2014) Digit patterning is controlled by a Bmp-Sox9-Wnt Turing network modulated by morphogen gradients. Science 345:566–570. https://doi.org/10.1126/science.1252960
Roussel MR (1996) The use of delay differential equations in chemical kinetics. J Phys Chem 100:8323–8330. https://doi.org/10.1021/jp9600672
Roussel MR (2019) A delayed mass-action model for the transcriptional control of Hmp, an NO detoxifying enzyme, by the iron-sulfur protein FNR. In: Valmorbida G, Seuret A, Boussaada I, Sipahi R (eds) Delays and interconnections: methodology, algorithms and applications. Advances in Delays and Dynamics, vol 10. Springer, Cham, pp 215–230. https://doi.org/10.1007/978-3-030-11554-8_14
Roussel MR (2019) Nonlinear dynamics: a hands-on introductory survey. Morgan & Claypool, San Rafael. https://doi.org/10.1088/2053-2571/ab0281
Roussel CJ, Roussel MR (2018) A mathematical model of the biochemical network underlying left-right asymmetry establishment in mammals. BioSystems 173:281–297. https://doi.org/10.1016/j.biosystems.2018.10.003
Schauer M, Heinrich R (1983) Quasi-steady-state approximation in the mathematical modeling of biochemical reaction networks. Math Biosci 65:155–170. https://doi.org/10.1016/0025-5564(83)90058-5
Schlosser PM, Feinberg M (1994) A theory of multiple steady states in isothermal homogeneous CFSTRs with many reactions. Chem Eng Sci 49:1749–1767. https://doi.org/10.1016/0009-2509(94)80061-8
Schuster S, Höfer T (1991) Determining all extreme semi-positive conservation relations in chemical reaction systems: a test criterion for conservativity. J Chem Soc Faraday Trans 87:2561–2566. https://doi.org/10.1039/FT9918702561
Segel LA, Slemrod M (1989) The quasi-steady-state assumption: a case study in perturbation. SIAM Rev 31:446–477. https://doi.org/10.1137/1031091
Slin’ko MG, Slin’ko MM (1978) Self-oscillations of heterogeneous catalytic reaction rates. Catal Rev Sci Eng 17:119–153. https://doi.org/10.1080/03602457808080880
Snowden TJ, van der Graaf PH, Tindall MJ (2017) Methods of model reduction for large-scale biological systems: a survey of current methods and trends. Bull Math Biol 79:1449–1486. https://doi.org/10.1007/s11538-017-0277-2
Sokolowski TR, Walczak AM, Bialek W, Tkačik G (2016) Extending the dynamic range of transcription factor action by translational regulation. Phys Rev E 93:022404. https://doi.org/10.1103/PhysRevE.93.022404
Soliman S (2013) A stronger necessary condition for the multistationarity of chemical reaction networks. Bull Math Biol 75:2289–2303. https://doi.org/10.1007/s11538-013-9893-7
Soranzo N, Altafini C (2009) ERNEST: a toolbox for chemical reaction network theory. Bioinformatics 21:2853–2854. https://doi.org/10.1093/bioinformatics/btp513
Stagni A, Cuoci A, Frassoldati A et al (2014) Lumping and reduction of detailed kinetic schemes: an effective coupling. Ind Eng Chem Res 53:9004–9016. https://doi.org/10.1021/ie403272f
Steuer R, Gross T, Selbig J, Blasius B (2006) Structural kinetic modeling of metabolic networks. Proc Natl Acad Sci USA 103:11868–11873. https://doi.org/10.1073/pnas.0600013103
Stiefenhofer M (1998) Quasi-steady-state approximation for chemical reaction networks. J Math Biol 36:593–609. https://doi.org/10.1007/s002850050116
Takashima Y, Ohtsuka T, González A, Kageyama R (2011) Intronic delay is essential for oscillatory expression in the segmentation clock. Proc Natl Acad Sci USA 108:3300–3305. https://doi.org/10.1073/pnas.1014418108
Thomas R (1994) The role of feedback circuits: positive feedback circuits are a necessary condition for positive real eigenvalues of the Jacobian matrix. Ber Bunsenges Phys Chem 98:1148–1151. https://doi.org/10.1002/bbpc.19940980916
Thomas R, Kaufman M (2001) Multistationarity, the basis of cell differentiation and memory. I. Structural conditions of multistationarity and other nontrivial behavior. Chaos 11:170–179. https://doi.org/10.1063/1.1350439
Tomlin AS, Pilling MJ, Turányi T et al (1992) Mechanism reduction for the oscillatory oxidation of hydrogen: sensitivity and quasi-steady-state analyses. Combust Flame 91:107–130. https://doi.org/10.1016/0010-2180(92)90094-6
Tóth J, Nagy AL, Papp D (2018) Reaction kinetics: exercises, programs and theorems. Springer, New York. https://doi.org/10.1007/978-1-4939-8643-9
