Abstract
A novel thermotolerant caproic acid-producing bacterial strain, Clostridium M1NH, was successfully isolated from sewage sludge. Ethanol and acetic acid at a molar ratio of 4:1 proved to be the optimal substrates, yielding a maximum caproic acid production of 3.5 g/L. Clostridium M1NH exhibited remarkable tolerance to high concentrations of ethanol (up to 5% v/v), acetic acid (up to 5% w/v), and caproic acid (up to 2% w/v). The strain also demonstrated a wide pH tolerance range (pH 5.5–7.5) and an elevated temperature optimum between 35 and 40 °C. Phylogenetic analysis based on 16S rRNA gene sequences revealed that Clostridium M1NH shares a 98% similarity with Clostridium luticellarii DSM 29923 T. The robustness of strain M1NH and its efficient caproic acid production from low-cost substrates highlight its potential for sustainable bio-based chemical production. The maximum caproic acid yield achieved by Clostridium M1NH was 1.6-fold higher than that reported for C. kluyveri under similar fermentation conditions. This study opens new avenues for valorizing waste streams and advancing a circular economy model in the chemical industry.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Caproic acid is a six-carbon saturated fatty acid that is colorless, odorless, and tasteless while soluble in water (9.7 g/L at 20 °C) and organic solvents. Caproic acid has broad potential applications. Its broad potential applications include direct use as feed additives at concentrations of 5–10 g/kg [1], antimicrobials at minimum inhibitory concentrations (MICs) ranging from 0.87 to 10.00 mg/mL [2], and plant growth promoters at concentrations of 20 mM [3]. Caproic acid can also serve as a precursor to various commodities, such as fragrances, lubricants, paint additives, and pharmaceuticals [4,5,6]. With a higher energy density (27.5 MJ/kg) and stronger hydrophobicity (log P 2.05) than its short-chain fatty acid precursors [7], caproic acid has traditionally been sourced from animal fats, plant oils, and petroleum. However, the shift toward bio-based production methods presents an eco-friendly alternative, reducing reliance on fossil resources and associated environmental impacts.
Bio-based caproic acid production is an attractive alternative to the conventional petrochemical route, as it can reduce dependency on fossil resources, lower greenhouse gas emissions, and utilize renewable biomass feedstocks. Recent advancements in bio-based caproic acid production emphasize the role of anaerobic fermentation by chain-elongating bacteria, such as Clostridium kluyveri, capable of producing caproic acid at concentrations up to 12.8 g/L [8, 9] and Megasphaera elsdenii, with reported yields of 9.7–11.4 g/L [10, 11]. Other promising strains include Caproiciproducens sp., achieving caproic acid concentrations of 3.93 g/L [12, 13], and Ruminococcaceae bacterium CPB6, producing up to 16.6 g/L [14]. These processes leverage the reverse β-oxidation pathway from short-chain fatty acids, highlighting the biotechnological potential of microbial synthesis. However, conventional fermentation processes, predominantly conducted at moderate temperatures between 25 and 35 °C, face limitations, including contamination risk and reduced efficiency.
Exploring thermotolerant microorganisms emerges as a promising frontier, potentially overcoming these challenges by facilitating fermentation at elevated temperatures. Thermotolerant yeast strains, such as Saccharomyces cerevisiae LBM-1, can improve ethanol fermentation efficiency, allowing fermentation to occur at temperatures up to 45 °C [15]. Thermotolerant bacteria could have high metabolic rates (up to threefold higher than mesophilic strains), higher end-product yields (10–15% increase), and stable enzymes (half-lives up to 10 times longer at elevated temperatures) [16]. These bacteria resist high temperatures by producing specialized proteins known as chaperonins, which are thermostable and can maintain their structural integrity at temperatures up to 70 °C [16]. Such conditions promise a reduced risk of contamination, enhanced metabolic rates (2–3 times higher), product yields (up to 20% increase), and process efficiencies (up to 30% improvement). Thermotolerant conditions also decrease the viscosity of the reaction medium by 20–50%, increase the bioavailability and solubility of organic compounds by 10–30%, increase the diffusion coefficient of substrates by 30–50%, and reduce cooling costs in commercial-scale operations by 15–25% [17]. Despite the potential, applying thermotolerant bacteria to caproic acid production remains underexplored, particularly in utilizing sewage sludge as a biomass feedstock.
Sewage sludge, rich in organic matter (60–80% of dry weight), nutrients (nitrogen 2–5%, phosphorus 1–2%), and microbial diversity (up to 1012 cells/g), represents an untapped resource for bio-based compound production [12]. Sewage sludge contains diverse caproic acid-producing bacteria, including potentially novel strains with thermotolerant capabilities that offer a sustainable bio-based production pathway. However, the functional capacity and diversity of caproic acid-producing thermotolerant bacteria within this matrix must be better understood. This study aims to screen, isolate, and characterize caproic acid-producing bacteria from the sewage sludge of the Ube Eastern Wastewater Treatment Plant in Japan, which processes 50,000 m3 of wastewater per day. The thermotolerance of isolated bacteria and their efficiency in caproic acid production were assessed to identify strains capable of producing caproic acid at concentrations exceeding 3 g/L at temperatures above 35 °C.
Materials and Methods
Microbial Source
The sewage sludge used in this study was collected from the Ube Eastern wastewater treatment plant in Japan, which treats municipal wastewater at a rate of 50,000 m3 per day. This sludge is a rich source of diverse microbial communities with potentially unique capabilities. The collected sludge samples were stored at 4 °C to preserve the microbial composition until further processing and experimentation.
