Abstract
A reduction in the use of agrochemicals requires the development of either alternatives or complementary control methods in order to limit their dangerousness. An alternative is the biological control of fungi by bacteria. The fungal cell wall is a unique structure of the fungi, composed of glucan, chitin, and glycoproteins. Therefore, bacteria producing mycolytic enzymes, like chitinases, are of great interest to degrade fungal cell-wall components. The objectives of this work were to isolate chitinolytic bacteria from the guano of insectivorous bats (Tadarida brasiliensis) and to verify the presence of antifungal activities against phytopathogenic fungi. From the guano samples, 28 bacterial isolates were obtained, 70% of which presented chitinolytic activity. Four isolates were selected since they showed the highest values of chitinase activity, and they were characterized as belonging to Bacillus genus, by analyzing the 16S ribosomal RNA gene sequence. Cell-free supernatants of bacterial cultures were used in inhibition tests on 16 fungi: Alternaria and Colletotrichum acutatum were the most affected. Chitinase and antifungal activities were observed in the cell-free supernatant regardless of the culture medium used. Both activities were stable to heat and proteinase K treatments. Finally, when the culture medium was supplemented with 1 ml of cell-free supernatants (0.33%) and incubated for 120 h, the inhibition of hyphae formation and germination spores of reporter fungus were observed under light microscopy. These results suggest the feasibility of using cell-free supernatants as eco-friendly fungicides. The use of them may contribute to reducing the dose of toxic chemicals.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In global agriculture, many microbial diseases are responsible for the deterioration of fruits, leaves, stems, and roots, and they are caused by invasive phytopathogenic fungi. More than 8000 fungal species are known to cause diseases in plants. The damage impacts on both economic and biological production since the growth and development of the host plants could be strongly affected. It is estimated that fungi cause losses of 5–25% in developed countries and of 20–50% in emerging countries [1].
At present, agricultural production depends mainly on chemical inputs such as fertilizers, pesticides, and herbicides. Excessive use and misuse of these chemicals have caused food contamination, weed and disease resistance, and negative environmental outcomes with a serious impact on human health. The application of these chemical inputs causes the accumulation of toxic compounds in the soil, harming the health of the environment in the long term [2]. To control phytopathogenic fungi, different strategies have been used, and some of them involve utilizing resistant plant varieties, planting transgenic seeds on suitable dates, or/and applying agrochemicals.
Since the 1940s, pest management in agricultural countries has been based mainly on the use of synthetic pesticides. About 2.3 billion kg of pesticides are used annually worldwide [Atwood and Paisley-Jones 2017 cited by 3]. The excessive use of these chemicals is accompanied by growing general concern and consumer awareness of the adverse impact of pesticides on the environment and animal and human health [3, 4]. Although the use of agrochemicals increases short-term crop profitability, it causes long-term environmental harm. Furthermore, their use gives rise to increasingly resistant phytopathogenic strains, which requires a higher dosage of agrochemicals as well as the search for new, more potent substances. These practices also impair the sustainable functioning of ecosystems because they eliminate beneficial soil organisms [5, 6].
In addition to their harmful effects on the soil, most agrochemicals are known to pose risks to human health. In particular, they cause skin and eye damage and may harm internal organs such as the lungs. Moreover, some of them are considered carcinogens by different international health organizations [7]. The reduction in agrochemical use requires the development of either alternatives or complementary control methods in order to limit its dangerousness. An interesting alternative to agrochemicals is the use of living organisms or products derived from them. This strategy, known as biological control or biocontrol, has been increasingly assessed as an environment-friendly methodology. One of the biocontrol approaches involves microorganisms producing mycolytic enzymes, often from bacterial sources (chitinases, glucanases), which are of great interest since they could degrade fungal cell-wall components. The fungal cell wall is a structure with high plasticity that protects the cells from different types of environmental stress. The structure of the fungal cell wall is unique to the fungi, and it is composed of glucan, chitin, and glycoproteins [8, 9]. Since humans lack the fungi cell-wall components, this structure is an excellent target for the development of antifungal substances that would not pose risks to humans.
Taking into account the great microbial and environmental diversity, the isolation of microorganisms in habitats that act as selection pressure and favor the expression of these enzymes is considered important since it will make it possible to find more competent microorganisms in relation to the desired characteristics. Whitaker et al. (2004) were the first to inform the presence of chitinolytic bacteria in the insectivorous bat guts, suggesting that these symbiotic bacteria help bats to digest their insect prey chitin [10]. For this reason, the aim of this work was to isolate chitinolytic bacteria from the guano of insectivorous bats to verify the presence of antifungal activities against phytopathogenic fungi of important crops. Also, this work attempted to elucidate whether there is a relationship between chitinolytic and antifungal activities.
Materials and Methods
Sampling Site and Bacterial Isolation
A total of three guano samples were aseptically collected from different areas inside Escaba dam vaults. Escaba dam is located in Juan Bautista Alberdi department, about 127 km southwest of San Miguel de Tucuman city (Argentina), at an altitude of 630 m.a.s.l. Samples were transported in sterile plastic bags and kept at 4 °C until use. Samples were aseptically homogenized; 1 g of each one was suspended in 50 ml of sterile distilled water and incubated at 30 °C with constant shaking at 200 rpm for 24 h. Then, the diluted samples were plated on standard nutrient (SN) agar (g L−1): NaCl, 6; Peptone, 15; yeast extract, 3; glucose, 1. After 72 h at 30 °C, well-isolated colonies were transferred to fresh plates to obtain pure cultures, which were maintained on SN agar slants and preserved at 4 °C. Also, their corresponding parallel duplicates were lyophilized.
