Abstract
This study aimed to compare the genotype diversity of C. jejuni isolates. From the total of 64 C. jejuni strains evaluated, 44 were isolated from broiler carcasses (2015–2016) and 20 from hospitalized patients with gastroenteritis caused by the microorganism (2000–2006). The strains were correlated for the presence of flaA, pldA, cadF, ciaB, cdtABC, luxS, dnaJ, cbrA, htrA, pVir, Hcp, cstII, and neuA genes by PCR (polymerase chain reaction) and for phylogenetic proximity by PFGE (pulsed-field gel electrophoresis). Of the total strains studied, 28 (43.7%) presented all the studied genes, except pVir. Among these strains, 25 (89.3%) were of poultry origin. Poultry strains showed a higher prevalence (P < 0.05) of genes linked to adhesion, colonization, invasion, cytotoxicity, biofilm formation, and adaptation to adverse conditions. Additionally, the profile that denotes the presence of all genes identified in the study (P1) was identified in 56.8% of poultry strains and in 15.0% of human strains. Molecular typing analysis identified five pulsotypes, none of which grouped strains from different origins. Although human strains were from hospitalized patients, they presented limited virulence capacity and adaptability to adverse conditions compared to chicken carcasses, besides being different in molecular typing. However, the ability to cause Guillain-Barré Syndrome is equal for both strains. In general, poultry strains, being more recent, are more specialized to adapt to the environment, invade, and cause disease in the human host.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Brazilian poultry is one of the most profitable agribusiness sectors for the country. Brazil’s prominent position as the third largest producer and largest exporter of chicken meat in the world promotes constant concerns about quality standards and ensuring consumer food safety [1].
Microbiological quality is one of the most important pillars for domestic market and export. Thus, the presence of zoonotic microorganisms must be constantly monitored throughout the poultry production process. Among these microorganisms, species of the genus Campylobacter have received special attention due to the growing number of cases of gastroenteritis in the world.
Data from epidemiological surveillance agencies such as the European Food Safety Authority (EFSA) in the European Union (EU), and Centers for Disease Control and Prevention (CDC) in the USA indicate that Campylobacter affects 1.3 million people per year in the USA, and nine million in the EU, with high costs associated with lost productivity and public health care [2, 3]. In the scientific literature, we find studies performed in Brazil that has been evaluating for over two decades the potential risks of campylobacteriosis in farm animals, mainly related to chicken meat [4], as well as in human clinical cases, due to the risk of hospitalization for Guillain-Barré Syndrome [5]. But, despite Brazil’s prominent position in poultry production [1], official cases of campylobacteriosis are underreported and molecular studies of C. jejuni diversity and virulence are still scarce in the country [6, 7], probably due to the absence of cheap and easy to implement isolation and identification methodologies, as well as a lack of legislation that mandates analysis and the maximum tolerated numbers for this microorganism in animal foods, mainly in chicken meat.
Campylobacteriosis is characterized by presenting from mild, self-limiting, watery diarrhea to bloody dysentery with mucus and white blood cells, and may be accompanied by headaches and abdominal pain, fever, malaise, nausea, and vomiting, symptomatology similar to that caused by several other enteric pathogens [8]. A low infecting dose of Campylobacter, about 400 to 500 cells, represents a higher risk of infection [9, 10]. Most infected people recover within 2 to 5 days. However, in some cases, infection can cause more serious illnesses such as sepsis, abortion, meningitis, abscesses, and complications such as Guillain-Barré Syndrome (GBS), characterized by flaccid paralysis that can cause death from respiratory failure [8].
Due to the large number of reported cases of campylobacteriosis in Europe and the USA and the official underreporting of cases in Brazil, it has become necessary to use molecular methods of genetic characterization and epidemiological typing that allow discrimination of bacterial strains and knowledge of virulence and adaptation potential. Data obtained from these tests can be used by public health surveillance to identify the causes of food outbreaks and to understand the risks [6, 11].
Among these methods, pulsed-field gel electrophoresis (PFGE) is considered the gold standard in bacterial epidemiological analyses. PFGE allows investigation of genomic variability of all genetic material among isolates of bacteria of the same species. Presence of insertions, deletions, or mutations can be detected between the genomes of bacterial isolates, allowing a high discriminatory power compared to other techniques [12].