Enrichment and Isolation of Caproic Acid-Producing Bacteria
Ten different substrate combinations were used to enrich caproic acid-producing bacteria from sewage sludge, as detailed in Table S1. These substrates were prepared to maintain a defined carbon source-to-electron donor ratio of 3:1 based on molarity. The composition of the basic anaerobic medium is modified based on Angelidaki et al. [18]. The medium was prepared per 1,000 mL with the following components: 1 g/L NH4Cl, 0.1 g/L NaCl, 0.1 g/L MgCl2·6H2O, 0.05 g/L CaCl2·2H2O, 0.3 K2HPO4·3H2O, 0.001 g/L C12H6NO4Na, 2.6 g/L NaHCO3, 1 mL trace-metal solution, 10 μL vitamin mixture solution, and 1 g yeast extract. 50 mL of the basal medium was prepared in 100-mL glass serum bottles (autoclaved at 121 °C for 15 min). The respective substrates (details in Table S1) were added aseptically to each bottle. The pH was then adjusted to 6.5 using a sterile solution of HCL or NaOH. For each substrate combination, triplicate enrichment cultures were prepared by inoculating 10% (w/v) of the sludge into the medium. The bottles were purged with a nitrogen-carbon dioxide mixture (80:20 v/v) to create an oxygen-free environment, sealed with rubber stoppers and aluminum crimps, and incubated at 35 °C for 7 days. Samples for high-performance liquid chromatography (HPLC) analysis were prepared by centrifugation at 10,000 rpm for 10 min using a Heraeus Multifuge X3R centrifuge (Thermo Fisher Scientific, USA) and filtration through a 0.45-μm pore-size nylon membrane filter (MilliporeSigma, USA) to remove suspended solids.
Enriched cultures with the highest caproic acid concentration were selected for isolation. The isolation process was conducted within an anaerobic chamber (Coy Laboratory Products, USA) filled with an inert gas mixture (90% N2, 5% CO2, and 5% H2) to maintain anaerobic conditions. Serial dilutions (10–1 to 10–9) of the enriched culture were prepared in an oxygen-free liquid medium containing ethanol and acetic acid (3:1 molar ratio). 0.5 mL of the enriched culture was transferred using a syringe into a serum bottle containing 4.5 mL of oxygen-free liquid medium containing ethanol and acetic acid. The culture was gently mixed for one minute to ensure a homogenous mixture. This process was repeated by serially transferring 0.5 mL aliquots of the diluted culture into new serum bottles containing fresh, oxygen-free liquid medium containing ethanol and acetic acid. Aliquots (0.1 mL) of diluted culture samples (10–6 to 10–9) were spread onto pre-reduced agar plates containing ethanol and acetic acid (3:1 molar ratio) and incubated at 35 °C for 7 days in an anaerobic jar with gas-generating packets (AnaeroGen, Oxoid). Single colonies were picked, subcultured three times on the same medium to ensure purity, and transferred to a liquid medium for further characterization.
Characterization and Identification of Strain M1NH
Strain M1NH, selected for its high caproic acid production, was characterized morphologically and biochemically. Colony morphology was observed using a stereomicroscope (Olympus SZX16, Japan). Gram staining was performed to determine the Gram reaction and cell morphology using a light microscope (Olympus BX51, Japan) at 1000 × magnification. Biochemical tests, including catalase, oxidase, and sugar fermentation, were conducted according to Bergey's Manual of Determinative Bacteriology [19]. The genomic DNA of strain M1NH was extracted using the NucleoSpin Microbial DNA kit (Macherey-Nagel, Germany) following the manufacturer's instructions. The 16S rRNA gene was amplified using universal primers 27F (5'-AGAGTTTGATCCTGGCTCAG-3') and 1492R (5'-GGTTACCTTGTTACGACTT-3'). PCR products were purified using the NucleoSpin Gel and PCR Clean-up kit (Macherey-Nagel, Germany) and sequenced at the Center for Gene Research (Yamaguchi University, Japan). The obtained 16S rRNA gene sequence was compared with the NCBI 16S rRNA gene database using the BLAST tool. A phylogenetic tree was constructed using the neighbor-joining method with 1,000 bootstrap replications in MEGA 7.0 software [20]. Reference sequences of related type strains were retrieved from the LPSN database [21].
Effect of Substrates and Environmental Conditions on M1NH Growth
The effects of temperature, pH, and substrate concentrations on the growth of strain M1NH were investigated using a factorial design. Fresh anaerobic medium (50 mL) was prepared in 100-mL glass serum bottles and autoclaved at 121 °C for 15 min for all experiments. The medium was inoculated with a 5% (v/v) inoculum of strain M1NH and flushed with a nitrogen-carbon dioxide mixture (80:20 v/v) to maintain anaerobic conditions. All experiments were conducted in triplicate. To evaluate the effect of temperature, bottles were incubated at 25 °C, 30 °C, 35 °C, 40 °C, 45 °C, 50 °C, and 55 °C, with the pH adjusted to 6.5 using sterile HCl or NaOH. For the pH effect, the medium was adjusted to pH 3, 4, 5, 6, 7, 8, 9, and 10 and incubated at the optimal temperature determined from the previous experiment. The impact of ethanol (1–7% v/v), acetic acid (1–7% w/v), and caproic acid (0.5–4% w/v) concentrations on M1NH growth was tested at the optimal pH and temperature. In all experiments, growth was monitored by measuring the optical density at 600 nm (OD600) every 6 h using a UV–Visible spectrophotometer (Genesys 20, Thermo Fisher Scientific Inc., USA). Growth data were fitted to the Gompertz model [22] to determine the maximum specific growth rate (μmax) and lag phase duration (λ).
Optimization Condition for Caproic Acid Production by Strain M1NH
The selected caproic acid-producing culture, strain M1NH, was subjected to adaptive one factor at a time. This method explored the impact of pH, temperature, and carbon source-to-electron donor ratio on caproic acid production. pH optimization involved testing levels of 5.5, 6.0, 6.5, 7.0, and 7.5 with an ethanol-to-acetic acid ratio of 3:1 (120 mM:40 mM) and a temperature of 40 °C. Temperature variations of 30 °C, 35 °C, 37 °C, and 40 °C were evaluated at the optimal pH with the same ethanol-to-acetic acid ratio. The ethanol-to-acetic acid ratios tested were 4:1 (160 mM:40 mM), 3:1 (120 mM:40 mM), 2:1 (80 mM:40 mM), and 1:1 (40 mM:40 mM) under the optimal pH and temperature conditions. Caproic acid concentrations were measured using HPLC. The optimized conditions were determined based on the highest caproic acid production.