Isolation of Chitinase-Producing Bacteria
For the isolation of chitinase-producing bacteria, SPI agar medium was used [11]. SPI agar was supplemented with colloidal chitin 0.5 (w/v) as sole carbon and energy source and labeled CC-SPI. Colloidal chitin was prepared from commercial chitin (Sigma Chemicals Co.) according to Hsu and Lockwood’s (1975) with minimal modifications [12]. In brief, chitin powder (10 g) was slowly added to 150 mL of concentrated HCl and kept for 60 min at 30 °C with vigorous stirring. The chitin colloidal suspension from the precipitate was then obtained by slowly adding 400 mL of water at 4–10 °C. The suspension was collected by filtration with suction on a paper filter and washed by suspending it in about 5 L of sterile distilled water until the neutral pH was reached.
Inoculum Preparation
Inoculum suspensions were obtained from the culture of microorganisms grown in SN liquid after 24 h incubation at 30 °C and 200 rpm. The cells were collected by centrifugation at 9000×g at 4 °C and washed with a sterile saline buffer. Finally, when microorganisms presented unicellular growth, the cell suspensions were adjusted to the turbidity of 2 McFarland standard with a sterile physiological solution (DO600 nm = 1). When microorganisms showed mycelial growth, a 2 mm diameter pellet was used.
Chitinolytic Activity
Qualitative and Semiquantitative Analysis of Chitinolytic Activity
The screening of chitinase-producing bacteria or qualitative assay was performed by inoculating 10 μl of each bacterial inoculum onto CC-SPI agar medium at 30 °C to evaluate the appearance of clear zones on the opaque matrix. A semiquantitative analysis of chitinase production was performed by seeding selected isolates onto CC-SPI plates with pH adjusted to 5.0, 7.0, or 9.0 with buffer solutions. Plates were incubated at 10, 20, 30, or 37 °C for eight days, and the following criteria were considered to further select the best producing isolates: growth and hydrolysis capacity of chitin. Growth was assessed by considering the diameter of the colonies measured in mm, whereas chitin hydrolysis capacity was estimated by measuring the distance between the outer edge of the colony and the outer edge of the clear hydrolysis zone or halo, in mm. Then, a hydrolysis factor (HF) was defined as:
where THH (total halo hydrolysis) is the distance from the outer edge of the colony to the outer edge of the hydrolysis halo (HH) multiplied by 2, while total size (TS) is the diameter of the colony plus THH (Fig. 1).
Quantitative Analysis of Chitinase Activity
Based on the HF results, the bacteria with the highest chitinase activity were selected to carry out the quantitative analysis. They were grown in three different culture media. Two of them contained chitin as the sole carbon source: CC-SPI and P-SPI. CC-SPI was prepared as mentioned above (Section see “Isolation of chitinase-producing bacteria”), while P-SPI was prepared by adding 100 mL of SPI 10 X to 900 ml of distilled water. Then, 10 g of fine powder of pupae (Drosophila melanogaster) crushed with mortar was added and sterilized by autoclaving. The powder of pupae was kindly provided by Dr. Obrutski. The third SN is a complex medium without chitin. These experiments were conducted to investigate whether the expression of chitinase activity needs chitin as an inductor and whether it would be possible to use an inexpensive and easier alternative to the commercial chitin.
Every 48 h for 8 days, 10 mL samples of the cultures were centrifuged at 10,000×g for 15 min at 4 °C, and the cell-free supernatants were used for chitinase activity assays, following the method of Yanai et al. (1992) [13]. The reaction mixture consisted of 250 µl of 0.5% colloidal chitin, 250 µl of 0.05 M potassium phosphate buffer (pH 7.0), and 500 µl of cell-free supernatant. The reaction mixture was incubated for 2 h at 37 °C, and after centrifugation at 4000×g for 2 min, 500 µl supernatant was mixed with 100 µl of 0.8 M boric acid. The pH of this mixed solution was adjusted to 10.2 with KOH. Next, the solution was heated for 3 min in boiling water. The mixture was cooled, and 3 ml of p-dimethyl aminobenzaldehyde (DMAB) (1 g of DMAB dissolved in 100 ml of glacial acetic acid containing 1% 6 N V/V hydrochloric acid) was added. Finally, the mixture was incubated for 20 min at 37 °C, and absorbance at 585 nm was measured against water as blank.
The results were expressed in units of chitinase activity: 1 milli Unit (mU) was defined as 1 µmol of N-Acetyl glucosamine released from colloidal chitin per minute. Specific chitinase activity was expressed per gram of protein (mUE) and total protein concentration was determined according to Bradford (1976) using bovine serum albumin as standard [14].
16S rRNA Gene Sequence Analysis
Total genomic DNA was extracted according to Martinez et al. 2002 [15], and PCR amplifications were performed in 25 µl reaction volume using 9700 Perkin-Elmer Thermal Cycler (Applied Biosystems, Foster City, CA, USA). Small subunit ribosomal genes sequences were amplified with the universal primers 27F (5′-AGA GTT TGA TCM TGG CTC AG-3′) and 1492R (5′-GGT TAC CTT GTT ACG ACT T-3′) using the following conditions: initial denaturation at 94 °C for 3 min; followed by 35 cycles of denaturation at 94 °C for 30 s; annealing at 58 °C for 30 s; and extension at 72 °C for 30 s. The final DNA concentration in the PCR products was made using Epoch (Biotek), and the integrity of the samples was evaluated through gel electrophoresis of 1.5% Agarose. The amplicons obtained were purified and sequenced by Macrogen Inc. (Seoul, Korea). The 16S rRNA gene sequences obtained from the selected isolates were edited with Mega 7. A comparison of these sequences with partial 16S rRNA gene sequences published in the GenBank was performed using the BLAST tool from the National Center for Biotechnology Information (NCBI) (http//: www.ncbi.nlm.nih.gov) to search for similar sequences in public databases.
