Abstract
In the present study, a comprehensive proteomic analysis of Brucella melitensis (B. melitensis) strain ATCC23457 was carried out to investigate proteome alterations in response to in vitro-induced nutrient stress. Our analysis resulted in the identification of 2440 proteins, including 365 hypothetical proteins and 850 potentially secretory proteins representing ~77.8% of the B. melitensis proteome. Utilizing a proteogenomics approach, we provide translational evidence for eight novel putative protein-coding genes and confirmed the coding potential of 31 putatively annotated pseudogenes, thus refining the existing genome annotation. Further, using a label-free quantitative proteomic approach, new insights into the cellular processes governed by nutrient stress, including enrichment of amino acid metabolism (E), transcription (K), energy production and conversion (C), and biogenesis (J) processes were obtained. Pathway analysis revealed the enrichment of survival and homeostasis maintenance pathways, including type IV secretion system, nitrogen metabolism, and urease pathways in response to nutrient limitation. To conclude, our analysis demonstrates the utility of in-depth proteomic analysis in enabling improved annotation of the B. melitensis genome. Further, our results indicate that B. melitensis undergoes metabolic adaptations during nutrient stress similar to other Brucella. sp, and adapts itself for long-term persistence and survival.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Brucellosis caused by Brucella spp is a zoonotic disease of significant public health importance, with over half a million cases reported annually [1]. Despite being recognized as a global health priority, brucellosis has remained a neglected zoonotic disease in the Indian sub-continent due to a lack of awareness and prioritization over other infectious diseases [2]. To date, about 39 strains of Brucella have been sequenced, which has dramatically accelerated studies on comparative genome analysis for their genetic conservation and variability. Additionally, such studies have provided key insights into Brucella virulence and its immune evasion mechanisms within the host. However, despite available genomic data on metabolic and protein transport mechanisms in the Brucella genome, information concerning their role in pathogenesis and their adaptability to the host environment remains poorly defined.
Recent advances in mass spectrometry-based proteomics have aided in the refinement and integration of protein-level information into the genome annotation process [3,4,5]. Although several studies have explored the host response to Brucella infection [6, 7]; very few studies have characterized the proteome repertoire of the organism enabling confirmation of several putative proteins and rectifying genome annotation errors in Brucella abortus (B. abortus) strain 2308 [8]. Despite best efforts to comprehensively characterize the proteome of Brucella melitensis (B. meletensis), limited evidence of the actual number of protein-coding genes has been obtained to date [9, 10]. To better understand the virulence mechanisms initiated by B. melitensis and ensure detailed functional studies, it is imperative to obtain a comprehensive proteomic expression profile as well as catalog changes in the proteome in response to various stimuli such as nutrient stress and antimicrobials. Towards this end, we carried out an in-depth proteomic analysis of B. melitensis strain ATCC23457. Additionally, we also investigated the global proteome changes in response to nutrient limitation.
Material and Methods
Culturing of B. melitensis
B. melitensis strain ATCC23457, previously isolated from human blood, was cultured in both protein-replete and protein-free media. For the nutrient-rich condition, culturing was performed in Brucella broth (BD Biosciences, USA), followed by incubation for two weeks at 37 °C in 5% CO2. For nutritional deprived conditions, the cells were cultured using BacT/Alert 3D Advance Automated Culture System (Biomeriux, France), which required Fan aerobic (PF) plus liquid culture bottles for bacterial growth without any growth supplements. For confirmation, aliquots of both cultures were subjected to Gram staining and Brucella Omp 31-based PCR (Supplementary Fig. 1). Both cultures were grown until O.D600 of around 0.8–1.0 was reached and harvested at 10,000×g for 10 min. The pellets were washed three times in sterile ice-cold 1×sterile phosphate-buffered saline (PBS) by centrifugation at 10,000 g for 10 min. The washed pellets were stored at – 80 °C until further downstream processing.
Protein Extraction and Sample Preparation for Mass Spectrometry
Proteins were extracted from culture pellets using SDS lysis buffer (2% SDS in 50 mM TEABC). Briefly, 500 µl lysis buffers were added to the cell pellets and sonicated using a probe sonicator (Fisher Scientific, USA) on ice for 20 min. The lysates were heated at 95 ºC for 10 min, allowed to cool to room temperature, and centrifuged at 12,000 rpm for 10 min at room temperature. Supernatants were transferred to fresh tubes, and protein estimation was carried out using bicinchoninic acid (BCA) assay (Pierce, Waltham, MA). A total of 200 µg protein from each condition were reduced and alkylated with 10 mM dithiothreitol (DTT) at 60 °C for 20 min and 20 mM iodoacetamide (IAA) at room temperature for 10 min in the dark, respectively. The protein samples were then subjected to acetone precipitation with five volumes of chilled acetone at – 20 ºC for 6 h. Proteins pellets were obtained by centrifugation at 12,000 rpm for 15 min at 4 ºC and subjected to trypsin digestion with TPCK-treated trypsin (1:20) (Worthington Biochemical Corporation, Lakewood, NJ, USA) overnight at 37 ºC. Peptide fractionation was carried out using basic pH Reversed-Phase Liquid Chromatography (bRPLC) and strong cation exchange (SCX) as described earlier [11]. The samples were desalted using C18 Stage Tips prior to LC–MS analysis.
To analyze the secretory proteins, the culture filtrate was collected by centrifugation of the Brucella culture grown in nutrient-deprived media and filtered through a 0.22 µm bacteriological filter to remove any bacterial traces. The filtered media was then concentrated using 3KDa MWCO filters (Millipore). Protein estimation and sample preparation for proteomic analysis were carried out using the methodology described above. The resultant peptide digest was not subjected to any fractionation, and the data were acquired in technical replicates.