Analytical Methods
Caproic acid and other metabolites were quantified by HPLC (Agilent Technologies 1200 series, USA) equipped with a refractive index detector and Aminex HPX-87H ion exclusion column (300 mm × 7.8 mm; Bio-Rad, USA). The column and detector temperatures were maintained at 65 °C and 50 °C, respectively. The mobile phase was 0.005 M H2SO4 at a 0.6 mL/min flow rate. Samples were filtered through 0.2 μm nylon membrane filters before analysis. Concentrations were calculated based on calibration curves prepared with authentic standards (Sigma-Aldrich, USA). All enrichment experiments were conducted in triplicate. The data were subjected to a one-way analysis of variance (ANOVA) followed by Tukey's honestly significant difference (HSD) post hoc test to determine significant differences among the substrate combinations for caproic acid production. The level of significance was set at P < 0.05. Statistical analyses were performed using R software (version 4.0.3).
Results
Enrichment and Isolation of Caproic Acid-Producing Bacteria
The enrichment of caproic acid-producing bacteria was performed using various substrate conditions, including lactic acid, glucose, xylose, ethanol, and acetic acid, either alone or in combination. The combination of ethanol and acetic acid yielded significantly higher caproic acid concentration (2.7 ± 0.2 g/L) and conversion efficiency (47 ± 3%) compared to the other substrate combinations (P < 0.05) (Fig. 1). Substrate combinations such as ethanol-glucose (1.2 ± 0.1 g/L), ethanol-xylose (0.8 ± 0.1 g/L), lactic acid–glucose (0.6 ± 0.1 g/L), and lactic acid-xylose (0.3 ± 0.1 g/L) also showed potential for enriching caproic acid-producing bacteria, with caproic acid concentrations significantly lower than the ethanol–acetic acid combination (P < 0.05). The ethanol–acetic acid combination also resulted in moderate valeric acid (2 ± 0.5%) and butyric acid (7 ± 1%) conversion efficiencies, as well as a high acetic acid conversion efficiency (12 ± 2%). The influence of different carbon sources was evident in the case of lactic acid. Combined with glucose, it resulted in a caproic acid conversion efficiency of 6 ± 1%, a butyric acid conversion efficiency of 28 ± 3%, and an acetic acid conversion efficiency of 7.37 ± 0.8%. In contrast, the lactic acid–xylose combination yielded a caproic acid conversion efficiency of 3 ± 0.5%, along with other volatile fatty acids (VFAs) such as butyric acid (17 ± 2%) and propionic acid (6.67 ± 0.9%). Standalone substrates, ethanol, and acetic acid did not yield caproic acid, emphasizing the importance of combining suitable carbon sources for productive fermentation. Among these, ethanol demonstrated a propensity for acetic acid conversion efficiency (7.14 ± 0.8%), while acetic acid favored butyric acid (2 ± 0.5%). Glucose favored acetic acid conversion efficiency (12.5 ± 1.5%), while xylose favored butyric acid (52 ± 4%) and displayed a higher proportion of acetic acid (7.59 ± 0.9%).
Enrichment cultures using ethanol and acetic acid yielded three distinct bacterial strains: M1NH, M2NH, and M3NH (Fig. 2). One-way ANOVA and Tukey’s HSD post hoc test revealed significant differences among the strains for caproic acid production (p < 0.001) and butyric acid production (P < 0.001). Strain M1NH emerged as the most promising candidate, achieving a caproic acid yield of 2.5 ± 0.2 g/L with a conversion efficiency of 44 ± 3%, significantly higher than those of M2NH (0.6 ± 0.1 g/L) and M3NH (0.2 ± 0.1 g/L) (P < 0.05). Conversely, strains M2NH (0.9 ± 0.1 g/L) and M3NH (0.8 ± 0.1 g/L) displayed significantly higher butyric acid levels compared to the minimal production by M1NH (0.1 ± 0.05 g/L) (P < 0.05). Therefore, strain M1NH was selected for further investigation.
Characteristics and Phylogenetic Classification of Strain M1NH
Strain M1NH, a gram-positive, rod-shaped bacterium, exhibited anaerobic growth and spore-forming capability. The morphology of the strain reveals a large, smooth, and cream-colored appearance. Biochemical tests showed that strain M1NH was favorable for urease, catalase, and oxidase activities but negative for hydrogen sulfide production. These findings suggest the strain’s adaptability to environments containing these compounds, which is essential for its potential application in caproic acid production. A phylogenetic analysis was conducted to determine the evolutionary relationships between strain M1NH and related type strains within the genus Clostridium. The 16S rRNA gene sequence of strain M1NH (1,452 bp) was compared with those of reference strains retrieved from the LPSN database. The phylogenetic tree (Fig. 3) showed that strain M1NH (accession number PP374094) formed a distinct cluster with Clostridium luticellarii DSM 29923 T, supported by a high bootstrap value of 98%. The 16S rRNA gene sequence similarity between strain M1NH and C. luticellarii DSM 29923 T was 98.2%, indicating a close phylogenetic relationship. The sequence similarities between strain M1NH and other related type strains ranged from 92.5 to 95.1%. Escherichia coli ATCC 11775 T was used as an outgroup and shared only 84.3% sequence similarity with strain M1NH. The pairwise distance matrix analysis and the corresponding statistical test (one-way ANOVA with Tukey's HSD post hoc test) further strengthen the phylogenetic evidence for the taxonomic position of strain M1NH within the genus Clostridium. Based on the morphological, biochemical, and phylogenetic analyses, strain M1NH represents a novel member of the genus Clostridium, closely related to C. luticellarii. The distinct clustering of strain M1NH with C. luticellarii DSM 29923 T in the neighbor-joining tree, supported by high bootstrap values and low evolutionary distances, provides strong evidence for its taxonomic position within the genus Clostridium.