In Vitro Antifungal Activity
The cell-free supernatants of selected chitinase-producing bacteria were assayed to evaluate their antifungal activity against 16 phytopathogenic fungi (Table 3). These fungi were plated onto potato dextrose agar (PDA) by spreading a volume (100 L) inoculum. Then, 6 mm diameter holes were punched aseptically, filled with 100 μl of sterile cell-free supernatants from chitinolytic bacteria, and incubated at 30 °C for 120 h. The antifungal activity was assessed by measuring (in mm) the fungal growth inhibition halo around the hole.
Effect of Heat and Proteinase K Treatment on Chitinase and Antifungal Activities
Aliquots of cell-free supernatants from the strains that showed chitinolytic and antifungal activities were subjected to thermal treatments at 50 °C and 100 °C for 60 min, while those supernatants incubated at 30 °C were used as reference. Replica aliquots were also processed with proteinase K for 60 min at 60 °C. After all treatments, the samples were used for measuring chitinase activity as described (Section see “Qualitative semiquantitative analysis of chitinolytic activity”). To evaluate the fungal growth inhibition, the treated supernatants were also used for performing the fungal growth inhibition assay on plate, using as reporter the most sensitive fungus according to the results in Section see “In vitro antifungal activity”. The methodology and criteria were the same as those used in the test (Section see “In vitro antifungal activity”).
Morphological Changes in Fungi Treated With Cell-Free Bacterial Supernatant
The cell-free supernatants obtained from cultures in SN medium after 120 h growth were used as antifungal. The cultivation conditions were those described in Section see “In vitro antifungal activity”. The reporter fungus was inoculated into 300 mL of PD (potato dextrose) liquid medium in duplicate. One of the cultures was used as a control by adding 1 mL of sterile SN medium, and the second one was supplemented with 1 mL of cell-free bacterial supernatant corresponding to a solution of 0.33% of the supernatant. The cultures were incubated at 30 °C and 200 rpm for eight days; aliquots were sampled every 48 h for macro and microscopic (500 x) evaluations.
Results
Sampling Site and Screening for Chitinase-Producing Bacteria
Samples were collected from bat guano deposits located in one of the nine Escaba dam vaults. The samples presented pH 7.0 and had a solid state with loose and disaggregated material composed mainly of insect remains.
A total of 28 morphologically different bacteria were isolated from guano samples. The isolated bacteria were labeled with a letter followed by a serial number. All of them were able to grow on plates with colloidal chitin as the only carbon source. However, only 20 isolates showed colonies surrounded by clear zones on CC-SPI plates after 72 h of incubation, indicating chitinase activity. These isolates were selected for further studies.
Semiquantitative Analysis of Chitinolytic Activity
To select the best conditions for microbial growth and chitin hydrolysis capacity, the semiquantitative assay was used at the three different pH and temperatures already mentioned. Tables 1 and 2 show the average results of three independent tests of the 20 isolates. Neither the isolates at pH 5.0 and at different temperatures nor those at 10 °C and at the different pH combinations presented growth and hydrolysis of chitin capacity, and therefore these results were not included in the tables.
Notably, the growth on CC-SPI agar was slow for all the isolates studied, showing signs of development after 48 h of incubation in some isolates and after 72 h in others. The growth was measured every 24 h for eight days. The colony diameter measured in mm was used to evaluate the growth of each bacterial isolate. The growth values found on day seven were the highest, and after that, no significant increase in growth was observed. As shown in Table 1, the highest growth was obtained at 37 °C at both pH 7.0 and pH 9.0.
Regarding chitin hydrolysis capacity, the distance in millimeters between the outer edge of the colony and the outer edge of the hydrolysis zone (halo) was considered. Only 40% of the isolates at pH 7.0 and 9.0 showed halo at 20 °C. In the remaining conditions, the isolates showed the highest hydrolysis capacity values and were similar among them. However, isolates A6 and A10 decreased their hydrolysis capacity at pH 9.0 at 37 °C and isolate A8 showed greater activity at pH 7.0 than at pH 9.0, regardless of the temperature.
Considering the two criteria used for this test, it was noted that, while the best temperature conditions for the growth of most of the isolates were observed at 37 °C, the best conditions in terms of hydrolysis capacity were similar at 30 °C and 37 °C. Based on these results, it was decided to use pH 7.0 and 30 °C in the subsequent tests, because it is a more convenient condition for laboratory work.
To select microorganisms with the best growth and chitinolytic capacity, a hydrolysis factor (HF) was estimated (Fig. 2). Accordingly, isolate A14 was the microorganism that presented the highest HF, followed by A8, A9, B2, B6, A6, A12, B3, and B4, which were selected for further studies.
Quantitative Chitinolytic Activity
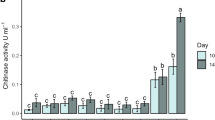
Cell-free supernatants from nine selected isolates grown in CC-SPI, P-SPI, and SN were assayed for chitinase activity (Fig. 3). All culture media were suitable for extracellular chitinase production for most of the isolates evaluated, except for B6, which did not show enzyme activity in SN medium, suggesting that the enzyme production would be substrate-dependent for this isolate. Remarkably, most of the cell-free supernatants obtained from SN media displayed higher activity values than those from the chitin-containing medium (CC-SPI), whereas the samples from P-SPI medium presented the lowest values. The cell-free supernatants retrieved from A9, B2, and B3 isolates produced similar chitinase activity in all the media tested. As a result, isolates A8, A9, A14, and B2 were selected for further characterization since they presented the highest chitinolytic activity.