Trofimenkoff EAM, Roussel MR (2020) Small binding-site clearance delays are not negligible in gene expression modeling. Math Biosci 325:108376. https://doi.org/10.1016/j.mbs.2020.108376
Turing AM (1952) The chemical basis of morphogenesis. Philos Trans R Soc London Ser B 237:37–72. https://doi.org/10.1098/rstb.1952.0012
Tyson JJ (1975) Classification of instabilities in chemical reaction systems. J Chem Phys 62:1010–1015. https://doi.org/10.1063/1.430567
Ünal HU, Roussel MR, Boussaada I, Niculescu SI (2021) Insights from a qualitative analysis of a gene expression model with delays. IFAC PaperOnLine 54(9):770–775. https://doi.org/10.1016/j.ifacol.2021.06.174
van Slyke DD, Cullen GE (1914) The mode of action of urease and of enzymes in general. J Biol Chem 19:141–180. https://doi.org/10.1016/S0021-9258(18)88300-4
Vassena N, Stadler PF (2024) Unstable cores are the source of instability in chemical reaction networks. Proc R Soc A 480:20230694. https://doi.org/10.1098/rspa.2023.0694
Vol’pert AI (1972) Differential equations on graphs. Math USSR Sb 17:571–582
Walsh CT (1984) Suicide substrates, mechanism-based enzyme inactivators: recent developments. Ann Rev Biochem 53:493–535. https://doi.org/10.1146/annurev.bi.53.070184.002425
Walther GR, Hartley M, Mincheva M (2014) GraTeLPy: graph-theoretic linear stability analysis. BMC Syst Biol 8:22. https://doi.org/10.1186/1752-0509-8-22
Wei J, Kuo JCW (1969) A lumping analysis in monomolecular reaction systems. Analysis of the exactly lumpable system. Ind Eng Chem Fundam 8:114–123. https://doi.org/10.1021/i160029a019
Yordanov P, Stelling J, Otero-Muras I (2020) BioSwitch: a tool for the detection of bistability and multi-steady state behaviour in signalling and gene regulatory networks. Bioinformatics 36:1640–1641. https://doi.org/10.1093/bioinformatics/btz746
Zeigarnik AV, Temkin ON (1994) A graph-theoretical model of complex reaction mechanisms: bipartite graphs and the stoichiometry of complex reactions. Kinet Catal 35:647–655
Acknowledgements
We would like to thank Dr Maya Mincheva of Northern Illinois University for useful conversations. This work was funded by the Natural Sciences and Engineering Research Council of Canada and by the University of Lethbridge Research Fund.
Author information
Authors and Affiliations
Contributions
MR Roussel: Conceptualization; Formal analysis; Funding acquisition; Resources; Supervision; Validation; Visualization; Writing—Review and Editing. T Soares: Formal analysis; Investigation; Visualization; Writing—Original Draft.
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Appendices
Ivanova’s terminology for bipartite graphs, subgraphs and fragments
This section outlines the terminology established by Mincheva and Roussel (2007a) based on the English translation of Ivanova’s original Russian terminology (Ivanova 1979b).
- (Reactant) Edge::
-
A directed arrow from a reactant species vertex A\(_k\) to a reaction vertex R\(_j\), denoted [A\(_k\),R\(_j\)]. In the classic version of the theory, these are just called ‘edges’, with no qualifier. Note that for the purposes of model reduction, we will also need to discuss product edges, i.e. edges from a reaction to a product vertex (Sect. 3).
- Edge weight::
-
The weight of an edge (reactant or product) is the corresponding stoichiometric coefficient.
- Positive path::
-
A sequence of directed arrows from a reactant species vertex to a reaction vertex and then from that same reaction vertex to a product species vertex, denoted [A\(_k\),R\(_j\),A\(_i\)].
- Negative path::
-
A sequence of directed arrows from a reactant species vertex to a reaction vertex, followed by an edge traversed against the direction of its arrow to another reactant species vertex, denoted \(\overline{[\text {A}_k,\text {R}_j,\text {A}_i]}\).
- Cycle::
-
A sequence of paths that start and end at the same species vertex. A cycle is said to be positive or to have positive parity if it contains an even number of negative paths, while a cycle is said to be negative or to have negative parity if it contains an odd number of negative paths.
- Subgraph::
-
A union of edges and cycles in which each species is at the origin of only one edge or path. Each subgraph has a coefficient \(K_g\) computed by Eq. (6).