Data Acquisition on Orbitrap Fusion Tribrid Mass Spectrometer
Mass spectrometric analysis of the samples derived from the culture pellets and the secretome fractions was carried out using an Orbitrap Fusion Tribrid mass spectrometer (Thermo Fisher Scientific, Bremen, Germany) coupled to Easy-nLC1200 nano-flow UHPLC (Thermo Scientific, Odense, Denmark). The data were acquired for each fraction in technical replicates. Briefly, tryptic peptides obtained from both high pH and SCX fractionation were reconstituted in 0.1% formic acid and loaded onto trap column nanoViper (75 µm × 2 cm, 3 µm C18 Aq) (Thermo Fisher Scientific, Bremen, Germany). The peptides were further resolved using EASY-Spray C18 analytical column (75 µm × 50 cm, PepMap C18, 2 µm C18 Aq) (Thermo Fisher Scientific, Bremen, Germany) set at 40 °C. The flow rate was set as 300 nl/min using a binary solvent system containing solvent A: 0.1% formic acid and solvent B: 0.1% formic acid in 80% acetonitrile. A linear gradient of 10–35% solvent B over 100 min followed by a linear gradient of 35–100% solvent B for 8 min was employed to resolve the peptides. The column was re-equilibrated to 5% solvent B for an additional 12 min. The total run time was 120 min. Data-dependent acquisition in positive ion mode was employed MS1 survey scans were performed in an Orbitrap mass analyzer with a scan range of 400–1600 m/z at a mass resolution of 120,000 mass resolution at 200 m/z. Peptide charge state was set to 2–6, and dynamic exclusion was set to 30 s along with an exclusion width of ± 20 ppm. MS/MS fragmentation was carried out for the most intense precursor ions selected at top speed data-dependent mode with a maximum cycle time of 3 s HCD fragmentation mode with a collision energy of 33% was employed for MS2 scans. The fragment ions were detected within the scan range of 110–2000 m/z at a mass resolution of 30,000. The maximum injection time was specified as 200 ms. Internal calibration was carried out using the lock mass option (m/z 445.1200025) from ambient air.
Mass Spectrometric Data Analysis
The mass spectrometry-derived data were analyzed using proteome discoverer software version 2.1 (Thermo Fisher Scientific, Bremen, Germany). The MS/MS data were searched against the following databases: (i) B. melitensis protein database (RefSeq version 82 containing 3254 entries; including 3138 Brucella proteins and 116 contaminants) and (ii) 6-frame translated genome database. The protein database and genome sequence were downloaded from RefSeq NCBI. The 6-frame translated genome database (NC_012441.1 Brucella melitensis ATCC 23457, GCF_000022625.1_ASM2262v1) was generated using in-house Perl scripts (https://github.com/beherasan/Sixframe). Using these scripts, the genome was translated between successive stop codons. The data were searched using both SEQUEST and Mascot search engines. The search parameters included trypsin specified as the protease, and a maximum of one allowed missed cleavage. The dynamic modifications used for the database search included oxidation of methionine and acetylation of protein at the N-terminus. The static modification was set as carbamidomethylation of cysteine. Precursor and fragment mass tolerance were set to 10 ppm and 0.05 Da, respectively. A false discovery rate (FDR) cut-off of 1% peptide spectral match and 1% peptide level was used for identifications using a decoy database.
Bioinformatics Analysis
For label-free quantitative proteomic comparison, Intensity-Based Absolute Quantification (iBAQ) values were calculated for the identified proteins [18]. The intensities of all peptides were extracted from the peptide-spectrum matches (PSM) tables from three searches on Proteome Discoverer 2.1 (PD). The theoretically observable peptides of lengths from 7 to 35 residues were generated from B. melitensis protein sequences using an in-house Perl script (https://github.com/beherasan/Sixframe). To obtain the iBAQ values, the total intensity of all the identified peptides for a given protein was divided by the total number of theoretically observable peptides for the same protein [12]. Fold change for each protein was calculated by dividing the average replicate iBAQ values in nutrient-rich conditions by that in nutrient-limited conditions and log2-transformed.
The Gene Ontology (GO)-based functional classification of proteins was carried out using eggNOG (evolutionary genealogy of genes: Non-supervised Orthologous Group) (v4.5, 2015) [13] (http://eggnog.embl.de). STRAP 1.5 (Software Tool for Researching Annotations of Proteins) programs [14, http://www.bumc.bu.edu/cardiovascularproteomics/cpctools/strap/]. SecretomeP2.0 [15, http://www.cbs.dtu.dk/services/SecretomeP] was employed to categorize identified proteins into classical and non-classical secretory proteins. PSORTB [16, http://www.psort.org/] was used to predict the subcellular localization of the identified secretory proteins. Sets of upregulated (fold change ≥ 2.0) and downregulated (fold change ≤ 0.5) genes along with their classification (molecular function, biological process) from the COG analysis were selected. An R package-GOPlot [17] was used to create the chord diagram representing a subset of differentially expressed proteins.
Proteogenomics Analysis
Peptides that mapped exclusively to the 6-frame translated genome database were referred to as Genome Search-Specific Peptides (GSSPs). Peptides that mapped to multiple genomic regions and those that did not qualify for the 1% FDR threshold at the peptide level were not considered for the analysis. Proteogenomic classification of peptides was manually ascertained using the Integrative Genomics Viewer [18]. GSSPs were used to either identify novel genes or refine the existing gene models by IGV-based visual comparison with the reference genome. The quality of MS/MS spectra for the GSSPs that supported the novel events was manually verified to ensure correct peptide assignments. Sequence alignment was performed using NCBI BLAST analysis to identify orthologous evidence [19]. Domain analysis was carried out using the SMART database [20]
Data Submission
The mass spectrometry proteomics data were deposited to the ProteomeXchange Consortium via the PRIDE partner repository with the dataset identifier PXD012396.