Taxonomic Characterization of Selected Isolates
Comparison of 16S rRNA gene sequences from A8 (GenBank AM778822.1), A9 (GenBank AM778819.1), A14 (GenBank AM778820.1), and B2 (GenBank AM 778823.1) were found to be closely related to species of the genus Bacillus, showing >99% identity values with B. atrophaeus NBRC 15539 (GenBank AB363731.1), B. vallismortis DSM 11031 (GenBank NR_024696.1), B. amyloliquefaciens DSM7 (FN597644.1), and B. inaquosus strain NRRL B-23052 (EU138467.1).
Distinctive features were observed through macroscopic and microscopic observations, such as the mucous consistency of colony A8, the production of black pigment by A14, and the star-shaped morphology of colony B2.
In Vitro Antifungal Activity
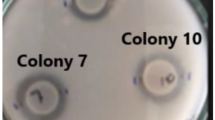
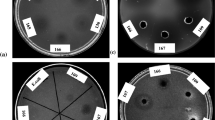
Antifungal activity against 16 fungal phytopathogenic species was evaluated by means of the diffusion well method, expressing the results according to the extent of inhibition of the fungal growth. Surprisingly, two types of halos were observed: a translucent halo corresponding to complete inhibition of fungal growth in this study was identified and named T, whereas a second type comprising opaque halos was observed, distinguished as O, which indicated a possible alteration of the mycelial development (Table 3).
Fungi Alternaria and C. acutatum were the most affected by all the cell-free supernatants, whereas fungi of the genus Fusarium were the most resistant. The cell-free extracts from Bacillus sp. A8 and A14 showed the highest antifungal activity on 12 out of the 16 tested fungi under the conditions used. By contrast, Bacillus sp. A9 and B2 showed a narrower spectrum of action, affecting 9 out of the 16 fungi. Pestalotia sp. (Palto phytopathogen) presented a complete growth inhibition with a translucent halo (Fig. 4B). This behavior was also observed in Geotrichum sp. and C. fragariae. C. acutatum showed both, translucent and opaque halos (Fig. 4C), while one strain of Fusarium exhibited only an opaque halo and the other Fusarium showed no inhibition (Figs. 4D and E, respectively).
Effect of Heat and Proteinase K Treatments on Chitinase and Antifungal Activities
Considering the results obtained, C. acutatum was selected to be used as reporter fungus of cell-free supernatants from Bacillus sp. A14. These supernatants have the potential to be used in pest control because this fungus is currently rotting many crops or causing anthracnose, mainly in blueberry and strawberry fields, leading to large economic losses when it is not controlled.
SN and CC-SPI were selected in this study because a difference was observed in chitinase activity in culture supernatants of Bacillus sp. A14. They were subjected to heat and proteinase K treatment in order to evaluate how these treatments affected both chitinase and antifungal activities.
Effect of Heat and Proteinase K Treatments on Chitinase Activity
The results obtained at 30 °C were taken as reference for this section. As shown in Fig. 4A, cell-free supernatants from SN showed the highest values of specific chitinase activity under all treatments, followed by CC-SPI (Fig. 5A). After heat and proteinase K treatments, the chitinase activity slightly decreased in the different cell-free bacterial supernatants, demonstrating high stability under working conditions.
Effect of Heat and Proteinase K Treatments on Antifungal Activity
The cell-free supernatants of Bacillus sp. A14 obtained from untreated SN produced the two types of halos on C. acutatum described in Section see “In vitro antifungal activity”. The opaque halo was constant when subjected to different treatments. However, the T halo decreased with heat treatments, reaching a complete reduction at 100 °C. By contrast, it was observed that the proteinase-treated supernatant produced a slight decrease in the T halo (Fig. 5B).
The supernatant obtained from untreated CC-SPI was also shown to produce two halos on C. acutatum. In this case, both halos were reduced by 50% with heat treatment at 50 °C. The T halo suffered a complete reduction, while the O halo underwent a slight reduction with respect to the treatment at 50 °C. In the case of the supernatant treated with proteinase k, the total T halo was completely reduced (Fig. 5C).
Morphological Changes on Fungi Treated with Cell-Free Bacterial Supernatant
As shown in Fig. 5, the culture of C. acutatum treated with the cell-free supernatant of Bacillus sp. A14 showed significant differences with respect to the reference culture. The latter presented low suspension of conidia and large pellet formation on the bottom of the Erlenmeyer. In the treated culture, abundant suspension of pink spores and small pellet formation were observed (data not shown).
Microscopic observations were carried out on fresh cultures after 120 h of incubation. Significant differences were observed in the abundance and development of reproductive structures between the reference and treated cultures. In the reference culture, there were few spores in suspension, a large number of well-formed budding structures, and an abundance of hyphae in the pellet (Fig. 6A). In contrast, a large number of suspended spores were observed in the treated culture, which provided a pink coloration to the medium, as well as numerous sporulation structures. There were also few sprouting structures and malformations (Fig. 6 B, C).