Given the national importance of the poultry market, the aim was to comparatively analyze C. jejuni strains isolated from carcasses of chickens destined for internal and external consumption and isolated from human clinical patients, regarding genetic proximity, the dissemination of different genetic profiles, and the pathogenic profile through the presence of virulence factors, adaptation factors, and GBS-related genes.
Materials and Methods
Bacterial Strains
We analyzed 64 strains of C. jejuni, 19 of which were received from the Fiocruz-RJ (CCAMP; source: human feces), one from Adolfo Lutz Institute of Ribeirão Preto (source: human feces), and 44 from Federal University of Uberlandia (CLEM; source: chicken carcass). The data referring to the origin of the strains are described in Table 1. Strains of human origin correspond to all the quantitative present in the culture bank of Fiocruz, which, due to the lack of active surveillance in Brazil, presents restrictions regarding the periodicity of isolations.
All strains were previously isolated and identified following the ISO isolation protocols [13] by Fiocruz and the Adolfo Lutz Institute for human strains and by Melo [14] for poultry strains. We cultured, confirmed, and restored with cryoprotectant in an ultrafreezer at −80 °C. The C. jejuni NCTC 11351 strain was used in all tests.
Species Confirmation
All strains were reactivated in Bolton selective enrichment broth (Oxoid®) with 5% defibrinated sheep blood (Laborclin®) under microaerophilic conditions at 37 °C for 44 h ± 4 h. Samples were then seeded on Campylobacter Blood-Free Selective Medium Agar (Modified CCDA-Preston) (Oxoid®). Bacterial colonies were used to identify C. jejuni biochemically (hippurate hydrolysis test) and by PCR multiplex.
DNA was obtained from the Wizard® Genomic DNA Purification Kit (Promega®), and PCR was performed with the GoTaq Green Master Mix Kit (Promega®) associated with C1 (5′CAAATAAAGTTAGAGGTAGAATGT3′)–C4 (5′GGATAAGCACTAGCTAGCTGAT3′) primers and pg3 (5′GAACTTGAACCGATTTG3′)–pg50 (5′ATGGGATTTCGTATTAAC3′) (Invitrogen®) [15].
Specific Gene Detection
A total of 13 adaptive virulence and resistance genes (flaA—motility, Hcp and pldA—extracellular colonization, ciaB and pVir—invasion, cadF—intracellular colonization, cdtABC—toxin production, luxS—quorum-sensing mechanism, dnaJ—thermotolerance, htrA—growth under stress, cbrA—resistance to osmotic shock, cstII and neuA—Guillain-Barré Syndrome (GBS) were evaluated in C. jejuni strains by PCR. All reactions were conducted with the GoTaq Green Master Mix Kit (Promega®). Genes were identified by primers and specific amplification conditions (Table 2).
PFGE
PFGE was conducted according to the CDC PulseNet standard [16] with CHEF Mapper III equipment (Bio-Rad). Genomic DNA was digested with restriction enzyme SmaI (Invitrogen®). The 1 Kb molecular weight marker (Promega®) was used to compare the formed bands.
Strains were packaged in solution containing SKG (SeaKem Gold) agarose and proteinase K (20 mg/mL). The agarose blocks were transferred to a cell lysis buffer at 54 °C for 15 min under orbital shaking. Four washes in ultrapure water and TE (Tris–EDTA) buffer were performed under the same conditions. The plugs were then digested with 40 U SmaI at room temperature for 2 h.
The DNA fragments were separated on 1% agarose gel in a 0.5X TBE (Tris–borate-EDTA) buffer for 18 h, with the following parameters: 200v, 120° angle, 6v/cm gradient, and buffer temperature of 14 °C.
Gels were stained with ethidium bromide and photographed under UV light. Analysis for dendrogram formation was performed using Gel Compare II software. The band patterns were compared using the DICE similarity coefficient in the UPGMA analysis method.
Statistical Analysis
A binomial test was used to compare proportions of studied genes present between the strains with 5% significance using GraphPad Prism 8.0.1 software.