Results
Proteomic Analysis of Brucella melitensis
In the present study, we report an in-depth proteome analysis of B. melitensis strain ATCC23457. Bacteria cultured in complete media (nutrient-rich) and media depleted with protein supplements (nutrient limitation) were considered for the analysis. Additionally, the conditioned media containing the secreted proteins were processed. Figure 1 demonstrates the schematic workflow of the experimental workflow employed. In all, 53 mass spectrometry analyses identified 27,279 peptides corresponding to 2440 proteins (Supplementary Table S1). The overall differences and overlap in the protein expression between the experimental conditions indicate protein-coding evidence for approximately 78% of the B. melitensis proteome, the largest reported to date (Fig. 2A). Of these, 2269 proteins were identified in nutrient-rich samples, while 2286 and 850 proteins were identified in nutrient-limiting conditions and culture filtrates, respectively (Fig. 2B).
Experimental design of the quantitative proteomics of B. melitensis ATCC23457. iBAQ based label-free quantitation was employed to compare the global proteomic changes of B. melitensis cultured in nutrient-rich and limited conditions. The conditioned media from the nutrient limited state was harvestedand proteins were extracted from all conditions and proteolytically digested using trypsin. Peptides were fractionated using high pH RPLC and StageTip-based SCX fractionation and analyzed on Orbitrap Fusion Tribrid mass spectrometer coupled online with nanoLC. The unassigned spectra were searched against a 6-frame translated genome database for the identification of novel genes and refining existing gene models
A Coverage of the B. melitensis proteome by high-resolution mass spectrometry. B Venn plot of protein identification overlaps among the proteins identified in control, nutrient stress and conditioned media. C Overview of the COG analysis of proteins identified in control and nutrient stress. Depicted in the graphical representation. D Summary of the workflow employed to analyze secreted proteins. SecretomeP analysis was employed to determine the secretory nature and localization of the proteins identified in conditioned media. E Categorization of proteins identified in the secretome fraction based on their secretory nature (i.e., classical, non-classical). F Categorization of proteins identified in the secretome fraction based on their localization as predicted by PSortB (i.e., cytoplasm, outer-membrane, and periplasm)
Functional Annotation of the Brucella melitensis Proteome
Functional categorization of the identified proteins was carried out using Clusters of Orthologous Groups (COG) analysis. Overall, 2203 were annotated in the COG database representing 20 of the 24 COG categories. The most frequent COGs observed apart from the function unknown category (S), were for proteins encoded by genes for amino acid transport and metabolism (E) (10.6%), transcription (K) (7.9%), energy production and conversion (C) (7.17%) and translation, ribosomal structure and biogenesis (J) (6.8%) (Fig. 2C). Classification based on cellular components revealed 30% (243) proteins localized in the cytoplasm and 20% (164) localized on the plasma membrane (Supplementary Fig. 2A). Classification based on molecular function revealed a vast majority of the proteins to be involved in the catalytic activity (49%, 989 proteins), binding activity (38%, 770 proteins) and structural molecule activity (3%, 53 proteins) (Supplementary Fig. 2B). Furthermore, several proteins described previously as putative with uncharacterized functions were identified. In all, we confirmed the expression of 365 proteins designated as hypothetical proteins (Supplementary Table S2). Of these, 60 proteins were assigned to COG categories, including cell wall/membrane/envelope biogenesis (M, 17 proteins), replication, recombination and repair (L, nine proteins), and transcription (K, six proteins), among others. 159 functionally uncharacterized hypothetical proteins were confirmed at the protein level in this study.
Identification of Potentially Secreted Proteins
To identify potentially secreted proteins in B. melitensis, we analyzed the identified proteins using SecretomeP (cut-off score > 0.5), which predicted 357 proteins to be secretory (Fig. 2D). Of these, 161 and 196 proteins were categorized into classical and non-classical secretory proteins, respectively, based on the presence or absence of a signal peptide sequence. Among the 161 classical secretory proteins, we identified 154 in the secretome fraction, confirming their secretory nature (Fig. 2E). We further investigated the predicted subcellular localization of the secretory proteins using PSORTb analysis. Our analysis revealed that ~20% localized to the membrane or extracellular space, 16% localized to the periplasm, and 21% were predicted with cytoplasmic localization (Fig. 2F). Several hypothetical proteins were also identified in the culture filtrate, of which 31 proteins were predicted to be secretory based on SecretomeP analysis including BMEA_RS07250 and BMEA_RS00375, which were identified with four and nine unique peptides, respectively (Supplementary Table S2). Representative MS/MS spectra of two unique peptides mapping to BMEA_RS00375 are provided in Supplementary Fig. 2.
Refinement of Annotated Gene Models Using Mass Spectrometry-Derived Data
We identified 241 Genome Search-Specific Peptides (GSSPs) from the mass spectrometry-derived data mapping to the six frame translated B. melitensis genome. These GSSPs were categorized into peptides mapping to intergenic regions, the alternative frame of translation, and N- and C-terminal extensions based on their positions on the reference genome. Altogether, our analysis identified eight novel ORFs and protein-coding evidence for 28 putatively annotated pseudogenes. Further, we also provide evidence of genome annotation errors and proof for correcting translational start sites for seven proteins (Supplementary Table S3). Of the eight novel genes identified, evidence from orthologous species supports the protein-coding potential of six genes. Among the novel ORFs, CSBMM_NG_4 encodes an ORF of 152 amino acids was identified with 2 unique peptides. The putative novel ORF is translated from the reverse strand. Orthology evidence revealed minimal conservation across bacterial species (Fig. 3A). Similarly, another ORF, CSBMM_NG_1, encodes a protein with 952 amino acids and was identified with six unique peptides. Orthology analysis revealed conservation across Brucella sp, and SMART analysis revealed the presence of transmembrane and major facilitator superfamily (MFS) domains. CSBMM_NG_3 encoding an ORF of 58 amino acids identified with two unique peptides. Domain analysis predicts that this short ORF is likely to be secreted as it contains a signal peptide. Orthology analysis further revealed conservation across Brucella sp. and Ochrobactrum sp suggesting that this protein may likely be a secretory virulence factor. (Supplementary Table S3).