Discussion
The colony of Tadarida brasiliensis bats, also known as the Brazilian free-tailed bat, sheltering in one of the vaults of the Escaba dam, was estimated at 10 million individuals in 1984 by the biologist Rubén Barquez from Miguel Lillo Institute of the National University of Tucumán. This colony is one of the largest in South America, and due to its ecological interest, it was declared a “natural monument” by Provincial Law N 7058/2001. Therefore, it was the selected source of guano sampling to isolate chitinolytic bacteria.
These mammals are primarily insectivores; they consume a significant number of insects every night, so they are valuable for insect control. Remains of exoskeletons that have not been digested by the digestive system of bats can be found in guano deposits. Some authors isolated and identified chitinase-producing bacteria from the gut content of insectivorous bats, which use these symbiotic bacteria to assimilate the chitin of their insect prey [10, 16]. To the best of our knowledge, no previous studies about chitinolytic microorganisms isolated from bat guano have been reported. The present study revealed that the guano of insectivorous bats is an appropriate environment for isolating chitinolytic bacteria without sacrificing the animal.
The use of culture media based on colloidal chitin as a screening method for chitinase-producing bacteria has been reported by several authors, such as Shang et al. (2004) [17], who recovered a strain of Aeromonas schubertii from the soil using 0.2% w/v colloidal chitin. On the same basis, Sastoque-Cala et al. (2007) isolated nine strains of chitinolytic microorganisms belonging to different morphological groups, from shrimp waste at a Colombian fishing industry [18]. However, in this case, they used concentrations of 0.5% w/v commercial colloidal chitin and alkaline pH (pH 9.2). In the present work, the strategies used by these authors were taken as reference for the selection of chitinolytic microorganisms. Following Sastoque-Cala et al. (2007), the colloidal chitin concentration was adjusted to 0.5% w/v but the pH was fixed at 7.0, respecting the pH value that the bat guano had. This technique made it possible to isolate several microorganisms capable of degrading chitin from bat guano. Four chitinolytic isolates selected for this study belonged to the genus Bacillus, which is the most widely used bacterial genus for metabolism production at the industry level as well as for biological control strategies [19]. For bacterial identification, the analysis of the 16S rRNA gene sequence is the most widely used methodology. However, this analysis is limited for certain bacterial groups that have a smaller variation between closely related species. A clear example of this is the group B. subtilis, whose members showed similarities of more than 99% [20]. Within this group, B. atrophaeus is characterized by the production of black pigments, like A14 isolate, a feature absents in the other species [21].
Chitinase activity is frequent in Bacillus species which could be induced by chitin oligomers in the culture media. For example, Mabuchi et al. (2000) showed that B. cereus produces two inducible chitinases, A (chia) and B (chib), possibly chib being responsible for extracellular chitinase activity [22]. These authors tested from monomer to hexamer molecules of GlcNAc to demonstrate that the enzyme activity grew as the chain length of chitin oligomer increased. Conversely, carbon sources such as glucose, N-acetyl-β-D-glucosamine, or deacetylated chitin and chitosan products may act as catabolic repressors of chitinases [23,24,25]. Furthermore, various additional carbon sources were shown to influence the chitinase production. Gomaa (2012) demonstrated that chitinase production was enhanced in B. thuringiensis when galactose was supplemented with colloidal chitin, whereas lactose increased chitinase production by B. licheniformis [26]. The effect of glucose was also tested, and it was found that 0.5% of this carbon source inhibited chitinase production by B. thuringiensis yet potentiated its production by B. licheniformis. In the present study, it was observed that Bacillus sp. A14 expressed lower chitinolytic activity when grown in media with chitin (CC-SPI, P-SPI) than in the rich medium source. Our results suggest that Bacillus sp. A 14 is able to express chitinases in the basal state without the presence of an inducer such as chitin. These results indicate that the repressor or inducer substance of chitinase expression differs among organisms.
Several results suggested that chitinase is directly associated with the antagonist activity of fungi. For example, it was reported that crude supernatants with chitinolytic activity from B. thuringiensis exhibited an inhibitory effect against Aspergillus niger [27]; and a chitinase of B. cereus, cloned and purified from the periplasmic fraction of Escherichia coli DH5a, inhibited conidial germination of Botrytis elliptica B061 effectively [28]. In addition, De la Vega et al. (2006) reported that a purified chitinase from B. thuringiensis subsp. aizawai showed lytic activity against the phytopathogenic fungi cell wall, inhibiting six mycelial growths of Fusarium sp. and Sclerotium rolfsii [29]. Also, crude and purified chitinase from B. pumilus revealed higher mycelium inhibition against Aspergillus and Fusarium strains [30, 31]. Recently, Dukare et al. (2020) identified, characterized, and evaluated native chitinolytic rhizobacteria with biocontrol potential against pigeon pea wilt disease, caused by F. udum [32]. Two promising bacterial strains (Bacillus spp. NS-22 and Pseudomonas spp. NS-1) were selected for their chitinolytic potential and possession of different antifungal traits. In this context, we checked whether cell-free supernatants containing chitinolytic activity also possess antifungal activity against the locally relevant phytopathogenic fungi. Our results showed that Fusarium sp. was not affected, although strong inhibitory effects were observed for members of Colletotrichum and Alternaria genera, which cause serious diseases to berry plants. Notably, the supernatants of Bacillus sp. that showed the highest chitinolytic activity were also those that showed the highest antifungal effect.
To inhibit the chitinase activity of Bacillus sp. A14 supernatants, they were exposed to various treatments (proteinase K and heat). However, conclusive evidence of inhibition was not reached because the chitinolytic activity was scarcely affected after the treatments.