Results and Discussion
The presence of genes associated with virulence and adaptation is described in Table 3. The studied genes can be divided into the pathogenicity categories, which includes the cadF, pldA, ciaB, flaA, and pVir genes; cytotoxicity, Hcp and cdtABC genes; formation of biofilms and adaptation to adverse conditions, luxS, htrA, cbrA, and dnaJ genes; and Guillain-Barré syndrome, cstII and neuA genes.
In general, our study demonstrated that the strains of poultry origin presented higher pathogenic and adaptive potential (P < 0.05), except for flaA, cstII, neuA, and Hcp genes, which showed no significant difference from human strains. None of the strains showed positivity for the pVir plasmidial gene that is related to factors that favor invasive Campylobacter infection [17]
Distinct results were found in a study by Oh et al. with strains from human and chicken feces isolated from 2007 to 2010 [18]. The authors did not identify significant differences in detection of flaA, cadF, racR, dnaJ, cdtA, cdtB, and cdtC genes for different strains. Rodrigues et al. also did not identify differences in C. jejuni virulence from children and dogs in 2010 and 2011 [19].
It is likely that the large difference related to the year of isolation of the strains influenced the results, since human strains were from 2000 to 2006 and poultry from 2015 and 2016. Some studies have already evaluated aspects of the evolutionary history of C. jejuni, and demonstrated that this microorganism uses mechanisms of mutation and gene recombination to create a more virulent population and adapt to different environments. These changes lead to the emergence of new strains that endanger the exposed population, render prevention techniques in industry and human medicine ineffective, and show the need for constant improvement of agent control forms [20, 21].
Absence of the pVir gene in all strains is consistent with other studies that also reported its absence or low prevalence in Campylobacter strains of different origins [17, 22,23,24]; this may also be due to the pVir gene being a plasmid, which may be lost during strain subcultures as well as during the DNA extraction process.
Despite being considered a virulence gene, the connection of pVir with the presence of bloody diarrhea could not be confirmed, since only 29% of C. jejuni obtained from bloody diarrhea samples contained this plasmid [17]. Additionally, studies by Marasini et al. [25] and Iglesias-Torrens et al. [26] considered that pVir, previously associated with virulence, is not necessary for C. jejuni to colonize birds or infect humans. Thus, the absence of pVir in the investigated strains does not infer variation in its virulence potential.
Prevalent presence of Hcp gene in both strains indicates the potent ability of these strains to express the possibility of colonizing and secreting substances that guarantee their survival and their ability to attack the host. These pathogenicity characteristics directly interfere with the clinical course of the disease, as the symptoms and outcome of infection depend on a number of factors that include host immunity, initiation of therapy, environmental factors, and, in particular, factors associated with the pathogenicity of the strain [27].
The presence of the flaA gene in both strains did not vary, showing the importance of motility as a relatively conserved trait in this species. The cadF, ciaB, and pldA virulence genes were found most frequently in carcass strains, confirming higher invasive potential, host adhesion, and colonization in establishment of the disease. The small number of human strains that presented cdtABC (3/20–15%) and luxS (9/20–45%) genes evidences the limited toxicity of these strains in causing invasive apoptosis-related host cells as well as a restricted ability to form biofilms in the gut and outdoors. At the same time, the fragility of human strains under adverse conditions of temperature, nutritional, and osmotic stress was higher than poultry strains [28,29,30,31,32].
The presence of GBS-linked genes did not differ between strains (P > 0.05). Identification of both genes (cstII and neuA) was detected in 47 (73.4%) strains, 31 (66.0%) from carcasses, and 16 (34.0%) from humans, and 53 (82.8%) had at least one of these genes. Similar results were found by Hardy et al. [33] and Amon et al. [34], who found no differences regarding the presence of these genes in human and bird strains.
Several studies have indicated that the terminal regions of the C. jejuni lipo-oligosaccharide are responsible for the production of autoimmune antibodies that attack human gangliosides responsible for GBS [35, 36]. Among these regions, the sialyltransferase enzyme encoded by the cstII gene and the sialic acid activation enzyme translated by the neuA gene showed a direct relationship with C. jejuni-associated GBS [34, 37].
A high number of strains with potential to cause neuropathy after the campylobacteriosis event in both human and poultry strains show the risk of GBS development in humans. However, statistical results suggest that isolates responsible for causing GBS in humans are not selected for environmental or host-specific factors and that the occurrence of autoimmune disease is likely to be mainly dependent on patient factors such as humoral and cellular immunity [34].