Phylogenetic tree based on 16S rRNA gene sequences showing the relationship between Clostridium strain M1NH and related type strains within the genus Clostridium. The tree was constructed using the neighbor-joining method with 1000 bootstrap replications. Bootstrap values greater than 50% are shown at the branch nodes. Escherichia coli ATCC 11775 T served as the outgroup. GenBank accession numbers for the 16S rRNA sequences are provided in parentheses. The scale bar represents 5% sequence divergence
The Growth and Inhibition of Clostridium M1NH
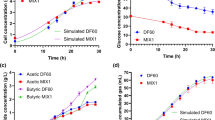
The growth of Clostridium M1NH significantly increased between 12 and 36 h, indicating exponential growth. Following rapid growth, the cell culture entered the stationary phase at 36 h, as evidenced by a stable growth rate. The growth of Clostridium M1NH was significantly influenced by pH. The optimal pH range for growth was 6.0–6.5, with the highest OD600 value (1.85 ± 0.10) observed at pH 6.5 after 36 h of incubation (Fig. 4a). The growth rate at pH 6.5 was 0.14 ± 0.01 h−1, and the doubling time was 5.3 ± 0.4 h, both significantly different from those at other pH levels (P < 0.05). Growth was severely inhibited at acidic pH levels (3–5) and alkaline pH levels (8–10), with minimal or no growth observed (OD600 < 0.5). Temperature significantly affected the growth of Clostridium M1NH. The optimal temperature range for growth was 35–40 °C, with a growth rate of 0.11 ± 0.01 h−1 and a doubling time of 6.5 ± 0.5 h (Fig. 4b). The strain could tolerate and grow within a temperature range of 30–41 °C. Still, the growth was significantly lower than the optimal range (P < 0.05). At temperatures below 30 °C and above 41 °C, the OD600 decreased significantly, and growth ceased entirely at 55 °C.
Ethanol concentration had a significant impact on the growth of Clostridium M1NH. High ethanol concentrations (> 1% v/v) inhibited the growth rate and increased the doubling time (Fig. 4c). At 0% ethanol, the OD600 was the highest (1.92 ± 0.12), indicating robust growth. As the ethanol concentration increased from 1 to 5%, the OD600 significantly decreased (P < 0.05), indicating reduced cell growth. At 6% ethanol, the OD600 dropped to 0, suggesting complete growth inhibition. The growth of Clostridium M1NH was significantly affected by acetic acid concentration. High acetic acid concentrations (> 1% w/v) inhibited the growth rate and increased the doubling time (Fig. 4d). At 0% acetic acid, the culture had the highest exponential growth rate (0.10 ± 0.01 h−1). As the acetic acid concentration increased from 1 to 5%, the growth rate significantly decreased from 0.08 ± 0.01 h−1 to 0.03 ± 0.01 h−1 (P < 0.05), suggesting nearly complete growth inhibition at 5% acetic acid. The doubling time increased from 7.38 ± 0.63 h at 0% acetic acid to 27.13 ± 2.31 h at 5% acetic acid. Caproic acid concentration significantly influenced the growth of Clostridium M1NH. At 0% caproic acid, the OD600 had the highest growth rate (0.10 ± 0.01 h−1 (Fig. 4e). As the caproic acid concentration increased from 0.5 to 2%, the OD600 progressively decreased, indicating reduced cell growth. The growth rate remained stable between 1.5 and 2% caproic acid concentrations. At concentrations higher than 2.5%, the OD600 dropped to 0, indicating complete growth inhibition.
Optimization for Caproic Acid Production from Clostridium M1NH
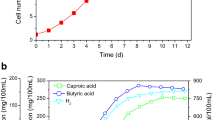
The initial pH significantly influenced caproic acid production by Clostridium M1NH. Caproic acid production was initiated at all pH levels on day 2 of fermentation, with pH 6.5 showing slightly higher initial production (0.82 ± 0.09 g/L) than acidic or alkaline conditions (Fig. 5a). On day 4, production reached a significant level at pH 6.0–7.5, with no significant differences among these pH levels (P > 0.05). However, pH 5.0 showed significantly lower production (1.25 ± 0.12 g/L) compared to the other pH levels (P < 0.05). On day 6, caproic acid concentration peaked at pH 6.0 (2.95 ± 0.18 g/L) and 7.0 (2.70 ± 0.15 g/L), which were significantly higher than those at other pH levels (P < 0.05). From day 8 to day 12, caproic acid production declined at pH 7.0 and 7.5, while it remained relatively stable at pH 6.0. Temperature had a significant effect on caproic acid production by Clostridium M1NH. At the early stage of fermentation (day 2), caproic acid production was generally low across all temperatures, with 35 °C showing slightly higher initial production (0.70 ± 0.08 g/L) (Fig. 5b). On day 4, caproic acid production began to increase noticeably, with 40 °C showing the highest concentration (2.70 ± 0.16 g/L), followed by 37 °C (2.35 ± 0.14 g/L). However, by day 10, the caproic acid concentration peaked at 30–35 °C, reaching the highest production of 3.10 ± 0.19 g/L and 3.20 ± 0.21 g/L, respectively. These values were significantly higher than those at 37 °C and 40 °C (P < 0.05). Caproic acid concentrations decreased at 37 °C and 40 °C after day 8. The ethanol-to-acetic acid ratio significantly affected caproic acid production by Clostridium M1NH. On day 2, caproic acid production was low across all ratios, with higher ethanol content ratios (4:1 and 3:1) showing slightly higher initial production (Fig. 5c). On day 6, production reached significant levels at 3:1 (2.85 ± 0.17 g/L) and 4:1 (3.10 ± 0.19 g/L) ratios, which were significantly higher than those at 2:1 and 1:1 ratio (P < 0.05). On day 10, caproic acid concentration peaked at the 4:1 ratio (3.49 ± 0.23 g/L), followed by the 3:1 ratio (3.21 ± 0.20 g/L). The 2:1 ratio showed a steady increase but at a slower rate, while the 1:1 ratio resulted in minimal further production after day 6. The 4:1 and 3:1 ratios maintained significantly higher caproic acid concentrations than the 2:1 and 1:1 ratios throughout the fermentation period (P < 0.05).