Biochemical studies on purified chitinases, frequently produced by Bacillus spp., showed that they present optimal activity at high temperature as well as a wide temperature range [26, 33, 34]. In fact, Yuli et al. (2004) noted that chitinase activity remained after five hours at 65 °C and after one hour at 80 °C [33]. Also, a chitinase from B. thuringiensis was active from 30 °C to 120 °C, and the partially purified enzyme showed maximum activity at 110 °C [35]. In agreement with these findings, the chitinolytic crude extract from Bacillus sp. A14 was not affected by incubation at 100 °C for one hour. On the other hand, the proteinase K treatment did not significantly alter the enzyme activity. Liu et al. (2011) reported a stable chitinase purified from Bacillus subtilis SL-13 with antifungal activity that endured sterilization conditions, UV, and treatment with 15 mg mL−1 proteinase K for 1.5 h [8].
Members of the Bacillus genus are also known to produce a wide range of antimicrobial compounds, the most bioactive molecules of which are synthesized non-ribosomal peptides and lipopeptides [36]. B. amyloliquefaciens PGPBacCA1 cells were applied on bean seeds and they were able to inhibit the development of several phytopathogenic fungi. The UV-MALDI TOF MS analysis showed that B. amyloliquefaciens PGPBacCA1 co-produces different homologs of the lipopeptides surfactin, iturin, and fengycin. These compounds were found to be responsible for the antagonistic effect [37]. Similarly, a study of B. methylotrophicus strain named XT1 CECT 8661 with activity against Botrytis cinerea detected several lipopeptides–surfactin, bacillomycin, and fengycin–in XT1 cultures using Q-TOF electrospray mass spectrometry analysis. In vitro antibiosis studies demonstrated the efficiency of the lipopeptide fraction in the inhibition of fungal growth [38]. Other authors reported that a purified peptide (designated as P657) from B. amyloliquefaciens showed antagonistic ability against B. cinerea. The biological activity of P657 was stable at a temperature as high as 100 °C for 20 min, and the treatment with trypsin and protease K had little effect on the inhibitory activity [39].
A key observation from this work is the sprouting inhibition shown for C. acutatum by the supernatant of the strain Bacillus sp. A14, since the germination of conidia represents the first step in the activation of an asexual life cycle and the spread of the disease of several aerial phytopathogenic fungi. Therefore, if it is possible to inhibit this stage, this disease may be successfully controlled. The evaluation of the antagonistic activity against Fusarium strains of the isolated extracellular culture supernatant of Bacillus showed that mycelial growth, conidial germination, and conidial production were effectively inhibited. Furthermore, the supernatant from the extracellular culture caused mycelial malformation and ultrastructural changes [37, 40, 41]. A bacterial strain, identified as B. amyloliquefaciens BA17, exhibited a strong antifungal effect against B. cinerea. The filtrate of BA17 liquid culture, when applied at 1%, 5%, and 10%, significantly reduced the spore germination, spore production, and mycelial growth of B. cinerea. The assessment of activities of intrasporic and extrasporic antagonistic substances of BA17 indicated that active substances were primarily produced outside the bacterial cell [42].
Contrary to the results presented here, the active substances produced by BA17 were sensitive to heat, while the antifungal activity of lipopeptides and P657 peptide mentioned above [38, 39] was thermostable according to the results of the present work.
Conclusion
Based on the results obtained, we speculate that the antifungal action of the extracellular culture supernatant of Bacillus sp. A14 is the result of the joint action of chitinase and other compounds like lipopeptides. Even though it was not possible in this work to attribute the antifungal activity observed solely to chitinases, the availability of a cell-free supernatant with hydrolytic activity and a thermostable antagonistic activity on phytopathogenic fungi of agricultural crops of interest provides a valuable product with low production and storage costs for future use in biocontrol processes. Working with cell-free supernatants for field applications has advantages over the use of living organisms, because a microorganism released to the environment will not respond in the same way as it does in the laboratory, where all parameters are controlled. Consequently, the whole repertoire of antifungal metabolites could be constrained in natural environments, or the microorganism can be displaced by competition with others present in the soil and vice versa, harming native microorganisms. Moreover, the thermostability of the antifungal activity suggests that the supernatants can function in harsh environments. In future studies, purifications will be carried out in order to accurately identify the bioactive molecules responsible for the antifungal activity of the supernatants Bacillus sp A14. It is also planned to carry out proteomic studies will extend the field of research to identify the proteins present in the cell-free supernatants.