A An example of a novel ORF – CSBMM_NG_4 encoded by B. melitensis ATCC 23457 genome using the proteogenomics approach. An ORF of 152 amino acids was identified with two unique peptides. B An example of protein-coding potential of a gene BMEA_RS1170 annotated as a pseudogene. The figure illustrates multiple peptide mapping to the genomic regions currently annotated as a pseudogene. The annotated frame of translation of pseudogene is +2 and the GSSPs identified in our study correspond to +3 frame of translation. Orthologous evidence suggests conservation across Brucella sp. Functional domain and a representative MS/MS spectra of one of the identified peptide “GAENKDAGLDFIAFASEPEHQVK” is provided
In the present study, 28 annotated pseudogenes were identified with several peptides, confirming them as protein-coding genes (Supplementary Table S3B). The current annotations in the genome were likely misread as pseudogenes owing to genome sequencing errors resulting in mutations introduced, causing frameshifts or premature stop codons. We identified peptide mapping to seven pseudogenes with in-frame stop codon and 21 with frameshift mutations. These include plasmid partitioning protein RepB, AraC family transcriptional regulator, ubiquinol oxidase subunit II, 3-hydroxyacyl-CoA dehydrogenase.BMEA_RS11770 (Fig. 3B) and BMEA_RS15875 (Supplementary Fig. 3) were identified with eight and ten unique peptides mapping to the genomic region. The current annotations are possible genome sequencing errors as we found orthologous evidence within B. melitensis spp. suggesting that these genes encode ABC transporter substrate-binding protein (WP_087932809.1) and plasmid partitioning protein RepB (WP_011005511.1), respectively. However, in these strains, the current annotation is based on computational prediction. Our analysis thus provides translational evidence of these proteins in B. melitensis spp. Similarly, pseudogene BMEA_RS06610 was identified as a protein-coding gene encoding for NAD (P)/FAD-dependent oxidoreductase (WP_006137125.1) with three peptides. Our analysis also resulted in the identification of peptides mapping to seven proteins with erroneously annotated TSS. For example, two peptides were observed with 4 and 2 PSMs, respectively, and mapped to the upstream region of an existing ORF that encodes 50S ribosomal protein L21. Since the time of the analysis, eight gene annotations including two novel genes, four pseudogenes and two events of N-terminal extensions, have been corrected and are now available in the latest version of the RefSeq database. The details are provided in Supplementary Table S3.
Label-Free Proteomic Analysis Reveals Differential Expression of Proteins Upon Nutrient Limitation
We next aimed to analyze the differential expression patterns of the proteome of B. melitensis upon nutrient limitation, as it is intuitive that for a facultative intracellular pathogen, adaptability to the host environment is vital for long-term persistence. Employing label-free quantitative proteomic analysis, differential expression of 270 proteins (P < 0.05) was observed. Of these, 104 were > twofold overexpressed, and 166 proteins were < twofold downregulated (Fig. 4A). A total of 210 and 86 proteins were identified with restricted expression in the nutrient-rich and nutrient-limited states, respectively, suggestive of stress response mediated differential expression. Additionally, several enzymes involved in metabolic pathways, such as hydA (dihydropyrimidinase), gltD (dihydropyrimidine dehydrogenase) responsible for pyrimidine metabolism, and lpdA (dihydrolipoyl dehydrogenase), a flavoprotein disulfide reductase enzyme (FDR) responsible for the conversion of dihydrolipoamide to lipoamide [21] were identified with restricted expression in the nutrient-rich state.
A Volcano plot of quantified proteins from cells under stress. Relative protein expression upon nutrient limitation was compared with nutrient-rich condition. B GOChordplot depicting selected differentially expressed proteins in response to nutrient limitation and their associated COG terms. The color coding red to blue depicting the fold change in response to nutrient limitation as compared to nutrient-rich state (Color figure online)
COG analysis of the differentially expressed proteins further revealed enrichment of amino acid metabolism (E), transcription (K), energy production and conversion (C), and biogenesis (J) in the limited nutrient states compared to the nutrient-rich state (Supplementary Fig. 4). A subset of differentially expressed genes and their association with COG terms in the two conditions are represented in Fig. 4B. Additionally, several proteins described previously as putative with uncharacterized functions including 66 and 23 proteins were identified with restricted expression in the nutrient-rich and nutrient-limited conditions, respectively. In comparison with the nutrient-rich state, 32 and 49 hypothetical proteins were overexpressed and downregulated, respectively, in the nutritionally deprived state, suggesting its role in the adaptation or survival of bacteria in the nutrient-limited/stressed environment.
Functional enrichment analysis of the differentially expressed proteins revealed significant alterations in the expression of proteins involved inorganic substance metabolic processes, transmembrane transporters, urease pathway, carbohydrate metabolism, nitrogen metabolism, pyrimidine metabolism, and type IV secretion system, among others. Notably, metabolic processes such as the TCA cycle, peptidoglycan metabolism, and folate synthesis were not altered in response to nutrient limitation. The significantly regulated pathways and the fold change expression of proteins upon nutrient limitation are listed in Table 1.