All 16 virulence profiles identified in Table 4 reinforce the higher pathogenicity and adaptive resistance of broiler strains, since the P1 profile (presence of all genes) was the most identified. Thus, even with strains isolated from humans with clinical symptoms, the greatest pathogenic potential of poultry strains is undeniable and denotes the danger posed by consumption of raw or undercooked chicken for the development of a severe and acute form of campylobacteriosis in the human host. In addition, they demonstrate the importance of practices that avoid cross contamination during preparation of these foods in homes and restaurants.
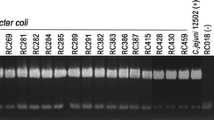
Five pulsotypes (genotypes) with more than 80% similarity were identified in the phylogenetic study between strains, four from chicken carcasses (A, B, C, and E) and one composed of two human strains (D) (Fig. 1). Each pulsotype grouped only two strains, showing the high genetic heterogeneity in C. jejuni.
All clusters presented strains isolated in the same year and with similar genetic characteristics. The pulsotypes B, C, and E showed the presence of all studied genes in common. In pulsotype A, one strain did not show flaA, and in D, one strain did not have cdtABC. This variation is consistent with a degree of homology of less than 100%, which allows for minor changes in the genome.
Absence of clusters with strains of humans and chickens makes clear the genetic distance between them, once again proving divergences related to the virulence and adaptation characteristics identified in these strains and the probable evolution that populations of C. jejuni suffered over time.
The similarity in strains of different origins was identified by Frazão et al. [20] in Brazil and by Oh et al. [18] in Korea. However, this homology was only detected in strains isolated in the same year or with up to 1 year of difference between them, which justifies the difference found in our study. Rapid molecular adaptation by genetic recombination in C. jejuni allows formation of quite phylogenetically distinct populations, preventing strains from grouping and allowing their constant evolution and specialization over time [21].
Conclusion
Our study proved that most of the studied genes (cadF, pldA, ciaB, cdtABC, luxS, htrA, cbrA, and dnaJ) were more prevalent in the strains of C. jejuni of poultry origin. The risk of developing GBS did not differ according to the origin and the absence of the pVir gene does not appear to interfere with the pathogenic potential. The phylogenetic heterogeneity between strains of human and poultry origin is consistent with the differences identified in the virulence profiles and with the temporal variation of isolation that shows that more recent strains (of poultry origin) are more specialized at the molecular level.
References
Associação Brasileira de Proteina Animal (2020) Relatórios anuais: Relatório anual 2020. Publishing ABPA, São Paulo, SP
Centers for Disease Control and Prevention (2020) National center for emerging and zoonotic infectious diseases. Campylobacteriosis. U.S. Department of Health and Human Services, Washington, DC
European Food Safety Authority (EFSA) (2020) The European Union Summary Report on Antimicrobial Resistance in zoonotic and indicator bacteria from humans, animals and food in 2017/2018. EFSA, Parma
Lopes GV, Landgraf M, Destro MT (2018) Occurrence of Campylobacter in raw chicken and beef from retail outlets in São Paulo. Brazil J Food Saf 38:e12442. https://doi.org/10.1111/jfs.12442
Souza CO, Vieira MACS, Batista FMA, Eulálio KD et al (2018) Serological markers of recent Campylobacter jejuni infection in patients with Guillain-Barré Syndrome in the State of Piauí, Brazil, 2014–2016. Am J Trop Med Hyg 98:586–588. https://doi.org/10.4269/ajtmh.17-0666