- Fragment::
-
A union of subgraphs that share the same vertices. Each fragment corresponds to a term in the characteristic polynomial of the form (3). The coefficient of a fragment, \(K_{S_k}\), is given by the sum of all of its subgraphs’ coefficients according to Eq. (5).
- Critical fragment::
-
A fragment with a negative coefficient.
- Order (of a cycle, subgraph or fragment)::
-
Number of chemical species vertices therein.
Graph-theoretic stability analysis: an example
As an illustration of the Ivanova style of graph-theoretical stability analysis, we consider the Brusselator (Prigogine and Lefever 1968), a model that displays positive-feedback oscillations. In its simplest form, the Brusselator can be written as the following mass-action system:




The law of mass action gives us monomial reaction rates of the form \(R_j=k_jx^{\alpha _{jx}}y^{\alpha _{jy}}\) for a two-variable system such as the Brusselator, where \(x=\mathrm {[X]}\), \(y=\mathrm {[Y]}\) and \(R_j\) is the rate of reaction \(\textrm{R}_j\). Due to this monomial form, the elements of the Jacobian matrix can be written
and similarly for Jacobian elements with respect to y. For the Brusselator, using the standard method of linear stability analysis for ODEs (Roussel 2019b), we get the characteristic polynomial
where \((x^*,y^*)\) is the unique steady state of this model. It is easy to show that the negative term \(-2R_3/x\) in the linear coefficient of this characteristic polynomial is necessary for the occurrence of an Andronov-Hopf bifurcation.
Bipartite graph corresponding to the Brusselator reactions (11)
The graph-theoretical analysis begins by building the bipartite graph associated with the set of reactions (11), which is shown in Fig. 19. Since there are two independent chemical species (no conservation relations), only fragments up to order 2 are relevant. Note that the characteristic polynomial is, correspondingly, of degree 2. Each fragment is in a one-to-one correspondence with a term in the characteristic polynomial. Fragments are unions of subgraphs that share a set of vertices. To identify fragments and their corresponding subgraphs, it is often easiest to start by choosing some vertices, adding all of the arrows that connect them in the bipartite graph, and then constructing all of the subgraphs that can be found in the chosen set. Consider for example the fragment \(S_2\) illustrated in Fig. 20. The subgraphs are unions of (reactant) edges and cycles that include all of the vertices in this fragment. For any given subgraph, we denote by \({\mathcal {E}}\) the set of edges and by \({\mathcal {C}}\) the set of cycles in the subgraph. Table 2 shows all of the subgraphs of the fragment \(S_2\). For example, subgraph \(g_1\) consists of the two edges \([\textrm{X},R_3]\) and \([\textrm{Y},R_3]\). \(K_g\) for this subgraph is calculated from the stoichiometric coefficients of these edges using Eq. (6a). Similarly, \(g_2\) consists of the cycle \([\textrm{X},R_3,\textrm{X}]\) and of the edge \([\textrm{Y},R_3]\). The coefficient of the cycle is calculated using Eq. (6b), and then again we apply Eq. (6a) to get the coefficient of the subgraph. Note that the negative value of \(K_C\) implies a positive cycle. The coefficients of the subgraphs are added to give the coefficient of the fragment according to Eq. (5). For this fragment, \(K_{S_2}=0\). Correspondingly, the characteristic polynomial does not contain a term \(\frac{R_3}{x}\,\frac{R_3}{y}\).
The only critical fragment for this mechanism is \(S_1\left( \begin{array}{c} \textrm{X}\\ R_3 \end{array}\right) \), illustrated in Fig. 21. This fragment has two subgraphs, the edge subgraph \(g_1=\left\{ [\textrm{X},R_3]\right\} \) for which \(K_{g_1}=4\) and the cyclic subgraph \(g_2=\left\{ [\textrm{X},R_3,\textrm{X}]\right\} \) for which \(K_{g_2}=-6\). The coefficient of the fragment is therefore \(K_{S_1}=-2\), the coefficient of the term \(R_3/x\) in the characteristic polynomial. Note that the critical fragment extracts from the mechanism the autocatalytic (positive feedback) loop that is responsible for the appearance of an Andronov-Hopf bifurcation and of the consequent limit-cycle oscillations in the Brusselator.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Roussel, M.R., Soares, T. Graph-based, dynamics-preserving reduction of (bio)chemical systems. J. Math. Biol. 89, 42 (2024). https://doi.org/10.1007/s00285-024-02138-0
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00285-024-02138-0
Keywords
- Model reduction
- Chemical reaction networks
- Mass-action modeling
- Graph-theoretical methods
- Control of gene expression
- Nitric oxide metabolism