Alterations in Carbohydrate Transport and Metabolism
We identified increased expression of several MFS transporters by over twofold. Several enzymes involved in the synthesis and utilization of carbohydrates, including galactose mutarotase (BMEA_RS04105) (0.13-fold), trehalose utilization protein ThuA (0.35-fold), inositol 2-dehydrogenase (0.15-fold) and other sugars were found to be downregulated suggesting its vital role in bacterial growth. Additionally, N-acetylmannosamine kinase was downregulated by 0.36-fold upon nutrient limitation. Interestingly, several enzymes of the glycolysis pathway were identified with similar expression patterns in both nutrient-rich and nutrient-limited states.
Differential Expression of Putative Virulence Factors Upon Nutrient Limitation
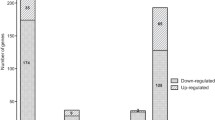
Our analysis demonstrates the repression of VirB operon in the nutrient-limited state compared to nutrient-rich conditions. Of the 12 virulence factors encoded by the VirB operon, we identified 9 proteins including VirB1, VirB4, VirB5, VirB6, VirB7, VirB8, VirB9, VirB10, and VirB11. Of these, VirB1, VirB4, VirB5, and VirB8 were identified with restricted expression in the nutrient-rich condition as shown in Fig. 5. In addition, we observed altered expression of quorum-sensing regulator VjbR (vacuolar hijacking Brucella regulator) (0.08-fold downregulated upon nutrient limitation), and GntR (Global Transcriptional regulator) in VirB operon (2.3-fold overexpressed).
In addition to the VirB operon, various Brucella flagellar proteins FlgH (L-ring protein), FlaF (Biosynthesis regulator), FliP (Biosynthesis protein),and MotB (motor) were identified with a restricted expression upon nutrient limitation,and FlgN was detected to be 5.38-fold overexpressed indicating that these proteins may be involved in stress response. Additionally, we identified heat shock proteins (Hsp) and molecular chaperones such as Hsp20 (2.03-fold up in response to nutrient limitation), while Tir and DnaJ were detected with > twofold expression in nutrient-rich conditions. We also identified RNA helicase to be upregulated over 11.3-fold upon nutrient limitation. We observed an increased expression of 3 cold shock proteins upon nutrient limitation.
Discussion
Brucella sp. possesses intrinsic mechanisms to adapt to environmental stress. Although the proteome profiles and their role in metabolic adaptation and pathogenicity have been reported in the case of B. suis and B. abortus, a detailed analysis of the proteome repertoire of B. melitensis has not been available to date. Towards this end, we performed an in-depth analysis of B. melitensis strain ATCC23457. We investigated the global proteome changes in response to nutrient limitation and also refined the existing genome annotation by providing translational evidence of several putative proteins. Additionally, expression profiles of several key proteins in terms of nutrient-rich and stress conditions were studied.
Pathogenic bacteria are known to contain specific secretory pathways that mediate the secretion of effectors and virulence factors. These factors are responsible for establishing host–pathogen interaction and its adaptation to the various environments within the host, sensing nutrient or environmental stress and are involved in substrate binding, adhesion, and cell–cell communication [22]. Employing in-depth proteome profiling, 2 440 proteins accounting for ~78% of the B. melitensis proteomewere identified. Of these we provided protein-coding evidence for 357 proteins corresponding to the secretory pathway, which included classical secretory proteins along with hypothetical proteins localized in periplasm, cytoplasm, and periplasmic space.
The utility of utilizing mass spectrometry-derived data to refine the genome annotation of organisms has been successfully demonstrated [11]. Employing a similar strategy, we identified 241 Genome Search-Specific Peptides (GSSPs) mapping to the six frame translated B. melitensis genome. These GSSPs were categorized into peptides mapping to intergenic regions, the alternative frame of translation, N- and C-terminal extensions based on their positions on the reference genome. Notably, the presence of pseudogenes is one of the characteristic features in the bacterial genome, as a majority of these are computationally annotated using comparative approaches such as aligning homologs, truncated, and disrupted sequence searching [23]. Previous comparative studies have reported around 5% of pseudogenes in the genome of Brucella sp, suggestive of genome reduction or reductive evolution [24]. In the present study, we provide protein-coding evidence for 28 annotated pseudogenes refuting their current status. It is likely that the current annotations are misread as pseudogenes owing to genome sequencing errors resulting in mutations introduced, causing frameshifts or premature stop codons.
We next aimed to study changes in Brucella proteins with respect to nutrient conditions. In the case of Brucella sp., primarily B. abortus, persistence within the host cells in conditions of extreme nutrient starvation is predominantly dependent on metabolic reprogramming [25, 26]. However, the effect of nutrient limitation on B. melitensis growth and proteome expression has not been investigated to a greater extent. Employing label-free quantitative proteomic analysis, 86 proteins were identified with restricted expression in the nutrient-rich and nutrient-limited states, respectively, suggestive of stress response mediated differential expression. Their decreased expression upon nutrient limitation suggests its dependence on protein-replete media for sustained growth and development [27]. COG analysis indicated that several proteins with restricted expression in the nutrient-rich and nutrient-limited conditions were described as putative with uncharacterized functions. We also found alterations with respect to several proteins involved in carbohydrate transport and metabolism. These proteins have also been demonstrated to be the prospective target for drug development against brucellosis [28, 29]. N-acetyl glucosamine (GlcNAc) and N-acetylmuramic acid (MurNAc) crosslinks are required to form the bacterial cell wall, and the enzymes needed for their synthesis were also identified in the present study with no significant differences in expression in the limited nutrient state [30]. Nutrient uptake by microorganisms from the external environment is essential for bacterial survival and persistence. Transporters such as Heme transporter proteins BhuA, multidrug transporter AcrB, and potassium transporter kup were overexpressed in response to nutrient limitation, indicating their role in bacterial survival. The ABC-type microcin C transport system has been reported to be involved in Brucella’s virulence via resistance to antimicrobial action [31].