BRASIL. Ministério da Saúde (2019) Surtos de doenças transmitidas por alimentos no Brasil, 2019. Available at https://antigo.saude.gov.br/images/pdf/2019/maio/17/Apresentacao-Surtos-DTA-Maio-2019.pdf. Acessed on 28 Jan 2021
Melo RT, Grazziotin AL, Valadares Junior EC et al (2019) Evolution of Campylobacter jejuni of poultry origin in Brazil. Food Microbiol 82:489–496. https://doi.org/10.1016/j.fm.2019.03.009
World Health Organization (2013) The global view of campylobacteriosis. WHO, Geneva
Al Amri A, Senok AC, Ismaeel AY et al (2007) Multiplex PCR for direct identification of Campylobacter spp. in human and chicken stools. J Med Microbiol. https://doi.org/10.1099/jmm.0.47220-0
Ivanovic S (2012) Campylobacter as a cause of gastroenteritis in humans and animals. Afr J Microbiol Res 6:1651–1657
Nakari UM, Hakkinen M, Siitonen A (2011) Identification of persistent subtypes of Campylobacter jejuni by pulsed-field gel electrophoresis in Finland. Foodborne Pathog Dis. https://doi.org/10.1089/fpd.2011.0882
Goering RV (2010) Pulsed field gel electrophoresis: a review of application and interpretation in the molecular epidemiology of infectious disease. Infect Genet Evol. https://doi.org/10.1016/j.meegid.2010.07.023
International Standards Organization (2006) ISO 10272-1: microbiology of food and animal feeding stuffs—horizontal method for detection and enumeration of Campylobacter spp. Part 1: detection method ISO 10272-1. ISO, Geneva
Melo RT (2017) Emergência de Campylobacter jejuni no setor avícola e na saúde pública do Brasil. Master dissertation, Federal University of Uberlandia, Uberlandia
Harmon KM, Ransom GM, Wesley IV (1997) Differentiation of Campylobacter jejuni and Campylobacter coli by polymerase chain reaction. Mol Cell Probes. https://doi.org/10.1006/mcpr.1997.0104
Centers for Disease Control and prevention (CDC) (2013) Standardized laboratory protocol for molecular subtyping of Campylobacter jejuni by pulsed field gel electrophoresis (PFGE): PulseNet USA, the national molecular subtyping network for foodborne disease surveillance. CDC, Atlanta
Bacon DJ, Alm R, Burr DH et al (2000) Involvement of a plasmid in virulence of Campylobacter jejuni. Infect Immun 68:4384–4390. https://doi.org/10.1128/IAI.68.8.4384-4390.2000
Oh JY, Kwon YK, Wei B et al (2017) Epidemiological relationships of Campylobacter jejuni strains isolated from humans and chickens in South Korea. J Microbiol. https://doi.org/10.1007/s12275-017-6308-8
Rodrigues CG, Melo RT, Fonseca BB et al (2015) Occurrence and characterization of Campylobacter spp. isolates in dogs, cats and children. Pesqui Vet Bras. https://doi.org/10.1590/S0100-736X2015000400009
Frazão MR, Medeiros MIC, Da Silva DS, Falcão JP (2017) Pathogenic potential and genotypic diversity of Campylobacter jejuni: a neglected food-borne pathogen in Brazil. J Med Microbiol. https://doi.org/10.1099/jmm.0.000424
Sheppard SK, Maiden MCJ (2015) The evolution of Campylobacter jejuni and Campylobacter coli. Cold Spring Harb Perspect Biol. https://doi.org/10.1101/cshperspect.a018119
de Nascimento Veras H, Medeiros PHQS, Ribeiro SA et al (2018) Campylobacter jejuni virulence genes and immune-inflammatory biomarkers association with growth impairment in children from Northeastern Brazil. Eur J Clin Microbiol Infect Dis. https://doi.org/10.1007/s10096-018-3337-0
Tracz DM, Keelan M, Ahmed-Bentley J et al (2005) pVir and bloody diarrhea in Campylobacter jejuni enteritis. Emerg Infect Dis. https://doi.org/10.3201/eid1106.041052
Schmidt-Ott R, Pohl S, Burghard S et al (2005) Identification and characterization of a major subgroup of conjugative Campylobacter jejuni plasmids. J Infect. https://doi.org/10.1016/j.jinf.2004.02.013
Marasini D, Karki AB, Buchheim MA, Fakhr MK (2018) Phylogenetic relatedness among plasmids harbored by Campylobacter jejuni and Campylobacter coli isolated from retail meats. Front Microbiol 9:1–14. https://doi.org/10.3389/fmicb.2018.02167