Comparison with Other Strains
The comparison of caproic acid production by Clostridium M1NH with previously reported strains is shown in Table 1. The table includes information on the substrate used, caproic acid production, fermentation time, temperature, and the corresponding references. Clostridium M1NH efficiently produces caproic acid utilizing a combination of ethanol and acetic acid as substrates. In contrast, other strains utilize various substrates, including lactate (Ruminococcaceae CPB6), fructose (Megasphaera sp. MH), glucose (Megasphaera elsdenii ATCC 25940), ethanol (C. kluyveri 3231B), and galactitol (Clostridium sp. BS-1). Clostridium M1NH achieves a caproic acid production of 3.5 g/L, comparable to the yield of C. kluyveri (2.5 g/L) when using ethanol and acetic acid as substrates. However, some strains, such as Ruminococcaceae CPB6 (16.6 g/L), Megasphaera sp. MH (9.7 g/L), M. elsdenii ATCC 25940 (11.4 g/L), and C. kluyveri 3231B (12.8 g/L), exhibit higher caproic acid production using different substrates. Clostridium M1NH requires a relatively long fermentation time of 12 days to achieve its maximum caproic acid production. In comparison, some strains, like Megasphaera sp. MH and C. kluyveri 3231B, achieve high caproic acid yields within a shorter fermentation period of 1 and 3 days, respectively. Clostridium M1NH demonstrates a unique thermotolerance, with an optimal temperature range of 35–40 °C for caproic acid production. Most other strains operate at 37 °C, while Ruminococcaceae CPB6 has a broader temperature range of 30–40 °C. Some studies such as those involving M. elsdenii ATCC 25940 and Clostridium sp. BS-1 employ fed-batch fermentation and product removal strategies to reduce product inhibition and enhance caproic acid production. Clostridium M1NH may not exhibit the highest caproic acid production among the compared strains, but its thermotolerance and ability to efficiently utilize ethanol and acetic acid as substrates highlight its potential for application in bioprocesses where elevated temperatures and specific substrate availability are advantageous.
Discussion
The results of this study provide valuable insights into the conditions that favor caproic acid production by enriched caproic acid-producing bacteria and the specific strain Clostridium M1NH. Different substrate combinations significantly influenced the enrichment and production of caproic acid. The combination of ethanol and acetic acid yielded the highest caproic acid concentration and conversion efficiency, highlighting the importance of selecting suitable carbon sources for optimizing production. This finding aligns with previous studies, such as those by Liu et al. [23], Bao et al. [24], and Steinbusch et al. [25], which reported varying degrees of success with similar substrates. The high conversion efficiency observed in this study (47%) suggests that ethanol–acetic acid is particularly effective for caproic acid production. Other combinations, such as lactic acid–glucose and lactic acid–xylose, also showed promise, indicating potential alternative substrates for caproic acid enrichment. The isolation of Clostridium M1NH from enrichment cultures and its subsequent characterization revealed essential aspects of its physiology and possible application. The strain's ability to produce significant amounts of caproic acid (3.5 g/L) and its high conversion efficiency (44%) position it as a strong candidate for industrial caproic acid production. The phylogenetic analysis indicated that Clostridium M1NH is closely related to Clostridium luticellarii DSM 29923 T, suggesting it may share metabolic pathways conducive to caproic acid synthesis. The strain's growth characteristics, including its optimal pH (6.5) and temperature (35–40 °C), provide critical information for designing fermentation processes. Understanding the factors that inhibit the growth of Clostridium M1NH is crucial for optimizing caproic acid production. High ethanol and acetic acid concentrations inhibited growth, with significant reductions in OD600 at concentrations above 1%. This finding emphasizes the need to maintain substrate concentrations within optimal ranges to avoid inhibitory effects. A study on Escherichia coli showed that ethanol at 30 g/L decreased the growth yield and experimentally enhanced the specific death rate [26]. Ethanol inhibition can be caused by end-product inhibition or chaotropic-induced stress, resulting in increased membrane fluidization and disruption of macromolecules [31]. A similar observation discussed the fermentation kinetics of Clostridium tyrobutyricum cultures after being classically adapted for growth at 26.3 g/L acetic acid. It showed a lag time of 25 h and log phase growth of 0.07 h−1 [32]. The mechanism of action of high concentrations of acetic acid as an inhibitor of bacterial growth is due to the acidification of the cytoplasm, which can lead to the disruption of cellular processes. Acetic acid can also inhibit the activity of enzymes involved in energy metabolism, such as ATP synthase and succinate dehydrogenase [33].
The inhibition observed with increasing caproic acid concentrations (> 2.5%) suggests that caproic acid toxicity could limit production yields. Strategies to mitigate these effects, such as in situ product removal, could enhance the strain's tolerance and productivity. A dose of 4 mM caproic acid is sufficient to decrease the growth rate of Saccharomyces cerevisiae by 85% [34], while a dose of 40 mM caproic acid completely inhibits the growth of Escherichia coli [35]. 1% caproic acid completely inhibits the growth of Staphylococcus aureus and Escherichia coli [36]. Furthermore, the inhibitory concentration of caproate is much lower than the inhibitory concentration of butyrate at neutral pH, as demonstrated with C. kluyveri [37]. The exact mechanism of action is not fully understood. However, it is suggested that the fatty acid derivatives might interact with the microbial cell membrane, leading to increased permeability and leakage of cell contents [35, 38]. These effects can lead to cell death or reduced growth. A caproic acid-producing strain increased membrane leakage as the product titer increased, but there was no change in membrane fluidity [35]. However, more research is needed to understand the underlying molecular mechanisms fully.