Quorum-sensing regulator VjbR (vacuolar hijacking Brucella regulator), and GntR (Global Transcriptional regulator) are involved in the regulation of the VirB operon system and are essential for Brucella intercellular survival [32]. Further, VjbR has been suggested to be involved in bacterial adaptations to various environmental conditions by regulating bacterial cell division until it reaches the endoplasmic reticulum [33]. Weeks et al. demonstrated its role in the growth of B. melitensis in nutrient-rich and nutrient-limited media, i.e., increased expression of vjbR in culture-rich media [34]. Our results agree with the published literature that suggests that the Quorum sensing gene (vjbR) aids Brucella sp in adapting to various environmental conditions. Moreover, it has been demonstrated that the regulator vjbR cross-talks with other components in Brucella, such as the phosphoenolpyruvate phosphotransferase system (PTS), two-component system BvrS/R [35]. The phosphoenolpyruvate phosphotransferase system (PTS) comprises of cytoplasmic energy-coupling proteins (Enzyme I and HPr) and several carbohydrate-specific enzymes II and regulates the carbohydrate, energy conditions, and QS gene in the bacterial cell [36]. It has been suggested as one of the factors responsible for the adaptation of Brucella to a stressful environment. Viadas et al. in a study, demonstrated the restricted growth of Brucella abortus in minimal nutrient media with a limited carbon source, confirming the role of PTS and two-component system based on the environmental condition [37]. The differential expression of PTS in the present study confirms its activity upon nutrient limitation.
Differential expression of several of Brucella flagellar proteins was also observed in response to nutrient limitation. Brucella flagellar proteins comprising of flagellum genes FlgI (P-ring protein), FliK (Hook-length control protein), MotA (Motor), MotB, FlgM (Biosynthesis protein), FlgH (L-ring protein), FliI (flagellum-specific ATP synthase), FliP (Biosynthesis protein), and FlaF (Biosynthesis regulator) have been categorized as vital virulence factors required for persistent infection. [38].
Heat shock proteins and molecular chaperones play an essential role as a part of the protein repair system and are induced upon stress response in bacteria to adapt to the hostile environment of the host cell [10]. It has been previously reported that the nucleic acid-binding protein-DEAD-box containing RNA helicase is essential for the growth of extremophiles in conditions of low temperature and is also frequently associated with cold shock responses in bacteria [39]. We observed increased expression of three cold shock proteins upon nutrient limitation including CspA, a known RNA chaperone in E. coli, responsible for the regulation of multiple genes such as flagellar operon [40]. Overexpression in our analysis confirms the role of CspA in virulence as well as in stress-tolerance response.
Additionally, several genes responsible for the adaptation to other stress responses [41] were identified to be differentially expressed in the present study. Haine et al. demonstrated B. melitensis survival in low-oxygen/anaerobic conditions with upregulation of enzymes involved in denitrification and anaerobic electron transport [42]. It has been previously reported that B. suis utilizes denitrification with nitrogen oxide as a terminal electron acceptor during low-oxygen tension and infection [43]. Furthermore, B. abortus uses denitrification enzymes to reduce nitric oxide and ROS production by the host, thereby increasing its survival inside the macrophage [44].
During the development of Brucella sp., the niche or compartment (macrophage, epithelial cells) occupied by the bacteria are acidic for its replication. This low pH also acts as the stimuli for the expression of the pathogenic pathway such as Type IV secretion system [45]. Several genes such as HdeA and urease complex facilitate the adaptation of bacteria to acid stress environment. The chaperone HdeA known to be activated at low pH, was downregulated in response to nutrient limitation (0.29-fold). HdeA has been demonstrated to play a critical role in the acid-stressed environment in E. coli [46], and B. abortus [47]. Interestingly, we observed upregulation of structural subunits of the urease complex with ureA and ureB in response to nutrient limitation, whereas ureC subunit alpha 1 was unaltered. On the contrary, ureC subunit alpha 2 was detected with restricted expression in the nutrient-limited state. The accessory subunits—ureE, ureF, and ureG2 were found to be downregulated. Our analysis is thus indicative of key adaptive processes of Brucella that are affected by nutrient limitation. This, in turn, provides a broad scope to target these proteins intracellular using limiting nutrients and, therefore, can serve as vital therapeutic targets.
Conclusions
The in-depth proteomic characterization of Brucella melitensis ATCC23457 demonstrates the utility of high throughput mass spectrometry analysis in providing translational evidence for the expression of several putative protein-coding genes along with enabling the refinement of genome annotation. Further, our analysis elucidates the effect of nutrient stress on the expression of proteins involved in key biochemical and cellular pathways, including those involved in the survival and adaptation of the pathogen. The putative novel proteins identified in this study and the data demonstrating differential regulation of several hypothetical proteins with uncharacterized function merits further investigation concerning their potential roles in virulence, survival, and adaptation to various stress responses. The high-resolution proteomic landscape of B. melitensis presented in the study will serve as a baseline for future studies and foster future research in this area. Also, this study provides clues about B. melitensis metabolism in nutrient-deficient and nutrient-rich conditions, which could lead to future avenues of antimicrobial therapy research.