References
FHIA (2007) http://www.fhia.org.hn/dowloads/fhia_informa/fhiainfdic2007.pdf. Accessed 2 Jun 2021
Alori ET, Babalola OO (2018) Microbial Inoculants for improving crop quality and human health in Africa. Front Microbiol. https://doi.org/10.3389/fmicb.2018.02213
Berini F, Katz C, Gruzdev N, Casartelli M, Tettamanti G, Marinelli F (2018) Microbial and viral chitinases: attractive biopesticides for integrated pest management. Biotechnol Adv 36(3):818–838. https://doi.org/10.1016/j.biotechadv.2018.01.002
Van Wambeke E (2007) Combinations of reduced rates of 1,3-dichloropropene and dazomet as a broad spectrum soil fumigation strategy in view of methyl bromide replacement. Commun Agric Appl Biol Sci 72:61–70
Paoletti MG (1999) The role of earthworms for assessment of sustainabilityand as bioindicators. Agric Ecosyst Environ 74:137–155. https://doi.org/10.1016/S0167-8809(99)00034-1
Frampton GK, Jansch S, Scott-Fordsmand JJ, Römbke J, Van den Brink PJ (2006) Effects of pesticides on soil invertebrates in laboratory studies: a review and analysis using species sensitivity distributions. Environ Toxicol Chem 25(9):2480–2489. https://doi.org/10.1897/05-438r.1
Siemiatycki J, Richardson L, Straif K, Latreille B, Lakhani R, Campbell S, Rousseau MC, Boffetta P (2004) Listing occupational carcinogens. Environ Health Perspect 112(15):1447–1459. https://doi.org/10.1289/ehp.7047
Liu Y, Tao J, Yan YJ, Li B, Li H, Li C (2011) Biocontrol efficiency of Bacillus subtilis SL-13 and characterization of an antifungal chitinase. Biotechnol Bioeng Chin J Chem Eng 19:128–134. https://doi.org/10.1016/S1004-9541(09)60188-9
Kim HJ, Choi HS, Yang SY, Kim IS, Yamaguchi T, Sohng JK, Park SK, Kim JC, Lee CH, Gardener BM, Kim YC (2014) Both extracellular chitinase and a new cyclic lipopeptide, chromobactomycin, contribute to the biocontrol activity of Chromobacterium sp. C61. Mol Plant Pathol 15(2):122–32. https://doi.org/10.1111/mpp.12070
Whitaker JO, Dannelly HK, Prentice DA (2004) Chitinase in insectivorous bats. J Mammal 85:15. https://doi.org/10.1644/1545-1542(2004)085%3c0015:CIIB%3e2.0.CO;2
Spizizen J (1958) Transformation of biochemically deficient strains of Bacillus subtilis by deoxyribonucleate. Proc Natl Acad Sci USA 44:1072–1078. https://doi.org/10.1073/pnas.44.10.1072
Hsu S, Lockwood J (1975) Powdered chitin agar as a selective medium for enumeration of actinomycetes in water and soil. Appl Microbiol 29:422–426
Yanai K, Takaya N, Kojima N, Horiuchi H, Ohta A, Takagi M (1992) Purification of two chitinases from Rhizopus oligo- sporus and isolation and sequencing of the encoding genes. J Bacteriol 174(22):7398–7406. https://doi.org/10.1128/jb.174.22.7398-7406.1992
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254. https://doi.org/10.1016/0003-2697(76)90527-3
Martínez MA, Delgado OD, Breccia JD, Baigorí MD, Siñeriz F (2002) Revision of the taxonomic position of the xylanolytic Bacillus sp MIR32 reidentified as Bacillus halodurans and plasmid-mediated transformation of B. halodurans species. Extremophiles 6(5):391–395. https://doi.org/10.1007/s00792-002-0269-4
Irulan A, Nathan YP, Priya G, Marimuthu G, Elangovan V (2011) Isolation and characterization of chitinase producing gut microflora of insectivorous bats. Trends Biosci 4:8–11
Shang-Hsin G, Jeen-Kuan C, Wen-Chien L (2004) Purification and characterization of extracellular chitinase from Aeromonas schubertii Taiwan. Enzyme Microb Technol 35:550–556. https://doi.org/10.1016/j.enzmictec.2004.08.025
Sastoque- Cala L, Mercado-Reyes M, Martínez-Salgado M, Quevedo-Hidalgo B, Pedroza-Rodríguez A (2007) Producción de quitinasas extracelulares con una cepa alcalófila halotolerante de Streptomyces sp. aislada de residuos de camarón. Rev Mex Ing Qca 6:137–146
Veliz E, Martínez-Hidalgo P, Hirsch AM (2017) Chitinase-producing bacteria and their role in biocontrol. AIMS Microbiol 3(3):689–705. https://doi.org/10.3934/microbiol.2017.3.689
Jeyaram K, Wahengbam R, Thangjam AS, Gbenga AA, Khundrakpam B, Folarin AO (2011) Distinct differentiation of closely related species of group with industrial importance. J Microbiol Methods 87(2):161–164. https://doi.org/10.1016/j.mimet.2011.08.011
Nakamura LK (1989) Taxonomic relationship of black pigmented Bacillus subtilis strains and a proposal for Bacillus atrophaeus sp. Int J Syst Bacteriol 39:295–300. https://doi.org/10.1099/00207713-39-3-295
Mabuchi N, Araki Y (2001) Cloning and sequencing of two genes encoding chitinases A and B from Bacillus cereus CH. Can J Microbiol 47(10):895–902. https://doi.org/10.1139/w01-093
Sato Y, Araki Y (2007) Analysis of ChiA and ChiB production by Bacillus cereus CH: induction, gene expression, and localization of two chitinase. J Environ Biotechnol 7:27–32
Sato Y, Araki Y (2008) Identification of Inducers for chitinase B (ChiB) production in Bacillus cereus CH and estimation of its induction mechanism. J Environ Biotechnol 8:119–121
Mayorga-Reyes L, Calderón-Garza E, Gutiérrez-Nava A, González-Cervantes R, Azaola-Espinosa A, Barranco-Florido E (2012) Characterization and expression of the chitinase chit II gene from Lecanicillium lecanii in solid-state fermetnation. Rev Mex Ing Quím 11:97–104. http://www.scielo.org.mx/scielo.php?script=sci_arttext&pid=S1665-27382012000100008&lng=es&tlng=. Accessed 2 Jun 2021