The optimization studies provided detailed insights into the conditions that maximize caproic acid production. The optimal initial pH (6.5) and temperature (30–35 °C) for caproic acid production align with the growth characteristics of Clostridium M1NH. The highest caproic acid yield (3.5 g/L) was achieved with 7.73 g/L (160 mM) ethanol and 2.36 g/L (40 mM) acetic acid, indicating that precise control of substrate concentrations is key to maximizing production. Yan et al. [39] found that the optimum initial pH for caproic acid production by C. kluyveri was determined to be 6.41. San-Valero et al. [40] found that C. kluyveri produced maximum caproic acid when the pH was maintained at 6.8. In addition, pH affected the dissolution state of CO2 in the fermentation system, and CO2 was a very important inorganic carbon source in the synthesis and metabolism of C. kluyveri. The influence of pH on metabolic pathways, enzyme activities, and microbial growth dynamics likely contributes to the observed differences in caproic acid production [41]. A strain of Ruminococcaceae bacterium CPB6, affiliated with Clostridium cluster IV, prefers a temperature range of 30 to 40 °C for caproic acid production [14].
The comparison of caproic acid production by Clostridium M1NH with other known strains highlights its unique characteristics and potential advantages. Although some strains, such as Ruminococcaceae bacterium CPB6 and C. kluyveri 3231B, achieved higher caproic acid concentrations, they required specific substrates (lactate and ethanol, respectively) and shorter fermentation times. In contrast, Clostridium M1NH demonstrated the ability to efficiently utilize a combination of ethanol and acetic acid, which are more readily available and cost-effective substrates. Additionally, the thermotolerance of Clostridium M1NH, with an optimal growth temperature range of 35–40 °C, is particularly advantageous for industrial-scale fermentations, where maintaining strict temperature control can be challenging and expensive. The strain's tolerance to higher concentrations of caproic acid (up to 2% w/v) compared to other strains is another desirable trait, as it could lead to more efficient downstream processing and reduced production costs.
The successful isolation and characterization of a novel caproic acid-producing strain from sewage sludge was Clostridium M1NH. The strain's ability to efficiently produce caproic acid using ethanol and acetic acid as substrates and its thermotolerance and resistance to high product concentrations make it a promising candidate for industrial applications. The optimization of fermentation conditions, including pH, temperature, and substrate ratios, provided valuable insights into maximizing caproic acid production. Further research should focus on scaling up the fermentation process, investigating the strain's performance in continuous or fed-batch systems, and exploring strategies for in situ product removal to minimize the inhibitory effects of caproic acid accumulation. Additionally, metabolic engineering approaches could enhance the strain's caproic acid production capacity and substrate utilization efficiency. The findings of this study contribute to the growing body of knowledge on microbial caproic acid production and offer new opportunities for the sustainable production of this valuable chemical from renewable resources.
Conclusion
This study uses a novel thermotolerant caproic acid-producing bacterial strain, Clostridium sp. M1NH was successfully isolated from sewage sludge. Substrate optimization revealed that ethanol and acetic acid, at a molar ratio of 4:1, were the most effective carbon sources for caproic acid production, resulting in a maximum yield of 3.49 ± 0.23 g/L after 10 days of fermentation. Clostridium sp. M1NH exhibited remarkable tolerance to high concentrations of ethanol (up to 5% v/v), acetate (up to 5% w/v), and caproic acid (up to 2% w/v). The strain also demonstrated a wide pH tolerance range (pH 5.5–7.5) and an elevated temperature optimum between 30 and − 35 °C, with a maximum specific growth rate of 0.14 ± 0.01 h−1 at pH 6.5 and 35 °C. Phylogenetic analysis based on 16S rRNA gene sequences revealed that Clostridium sp. M1NH shares a 98.2% similarity with the type strain Clostridium luticellarii DSM 29923 T, indicating a close evolutionary relationship. Optimization of fermentation conditions further enhanced caproic acid production, with the highest yield achieved at an initial pH of 6.0–6.5, a temperature of 30–35 °C, and an ethanol-to-acetate ratio of 4:1. The maximum caproic acid yield obtained by Clostridium sp. M1NH (3.49 ± 0.23 g/L) was 1.6-fold higher than that reported for C. kluyveri (2.5 g/L) under similar fermentation conditions, highlighting the superior performance of the newly isolated strain. Furthermore, the thermotolerance and wide substrate utilization range of Clostridium sp. M1NH make it an attractive candidate for industrial-scale caproic acid production from diverse low-cost feedstocks.