References
Corbel MJ (1997) Brucellosis: an overview. Emerg Infect Dis 3(2):213
Majalija S, Luyombo P, Tumwine G (2018) Sero-prevalence and associated risk factors of Brucellosis among Malaria negative febrile out-patients in Wakiso district,Central Uganda. BMC Res Notes 11(1):1–6
Subbannayya Y, Pinto SM, Gowda H, Prasad TK (2016) Proteogenomics for understanding oncology: recent advances and future prospects. Expert Rev Proteom 13(3):297–308
Kucharova V, Wiker HG (2014) Proteogenomics in microbiology: taking the right turn at the junction of genomics and proteomics. Proteomics 14(23–24):2360–2675
Yang Y, Hu M, Yu K, Zeng X, Liu X (2015) Mass spectrometry-based proteomic approaches to study pathogenic bacteria-host interactions. Protein Cell 6(4):265–274
Mol JP, Pires SF, Chapeaurouge AD, Perales J, Santos RL, Andrade HM, Lage AP (2016) Proteomic profile of Brucella abortus-infected bovine chorioallantoic membrane explants. PLoS ONE 11(4):e0154209
Lauer SA, Iyer S, Sanchez T, Forst CV, Bowden B, Carlson K et al (2014) Proteomic analysis of detergent resistant membrane domains during early interaction of macrophages with rough and smooth Brucella melitensis. PLoS ONE 9(3):e91706
Connolly JP, Comerci D, Alefantis TG, Walz A, Quan M, Chafin R et al (2006) Proteomic analysis of Brucella abortus cell envelope and identification of immunogenic candidate proteins for vaccine development. Proteomics 6(13):3767–3780
Wagner MA, Eschenbrenner M, Horn TA, Kraycer JA, Mujer CV, Hagius S et al (2002) Global analysis of the Brucella melitensis proteome: identification of proteins expressed in laboratory-grown culture. Proteomics 2(8):1047–1060
Wang Y, Chen Z, Qiao F, Ying T, Yuan J, Zhong Z, Huang L (2009) Comparative proteomics analyses reveal the vir B of B. melitensis affects expression of intracellular survival related proteins. PLoS ONE 4(4):e5368
Pinto SM, Verma R, Advani J, Chatterjee O, Patil AH, Kapoor S et al (2018) Integrated multi-omic analysis of Mycobacterium tuberculosis H37Ra redefines virulence attributes. Front Microbiol 9:1314
Schwanhäusser B, Busse D, Li N, Dittmar G, Schuchhardt J, Wolf J et al (2011) Global quantification of mammalian gene expression control. Nature 473(7347):337–342
Tatusov RL, Koonin EV, Lipman DJ (1997) A genomic perspective on protein families. Science 278(5338):631–637
Bhatia VN, Perlman DH, Costello CE, McComb ME (2009) Software tool for researching annotations of proteins: open-source protein annotation software with data visualization. Anal Chem 81(23):9819–9823
Bendtsen JD, Jensen LJ, Blom N, Von Heijne G, Brunak S (2004) Feature-based prediction of non-classical and leaderless protein secretion. Protein Eng Des Sel 17(4):349–356
Yu NY, Wagner JR, Laird MR, Melli G, Rey S, Lo R, Brinkman FS (2010) PSORTb 3.0: improved protein subcellular localization prediction with refined localization subcategories and predictive capabilities for all prokaryotes. Bioinformatics 26(13):1608–1615
Walter W, Sánchez-Cabo F, Ricote M (2015) GOplot: an R package for visually combining expression data with functional analysis. Bioinformatics 31(17):2912–2914
Prasad TK, Mohanty AK, Kumar M, Sreenivasamurthy SK, Dey G, Nirujogi RS et al (2017) Integrating transcriptomic and proteomic data for accurate assembly and annotation of genomes. Genome Res 27(1):133–144
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215(3):403–410
Letunic I, Doerks T, Bork P (2015) SMART: recent updates, new developments and status in 2015. Nucleic Acids Res 43(D1):D257–D260
Argyrou A, Vetting MW, Blanchard JS (2004) Characterization of a new member of the flavoprotein disulfide reductase family of enzymes from Mycobacterium tuberculosis. J Biol Chem 279(50):52694–52702
Gagic D, Ciric M, Wen WX, Ng F, Rakonjac J (2016) Exploring the secretomes of microbes and microbial communities using filamentous phage display. Front Microbiol 7:429
Lerat E, Ochman H (2005) Recognizing the pseudogenes in bacterial genomes. Nucleic Acids Res 33(10):3125–3132
Hong PC, Tsolis RM, Ficht TA (2000) Identification of genes required for chronic persistence of Brucella abortus in mice. Infect Immun 68(7):4102–4107
Roop RM, Gaines JM, Anderson ES, Caswell CC, Martin DW (2009) Survival of the fittest: how Brucella strains adapt to their intracellular niche in the host. Med Microbiol Immunol 198(4):221–238
Hidese R, Mihara H, Kurihara T, Esaki N (2012) Pseudomonas putida PydR, a RutR-like transcriptional regulator, represses the dihydropyrimidine dehydrogenase gene in the pyrimidine reductive catabolic pathway. J Biochem 152(4):341–346
Bao Y, Tian M, Li P, Liu J, Ding C, Yu S (2017) Characterization of Brucella abortus mutant strain Δ22915, a potential vaccine candidate. Vet Res 48(1):1–13
Singh AK, Balakrishna K, Sripathy MH, Batra HV (2014) Studies on recombinant glucokinase (r-glk) protein of Brucella abortus as a candidate vaccine molecule for brucellosis. Vaccine 32(43):5600–5606
Rest RF, Robertson DC (1974) Glucose transport in Brucella abortus. J Bacteriol 118(1):250–258
Vollmer W, Blanot D, De Pedro MA (2008) Peptidoglycan structure and architecture. FEMS Microbiol Rev 32(2):149–167
Wang Z, Bie P, Cheng J, Lu L, Cui B, Wu Q (2016) The ABC transporter YejABEF is required for resistance to antimicrobial peptides and the virulence of Brucella melitensis. Sci Rep 6(1):1–10