Iglesias-Torrens Y, Miró E, Guirado P, Llovet T, Muñoz C, Cerdà-Cuéllar M, Madrid C, Balsalobre C, Navarro F (2018) Population structure, antimicrobial resistance, and virulence-associated genes in Campylobacter jejuni isolated from three ecological niches: gastroenteritis patients, broilers, and wild birds. Front Microbiol 9:1676. https://doi.org/10.3389/fmicb.2018.01676
Agnetti J, Seth-Smith HMB, Ursich S et al (2019) Clinical impact of the type VI secretion system on virulence of Campylobacter species during infection. BMC Infect Dis. https://doi.org/10.1186/s12879-019-3858-x
Eucker TP, Konkel ME (2012) The cooperative action of bacterial fibronectin-binding proteins and secreted proteins promote maximal Campylobacter jejuni invasion of host cells by stimulating membrane ruffling. Cell Microbiol 14:226
Poly F, Ewing C, Goon S et al (2007) Heterogeneity of a Campylobacter jejuni protein that is secreted through the flagellar filament. Infect Immun. https://doi.org/10.1128/IAI.00159-07
Feodoroff B, Ellström P, Hyytiäinen H et al (2010) Campylobacter jejuni isolates in Finnish patients differ according to the origin of infection. Gut Pathog. https://doi.org/10.1186/1757-4749-2-22
Dasti JI, Tareen AM, Lugert R et al (2010) Campylobacter jejuni: a brief overview on pathogenicity-associated factors and disease-mediating mechanisms. Int J Med Microbiol. https://doi.org/10.1016/j.ijmm.2009.07.002
Cróinín TÓ, Backert S, (2012) Host epithelial cell invasion by Campylobacter jejuni: trigger or zipper mechanism? Front Cell Infect Microbiol. https://doi.org/10.3389/fcimb.2012.00025
Hardy CG, Lackey LG, Cannon J et al (2011) Prevalence of potentially neuropathic Campylobacter jejuni strains on commercial broiler chicken products. Int J Food Microbiol. https://doi.org/10.1016/j.ijfoodmicro.2010.12.027
Amon P, Klein D, Springer B et al (2012) Analysis of Campylobacter jejuni isolates of various sources for loci associated with Guillain-Barré syndrome. Eur J Microbiol Immunol. https://doi.org/10.1556/eujmi.2.2012.1.4
Ang CW, Laman JD, Willison HJ et al (2002) Structure of Campylobacter jejuni lipopolysaccharides determines antiganglioside specificity and clinical features of Guillain-Barré and Miller Fisher patients. Infect Immun 70:1202–1208. https://doi.org/10.1128/IAI.70.3.1202-1208.2002
Prendergast MM, Moran AP (2000) Lipopolysaccharides in the development of the Guillain-Barré syndrome and Miller Fisher syndrome forms of acute inflammatory peripheral neuropathies. J Endotoxin Res 6:341–359
Van Belkum A, Van Den Braak N, Godschalk P et al (2001) A Campylobacter jejuni gene associated with immune-mediated neuropathy. Nat Med. https://doi.org/10.1038/89831
Acknowledgements
To Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq) and Fundação de Amparo à Pesquisa do Estado de Minas Gerais (FAPEMIG) for financial support for the execution of the study.
Funding
This research was funded by Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq) and Fundação de Amparo à Pesquisa do Estado de Minas Gerais (FAPEMIG).
Author information
Authors and Affiliations
Contributions
Conceptualization, R.T.M and R.F.B.; methodology, R.T.M., C.F.D., R.F.B. and E.C.A.L; software, G.P.M. and J.P.S; validation, R.T.M. and D.A.R.; formal analysis, R.T.M. and J.P.S.; investigation, G.P.M.; resources, R.T.M. and D.A.R.; data curation, E.C.A.L.; writing-original draft preparation, R.T.M., M.G.T., C.F.D., and R.F.B.; writing-review and editing, R.T.M., M.G.T., J.P.S. and D.A.R.; visualization, G.P.M., M.G.T. and E.C.A.L.; supervision, R.T.M and D.A.R.; project administration, R.T.M. and D.A.R.; funding acquisition, R.T.M. and D.A.R. All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
de Melo, R.T., Dumont, C.F., Braz, R.F. et al. Genotypical Relationship Between Human and Poultry Strains of Campylobacter jejuni. Curr Microbiol 78, 2980–2988 (2021). https://doi.org/10.1007/s00284-021-02553-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00284-021-02553-2