References
Zentek J, Buchheit-Renko S, Ferrara F, Vahjen W, Van Kessel A, Pieper R (2011) Nutritional and physiological role of medium-chain triglycerides and medium-chain fatty acids in piglets. Anim Health Res Rev 12:83–93. https://doi.org/10.1017/S1466252311000089
Woolford MK (1975) Microbiological screening of the straight chain fatty acids (C1–C12) as potential silage additives. J Sci Food Agric 26:219–228. https://doi.org/10.1002/jsfa.2740260213
Scalschi L, Vicedo B, Camañes G, Fernandez-Crespo E, Lapeña L, González-Bosch C, García-Agustín P (2013) Hexanoic acid is a resistance inducer that protects tomato plants against Pseudomonas syringae by priming the jasmonic acid and salicylic acid pathways. Mol Plant Pathol 14:342–355. https://doi.org/10.1111/mpp.12010
Levy P, Sanderson J, Kispert R, Wise D (1981) Biorefining of biomass to liquid fuels and organic chemicals. Enzyme Microb Technol 3:207–215. https://doi.org/10.1016/0141-0229(81)90087-9
Angenent LT, Richter H, Buckel W, Spirito CM, Steinbusch KJ, Plugge CM, Strik DP, Grootscholten TI, Buisman CJ, Hamelers HV (2016) Chain elongation with reactor microbiomes: open-culture biotechnology to produce biochemicals. Environ Sci Technol 50:2796–2810. https://doi.org/10.1021/acs.est.5b04847
Agler MT, Wrenn BA, Zinder SH, Angenent LT (2011) Waste to bioproduct conversion with undefined mixed cultures: the carboxylate platform. Trends Biotechnol 29:70–78. https://doi.org/10.1016/j.tibtech.2010.11.006
O-Thong S, Zhu X, Angelidaki I, Zhang S, Luo G (2020) Chapter two—medium chain fatty acids production by microbial chain elongation: recent advances. In: Li Y, Khanal SK (eds) Advances in bioenergy. Elsevier, Amsterdam, pp 63–99
Hu X-l, Du H, Xu Y (2015) Identification and quantification of the caproic acid-producing bacterium Clostridium kluyveri in the fermentation of pit mud used for Chinese strong-aroma type liquor production. Int J Food Microbiol 214:116–122. https://doi.org/10.1016/j.ijfoodmicro.2015.07.032
Reddy MV, Mohan SV, Chang Y-C (2018) Medium-chain fatty acids (MCFA) production through anaerobic fermentation using Clostridium kluyveri: effect of ethanol and acetate. Appl Biochem Biotechnol 185:594–605. https://doi.org/10.1007/s12010-017-2674-2
Choi K, Jeon BS, Kim B-C, Oh M-K, Um Y, Sang B-I (2013) In situ biphasic extractive fermentation for hexanoic acid production from sucrose by Megasphaera elsdenii NCIMB 702410. Appl Biochem Biotechnol 171:1094–1107. https://doi.org/10.1007/s12010-013-0310-3
Lee N-R, Lee CH, Lee D-Y, Park J-B (2020) Genome-scale metabolic network reconstruction and in silico analysis of hexanoic acid producing Megasphaera elsdenii. Microorganisms 8:539. https://doi.org/10.3390/microorganisms8040539
Kurniawan E, Gagnon Y, Cheirsilp B, O-Thong S (2023) Effect of inoculum source, electron donors, and substrates on medium-chain fatty acid production from palm oil mill effluent. Ind Crops Prod 206:117654. https://doi.org/10.1016/j.indcrop.2023.117654
Esquivel-Elizondo S, Bağcı C, Temovska M, Jeon BS, Bessarab I, Williams RB, Huson DH, Angenent LT (2021) The isolate Caproiciproducens sp. 7D4C2 produces n-caproate at mildly acidic conditions from hexoses: genome and rBOX comparison with related strains and chain-elongating bacteria. Front Microbiol 11:594524. https://doi.org/10.18174/icec2020.18020
Zhu X, Zhou Y, Wang Y, Wu T, Li X, Li D, Tao Y (2017) Production of high-concentration n-caproic acid from lactate through fermentation using a newly isolated Ruminococcaceae bacterium CPB6. Biotechnol Biofuels 10:102. https://doi.org/10.1186/s13068-017-0788-y
Prado CD, Mandrujano GPL, Souza JP, Sgobbi FB, Novaes HR, da Silva JPMO, Alves MHR, Eliodório KP, Cunha GCG, Giudici R, Procópio DP, Basso TO, Malavazi I, Cunha AF (2020) Physiological characterization of a new thermotolerant yeast strain isolated during Brazilian ethanol production, and its application in high-temperature fermentation. Biotechnol Biofuels Bioprod 13:178. https://doi.org/10.1186/s13068-020-01817-6
Kanekar PP, Kanekar SP (2022) Thermophilic, thermotolerant microorganisms. In: Kanekar PP, Kanekar SP (eds) Diversity and biotechnology of extremophilic microorganisms from India. Springer, Singapore, pp 117–153
Gomes E, de Souza AR, Orjuela GL, Da Silva R, de Oliveira TB, Rodrigues A (2016) Applications and benefits of thermophilic microorganisms and their enzymes for industrial biotechnology. In: Schmoll M, Dattenböck C (eds) Gene expression systems in fungi: advancements and applications. Springer, Cham, pp 459–492
Angelidaki I, Alves M, Bolzonella D, Borzacconi L, Campos J, Guwy A, Kalyuzhnyi S, Jenicek P, Van Lier J (2009) Defining the biomethane potential (BMP) of solid organic wastes and energy crops: a proposed protocol for batch assays. Water Sci Technol 59:927–934. https://doi.org/10.2166/wst.2009.040
Vos P, Garrity G, Jones D, Krieg NR, Ludwig W, Rainey FA, Schleifer K-H, Whitman WB (2011) Bergey’s manual of systematic bacteriology; volume 3: The Firmicutes. Springer, New York
Kumar S, Stecher G, Li M, Knyaz C, Tamura K (2018) MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol 35(6):1547–1549. https://doi.org/10.1093/molbev/msy096
Parte AC, Sardà Carbasse J, Meier-Kolthoff JP, Reimer LC, Göker M (2020) List of prokaryotic names with standing in nomenclature (LPSN) moves to the DSMZ. Int J Syst Evol Microbiol 70(11):5607–5612. https://doi.org/10.1099/ijsem.0.004332
Zwietering MH, Jongenburger I, Rombouts FM, van’t Riet K (1990) Modeling of the bacterial growth curve. Appl Environ Microbiol 56(6):1875–1881. https://doi.org/10.1128/aem.56.6.1875-1881.1990