Haine V, Sinon A, Van Steen F, Rousseau S, Dozot M, Lestrate P et al (2005) Systematic targeted mutagenesis of Brucella melitensis 16M reveals a major role for GntR regulators in the control of virulence. Infect Immun 73(9):5578–5586
Delrue RM, Deschamps C, Léonard S, Nijskens C, Danese I, Schaus JM et al (2005) A quorum-sensing regulator controls expression of both the type IV secretion system and the flagellar apparatus of Brucella melitensis. Cell Microbiol 7(8):1151–1161
Weeks JN, Galindo CL, Drake KL, Adams GL, Garner HR, Ficht TA (2010) Brucella melitensis VjbR and C12-HSL regulons: contributions of the N-dodecanoyl homoserine lactone signaling molecule and LuxR homologue VjbR to gene expression. BMC Microbiol 10(1):1–20
Dozot M, Poncet S, Nicolas C, Copin R, Bouraoui H, Mazé A et al (2010) Functional characterization of the incomplete phosphotransferase system (PTS) of the intracellular pathogen Brucella melitensis. PLoS ONE 5(9):e12679
Deutscher J, Francke C, Postma PW (2006) How phosphotransferase system-related protein phosphorylation regulates carbohydrate metabolism in bacteria. Microbiol Mol Biol Rev 70(4):939–1031
Stülke J (2007) Regulation of virulence in Bacillus anthracis: the phosphotransferase system transmits the signals. Mol Microbiol 63(3):626–628
Sakaizawa T, Matsumura T, Fujii C, Hida S, Toishi M, Shiina T et al (2018) Potential role of ASC, a proapoptotic protein, for determining the cisplatin susceptibility of lung cancer cells. Tohoku J Exp Med 244(2):133–144
Wang Z, Wang S, Wu Q (2014) Cold shock protein A plays an important role in the stress adaptation and virulence of Brucella melitensis. FEMS Microbiol Lett 354(1):27–36
Jiang W, Hou Y, Inouye M (1997) CspA, the major cold-shock protein of Escherichia coli, is an RNA chaperone. J Biol Chem 272(1):196–202
Robertson GT, Roop RM (1999) The Brucella abortus host factor I (HF-I) protein contributes to stress resistance during stationary phase and is a major determinant of virulence in mice. Mol Microbiol 34(4):690–700
Haine V, Dozot M, Dornand J, Letesson JJ, De Bolle X (2006) NnrA is required for full virulence and regulates several Brucella melitensis denitrification genes. J Bacteriol 188(4):1615–1619
Köhler S, Foulongne V, Ouahrani-Bettache S, Bourg G, Teyssier J, Ramuz M, Liautard JP (2002) The analysis of the intramacrophagic virulome of Brucella suis deciphers the environment encountered by the pathogen inside the macrophage host cell. Proc Natl Acad Sci USA 99(24):15711–15716
Wang M, Qureshi N, Soeurt N, Splitter G (2001) High levels of nitric oxide production decrease early but increase late survival of Brucella abortus in macrophages. Microb Pathog 31(5):221–230
Porte F, Liautard JP, Köhler S (1999) Early acidification of phagosomes containing Brucella suis is essential for intracellular survival in murine macrophages. Infect Immun 67(8):4041–4047
Gajiwala KS, Burley SK (2000) HDEA, a periplasmic protein that supports acid resistance in pathogenic enteric bacteria. J Mol Biol 295(3):605–612
Valderas MW, Alcantara RB, Baumgartner JE, Bellaire BH, Robertson GT, Ng WL et al (2005) Role of HdeA in acid resistance and virulence in Brucella abortus 2308. Vet Microbiol 107(3–4):307–312
Acknowledgements
We thank Karnataka Biotechnology and Information Technology Services (KBITS), Government of Karnataka, for support to the Center for Systems Biology and Molecular Medicine at Yenepoya (Deemed to be University) under the Biotechnology Skill Enhancement Programme (BiSEP) in Multiomics Technology. SMP is a recipient of INSPIRE Faculty Award from the Department of Science and Technology (DST), Government of India. SKB is a recipient of Bioinformatics National Certification (BINC)-Junior Research Fellowship from the Department of Biotechnology (DBT), Government of India. NA is a recipient of research fellowship assistance from YU. We thank Saketh Kapoor for his assistance in sample processing and data analysis.
Funding
This study was funded by Indian Council of Medical Research, Govt. of India [Grant No. Zon.15/11/2014-ECD-II].
Author information
Authors and Affiliations
Contributions
AAH, SMP, TSKP, HFD and RSK contributed to conception and design of the study. AAH, PRK, NMB, LRS, RSK provided samples for the study. SMP, and NA processed the samples. SMP, YS and SKB acquired and analyzed the data. AAH, SMP, YS and SKB prepared tables and figures. AAH, SMP, YS carried out the literature search and wrote sections of the manuscript. All authors contributed to manuscript revision, read and approved the submitted version.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Ethical Approval
All protocols for sample collection from humans were approved by the Ethical Committee of the Dr. G.M. Taori Central India Institute of Medical Sciences (CIIMS).
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.



Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Husain, A.A., Pinto, S.M., Agarwal, N. et al. Comprehensive Proteomic Analysis of Brucella melitensis ATCC23457 Strain Reveals Metabolic Adaptations in Response to Nutrient Stress. Curr Microbiol 80, 20 (2023). https://doi.org/10.1007/s00284-022-03105-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00284-022-03105-y