Gomaa EZ (2012) Chitinase Production by Bacillus thuringiensis and Bacillus licheniformis: their potential in antifungal biocontrol. J. Microbiol 50(1):103–111. https://doi.org/10.1007/s12275-012-1343-y
Driss F, Kallassy-Awad M, Zouari N, Jaoua S (2005) Molecular characterization of a novel chitinase from Bacillus thuringiensis subsp. kurstaki. J Appl Microbiol 99(4):945–953. https://doi.org/10.1111/j.1365-2672.2005.02639.x
Huang CJ, Wang TK, Chung SC, Chen CY (2005) Identification of an antifungal chitinase from a potential biocontrol agent, Bacillus cereus 28–9. J Biochem Mol Biol 38(1):82–8. https://doi.org/10.5483/BMBRep.2005.38.1.082
De la Vega LM, Barboza-Corona JE, Aguilar-Uscanga MG, Ramirez-Lepe M (2006) Purification and characterization of an exochitinase from Bacillus thuringiensis subsp. Aizawai and its action against phytopathogenic fungi. Can J Microbiol 52(7):651–657. https://doi.org/10.1139/w06-019
Gurav R, Tang J, Jadhav J (2017) Novel chitinase producer Bacillus pumilus RST25 isolated from the shellfish processing industry revealed antifungal potential against phyto-pathogens. Int Biodeterior Biodegrad 125:228–234. https://doi.org/10.1016/j.ibiod.2017.09.015
Rishad KS, Rebello S, Shabanamol PS, Jisha MS (2017) Biocontrol potential of, halotolerant bacterial chitinase from high yielding novel Bacillus pumilus MCB-7 autochthonous to mangrove ecosystem. Pest Biochem Physiol 137:36–41. https://doi.org/10.1016/j.pestbp.2016.09.005
Dukare AS, Paul S, Asha A (2020) Isolation and efficacy of native chitinolytic rhizobacteria for biocontrol activities against Fusarium wilt and plant growth promotion in pigeon pea (Cajanus cajan L.). Egypt J Biol Pest Control 30(1):56. https://doi.org/10.1186/s41938-020-00256-7
Yuli PE, Suhartono MT, Rukayadi Y, Hwang JK, Pyun YR (2004) Characteristics of thermostable chitinase enzymes from the Indonesian Bacillus sp. 13.26. Enzyme Microb Technol 35:147–215. https://doi.org/10.1016/j.enzmictec.2004.03.017
Rosas-García NM, Fortuna-González JM, Barboza-Corona JE (2013) Characterization of the chitinase gene in Bacillus thuringiensis Mexican isolates. Folia Microbiol 58(6):483–90. https://doi.org/10.1007/s12223-013-0233-y
Kuzu SB, Güvenmez HK, Denizci AA (2012) Production of a thermostable and alkaline chitinase by Bacillus thuringiensis subsp. kurstaki strain HBK-51. Biotechnol Res Int. https://doi.org/10.1155/2012/135498
Fira D, Dimkić I, Berić T, Lozo J, Stanković S (2018) Biological control of plant pathogens by Bacillus species. J Biotechnol 285:44–55. https://doi.org/10.1016/j.jbiotec.2018.07.044
Torres MJ, Pérez Brandan C, Sabaté DC, Petroselli G, Erra-Balsells R, Audisio MC (2017) Biological activity of the lipopeptide-producing Bacillus amyloliquefaciens PGPBacCA1 on common bean Phaseolus vulgaris L. pathogens. Biol Control 105:93–99. https://doi.org/10.1016/j.biocontrol.2016.12.001
Toral L, Rodríguez M, Béjar V, Sampedro I (2018) Antifungal activity of lipopeptides from Bacillus XT1 CECT 8661 against Botrytis cinerea. Front Microbiol 9:1315. https://doi.org/10.3389/fmicb.2018.01315
Zhang QX, Zhang Y, He LL, Ji ZL, Tong YH (2018) Identification of a small antimycotic peptide produced by Bacillus amyloliquefaciens 6256. Pestic Biochem Physiol 150:78–82. https://doi.org/10.1016/j.pestbp.2018.07.006
Li YG, Wang RT, Liu JX, Xu LK, Ji PS, Sun L, Pan HY, Jiang BW, Li LR (2019) Identification of a biocontrol agent Bacillus vallismortis BV23 and assessment of effects of its metabolites on Fusarium graminearum causing corn stalk rot. Biocontrol Sci Techn 29:263–275. https://doi.org/10.1080/09583157.2018.1548575
Wang S, Sun L, Zhang W, Chi F, Hao X, Bian J, Li Y (2020) Bacillus velezensis BM21, a potential and efficient biocontrol agent in control of corn stalk rot caused by Fusarium graminearum. Egypt J Biol Pest Control 30(1):9. https://doi.org/10.1186/s41938-020-0209-6
Li Y, Cai Y, Liang Y, Ji P, Xu L (2020) Assessment of antifungal activities of a biocontrol bacterium BA17 for managing postharvest gray mold of green bean caused by Botrytis cinerea. Postharvest Biol Technol. https://doi.org/10.1016/j.postharvbio.2019.111086
Acknowledgements
This work was supported by the financial assistance of CONICET. The authors also thank the Academic style (https://www.academicstyle.com.ar/correcion-estilos) for the English revision of this article.
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design of the work. BM and VL: designed and supervised all the work. DC: carried out the sampling and all the experimental part. MM: identified the microorganisms. DD, MM and VL: drafted and prepared the original draft. The final manuscript has been read, modified and approved by all named authors.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
CD, D., LB, V., MA, M. et al. Extracellular Antifungal Activity of Chitinase-Producing Bacteria Isolated From Guano of Insectivorous Bats. Curr Microbiol 78, 2787–2798 (2021). https://doi.org/10.1007/s00284-021-02555-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00284-021-02555-0