Liu Y, Lü F, Shao L, He P (2016) Alcohol-to-acid ratio and substrate concentration affect product structure in chain elongation reactions initiated by unacclimatized inoculum. Bioresour Technol 218:1140–1150. https://doi.org/10.1016/j.biortech.2016.07.067
Bao S, Wang Q, Zhang P, Zhang Q, Wu Y, Li F, Tao X, Wang S, Nabi M, Zhou Y (2019) Effect of acid/ethanol ratio on medium chain carboxylate production with different VFAs as the electron acceptor: insight into carbon balance and microbial community. Energies 12:3720. https://doi.org/10.3390/en12193720
Steinbusch KJ, Hamelers HV, Plugge CM, Buisman CJ (2011) Biological formation of caproate and caprylate from acetate: fuel and chemical production from low grade biomass. Energy Environ Sci 4:216–224. https://doi.org/10.1039/c0ee00282h
Sawada T, Nakamura Y (1987) Growth inhibitory and lethal effects of ethanol on Escherichia coli. Biotechnol Bioeng 29:742–746. https://doi.org/10.1002/bit.260290611
Jeon BS, Choi O, Um Y, Sang BI (2016) Production of medium-chain carboxylic acids by Megasphaera sp. MH with supplemental electron acceptors. Biotechnol Biofuels 9:129. https://doi.org/10.1186/s13068-016-0549-3
Roddick FA, Britz ML (1997) Production of hexanoic acid by free and immobilised cells of Megasphaera elsdenii: influence of in-situ product removal using ion exchange resin. J Chem Technol Biotechnol 69:383–391
Weimer PJ, Stevenson DM (2012) Isolation, characterization, and quantification of Clostridium kluyveri from the bovine rumen. Appl Microbiol Biotechnol 94:461–466. https://doi.org/10.1007/s00253-011-3751-z
Jeon BS, Moon C, Kim B-C, Kim H, Um Y, Sang B-I (2013) In situ extractive fermentation for the production of hexanoic acid from galactitol by Clostridium sp. BS-1. Enzyme Microb Tecshnol 53:143–151. https://doi.org/10.1016/j.enzmictec.2013.02.008
Kuil T, Yayo J, Pechan J, Küchler J, van Maris AJA (2022) Ethanol tolerance of Clostridium thermocellum: the role of chaotropicity, temperature and pathway thermodynamics on growth and fermentative capacity. Microb Cell Fact 21:273. https://doi.org/10.1186/s12934-022-01999-8
Jaros AM, Rova U, Berglund KA (2013) Acetate adaptation of Clostridia tyrobutyricum for improved fermentation production of butyrate. Springerplus 2:47. https://doi.org/10.1186/2193-1801-2-47
Pinhal S, Ropers D, Geiselmann J, De Jong H (2019) Acetate metabolism and the inhibition of bacterial growth by acetate. J Bacteriol 201:e00147-19201. https://doi.org/10.1128/jb.00147-00119
Liu P, Chernyshov A, Najdi T, Fu Y, Dickerson J, Sandmeyer S, Jarboe L (2013) Membrane stress caused by octanoic acid in Saccharomyces cerevisiae. Appl Microbiol Biotechnol 97:3239–3251. https://doi.org/10.1007/s00253-013-4773-5
Royce LA, Liu P, Stebbins MJ, Hanson BC, Jarboe LR (2013) The damaging effects of short chain fatty acids on Escherichia coli membranes. Appl Microbiol Biotechnol 97(18):8317–8327. https://doi.org/10.1007/s00253-013-5113-5
Hismiogullari S, Elyurek E, Hismiogullari A, Sahin F, Basalan M, Yenice S (2008) Effects of caproic and caprylic acids on microbial growth and cytotoxicity. J Anim Vet Adv 7:731–735
Candry P, Van Daele T, Denis K, Amerlinck Y, Andersen SJ, Ganigué R, Arends JBA, Nopens I, Rabaey K (2018) A novel high-throughput method for kinetic characterisation of anaerobic bioproduction strains, applied to Clostridium kluyveri. Sci Rep 8:9724. https://doi.org/10.1038/s41598-018-27594-9
Nobmann P, Bourke P, Dunne J, Henehan G (2010) In vitro antimicrobial activity and mechanism of action of novel carbohydrate fatty acid derivatives against Staphylococcus aureus and MRSA. J Appl Microbiol 108:2152–2161. https://doi.org/10.1111/j.1365-2672.2009.04622.x
Yan S-B, Wang S-C, Qiu Z-F, Wei G-G, Zhang K-G (2015) Optimization of caproic acid production from Clostridium kluyveri H588 and its application in Chinese luzhou-flavor liquor brewing. Adv J Food Sci Technol 7:614–626. https://doi.org/10.19026/ajfst.7.1368
San-Valero P, Fernández-Naveira Á, Veiga M, Kennes C (2019) Influence of electron acceptors on hexanoic acid production by Clostridium kluyveri. J Environ Manage 242:515–521. https://doi.org/10.1016/j.jenvman.2019.04.093
Candry P, Radić L, Favere J, Carvajal-Arroyo J-M, Rabaey K, Ganigué R (2020) Mildly acidic pH selects for chain elongation to caproic acid over alternative pathways during lactic acid fermentation. Water Res 186:116396. https://doi.org/10.1016/j.watres.2020.116396
Acknowledgments
The authors would like to thank the Research Center for Thermotolerant Microbial Resources and Faculty of Engineering, Yamaguchi University, Japan, and the National Research Council of Thailand (NRCT) through the Talented Mid-Career Research Grant (Grant No. N41A640088) for the financial support.
Funding
This study was supported by grant No. N41A640088 from the National Research Council of Thailand (NRCT).
Author information
Authors and Affiliations
Contributions
Edy Kurniawan contributed toward visualization, conceptualization, formal analysis, investigation, methodology, software, writing—original draft, and writing—review and editing. Chonticha Leamdum contributed toward data curation and data analysis. Sompong O-Thong contributed toward conceptualization, data curation, funding acquisition, project administration, resources, supervision, validation, and writing—review and editing. Tsuyoshi Imai contributed toward funding acquisition, review, and editing.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Kurniawan, E., Leamdum, C., Imai, T. et al. Exploiting the Thermotolerance of Clostridium Strain M1NH for Efficient Caproic Acid Fermentation from Ethanol and Acetic Acid. Curr Microbiol 81, 244 (2024). https://doi.org/10.1007/s00284-024-03780-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00284-024-03780-z